Supplementary Figure 1: sgrna library generation and the length of sgrnas for the functional screen. (a) A diagram of the retroviral vector for sgrna
|
|
|
- Anastasia Roberts
- 6 years ago
- Views:
Transcription
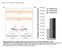
1 Supplementary Figure 1: sgrna library generation and the length of sgrnas for the functional screen. (a) A diagram of the retroviral vector for sgrna expression. It contains a U6-promoter-driven sgrna expression cassette, and a PGK-promoter-driven puromycin resistance gene. Restriction enzyme sites for cloning of the sgrna library are also indicated. (b) A list of 17 mouse mirna or mirna clusters included as input for the Molecular Chipper procedure. (c) An empty control vector (Ctrl), an sgrna vector with G+19mer design (G+19) and an sgrna vector with G+20mer design (G+20) were transduced into the mir-142-3p reporter cell line. Top: representative FACS plot demonstrating efficient inhibition of mir-142 activity. Bottom: Design of the G+19 and G+20 sgrnas, with mir-142 genomic sequence in the middle. Yellow highlighted regions reflect mature mir-142 mirna sequences. NGG PAM (CCA on the antisense strand) is shown in red. Note that the first base G in both G+19mer and G+20mer is a mismatch on the target sequence. 1
2 Supplementary Figure 2: Additional data for the sgrna-based functional screen. (a) Enrichment plots similar to those in Fig 3a is shown. Left panel, sgrna enrichments in mir- 142 are shown for high- and med-gfp populations (both representing near complete ablation of mir-142 expression) in each of the three biological triplicates. X-axis indicates position in bp. Horizontal black bars indicate the locations of mature mirnas. Blue and red indicate enriched sgrnas that were mapped to sense and antisense strand, respectively. The positions of the last targeting-domain base are used to reflect the positions of sgrnas. Blue and red boxes indicate 5 - and 3 -hit regions. Yellow box highlights sgrnas mapped to the pre-mir-142 region (including mature mir-142-5p and mir-142-3p, and loop region in between). (b) Enrichment plots for sgrnas mapped to mir-126 from the high-gfp populations are shown, in biological triplicates. 2
3 Supplementary Figure 3: ESCScanner analysis of sgrna enrichment in the functional screen. (a) Diagram for the principle of the ESCScanner algorithm, which is used to scan for enriched clusters of sgrnas from the screen. ESCScanner calculates the probability of observing a pattern of sgrna enrichment within a scanning window. Detected sgrnas are shown in orange and enriched sgrnas are shown in blue peaks. (b) ESCScanner plots are shown for the mir- 142 region for the three biological replicates. Data for the Low-GFP samples are shown. Specifically, a 21-bp scanning window was used and the algorithm was applied independently to the three biological replicates. X-axis indicates the position of the center of the scanning window. Y-axis indicates the log10 (probability). Horizontal black bars indicate the locations of mature mirnas. Blue and red boxes indicate 5 - and 3 -hit regions. Yellow box highlights sgrnas mapped to the pre-mir-142 region (including mature mir-142-5p and mir-142-3p, and loop region in between). Pink box indicates the position of the control 3 deletion (CtrlΔ3 ) for Supplementary Fig. 4. (c) ESCScanner plots are shown for the mir-126 region for the three biological replicates. Data for the Low-GFP samples are shown. 3
4 Supplementary Figure 4. mirna processing reporter activities for additional mir-142 mutants. (a) A Genome Browser snap shot of RNA seq data of murine spleen cells is shown. Data are from the CSHL long RNA-seq trace. The black bars at the bottom indicate relative positions of the 5 - hit region, mir-142-5p, mir-142-3p and the 3 - hit region. Note that abundant RNA signals can be observed from >1kb upstream of the hit regions on mir-142. (b) Designs of additional mir- 142 mutants cloned into a mirna processing reporter. Designs are similar to those in Fig. 4a. Additional mutants include a control 3 deletion (CtrlΔ3, 20 bp deletion), deleting hairpin from wildtype pri-mir-142 (ΔH), deleting hairpin from the three mutant pri-mir-142 constructs as in Fig 4a (LΔ3 ΔH, SΔ3 ΔH, and Δ5 ΔH). The narrow vertical blue bar upstream of the 3 -hit region depicts a putative CNNC site, which was not disrupted by the deletions. (c) Processing reporter activities of the indicated reporter constructs were determined in the BaF3 cells. * P<0.05; **P<0.01; NS: not significant. N=3 biological replicates. Data from a representative experiment out of two performed. Error bars represent s.d. (d) GFP fluorescence of the indicated processing reporters were determined in the indicated cell lines. N=3 biological replicates. Data were normalized to the mean GFP level in WT construct, and are from a representative experiment out of two performed. Error bars represent s.d. (e) Processing efficiencies of the indicated human primir-142 mutant reporters were determined in BaF3 cells. **P<0.01; student s t-test. N=3 4
5 biological replicates. Data from a representative experiment out of two performed. Error bars represent s.d. 5
6 Supplementary Figure 5: Compatibility of Molecular-Chipper-generated sgrna library with VQR-Cas9. (a) A NGA-PAM sgrna in our library, mapped to the mir-142 loop, was cloned and transduced into BaF3 mir-142-3p reporter cells (no Cas9), reporter cells expressing the wildtype Cas9 (WT-Cas9) or reporter cells expressing VQR-Cas9 (with dsredexpress as selection marker (VQR-Cas9-dsRed)). GFP and mcherry expression levels were determined by flow cytometry. Representative flow cytometry plots are shown, with numbers indicating the percentage of cells within the gates. (b) Representative flow cytometry plots are shown for sgrna library transduced BaF3 mir-142-3p reporter cells (no Cas9), the reporter cells expressing only WT- Cas9, the reporter cells expressing only VQR-Cas9 (with dsredexpress as a marker), or the reporter cells expressing both WT-Cas9 and VQR-Cas9. Number indicates the frequency of the gated populations. (c) Quantified % of GFP+ cells (reflecting low mir-142 activity) for data in b. N=3 biological replicates. Error bars represent s.d. Numbers at each bar indicate average % of GFP+ cells. (d-e) Experiments similar to (a-c) were performed, except that a lentiviral vector with zeocin as selection marker was used. Similar results were observed. 6
7 Supplementary Figure 6: Additional data of the sgrna library complexity and enrichment. (a) The saturation effect of the Molecular-Chipper-generated sgrna library. The indicated number of mapped sequence reads were randomly selected from all sequence reads, and the number of unique sgrnas was determined in each set of randomly selected sequences. Note that unique sgrnas were observed with 160,000 randomly selected mapped deep sequencing reads. At this level, the median distance of neighboring NGG-PAM sgrnas is 10 bp. (b) A histogram of the log2(enrichment) scores of all NGG-PAM sgrnas in our library is shown. Data are based on all experimental pairs of samples (low-gfp, med-gfp and high-gfp versus neg-gfp). There were 2.1% of NGG-PAM sgrnas with >1 of log2(enrichment). 7
8 Supplementary Table 1. Primer Sequences for mirnas and their Flanking Regions mirna construct Primer sequences to amplify mirna genomic fragments Sense Antisense Size of PCR (bp) mmu mir 16 1 TACTATTGAGGTGCTAGGAG CTAAAACCCATGGCCTTGTG 484 mmu mir 23b GGTCCCTAAGGTATTGGTCT CGGCCATAATTAAAGGGCTG 408 mmu mir 23a_mir27a_miR 24 2_cluster TGTGTGGTGAGGTGTACCTA TTAATGCAAGGGTTACCCGC 780 hsa mir 24 1 AAACCCAGGTGCATCAAGGA AACACGTGGCAAATGCTCAG 495 hsa mir 26a 2 AAGGAACACTTGTGCAGGCT CTGTGCACTGACAGAAAAGC 485 hsa mir 26b TCTGCACTACACCCAGGTT CAGGAACAGTGGAGGAGGA 467 hsa mir 27b GGTTCCTGGCATGCTGATTT TTCTGTGACTCCCAATACAC 506 hsa mir 29a CCTGAAGTAAGTGTCCAGTG ACACACCCACCATCACTATG 488 hsa mir 29b 1 GAGGGCTCAGTTACCATTTG ATGTAGGTCTTCATCCGAGC 471 hsa mir 29b 2_mir 29c_cluster ATGTTGAATGGATTTGGTTCTT TAACAGTACTACATGTCAGAAA 1026 hsa mir 125b 2 GTTCTGATCTATAGGTGGCC GCCATATCTTTAGAGTACCG 433 hsa mir 126 GCTGAGCTGAACATGGAAGA AGATCCAGGCGCTAGTCAG 524 mmu mir 142 ATGTTGAGTCACCACCCACA TATCCTCACGGACGTGCATT 458 hsa mir 146 CCACATGCCCAGCATGTTTA CAATCCAACATGACTCCCTC 402 hsa mir 155 CTGAAGTCTACCTTGCCTTC CATGTGGGCTTGAAGTTGAG 472 hsa mir 222 AGGAAGTGAATCTAAAGGTAG GACTTCATCCTTCATCAAACT 426 mmu mir 451 GACCTTGGCTGGGATATCAT TCTTTGGCACAGTGAAGAGG 362 mirna Constructs: the murine mirna(s) included as input for sgrna library production. Sense: Sequences of sense primer in PCR from murine genomic DNA to amplify the corresponding mirna. Antisense: Sequences of antisense primer in PCR from murine genomic DNA to amplify the corresponding mirna. Size: Size of mirna plus flanking regions for each mirna or mirna cluster in bp.
