Sanger Sequencing: Troubleshooting Guide
|
|
|
- Reginald Wade
- 6 years ago
- Views:
Transcription
1 If you need help analysing your Sanger sequencing output, this guide can help. CONTENTS 1 Introduction Sequence Data Evaluation Troubleshooting Reviewing the Sequence Electropherogram Raw Sequence Result Evaluation Failed sequence Weak sequence Short sequence (or shorter than expected) Multiple sequences Artifacts Review AGRF submission Reviewing Experimental Setup of SEQ Reaction DNA Template Review Primer Design Review Contact AGRF Sequencing Page 1 of 11
2 1 Introduction This document highlights some common problems associated with DNA sequencing as well as the possible causes and solutions for these problems. Pictures of sequence traces are provided where possible along with the information describing the problem, how to identify the problem, the cause, and the potential solution for the problem. Other problems can occur with sequence data, but the following are those seen most commonly. Use this guide as the AGRF recommended data review/troubleshooting process. 2 Sequence Data Evaluation For each sample processed, the following are provided: Filename.ab1: The raw chromatogram trace file Filename.seq: A text file of the sequence, as generated by the sequencing instruments Filename.fa: A quality trimmed FASTA formatted text file Filename.bn: A BLAST file (GenBank) of the quality trimmed FASTA file The filename.ab1 file contains annotation of the sample, the raw data trace and the analysed electropherogram. Basecalling and analysis algorithms are applied to the raw data to create the analysed data trace. When evaluating or trouble-shooting sequence data, it is important to look at the raw, and analysed data traces. The raw data trace should show an even distribution of peaks across the read and no residual dyes (Figure 1). The analysed data trace should show sharp, evenly spaced peaks across the read and a clear baseline (Figure 2). AGRF recommends the use of Applied Biosystem s free Sequence Scanner software (available for download at - sequencing/sanger-sequencing/sanger-dna-sequencing/sanger-sequencing-dataanalysis.html) Figure 1: Raw Data Page 2 of 11
3 Figure 2: Analysed data It is equally important to look at data values displayed in the annotation file (Figure 3). It is useful to check the following: Average signal to noise ratio indicates labelling efficiency and should fall between be 100 and 750. The base call start indicates the scan point at which the read commences at and should be ~600 to 800. The end point should be ~13,000 to 14,000 or at the end of the read. The number of QV bases >=20 should be ~950 to 1000 (less for shorter PCR fragments) Figure 3: Annotation file which shows values for signal strength and start/end points Page 3 of 11
4 3 Troubleshooting When troubleshooting sequencing data, follow the workflow below to try to identify the cause of your problem. The following steps in this section use Sequencing Analysis Software or Sequence Scanner Software. 3.1 Reviewing the Sequence Electropherogram Select the Electropherogram tab and review the sequence for data quality. Check the following: Well-defined peak resolution minimal fluorescence overlap from one peak to the next with a sharp peak top. Uniform peak spacing peak spacing is consistent throughout the trace. Signal-to-noise ratios and variation in peak heights High signal to noise ratio and even peak height characterize good quality sequence. (Please note: the analysed view is re-scaled, the peak heights are not representative of raw fluorescence detected by the AB 3730xl.) Raw Sequence Select the Raw tab and review the unprocessed fluorescence data to assess the signal quality. Check the following: Artifacts Are there any artifacts, such as four-color spikes? Peak heights Are peaks well-resolved, with acceptable heights? Data start points Do any data start points deviate from others in the same submission? Length of read Was the expected length of read obtained? Does the signal stop suddenly? Baseline Is there background noise for all the peaks? Zoom in horizontally and vertically to verify the baseline noise. Page 4 of 11
5 3.2 Result Evaluation Use the following examples as a guide to try to identify an explanation of your results. Please note that this list is not exhaustive, but include the most common results seen at the AGRF Failed sequence No noticeable fluorescence peaks in raw data Only background noise seen in electropherogram Problem Probable Cause Solution No sequence detected No priming site present Confirm the primer site is present in the template. Redesign or use a different primer Primers have degraded through freeze-thaw cycles Make up new primer stocks Inefficient primer binding Redesign primer Insufficient amount of DNA template Re-quantify DNA and increase the amount of DNA if required DNA template has degraded or Inhibitory contaminant in your samples e.g. salts, phenol, EDTA, ethanol Re-extract DNA template or clean-up template. Page 5 of 11
6 3.2.2 Weak sequence Very low peak height in the raw data trace Base calls fade before the end of the read and the signal-to-noise ratios are very low Problem Probable Cause Solution Low peaks throughout trace Insufficient amount of DNA template Inhibitory contaminant in your samples (e.g. salts, phenol, EDTA, ethanol) Quantitate the DNA Increase the amount of DNA template, clean-up DNA template Insufficient amount of primer or inefficient primer binding Check primer dilution and/or redesign primer Page 6 of 11
7 3.2.3 Short sequence (or shorter than expected) Very high peaks in the raw data trace that fade off abruptly Poor quality sequence at start leading to shorter than expected sequence length Problem Probable Cause Solution Sequence starts well but signal drops gradually (Ski-sloping) Primer or Template ratio is incorrect or contaminant is present in template Re-examine template and primer concentration Re-extract or clean-up template Repetitive region - Repeat regions, especially GC and GT repeats, can cause the signal to fade either due to depletion or slippage or secondary structure Add (1ul) DMSO to the sequencing reaction Sequence complementary strand the Sequence starts well but signal stops abruptly Secondary structure - GC and AT rich templates can cause the DNA to loop and form hairpins Linearized DNA - restriction enzymes may have cut the template Add (1ul) DMSO to the sequencing reaction to help relax the structure Design primers close to the hairpin Run product out on an agarose gel to check Page 7 of 11
8 3.2.4 Multiple sequences Overlapping peaks in all or part of the electropherogram that maintain correct base-spacing Overlapping peaks after a homopolymer region Problem Probable Cause Solution Overlapping peaks in all or part of the sequence Overlapping peaks following stretch of mononucleotide sequence Mixed plasmid preparation Multiple PCR products Re-isolate the the DNA from a pure colony and re-sequence Check PCR template on gel for single band Frame shift mutation Use a different primer after the mutation or sequence the complementary strand Primer-dimer contamination Multiple priming sites Multiple primers in reaction Optimise PCR amplification or redesign primer Make sure primer only has one priming site Ensure only one primer has been used Primer with N-1 contamination Re-synthesize primer with PAGE purification Enzyme slippage occurs giving varying lengths of the same sequence after this region (n-1, n-2 and n-3 populations) Sequence the complementary strand Page 8 of 11
9 3.2.5 Artifacts Peaks of excess dye present in the raw data trace Large broadened peaks that obscure the sequence Problem Probable Cause Solution Large peaks obscuring the real sequence Sample peaks become lumpy and increasingly unreadable early in the sequence (before 500bp) Dye blobs caused by unincorporated BigDye Terminator (BDT) and are typically seen at 70bp and 120bp. Usually seen in failed or weak sequences. Real sequence can still be read underneath these blobs If related to individual samples this is due to a contaminant in the sample For CS submissions, review cleanup method used and/or add more DNA template and less BDT. For PD submissions, please notify AGRF staff and a free re-run will be provided. Clean up template DNA Page 9 of 11
