CATALYTIC MECHANISM AND EVOLUTION OF AN ENZYME RESPONSIBLE FOR NYLON-6 BYPRODUCT DEGRADATION
|
|
|
- Frederick Boyd
- 6 years ago
- Views:
Transcription
1 THE JOURNAL OF BIOLOGICAL CHEMISTRY VOL. 285, NO. 2, pp , January 8, by The American Society for Biochemistry and Molecular Biology, Inc. Printed in the U.S.A. X-ray Crystallographic Analysis of the 6-Aminohexanoate Cyclic Dimer Hydrolase CATALYTIC MECHANISM AND EVOLUTION OF AN ENZYME RESPONSIBLE FOR NYLON-6 BYPRODUCT DEGRADATION * S Received for publication, July 13, 2009, and in revised form, October 14, 2009 Published, JBC Papers in Press, November 3, 2009, DOI /jbc.M Kengo Yasuhira 1, Naoki Shibata 1, Go Mongami, Yuki Uedo, Yu Atsumi, Yasuyuki Kawashima, Atsushi Hibino, Yusuke Tanaka, Young-Ho Lee, Dai-ichiro Kato, Masahiro Takeo, Yoshiki Higuchi 2, and Seiji Negoro 3 From the Department of Materials Science and Chemistry, Graduate School of Engineering, University of Hyogo, Hyogo , the Department of Life Science, Graduate School of Life Science, University of Hyogo, Hyogo , the RIKEN Harima Institute, SPring-8 Center, Hyogo , and the Institute for Protein Research, Osaka University, Osaka , Japan We performed x-ray crystallographic analyses of the 6-aminohexanoate cyclic dimer (Acd) hydrolase (NylA) from Arthrobacter sp., an enzyme responsible for the degradation of the nylon-6 industry byproduct. The fold adopted by the 472- amino acid polypeptide generated a compact mixed / fold, typically found in the amidase signature superfamily; this fold was especially similar to the fold of glutamyl-trna Gln amidotransferase subunit A (z score, 49.4) and malonamidase E2 (z score, 44.8). Irrespective of the high degree of structural similarity to the typical amidase signature superfamily enzymes, the specific activity of NylA for glutamine, malonamide, and indoleacetamide was found to be lower than 0.5% of that for Acd. However, NylA possessed carboxylesterase activity nearly equivalent to the Acd hydrolytic activity. Structural analysis of the inactive complex between the activity-deficient S174A mutant of NylA and Acd, performed at 1.8 Å resolution, suggested the following enzyme/substrate interactions: a Ser 174 -cis-ser 150 -Lys 72 triad constitutes the catalytic center; the backbone N in Ala 171 and Ala 172 are involved in oxyanion stabilization; Cys 316 -S forms a hydrogen bond with nitrogen (Acd-N 7 ) at the uncleaved amide bond in two equivalent amide bonds of Acd. A single S174A, S150A, or K72A substitution in NylA by site-directed mutagenesis decreased the Acd hydrolytic and esterolytic activities to undetectable levels, indicating that Ser 174 -cis-ser 150 -Lys 72 is essential for catalysis. In * This work was supported in part by a grant-in-aid for scientific research from the Japan Society for Promotion of Science and by grants from the Global Centers of Excellence Program, the National Project on Protein Structural and Functional Analyses, the Core Research for Evolutional Science and Technology Program, Development of the Foundation for Nano-Interface Technology from JST, and the Japan Aerospace Exploration Agency project. S The on-line version of this article (available at contains supplemental Table S1 and Figs. S1 S4. The atomic coordinates and structure factors (codes 3A2P and 3A2Q) have been deposited in the Protein Data Bank, Research Collaboratory for Structural Bioinformatics, Rutgers University, New Brunswick, NJ ( 1 These authors contributed equally to this work. 2 To whom correspondence may be addressed: Dept. of Life Science, Graduate School of Life Science, University of Hyogo, Koto, Kamigori-cho, Ako-gun, Hyogo , Japan. Tel. and Fax: ; hig@sci.u-hyogo.ac.jp. 3 To whom correspondence may be addressed: Dept. of Materials Science and Chemistry, Graduate School of Engineering, University of Hyogo, 2167 Shosha, Himeji, Hyogo , Japan. Tel. and Fax: ; negoro@eng.u-hyogo.ac.jp. contrast, substitutions at position 316 specifically affected Acd hydrolytic activity, suggesting that Cys 316 is responsible for Acd binding. On the basis of the structure and functional analysis, we discussed the catalytic mechanisms and evolution of NylA in comparison with other Ser-reactive hydrolases. Nylon-6 is produced by ring cleavage polymerization of -caprolactam and consists of more than 100 units of 6-aminohexanoate. During the polymerization reaction, however, some molecules fail to polymerize and remain oligomers, whereas others undergo head-to-tail condensation to form cyclic oligomers (1, 2). These byproducts (designated nylon oligomers) contribute to an increase in industrial waste material into the environment. Therefore, biodegradation of the xenobiotic compounds is important in terms of environmental points of view. In addition, the biological system for degradation provides us with a suitable system to study how microorganisms have evolved specific enzymes responsible for this degradation (1, 2). Previous biochemical studies in a nylon oligomer-degrading bacterium Arthrobacter sp. strain KI72 have revealed that three enzymes, 6-aminohexanoate cyclic dimer hydrolase (NylA), 4 6-aminohexanoate dimer hydrolase (NylB), and endo-type 6-aminohexanoate oligomer hydrolase (NylC), are responsible for the degradation of nylon oligomers (1, 2). NylA specifically hydrolyzes one of the two equivalent amide bonds in Acd, generating a 6-aminohexanoate linear dimer (Ald) (see Fig. 1A). However, NylA is barely active on Ald (substrate for NylB), 6-aminohexanoate cyclic oligomers (degree of polymerization 3, substrates for NylC), and more than 60 kinds of various amide compounds, including peptides and -lactams (1 3). NylA is a homodimeric enzyme, and Acd hydrolytic activity is inhibited by diisopropylfluorophosphate, a Ser-enzyme-specific inhibitor, suggesting that Ser is involved in the catalytic function (3). NylA has also been found in Pseudomonas sp. NK87 and six alkalophilic strains, including Kocuria sp. KY2 (4 6). All of the NylA proteins identified so far have been 4 The abbreviations used are: NylA, 6-aminohexanoate cyclic dimer hydrolase; Acd, 6-aminohexanoate cyclic dimer; Ald, 6-aminohexanoate linear dimer; NylA-A 174, NylA containing the S174A substitution; AS, amidase signature; GatA, glutamyl-trna Gln amidotransferase subunit A; MAE2, malonamidase E2; PAM, peptide amidase. JANUARY 8, 2010 VOLUME 285 NUMBER 2 JOURNAL OF BIOLOGICAL CHEMISTRY 1239
2 found to be immunologically identical and possess similar subunit molecular masses of 52 kda (5, 6). NylA from Pseudomonas is 99% identical in amino acid sequence to the homolog from Arthrobacter, and both nyla genes are encoded on a plasmid (4, 7). NylA is classified as an amidase signature (AS) superfamily protein, characterized by the presence of a conserved stretch of 130 amino acid residues (AS sequence) (8 21). AS superfamily enzymes are widely distributed among microorganisms, animals, and plants; they are involved in various biological functions, including transamidation of misacylated Glu-tRNA Gln (8, 9), formation of the plant hormone indoleacetic acid (10, 11), and hydrolysis of malonamide (12, 13), fatty acid amides (14, 15), and peptide amides (16, 17). In addition, amidases from Rhodococcus rhodochrous and Sulfolobus solfataricus possess nitrilase activity, catalyzing the cleavage of the carbon-nitrogen triple bond in a nitrile (18, 19). Recently, NylA from Arthrobacter was also shown to exhibit 98% homology to -laurolactam hydrolase from Rhodococcus (dbj BAG ), Cupriavidus (dbj BAH ), and Sphingomonas (dbj BAH ), which are potentially used for production of 12-aminolauryl acid (material for nyon-12 production) (20). Moreover, NylA is closely related to an aryl acylamidase from Nocardia farcinica, which cleaves the amide bond in a polyamide model compound (adipic acid bishexylamide) and the ester bond in bis(benzoyloxyethyl)terephthalate and increases the hydrophilicity of polyamide 6 (21). Thus, from fundamental, industrial, and environmental points of view, it would be interesting to know the structural factors that determine the unique substrate specificity of NylA recognizing a cyclic amide compound having 6-aminohexanoate as a monomeric unit. Recently, we have reported the crystallization conditions of NylA from strain KI72 (22). In this paper, we performed x-ray crystallographic analysis of NylA, identified the amino acid residues responsible for catalytic function based on the three-dimensional structures, estimated the catalytic mechanisms, and discussed the evolution of nylon oligomer-degrading enzymes. EXPERIMENTAL PROCEDURES Plasmids and Site-directed Mutagenesis The hybrid plasmid puefa, containing a wild-type nyla gene in the vector puc18, was used as the parental plasmid DNA (22). To obtain mutant enzyme versions of NylA, site-directed mutagenesis was carried out by the modification of restriction site method (23) (for K72A, S150A, S174A, C316G, C316D, C316E, and C316S substitutions) or PrimeSTAR mutagenesis kit (Takara Bio Inc., Otsu, Japan) (for N125A, C316A, C316K, and C316N substitutions) using the primers shown in supplemental Table S1 and plasmid puefa as a template DNA. After sequencing the entire nyla region, we confirmed that a single desired substitution was integrated in the wild-type nyla sequence, and recombinant plasmid expressing the mutant nyla genes were constructed using the puc18 vector, as described previously (22). Enzyme Purification Because cultivation at temperatures 30 C or higher results in insoluble NylA, Escherichia coli JM109 cells expressing NylA and mutant enzymes were grown at 25 C, as described previously (22). Buffer A (20 mm phosphate buffer containing 10% glycerol, ph7.0) was used throughout for purification. Cell extracts obtained by ultrasonication were used as crude enzyme. NylA and its mutant enzymes were fractionated by (NH 4 ) 2 SO 4 (25 55% saturated). The enzyme sample dissolved in buffer A was desalted using a PD-10 desalting column (GE Healthcare), and the protein-containing fractions were loaded onto a Hi-Trap Q-Sepharose (GE Healthcare) column equilibrated with buffer A. After the column was washed with buffer A, the enzyme was eluted by increasing the NaCl concentration from 0 to 0.25 M (22). Fractions containing NylA or its mutants were identified as highly expressed 52-kDa bands on SDS- PAGE, which were absent in cell extracts of E. coli JM109 harboring puc18. At the final stage, the enzymes were judged to be homogeneous by analyzing the SDS-PAGE, native PAGE, and light-scattering diffraction patterns. Assay To measure Acd hydrolase activity, enzyme reactions were performed at 30 C using 10 mm Acd (in buffer A) (standard assay condition). Amino groups liberated by the reaction were quantified by colorimetry using trinitrobenzenesulfonic acid (3). Enzyme reactions were similarly performed using 20 mm malonamide (malonamidase activity) or 20 mm indoleacetamide (indoleacetamide hydrolase activity), and release of ammonia formed by the hydrolytic reactions was quantified by colorimetry using trinitrobenzenesulfonic acid (3). For kinetic study, Acd hydrolytic activity was assayed under standard assay conditions, except that various concentrations of Acd were used. Kinetic parameters (k cat and K m values) were evaluated by directly fitting the Michaelis-Menten equation to the data using GraphPad prism, version 5.01 (GraphPad, San Diego, CA). The k cat values were expressed as turnover numbers/subunit (M r of the subunit: 53,000). For semi-quantitative detection of Acd hydrolase and glutaminase activity, the enzyme reactions were performed using purified NylA (0.1 1 mg/ml) and 10 mm Acd or 20 mm Gln. After the reaction proceeded at 30 C for 2 48 h, 10- l aliquots were sampled, and the reactions were stopped by incubating the aliquots in boiling water for 3 min. The reaction mixtures (1 l) were spotted onto a silica gel plate. The samples were developed by solvent mixture (1-propanol:water:ethyl acetate:ammonia 24:12:4:1.3), and degradation products were detected by spraying with 0.2% ninhydrin solution (in butanol saturated with water) (6). To measure carboxylesterase activity, enzyme reactions were performed at 30 C using 0.2 mm each of p-nitrophenylacetate (C2-ester), p-nitrophenylbutyrate (C4-ester), and p-nitrophenyloctanoate (C8-ester) dissolved in 50 mm phosphate buffer containing 0.43% Tween 20 (ph 7.0). The release of p-nitrophenol ( 400 nm 6,710 M 1 cm 1 ) was assayed by absorbance at 400 nm. As a control, we confirmed that spontaneous hydrolysis of p-nitrophenylesters in buffer A is not observed during the 30 min of the reaction. Carboxylesterase activities were assayed in triplicate, and the average values and standard errors were calculated. Crystallographic Analysis Crystallization and Data Collection of Diffraction Hexagonal prismatic crystals ( mm) were obtained by the 1240 JOURNAL OF BIOLOGICAL CHEMISTRY VOLUME 285 NUMBER 2 JANUARY 8, 2010
