Heme utilization in the Caenorhabditis elegans hypodermal cells is facilitated by hemeresponsive
|
|
|
- Matilda Harrison
- 6 years ago
- Views:
Transcription
1 Supplemental Data Heme utilization in the Caenorhabditis elegans hypodermal cells is facilitated by hemeresponsive gene-2 Caiyong Chen 1, Tamika K. Samuel 1, Michael Krause 2, Harry A. Dailey 3, and Iqbal Hamza 1 1 From the Department of Animal & Avian Sciences and Department of Cell Biology & Molecular Genetics, University of Maryland, College Park, Maryland Laboratory of Molecular Biology, National Institute of Diabetes and Digestive and Kidney Diseases, National Institutes of Health, Bethesda, Maryland Biomedical and Health Sciences Institute, Department of Microbiology and the Department of Biochemistry and Molecular Biology, University of Georgia, Athens, Georgia Table of Contents S-1 Cover page S-2 Supplemental Methods S-3 Table S1 S-4 Table S2 S-5 Figure S1 S-6 Figure S2 S-7 Figure S3 S-8 Figure S4 S-9 Figure S5 S-10 Figure S6 S-11 Figure S7 S-1
2 Supplemental Method: Microarray Analysis C. elegans whole genome expression array chips (Affymetrix) were probed and the data analyzed using both MAS5.0 and Robust Multichip Analysis (RMA). For MAS5.0 analysis, a pseudochip was created from the three wildtype broodmate controls and used as a baseline file to which each of the experimental hrg-2 mutant samples were compared. For both heme concentration growth conditions (4 µm and 20 µm), the MAS5.0 detection and change in expression calls were used to narrow the list of genes of interest. For 4 µm heme, a comparison between hrg-2 mutants and the wildtype control resulted in 1,050 probe sets with decreased and 49 probe sets with increased expression across all three biological replicates. For 20 µm heme, this same comparison resulted in 273 probe sets with decreased and 35 probe sets with increased expression across all replicates. All probe sets resulting in altered expression across all three biological replicates, the most stringent condition, were subjected to Gene Ontology (GO) analysis using the DAVID functional annotation tools ( That analysis was most informative for the downregulated genes in 4 µm heme for hrg-2 mutants. Biological processes highly overrepresented in that dataset were cuticle collagen/molting (enrichment score 7.01), locomotion/larval growth (enrichment score 6.31), growth (enrichment score 4.99), hexose metabolism (enrichment score 2.78), fatty acid/lipid biosynthesis (enrichment score 2.48), and ER/cellular stress (enrichment score 2.47). To identify all Affymetrix array probe sets related to genes encoding cytochromes, we searched the C. elegans Wormbase genome database (WS195) for PFAM motifs related to cytochrome c (PF00034) and p450 (PF00067) and recovered 81 Affymetrix probe sets representing 75 genes. Among these probe sets at 4 µm heme, a comparison between hrg-2 mutants and the wildtype control resulted in one gene (cyp-37b1) that was consistently downregulated and no genes upregulated across all three biological replicates. For 20 µm heme, this same comparison resulted in three genes downregulated (cyp-25a1, cyp-29a3, cyp-36a1) and no genes upregulated (supplemental Fig S7). S-2
3 Supplemental Table S1. Transgenic C. elegans strains used in this study Number of Strain Genotype Method analyzed lines IQ8021 hrg-2 1.5kb ::gfp, rol-6+ microinjection 7 IQ8023 hrg-2 0.5kb ::gfp, rol-6+ microinjection 2 unc-119(ed3) III; hrg-2 1.5kb ::hrg-2::yfp, microparticle IQ unc-119+ bombardment IQ : TRAM: translocating chain associated membrane protein. unc-119(ed3) III; hrg-2 1.5kb ::hrg-2::yfp, microparticle dpy-7::mcherry::tram a, unc-119+ bombardment S-3
4 Supplemental Table S2. Oxygen consumption rates of transformed wildtype W303 and hem1δ yeast grown in 5 μm heme. Strains μm/min/mg Wildtype vector ± 1.05 hem1δ vector 2.52 ± 0.25 hem1δ cdr ± 0.23 ns hem1δ hrg ± 0.11 *** hem1δ hrg ± 0.37 *** ns: not significant when compared to hem1δ vector ***: P<0.001 compared to hem1δ vector S-4