10 4 Review AGRF submission Take some time to review the samples in each submission batch. For example, does the problem occur in: o Specific samples o Specific submissions o Samples extracted during the same process, or different processes o Samples stored under similar conditions or under different conditions etc. Is the symptom present in other samples of the same submission? Are there any differences in how the templates of other samples in the same submission were prepared? Was the same primer used in all samples of the submission? How was the template stored after preparation for this submission? 5 Reviewing Experimental Setup of SEQ Reaction Based on the results from Sequence Data Evaluation, use the tables below to review your experimental setup. 5.1 DNA Template Review Recommendation Run an agarose gel to detect any contaminating DNA or RNA. Measure the A260/A280 ratio of your samples. Quantitate the DNA template using the absorbance at 260 nm (A260). Dilute or concentrate the DNA as needed to obtain an A260 reading between 0.05 and Comment Purified DNA should run as a single band on an agarose gel. Note: Uncut plasmid DNA can run as three bands: supercoiled, nicked, and linear. Note: RNA contamination up to 1 μg can be tolerated in the sequencing reaction, but it affects DNA quantitation greatly. For pure preparations of DNA (in TE), the A260/A280* ratio is 1.8. Very clean samples in pure water can give a ratio of 1.5 to 1.6. Smaller ratios may indicate the presence of protein or organic contaminants. Ratios less than 1.8 may still produce high quality results. Quantitation by agarose gel electrophoresis may not be accurate because ethidium bromide incorporation is not consistent and the method of comparing the standard and sample brightness is subjective. A260 values below 0.05 or above 1.00 are not accurate because Beer s law generally applies only within a certain concentration range. Outside of this concentration range, the relationship between absorbance and concentration is nonlinear. *A260 and A280 are the optical spectrometer measurement of absorbance at the wavelengths of 260 nm and 280 nm respectively. A260 is frequently used to measure DNA/RNA concentration and A280 is used to measure protein concentration. A ratio of A260/A280 > 1.8 suggests little protein contamination in a DNA/RNA sample. Page 10 of 11
11 5.2 Primer Design Review Recommendation Ensure that the primer has Tm >45 C. Ensure that primers are at least 18 bases long. Ensure that there are no known secondary hybridization sites on the target DNA. Choose primers that do not have runs of identical nucleotides, especially 4 or more Gs. Choose primers with G-C content in the range of 30 to 80%, preferably 50 to 55%. Design primers to minimize the potential for secondary structure and/or hybridization Purify primers by HPLC to reduce the quantity of n-1 primers. Comment If the Tm is too low, it may result in poor priming and low or no signal Primers that are too short may have Tms that are too low. Secondary hybridization sites on the target DNA can result in double peaks throughout the sequence Runs of identical nucleotides in primers can cause n+1 or n-1 effects. Also, these primers may be more difficult to synthesize. If the G-C content is too low, the Tm may be too low. If so, increase the primer length beyond 18 bases to obtain a Tm>45 C. Primer-dimer formation from hybridization can result in mixed sequence at the beginning of the sequence. Secondary structure in the primer, particularly at the 3 end can result in poor priming and low or no signal. Primers containing contaminants or synthesized primers of the wrong length can cause problems in sequencing reactions, such as failed reactions, noisy data, or poor sequencing results. If the primer is a short oligo that contains n-1 primers, HPLC cannot always remove the n-1 contaminants. 6 Contact AGRF Sequencing If you have not resolved your problem, please contact AGRF Sequencing for further support. Contact Details sequencing@agrf.org.au Phone: Page 11 of 11
Sanger sequencing troubleshooting guide. GATC Biotech AG
 Sanger sequencing troubleshooting guide GATC Biotech AG April, 2017 Introduction All sequencing data generated at GATC Biotech is carefully analysed before it is delivered to the customer. In cases where
Sanger sequencing troubleshooting guide GATC Biotech AG April, 2017 Introduction All sequencing data generated at GATC Biotech is carefully analysed before it is delivered to the customer. In cases where
Diagnosis Sanger. Interpreting and Troubleshooting Chromatograms. Volume 1: Help! No Data! GENEWIZ Technical Support
 Diagnosis Sanger Interpreting and Troubleshooting Chromatograms GENEWIZ Technical Support DNAseq@genewiz.com Troubleshooting This troubleshooting guide is based on common issues seen from samples within
Diagnosis Sanger Interpreting and Troubleshooting Chromatograms GENEWIZ Technical Support DNAseq@genewiz.com Troubleshooting This troubleshooting guide is based on common issues seen from samples within
Quick Reference Guide
 BrilliantDye Terminator v3.1 Cycle Sequencing Kit Quick Reference Guide Version: 3.0 Revision date: 23-01-2018 Product and Company Information Product name: BrilliantDye v3.1 Terminator Cycle Sequencing
BrilliantDye Terminator v3.1 Cycle Sequencing Kit Quick Reference Guide Version: 3.0 Revision date: 23-01-2018 Product and Company Information Product name: BrilliantDye v3.1 Terminator Cycle Sequencing
Tips DNA Sequencing (Revision Date: 09 July 2007)
 Tips DNA Sequencing (Revision Date: 09 July 2007) 1. PCR product: clone, or directly sequence? 2. How do I clean DNA template for sequencing? 3. How much DNA should I use in a sequencing reaction? 4. Why
Tips DNA Sequencing (Revision Date: 09 July 2007) 1. PCR product: clone, or directly sequence? 2. How do I clean DNA template for sequencing? 3. How much DNA should I use in a sequencing reaction? 4. Why
2. Pyrosequencing Assay Design
 2. Pyrosequencing Assay Design 2.1 Guidelines for PCR set-up and primer design 2.1.1 PCR primer design Design of PCR primers follows standard rules, i.e. calculated Tm of 62-65 C, primer length of about
2. Pyrosequencing Assay Design 2.1 Guidelines for PCR set-up and primer design 2.1.1 PCR primer design Design of PCR primers follows standard rules, i.e. calculated Tm of 62-65 C, primer length of about
Tips for sequencing DNA
 Sequencing tips and trouble shooting from website http://www.nucleics.com/dna_sequencing_support/dna-sequencing-protocol-tips.html Tips for sequencing DNA Use clean DNA The cleanliness of the DNA is the
Sequencing tips and trouble shooting from website http://www.nucleics.com/dna_sequencing_support/dna-sequencing-protocol-tips.html Tips for sequencing DNA Use clean DNA The cleanliness of the DNA is the
GENOMICS RESEARCH LABORATORY Sample Submission Requirements
 Sanger Sequencing GENOMICS RESEARCH LABORATORY Sample Submission Requirements The quality of the starting template is the key factor in the quality of DNA sequence data. Sequencing by capillary electrophoresis
Sanger Sequencing GENOMICS RESEARCH LABORATORY Sample Submission Requirements The quality of the starting template is the key factor in the quality of DNA sequence data. Sequencing by capillary electrophoresis
The Agilent Total RNA Isolation Kit
 Better RNA purity. Better data. Without DNase treatment. The Agilent Total RNA Isolation Kit Now you can isolate highly purified, intact RNA without DNase treatment Introducing the Agilent Total RNA Isolation
Better RNA purity. Better data. Without DNase treatment. The Agilent Total RNA Isolation Kit Now you can isolate highly purified, intact RNA without DNase treatment Introducing the Agilent Total RNA Isolation
7. Troubleshooting Guidelines
 7. Troubleshooting Guidelines The Pyrosequencing TM technique provides the user with sequence information for several nucleotides 3 of the sequencing primer apart from the polymorphic site. This gives
7. Troubleshooting Guidelines The Pyrosequencing TM technique provides the user with sequence information for several nucleotides 3 of the sequencing primer apart from the polymorphic site. This gives
FastFire qpcr PreMix (Probe)
 FastFire qpcr PreMix (Probe) For fast, quantitative, specific real-time PCR using sequence-specific probe www.tiangen.com/en FP130820 Kit Contents FastFire qpcr PreMix (Probe) Contents 2 FastFire qpcr
FastFire qpcr PreMix (Probe) For fast, quantitative, specific real-time PCR using sequence-specific probe www.tiangen.com/en FP130820 Kit Contents FastFire qpcr PreMix (Probe) Contents 2 FastFire qpcr
Comparison of ExoSAP-IT and ExoSAP-IT Express reagents to alternative PCR cleanup methods
 WHITE PAPER ExoSAP-IT PCR cleanup reagents Comparison of ExoSAP-IT and ExoSAP-IT Express reagents to alternative PCR cleanup methods Abstract Here we present superior workflow advantages of enzymatic PCR
WHITE PAPER ExoSAP-IT PCR cleanup reagents Comparison of ExoSAP-IT and ExoSAP-IT Express reagents to alternative PCR cleanup methods Abstract Here we present superior workflow advantages of enzymatic PCR
Interpretation of sequence results
 Interpretation of sequence results An overview on DNA sequencing: DNA sequencing involves the determination of the sequence of nucleotides in a sample of DNA. It use a modified PCR reaction where both
Interpretation of sequence results An overview on DNA sequencing: DNA sequencing involves the determination of the sequence of nucleotides in a sample of DNA. It use a modified PCR reaction where both
Guide to Sanger Sequencing at RAMAC
 Guide to Sanger Sequencing at RAMAC CONTENTS 1 Introduction... 3 2 Options for your sequencing needs... 3 3 What is Sanger sequencing?... 4 4 Template preparation... 5 4.1 Plasmid template... 5 4.2 PCR
Guide to Sanger Sequencing at RAMAC CONTENTS 1 Introduction... 3 2 Options for your sequencing needs... 3 3 What is Sanger sequencing?... 4 4 Template preparation... 5 4.1 Plasmid template... 5 4.2 PCR
GenBuilder TM Plus Cloning Kit User Manual
 GenBuilder TM Plus Cloning Kit User Manual Cat. No. L00744 Version 11242017 Ⅰ. Introduction... 2 I.1 Product Information... 2 I.2 Kit Contents and Storage... 2 I.3 GenBuilder Cloning Kit Workflow... 2
GenBuilder TM Plus Cloning Kit User Manual Cat. No. L00744 Version 11242017 Ⅰ. Introduction... 2 I.1 Product Information... 2 I.2 Kit Contents and Storage... 2 I.3 GenBuilder Cloning Kit Workflow... 2
GenBuilder TM Plus Cloning Kit User Manual
 GenBuilder TM Plus Cloning Kit User Manual Cat.no L00744 Version 11242017 Ⅰ. Introduction... 2 I.1 Product Information... 2 I.2 Kit Contents and Storage... 2 I.3 GenBuilder Cloning Kit Workflow... 2 Ⅱ.