3 sitting drop vapor diffusion method as reported previously (22). The droplets were prepared by mixing 2 3 l of purified NylA solution (10 mg ml 1 protein in 20 mm phosphate buffer containing 10% glycerol, ph 7.3) and the same volume of reservoir solution containing 1.0 M sodium citrate as a precipitant in 0.1 M imidazole buffer (ph 8.0) and 12.5% glycerol and equilibrated against 100 l of reservoir solution at 10 C. The crystal belonged to space group P6 2, with unit cell parameters a b , c Å (see Table 1). For native crystals, the crystals were soaked for 24 h in cryoprotectant solution (1.0 M sodium citrate, 0.1 M imidazole, ph 8.0, 25% glycerol). Heavy atom derivatives were prepared by soaking the crystals for 24 h in cryoprotectant solution containing 1 mm HgCl 2. For analysis of enzyme substrate complexes, the activity-deficient S174A mutant of NylA (NylA-A 174 ) was used to prevent hydrolysis of the substrate during crystallographic analysis. the NylA- A 174 Acd complex was prepared by soaking the crystals in the cryoprotectant solution containing 20 mm Acd for 3 h. Cryocooling was performed by blowing cold nitrogen steam onto the crystals at 100 K. The diffraction data sets of the NylA crystal and NylA-A 174 Acd complex were collected at the SPring-8 (Hyogo, Japan) Beamlines BL38B1 (equipped with a Rigaku Jupiter CCD detector system) and BL41XU (equipped with an ADSC Quantum 315 detector system), respectively. For HgCl 2 derivatives, diffraction data were recorded on Beamline BL-5A at the Photon Factory (Tsukuba, Japan) using an ADSC Quantum 315 detector system. The following parameters were chosen for data collection: wavelength, Å; crystal to detector distance, 180 mm for native crystals, 300 mm for mercury derivatives, and 200 mm for NylA-A 174 Acd complexes; oscillation range/image, 0.5 for native crystals and mercury derivatives or 1.0 for NylA-A 174 Acd complexes. Indexing, integration, and scaling of reflections were performed using the HKL2000 program package (24). The diffraction data were collected from native NylA crystal, NylA-mercury (II) dichloride derivative, and NylA-A 174 Acd complex crystal to resolutions of 1.90, 2.06, and 1.80 Å, respectively (see Table 1). Phase Determination, Model Building, and Crystallographic Refinement The NylA structure was determined by the single wavelength anomalous diffraction method using the HgCl 2 derivative data. The mercury substructure for the derivative crystal was solved at 2.1 Å resolution by the program BnP (25) using the anomalous signal of the mercury atoms. Initial phase parameters were determined using the program SHARP (26). The electron density map was automatically traced with ARP/ warp. The models for the untraced regions, except for amino acids at position 1 4, 19 26, , and , were constructed by manual model building using XFIT (27). Rigid body refinement was performed using the coordinates of the initial model to fit the unit cell of the NylA crystal, followed by positional and B-factor refinement with the program CNS (28). For the NylA-A 174 Acd complex, rigid body refinement was performed using the coordinates of the refined NylA structure to fit the unit cell of the NylA-A 174 Acd complex, followed by positional and B-factor refinement with the program CNS. With several cycles of a manual model rebuilding by XFIT, R-factor and R free were 18.1 and 19.5% (for NylA) and 18.3 and 19.5% (for the NylA-A 174 Acd complex), respectively. The Molprobity (29) Ramachandran analysis showed 467 residues (97.1%) in favored regions, 481 residues (99.8%) in allowed regions, and 1 residue (Thr 75 ) (0.2%) in outliers for NylA and 465 residues (96.9%) in favored regions, 479 residues (99.8%) in allowed regions, and 1 residue (Ser 416 ) (0.2%) in outliers for the NylA-A 174 Acd complex. The electron density map of Thr 75 and Ser 416, located in outliers, was clear enough to assign the structures without ambiguity. The results of the crystal structure analysis are summarized in Table 1. Figures of three-dimensional models of proteins were generated with the program MolFeat (version 3.6; FiatLux Co., Tokyo, Japan). RESULTS AND DISCUSSION Overall Structure of NylA and Structural Comparison with Proteins in Protein Data Bank The structure of NylA from Arthrobacter sp. KI72 was solved with single anonymous dispersion phasing using a mercury derivative crystal of the enzyme at 2.2 Å resolution, and models of the native enzymes were constructed at 1.9 Å resolution (Table 1). NylA consists of a single polypeptide chain of 472 amino acid residues (3). The overall structure of the molecule is a compact mixed / fold containing a central irregular -sheet composed of 11 -strands surrounded by 20 -helices. Each -strand is sequentially rotated in the same direction with respect to an adjacent -strand, such that the -strand at one end of the molecule is rotated 180 o relative to the -strand at the other end of the molecule (see Fig. 2A). A homology search based on the NylA structure was carried out using the DALI program. Among the 55,000 proteins registered in the Protein Data Bank, five proteins classified as AS superfamily, glutamyl-trna Gln amidotransferase subunit A (GatA) (z score, 49.4) (Protein Data Bank code 2df4) (9), probable amidase (z score, 45.0) (Protein Data Bank code 2dc0), malonamidase E2 (MAE2) (z score, 44.8) (Protein Data Bank code 1ocm) (12), peptide amidase (PAM) (z score, 44.0) (Protein Data Bank code 1m21) (16), and fatty acid amide hydrolase (z score, 38.2) (Protein Data Bank code 1mt5) (14), and their mutants were extracted as proteins with high z scores. The z scores of other proteins in Protein Data Bank were found to be lower than 4.5. Bacterial glutamyl-trna Gln amidotransferase is a heterotrimer (GatCAB). GatA receives top scores in a Protein Data Bank homology search; the protein possesses glutaminase activity for generating an ammonia donor for recruitment of the amidotransferase reaction of misacylated Glu-tRNA Gln to a corrected Gln-tRNA Gln. MAE2 and PAM catalyze the hydrolysis of malonamide and peptide amide, respectively. Fatty acid amide hydrolase is a transmembrane protein that catalyzes the hydrolysis of fatty-acid amide. Functionally, GatA, MAE2, and PAM catalyze the hydrolysis of terminal amide bonds, releasing ammonia, whereas NylA specifically hydrolyzes internal amide linkages of Acd (Fig. 1). Although probable amidase (Protein Data Bank code 2dc0) was extracted as the second highest z score, its function has not been identified. Accordingly, structural and functional comparison mainly focuses on GatA, MAE2, and PAM, which have z scores higher than 40. The structurally superimposable regions comprise 451 residues (GatA), 405 residues (MAE2), or 439 residues (PAM), and JANUARY 8, 2010 VOLUME 285 NUMBER 2 JOURNAL OF BIOLOGICAL CHEMISTRY 1241
4 TABLE 1 Data collection statistics The values in parentheses are for the outer resolution shell. Data collection NylA (native) NylA-A 174 Acd complex Mercury (II) chloride derivative Space group P6 2 P6 2 P6 2 Unit cell a b (Å) c (Å) Wavelength (Å) Resolution (Å) ( ) ( ) ( ) Total reflections 485, , ,039 Unique reflections 44,300 (4,021) 52,953 (5,212) 34,955 (2,980) Completeness (%) 98.6 (90.4) 99.7 (99.1) 97.2 (82.3) R merge (%) 3.8 (30.4) 5.4 (48.9) 5.8 (33.4) I / (I) 24.5 (5.2) 22.3 (5.0) 16.5 (6.6) Phasing Figure of merit, centric/acentric 0.106/0.380 Phasing power 1.67 R Curris Refinement Resolution range (Å) ( ) ( ) R work (%) a 18.1 (26.2) 18.7 (27.9) R free (%) b 21.4 (28.4) 22.2 (33.0) No. of protein atoms 3,602 3,592 No. of ligand atoms 0 16 No. of water molecules Root mean square bond distances (Å) Root mean square bond angles ( ) Average B-factor Protein Ligand 45.7 Solvent a R work hkl F obs k F calc ( hkl F obs ) 1 ; k, scaling factor. b R work hkl F obs k F calc ( hkl F obs ) 1, where all of the reflections belong to a test set of randomly selected data. the root mean square deviations of the superimposed C atoms were calculated to be Å. The loop region flanked by 9 and H17 in NylA was residues longer than the corresponding region in the other AS superfamily enzymes (supplemental Figs. S1 and S2). As discussed below, we suggest that dynamic motion of this flexible loop is important for the catalytic function of NylA. FIGURE 1. Hydrolysis of various amide and ester compounds. A, Acd. B, Gln. C, malonamide. D, indoleacetamide. E, p-nitrophenylacetate (C2-ester). Functional Comparison with Typical AS Superfamily Enzymes To examine the activity of NylA on substrates recognized by typical AS superfamily enzymes, we examined glutaminase, malonamidase, and indoleacetamide hydrolase activity in NylA (Fig. 1, B D). TLC analyses demonstrated that the product (Ald) was detected from Acd within 2hofthereaction using purified NylA (0.1 mg ml 1 ). In contrast, no detectable amount of product (Glu) was obtained from the substrate (Gln), even after continuing the reaction for 48 h using 1 mg ml 1 of NylA (data not shown). This suggests that the glutaminase activity caused by NylA was less than 0.5% of the level of Acd hydrolytic activity. Moreover, the hydrolytic activity of NylA for malonamide and indoleacetamide is 0.1% of the level of its activity for Acd. These results demonstrate that NylA specifically recognizes Acd as a preferred substrate irrespective of the structural conservations with other AS superfamily enzymes. In contrast, NylA had hydrolytic activity for p-nitrophenylacetate (C2-ester), p-nitrophenylbutyrate (C4-ester), and p-nitrophenyloctanoate (C8-ester), which are almost equivalent to the level of activity for Acd (Fig. 1E and Table 2). The structural basis for substrate recognition, on the basis of the enzyme/substrate interaction, is discussed below. Enzyme/Substrate Interaction Catalytic Residues In AS superfamily enzymes, the Ser-cis- Ser-Lys triad has been proposed to make up the catalytic residues (8 21). The NylA triad (Ser 174 -cis-ser 150 -Lys 72 ) spatially shares a similar position with that of MAE2 (Ser 155 -cis-ser JOURNAL OF BIOLOGICAL CHEMISTRY VOLUME 285 NUMBER 2 JANUARY 8, 2010
5 TABLE 2 Effect of amino acid substitution in NylA on Acd hydrolytic and esterolytic activity The effects of amino acid substitutions at positions 72, 150, 174, 125, and 316 were tested using purified NylA and its mutant enzymes. Enzyme activities for 10 mm Acd, 0.2 mm p-nitrophenylacetate (C2-ester), 0.2 mm p-nitrophenylbutyrate (C4-ester), and 0.2 mm p-nitrophenyloctanoate (C8-ester) were assayed. The specific activity ( mol min 1 (units) mg 1 ) was estimated for each enzyme. The numbers in parentheses indicate the activity relative to the specific activity of wild-type NylA. Acd hydrolytic activity was assayed in duplicate, and the average value was calculated. The deviations were found to be less than 2%. Acd hydrolytic activity Esterase activity C2-ester C4-ester C8-ester units mg 1 units mg 1 Wild type 4.17 (100) (100) (100) (100) K72A S150A S174A N125A 0.57 (13.7) (71.7) (62.3) (70.9) C316A 0.73 (17.5) (90.6) (108) (101) C316S 4.46 (107) (162) (142) (225) C316G 7.70 (185) (125) (129) (124) C316D (1.51) (42.9) (83.1) (68.2) C316E (0.96) (59.2) (84.3) (64.9) C316N 0.18 (4.32) (43.5) (62.9) (50.7) C316K 0.15 (3.60) (44.0) (62.4) (55.4) FIGURE 2. Stereoviews of NylA structures. A, ribbon diagram of the overall structure of NylA. -Helices and -strands are colored in green and orange, respectively, with the number shown in Fig. S1. Substrate Acd and catalytic residues (Lys 72, cis-ser 150, and Ser 174 ) are shown by a ball and stick diagram colored in red and blue, respectively. B, superimposition of NylA structure (green) with glutamyl-trna Gln amidotransferase subunit A (Protein Data Bank code 2df4; blue), malonamidase E2 (Protein Data Bank code 1ocm; purple), and peptide amidase (Protein Data Bank code 1m21; olive). Superimposition was carried out for the Ser-cis-Ser-Lys catalytic triad and its linked region on the transformation matrix generated by secondary structure matching (45). Lys 62 ), GatA (Ser 178 -cis-ser 154 -Lys 79 ), and PAM (Ser 226 -cis- Ser 202 -Lys 123 ) (Fig. 2B). To examine whether the NylA triad is responsible for catalytic function, we individually replaced Ser 174, cis-ser 150, and Lys 72 with Ala by site-directed mutagenesis, purified the mutant enzymes to homogeneity, and assayed the enzymatic activities under standard assay conditions (10 Three-dimensional Structure of Nylon Oligomer Hydrolase mm Acd). Single S174A, S150A, or K72A mutations resulted in drastic decreases in Acd hydrolytic and esterolytic activities to undetectable levels ( unit mg 1 ) (Table 2), indicating that the Ser 174 -cis- Ser 150 -Lys 72 triad is responsible for NylA function. To analyze the enzyme/substrate interactions, we performed x-ray crystallographic analysis of NylA- A 174 (S174A mutant of NylA) Acd complex at 1.80 Å resolution (Table 1). Amino acid residues and Acd substrate in the catalytic cleft gave a clear electron density distribution for which structural models could be determined (Fig. 3). Superimposition of the Acd-bound structure with the unbound structure revealed that the root mean square deviation at C is smaller than 0.1 Å for the overall structures. Interestingly, Acd is entirely trapped inside the protein molecules upon Acd binding (Figs. 2A, 3B, and 4). Accessibility of the substrate and water molecules to the catalytic center is discussed below. In most Ser-reactive hydrolases, the reaction steps of catalysis are divided into acylation and deacylation steps (30, 31). During acylation, the catalytic reactions are initiated by nucleophilic attack of Ser-O to the electrophilic carbonyl carbons of amide bonds in the substrate. Therefore, questions are raised as to what residue functions as a nucleophile. X-ray crystallographic analysis of NylA and the NylA-A 174 Acd complex suggested that the environment around Ser 174 -O is not sufficiently polar to promote deprotonation of Ser 174 -O H. In JANUARY 8, 2010 VOLUME 285 NUMBER 2 JOURNAL OF BIOLOGICAL CHEMISTRY 1243