9 Supplementary Table 2. Candidate sgrnas that disrupt mir 142 expression samples sgrna Gene Strand Position Length PAM Sam1HighGFP Sam1MedGFP Sam1LowGFP Sam2HighGFP Sam2MedGFP Sam2LowGFP Sam3HighGFP Sam3MedGFP Sam3LowGFP gtcaccacccacaaggccca mir NGG ccacccacaaggcccaggg mir NGG cacccacaaggcccaggg mir NGG ccagggcgggccctctagg mir NGG accgctccaccctgcctg mir NGG gaccgctccaccctgcctg mir NGG ggaccgctccaccctgcctg mir NGG cccctccgtgtaacttccca mir NGG ccctccgtgtaacttccca mir NGG tagtagtgctttctacttta mir NGG cactactaacagcactgga mir NGG agtgcactcatccataaagt mir NGG gatgagtgcactgtgggctt mir NGG atgagtgcactgtgggctt mir NGG cggagaccacgccacgccg mir NGG agggggccgcggcgtggcg mir NGG ggtggcagggggccgcggcg mir NGG gtggcagggggccgcggcg mir NGG ggggagcctggccaaatga mir NGG gcattcgagagagcggctg mir NGG gggcaggggcctttattaa mir NGG cggagaccacgccacgcc mir NGG agaccacgccacgccgcgg mir NGG gtggcagggggccgcggc mir NGG gaccgctccaccctgcct mir GTGG accgctccaccctgcct mir GTGG Candidate sgrnas were derived by the criteria in Methods. sgrna: Sequences of sgrnas, without first base of G. Only sgrnas located before "NGG" PAM (hiligted in yellow) or "GTGG" PAM (hilighted in blue) are shown. Gene: The mirna gene in which the sgrna is located. Strand: The strand to which the sgrna is mapped. "1" stands for "+" strand, "0" stands for " " strand. Position: The position of sgrna relative to the input mirna sequence. Position is calculated by the position of the last base of the targeting domain in sgrnas. Length: The length of sgrna targeting domain (including the first base of G, which is not shown in the sequence). Samples: The log2 enrichment level for each sgrna in each indicated sample is shown. Log2 enrichment level below 0 is shown as 0.
10 Supplementary Table 3. Samples and Barcodes for Deep Sequencing Barcode Primer Barcode Screen populations CGTGAT ATCACG Sam1NegGFP CTGATC GATCAG Sam1LowGFP GGACGG CCGTCC Sam1MedGFP CTCTAC GTAGAG Sam1HighGFP CACTGT ACAGTG Sam2NegGFP TTGACT AGTCAA Sam2LowGFP GTAGCC GGCTAC Sam2MedGFP TACAAG CTTGTA Sam2HighGFP TTTCAC GTGAAA Sam3NegGFP CGAAAC GTTTCG Sam3LowGFP CGTACG CGTACG Sam3MedGFP CCACTC GAGTGG Sam3HighGFP GCGGAC GTCCGC Unsorted Barcode: Barcode sequence in sequencing reads. Primer Barcode: Barcode sequence in reverse PCR primer to amplily sgrnas for deep sequencing. These barcodes are reverse complement of those listed in "Barcode". Screen populations: Cells with different GFP levels were sorted and/or re sorted from the BaF3 mir 142 3p reporter cell line transduced by the sgrna library in 3 biological replicates, sample 1, sample 2 and sample 3.
Supplementary Figure 1
 Supplementary Figure 1 CITE-seq library preparation. (a) Illustration of the DNA-barcoded antibodies used in CITE-seq. (b) Antibody-oligonucleotide complexes appear as a high-molecular-weight smear when
Supplementary Figure 1 CITE-seq library preparation. (a) Illustration of the DNA-barcoded antibodies used in CITE-seq. (b) Antibody-oligonucleotide complexes appear as a high-molecular-weight smear when
TailorMix Stranded mrna Sample Preparation Kit
 TailorMix Stranded mrna Sample Preparation Kit Catalog Numbers: TM200-A and TM200-B Introduction The TailorMix Stranded mrna Sample Preparation Kit from SeqMatic is a comprehensive solution for generating
TailorMix Stranded mrna Sample Preparation Kit Catalog Numbers: TM200-A and TM200-B Introduction The TailorMix Stranded mrna Sample Preparation Kit from SeqMatic is a comprehensive solution for generating
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Endogenous gene tagging to study subcellular localization and chromatin binding. a, b, Schematic of experimental set-up to endogenously tag RNAi factors using the CRISPR Cas9 technology,
Supplementary Figure 1 Endogenous gene tagging to study subcellular localization and chromatin binding. a, b, Schematic of experimental set-up to endogenously tag RNAi factors using the CRISPR Cas9 technology,
File name: Supplementary Information Description: Supplementary figures and supplementary tables. File name: Peer review file Description:
 File name: Supplementary Information Description: Supplementary figures and supplementary tables. File name: Peer review file Description: Supplementary Figure 1. dcas9-mq1 fusion protein induces de novo
File name: Supplementary Information Description: Supplementary figures and supplementary tables. File name: Peer review file Description: Supplementary Figure 1. dcas9-mq1 fusion protein induces de novo
i-stop codon positions in the mcherry gene
 Supplementary Figure 1 i-stop codon positions in the mcherry gene The grnas (green) that can potentially generate stop codons from Trp (63 th and 98 th aa, upper panel) and Gln (47 th and 114 th aa, bottom
Supplementary Figure 1 i-stop codon positions in the mcherry gene The grnas (green) that can potentially generate stop codons from Trp (63 th and 98 th aa, upper panel) and Gln (47 th and 114 th aa, bottom
Guidelines for sequencing SCC libraries All libraries made using SCC supplied hydrogel after are V3 libraries
 2018-6-20 Guidelines for sequencing SCC libraries All libraries made using SCC supplied hydrogel after 1-1-2017 are V3 libraries Upon receiving your libraries: When your libraries are delivered you will
2018-6-20 Guidelines for sequencing SCC libraries All libraries made using SCC supplied hydrogel after 1-1-2017 are V3 libraries Upon receiving your libraries: When your libraries are delivered you will
GENERAL INFORMATION...
 BIOO LIFE SCIENCE PRODUCTS NEXTflex TM ChIP-Seq Barcodes - 6 (Illumina Compatible) Catalog #: 514120 (48 reactions) BIOO Scientific Corp. 2012 V13.05 TABLE OF CONTENTS GENERAL INFORMATION... 1 Product
BIOO LIFE SCIENCE PRODUCTS NEXTflex TM ChIP-Seq Barcodes - 6 (Illumina Compatible) Catalog #: 514120 (48 reactions) BIOO Scientific Corp. 2012 V13.05 TABLE OF CONTENTS GENERAL INFORMATION... 1 Product
NEXTFLEX Small RNA Barcode Primers - Set A (For Illumina Platforms) Catalog #NOVA (Kit contains 96 reactions)
 NEXTFLEX Small RNA Barcode Primers - Set A (For Illumina Platforms) Catalog #NOVA-513305 (Kit contains 96 reactions) Bioo Scientific Corp. 2014-2018 V14.07 This product is for research use only. Not for
NEXTFLEX Small RNA Barcode Primers - Set A (For Illumina Platforms) Catalog #NOVA-513305 (Kit contains 96 reactions) Bioo Scientific Corp. 2014-2018 V14.07 This product is for research use only. Not for
NEXTflex PCR-Free Barcodes - 6 (For Illumina Platforms) Catalog # (Kit contains 48 reactions) Bioo Scientific Corp V12.