GenBuilder TM Plus Cloning Kit User Manual Cat.no L00744 Version 11242017 Ⅰ. Introduction... 2 I.1 Product Information... 2 I.2 Kit Contents and Storage... 2 I.3 GenBuilder Cloning Kit Workflow... 2 Ⅱ.
ZR-96 Genomic DNA Clean & Concentrator -5 Catalog Nos. D4066 & D4067
 INSTRUCTION MANUAL ZR-96 Genomic DNA Clean & Concentrator -5 Catalog Nos. D4066 & D4067 Highlights 96-well plate recovery of large-sized DNA (e.g., genomic, mitochondrial, plasmid (BAC/PAC), viral, phage,
INSTRUCTION MANUAL ZR-96 Genomic DNA Clean & Concentrator -5 Catalog Nos. D4066 & D4067 Highlights 96-well plate recovery of large-sized DNA (e.g., genomic, mitochondrial, plasmid (BAC/PAC), viral, phage,
Roche Molecular Biochemicals Technical Note No. LC 9/2000
 Roche Molecular Biochemicals Technical Note No. LC 9/2000 LightCycler Optimization Strategy Introduction Purpose of this Note Table of Contents The LightCycler system provides different detection formats
Roche Molecular Biochemicals Technical Note No. LC 9/2000 LightCycler Optimization Strategy Introduction Purpose of this Note Table of Contents The LightCycler system provides different detection formats
Genomic DNA Clean & Concentrator -25 Catalog Nos. D4064 & D4065
 INSTRUCTION MANUAL Genomic DNA Clean & Concentrator -25 Catalog Nos. D4064 & D4065 Highlights Quick (5 minute) spin column recovery of large-sized DNA (e.g., genomic, mitochondrial, plasmid (BAC/PAC),
INSTRUCTION MANUAL Genomic DNA Clean & Concentrator -25 Catalog Nos. D4064 & D4065 Highlights Quick (5 minute) spin column recovery of large-sized DNA (e.g., genomic, mitochondrial, plasmid (BAC/PAC),
GenBuilder TM Cloning Kit User Manual
 GenBuilder TM Cloning Kit User Manual Cat.no L00701 Version 11242017 Ⅰ. Introduction... 2 I.1 Product Information... 2 I.2 Kit Contents and Storage... 2 I.3 GenBuilder Cloning Kit Workflow... 2 Ⅱ. DNA
GenBuilder TM Cloning Kit User Manual Cat.no L00701 Version 11242017 Ⅰ. Introduction... 2 I.1 Product Information... 2 I.2 Kit Contents and Storage... 2 I.3 GenBuilder Cloning Kit Workflow... 2 Ⅱ. DNA
User Bulletin. ABI PRISM 373 DNA Sequencer. Using the ABI PRISM 373 BigDye Filter Wheel. Contents. Introduction. February 16, 1998 (updated 06/2001)
 User Bulletin ABI PRISM 373 DNA Sequencer February 16, 1998 (updated 06/2001) SUBJECT: Using the ABI PRISM 373 BigDye Filter Wheel Contents The following topics are discussed in this bulletin: Topic See
User Bulletin ABI PRISM 373 DNA Sequencer February 16, 1998 (updated 06/2001) SUBJECT: Using the ABI PRISM 373 BigDye Filter Wheel Contents The following topics are discussed in this bulletin: Topic See
1
 1 2 3 4 5 Cosmids are plasmid vectors that contain cos sites. The cos site is the only requirement for DNA to be packaged into a phage particle 6 7 8 9 10 11 12 13 14 15 16 For de novo sequencing using
1 2 3 4 5 Cosmids are plasmid vectors that contain cos sites. The cos site is the only requirement for DNA to be packaged into a phage particle 6 7 8 9 10 11 12 13 14 15 16 For de novo sequencing using
Guidelines for Developing Robust and Reliable PCR Assays
 Guidelines for Developing Robust and Reliable PCR Assays Leta Steffen, PhD Applications Scientist Promega Corporation Outline 1) PCR reaction components What is in the reaction? How does it affect assay
Guidelines for Developing Robust and Reliable PCR Assays Leta Steffen, PhD Applications Scientist Promega Corporation Outline 1) PCR reaction components What is in the reaction? How does it affect assay
Cold Fusion Cloning Kit. Cat. #s MC100A-1, MC101A-1. User Manual
 Fusion Cloning technology Cold Fusion Cloning Kit Store the master mixture and positive controls at -20 C Store the competent cells at -80 C. (ver. 120909) A limited-use label license covers this product.
Fusion Cloning technology Cold Fusion Cloning Kit Store the master mixture and positive controls at -20 C Store the competent cells at -80 C. (ver. 120909) A limited-use label license covers this product.
Guide to Sanger Sequencing at RAMAC
 Guide to Sanger Sequencing at RAMAC CONTENTS 1 Introduction... 3 2 Options for your sequencing needs... 3 3 What is Sanger sequencing?... 4 4 Template preparation... 5 4.1 Plasmid template... 5 4.2 PCR
Guide to Sanger Sequencing at RAMAC CONTENTS 1 Introduction... 3 2 Options for your sequencing needs... 3 3 What is Sanger sequencing?... 4 4 Template preparation... 5 4.1 Plasmid template... 5 4.2 PCR
1. COMPONENTS. PyroStart Fast PCR Master Mix (2X) (#K0211 for 250 reactions of 20µl) 2. STORAGE 3. DESCRIPTION
 1 2 1 PyroStart Fast PCR Master Mix (2X) (#K0211 for 250 reactions of 20µl) TABLE OF CONTENTS 1. COMPONENTS... 2 2. STORAGE... 2 3. DESCRIPTION... 2 4. PROTOCOL FOR FAST PCR... 3 4.1. General Considerations...
1 2 1 PyroStart Fast PCR Master Mix (2X) (#K0211 for 250 reactions of 20µl) TABLE OF CONTENTS 1. COMPONENTS... 2 2. STORAGE... 2 3. DESCRIPTION... 2 4. PROTOCOL FOR FAST PCR... 3 4.1. General Considerations...