6 FIGURE 3. Stereoviews of catalytic cleft of NylA and NylA-A 174 Acd complex. A and B, 2Fo Fc electron density maps of NylA (green) (A) and the NylA-A 174 Acd complex (orange) (B) contoured at 1.0. Side chain atoms of catalytic/binding residues (Lys 72, Gly 124, Asn 125, cis-ser 150, Ala 171, Ala 172, Ser 174 (Ala 174 ), and Cys 316 ), water molecules, and substrate Acd are shown. C, structure around catalytic residues of NylA (green) was superimposed with that around corresponding residues of the NylA-A 174 Acd complex (orange). Carbon, nitrogen, and oxygen atoms in substrate Acd are shown in yellow, blue, and red, respectively. Possible hydrogen bonds are indicated as red dotted lines with the distances in angstroms. addition, because Ser 174 -O His 4.8 Å away from Lys 72 -N,it is unlikely that Lys 72 directly participates in eliminating the proton from Ser 174 -O H (Fig. 3). However, Ser 150 exhibited an unusual cis-conformation. We estimate that low steric hindrance of Ser 150 for the backbone structures, because of the presence of consecutive Gly 148 /Gly 149 residues, should allow the unusual cis-conformation of Ser 150. Ser 174 -O is spatially 2.8 Å from either cis-ser 150 -O or cis-ser 150 -N (backbone) (Fig. 3C). This spatial location should enable Ser 174 -O to form hydrogen bonds simultaneously with cis- Ser 150 -O and cis-ser 150 -N. Because of the dual hydrogen bonding, it is likely that Ser 174 -O favors a deprotonated form even at physiological ph, resulting in the increase in nucleophilicity of Ser 174 (Fig. 5, A and B). It should be noted that the CD spectrum of the Ala 150 mutant was found to be similar to that of wildtype NylA (supplemental Fig. S3A). This result indicates that substitution of Ser 150, which has an unusual cis-conformer, with Ala does not cause misfolded proteins. Moreover, the Ala 150 mutant has no detectable activity ( unit mg 1 ) under any assay conditions using different Acd concentrations (0.625, 1.25, 2.5, 5.0, and 10.0 mm; almost maximum solubility in water) and an enzyme sample that has been concentrated 20-fold (2.07 mg ml 1 ). On the basis of these findings, we estimated that turnover (k cat ) of the Ala 150 mutant is s 1. Similar roles of cis-ser for increasing the nucleophilicity of catalytic Ser have been proposed for PAM and MAE2 (12, 16). Substrate-binding Residues Structural analysis of NylA and the NylA-A 174 Acd complex revealed that six hydrogen bonds are possible between NylA and Acd (Fig. 3B). Among the four hydrogen bonds connecting to cleaved amide linkages in Acd, two bonds, from Acd- C 1 -carbonyl O to Ala 171 -N (backbone) and to Ala 172 -N (backbone), are estimated to be responsible for oxyanion stabilization (see Proposed Catalytic Mechanisms ), whereas the other two bonds, from Acd-N 14 to Gly 124 -O (backbone) (3.0 Å) and to cis-ser 150 -O (2.8 Å), are estimated to be involved in Acd binding (Fig. 3C). In addition, at the uncleaved amide linkage in Acd, two hydrogen bonds, from Acd-N 7 to Cys 316 -S (3.2 Å) and from Acd-C 8 -carbonyl O to Asn 125 -N (2.7 Å), were identified (Fig. 3B). In the multiple three-dimensional structural alignments, Cys 316 in NylA is altered to Ala 313 (GatA), Gln 277 (MAE2), or Leu363 (PAM), suggesting that the interaction at position 316 is unique for NylA. Similarly, Asn 125 in NylA is altered to Met 129 (GatA) and Ser JOURNAL OF BIOLOGICAL CHEMISTRY VOLUME 285 NUMBER 2 JANUARY 8, 2010
7 FIGURE 4. Surface structures of the entrance of the catalytic cleft of NylA-A 174 Acd complex. A, stereoview of the entrance of the catalytic cleft. The flexible loop region (positions ) is shown in yellow, whereas rest of the region is shown in gray. B, stereoview of cross-section, including catalytic/binding residues and substrate Acd. The main chain folding (ribbon diagram) and side chain (ball and stick diagram) of catalytic/ binding residues (Lys 72, Gly 124, Asn 125, cis-ser 150, Ala 171, Ala 172, Ser 174 (Ala 174 ), and Cys 316 ) are shown. The surface is shown as a gray layer. Carbon, nitrogen, and oxygen atoms in substrate Acd are shown in yellow, blue, and red, respectively. (MAE2), although Asn 125 is conserved as Asn 172 in PAM (supplemental Fig. S1). To examine the effect of amino acid substitutions at positions 125 and 316 on catalytic function, we constructed the N125A and C316A mutants by site-directed mutagenesis. Assays under standard conditions showed that hydrolytic activity was 14% (Ala 125 mutant) and 18% (Ala 316 mutant) of the wild-type NylA level (Table 2). A kinetic study showed that the N125A mutation increased the K m 4-fold, suggesting that the decrease in substrate-binding is due to the loss of hydrogen bonding between the Acd-C 8 -carbonyl O and the Asn 125 -N. Similarly, the C316A substitution should result in the loss of hydrogen bonding between Acd-N 7 and Cys 316 -S. However, the C316A substitution decreased the K m to approximately one-quarter the level of wild-type NylA (Table 3). Moreover, these two mutations also decreased the k cat values to 23% (Ala 125 enzyme) and 14% (Ala 316 enzyme) of the wild-type NylA level. The effects of amino acid substitutions at position 316 were more extensively examined. It should be noted that the overall Three-dimensional Structure of Nylon Oligomer Hydrolase protein structures were not affected much by substitutions at the substrate-binding sites, because the CD spectra of the three mutants (N125A, C316S, and C316D) were very similar to the spectrum of wildtype enzyme (supplemental Fig. S3B). Substitution to acidic (Glu and Asp), basic (Lys), or other polar (Asn) residues decreased the activity to 1 4% of the wild-type level (Table 3). Although the C316S substitution had no significant effect on activity under standard assay conditions, a kinetic study showed that the K m for Acd had decreased from 3.5 mm (wild-type NylA) to 1.8 mm (Ser 316 mutant) without affecting the k cat. It is likely that possible hydrogen bonding between the Acd-N 7 and Ser 316 -O more stably positions the substrate than hydrogen bonding with Cys 316 -S. Unexpectedly, k cat was enhanced in the Gly 316 mutant without affecting the K m, even though the C316G mutation should have resulted in the loss of hydrogen bonding at position 316. In contrast, the C316D mutation significantly decreased activity. Because of the limited solubility of Acd in water, it is possible to perform kinetic studies only at concentrations lower than 10 mm Acd. Although the standard error for the calculated K m value is quite large because of extrapolation to higher concentrations, we found that the calculated K m value of the Asp 316 mutant was 20 mm, which exceeds the saturated Acd concentration, and the k cat value was 3% the level of wild-type NylA (Table 3). Thus, the binding affinity and turnover of catalysis are estimated to be drastically decreased by C316D substitutions. However, it should be demonstrated that amino acid substitutions at position 316 had less effect on carboxylesterase activity than Acd hydrolytic activity. Namely, the Acd hydrolytic activity varied 200-fold ( % the activity of wild-type NylA) among the mutants, whereas the activity for C2-, C4-, and C8-esters was only within the range of % (Table 2). Substrate recognition between amides and carboxylesters is discussed below with other Ser-reactive hydrolases. Proposed Catalytic Mechanisms From the spatial location of catalytic/binding residues and functional analysis by site-directed mutagenesis, we estimated the catalytic mechanisms of NylA (Fig. 5). Nucleophilic Attack of Ser 174 and Oxyanion Stabilization Nucleophilic Ser 174 -O activated by cis-ser 150 should attack JANUARY 8, 2010 VOLUME 285 NUMBER 2 JOURNAL OF BIOLOGICAL CHEMISTRY 1245
8 FIGURE 5. Proposed catalytic mechanism of NylA. The steps in the catalytic reaction are shown. A, free enzyme. B, enzyme substrate. C, tetrahedral intermediate. D, acyl-enzyme. E, tetrahedral intermediate. TABLE 3 Kinetic parameters of NylA mutant enzymes Acd hydrolytic activity was assayed using the purified enzymes under standard assay conditions, except that various concentrations of Acd were used. Mutation Kinetic study K m k cat k cat /K m mm s 1 s 1 mm 1 Wild type N125A C316A C316S C316G C316D the electrophilic carbonyl carbon of the substrate (Fig. 5B), with the subsequent formation of a tetrahedral intermediate containing an unstable oxyanion (Fig. 5C). Therefore, stabilization of the oxyanion is important for catalytic function. The carbonyl oxygen in Acd is spatially 2.9 and 3.0 Å from the main chain nitrogens of Ala 171 and Ala 172, respectively (Fig. 3C). Therefore, we estimate that the oxyanion formed during Acd hydrolytic reaction is stabilized by the positively charged nitrogens at Ala 171 and Ala 172 (Fig. 5C). Formation of Acyl-enzyme To break down the tetrahedral intermediate to the acyl-enzyme, the tetrahedral intermediate should accept a proton from a general acid. Because cis-ser 150 -O is close to Acd-N 14 (nitrogen of cleaved amide linkage in Acd) (distance, 2.8 Å) (Fig. 3C), it should be possible that cis-ser 150 -O H functions as the general acid, providing a proton to the departing amino group (Fig. 5C). The proton transfer should be facilitated by the amino group of Lys 72 -NH 3, which can stabilize the deprotonated Ser 150 -O (2.7 Å from Lys 72 -NH 3 ) (Figs. 3C and 5C). Thus, cis-ser 150 and Lys 72 should be involved in facilitating proton transfer and maintaining the optimum electrostatic environment to promote the acylation of Ser 174. Deacylation Conversion of the acyl-enzyme to free enzyme (deacylation) generally requires two steps, i.e. formation of a tetrahedral intermediate through nucleophilic attack of a water molecule (Fig. 5D) and regeneration of free enzyme (Fig. 5E). In NylA, water molecules responsible for deacylation have not been found around the catalytic center in the Acd-bound structure (Fig. 3B). However, as discussed below, we estimate that the enzyme accepts water molecules for deacylation from solvent water through the dynamic motion of a flexible loop region (Phe 400 Phe 414 ) (Fig. 4 and supplemental Fig. S4). The incorporated water molecule should function as a nucleophile to form a tetrahedral intermediate, and the resulting tetrahedral intermediate should be stabilized by Ala 171 -N and Ala 172 -N (Fig. 5E), in a similar fashion as the acylation step. Finally, the intermediate should be hydrolyzed to Ald plus free enzyme, which reinitiates the successive catalytic cycle. Accessibility of Substrate to Catalytic Center The catalytic (Ser 174, cis-ser 150, Lys 72 ) and binding (Asn 125 and Cys 316 ) residues in NylA are located at 2 (Lys 72 ), H7 (cis- Ser 150 ), H8 (Ser 174 ), H6 (Asn 125 ), and H14 (Cys 316 )(supplemental Fig. S1). Through these interactions, NylA should hold Acd inside the catalytic cleft, surrounded by stable secondary structures (Fig. 4B). Therefore, questions are raised regarding how the Acd substrate can penetrate close to the catalytic residues located near the center of the globular protein molecule. As stated above, the loop region (Thr 388 Phe 414 ), flanked by 9 and H17 in NylA, is residues longer than the corresponding regions in MAE2, PAM, and GatA (supplemental Fig JOURNAL OF BIOLOGICAL CHEMISTRY VOLUME 285 NUMBER 2 JANUARY 8, 2010