 NEXTflex PCR-Free Barcodes - 6 (For Illumina Platforms) Catalog #514110 (Kit contains 48 reactions) Bioo Scientific Corp. 2014-2018 V12.10 This product is for research use only. Not for use in diagnostic
NEXTflex PCR-Free Barcodes - 6 (For Illumina Platforms) Catalog #514110 (Kit contains 48 reactions) Bioo Scientific Corp. 2014-2018 V12.10 This product is for research use only. Not for use in diagnostic
Zhang et al., RepID facilitates replication Initiation. Supplemental Information:
 Supplemental Information: a b 1 Supplementary Figure 1 (a) DNA sequence of all the oligonucleotides used in this study. Only one strand is shown. The unshaded nucleotide sequences show changes from the
Supplemental Information: a b 1 Supplementary Figure 1 (a) DNA sequence of all the oligonucleotides used in this study. Only one strand is shown. The unshaded nucleotide sequences show changes from the
sherwood - UltramiR shrna Collections
 sherwood - UltramiR shrna Collections Incorporating advances in shrna design and processing for superior potency and specificity sherwood - UltramiR shrna Collections Enabling Discovery Across the Genome
sherwood - UltramiR shrna Collections Incorporating advances in shrna design and processing for superior potency and specificity sherwood - UltramiR shrna Collections Enabling Discovery Across the Genome
Nature Biotechnology: doi: /nbt Supplementary Figure 1
 Supplementary Figure 1 Negative selection analysis of sgrnas targeting all Brd4 exons comparing day 2 to day 10 time points. Systematic evaluation of 64 Brd4 sgrnas in negative selection experiments, targeting
Supplementary Figure 1 Negative selection analysis of sgrnas targeting all Brd4 exons comparing day 2 to day 10 time points. Systematic evaluation of 64 Brd4 sgrnas in negative selection experiments, targeting
Nature Genetics: doi: /ng Supplementary Figure 1. ChIP-seq genome browser views of BRM occupancy at previously identified BRM targets.
 Supplementary Figure 1 ChIP-seq genome browser views of BRM occupancy at previously identified BRM targets. Gene structures are shown underneath each panel. Supplementary Figure 2 pref6::ref6-gfp complements
Supplementary Figure 1 ChIP-seq genome browser views of BRM occupancy at previously identified BRM targets. Gene structures are shown underneath each panel. Supplementary Figure 2 pref6::ref6-gfp complements
Supporting Information
 Supporting Information Gómez-Marín et al. 10.1073/pnas.1505463112 SI Materials and Methods Generation of BAC DK74B2-six2a::GFP-six3a::mCherry-iTol2. BAC clone (number 74B2) from DanioKey zebrafish BAC
Supporting Information Gómez-Marín et al. 10.1073/pnas.1505463112 SI Materials and Methods Generation of BAC DK74B2-six2a::GFP-six3a::mCherry-iTol2. BAC clone (number 74B2) from DanioKey zebrafish BAC
Supplementary Figure 1
 Supplementary Figure 1 Concept of barcoding to suppress error in sequencing. Each template DNA molecule is barcoded with a random and unique sequence (marked as red, turquoise and green). All PCR generated
Supplementary Figure 1 Concept of barcoding to suppress error in sequencing. Each template DNA molecule is barcoded with a random and unique sequence (marked as red, turquoise and green). All PCR generated
Nature Biotechnology: doi: /nbt Supplementary Figure 1. Number and length distributions of the inferred fosmids.
 Supplementary Figure 1 Number and length distributions of the inferred fosmids. Fosmid were inferred by mapping each pool s sequence reads to hg19. We retained only those reads that mapped to within a
Supplementary Figure 1 Number and length distributions of the inferred fosmids. Fosmid were inferred by mapping each pool s sequence reads to hg19. We retained only those reads that mapped to within a
Percent survival. Supplementary fig. S3 A.
 Supplementary fig. S3 A. B. 100 Percent survival 80 60 40 20 Ml 0 0 100 C. Fig. S3 Comparison of leukaemia incidence rate in the triple targeted chimaeric mice and germline-transmission translocator mice
Supplementary fig. S3 A. B. 100 Percent survival 80 60 40 20 Ml 0 0 100 C. Fig. S3 Comparison of leukaemia incidence rate in the triple targeted chimaeric mice and germline-transmission translocator mice
Supplementary Figure 1
 number of cells, normalized number of cells, normalized number of cells, normalized Supplementary Figure CD CD53 Cd3e fluorescence intensity fluorescence intensity fluorescence intensity Supplementary
number of cells, normalized number of cells, normalized number of cells, normalized Supplementary Figure CD CD53 Cd3e fluorescence intensity fluorescence intensity fluorescence intensity Supplementary
Supplementary Figure 1 An overview of pirna biogenesis during fetal mouse reprogramming. (a) (b)
 Supplementary Figure 1 An overview of pirna biogenesis during fetal mouse reprogramming. (a) A schematic overview of the production and amplification of a single pirna from a transposon transcript. The
Supplementary Figure 1 An overview of pirna biogenesis during fetal mouse reprogramming. (a) A schematic overview of the production and amplification of a single pirna from a transposon transcript. The
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Origin use and efficiency are similar among WT, rrm3, pif1-m2, and pif1-m2; rrm3 strains. A. Analysis of fork progression around confirmed and likely origins (from cerevisiae.oridb.org).
Supplementary Figure 1 Origin use and efficiency are similar among WT, rrm3, pif1-m2, and pif1-m2; rrm3 strains. A. Analysis of fork progression around confirmed and likely origins (from cerevisiae.oridb.org).
Schematic representation of the endogenous PALB2 locus and gene-disruption constructs
 Supplementary Figures Supplementary Figure 1. Generation of PALB2 -/- and BRCA2 -/- /PALB2 -/- DT40 cells. (A) Schematic representation of the endogenous PALB2 locus and gene-disruption constructs carrying
Supplementary Figures Supplementary Figure 1. Generation of PALB2 -/- and BRCA2 -/- /PALB2 -/- DT40 cells. (A) Schematic representation of the endogenous PALB2 locus and gene-disruption constructs carrying
SUPPLEMENTARY INFORMATION
 AS-NMD modulates FLM-dependent thermosensory flowering response in Arabidopsis NATURE PLANTS www.nature.com/natureplants 1 Supplementary Figure 1. Genomic sequence of FLM along with the splice sites. Sequencing
AS-NMD modulates FLM-dependent thermosensory flowering response in Arabidopsis NATURE PLANTS www.nature.com/natureplants 1 Supplementary Figure 1. Genomic sequence of FLM along with the splice sites. Sequencing
Parts of a standard FastQC report
 FastQC FastQC, written by Simon Andrews of Babraham Bioinformatics, is a very popular tool used to provide an overview of basic quality control metrics for raw next generation sequencing data. There are
FastQC FastQC, written by Simon Andrews of Babraham Bioinformatics, is a very popular tool used to provide an overview of basic quality control metrics for raw next generation sequencing data. There are
Supporting Information
 Supporting Information Kilian et al. 10.1073/pnas.1105861108 SI Materials and Methods Determination of the Electric Field Strength Required for Successful Electroporation. The transformation construct
Supporting Information Kilian et al. 10.1073/pnas.1105861108 SI Materials and Methods Determination of the Electric Field Strength Required for Successful Electroporation. The transformation construct
Protocol for cloning SEC-based repair templates using Gibson assembly and ccdb negative selection
 Protocol for cloning SEC-based repair templates using Gibson assembly and ccdb negative selection Written by Dan Dickinson (daniel.dickinson@austin.utexas.edu) and last updated January 2018. A version
Protocol for cloning SEC-based repair templates using Gibson assembly and ccdb negative selection Written by Dan Dickinson (daniel.dickinson@austin.utexas.edu) and last updated January 2018. A version
Supplementary Figure 1 Activities of ABEs using extended sgrnas in HEK293T cells.