Sanger Sequencing Portfolio
 Molecular Cloning Laboratories Sanger Sequencing Portfolio Significant Cost Savings vs ABI Uncompromised Performance Longer Read Length Longer Shelf Life No Recalibrations Necessary PAGE 1 BrightDye Terminator
Molecular Cloning Laboratories Sanger Sequencing Portfolio Significant Cost Savings vs ABI Uncompromised Performance Longer Read Length Longer Shelf Life No Recalibrations Necessary PAGE 1 BrightDye Terminator
GeneCopoeia TM. All-in-One qpcr Mix For universal quantitative real-time PCR. User Manual
 GeneCopoeia TM Expressway to Discovery All-in-One qpcr Mix For universal quantitative real-time PCR Cat. No. AOPR-0200 (200 qpcr reactions) Cat. No. AOPR-0600 (600 qpcr reactions) Cat. No. AOPR-1000 (1000
GeneCopoeia TM Expressway to Discovery All-in-One qpcr Mix For universal quantitative real-time PCR Cat. No. AOPR-0200 (200 qpcr reactions) Cat. No. AOPR-0600 (600 qpcr reactions) Cat. No. AOPR-1000 (1000
Genomic Sequencing. Genomic Sequencing. Maj Gen (R) Suhaib Ahmed, HI (M)
 Maj Gen (R) Suhaib Ahmed, HI (M) The process of determining the sequence of an unknown DNA is called sequencing. There are many approaches for DNA sequencing. In the last couple of decades automated Sanger
Maj Gen (R) Suhaib Ahmed, HI (M) The process of determining the sequence of an unknown DNA is called sequencing. There are many approaches for DNA sequencing. In the last couple of decades automated Sanger
Efficient Method for Isolation of High Quality Concentrated Cellular RNA with Extremely Low Levels of Genomic DNA Contamination Application
 Efficient Method for Isolation of High Quality Concentrated Cellular RNA with Extremely Low Levels of Genomic DNA Contamination Application Gene Expression Authors Ilgar Abbaszade, Claudia Robbins, John
Efficient Method for Isolation of High Quality Concentrated Cellular RNA with Extremely Low Levels of Genomic DNA Contamination Application Gene Expression Authors Ilgar Abbaszade, Claudia Robbins, John
 PRODUCT INFORMATION Long PCR Enzyme Mix #K0182 500 u Lot Exp. 00.0000 Store at -20 C. CERTIFICATE OF ANALYSIS Long PCR Enzyme Mix is functionally tested in PCR amplification of 47.4 kb fragment from lambda
PRODUCT INFORMATION Long PCR Enzyme Mix #K0182 500 u Lot Exp. 00.0000 Store at -20 C. CERTIFICATE OF ANALYSIS Long PCR Enzyme Mix is functionally tested in PCR amplification of 47.4 kb fragment from lambda
QUANTITATIVE RT-PCR PROTOCOL (SYBR Green I) (Last Revised: April, 2007)
 QUANTITATIVE RT-PCR PROTOCOL (SYBR Green I) (Last Revised: April, 007) Please contact Center for Plant Genomics (CPG) facility manager Hailing Jin (hljin@iastate.edu) regarding questions or corrections.
QUANTITATIVE RT-PCR PROTOCOL (SYBR Green I) (Last Revised: April, 007) Please contact Center for Plant Genomics (CPG) facility manager Hailing Jin (hljin@iastate.edu) regarding questions or corrections.
THUNDERBIRD SYBR qpcr Mix
 Instruction manual THUNDERBIRD SYBR qpcr Mix 1304 A4251K THUNDERBIRD SYBR qpcr Mix QPS-201T 1 ml x 1 QPS-201 1.67 ml x 3 Contents [1] Introduction [2] Components [3] Primer design [4] Template DNA [5]
Instruction manual THUNDERBIRD SYBR qpcr Mix 1304 A4251K THUNDERBIRD SYBR qpcr Mix QPS-201T 1 ml x 1 QPS-201 1.67 ml x 3 Contents [1] Introduction [2] Components [3] Primer design [4] Template DNA [5]
BENG 183 Trey Ideker (the details )
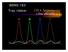 BENG 183 Trey Ideker (the details ) (1) Devils in the details: Sequencing topics to be covered in today s lecture DNA preparation prior to sequencing Amplification: vectors or cycle sequencing PAGE and
BENG 183 Trey Ideker (the details ) (1) Devils in the details: Sequencing topics to be covered in today s lecture DNA preparation prior to sequencing Amplification: vectors or cycle sequencing PAGE and
Bead Type (NaI) Gel Extraction Kits
 Bead Type (NaI) Contents Kit Contents Principle Important Notes Bead Type (NaI) Gel Extraction Kit Protocol Troubleshooting Guide 2 3 3 4 5 Ordering Information 6 Kit Contents Catalog No. Number of preparations
Bead Type (NaI) Contents Kit Contents Principle Important Notes Bead Type (NaI) Gel Extraction Kit Protocol Troubleshooting Guide 2 3 3 4 5 Ordering Information 6 Kit Contents Catalog No. Number of preparations
Design. Construction. Characterization
 Design Construction Characterization DNA mrna (messenger) A C C transcription translation C A C protein His A T G C T A C G Plasmids replicon copy number incompatibility selection marker origin of replication
Design Construction Characterization DNA mrna (messenger) A C C transcription translation C A C protein His A T G C T A C G Plasmids replicon copy number incompatibility selection marker origin of replication
Real Time Quantitative PCR Assay Validation, Optimization and Troubleshooting
 Real Time Quantitative PCR Assay Validation, Optimization and Troubleshooting Dr Steffen Muller Field Applications Scientist Stratagene Europe Overview The Importance of Controls Validation and Optimization
Real Time Quantitative PCR Assay Validation, Optimization and Troubleshooting Dr Steffen Muller Field Applications Scientist Stratagene Europe Overview The Importance of Controls Validation and Optimization
GenepHlow Gel Extraction Kit
 Instruction Manual Ver. 02.10.17 For Research Use Only GenepHlow Gel Extraction Kit DFG004 (4 Preparation Sample Kit) DFG100 (100 Preparation Kit) DFG300 (300 Preparation Kit) Advantages Convenient: includes
Instruction Manual Ver. 02.10.17 For Research Use Only GenepHlow Gel Extraction Kit DFG004 (4 Preparation Sample Kit) DFG100 (100 Preparation Kit) DFG300 (300 Preparation Kit) Advantages Convenient: includes
Assay Design Considerations, Optimization and Validation
 Assay Design Considerations, Optimization and Validation Ray Meng, Ph.D. International Field Applications Specialist Gene Expression Division Bio-Rad Laboratories, Inc. Assay Design Considerations Experiment
Assay Design Considerations, Optimization and Validation Ray Meng, Ph.D. International Field Applications Specialist Gene Expression Division Bio-Rad Laboratories, Inc. Assay Design Considerations Experiment
Zymo-Spin Technology. The microcentrifuge column that revolutionized nucleic acid clean-up
 Zymo-Spin Technology The microcentrifuge column that revolutionized nucleic acid clean-up Overview A Comprehensive Solution for DNA and RNA Clean-Up High-quality, inhibitor-free DNA and RNA is crucial
Zymo-Spin Technology The microcentrifuge column that revolutionized nucleic acid clean-up Overview A Comprehensive Solution for DNA and RNA Clean-Up High-quality, inhibitor-free DNA and RNA is crucial
Polymerase Chain Reaction-361 BCH
 Polymerase Chain Reaction-361 BCH 1-Polymerase Chain Reaction Nucleic acid amplification is an important process in biotechnology and molecular biology and has been widely used in research, medicine, agriculture
Polymerase Chain Reaction-361 BCH 1-Polymerase Chain Reaction Nucleic acid amplification is an important process in biotechnology and molecular biology and has been widely used in research, medicine, agriculture
Seminar: Sanger Sequencing
 Seminar: Sanger Sequencing 04. August 2010 Dr. René Gödde Produktspezialist Laborautomation Seminar: Sanger Sequencing Beckman Coulter (Genomics) Agencourt CleanSEQ kit Requirements for a good sequencing
Seminar: Sanger Sequencing 04. August 2010 Dr. René Gödde Produktspezialist Laborautomation Seminar: Sanger Sequencing Beckman Coulter (Genomics) Agencourt CleanSEQ kit Requirements for a good sequencing
Presto Stool DNA Extraction Kit
 Instruction Manual Ver. 10.21.17 For Research Use Only Presto Stool DNA Extraction Kit Advantages STLD004 (4 Preparation Sample Kit) STLD050 (50 Preparation Kit) STLD100 (100 Preparation Kit) Sample: 180-200
Instruction Manual Ver. 10.21.17 For Research Use Only Presto Stool DNA Extraction Kit Advantages STLD004 (4 Preparation Sample Kit) STLD050 (50 Preparation Kit) STLD100 (100 Preparation Kit) Sample: 180-200
Troubleshooting of Real Time PCR Ameer Effat M. Elfarash
 Troubleshooting of Real Time PCR Ameer Effat M. Elfarash Dept. of Genetics Fac. of Agriculture, Assiut Univ. aelfarash@aun.edu.eg What is Real-Time PCR used for? Gene expression analysis Disease diagnosis
Troubleshooting of Real Time PCR Ameer Effat M. Elfarash Dept. of Genetics Fac. of Agriculture, Assiut Univ. aelfarash@aun.edu.eg What is Real-Time PCR used for? Gene expression analysis Disease diagnosis
Description...1 Components...1 Storage... 1 Technical Information...1 Protocol...2 Examples using the kit...4 Troubleshooting...