9 S1). It is likely that the longer loop makes it more flexible. Actually, crystallographic analysis of NylA revealed that the atomic B-factors of amino acid residues are significantly large (80 95 Å 2 ), especially at Phe 400 Arg 410, compared with helices, -strands, and the other loop regions (supplemental Fig. S4). Because the atomic B-factors correlate with the degree of thermal motion and spatial disorder in the crystal structures, the structure of the loop should significantly deviate during the enzymatic reaction. Interestingly, the flexible loop (Phe 410 Phe 414 ) region forms a catalytic cleft with a long helix (H14) and short helix (H15), including catalytic/binding residues (Fig. 4). We suggest that the dynamic motion of the loop located between -strand 9 ( 9) and Helix 17 (H17) allows Acd to incorporate into the cleft, including the catalytic center. Assuming that the substrate is incorporated in this manner, the catalytic triads (Ser 174 -cis-ser 150 -Lys 72 ) are located at the bottom of the catalytic cleft, whereas the binding residues (Asn 125 and Cys 316 ) are localized closer to the entrance of the cleft. Water Molecules In most hydrolases, the catalytic sites are open to solvent, so the enzyme can accept water molecules for deacylation from the massive solvent environment. In class A -lactamase, hydrolytic water, activated by Glu 166, is considered to be responsible for either the acylation or deacylation step (32 34). In NylA, 521 and 445 water molecules were identified for Acdbound and unbound structures, respectively. Larger numbers of water molecules in substrate-bound structures are thought to be due to the higher resolutions. In unbound structures, seven water molecules (Wat34, Wat181, Wat230, Wat231, Wat232, Wat233, and Wat260) were identified in the cleft around the catalytic residues (Fig. 3A). However, in the enzyme substrate complex, these water molecules are replaced with Acd, and no water molecules generating hydrogen bonds with substrate or catalytic residues can be identified (Fig. 3B). Wat354 in the NylA-A 174 Acd complex (Wat361 in the unbound enzyme) is 3.5 Å from Asn 125 -N. However, Wat354 is close to uncleaved amide linkages in Acd and therefore is not considered to be responsible for deacylation. These results suggest that the enzyme accepts water molecules responsible for deacylation during the dynamic motion of the flexible loop regions. Even after completing the stage of acylation, no reaction product should be released from NylA utilizing cyclic amide compounds as substrate (Fig. 5D), whereas first reaction product is released from ordinary protease/esterase utilizing the linear amide/ester compounds. Therefore, the bulky N-(6-aminohexanoyl)-6-aminohexanoyl group in the acyl-nyla enzyme transiently alters the position of the flexible loop, which allows the penetration of water molecules to the catalytic center. Alternatively, the loop region flexibly fluctuates regardless of the presence or absence of substrate and freely accepts water molecules from the solvent environment. In either case, dynamic motion of the flexible loop associated with the uptake of substrate/water molecules and release of product should be essential for the catalytic reaction of NylA. Comparison with Other Ser-reactive Hydrolases and Evolutionary Implications Typical Ser-proteases (trypsin, etc.) and lipase/esterases possess a classical Ser-His-Asp(Glu) triad, in which His abstracts the proton of Ser-OH and Asp (Glu) promotes the role of His by stabilizing the positively charged His-imidazole (30, 31). Another catalytic triad, Ser-Tyr-Lys, has been identified in NylB (Ald-hydrolase) (37 41), NylB carboxylesterase (37 41), EstB carboxylesterase (42), DD-peptidase (43), and class C -lactamase (44). NylB and a NylB/NylB hybrid protein (Hyb24DN; having a NylB-level of Ald hydrolytic activity) effectively recognize substrates having 6-aminohexanoate (nylon unit) as N-terminal residues in the substrate but possess activities for various carboxylesters (37 41). X-ray crystallographic analysis of Hyb-24DN (a NylB/NylB hybrid) and its Ald-bound structure demonstrates that Ald binding induced the transition from the open (substrate-unbound) form to the closed (substrate-bound) form through the dynamic motion of the loop region and the flip-flop of Tyr 170, whereas the enzyme performs the catalytic function as a relaxed open form during the entire catalytic cycle as a carboxylesterase (38). This model is consistent with our finding that Ald hydrolytic activity is significantly affected by amino acid substitutions at positions 170, 181, 187, 264, 266, and 370, responsible for Ald binding, whereas amino acid substitutions at these positions barely affect the esterase activity (37 41). A molecular dynamic simulation for NylB/NylB has suggested that the substrateunbound open form is maintained at an energy minimum and requires activation energy to transition to the substrate-bound closed form (38). In addition, water molecules incorporated into the catalytic cleft are excluded in the closed form but regained from the solvent water at deacylation with a back transition to the open form. We have speculated that the release of 6-aminohexanoate (corresponding to the C-terminal half of Ald) triggers the penetration of water molecules to the catalytic center (38). On the basis of these results, we have proposed that NylB has evolved from a pre-existing esterase with a -lactamase fold (37 41). Present x-ray crystallographic analysis revealed that NylA is classified as a member of the AS superfamily and utilizes the distinct catalytic triad Ser-cis-Ser-Lys. Although NylA possesses broad specificity for carboxylesters, we estimate that the unique substrate specificity of NylA recognizing the nylon-6 byproduct should be achieved by incorporating amino acid residues responsible for Acd-binding while utilizing the common Ser-cis-Ser-Lys catalytic triad conserved in the AS superfamily. Thus, the new enzyme acting on the nylon-6 byproduct seems to diverge from ancestral proteins to create families composed of highly specialized enzymes. In malonamidase E2 (MAE2, an AS family enzyme), the catalytic activity is drastically decreased by R158E and R158Q mutations, and the k cat value of the R158K mutant is similar to that of wild type with a K m value increased by 100-fold (13). In Hyb-24DN, Asp 181 -COO is responsible for electrostatic stabilization of Ald-NH 3, and D181K and D181H substitutions decreased the activity by 10,000-fold (37, 38). In contrast to the drastic effect by electrostatic stabilization, D370Y substitu- JANUARY 8, 2010 VOLUME 285 NUMBER 2 JOURNAL OF BIOLOGICAL CHEMISTRY 1247
10 tion in NylB -type carboxylesterase (Hyb-24) increased the Ald hydrolytic activity by 8.2-fold, concomitant with formation of a hydrogen bond with substrate Ald. Thus, we roughly estimate that the energetic contribution of a hydrogen bond contributes a 10-fold difference in binding affinity (39 41). However, the improvements in NylA activity caused by substitution at position 316 are extremely modest. We suggest that that the effects of amino acid substitution at the Acd-binding sites in NylA are summarized as follows: (i) Even if the hydrogen bond at Acd-N 7 is eliminated, close contact with the substrate and inner surface of the protein molecule still stably holds the substrate inside the globular protein. (ii) Because of the inflexible compact structure of Acd trapped in the catalytic cleft, amino acid substitutions interacting at the uncleaved site (Acd-N 7 ) affect the suitable positioning of the cleaved site (Acd-C 1 /Acd-N 14 ) against the catalytic residues, resulting in the decrease in k cat. (iii) Substitution of Cys 316 to a bulky/polar residue (Glu, Asp, Asn, and Lys), located at the entrance of the catalytic cleft, has a negative effect on the incorporation of the substrate into the catalytic cleft, resulting in a decrease in enzyme activity. In particular, replacement to acidic residues (Glu and Asp) significantly decreases the activity, probably because of an electrostatic effect. Finally, it should be noted that plasmid-encoded NylA from Arthrobacter and Pseudomonas and -laurolactam hydrolase from Rhodococcus, Cupriavidus, and Sphingomonas exhibit 98 99% overall homology, although these strains have been isolated independently and classified as different genera. These results may imply that the nyla-related genes have been recently distributed among microorganisms in the course of evolution, probably by plasmid-mediated gene transfer (4 7, 35, 36). Acknowledgment We thank Prof. Yuji Goto (Osaka University) for analyzing the CD spectra of NylA and its mutant enzymes. REFERENCES 1. Negoro, S. (2000) Appl. Microbiol. Biotechnol. 54, Negoro, S. (2002) Biopolymers 9, Kinoshita, S., Negoro, S., Muramatsu, M., Bisaria, V. S., Sawada, S., and Okada, H. (1977) Eur. J. Biochem. 80, Kanagawa, K., Negoro, S., Takada, N., and Okada, H. (1989) J. Bacteriol. 171, Tsuchiya, K., Fukuyama, S., Kanzaki, N., Kanagawa, K., Negoro, S., and Okada, H. (1989) J. Bacteriol. 171, Yasuhira, K., Tanaka, Y., Shibata, H., Kawashima, Y., Ohara, A., Kato, D., Takeo, M., and Negoro, S. (2007) Appl. Environ. Microbiol. 73, Kato, K., Ohtsuki, K., Koda, Y., Maekawa, T., Yomo, T., Negoro, S., and Urabe, I. (1995) Microbiology 141, Schmitt, E., Panvert, M., Blanquet, S., and Mechulam, Y. (2005) Structure 13, Nakamura, A., Yao, M., Chimnaronk, S., Sakai, N., and Tanaka, I. (2006) Science 312, Neu, D., Lehmann, T., Elleuche, S., and Pollmann, S. (2007) FEBS J. 274, Yamada, T., Palm, C. J., Brooks, B., and Kosuge, T. (1985) Proc. Natl. Acad. Sci. U.S.A. 82, Shin, S., Lee, T. H., Ha, N. C., Koo, H. M., Kim, S. Y., Lee, H. S., Kim, Y. S., and Oh, B. H. (2002) EMBO J. 21, Yun, Y. S., Lee, W., Shin, S., Oh, B. H., and Choi, K. Y. (2006) J. Biol. Chem. 281, Bracey, M. H., Hanson, M. A., Masuda, K. R., Stevens, R. C., and Cravatt, B. F. (2002) Science 298, Wei, B. Q., Mikkelsen, T. S., McKinney, M. K., Lander, E. S., and Cravatt, B. F. (2006) J. Biol. Chem. 281, Labahn, J., Neumann, S., Büldt, G., Kula, M. R., and Granzin, J. (2002) J. Mol. Biol. 322, Valiña, A. L., Mazumder-Shivakumar, D., and Bruice, T. C. (2004) Biochemistry 43, Kobayashi, M., Goda, M., and Shimizu, S. (1998) FEBS Lett. 439, Cilia, E., Fabbri, A., Uriani, M., Scialdone, G. G., and Ammendola, S. (2005) FEBS J. 272, Asano, Y., Fukuta, Y., Yoshida, Y., and Komeda, H. (2008) Biosci. Biotechnol. Biochem. 72, Heumann, S., Eberl, A., Fischer-Colbrie, G., Pobeheim, H., Kaufmann, F., Ribitsch, D., Cavaco-Paulo, A., and Guebitz, G. M. (2009) Biotechnol. Bioeng. 102, Yasuhira, K., Uedo, Y., Shibata, N., Negoro, S., Takeo, M., and Higuchi, Y. (2006) Acta Crystallogr. Sect. F Struct. Biol. Cryst. Commun. 62, Ito, W., Ishiguro, H., and Kurosawa, Y. (1991) Gene 102, Otwinowski, Z., and Minor, W. (1997) Methods Enzymol. 276, Weeks, C. M., Blessing, R. H., Miller, R., Mungee, R., Potter, S. A., Rappleye, J., Smith, G. D., Xu, H., and Furey, W. (2002) Z. Kristallogr. 217, de La Fortelle, E., and Bricogne, G. (1997) Methods Enzymol. 276, McRee, D. E. (1993) Practical Protein Crystallography, Academic Press, San Diego, CA 28. Brünger, A. T., Adams, P. D., Clore, G. M., DeLano, W. L., Gros, P., Grosse-Kunstleve, R. W., Jiang, J. S., Kuszewski, J., Nilges, N., Pannu, N. S., Read, R. J., Rice, L. M., Simonson, T., and Warren, G. L. (1998) Acta Crystallogr. D Biol. Crystallogr. 54, Davis, I. W., Leaver-Fay, A., Chen, V. B., Block, J. N., Kapral, G. J., Wang, X., Murray, L. W., Arendall, W. B., 3rd, Snoeyink, J., Richardson, J. S., and Richardson, D. C. (2007) Nucleic Acids Res. 35, W375 W Fersht, A. R. (1984) Enzyme Structure and Mechanism (2nd Ed), pp , W. H. Freeman and Co., New York 31. Arpigny, J. L., and Jaeger, K. E. (1999) Biochem. J. 343, Zawadzke, L. E., Chen, C. C., Banerjee, S., Li, Z., Wäsch, S., Kapadia, G., Moult, J., and Herzberg, O. (1996) Biochemistry 35, Minasov, G., Wang, X., and Shoichet, B. K. (2002) J. Am. Chem. Soc. 124, Hermann, J. C., Ridder, L., Mulholland, A. J., and Höltje, H. D. (2003) J. Am. Chem. Soc. 125, Okada, H., Negoro, S., Kimura, H., and Nakamura, S. (1983) Nature 306, Yasuhira, K., Uedo, Y., Takeo, M., Kato, D., and Negoro, S. (2007) J. Biosci. Bioeng. 104, Negoro, S., Ohki, T., Shibata, N., Mizuno, N., Wakitani, Y., Tsurukame, J., Matsumoto, K., Kawamoto, I., Takeo, M., and Higuchi, Y. (2005) J. Biol. Chem. 280, Negoro, S., Ohki, T., Shibata, N., Sasa, K., Hayashi, H., Nakano, H., Yasuhira, K., Kato, D., Takeo, M., and Higuchi, Y. (2007) J. Mol. Biol. 370, Ohki, T., Wakitani, Y., Takeo, M., Yasuhira, K., Shibata, N., Higuchi, Y., and Negoro, S. (2006) FEBS Lett. 580, Kawashima, Y., Ohki, T., Shibata, N., Higuchi, Y., Wakitani, Y., Matsuura, Y., Nakata, Y., Takeo, M., Kato, D., and Negoro, S. (2009) FEBS J. 276, Ohki, T., Shibata, N., Higuchi, Y., Kawashima, Y., Takeo, M., Kato, D., and Negoro, S. (2009) Protein Sci. 18, Wagner, U. G., Petersen, E. I., Schwab, H., and Kratky, C. (2002) Protein Sci. 11, Silvaggi, N. R., Anderson, J. W., Brinsmade, S. R., Pratt, R. F., and Kelly, J. A. (2003) Biochemistry 42, Nukaga, M., Kumar, S., Nukaga, K., Pratt, R. F., and Knox, J. R. (2004) J. Biol. Chem. 279, Laskowski, R. A., McArthur, M. W., Moss, D. S., and Thornton, J. M. (1993) J. Appl. Crystallogr. 265, JOURNAL OF BIOLOGICAL CHEMISTRY VOLUME 285 NUMBER 2 JANUARY 8, 2010
X-ray structures of fructosyl peptide oxidases revealing residues responsible for gating oxygen access in the oxidative half reaction
 X-ray structures of fructosyl peptide oxidases revealing residues responsible for gating oxygen access in the oxidative half reaction Tomohisa Shimasaki 1, Hiromi Yoshida 2, Shigehiro Kamitori 2 & Koji
X-ray structures of fructosyl peptide oxidases revealing residues responsible for gating oxygen access in the oxidative half reaction Tomohisa Shimasaki 1, Hiromi Yoshida 2, Shigehiro Kamitori 2 & Koji
Appendix B Dansyl probe syntheses and characterization and D-8-Ad:P450cam structure determination
 201 Appendix B Dansyl probe syntheses and characterization and D-8-Ad:P450cam structure determination Acknowlegements. The structure of the D-8-Ad:P450cam conjugate was determined by Anna-Maria A. Hays.