5 Supplemental Figure S1. Chen et al., Supplemental FIGURE S1. Multiple sequence alignment of HRG-2 proteins among C. elegans, C. briggsae, and C. remanei. C. briggsae has two putative HRG-2 orthologs, of which the candidate with higher homology (WormBase protein ID CBP20711) is shown in the alignment. The protein ID for C. remanei HRG-2 is RP Transmembrane domains (TMD) are marked with gray filled boxes. GST-like domains were drawn based on the amino acid positions in C. elegans HRG-2 and are marked with unfilled boxes. The numbers at the end of alignment indicate the percentage identity between C. elegans HRG-2 and its homologs. S-5
6 Supplemental Figure S2. Chen et al., Supplemental FIGURE S2. Evolutionary relationships of HRG-2, CDRs, and their homologs in nematodes. Protein sequences were aligned using the ClustalW program, and the phylogenetic tree was constructed with the neighbor-joining method in MEGA 4. The branch lengths of the tree reflect the evolutionary distances, which are in the units of the number of amino acid substitutions. The scale bar represents 10% sequencee divergence. HRG-2 and CDRs in C. elegans are marked with a box or asterisks. Cel: C. elegans. CBP: C. briggsae. CN: C. brenneri. RP: C. remanei. PP: P. pacificus. S-6
7 Supplemental Figure S3. Chen et al., Supplemental FIGURE S3. Multiple sequence alignment of HRG-2 and the putative homolgs in vertebrate species. The accession numbers of the putative homologs are NP_ (human), NP (mouse), XP_ (chick), and XP_ (zebrafish). The numbers at the end of alignment indicate the percentage identities between C. elegans HRG-2 and its homologs. S-7
8 Supplemental Figure S4. Chen et al., Supplemental FIGURE S4. The 0.5-kb intergenic region is responsible for the spatial expression and heme response of hrg-2. A, Schematic representation of the hrg-2::gfp 0.5 reporter construct. Genomic structures of hrg-2 and cdr-7 are shown on the top. NLS: nuclear localization signal. B, hrg-2::gfpattern is similar to that of hrg-2::gfp 1.5. Arrows indicate GFP expression in hypodermal cells. C, 0.5 is predominantly expressed in hypodermal cells in C. elegans. This expression The hrg-2::gfp 0.5 reporter is not expressed when the worms were exposed to 20 µm heme. Asterisk indicates the non-specific autofluorescence of gut granules. Scale bars, 20 µm. S-8
9 Supplemental Figure S5. Chen et al., Supplemental FIGURE S5. Immunofluorescence microscopy of HEK-293 cells expressing HRG-2. Cells co-transfected with HA-tagged HRG-2 and the mitochondrial marker mitoyfp were subjected to epifluorescence (top) or confocal (bottom) microscopy using anti-ha primary antibody and an Alexa 568 conjugated secondary antibody. Scale bar indicates 10 μm. S-9
10 Supplemental Figure S6. Chen et al., Supplemental FIGURE S6. hrg-2 deletion does not affect the growth of C. elegans. Both hrg-2 (tm3798) and its wildtype broodmate worms were first grown at 2 μm heme for one generation in mcehr-2 medium. Synchronized L1 larvae were inoculated into the liquid medium containing 0, 4, and 20 μm heme. After 9 days of growth, worms in all treatments were counted, and the numbers were normalized to the actual input (0 μm).. The experiment was performed in duplicate. A total of 268 wildtype and 206 hrg-2 worms were scored for the number of their progeny at each heme concentration. No statistical difference was observed between wildtype and hrg-2 animals (P >0.05). S-10
11 Supplemental Figure S7. Chen et al., Normalized Fold Change cyp 25A1 cyp 29A3 cyp 36A1 cyp 37B1 4 μm Heme 20 μm Heme Supplemental FIGURE S7. Changes in cytochrome gene expression in hrg-2 mutants. Microarrays performed on hrg-2 and wildtype broodmate controls showed that one gene at 4 µm heme (cyp-37b1) and three genes at 20 µm heme (cyp-25a1, cyp-29a3, cyp-36a1) were downregulated from the 75 cytochrome encoding genes represented on the expression arrays Consistent changes in expression for these four genes were observed across all three biological replicates. S-11
Supplementary Figure S1. Immunodetection of full-length XA21 and the XA21 C-terminal cleavage product.