 QuickClean II Gel Extraction Kit Cat. No. L00418 Technical Manual No. TM0594 Version: 03042011 I II III IV V VI VII VIII Description......1 Components.....1 Storage.... 1 Technical Information....1 Protocol.....2
QuickClean II Gel Extraction Kit Cat. No. L00418 Technical Manual No. TM0594 Version: 03042011 I II III IV V VI VII VIII Description......1 Components.....1 Storage.... 1 Technical Information....1 Protocol.....2
BIOO LIFE SCIENCE PRODUCTS
 BIOO LIFE SCIENCE PRODUCTS FOR REFERENCE PURPOSES This manual is for Reference Purposes Only. DO NOT use this protocol to run your assays. Periodically, optimizations and revisions are made to the kit
BIOO LIFE SCIENCE PRODUCTS FOR REFERENCE PURPOSES This manual is for Reference Purposes Only. DO NOT use this protocol to run your assays. Periodically, optimizations and revisions are made to the kit
SEQUENCING SERVICES. McGill University and Génome Québec Innovation Centre FEBRUARY 28, Version 3.5
 FEBRUARY 28, 2018 SEQUENCING SERVICES McGill University and Génome Québec Innovation Centre Sanger Sequencing User Guide Version 3.5 Copyright 2010 McGill University and Génome Québec Innovation Centre
FEBRUARY 28, 2018 SEQUENCING SERVICES McGill University and Génome Québec Innovation Centre Sanger Sequencing User Guide Version 3.5 Copyright 2010 McGill University and Génome Québec Innovation Centre
Executive Summary. clinical supply services
 clinical supply services case study Development and NDA-level validation of quantitative polymerase chain reaction (qpcr) procedure for detection and quantification of residual E.coli genomic DNA Executive
clinical supply services case study Development and NDA-level validation of quantitative polymerase chain reaction (qpcr) procedure for detection and quantification of residual E.coli genomic DNA Executive
Genetics and Genomics in Medicine Chapter 3. Questions & Answers
 Genetics and Genomics in Medicine Chapter 3 Multiple Choice Questions Questions & Answers Question 3.1 Which of the following statements, if any, is false? a) Amplifying DNA means making many identical
Genetics and Genomics in Medicine Chapter 3 Multiple Choice Questions Questions & Answers Question 3.1 Which of the following statements, if any, is false? a) Amplifying DNA means making many identical
Presto Food DNA Extraction Kit
 Instruction Manual Ver. 06.28.17 For Research Use Only Presto Food DNA Extraction Kit Advantages FGD004 (4 Preparation Sample Kit) FGD100 (100 Preparation Kit) FGD300 (300 Preparation Kit) Sample: 200
Instruction Manual Ver. 06.28.17 For Research Use Only Presto Food DNA Extraction Kit Advantages FGD004 (4 Preparation Sample Kit) FGD100 (100 Preparation Kit) FGD300 (300 Preparation Kit) Sample: 200
Roche Molecular Biochemicals Technical Note No. LC 10/2000
 Roche Molecular Biochemicals Technical Note No. LC 10/2000 LightCycler Overview of LightCycler Quantification Methods 1. General Introduction Introduction Content Definitions This Technical Note will introduce
Roche Molecular Biochemicals Technical Note No. LC 10/2000 LightCycler Overview of LightCycler Quantification Methods 1. General Introduction Introduction Content Definitions This Technical Note will introduce
qpcr Quantitative PCR or Real-time PCR Gives a measurement of PCR product at end of each cycle real time
 qpcr qpcr Quantitative PCR or Real-time PCR Gives a measurement of PCR product at end of each cycle real time Differs from endpoint PCR gel on last cycle Used to determines relative amount of template
qpcr qpcr Quantitative PCR or Real-time PCR Gives a measurement of PCR product at end of each cycle real time Differs from endpoint PCR gel on last cycle Used to determines relative amount of template
Functional Genomics Research Stream. Research Meeting: June 19, 2012 SYBR Green qpcr, Research Update
 Functional Genomics Research Stream Research Meeting: June 19, 2012 SYBR Green qpcr, Research Update Updates Alternate Lab Meeting Fridays 11:30-1:00 WEL 4.224 Welcome to attend either one Lab Log thanks
Functional Genomics Research Stream Research Meeting: June 19, 2012 SYBR Green qpcr, Research Update Updates Alternate Lab Meeting Fridays 11:30-1:00 WEL 4.224 Welcome to attend either one Lab Log thanks
SunScript TM One Step RT-qPCR Kit
 INDEX Ordering Information...3 Kit Contents...3 Shipping and Storage...3 Handling...3 Quality Control...3 Reagents and Equipment to be Supplied by the User...3 Description...4 Protocol...4 Troubleshooting
INDEX Ordering Information...3 Kit Contents...3 Shipping and Storage...3 Handling...3 Quality Control...3 Reagents and Equipment to be Supplied by the User...3 Description...4 Protocol...4 Troubleshooting
Accura High-Fidelity Polymerase
 Accura High-Fidelity Polymerase FOR RESEARCH USE ONLY. NOT FOR HUMAN OR DIAGNOSTIC USE Lucigen Corporation 2905 Parmenter St, Middleton, WI 53562 USA Toll Free: (888) 575-9695 (608) 831-9011 FAX: (608)
Accura High-Fidelity Polymerase FOR RESEARCH USE ONLY. NOT FOR HUMAN OR DIAGNOSTIC USE Lucigen Corporation 2905 Parmenter St, Middleton, WI 53562 USA Toll Free: (888) 575-9695 (608) 831-9011 FAX: (608)
SANGER SEQUENCING WHITE PAPER
 AAAT T C SANGER SEQUENCING WHITE PAPER FACTORS AFFECTING SANGER SEQUENCING ROBUSTNESS AND DATA QUALITY PATRICK WARNER, SHEA ANDERSON, ARCHANA DESHPANDE, DARYL M. GOHL, KENNETH B. BECKMAN UNIVERSITY OF
AAAT T C SANGER SEQUENCING WHITE PAPER FACTORS AFFECTING SANGER SEQUENCING ROBUSTNESS AND DATA QUALITY PATRICK WARNER, SHEA ANDERSON, ARCHANA DESHPANDE, DARYL M. GOHL, KENNETH B. BECKMAN UNIVERSITY OF
Roche Molecular Biochemicals Technical Note No. LC 12/2000
 Roche Molecular Biochemicals Technical Note No. LC 12/2000 LightCycler Absolute Quantification with External Standards and an Internal Control 1. General Introduction Purpose of this Note Overview of Method
Roche Molecular Biochemicals Technical Note No. LC 12/2000 LightCycler Absolute Quantification with External Standards and an Internal Control 1. General Introduction Purpose of this Note Overview of Method
Presto Soil DNA Extraction Kit
 Instruction Manual Ver. 02.23.17 For Research Use Only Presto Soil DNA Extraction Kit Advantages SLD004 (4 Preparation Sample Kit) SLD050 (50 Preparation Kit) SLD100 (100 Preparation Kit) Sample: 250-500
Instruction Manual Ver. 02.23.17 For Research Use Only Presto Soil DNA Extraction Kit Advantages SLD004 (4 Preparation Sample Kit) SLD050 (50 Preparation Kit) SLD100 (100 Preparation Kit) Sample: 250-500
Quick and easy 7 minute protocol to select for 300 bp, 200 bp, 150 bp, 100 bp, 50 bp DNA fragments or perform a double size selection
 INSTRUCTION MANUAL Select-a-Size DNA Clean & Concentrator Catalog No. D4080 Highlights Quick and easy 7 minute protocol to select for 300 bp, 200 bp, 150 bp, 100 bp, 50 bp DNA fragments or perform a double
INSTRUCTION MANUAL Select-a-Size DNA Clean & Concentrator Catalog No. D4080 Highlights Quick and easy 7 minute protocol to select for 300 bp, 200 bp, 150 bp, 100 bp, 50 bp DNA fragments or perform a double
Quick and easy 7 minute protocol to select for 300 bp, 200 bp, 150 bp, 100 bp, 50 bp DNA fragments or perform a double size selection
 INSTRUCTION MANUAL Select-a-Size DNA Clean & Concentrator Catalog No. D4080 Highlights Quick and easy 7 minute protocol to select for 300 bp, 200 bp, 150 bp, 100 bp, 50 bp DNA fragments or perform a double
INSTRUCTION MANUAL Select-a-Size DNA Clean & Concentrator Catalog No. D4080 Highlights Quick and easy 7 minute protocol to select for 300 bp, 200 bp, 150 bp, 100 bp, 50 bp DNA fragments or perform a double
RTS Wheat Germ LinTempGenSet, His 6 -tag Manual
 RTS Wheat Germ LinTempGenSet, His 6 -tag Manual For rapid production of linear expression templates using PCR RTS Wheat Germ LinTempGenSet, His6-tag, April, 2015 2015 biotechrabbit, all rights reserved.