201 Appendix B Dansyl probe syntheses and characterization and D-8-Ad:P450cam structure determination Acknowlegements. The structure of the D-8-Ad:P450cam conjugate was determined by Anna-Maria A. Hays.
Supporting Online Material. Av1 and Av2 were isolated and purified under anaerobic conditions according to
 Supporting Online Material Materials and Methods Av1 and Av2 were isolated and purified under anaerobic conditions according to published protocols (S1). Crystals of nf-, pcp- and adp-av2:av1 complexes
Supporting Online Material Materials and Methods Av1 and Av2 were isolated and purified under anaerobic conditions according to published protocols (S1). Crystals of nf-, pcp- and adp-av2:av1 complexes
Suppl. Figure 1: RCC1 sequence and sequence alignments. (a) Amino acid
 Supplementary Figures Suppl. Figure 1: RCC1 sequence and sequence alignments. (a) Amino acid sequence of Drosophila RCC1. Same colors are for Figure 1 with sequence of β-wedge that interacts with Ran in
Supplementary Figures Suppl. Figure 1: RCC1 sequence and sequence alignments. (a) Amino acid sequence of Drosophila RCC1. Same colors are for Figure 1 with sequence of β-wedge that interacts with Ran in
Supplementary Note 1. Enzymatic properties of the purified Syn BVR
 Supplementary Note 1. Enzymatic properties of the purified Syn BVR The expression vector pet15b-syn bvr allowed us to routinely prepare 15 mg of electrophoretically homogenous Syn BVR from 2.5 L of TB-medium
Supplementary Note 1. Enzymatic properties of the purified Syn BVR The expression vector pet15b-syn bvr allowed us to routinely prepare 15 mg of electrophoretically homogenous Syn BVR from 2.5 L of TB-medium
Solutions to 7.02 Quiz II 10/27/05
 Solutions to 7.02 Quiz II 10/27/05 Class Average = 83 Standard Deviation = 9 Range Grade % 87-100 A 43 74-86 B 39 55-73 C 17 > 54 D 1 Question 1 (56 points) While studying deep sea bacteria, you discover
Solutions to 7.02 Quiz II 10/27/05 Class Average = 83 Standard Deviation = 9 Range Grade % 87-100 A 43 74-86 B 39 55-73 C 17 > 54 D 1 Question 1 (56 points) While studying deep sea bacteria, you discover
The Skap-hom Dimerization and PH Domains Comprise
 Molecular Cell, Volume 32 Supplemental Data The Skap-hom Dimerization and PH Domains Comprise a 3 -Phosphoinositide-Gated Molecular Switch Kenneth D. Swanson, Yong Tang, Derek F. Ceccarelli, Florence Poy,
Molecular Cell, Volume 32 Supplemental Data The Skap-hom Dimerization and PH Domains Comprise a 3 -Phosphoinositide-Gated Molecular Switch Kenneth D. Swanson, Yong Tang, Derek F. Ceccarelli, Florence Poy,
Protein Structure/Function Relationships
 Protein Structure/Function Relationships W. M. Grogan, Ph.D. OBJECTIVES 1. Describe and cite examples of fibrous and globular proteins. 2. Describe typical tertiary structural motifs found in proteins.
Protein Structure/Function Relationships W. M. Grogan, Ph.D. OBJECTIVES 1. Describe and cite examples of fibrous and globular proteins. 2. Describe typical tertiary structural motifs found in proteins.
Case 7 A Storage Protein From Seeds of Brassica nigra is a Serine Protease Inhibitor
 Case 7 A Storage Protein From Seeds of Brassica nigra is a Serine Protease Inhibitor Focus concept Purification of a novel seed storage protein allows sequence analysis and determination of the protein
Case 7 A Storage Protein From Seeds of Brassica nigra is a Serine Protease Inhibitor Focus concept Purification of a novel seed storage protein allows sequence analysis and determination of the protein
Case 7 A Storage Protein From Seeds of Brassica nigra is a Serine Protease Inhibitor Last modified 29 September 2005
 Case 7 A Storage Protein From Seeds of Brassica nigra is a Serine Protease Inhibitor Last modified 9 September 005 Focus concept Purification of a novel seed storage protein allows sequence analysis and
Case 7 A Storage Protein From Seeds of Brassica nigra is a Serine Protease Inhibitor Last modified 9 September 005 Focus concept Purification of a novel seed storage protein allows sequence analysis and
Supporting Information
 Title Structure of bacterial cellulose synthase subunit D Hu, Song-Qing; Gao, Yong-Gui; Tajima, Kenji; Sunagaw Author(s) Yoda, Takanori; Shimura, Daisuke; Satoh, Yasuharu; M CitationProceedings of the
Title Structure of bacterial cellulose synthase subunit D Hu, Song-Qing; Gao, Yong-Gui; Tajima, Kenji; Sunagaw Author(s) Yoda, Takanori; Shimura, Daisuke; Satoh, Yasuharu; M CitationProceedings of the
BC 367, Exam 2 November 13, Part I. Multiple Choice (3 pts each)- Please circle the single best answer.
 Name BC 367, Exam 2 November 13, 2008 Part I. Multiple Choice (3 pts each)- Please circle the single best answer. 1. The enzyme pyruvate dehydrogenase catalyzes the following reaction. What kind of enzyme
Name BC 367, Exam 2 November 13, 2008 Part I. Multiple Choice (3 pts each)- Please circle the single best answer. 1. The enzyme pyruvate dehydrogenase catalyzes the following reaction. What kind of enzyme
Proteins were extracted from cultured cells using a modified buffer, and immunoprecipitation and
 Materials and Methods Immunoprecipitation and immunoblot analysis Proteins were extracted from cultured cells using a modified buffer, and immunoprecipitation and immunoblot analyses with corresponding
Materials and Methods Immunoprecipitation and immunoblot analysis Proteins were extracted from cultured cells using a modified buffer, and immunoprecipitation and immunoblot analyses with corresponding
Protein analysis. Dr. Mamoun Ahram Summer semester, Resources This lecture Campbell and Farrell s Biochemistry, Chapters 5
 Protein analysis Dr. Mamoun Ahram Summer semester, 2015-2016 Resources This lecture Campbell and Farrell s Biochemistry, Chapters 5 Bases of protein separation Proteins can be purified on the basis Solubility
Protein analysis Dr. Mamoun Ahram Summer semester, 2015-2016 Resources This lecture Campbell and Farrell s Biochemistry, Chapters 5 Bases of protein separation Proteins can be purified on the basis Solubility
Dr. R. Sankar, BSE 631 (2018)
 Pauling, Corey and Branson Diffraction of DNA http://www.nature.com/scitable/topicpage/dna-is-a-structure-that-encodes-biological-6493050 In short, stereochemistry is important in determining which helices
Pauling, Corey and Branson Diffraction of DNA http://www.nature.com/scitable/topicpage/dna-is-a-structure-that-encodes-biological-6493050 In short, stereochemistry is important in determining which helices
absorption spectra were measured on a Hewlett Packard 8452A diode array spectrophotometer.
 S1 Experimental General: P450cam was expressed and purified as previously described. 1 Steady state UV-visible absorption spectra were measured on a Hewlett Packard 8452A diode array spectrophotometer.
S1 Experimental General: P450cam was expressed and purified as previously described. 1 Steady state UV-visible absorption spectra were measured on a Hewlett Packard 8452A diode array spectrophotometer.
6-Foot Mini Toober Activity
 Big Idea The interaction between the substrate and enzyme is highly specific. Even a slight change in shape of either the substrate or the enzyme may alter the efficient and selective ability of the enzyme
Big Idea The interaction between the substrate and enzyme is highly specific. Even a slight change in shape of either the substrate or the enzyme may alter the efficient and selective ability of the enzyme
CFSSP: Chou and Fasman Secondary Structure Prediction server
 Wide Spectrum, Vol. 1, No. 9, (2013) pp 15-19 CFSSP: Chou and Fasman Secondary Structure Prediction server T. Ashok Kumar Department of Bioinformatics, Noorul Islam College of Arts and Science, Kumaracoil
Wide Spectrum, Vol. 1, No. 9, (2013) pp 15-19 CFSSP: Chou and Fasman Secondary Structure Prediction server T. Ashok Kumar Department of Bioinformatics, Noorul Islam College of Arts and Science, Kumaracoil
Extracting Pure Proteins from Cells
 Extracting Pure Proteins from Cells 0 Purification techniques focus mainly on size & charge 0 The first step is homogenization (grinding, Potter Elvejhem homogenizer, sonication, freezing and thawing,
Extracting Pure Proteins from Cells 0 Purification techniques focus mainly on size & charge 0 The first step is homogenization (grinding, Potter Elvejhem homogenizer, sonication, freezing and thawing,
Supplementary Figure 1. Electron microscopy of gb-698glyco/1g2 Fab complex. a)
 Supplementary Figure 1. Electron microscopy of gb-698glyco/1g2 Fab complex. a) Representative images of 2D class averages of gb-698glyc bound to 1G2 Fab. Top views of the complex were underrepresented
Supplementary Figure 1. Electron microscopy of gb-698glyco/1g2 Fab complex. a) Representative images of 2D class averages of gb-698glyc bound to 1G2 Fab. Top views of the complex were underrepresented
SUPPLEMENTARY INFORMATION
 Molecular basis of RNA-dependent RNA polymerase II activity Elisabeth Lehmann, Florian Brueckner, and Patrick Cramer Gene Center Munich and Center for integrated Protein Science CiPS M, Department of Chemistry
Molecular basis of RNA-dependent RNA polymerase II activity Elisabeth Lehmann, Florian Brueckner, and Patrick Cramer Gene Center Munich and Center for integrated Protein Science CiPS M, Department of Chemistry
Protein expression and purification from Hi5 insect cells is as described (1, 2).
 Materials & methods Protein expression and purification Protein expression and purification from Hi5 insect cells is as described (1, 2). The purified complex exhibited a 2:2:2 stoichiometry as determined
Materials & methods Protein expression and purification Protein expression and purification from Hi5 insect cells is as described (1, 2). The purified complex exhibited a 2:2:2 stoichiometry as determined
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10963 Supplementary Table 1 Data collection, phasing and refinement statistics. Crystal Native Derivative-1 (OsO 4 ) Derivative-2 (Orange-Pt) Data collection Space group C2 C2 C2 Cell
doi:10.1038/nature10963 Supplementary Table 1 Data collection, phasing and refinement statistics. Crystal Native Derivative-1 (OsO 4 ) Derivative-2 (Orange-Pt) Data collection Space group C2 C2 C2 Cell
Supplementary Information for. Structure of human tyrosylprotein sulfotransferase-2 reveals the mechanism of protein tyrosine sulfation reaction
 Supplementary Information for Structure of human tyrosylprotein sulfotransferase-2 reveals the mechanism of protein tyrosine sulfation reaction Takamasa Teramoto, Yukari Fujikawa, Yoshirou Kawaguchi, Katsuhisa
Supplementary Information for Structure of human tyrosylprotein sulfotransferase-2 reveals the mechanism of protein tyrosine sulfation reaction Takamasa Teramoto, Yukari Fujikawa, Yoshirou Kawaguchi, Katsuhisa
Purification: Step 1. Protein and Peptide Chemistry. Lecture 11. Big Problem: Crude extract is not the natural environment. Cells: Break them open!