 Supplementary Figure 1 Activities of ABEs using extended sgrnas in HEK293T cells. Base editing efficiencies of ABEs with extended sgrnas at Site 18 (a), Site 19 (b), the HBB-E2 site (c), and the HBB-E3
Supplementary Figure 1 Activities of ABEs using extended sgrnas in HEK293T cells. Base editing efficiencies of ABEs with extended sgrnas at Site 18 (a), Site 19 (b), the HBB-E2 site (c), and the HBB-E3
Student Learning Outcomes (SLOS)
 Student Learning Outcomes (SLOS) KNOWLEDGE AND LEARNING SKILLS USE OF KNOWLEDGE AND LEARNING SKILLS - how to use Annhyb to save and manage sequences - how to use BLAST to compare sequences - how to get
Student Learning Outcomes (SLOS) KNOWLEDGE AND LEARNING SKILLS USE OF KNOWLEDGE AND LEARNING SKILLS - how to use Annhyb to save and manage sequences - how to use BLAST to compare sequences - how to get
Nature Immunology: doi: /ni.3694
 Supplementary Figure 1 Expression of Bhlhe41 and Bhlhe40 in B cell development and mature B cell subsets. (a) Scatter plot showing differential expression of genes between splenic B-1a cells and follicular
Supplementary Figure 1 Expression of Bhlhe41 and Bhlhe40 in B cell development and mature B cell subsets. (a) Scatter plot showing differential expression of genes between splenic B-1a cells and follicular
NEBNext Multiplex Small RNA Library Prep Set for Illumina Set 1, Set 2, Index Primers 1 48 and Multiplex Compatible
 LIBRARY PREPARATION NEBNext Multiplex Small RNA Library Prep Set for Illumina Set 1, Set 2, Index Primers 1 48 and Multiplex Compatible Instruction Manual NEB #E7300S/L, #E7580S/L, #E7560S, #E7330S/L 24/96
LIBRARY PREPARATION NEBNext Multiplex Small RNA Library Prep Set for Illumina Set 1, Set 2, Index Primers 1 48 and Multiplex Compatible Instruction Manual NEB #E7300S/L, #E7580S/L, #E7560S, #E7330S/L 24/96
Supplementary Information
 Supplementary Information Vidigal and Ventura a wt locus 5 region 3 region CCTCTGCCACTGCGAGGGCGTCCAATGGTGCTTG(...)AACAGGTGGAATATCCCTACTCTA predicted deletion clone 1 clone 2 clone 3 CCTCTGCCACTGCGAGGGCGTC-AGGTGGAATATCCCTACTCTA
Supplementary Information Vidigal and Ventura a wt locus 5 region 3 region CCTCTGCCACTGCGAGGGCGTCCAATGGTGCTTG(...)AACAGGTGGAATATCCCTACTCTA predicted deletion clone 1 clone 2 clone 3 CCTCTGCCACTGCGAGGGCGTC-AGGTGGAATATCCCTACTCTA
of NOBOX. Supplemental Figure S1F shows the staining profile of LHX8 antibody (Abcam, ab41519), which is a
 SUPPLEMENTAL MATERIALS AND METHODS LHX8 Staining During the course of this work, it came to our attention that the NOBOX antibody used in this study had been discontinued. There are a number of other nuclear
SUPPLEMENTAL MATERIALS AND METHODS LHX8 Staining During the course of this work, it came to our attention that the NOBOX antibody used in this study had been discontinued. There are a number of other nuclear
B. Transgenic plants with strong phenotype (%)
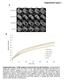
 A. TCTAGTTGTTGTTGTTATGGTCTAGTTGTTGTTGTTATGGTCTAATTT AAATATGGTCTAAAGAAGAAGAATATGGTCTAAAGAAGAAGAATATGG 2XP35S STTM165 5 GGGGGATGAAGctaCCTGGTCCGA3 3 CCCCCUACUUC---GGACCAGGCU5 mir165 HindIII mir165 96 nt GTTGTTGTTGTTATGGTCTAGTTGTTGTTGTTATGGTCTAATTT
A. TCTAGTTGTTGTTGTTATGGTCTAGTTGTTGTTGTTATGGTCTAATTT AAATATGGTCTAAAGAAGAAGAATATGGTCTAAAGAAGAAGAATATGG 2XP35S STTM165 5 GGGGGATGAAGctaCCTGGTCCGA3 3 CCCCCUACUUC---GGACCAGGCU5 mir165 HindIII mir165 96 nt GTTGTTGTTGTTATGGTCTAGTTGTTGTTGTTATGGTCTAATTT
Dharmacon Edit-R Lentiviral sgrna Pooled Screening Libraries
 GE Healthcare Dharmacon Edit-R Lentiviral sgrna Pooled Screening Libraries Contents 1. Introduction...3 A. CRISPR-Cas: An adaptive immunity defense mechanism in bacteria and archaea... 3 B. Engineering
GE Healthcare Dharmacon Edit-R Lentiviral sgrna Pooled Screening Libraries Contents 1. Introduction...3 A. CRISPR-Cas: An adaptive immunity defense mechanism in bacteria and archaea... 3 B. Engineering
Supplementary Information
 Supplementary Information Super-resolution imaging of fluorescently labeled, endogenous RNA Polymerase II in living cells with CRISPR/Cas9-mediated gene editing Won-Ki Cho 1, Namrata Jayanth 1, Susan Mullen
Supplementary Information Super-resolution imaging of fluorescently labeled, endogenous RNA Polymerase II in living cells with CRISPR/Cas9-mediated gene editing Won-Ki Cho 1, Namrata Jayanth 1, Susan Mullen
SUPPLEMENTARY INFORMATION
 doi:.38/nature899 Supplementary Figure Suzuki et al. a c p7 -/- / WT ratio (+)/(-) p7 -/- / WT ratio Log X 3. Fold change by treatment ( (+)/(-)) Log X.5 3-3. -. b Fold change by treatment ( (+)/(-)) 8
doi:.38/nature899 Supplementary Figure Suzuki et al. a c p7 -/- / WT ratio (+)/(-) p7 -/- / WT ratio Log X 3. Fold change by treatment ( (+)/(-)) Log X.5 3-3. -. b Fold change by treatment ( (+)/(-)) 8
Nature Methods: doi: /nmeth Supplementary Figure 1. Pilot CrY2H-seq experiments to confirm strain and plasmid functionality.
 Supplementary Figure 1 Pilot CrY2H-seq experiments to confirm strain and plasmid functionality. (a) RT-PCR on HIS3 positive diploid cell lysate containing known interaction partners AT3G62420 (bzip53)
Supplementary Figure 1 Pilot CrY2H-seq experiments to confirm strain and plasmid functionality. (a) RT-PCR on HIS3 positive diploid cell lysate containing known interaction partners AT3G62420 (bzip53)
embryos. Asterisk represents loss of or reduced expression. Brackets represent
 Supplemental Figures Supplemental Figure 1. tfec expression is highly enriched in tail endothelial cells (A- B) ISH of tfec at 15 and 16hpf in WT embryos. (C- D) ISH of tfec at 36 and 38hpf in WT embryos.
Supplemental Figures Supplemental Figure 1. tfec expression is highly enriched in tail endothelial cells (A- B) ISH of tfec at 15 and 16hpf in WT embryos. (C- D) ISH of tfec at 36 and 38hpf in WT embryos.
Revised: RG-RV2 by Fukuhara et al.