 Supplementary Information Supplementary Figure S1. Immunodetection of full-length XA21 and the XA21 C-terminal cleavage product. Total protein extracted from Kitaake wild type and rice plants carrying
Supplementary Information Supplementary Figure S1. Immunodetection of full-length XA21 and the XA21 C-terminal cleavage product. Total protein extracted from Kitaake wild type and rice plants carrying
SUPPLEMENTARY INFORMATION
 Figure S1. lev-9 mutants are resistant to levamisole. The levamisole dose-response curves indicate that lev-9 mutants are partially resistant to levamisole similar to lev-10(kr26) mutants. unc-29(x29)
Figure S1. lev-9 mutants are resistant to levamisole. The levamisole dose-response curves indicate that lev-9 mutants are partially resistant to levamisole similar to lev-10(kr26) mutants. unc-29(x29)
A Nucleus-Encoded Chloroplast Protein YL1 Is Involved in Chloroplast. Development and Efficient Biogenesis of Chloroplast ATP Synthase in Rice
 A Nucleus-Encoded Chloroplast Protein YL1 Is Involved in Chloroplast Development and Efficient Biogenesis of Chloroplast ATP Synthase in Rice Fei Chen 1,*, Guojun Dong 2,*, Limin Wu 1, Fang Wang 3, Xingzheng
A Nucleus-Encoded Chloroplast Protein YL1 Is Involved in Chloroplast Development and Efficient Biogenesis of Chloroplast ATP Synthase in Rice Fei Chen 1,*, Guojun Dong 2,*, Limin Wu 1, Fang Wang 3, Xingzheng
pmyrsaur78 psosein4+ psosetr1+ psosetr2+
 SAUR77 SAUR78 SAUR76 Figure S1 Phylogenetic analysis of Arabidopsis SAUR proteins. The analyses were conducted in MEGA5, using the Neighbor-Joining method. The percentage of replicate trees in which the
SAUR77 SAUR78 SAUR76 Figure S1 Phylogenetic analysis of Arabidopsis SAUR proteins. The analyses were conducted in MEGA5, using the Neighbor-Joining method. The percentage of replicate trees in which the
(A) Schematic depicting morphology of cholinergic DA and DB motor neurons in an L1
 SUPPLEMENTAL MATERIALS Supplemental Figure 1 Figure S1. Expression of the non-alpha nachr subunit ACR-2. (A) Schematic depicting morphology of cholinergic DA and DB motor neurons in an L1 animal. Wide-field
SUPPLEMENTAL MATERIALS Supplemental Figure 1 Figure S1. Expression of the non-alpha nachr subunit ACR-2. (A) Schematic depicting morphology of cholinergic DA and DB motor neurons in an L1 animal. Wide-field
Figure S1. Figure S2. Figure S3 HB Anti-FSP27 (COOH-terminal peptide) Ab. Anti-GST-FSP27(45-127) Ab.
 / 36B4 mrna ratio Figure S1 * 2. 1.6 1.2.8 *.4 control TNFα BRL49653 Figure S2 Su bw AT p iw Anti- (COOH-terminal peptide) Ab Blot : Anti-GST-(45-127) Ab β-actin Figure S3 HB2 HW AT BA T Figure S4 A TAG
/ 36B4 mrna ratio Figure S1 * 2. 1.6 1.2.8 *.4 control TNFα BRL49653 Figure S2 Su bw AT p iw Anti- (COOH-terminal peptide) Ab Blot : Anti-GST-(45-127) Ab β-actin Figure S3 HB2 HW AT BA T Figure S4 A TAG
Supporting Online Material for
 www.sciencemag.org/cgi/content/full/1119481/dc1 Supporting Online Material for LIN-12/Notch Activation Leads to MicroRNA-Mediated Down-Regulation of Vav in C. elegans Andrew S. Yoo and Iva Greenwald* *To
www.sciencemag.org/cgi/content/full/1119481/dc1 Supporting Online Material for LIN-12/Notch Activation Leads to MicroRNA-Mediated Down-Regulation of Vav in C. elegans Andrew S. Yoo and Iva Greenwald* *To
An antibiotic selection marker for nematode transgenesis
 nature methods An antibiotic selection marker for nematode transgenesis Rosina Giordano-Santini, Stuart Milstein, Nenad Svrzikapa, Domena Tu, Robert Johnsen, David Baillie, Marc Vidal & Denis Dupuy Supplementary
nature methods An antibiotic selection marker for nematode transgenesis Rosina Giordano-Santini, Stuart Milstein, Nenad Svrzikapa, Domena Tu, Robert Johnsen, David Baillie, Marc Vidal & Denis Dupuy Supplementary