 Lecture 11 Protein and Peptide Chemistry Margaret A. Daugherty Fall 2003 Purification: Step 1 Cells: Break them open! Crude Extract Total contents of cell Big Problem: Crude extract is not the natural
Lecture 11 Protein and Peptide Chemistry Margaret A. Daugherty Fall 2003 Purification: Step 1 Cells: Break them open! Crude Extract Total contents of cell Big Problem: Crude extract is not the natural
Purification: Step 1. Lecture 11 Protein and Peptide Chemistry. Cells: Break them open! Crude Extract
 Purification: Step 1 Lecture 11 Protein and Peptide Chemistry Cells: Break them open! Crude Extract Total contents of cell Margaret A. Daugherty Fall 2003 Big Problem: Crude extract is not the natural
Purification: Step 1 Lecture 11 Protein and Peptide Chemistry Cells: Break them open! Crude Extract Total contents of cell Margaret A. Daugherty Fall 2003 Big Problem: Crude extract is not the natural
Ali Yaghi. Tamara Wahbeh. Mamoun Ahram
 28 Ali Yaghi Tamara Wahbeh Mamoun Ahram This sheet is a continuation of protein purification methods. Isoelectric focusing Separation of proteins based on Isoelectric points(charge),and it is a horizontal
28 Ali Yaghi Tamara Wahbeh Mamoun Ahram This sheet is a continuation of protein purification methods. Isoelectric focusing Separation of proteins based on Isoelectric points(charge),and it is a horizontal
Structural bioinformatics
 Structural bioinformatics Why structures? The representation of the molecules in 3D is more informative New properties of the molecules are revealed, which can not be detected by sequences Eran Eyal Plant
Structural bioinformatics Why structures? The representation of the molecules in 3D is more informative New properties of the molecules are revealed, which can not be detected by sequences Eran Eyal Plant
Supplementary information, Figure S1A ShHTL7 interacted with MAX2 but not another F-box protein COI1.
 GR24 (μm) 0 20 0 20 GST-ShHTL7 anti-gst His-MAX2 His-COI1 PVDF staining Supplementary information, Figure S1A ShHTL7 interacted with MAX2 but not another F-box protein COI1. Pull-down assays using GST-ShHTL7
GR24 (μm) 0 20 0 20 GST-ShHTL7 anti-gst His-MAX2 His-COI1 PVDF staining Supplementary information, Figure S1A ShHTL7 interacted with MAX2 but not another F-box protein COI1. Pull-down assays using GST-ShHTL7
SUPPLEMENTARY INFORMATION
 doi: 10.1038/nature06147 SUPPLEMENTARY INFORMATION Figure S1 The genomic and domain structure of Dscam. The Dscam gene comprises 24 exons, encoding a signal peptide (SP), 10 IgSF domains, 6 fibronectin
doi: 10.1038/nature06147 SUPPLEMENTARY INFORMATION Figure S1 The genomic and domain structure of Dscam. The Dscam gene comprises 24 exons, encoding a signal peptide (SP), 10 IgSF domains, 6 fibronectin
Molecular design principles underlying β-strand swapping. in the adhesive dimerization of cadherins
 Supplementary information for: Molecular design principles underlying β-strand swapping in the adhesive dimerization of cadherins Jeremie Vendome 1,2,3,5, Shoshana Posy 1,2,3,5,6, Xiangshu Jin, 1,3 Fabiana
Supplementary information for: Molecular design principles underlying β-strand swapping in the adhesive dimerization of cadherins Jeremie Vendome 1,2,3,5, Shoshana Posy 1,2,3,5,6, Xiangshu Jin, 1,3 Fabiana
Solutions to Problem Set 1
 MIT Department of Biology 7.014 Introductory Biology, Spring 004 Question 1 Solutions to 7.014 Problem Set 1 a) Describe the conditions of the atmosphere on prebiotic earth and how these conditions differ
MIT Department of Biology 7.014 Introductory Biology, Spring 004 Question 1 Solutions to 7.014 Problem Set 1 a) Describe the conditions of the atmosphere on prebiotic earth and how these conditions differ
AP Biology Book Notes Chapter 3 v Nucleic acids Ø Polymers specialized for the storage transmission and use of genetic information Ø Two types DNA
 AP Biology Book Notes Chapter 3 v Nucleic acids Ø Polymers specialized for the storage transmission and use of genetic information Ø Two types DNA Encodes hereditary information Used to specify the amino
AP Biology Book Notes Chapter 3 v Nucleic acids Ø Polymers specialized for the storage transmission and use of genetic information Ø Two types DNA Encodes hereditary information Used to specify the amino
has only one nucleotide, U20, between G19 and A21, while trna Glu CUC has two
 SPPLEMENTRY INFORMTION doi:1.138/nature9411 Supplementary Discussion The structural characteristics of trn Gln G in comparison to trn Glu 16 in trn Gln G is directed towards the G19 56 pair, or the outer
SPPLEMENTRY INFORMTION doi:1.138/nature9411 Supplementary Discussion The structural characteristics of trn Gln G in comparison to trn Glu 16 in trn Gln G is directed towards the G19 56 pair, or the outer
Supplementary Fig. 1. Initial electron density maps for the NOX-D20:mC5a complex obtained after SAD-phasing. (a) Initial experimental electron
 Supplementary Fig. 1. Initial electron density maps for the NOX-D20:mC5a complex obtained after SAD-phasing. (a) Initial experimental electron density map obtained after SAD-phasing and density modification
Supplementary Fig. 1. Initial electron density maps for the NOX-D20:mC5a complex obtained after SAD-phasing. (a) Initial experimental electron density map obtained after SAD-phasing and density modification
Basic concepts of molecular biology
 Basic concepts of molecular biology Gabriella Trucco Email: gabriella.trucco@unimi.it Life The main actors in the chemistry of life are molecules called proteins nucleic acids Proteins: many different
Basic concepts of molecular biology Gabriella Trucco Email: gabriella.trucco@unimi.it Life The main actors in the chemistry of life are molecules called proteins nucleic acids Proteins: many different
Speaker: Yu-Chen Ku Professor: Ching-Tsan Huang, Ph. D. Source: Biochemical Journal (2007) 402, Date: December 4th, 2007
 Directed evolution and structural analysis of N-carbamoyl-D-amino acid amidohydrolase provide insights into recombinant protein solubility in Escherichia coli Speaker: Yu-Chen Ku Professor: Ching-Tsan
Directed evolution and structural analysis of N-carbamoyl-D-amino acid amidohydrolase provide insights into recombinant protein solubility in Escherichia coli Speaker: Yu-Chen Ku Professor: Ching-Tsan
Structure and Function of the First Full-Length Murein Peptide Ligase (Mpl) Cell Wall Recycling Protein
 Paper Presentation PLoS ONE 2011 Structure and Function of the First Full-Length Murein Peptide Ligase (Mpl) Cell Wall Recycling Protein Debanu Das, Mireille Herve, Julie Feuerhelm, etc. and Dominique
Paper Presentation PLoS ONE 2011 Structure and Function of the First Full-Length Murein Peptide Ligase (Mpl) Cell Wall Recycling Protein Debanu Das, Mireille Herve, Julie Feuerhelm, etc. and Dominique
7.014 Solution Set 4
 7.014 Solution Set 4 Question 1 Shown below is a fragment of the sequence of a hypothetical bacterial gene. This gene encodes production of HWDWN, protein essential for metabolizing sugar yummose. The
7.014 Solution Set 4 Question 1 Shown below is a fragment of the sequence of a hypothetical bacterial gene. This gene encodes production of HWDWN, protein essential for metabolizing sugar yummose. The
MOLECULAR BASIS FOR THE BIRTH OF A NYLON OLIGOMER-DEGRADING ENZYME
 THE JOURNAL OF BIOLOGICAL CHEMISTRY VOL. 280, NO. 47, pp. 39644 39652, November 25, 2005 2005 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in the U.S.A. X-ray Crystallographic
THE JOURNAL OF BIOLOGICAL CHEMISTRY VOL. 280, NO. 47, pp. 39644 39652, November 25, 2005 2005 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in the U.S.A. X-ray Crystallographic
Zool 3200: Cell Biology Exam 3 3/6/15
 Name: Trask Zool 3200: Cell Biology Exam 3 3/6/15 Answer each of the following questions in the space provided; circle the correct answer or answers for each multiple choice question and circle either
Name: Trask Zool 3200: Cell Biology Exam 3 3/6/15 Answer each of the following questions in the space provided; circle the correct answer or answers for each multiple choice question and circle either
Figure A summary of spontaneous alterations likely to require DNA repair.
 DNA Damage Figure 5-46. A summary of spontaneous alterations likely to require DNA repair. The sites on each nucleotide that are known to be modified by spontaneous oxidative damage (red arrows), hydrolytic
DNA Damage Figure 5-46. A summary of spontaneous alterations likely to require DNA repair. The sites on each nucleotide that are known to be modified by spontaneous oxidative damage (red arrows), hydrolytic
SUPPLEMENTARY INFORMATION
 Structure of a tyrosyl-trna synthetase splicing factor bound to a group I intron RNA Paul J. Paukstelis 1, Jui-Hui Chen 2, Elaine Chase 2, Alan M. Lambowitz 1,*, and Barbara L. Golden 2,*,. 1 Institute
Structure of a tyrosyl-trna synthetase splicing factor bound to a group I intron RNA Paul J. Paukstelis 1, Jui-Hui Chen 2, Elaine Chase 2, Alan M. Lambowitz 1,*, and Barbara L. Golden 2,*,. 1 Institute
Chapter 5: Proteins: Primary Structure
 Instant download and all chapters Test Bank Fundamentals of Biochemistry Life at the Molecular Level 4th Edition Donald Voet https://testbanklab.com/download/test-bank-fundamentals-biochemistry-life-molecular-level-
Instant download and all chapters Test Bank Fundamentals of Biochemistry Life at the Molecular Level 4th Edition Donald Voet https://testbanklab.com/download/test-bank-fundamentals-biochemistry-life-molecular-level-
Supporting Information
 Supporting Information Nishimura et al. 10.1073/pnas.1003553107 SI Text SI Materials and Methods. Plasmid construction. The cdnas of human Gα q were amplified and subcloned into pcmv5. Mutants of Gα q
Supporting Information Nishimura et al. 10.1073/pnas.1003553107 SI Text SI Materials and Methods. Plasmid construction. The cdnas of human Gα q were amplified and subcloned into pcmv5. Mutants of Gα q
Supplementary Figure 1
 Supplementary Figure 1 2 Supplementary Figure 1: Sequence alignment of HsHSD17B8 and HsCBR4 of with KAR orthologs. The secondary structure elements as calculated by DSSP and residue numbers are displayed
Supplementary Figure 1 2 Supplementary Figure 1: Sequence alignment of HsHSD17B8 and HsCBR4 of with KAR orthologs. The secondary structure elements as calculated by DSSP and residue numbers are displayed
 Problem: The GC base pairs are more stable than AT base pairs. Why? 5. Triple-stranded DNA was first observed in 1957. Scientists later discovered that the formation of triplestranded DNA involves a type
Problem: The GC base pairs are more stable than AT base pairs. Why? 5. Triple-stranded DNA was first observed in 1957. Scientists later discovered that the formation of triplestranded DNA involves a type
BACTERIAL PRODUCTION EXPRESSION METHOD OVERVIEW: PEF # GENE NAME EXPRESSION VECTOR MOLECULAR WEIGHT kda (full-length) 34.