 Supplemental Figure 1 The generation of Spns2 conditional knockout mice. (A) Schematic representation of the wild type Spns2 locus (Spns2 + ), the targeted allele, the floxed allele (Spns2 f ) and the
Supplemental Figure 1 The generation of Spns2 conditional knockout mice. (A) Schematic representation of the wild type Spns2 locus (Spns2 + ), the targeted allele, the floxed allele (Spns2 f ) and the
Integrated Omics Study Delineates the Dynamics of Lipid Droplets in Rhodococcus Opacus PD630
 School of Natural Sciences and Mathematics 2013-10-22 Integrated Omics Study Delineates the Dynamics of Lipid Droplets in Rhodococcus Opacus PD630 UTD AUTHOR(S): Michael Qiwei Zhang 2013 The Authors This
School of Natural Sciences and Mathematics 2013-10-22 Integrated Omics Study Delineates the Dynamics of Lipid Droplets in Rhodococcus Opacus PD630 UTD AUTHOR(S): Michael Qiwei Zhang 2013 The Authors This
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION Supplementary figure 1: List of primers/oligonucleotides used in this study. 1 Supplementary figure 2: Sequences and mirna-targets of i) mcherry expresses in transgenic fish used
SUPPLEMENTARY INFORMATION Supplementary figure 1: List of primers/oligonucleotides used in this study. 1 Supplementary figure 2: Sequences and mirna-targets of i) mcherry expresses in transgenic fish used
Supplementary Figure 1. Phenotype and genotype of cultured and transplanted S1 KCST (A) Brightfield and mcherry fluorescence images of the spheres
 Supplementary Figure 1. Phenotype and genotype of cultured and transplanted S1 KCST (A) Brightfield and mcherry fluorescence images of the spheres generated from the CD133-positive cells infected with
Supplementary Figure 1. Phenotype and genotype of cultured and transplanted S1 KCST (A) Brightfield and mcherry fluorescence images of the spheres generated from the CD133-positive cells infected with
CloneTracker XP 10M Barcode-3 Library with RFP-Puro
 CloneTracker XP 10M Barcode-3 Library with RFP-Puro Shipment Contents: Store at -20 C Description: Cellecta s CloneTracker XP 10M Barcode-3 Library with RFP-Puro is a pooled barcode library that enables
CloneTracker XP 10M Barcode-3 Library with RFP-Puro Shipment Contents: Store at -20 C Description: Cellecta s CloneTracker XP 10M Barcode-3 Library with RFP-Puro is a pooled barcode library that enables
AD BD TOC1. Supplementary Figure 1: Yeast two-hybrid assays showing the interaction between
 AD X BD TOC1 AD BD X PIFΔAD PIF TOC1 TOC1 PIFΔAD PIF N TOC1 TOC1 C1 PIFΔAD PIF C1 TOC1 TOC1 C PIFΔAD PIF C TOC1 Supplementary Figure 1: Yeast two-hybrid assays showing the interaction between PIF and TOC1
AD X BD TOC1 AD BD X PIFΔAD PIF TOC1 TOC1 PIFΔAD PIF N TOC1 TOC1 C1 PIFΔAD PIF C1 TOC1 TOC1 C PIFΔAD PIF C TOC1 Supplementary Figure 1: Yeast two-hybrid assays showing the interaction between PIF and TOC1
Cleanup. Total Time 2.5 hr
 sparq DNA Library Prep Kit Cat. No. 95191-024 Size: 24 reactions Store at -25 C to -15 C 95191-096 96 reactions Description The sparq DNA Library Prep Kit provides components for the rapid construction
sparq DNA Library Prep Kit Cat. No. 95191-024 Size: 24 reactions Store at -25 C to -15 C 95191-096 96 reactions Description The sparq DNA Library Prep Kit provides components for the rapid construction
Spermatazoa. Sertoli cells. Morc1 WT adult. Morc1 KO adult. Morc1 WT P14.5. Morc1 KO P14.5. Germ cells. Germ cells. Spermatids.
 a. Spermatazoa Sertoli cells Morc1 WT adult c. Morc1 KO adult Morc1 WT P1.5 Morc1 KO P1.5 Morc1 WT adult Morc1 KO adult Germ cells Germ cells Spermatids Spermatozoa Supplementary Figure 1. Confirmation
a. Spermatazoa Sertoli cells Morc1 WT adult c. Morc1 KO adult Morc1 WT P1.5 Morc1 KO P1.5 Morc1 WT adult Morc1 KO adult Germ cells Germ cells Spermatids Spermatozoa Supplementary Figure 1. Confirmation
Supplementary Methods
 Supplementary Methods Reverse transcribed Quantitative PCR. Total RNA was isolated from bone marrow derived macrophages using RNeasy Mini Kit (Qiagen), DNase-treated (Promega RQ1), and reverse transcribed
Supplementary Methods Reverse transcribed Quantitative PCR. Total RNA was isolated from bone marrow derived macrophages using RNeasy Mini Kit (Qiagen), DNase-treated (Promega RQ1), and reverse transcribed
Supplementary Figure 1
 Supplementary Figure 1 Supplementary Fig. 1 shrna mediated knockdown of ZRSR2 in K562 and 293T cells. (a) ZRSR2 transcript levels in stably transduced K562 cells were determined using qrt-pcr. GAPDH was
Supplementary Figure 1 Supplementary Fig. 1 shrna mediated knockdown of ZRSR2 in K562 and 293T cells. (a) ZRSR2 transcript levels in stably transduced K562 cells were determined using qrt-pcr. GAPDH was
Nature Methods: doi: /nmeth Supplementary Figure 1. Construction of a sensitive TetR mediated auxotrophic off-switch.
 Supplementary Figure 1 Construction of a sensitive TetR mediated auxotrophic off-switch. A Production of the Tet repressor in yeast when conjugated to either the LexA4 or LexA8 promoter DNA binding sequences.
Supplementary Figure 1 Construction of a sensitive TetR mediated auxotrophic off-switch. A Production of the Tet repressor in yeast when conjugated to either the LexA4 or LexA8 promoter DNA binding sequences.
Supplementary Figure 1
 Supplementary Figure 1 Supplementary Figure 1: Vector maps of TRMPV and TRMPVIR variants. Many derivatives of TRMPV have been generated and tested. Unless otherwise noted, experiments in this paper use
Supplementary Figure 1 Supplementary Figure 1: Vector maps of TRMPV and TRMPVIR variants. Many derivatives of TRMPV have been generated and tested. Unless otherwise noted, experiments in this paper use
Donor DNA Utilization during Gene Targeting with Zinc- finger Nucleases
 Donor DNA Utilization during Gene Targeting with Zinc- finger Nucleases Kelly J. Beumer, Jonathan K. Trautman, Kusumika Mukherjee and Dana Carroll Department of Biochemistry University of Utah School of
Donor DNA Utilization during Gene Targeting with Zinc- finger Nucleases Kelly J. Beumer, Jonathan K. Trautman, Kusumika Mukherjee and Dana Carroll Department of Biochemistry University of Utah School of
Supplementary Figure 1.
 Supplementary Figure 1. (A) UCSC Genome Browser view of region immediately downstream of the NEUROG3 start codon. All candidate grna target sites which meet the G(N 19 )NGG constraint are aligned to illustrate
Supplementary Figure 1. (A) UCSC Genome Browser view of region immediately downstream of the NEUROG3 start codon. All candidate grna target sites which meet the G(N 19 )NGG constraint are aligned to illustrate
Supplemental Figure 1 A
 Supplemental Figure A prebleach postbleach 2 min 6 min 3 min mh2a.-gfp mh2a.2-gfp mh2a2-gfp GFP-H2A..9 Relative Intensity.8.7.6.5 mh2a. GFP n=8.4 mh2a.2 GFP n=4.3 mh2a2 GFP n=2.2 GFP H2A n=24. GFP n=7.
Supplemental Figure A prebleach postbleach 2 min 6 min 3 min mh2a.-gfp mh2a.2-gfp mh2a2-gfp GFP-H2A..9 Relative Intensity.8.7.6.5 mh2a. GFP n=8.4 mh2a.2 GFP n=4.3 mh2a2 GFP n=2.2 GFP H2A n=24. GFP n=7.