RTS Wheat Germ LinTempGenSet, His 6 -tag Manual For rapid production of linear expression templates using PCR RTS Wheat Germ LinTempGenSet, His6-tag, April, 2015 2015 biotechrabbit, all rights reserved.
PureSpin DNA Clean Up Kit
 PureSpin DNA Clean Up Kit Micro Columns INSTRUCTION MANUAL KIT COMPONENTS For Research Use Only PureSpin DNA Clean Up Kit, Micro Columns w/out Caps (Kit Size) OD2080 (50 Preps.) OD2080-2 (200 Preps.) Storage
PureSpin DNA Clean Up Kit Micro Columns INSTRUCTION MANUAL KIT COMPONENTS For Research Use Only PureSpin DNA Clean Up Kit, Micro Columns w/out Caps (Kit Size) OD2080 (50 Preps.) OD2080-2 (200 Preps.) Storage
Polymerase Chain Reaction (PCR)
 Laboratory for Environmental Pathogens Research Department of Environmental Sciences University of Toledo Polymerase Chain Reaction (PCR) Background information The polymerase chain reaction (PCR) is an
Laboratory for Environmental Pathogens Research Department of Environmental Sciences University of Toledo Polymerase Chain Reaction (PCR) Background information The polymerase chain reaction (PCR) is an
Lecture Four. Molecular Approaches I: Nucleic Acids
 Lecture Four. Molecular Approaches I: Nucleic Acids I. Recombinant DNA and Gene Cloning Recombinant DNA is DNA that has been created artificially. DNA from two or more sources is incorporated into a single
Lecture Four. Molecular Approaches I: Nucleic Acids I. Recombinant DNA and Gene Cloning Recombinant DNA is DNA that has been created artificially. DNA from two or more sources is incorporated into a single
P HENIX. PHENIX PCR Enzyme Guide Tools For Life Science Discovery RESEARCH PRODUCTS
 PHENIX PCR Enzyme Guide PHENIX offers a broad line of premium quality PCR Enzymes. This PCR Enzyme Guide will help simplify your polymerase selection process. Each DNA Polymerase has different characteristics
PHENIX PCR Enzyme Guide PHENIX offers a broad line of premium quality PCR Enzymes. This PCR Enzyme Guide will help simplify your polymerase selection process. Each DNA Polymerase has different characteristics
HiPer Real-Time PCR Teaching Kit
 HiPer Real-Time PCR Teaching Kit Product Code: HTBM032 Number of experiments that can be performed: 10 Duration of Experiment Protocol: 1.5 hours Storage Instructions: The kit is stable for 12 months from
HiPer Real-Time PCR Teaching Kit Product Code: HTBM032 Number of experiments that can be performed: 10 Duration of Experiment Protocol: 1.5 hours Storage Instructions: The kit is stable for 12 months from
Dig System for Starters
 Dig System for Starters Content 1. Powerful and Versatile DIG System 2. Labeling Nucleic Acids using the DIG System 3. Critical Hints for PCR Labeling 1 2 3 1. Powerful and Versatile DIG System Powerful
Dig System for Starters Content 1. Powerful and Versatile DIG System 2. Labeling Nucleic Acids using the DIG System 3. Critical Hints for PCR Labeling 1 2 3 1. Powerful and Versatile DIG System Powerful
Bootcamp: Molecular Biology Techniques and Interpretation
 Bootcamp: Molecular Biology Techniques and Interpretation Bi8 Winter 2016 Today s outline Detecting and quantifying nucleic acids and proteins: Basic nucleic acid properties Hybridization PCR and Designing
Bootcamp: Molecular Biology Techniques and Interpretation Bi8 Winter 2016 Today s outline Detecting and quantifying nucleic acids and proteins: Basic nucleic acid properties Hybridization PCR and Designing
Optimizing crna fragmentation for microarray experiments using the Agilent 2100 bioanalyzer. Application Deborah Vitale.
 Optimizing crna fragmentation for microarray experiments using the Agilent 2100 bioanalyzer Application Deborah Vitale Introduction Oligonucleotide arrays are a powerful tool for gene expression studies.
Optimizing crna fragmentation for microarray experiments using the Agilent 2100 bioanalyzer Application Deborah Vitale Introduction Oligonucleotide arrays are a powerful tool for gene expression studies.
Presto Mini Plasmid Kit
 Instruction Manual Ver. 03.06.17 For Research Use Only Presto Mini Plasmid Kit PDH004 (4 Preparation Sample Kit) PDH100 (100 Preparation Kit) PDH300 (300 Preparation Kit) Advantages Sample: 1-7 ml of cultured
Instruction Manual Ver. 03.06.17 For Research Use Only Presto Mini Plasmid Kit PDH004 (4 Preparation Sample Kit) PDH100 (100 Preparation Kit) PDH300 (300 Preparation Kit) Advantages Sample: 1-7 ml of cultured
TB Green Premix Ex Taq II (Tli RNaseH Plus)
 Cat. # RR820A For Research Use TB Green Premix Ex Taq II (Tli RNaseH Plus) Product Manual The long-term storage temperature of this product has been changed to -20 since Lot. #AGY1013N. See section IV.
Cat. # RR820A For Research Use TB Green Premix Ex Taq II (Tli RNaseH Plus) Product Manual The long-term storage temperature of this product has been changed to -20 since Lot. #AGY1013N. See section IV.