 BACTERIAL PRODUCTION PEF # GENE NAME EXPRESSION VECTOR MOLECULAR WEIGHT 2015-XXXX XXXX pet-32a 50.9 kda (full-length) 34.0 kda (cleaved) EXPRESSION METHOD OVERVIEW: Plasmid DNA was transformed into BL21
BACTERIAL PRODUCTION PEF # GENE NAME EXPRESSION VECTOR MOLECULAR WEIGHT 2015-XXXX XXXX pet-32a 50.9 kda (full-length) 34.0 kda (cleaved) EXPRESSION METHOD OVERVIEW: Plasmid DNA was transformed into BL21
SUPPLEMENTARY INFORMATION. Reengineering Protein Interfaces Yields Copper-Inducible Ferritin Cage Assembly
 SUPPLEMENTARY INFORMATION Reengineering Protein Interfaces Yields Copper-Inducible Ferritin Cage Assembly Dustin J. E. Huard, Kathleen M. Kane and F. Akif Tezcan* Department of Chemistry and Biochemistry,
SUPPLEMENTARY INFORMATION Reengineering Protein Interfaces Yields Copper-Inducible Ferritin Cage Assembly Dustin J. E. Huard, Kathleen M. Kane and F. Akif Tezcan* Department of Chemistry and Biochemistry,
Crystal Structure of a Self-Splicing Group I Intron with Both Exons
 Supplementary Data for: Crystal Structure of a Self-Splicing Group I Intron with Both Exons Peter L. Adams, Mary R. Stahley, Anne B. Kosek, Jimin Wang and Scott A. Strobel The supplementary material includes
Supplementary Data for: Crystal Structure of a Self-Splicing Group I Intron with Both Exons Peter L. Adams, Mary R. Stahley, Anne B. Kosek, Jimin Wang and Scott A. Strobel The supplementary material includes
Supplemental Information. The structural basis of R Spondin recognition by LGR5 and RNF43
 Supplemental Information The structural basis of R Spondin recognition by LGR5 and RNF43 Po Han Chen 1, Xiaoyan Chen 1, Deyu Fang 2, Xiaolin He 1* 1 Department of Molecular Pharmacology and Biological
Supplemental Information The structural basis of R Spondin recognition by LGR5 and RNF43 Po Han Chen 1, Xiaoyan Chen 1, Deyu Fang 2, Xiaolin He 1* 1 Department of Molecular Pharmacology and Biological
A Brief Introduction to Structural Biology and Protein Crystallography
 A Brief Introduction to Structural Biology and Protein Crystallography structural biology of H2O http://courses.cm.utexas.edu/jrobertus/ch339k/overheads-1/water-structure.jpg Protein polymers fold up into
A Brief Introduction to Structural Biology and Protein Crystallography structural biology of H2O http://courses.cm.utexas.edu/jrobertus/ch339k/overheads-1/water-structure.jpg Protein polymers fold up into
D-3-Phosphoglycerate Dehydrogenase* Gregory A. Grant, Zhiqin Hu, and Xiao Lan Xu
 THE JOURNAL OF BIOLOGICAL CHEMISTRY Vol. 276, No. 2, Issue of January 12, pp. 1078 1083, 2001 2001 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in U.S.A. Specific Interactions
THE JOURNAL OF BIOLOGICAL CHEMISTRY Vol. 276, No. 2, Issue of January 12, pp. 1078 1083, 2001 2001 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in U.S.A. Specific Interactions
Alpha-helices, beta-sheets and U-turns within a protein are stabilized by (hint: two words).
 1 Quiz1 Q1 2011 Alpha-helices, beta-sheets and U-turns within a protein are stabilized by (hint: two words) Value Correct Answer 1 noncovalent interactions 100% Equals hydrogen bonds (100%) Equals H-bonds
1 Quiz1 Q1 2011 Alpha-helices, beta-sheets and U-turns within a protein are stabilized by (hint: two words) Value Correct Answer 1 noncovalent interactions 100% Equals hydrogen bonds (100%) Equals H-bonds
SUPPLEMENTARY MATERIAL
 SUPPLEMENTARY MATERIAL Purification and biochemical characterization of acid phosphatase-i from seeds of Nelumbo nucifera Sanaullah Khan a*, Shahnaz Asmat c, Sajida Batool a, Mushtaq Ahmed b a Department
SUPPLEMENTARY MATERIAL Purification and biochemical characterization of acid phosphatase-i from seeds of Nelumbo nucifera Sanaullah Khan a*, Shahnaz Asmat c, Sajida Batool a, Mushtaq Ahmed b a Department
1) The penicillin family of antibiotics, discovered by Alexander Fleming in 1928, has the following general structure: O O
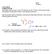 ame: TF ame: LS1a Fall 06 Problem Set #3 Due Friday 10/13 at noon in your TF s drop box on the 2 nd floor of the Science Center All questions including the (*extra*) ones should be turned in 1) The penicillin
ame: TF ame: LS1a Fall 06 Problem Set #3 Due Friday 10/13 at noon in your TF s drop box on the 2 nd floor of the Science Center All questions including the (*extra*) ones should be turned in 1) The penicillin
BIO303, Genetics Study Guide II for Spring 2007 Semester
 BIO303, Genetics Study Guide II for Spring 2007 Semester 1 Questions from F05 1. Tryptophan (Trp) is encoded by the codon UGG. Suppose that a cell was treated with high levels of 5- Bromouracil such that
BIO303, Genetics Study Guide II for Spring 2007 Semester 1 Questions from F05 1. Tryptophan (Trp) is encoded by the codon UGG. Suppose that a cell was treated with high levels of 5- Bromouracil such that
Analysis of the AcrB binding domain of the membrane fusion protein AcrA. Introduction
 Analysis of the AcrB binding domain of the membrane fusion protein AcrA Introduction Antibiotic resistance in bacteria is due in large part to multidrug efflux complexes like the TolC-AcrAB complex in
Analysis of the AcrB binding domain of the membrane fusion protein AcrA Introduction Antibiotic resistance in bacteria is due in large part to multidrug efflux complexes like the TolC-AcrAB complex in
Supporting Information
 Supporting Information Chan et al. 10.1073/pnas.0903849106 SI Text Protein Purification. PCSK9 proteins were expressed either transiently in 2936E cells (1), or stably in HepG2 cells. Conditioned culture
Supporting Information Chan et al. 10.1073/pnas.0903849106 SI Text Protein Purification. PCSK9 proteins were expressed either transiently in 2936E cells (1), or stably in HepG2 cells. Conditioned culture
Rajan Sankaranarayanan
 Proofreading during translation of the genetic code Rajan Sankaranarayanan Centre for Cellular and Molecular Biology (CCMB) Council for Scientific and Industrial Research (CSIR) Hyderabad, INDIA Accuracy
Proofreading during translation of the genetic code Rajan Sankaranarayanan Centre for Cellular and Molecular Biology (CCMB) Council for Scientific and Industrial Research (CSIR) Hyderabad, INDIA Accuracy
Supplementary Information. Single-molecule analysis reveals multi-state folding of a guanine. riboswitch
 Supplementary Information Single-molecule analysis reveals multi-state folding of a guanine riboswitch Vishnu Chandra 1,4,#, Zain Hannan 1,5,#, Huizhong Xu 2,# and Maumita Mandal 1,2,3,6* Department of
Supplementary Information Single-molecule analysis reveals multi-state folding of a guanine riboswitch Vishnu Chandra 1,4,#, Zain Hannan 1,5,#, Huizhong Xu 2,# and Maumita Mandal 1,2,3,6* Department of
Nanobiotechnology. Place: IOP 1 st Meeting Room Time: 9:30-12:00. Reference: Review Papers. Grade: 50% midterm, 50% final.
 Nanobiotechnology Place: IOP 1 st Meeting Room Time: 9:30-12:00 Reference: Review Papers Grade: 50% midterm, 50% final Midterm: 5/15 History Atom Earth, Air, Water Fire SEM: 20-40 nm Silver 66.2% Gold
Nanobiotechnology Place: IOP 1 st Meeting Room Time: 9:30-12:00 Reference: Review Papers Grade: 50% midterm, 50% final Midterm: 5/15 History Atom Earth, Air, Water Fire SEM: 20-40 nm Silver 66.2% Gold
Electronic Supplementary Information. Simultaneously sensitive detection of multiple DNA glycosylases from lung
 Electronic Supplementary Material (ESI) for Chemical Science. This journal is The Royal Society of Chemistry 2017 Electronic Supplementary Information Simultaneously sensitive detection of multiple DNA
Electronic Supplementary Material (ESI) for Chemical Science. This journal is The Royal Society of Chemistry 2017 Electronic Supplementary Information Simultaneously sensitive detection of multiple DNA
Molekulare Mechanismen der Signaltransduktion
 Molekulare Mechanismen der Signaltransduktion 12 Mechanism of auxin perception slides: http://tinyurl.com/modul-mms doi: 10.1038/nature05731 previous model http://www.plantcell.org/content/17/9/2425/f1.large.jpg
Molekulare Mechanismen der Signaltransduktion 12 Mechanism of auxin perception slides: http://tinyurl.com/modul-mms doi: 10.1038/nature05731 previous model http://www.plantcell.org/content/17/9/2425/f1.large.jpg
Structure formation and association of biomolecules. Prof. Dr. Martin Zacharias Lehrstuhl für Molekulardynamik (T38) Technische Universität München
 Structure formation and association of biomolecules Prof. Dr. Martin Zacharias Lehrstuhl für Molekulardynamik (T38) Technische Universität München Motivation Many biomolecules are chemically synthesized
Structure formation and association of biomolecules Prof. Dr. Martin Zacharias Lehrstuhl für Molekulardynamik (T38) Technische Universität München Motivation Many biomolecules are chemically synthesized
Supplementary Table 1: List of CH3 domain interface residues in the first chain (A) and
 Supplementary Tables Supplementary Table 1: List of CH3 domain interface residues in the first chain (A) and their side chain contacting residues in the second chain (B) a Interface Res. in Contacting
Supplementary Tables Supplementary Table 1: List of CH3 domain interface residues in the first chain (A) and their side chain contacting residues in the second chain (B) a Interface Res. in Contacting
Lecture 8: Affinity Chromatography-III
 Lecture 8: Affinity Chromatography-III Key words: Chromatography; Affinity chromatography; Protein Purification During this lecture, we shall be studying few more examples of affinity chromatography. The
Lecture 8: Affinity Chromatography-III Key words: Chromatography; Affinity chromatography; Protein Purification During this lecture, we shall be studying few more examples of affinity chromatography. The
Structural Bioinformatics (C3210) Conformational Analysis Protein Folding Protein Structure Prediction
 Structural Bioinformatics (C3210) Conformational Analysis Protein Folding Protein Structure Prediction Conformational Analysis 2 Conformational Analysis Properties of molecules depend on their three-dimensional
Structural Bioinformatics (C3210) Conformational Analysis Protein Folding Protein Structure Prediction Conformational Analysis 2 Conformational Analysis Properties of molecules depend on their three-dimensional
蛋白質體學. Proteomics Amino acids, Peptides and Proteins 陳威戎 & 21
 蛋白質體學 Proteomics 2015 Amino acids, Peptides and Proteins 陳威戎 2015. 09. 14 & 21 Outline 1. Amino Acids 2. Peptides and Proteins 3. Covalent Structure of Proteins Amino Acids Proteins are polymers of amino
蛋白質體學 Proteomics 2015 Amino acids, Peptides and Proteins 陳威戎 2015. 09. 14 & 21 Outline 1. Amino Acids 2. Peptides and Proteins 3. Covalent Structure of Proteins Amino Acids Proteins are polymers of amino
Fundamentals of Biochemistry
 Donald Voet Judith G. Voet Charlotte W. Pratt Fundamentals of Biochemistry Second Edition Chapter 6: Proteins: Three-Dimensional Structure Copyright 2006 by John Wiley & Sons, Inc. 1958, John Kendrew Any
Donald Voet Judith G. Voet Charlotte W. Pratt Fundamentals of Biochemistry Second Edition Chapter 6: Proteins: Three-Dimensional Structure Copyright 2006 by John Wiley & Sons, Inc. 1958, John Kendrew Any
CSE : Computational Issues in Molecular Biology. Lecture 19. Spring 2004
 CSE 397-497: Computational Issues in Molecular Biology Lecture 19 Spring 2004-1- Protein structure Primary structure of protein is determined by number and order of amino acids within polypeptide chain.
CSE 397-497: Computational Issues in Molecular Biology Lecture 19 Spring 2004-1- Protein structure Primary structure of protein is determined by number and order of amino acids within polypeptide chain.
Cristian Micheletti SISSA (Trieste)
 Cristian Micheletti SISSA (Trieste) michelet@sissa.it Mar 2009 5pal - parvalbumin Calcium-binding protein HEADER CALCIUM-BINDING PROTEIN 25-SEP-91 5PAL 5PAL 2 COMPND PARVALBUMIN (ALPHA LINEAGE) 5PAL 3
Cristian Micheletti SISSA (Trieste) michelet@sissa.it Mar 2009 5pal - parvalbumin Calcium-binding protein HEADER CALCIUM-BINDING PROTEIN 25-SEP-91 5PAL 5PAL 2 COMPND PARVALBUMIN (ALPHA LINEAGE) 5PAL 3
CRYSTALLIZATION NOTE Crystallization of Restriction Endonuclease BamHI with Nonspecific DNA
 Journal of Structural Biology 130, 81 85 (2000) doi:10.1006/jsbi.2000.4235, available online at http://www.idealibrary.com on CRYSTALLIZATION NOTE Crystallization of Restriction Endonuclease BamHI with
Journal of Structural Biology 130, 81 85 (2000) doi:10.1006/jsbi.2000.4235, available online at http://www.idealibrary.com on CRYSTALLIZATION NOTE Crystallization of Restriction Endonuclease BamHI with
Supplementary Results Supplementary Table 1. P1 and P2 enrichment scores for wild-type subtiligase.
 Supplementary Results Supplementary Table 1. P1 and P2 enrichment scores for wild-type subtiligase. Supplementary Table 2. Masses of subtiligase mutants measured by LC-MS. Supplementary Table 2 (cont d).
Supplementary Results Supplementary Table 1. P1 and P2 enrichment scores for wild-type subtiligase. Supplementary Table 2. Masses of subtiligase mutants measured by LC-MS. Supplementary Table 2 (cont d).