Nature Genetics: doi: /ng.3556 INTEGRATED SUPPLEMENTARY FIGURE TEMPLATE. Supplementary Figure 1
 INTEGRATED SUPPLEMENTARY FIGURE TEMPLATE Supplementary Figure 1 REF6 expression in transgenic lines. (a,b) Expression of REF6 in REF6-HA ref6 and REF6ΔZnF-HA ref6 plants detected by RT qpcr (a) and immunoblot
INTEGRATED SUPPLEMENTARY FIGURE TEMPLATE Supplementary Figure 1 REF6 expression in transgenic lines. (a,b) Expression of REF6 in REF6-HA ref6 and REF6ΔZnF-HA ref6 plants detected by RT qpcr (a) and immunoblot
Supplementary Information Targeting fidelity of adenine and cytosine base editors in mouse embryos
 Supplementary Information ing fidelity of adenine and cytosine base s in mouse embryos Lee et al. a P = 1.012e-14 b Frequency (%) 100% 80% 60% 40% 20% 0% CB AB On-target Bystander Proximal Indels Frequency
Supplementary Information ing fidelity of adenine and cytosine base s in mouse embryos Lee et al. a P = 1.012e-14 b Frequency (%) 100% 80% 60% 40% 20% 0% CB AB On-target Bystander Proximal Indels Frequency
Nature Methods: doi: /nmeth.4396
 Supplementary Figure 1 Comparison of technical replicate consistency between and across the standard ATAC-seq method, DNase-seq, and Omni-ATAC. (a) Heatmap-based representation of ATAC-seq quality control
Supplementary Figure 1 Comparison of technical replicate consistency between and across the standard ATAC-seq method, DNase-seq, and Omni-ATAC. (a) Heatmap-based representation of ATAC-seq quality control
Nature Genetics: doi: /ng Supplementary Figure 1
 Supplementary Figure 1 Ihh interacts preferentially with its upstream neighboring gene Nhej1. Genes are indicated by gray lines, and Ihh and Nhej1 are highlighted in blue. 4C seq performed in E14.5 limbs
Supplementary Figure 1 Ihh interacts preferentially with its upstream neighboring gene Nhej1. Genes are indicated by gray lines, and Ihh and Nhej1 are highlighted in blue. 4C seq performed in E14.5 limbs
Nature Biotechnology: doi: /nbt.4166
 Supplementary Figure 1 Validation of correct targeting at targeted locus. (a) by immunofluorescence staining of 2C-HR-CRISPR microinjected embryos cultured to the blastocyst stage. Embryos were stained
Supplementary Figure 1 Validation of correct targeting at targeted locus. (a) by immunofluorescence staining of 2C-HR-CRISPR microinjected embryos cultured to the blastocyst stage. Embryos were stained
b number of motif occurrences
 supplementary information DOI: 1.138/ncb194 a b number of motif occurrences number of motif occurrences number of motif occurrences I II III IV V X TTGGTCAGTGCA 3 8 23 4 13 239 TTGGTCAGTGCT 7 5 8 1 3 83
supplementary information DOI: 1.138/ncb194 a b number of motif occurrences number of motif occurrences number of motif occurrences I II III IV V X TTGGTCAGTGCA 3 8 23 4 13 239 TTGGTCAGTGCT 7 5 8 1 3 83
A CRISPR/Cas9 Vector System for Tissue-Specific Gene Disruption in Zebrafish
 Developmental Cell Supplemental Information A CRISPR/Cas9 Vector System for Tissue-Specific Gene Disruption in Zebrafish Julien Ablain, Ellen M. Durand, Song Yang, Yi Zhou, and Leonard I. Zon % larvae
Developmental Cell Supplemental Information A CRISPR/Cas9 Vector System for Tissue-Specific Gene Disruption in Zebrafish Julien Ablain, Ellen M. Durand, Song Yang, Yi Zhou, and Leonard I. Zon % larvae
Supplemental Data. Zhou et al. (2016). Plant Cell /tpc
 Supplemental Figure 1. Confirmation of mutant mapping results. (A) Complementation assay with stably transformed genomic fragments (ComN-N) (2 kb upstream of TSS and 1.5 kb downstream of TES) and CaMV
Supplemental Figure 1. Confirmation of mutant mapping results. (A) Complementation assay with stably transformed genomic fragments (ComN-N) (2 kb upstream of TSS and 1.5 kb downstream of TES) and CaMV
Supplementary Figure 1 Generation of migg1-yf mice. (A) Targeting strategy. Upper panel: schematic organization of the murine ɣ1 immunoglobulin
 Supplementary Figure 1 Generation of migg1-yf mice. (A) Targeting strategy. Upper panel: schematic organization of the murine ɣ1 immunoglobulin locus. The EcoRI restriction site between exons M1 and M2
Supplementary Figure 1 Generation of migg1-yf mice. (A) Targeting strategy. Upper panel: schematic organization of the murine ɣ1 immunoglobulin locus. The EcoRI restriction site between exons M1 and M2
Supplementary Information Design of small molecule-responsive micrornas based on structural requirements for Drosha processing
 Supplementary Information Design of small molecule-responsive micrornas based on structural requirements for Drosha processing Chase L. Beisel, Yvonne Y. Chen, Stephanie J. Culler, Kevin G. Hoff, & Christina
Supplementary Information Design of small molecule-responsive micrornas based on structural requirements for Drosha processing Chase L. Beisel, Yvonne Y. Chen, Stephanie J. Culler, Kevin G. Hoff, & Christina
Analyzing an individual sequence in the Sequence Editor
 BioNumerics Tutorial: Analyzing an individual sequence in the Sequence Editor 1 Aim The Sequence editor window is a convenient tool implemented in BioNumerics to edit and analyze nucleotide and amino acid
BioNumerics Tutorial: Analyzing an individual sequence in the Sequence Editor 1 Aim The Sequence editor window is a convenient tool implemented in BioNumerics to edit and analyze nucleotide and amino acid
Nature Structural and Molecular Biology: doi: /nsmb Supplementary Figure 1. Validation of CDK9-inhibitor treatment.
 Supplementary Figure 1 Validation of CDK9-inhibitor treatment. (a) Schematic of GAPDH with the middle of the amplicons indicated in base pairs. The transcription start site (TSS) and the terminal polyadenylation
Supplementary Figure 1 Validation of CDK9-inhibitor treatment. (a) Schematic of GAPDH with the middle of the amplicons indicated in base pairs. The transcription start site (TSS) and the terminal polyadenylation
SUPPLEMENTARY FIGURES AND TABLES
 SUPPLEMENTARY FIGURES AND TABLES Suppl. Table 1 Oligonucleotides used in the study The table shows the structure and nucleotide position of the oligonucleotides used in the study along with their application
SUPPLEMENTARY FIGURES AND TABLES Suppl. Table 1 Oligonucleotides used in the study The table shows the structure and nucleotide position of the oligonucleotides used in the study along with their application
Bootcamp: Molecular Biology Techniques and Interpretation
 Bootcamp: Molecular Biology Techniques and Interpretation Bi8 Winter 2016 Today s outline Detecting and quantifying nucleic acids and proteins: Basic nucleic acid properties Hybridization PCR and Designing
Bootcamp: Molecular Biology Techniques and Interpretation Bi8 Winter 2016 Today s outline Detecting and quantifying nucleic acids and proteins: Basic nucleic acid properties Hybridization PCR and Designing
Supplemental Information
 Supplemental Information Itemized List Materials and Methods, Related to Supplemental Figures S5A-C and S6. Supplemental Figure S1, Related to Figures 1 and 2. Supplemental Figure S2, Related to Figure
Supplemental Information Itemized List Materials and Methods, Related to Supplemental Figures S5A-C and S6. Supplemental Figure S1, Related to Figures 1 and 2. Supplemental Figure S2, Related to Figure
Mapping long-range promoter contacts in human cells with high-resolution capture Hi-C
 CORRECTION NOTICE Nat. Genet. 47, 598 606 (2015) Mapping long-range promoter contacts in human cells with high-resolution capture Hi-C Borbala Mifsud, Filipe Tavares-Cadete, Alice N Young, Robert Sugar,
CORRECTION NOTICE Nat. Genet. 47, 598 606 (2015) Mapping long-range promoter contacts in human cells with high-resolution capture Hi-C Borbala Mifsud, Filipe Tavares-Cadete, Alice N Young, Robert Sugar,
Supplementary Fig. S1. SAMHD1c has a more potent dntpase activity than. SAMHD1c. Purified recombinant SAMHD1c and SAMHD1c proteins (with
 Supplementary Fig. S1. SAMHD1c has a more potent dntpase activity than SAMHD1c. Purified recombinant SAMHD1c and SAMHD1c proteins (with concentration of 800nM) were incubated with 1mM dgtp for the indicated
Supplementary Fig. S1. SAMHD1c has a more potent dntpase activity than SAMHD1c. Purified recombinant SAMHD1c and SAMHD1c proteins (with concentration of 800nM) were incubated with 1mM dgtp for the indicated
Nature Genetics: doi: /ng.2949
 Supplementary Figure 1. Abnormal differentiation in vivo and colony growth in vitro of B cells with triplication of chr.21 orthologs. (a) (Top) B220 and CD43 staining of bone marrow from Ts1Rhr and wild-type
Supplementary Figure 1. Abnormal differentiation in vivo and colony growth in vitro of B cells with triplication of chr.21 orthologs. (a) (Top) B220 and CD43 staining of bone marrow from Ts1Rhr and wild-type
Rassf1a -/- Sav1 +/- mice. Supplementary Figures:
 1 Supplementary Figures: Figure S1: Genotyping of Rassf1a and Sav1 mutant mice. A. Rassf1a knockout mice lack exon 1 of the Rassf1 gene. A diagram and typical PCR genotyping of wildtype and Rassf1a -/-
1 Supplementary Figures: Figure S1: Genotyping of Rassf1a and Sav1 mutant mice. A. Rassf1a knockout mice lack exon 1 of the Rassf1 gene. A diagram and typical PCR genotyping of wildtype and Rassf1a -/-
Allele-specific locus binding and genome editing by CRISPR at the
 Supplementary Information Allele-specific locus binding and genome editing by CRISPR at the p6ink4a locus Toshitsugu Fujita, Miyuki Yuno, and Hodaka Fujii Supplementary Figure Legends Supplementary Figure
Supplementary Information Allele-specific locus binding and genome editing by CRISPR at the p6ink4a locus Toshitsugu Fujita, Miyuki Yuno, and Hodaka Fujii Supplementary Figure Legends Supplementary Figure
Supplementary Figure S1. Immunodetection of full-length XA21 and the XA21 C-terminal cleavage product.