Certificate of Analysis. Quality Control Parameters. 2 -deoxyadenosine-5 -triphosphate. MW = g /mol
 COA No: CA BN-0006 datp 100mM Storage Conditions: Lot number: -20 C DA-516110 For Research Use Only Expiry date: November 2018 Quality Control Parameters 2 -deoxyadenosine-5 -triphosphate C10H12N5O12P3Li4
COA No: CA BN-0006 datp 100mM Storage Conditions: Lot number: -20 C DA-516110 For Research Use Only Expiry date: November 2018 Quality Control Parameters 2 -deoxyadenosine-5 -triphosphate C10H12N5O12P3Li4
SYBR Premix Ex Taq II (Tli RNaseH Plus), ROX plus
 Cat. # RR82LR For Research Use SYBR Premix Ex Taq II (Tli RNaseH Plus), ROX plus Product Manual The long-term storage temperature of this product has been changed to -20 since Lot. A1901A. See section
Cat. # RR82LR For Research Use SYBR Premix Ex Taq II (Tli RNaseH Plus), ROX plus Product Manual The long-term storage temperature of this product has been changed to -20 since Lot. A1901A. See section
RTS 100 E. coli LinTempGenSet, Histag
 RTS 100 E. coli LinTempGenSet, Histag Manual For rapid production of linear expression templates using PCR RTS 100 E. coli LinTempGenSet, His-tag Manual, October, 2009 2009 5 PRIME, all rights reserved.
RTS 100 E. coli LinTempGenSet, Histag Manual For rapid production of linear expression templates using PCR RTS 100 E. coli LinTempGenSet, His-tag Manual, October, 2009 2009 5 PRIME, all rights reserved.
Performance characteristics of the High Sensitivity DNA kit for the Agilent 2100 Bioanalyzer
 Performance characteristics of the High Sensitivity DNA kit for the Agilent 2100 Bioanalyzer Technical Note 10 Measured conc. [ng/µl] 1 Y intercept = 0.09 r 2 = 0.993 0.1 0.1 1 10 Reference concentration
Performance characteristics of the High Sensitivity DNA kit for the Agilent 2100 Bioanalyzer Technical Note 10 Measured conc. [ng/µl] 1 Y intercept = 0.09 r 2 = 0.993 0.1 0.1 1 10 Reference concentration
AxyPrep PCR Clean-up Kit AxyPrep-96 PCR Clean-up Kit
 AxyPrep PCR Clean-up Kit AxyPrep-96 PCR Clean-up Kit For the purification of amplicons from PCRs Kit contents, storage and stability AxyPrep PCR Clean-up Kit Cat. No. AP-PCR-4 AP-PCR-50 AP-PCR-250 Kit
AxyPrep PCR Clean-up Kit AxyPrep-96 PCR Clean-up Kit For the purification of amplicons from PCRs Kit contents, storage and stability AxyPrep PCR Clean-up Kit Cat. No. AP-PCR-4 AP-PCR-50 AP-PCR-250 Kit
Polymerase Chain Reaction: Application and Practical Primer Probe Design qrt-pcr
 Polymerase Chain Reaction: Application and Practical Primer Probe Design qrt-pcr review Enzyme based DNA amplification Thermal Polymerarase derived from a thermophylic bacterium DNA dependant DNA polymerase
Polymerase Chain Reaction: Application and Practical Primer Probe Design qrt-pcr review Enzyme based DNA amplification Thermal Polymerarase derived from a thermophylic bacterium DNA dependant DNA polymerase
Functional Genomics Research Stream. Research Meetings: November 2 & 3, 2009 Next Generation Sequencing
 Functional Genomics Research Stream Research Meetings: November 2 & 3, 2009 Next Generation Sequencing Current Issues Research Meetings: Meet with me this Thursday or Friday. (bring laboratory notebook
Functional Genomics Research Stream Research Meetings: November 2 & 3, 2009 Next Generation Sequencing Current Issues Research Meetings: Meet with me this Thursday or Friday. (bring laboratory notebook
Introduction. Kit components. 50 Preps GF-PC-050. Product Catalog No. 200 Preps GF-PC Preps GF-PC Preps SAMPLE
 Introduction The GF-1 PCR Clean Up Kit is a system designed for rapid clean up of DNA bands ranging from 100bp to 20kb. The GF-1 PCR Clean Up Kit contains special buffers to provide the correct salt concentration
Introduction The GF-1 PCR Clean Up Kit is a system designed for rapid clean up of DNA bands ranging from 100bp to 20kb. The GF-1 PCR Clean Up Kit contains special buffers to provide the correct salt concentration
CloneDirect Rapid Ligation Kit
 CloneDirect Rapid Ligation Kit Lucigen Corporation 2905 Parmenter St, Middleton, WI 53562 USA Toll Free: (888) 575-9695 (608) 831-9011 FAX: (608) 831-9012 lucigen@lucigen.com www.lucigen.com FOR RESEARCH
CloneDirect Rapid Ligation Kit Lucigen Corporation 2905 Parmenter St, Middleton, WI 53562 USA Toll Free: (888) 575-9695 (608) 831-9011 FAX: (608) 831-9012 lucigen@lucigen.com www.lucigen.com FOR RESEARCH
SYBR Premix Ex Taq II (Tli RNaseH Plus), Bulk
 Cat. # RR820L For Research Use SYBR Premix Ex Taq II (Tli RNaseH Plus), Bulk Product Manual The long-term storage temperature of this product has been changed to -20 since Lot. AK9104. See section IV.
Cat. # RR820L For Research Use SYBR Premix Ex Taq II (Tli RNaseH Plus), Bulk Product Manual The long-term storage temperature of this product has been changed to -20 since Lot. AK9104. See section IV.
Biol/Chem 475 Spring 2007
 Biol/Chem 475 Spring 2007 Goal of lab: For most of the quarter, we will be exploring a gene family that was first discovered in fruitlfies and then found to be present in humans and worms and fish and
Biol/Chem 475 Spring 2007 Goal of lab: For most of the quarter, we will be exploring a gene family that was first discovered in fruitlfies and then found to be present in humans and worms and fish and
Real Time PCR (qpcr) Dott. Finetti Luca
 Real Time PCR (qpcr) Dott. Finetti Luca fntlcu1@unife.it PCR (Polymerase Chain Reaction) PCR (Polymerase Chain Reaction) Theoretically: 2 n PCR (Polymerase Chain Reaction) It often used as a qualitative
Real Time PCR (qpcr) Dott. Finetti Luca fntlcu1@unife.it PCR (Polymerase Chain Reaction) PCR (Polymerase Chain Reaction) Theoretically: 2 n PCR (Polymerase Chain Reaction) It often used as a qualitative
7 Synthesizing the ykkcd Mutant Toxin Sensor RNA in vitro
 7 Synthesizing the ykkcd Mutant Toxin Sensor RNA in vitro 7.1 Learning Objective In the quest toward understanding how the ykkcd toxin sensor recognizes the antibiotic tetracycline you thus far designed
7 Synthesizing the ykkcd Mutant Toxin Sensor RNA in vitro 7.1 Learning Objective In the quest toward understanding how the ykkcd toxin sensor recognizes the antibiotic tetracycline you thus far designed
DNA Labeling Kits Fluorescein-dCTP and -dutp Instruction Manual
 DNA Labeling Kits Fluorescein-dCTP and -dutp Instruction Manual Catalog Number 170-8223 170-8224 For Technical Service Call Your Local Bio-Rad Office or in the U.S. Call 1-800-4BIORAD (1-800-424-6723)
DNA Labeling Kits Fluorescein-dCTP and -dutp Instruction Manual Catalog Number 170-8223 170-8224 For Technical Service Call Your Local Bio-Rad Office or in the U.S. Call 1-800-4BIORAD (1-800-424-6723)
PCR OPTIMIZATION AND TROUBLESHOOTING
 PCR OPTIMIZATION AND TROUBLESHOOTING Amplification of each DNA fragment can occur only under the defined conditions which are provided by a reaction mixture. If no positive PCR result can be obtained,