Colchicine. Colchicine. a b c d e
 α1-tub Colchicine Laulimalide RB3 β1-tub Vinblastine α2-tub Taxol Colchicine β2-tub Maytansine a b c d e Supplementary Figure 1 Structures of microtubule and tubulin (a)the cartoon of head to tail arrangement
α1-tub Colchicine Laulimalide RB3 β1-tub Vinblastine α2-tub Taxol Colchicine β2-tub Maytansine a b c d e Supplementary Figure 1 Structures of microtubule and tubulin (a)the cartoon of head to tail arrangement
SUPPLEMENTARY INFORMATION Figures. Supplementary Figure 1 a. Page 1 of 30. Nature Chemical Biology: doi: /nchembio.2528
 SUPPLEMENTARY INFORMATION Figures Supplementary Figure 1 a b c Page 1 of 0 11 Supplementary Figure 1: Biochemical characterisation and binding validation of the reversible USP inhibitor 1. a, Biochemical
SUPPLEMENTARY INFORMATION Figures Supplementary Figure 1 a b c Page 1 of 0 11 Supplementary Figure 1: Biochemical characterisation and binding validation of the reversible USP inhibitor 1. a, Biochemical
The Study of NF-κB Peptide Mimics and How Proteins Bind DNA
 Georgia Southern University Digital Commons@Georgia Southern University Honors Program Theses Student Research Papers 2016 The Study of NF-κB Peptide Mimics and How Proteins Bind DNA Allee M. Murray Georgia
Georgia Southern University Digital Commons@Georgia Southern University Honors Program Theses Student Research Papers 2016 The Study of NF-κB Peptide Mimics and How Proteins Bind DNA Allee M. Murray Georgia
Model Peptides Reveal Specificity of IV -Acetyltransferase from Saccharomyces cerevisiae*
 THE JOURNAL OF B~OLOCICAL CHEMISTRY 0 1990 by The American Society for Biochemistry and Molecular Biology, Inc. Vol. 265, No. 20, Issue of July 15, pp. 11576-11580,19!30 Printed in U.S. A. Model Peptides
THE JOURNAL OF B~OLOCICAL CHEMISTRY 0 1990 by The American Society for Biochemistry and Molecular Biology, Inc. Vol. 265, No. 20, Issue of July 15, pp. 11576-11580,19!30 Printed in U.S. A. Model Peptides
Sequence Determinants of a Conformational Switch in a Protein
 Sequence Determinants of a Conformational Switch in a Protein Thomas A. Anderson, Matthew H. J. Cordes, and Robert T. Sauer Kyle Skalenko 4/17/09 Introduction Protein folding is guided by the following
Sequence Determinants of a Conformational Switch in a Protein Thomas A. Anderson, Matthew H. J. Cordes, and Robert T. Sauer Kyle Skalenko 4/17/09 Introduction Protein folding is guided by the following
Supporting Information
 Supporting Information Slep et al. 10.1073/pnas.0801569105 SI Materials and Methods Mouse RGS16, residues 53 180, was subcloned into a modified pgex-2t vector (GE Healthcare) to yield a thrombin-cleavable
Supporting Information Slep et al. 10.1073/pnas.0801569105 SI Materials and Methods Mouse RGS16, residues 53 180, was subcloned into a modified pgex-2t vector (GE Healthcare) to yield a thrombin-cleavable
Hmwk # 8 : DNA-Binding Proteins : Part II
 The purpose of this exercise is : Hmwk # 8 : DNA-Binding Proteins : Part II 1). to examine the case of a tandem head-to-tail homodimer binding to DNA 2). to view a Zn finger motif 3). to consider the case
The purpose of this exercise is : Hmwk # 8 : DNA-Binding Proteins : Part II 1). to examine the case of a tandem head-to-tail homodimer binding to DNA 2). to view a Zn finger motif 3). to consider the case
BASIC MOLECULAR GENETIC MECHANISMS Introduction:
 BASIC MOLECULAR GENETIC MECHANISMS Introduction: nucleic acids. (1) contain the information for determining the amino acid sequence & the structure and function of proteins (1) part of the cellular structures:
BASIC MOLECULAR GENETIC MECHANISMS Introduction: nucleic acids. (1) contain the information for determining the amino acid sequence & the structure and function of proteins (1) part of the cellular structures:
The GeneEditor TM in vitro Mutagenesis System: Site- Directed Mutagenesis Using Altered Beta-Lactamase Specificity
 Promega Notes Magazine Number 62, 1997, p. 02 The GeneEditor TM in vitro Mutagenesis System: Site- Directed Mutagenesis Using Altered Beta-Lactamase Specificity By Christine Andrews and Scott Lesley Promega
Promega Notes Magazine Number 62, 1997, p. 02 The GeneEditor TM in vitro Mutagenesis System: Site- Directed Mutagenesis Using Altered Beta-Lactamase Specificity By Christine Andrews and Scott Lesley Promega
7.014 Problem Set 4 Answers to this problem set are to be turned in. Problem sets will not be accepted late. Solutions will be posted on the web.
 MIT Department of Biology 7.014 Introductory Biology, Spring 2005 Name: Section : 7.014 Problem Set 4 Answers to this problem set are to be turned in. Problem sets will not be accepted late. Solutions
MIT Department of Biology 7.014 Introductory Biology, Spring 2005 Name: Section : 7.014 Problem Set 4 Answers to this problem set are to be turned in. Problem sets will not be accepted late. Solutions
1/4/18 NUCLEIC ACIDS. Nucleic Acids. Nucleic Acids. ECS129 Instructor: Patrice Koehl
 NUCLEIC ACIDS ECS129 Instructor: Patrice Koehl Nucleic Acids Nucleotides DNA Structure RNA Synthesis Function Secondary structure Tertiary interactions Wobble hypothesis DNA RNA Replication Transcription
NUCLEIC ACIDS ECS129 Instructor: Patrice Koehl Nucleic Acids Nucleotides DNA Structure RNA Synthesis Function Secondary structure Tertiary interactions Wobble hypothesis DNA RNA Replication Transcription
NUCLEIC ACIDS. ECS129 Instructor: Patrice Koehl
 NUCLEIC ACIDS ECS129 Instructor: Patrice Koehl Nucleic Acids Nucleotides DNA Structure RNA Synthesis Function Secondary structure Tertiary interactions Wobble hypothesis DNA RNA Replication Transcription
NUCLEIC ACIDS ECS129 Instructor: Patrice Koehl Nucleic Acids Nucleotides DNA Structure RNA Synthesis Function Secondary structure Tertiary interactions Wobble hypothesis DNA RNA Replication Transcription
His-Spin Protein Miniprep
 INSTRUCTIONS His-Spin Protein Miniprep Catalog No. P2001 (10 purifications) and P2002 (50 purifications). Highlights Fast 5 minute protocol to purify His-tagged proteins from cell-free extracts Screen
INSTRUCTIONS His-Spin Protein Miniprep Catalog No. P2001 (10 purifications) and P2002 (50 purifications). Highlights Fast 5 minute protocol to purify His-tagged proteins from cell-free extracts Screen
Programme Good morning and summary of last week Levels of Protein Structure - I Levels of Protein Structure - II
 Programme 8.00-8.10 Good morning and summary of last week 8.10-8.30 Levels of Protein Structure - I 8.30-9.00 Levels of Protein Structure - II 9.00-9.15 Break 9.15-11.15 Exercise: Building a protein model
Programme 8.00-8.10 Good morning and summary of last week 8.10-8.30 Levels of Protein Structure - I 8.30-9.00 Levels of Protein Structure - II 9.00-9.15 Break 9.15-11.15 Exercise: Building a protein model
Packing of Secondary Structures
 7.88 Lecture Notes - 5 7.24/7.88J/5.48J The Protein Folding and Human Disease Packing of Secondary Structures Packing of Helices against sheets Packing of sheets against sheets Parallel Orthogonal Table:
7.88 Lecture Notes - 5 7.24/7.88J/5.48J The Protein Folding and Human Disease Packing of Secondary Structures Packing of Helices against sheets Packing of sheets against sheets Parallel Orthogonal Table:
Structure and Possible Mechanism of the CcbJ Methyltransferase from Streptomyces caelestis
 Supplemental material to accompany Structure and Possible Mechanism of the CcbJ Methyltransferase from Streptomyces caelestis Jacob Bauer, a Gabriela Ondrovičová, a Lucie Najmanová, b Vladimír Pevala,
Supplemental material to accompany Structure and Possible Mechanism of the CcbJ Methyltransferase from Streptomyces caelestis Jacob Bauer, a Gabriela Ondrovičová, a Lucie Najmanová, b Vladimír Pevala,
SUPPLEMENTARY INFORMATION
 This supplementary information is an extension of the letter with the same title and includes further discussion on the comparison of our designed Fe B Mb (computer model and crystal structure) with the
This supplementary information is an extension of the letter with the same title and includes further discussion on the comparison of our designed Fe B Mb (computer model and crystal structure) with the
Supplementary Information
 Supplementary Information Supplementary Figures Figure S1. Study of mgtl translation in vitro. (A) Detection of 5 LR RNA using wild-type and anti-sd (91-95) substituted templates in a transcription-translation
Supplementary Information Supplementary Figures Figure S1. Study of mgtl translation in vitro. (A) Detection of 5 LR RNA using wild-type and anti-sd (91-95) substituted templates in a transcription-translation
Supporting Information. A general chemiluminescence strategy. for measuring aptamer-target binding and target concentration
 Supporting Information A general chemiluminescence strategy for measuring aptamer-target binding and target concentration Shiyuan Li, Duyu Chen, Qingtong Zhou, Wei Wang, Lingfeng Gao, Jie Jiang, Haojun
Supporting Information A general chemiluminescence strategy for measuring aptamer-target binding and target concentration Shiyuan Li, Duyu Chen, Qingtong Zhou, Wei Wang, Lingfeng Gao, Jie Jiang, Haojun
Proteins Amides from Amino Acids
 Chapter 26 and Chapter 28 Proteins Amides from Amino Acids Amino acids contain a basic amino group and an acidic carboxyl group Joined as amides between the ¾NH 2 of one amino acid and the ¾CO 2 H to the
Chapter 26 and Chapter 28 Proteins Amides from Amino Acids Amino acids contain a basic amino group and an acidic carboxyl group Joined as amides between the ¾NH 2 of one amino acid and the ¾CO 2 H to the
Lecture for Wednesday. Dr. Prince BIOL 1408
 Lecture for Wednesday Dr. Prince BIOL 1408 THE FLOW OF GENETIC INFORMATION FROM DNA TO RNA TO PROTEIN Copyright 2009 Pearson Education, Inc. Genes are expressed as proteins A gene is a segment of DNA that
Lecture for Wednesday Dr. Prince BIOL 1408 THE FLOW OF GENETIC INFORMATION FROM DNA TO RNA TO PROTEIN Copyright 2009 Pearson Education, Inc. Genes are expressed as proteins A gene is a segment of DNA that
Comparative Modeling Part 1. Jaroslaw Pillardy Computational Biology Service Unit Cornell Theory Center
 Comparative Modeling Part 1 Jaroslaw Pillardy Computational Biology Service Unit Cornell Theory Center Function is the most important feature of a protein Function is related to structure Structure is
Comparative Modeling Part 1 Jaroslaw Pillardy Computational Biology Service Unit Cornell Theory Center Function is the most important feature of a protein Function is related to structure Structure is
Supporting Information Contents
 Supporting Information Choy Theng Loh, Kiyoshi Ozawa, Kellie L. Tuck, Nicholas Barlow, Thomas Huber, Gottfried Otting, and Bim Graham Lanthanide tags for site-specific ligation to an unnatural amino acid
Supporting Information Choy Theng Loh, Kiyoshi Ozawa, Kellie L. Tuck, Nicholas Barlow, Thomas Huber, Gottfried Otting, and Bim Graham Lanthanide tags for site-specific ligation to an unnatural amino acid
Bioinformatics. ONE Introduction to Biology. Sami Khuri Department of Computer Science San José State University Biology/CS 123A Fall 2012
 Bioinformatics ONE Introduction to Biology Sami Khuri Department of Computer Science San José State University Biology/CS 123A Fall 2012 Biology Review DNA RNA Proteins Central Dogma Transcription Translation
Bioinformatics ONE Introduction to Biology Sami Khuri Department of Computer Science San José State University Biology/CS 123A Fall 2012 Biology Review DNA RNA Proteins Central Dogma Transcription Translation
Computational Methods for Protein Structure Prediction
 Computational Methods for Protein Structure Prediction Ying Xu 2017/12/6 1 Outline introduction to protein structures the problem of protein structure prediction why it is possible to predict protein structures
Computational Methods for Protein Structure Prediction Ying Xu 2017/12/6 1 Outline introduction to protein structures the problem of protein structure prediction why it is possible to predict protein structures
Bi 8 Lecture 7. Ellen Rothenberg 26 January Reading: Ch. 3, pp ; panel 3-1
 Bi 8 Lecture 7 PROTEIN STRUCTURE, Functional analysis, and evolution Ellen Rothenberg 26 January 2016 Reading: Ch. 3, pp. 109-134; panel 3-1 (end with free amine) aromatic, hydrophobic small, hydrophilic
Bi 8 Lecture 7 PROTEIN STRUCTURE, Functional analysis, and evolution Ellen Rothenberg 26 January 2016 Reading: Ch. 3, pp. 109-134; panel 3-1 (end with free amine) aromatic, hydrophobic small, hydrophilic