 Supplementary Information Supplementary Figure S1. Immunodetection of full-length XA21 and the XA21 C-terminal cleavage product. Total protein extracted from Kitaake wild type and rice plants carrying
Supplementary Information Supplementary Figure S1. Immunodetection of full-length XA21 and the XA21 C-terminal cleavage product. Total protein extracted from Kitaake wild type and rice plants carrying
Somatic Primary pirna Biogenesis Driven by cis-acting RNA Elements and Trans-Acting Yb
 Cell Reports Supplemental Information Somatic Primary pirna Biogenesis Driven by cis-acting RNA Elements and Trans-Acting Yb Hirotsugu Ishizu, Yuka W. Iwasaki, Shigeki Hirakata, Haruka Ozaki, Wataru Iwasaki,
Cell Reports Supplemental Information Somatic Primary pirna Biogenesis Driven by cis-acting RNA Elements and Trans-Acting Yb Hirotsugu Ishizu, Yuka W. Iwasaki, Shigeki Hirakata, Haruka Ozaki, Wataru Iwasaki,
Supplemental Table/Figure Legends
 MiR-26a is required for skeletal muscle differentiation and regeneration in mice Bijan K. Dey, Jeffrey Gagan, Zhen Yan #, Anindya Dutta Supplemental Table/Figure Legends Suppl. Table 1: Effect of overexpression
MiR-26a is required for skeletal muscle differentiation and regeneration in mice Bijan K. Dey, Jeffrey Gagan, Zhen Yan #, Anindya Dutta Supplemental Table/Figure Legends Suppl. Table 1: Effect of overexpression
High-resolution Identification and Separation of Living Cell Types by Multiple microrna-responsive Synthetic mrnas
 Supplementary information High-resolution Identification and Separation of Living Cell Types by Multiple microrna-responsive Synthetic mrnas Kei Endo *, Karin Hayashi, Hirohide Saito * Contents: 1. Supplementary
Supplementary information High-resolution Identification and Separation of Living Cell Types by Multiple microrna-responsive Synthetic mrnas Kei Endo *, Karin Hayashi, Hirohide Saito * Contents: 1. Supplementary
SUPPLEMENTARY FIGURES
 SUPPLEMENTARY FIGURES Supplementary Figure 1 Histogram displaying the distribution of DNA barcode copy numbers for each virus library. 100,000 BB88 cells were infected by lentivirus that carried the corresponding
SUPPLEMENTARY FIGURES Supplementary Figure 1 Histogram displaying the distribution of DNA barcode copy numbers for each virus library. 100,000 BB88 cells were infected by lentivirus that carried the corresponding
SUPPLEMENTARY INFORMATION
 Gene replacements and insertions in rice by intron targeting using CRISPR Cas9 Table of Contents Supplementary Figure 1. sgrna-induced targeted mutations in the OsEPSPS gene in rice protoplasts. Supplementary
Gene replacements and insertions in rice by intron targeting using CRISPR Cas9 Table of Contents Supplementary Figure 1. sgrna-induced targeted mutations in the OsEPSPS gene in rice protoplasts. Supplementary
Nature Biotechnology: doi: /nbt Supplementary Figure 1. In vitro validation of OTC sgrnas and donor template.
 Supplementary Figure 1 In vitro validation of OTC sgrnas and donor template. (a) In vitro validation of sgrnas targeted to OTC in the MC57G mouse cell line by transient transfection followed by 4-day puromycin
Supplementary Figure 1 In vitro validation of OTC sgrnas and donor template. (a) In vitro validation of sgrnas targeted to OTC in the MC57G mouse cell line by transient transfection followed by 4-day puromycin
Nature Biotechnology: doi: /nbt Supplementary Figure 1
 Supplementary Figure 1 Schematic and results of screening the combinatorial antibody library for Sox2 replacement activity. A single batch of MEFs were plated and transduced with doxycycline inducible
Supplementary Figure 1 Schematic and results of screening the combinatorial antibody library for Sox2 replacement activity. A single batch of MEFs were plated and transduced with doxycycline inducible
A Guide to CRISPR/Cas9
 Genome editing and beyond freepik A Guide to CRISPR/Cas9 The latest advance in genomic DNA editing is the Clustered Regularly Interspaced Short Palindromic Repeat (CRISPR)/Cas9 system. This simple-touse
Genome editing and beyond freepik A Guide to CRISPR/Cas9 The latest advance in genomic DNA editing is the Clustered Regularly Interspaced Short Palindromic Repeat (CRISPR)/Cas9 system. This simple-touse
Supplementary Figure 1. Nature Structural & Molecular Biology: doi: /nsmb.3494
 Supplementary Figure 1 Pol structure-function analysis (a) Inactivating polymerase and helicase mutations do not alter the stability of Pol. Flag epitopes were introduced using CRISPR/Cas9 gene targeting
Supplementary Figure 1 Pol structure-function analysis (a) Inactivating polymerase and helicase mutations do not alter the stability of Pol. Flag epitopes were introduced using CRISPR/Cas9 gene targeting
Chemical and/or enzymatic deamination across various sequencing methods to localize modified cytosines.
 Supplementary Figure 1 Chemical and/or enzymatic deamination across various sequencing methods to localize modified cytosines. (a) Upon treatment with bisulfite under acidic conditions and at elevated
Supplementary Figure 1 Chemical and/or enzymatic deamination across various sequencing methods to localize modified cytosines. (a) Upon treatment with bisulfite under acidic conditions and at elevated
Cell line CDH1 mrna AS-paRNA S-paRNA
 a b Cell line mrna AS-paRNA S-paRNA HCT116 P53 +/+ 39.47 + + HCT116 P53 -/- 35.36 + + LNCAP 1562.4 + - MCF7 C1 1864.76 + - MCF7 C2 1812.81 + - Supplementary Figure 1. Promoter-associated transcripts at
a b Cell line mrna AS-paRNA S-paRNA HCT116 P53 +/+ 39.47 + + HCT116 P53 -/- 35.36 + + LNCAP 1562.4 + - MCF7 C1 1864.76 + - MCF7 C2 1812.81 + - Supplementary Figure 1. Promoter-associated transcripts at
Molecular Biology: DNA sequencing
 Molecular Biology: DNA sequencing Author: Prof Marinda Oosthuizen Licensed under a Creative Commons Attribution license. SEQUENCING OF LARGE TEMPLATES As we have seen, we can obtain up to 800 nucleotides
Molecular Biology: DNA sequencing Author: Prof Marinda Oosthuizen Licensed under a Creative Commons Attribution license. SEQUENCING OF LARGE TEMPLATES As we have seen, we can obtain up to 800 nucleotides
Supplementary Figure 1
 Supplementary Figure 1 Product purity of cytosine base editing for the wheat genomic loci tested. Product distributions and indels frequencies at four representative wheat genomic DNA sites in wheat protoplasts
Supplementary Figure 1 Product purity of cytosine base editing for the wheat genomic loci tested. Product distributions and indels frequencies at four representative wheat genomic DNA sites in wheat protoplasts
Non-coding Function & Variation, MPRAs II. Mike White Bio /5/18
 Non-coding Function & Variation, MPRAs II Mike White Bio 5488 3/5/18 MPRA Review Problem 1: Where does your CRE DNA come from? DNA synthesis Genomic fragments Targeted regulome capture Problem 2: How do
Non-coding Function & Variation, MPRAs II Mike White Bio 5488 3/5/18 MPRA Review Problem 1: Where does your CRE DNA come from? DNA synthesis Genomic fragments Targeted regulome capture Problem 2: How do
Supplementary Figure 1 qrt-pcr expression analysis of NLP8 with and without KNO 3 during germination.