PCR OPTIMIZATION AND TROUBLESHOOTING Amplification of each DNA fragment can occur only under the defined conditions which are provided by a reaction mixture. If no positive PCR result can be obtained,
PCR PRIMER DESIGN SARIKA GARG SCHOOL OF BIOTECHNOLGY DEVI AHILYA UNIVERSITY INDORE INDIA
 PCR PRIMER DESIGN SARIKA GARG SCHOOL OF BIOTECHNOLGY DEVI AHILYA UNIVERSITY INDORE-452017 INDIA BIOINFORMATICS Bioinformatics is considered as amalgam of biological sciences especially Biotechnology with
PCR PRIMER DESIGN SARIKA GARG SCHOOL OF BIOTECHNOLGY DEVI AHILYA UNIVERSITY INDORE-452017 INDIA BIOINFORMATICS Bioinformatics is considered as amalgam of biological sciences especially Biotechnology with
DNA Labeling Kits Texas Red -dctp and -dutp Instruction Manual
 DNA Labeling Kits Texas Red -dctp and -dutp Instruction Manual Catalog Number 170-8221 170-8222 For Technical Service Call Your Local Bio-Rad Office or in the U.S. Call 1-800-4BIORAD (1-800-424-6723) Bio-Rad
DNA Labeling Kits Texas Red -dctp and -dutp Instruction Manual Catalog Number 170-8221 170-8222 For Technical Service Call Your Local Bio-Rad Office or in the U.S. Call 1-800-4BIORAD (1-800-424-6723) Bio-Rad
Finishing Drosophila Ananassae Fosmid 2728G16
 Finishing Drosophila Ananassae Fosmid 2728G16 Kyle Jung March 8, 2013 Bio434W Professor Elgin Page 1 Abstract For my finishing project, I chose to finish fosmid 2728G16. This fosmid carries a segment of
Finishing Drosophila Ananassae Fosmid 2728G16 Kyle Jung March 8, 2013 Bio434W Professor Elgin Page 1 Abstract For my finishing project, I chose to finish fosmid 2728G16. This fosmid carries a segment of
Genomic DNA Clean & Concentrator -10 Catalog Nos. D4010 & D4011
 INSTRUCTION MANUAL Genomic DNA Clean & Concentrator -10 Catalog Nos. D4010 & D4011 Highlights Quick (5 minute) spin column recovery of large-sized DNA (e.g., genomic, mitochondrial, plasmid (BAC/PAC),
INSTRUCTION MANUAL Genomic DNA Clean & Concentrator -10 Catalog Nos. D4010 & D4011 Highlights Quick (5 minute) spin column recovery of large-sized DNA (e.g., genomic, mitochondrial, plasmid (BAC/PAC),
Gene-Spin DNA Purification Kit - V3
 Protech Technology 波仕特生物科技 Gene-Spin 1-4-3 DNA Purification Kit - V3 (For PCR/DNA clean-up & Gel Extraction) Innovative Tools for Nucleic Acid Purification This Kit is for research purposes only. Not for
Protech Technology 波仕特生物科技 Gene-Spin 1-4-3 DNA Purification Kit - V3 (For PCR/DNA clean-up & Gel Extraction) Innovative Tools for Nucleic Acid Purification This Kit is for research purposes only. Not for
Experiment (5): Polymerase Chain Reaction (PCR)
 BCH361 [Practical] Experiment (5): Polymerase Chain Reaction (PCR) Aim: Amplification of a specific region on DNA. Primer design. Determine the parameters that may affect he specificity, fidelity and efficiency
BCH361 [Practical] Experiment (5): Polymerase Chain Reaction (PCR) Aim: Amplification of a specific region on DNA. Primer design. Determine the parameters that may affect he specificity, fidelity and efficiency
We begin with a high-level overview of sequencing. There are three stages in this process.
 Lecture 11 Sequence Assembly February 10, 1998 Lecturer: Phil Green Notes: Kavita Garg 11.1. Introduction This is the first of two lectures by Phil Green on Sequence Assembly. Yeast and some of the bacterial
Lecture 11 Sequence Assembly February 10, 1998 Lecturer: Phil Green Notes: Kavita Garg 11.1. Introduction This is the first of two lectures by Phil Green on Sequence Assembly. Yeast and some of the bacterial
AMPURE PCR PURIFICATION PAGE 1 OF 7
 PCR PURIFICATION PAGE 1 OF 7 Please refer to http://www.agencourt.com/technical/reagent_information/ for updated protocols. AMPure is a registered trademark of Agencourt Bioscience and is for laboratory
PCR PURIFICATION PAGE 1 OF 7 Please refer to http://www.agencourt.com/technical/reagent_information/ for updated protocols. AMPure is a registered trademark of Agencourt Bioscience and is for laboratory
Real-Time PCR Principles and Applications
 Real-Time PCR Principles and Applications Dr Esam Ibraheem Azhar (BSc, MSc, Ph.D Molecular Medical Virology) Asst. Prof. Medical Laboratory Technology Department Objectives Real-Time PCR Principles and
Real-Time PCR Principles and Applications Dr Esam Ibraheem Azhar (BSc, MSc, Ph.D Molecular Medical Virology) Asst. Prof. Medical Laboratory Technology Department Objectives Real-Time PCR Principles and
BIOO LIFE SCIENCE PRODUCTS
 BIOO LIFE SCIENCE PRODUCTS FOR REFERENCE PURPOSES This manual is for Reference Purposes Only. DO NOT use this protocol to run your assays. Periodically, optimizations and revisions are made to the kit
BIOO LIFE SCIENCE PRODUCTS FOR REFERENCE PURPOSES This manual is for Reference Purposes Only. DO NOT use this protocol to run your assays. Periodically, optimizations and revisions are made to the kit
EZ-10 SPIN COLUMN HANDBOOK
 EZ-0 SPIN COLUMN HANDBOOK EZ-0 Spin Column Plasmid DNA Mini Kit EZ-0 Spin Column PCR Products Purification Kit EZ-0 Spin Column DNA Gel Extraction Kit Version 20.0 Rev 3/23/205 ISO900 Certified 20 Konrad
EZ-0 SPIN COLUMN HANDBOOK EZ-0 Spin Column Plasmid DNA Mini Kit EZ-0 Spin Column PCR Products Purification Kit EZ-0 Spin Column DNA Gel Extraction Kit Version 20.0 Rev 3/23/205 ISO900 Certified 20 Konrad
 PRODUCT INFORMATION Thermo Scientific Luminaris Probe Low ROX qpcr Master Mix #K0944 For 5000 rxns Lot Expiry Date Store at -20 C in the dark CERTIFICATE OF ANALYSIS The absence of endo-, exodeoxyribonucleases
PRODUCT INFORMATION Thermo Scientific Luminaris Probe Low ROX qpcr Master Mix #K0944 For 5000 rxns Lot Expiry Date Store at -20 C in the dark CERTIFICATE OF ANALYSIS The absence of endo-, exodeoxyribonucleases
All-in-One qpcr Mix. User Manual. For universal quantitative real-time PCR
 All-in-One qpcr Mix For universal quantitative real-time PCR Cat. No. AOPR-0200 (200 qpcr reactions) Cat. No. AOPR-0600 (600 qpcr reactions) Cat. No. AOPR-1000 (1000 qpcr reactions) Cat. No. AOPR-1200
All-in-One qpcr Mix For universal quantitative real-time PCR Cat. No. AOPR-0200 (200 qpcr reactions) Cat. No. AOPR-0600 (600 qpcr reactions) Cat. No. AOPR-1000 (1000 qpcr reactions) Cat. No. AOPR-1200
AmpliTaq 360 DNA Polymerase
 Product Bulletin AmpliTaq 360 AmpliTaq 360 360 Coverage for a Full Range of Targets AmpliTaq 360 Benefits Optimized for the broadest range of targets (
Product Bulletin AmpliTaq 360 AmpliTaq 360 360 Coverage for a Full Range of Targets AmpliTaq 360 Benefits Optimized for the broadest range of targets (
SunScript One Step RT-PCR Kit
 SunScript ONE STEP R T-PCR KIT HANDBOOK SunScript One Step RT-PCR Kit INDEX Legal... 4 Intended use... 4 Kit contents... 5 Shipping and storage... 5 Handling... 6 Quality control... 6 Reagents and equipment...
SunScript ONE STEP R T-PCR KIT HANDBOOK SunScript One Step RT-PCR Kit INDEX Legal... 4 Intended use... 4 Kit contents... 5 Shipping and storage... 5 Handling... 6 Quality control... 6 Reagents and equipment...