 Supplementary Figure 1 qrt-pcr expression analysis of NLP8 with and without KNO 3 during germination. Seeds of Col-0 were harvested from plants grown at 16 C, stored for 2 months, imbibed for indicated
Supplementary Figure 1 qrt-pcr expression analysis of NLP8 with and without KNO 3 during germination. Seeds of Col-0 were harvested from plants grown at 16 C, stored for 2 months, imbibed for indicated
Supplementary Figure S1. The tetracycline-inducible CRISPR system. A) Hela cells stably
 Supplementary Information Supplementary Figure S1. The tetracycline-inducible CRISPR system. A) Hela cells stably expressing shrna sequences against TRF2 were examined by western blotting. shcon, shrna
Supplementary Information Supplementary Figure S1. The tetracycline-inducible CRISPR system. A) Hela cells stably expressing shrna sequences against TRF2 were examined by western blotting. shcon, shrna
1. Primers for PCR to amplify hairpin stem-loop precursor mir-145 plus different flanking sequence from human genomic DNA.
 Supplemental data: 1. Primers for PCR to amplify hairpin stem-loop precursor mir-145 plus different flanking sequence from human genomic DNA. Strategy#1: 20nt at both sides: #1_BglII-Fd primer : 5 -gga
Supplemental data: 1. Primers for PCR to amplify hairpin stem-loop precursor mir-145 plus different flanking sequence from human genomic DNA. Strategy#1: 20nt at both sides: #1_BglII-Fd primer : 5 -gga
Supplementary Figure 1. NORAD expression in mouse (A) and dog (B). The black boxes indicate the position of the regions alignable to the 12 repeat
 Supplementary Figure 1. NORAD expression in mouse (A) and dog (B). The black boxes indicate the position of the regions alignable to the 12 repeat units in the human genome. Annotated transposable elements
Supplementary Figure 1. NORAD expression in mouse (A) and dog (B). The black boxes indicate the position of the regions alignable to the 12 repeat units in the human genome. Annotated transposable elements
Supplemental Data. Na Xu et al. (2016). Plant Cell /tpc
 Supplemental Figure 1. The weak fluorescence phenotype is not caused by the mutation in At3g60240. (A) A mutation mapped to the gene At3g60240. Map-based cloning strategy was used to map the mutated site
Supplemental Figure 1. The weak fluorescence phenotype is not caused by the mutation in At3g60240. (A) A mutation mapped to the gene At3g60240. Map-based cloning strategy was used to map the mutated site
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature12119 SUPPLEMENTARY FIGURES AND LEGENDS pre-let-7a- 1 +14U pre-let-7a- 1 Ddx3x Dhx30 Dis3l2 Elavl1 Ggt5 Hnrnph 2 Osbpl5 Puf60 Rnpc3 Rpl7 Sf3b3 Sf3b4 Tia1 Triobp U2af1 U2af2 1 6 2 4 3
doi:10.1038/nature12119 SUPPLEMENTARY FIGURES AND LEGENDS pre-let-7a- 1 +14U pre-let-7a- 1 Ddx3x Dhx30 Dis3l2 Elavl1 Ggt5 Hnrnph 2 Osbpl5 Puf60 Rnpc3 Rpl7 Sf3b3 Sf3b4 Tia1 Triobp U2af1 U2af2 1 6 2 4 3
TECH NOTE Pushing the Limit: A Complete Solution for Generating Stranded RNA Seq Libraries from Picogram Inputs of Total Mammalian RNA
 TECH NOTE Pushing the Limit: A Complete Solution for Generating Stranded RNA Seq Libraries from Picogram Inputs of Total Mammalian RNA Stranded, Illumina ready library construction in
TECH NOTE Pushing the Limit: A Complete Solution for Generating Stranded RNA Seq Libraries from Picogram Inputs of Total Mammalian RNA Stranded, Illumina ready library construction in
Tutorial. In Silico Cloning. Sample to Insight. March 31, 2016
 In Silico Cloning March 31, 2016 Sample to Insight CLC bio, a QIAGEN Company Silkeborgvej 2 Prismet 8000 Aarhus C Denmark Telephone: +45 70 22 32 44 www.clcbio.com support-clcbio@qiagen.com In Silico Cloning
In Silico Cloning March 31, 2016 Sample to Insight CLC bio, a QIAGEN Company Silkeborgvej 2 Prismet 8000 Aarhus C Denmark Telephone: +45 70 22 32 44 www.clcbio.com support-clcbio@qiagen.com In Silico Cloning
Construct Design and Cloning Guide for Cas9-triggered homologous recombination
 Construct Design and Cloning Guide for Cas9-triggered homologous recombination Written by Dan Dickinson (ddickins@live.unc.edu) and last updated December 2013. Reference: Dickinson DJ, Ward JD, Reiner
Construct Design and Cloning Guide for Cas9-triggered homologous recombination Written by Dan Dickinson (ddickins@live.unc.edu) and last updated December 2013. Reference: Dickinson DJ, Ward JD, Reiner
Supplemental Figure 1.
 Supplemental Data. Charron et al. Dynamic landscapes of four histone modifications during de-etiolation in Arabidopsis. Plant Cell (2009). 10.1105/tpc.109.066845 Supplemental Figure 1. Immunodetection
Supplemental Data. Charron et al. Dynamic landscapes of four histone modifications during de-etiolation in Arabidopsis. Plant Cell (2009). 10.1105/tpc.109.066845 Supplemental Figure 1. Immunodetection
Fig. S1. Clustering analysis of expression array and ChIP-PCR assay in the ARF3 locus. (A) Typical examples of the transgenic plants used for
 Fig. S1. Clustering analysis of expression array and ChIP-PCR assay in the ARF3 locus. (A) Typical examples of the transgenic plants used for ChIP-chip and ChIP-PCR assays. The presence of pas1:t7:as1
Fig. S1. Clustering analysis of expression array and ChIP-PCR assay in the ARF3 locus. (A) Typical examples of the transgenic plants used for ChIP-chip and ChIP-PCR assays. The presence of pas1:t7:as1
CRISPR-dCas9 mediated TET1 targeting for selective DNA demethylation at BRCA1 promoter
 CRISPR-dCas9 mediated TET1 targeting for selective DNA demethylation at BRCA1 promoter SUPPLEMENTARY DATA See Supplementary Sequence File: 1 Supplementary Figure S1: The total protein was extracted from
CRISPR-dCas9 mediated TET1 targeting for selective DNA demethylation at BRCA1 promoter SUPPLEMENTARY DATA See Supplementary Sequence File: 1 Supplementary Figure S1: The total protein was extracted from
Chinese Academy of Sciences, Datun Road, Chaoyang District, Beijing, 10101, China
 SUPPLEMENTARY INFORMATION Title: Bi-directional processing of pri-mirnas with branched terminal loops by Arabidopsis Dicer-like1 Hongliang Zhu 1,2,3, Yuyi Zhou 1,2,4, Claudia Castillo-González 1,2, Amber
SUPPLEMENTARY INFORMATION Title: Bi-directional processing of pri-mirnas with branched terminal loops by Arabidopsis Dicer-like1 Hongliang Zhu 1,2,3, Yuyi Zhou 1,2,4, Claudia Castillo-González 1,2, Amber