Supplemental Figure 1 Human REEP family of proteins can be divided into two distinct subfamilies. Residues (single letter amino acid code) identical
 Supplemental Figure Human REEP family of proteins can be divided into two distinct subfamilies. Residues (single letter amino acid code) identical in all six REEPs are highlighted in green. Additional
Supplemental Figure Human REEP family of proteins can be divided into two distinct subfamilies. Residues (single letter amino acid code) identical in all six REEPs are highlighted in green. Additional
Nature Genetics: doi: /ng.3556 INTEGRATED SUPPLEMENTARY FIGURE TEMPLATE. Supplementary Figure 1
 INTEGRATED SUPPLEMENTARY FIGURE TEMPLATE Supplementary Figure 1 REF6 expression in transgenic lines. (a,b) Expression of REF6 in REF6-HA ref6 and REF6ΔZnF-HA ref6 plants detected by RT qpcr (a) and immunoblot
INTEGRATED SUPPLEMENTARY FIGURE TEMPLATE Supplementary Figure 1 REF6 expression in transgenic lines. (a,b) Expression of REF6 in REF6-HA ref6 and REF6ΔZnF-HA ref6 plants detected by RT qpcr (a) and immunoblot
Supplemental Data. Farmer et al. (2010) Plant Cell /tpc
 Supplemental Figure 1. Amino acid sequence comparison of RAD23 proteins. Identical and similar residues are shown in the black and gray boxes, respectively. Dots denote gaps. The sequence of plant Ub is
Supplemental Figure 1. Amino acid sequence comparison of RAD23 proteins. Identical and similar residues are shown in the black and gray boxes, respectively. Dots denote gaps. The sequence of plant Ub is
T H E J O U R N A L O F C E L L B I O L O G Y
 Supplemental material Wang et al., http://www.jcb.org/cgi/content/full/jcb.201405026/dc1 T H E J O U R N A L O F C E L L B I O L O G Y Figure S1. Generation and characterization of unc-40 alleles. (A and
Supplemental material Wang et al., http://www.jcb.org/cgi/content/full/jcb.201405026/dc1 T H E J O U R N A L O F C E L L B I O L O G Y Figure S1. Generation and characterization of unc-40 alleles. (A and
Liu et al., http :// /cgi /content /full /jcb /DC1
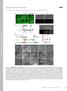 Supplemental material JCB Liu et al., http ://www.jcb.org /cgi /content /full /jcb.201506081 /DC1 THE JOU RNAL OF CELL BIO LOGY Figure S1. Identification of sorf-1 and sorf-2. (A) RNAi of ZK563.5 increases
Supplemental material JCB Liu et al., http ://www.jcb.org /cgi /content /full /jcb.201506081 /DC1 THE JOU RNAL OF CELL BIO LOGY Figure S1. Identification of sorf-1 and sorf-2. (A) RNAi of ZK563.5 increases
Two classes of silencing RNAs move between Caenorhabditis elegans tissues.
 Two classes of silencing RNAs move between Caenorhabditis elegans tissues. Antony M Jose, Giancarlo A Garcia, and Craig P Hunter. Supplementary Figures, Figure Legends, and Tables. Supplementary Figure
Two classes of silencing RNAs move between Caenorhabditis elegans tissues. Antony M Jose, Giancarlo A Garcia, and Craig P Hunter. Supplementary Figures, Figure Legends, and Tables. Supplementary Figure
Figure S1 Correlation in size of analogous introns in mouse and teleost Piccolo genes. Mouse intron size was plotted against teleost intron size for t
 Figure S1 Correlation in size of analogous introns in mouse and teleost Piccolo genes. Mouse intron size was plotted against teleost intron size for the pcloa genes of zebrafish, green spotted puffer (listed
Figure S1 Correlation in size of analogous introns in mouse and teleost Piccolo genes. Mouse intron size was plotted against teleost intron size for the pcloa genes of zebrafish, green spotted puffer (listed
Identification of biological themes in microarray data from a mouse heart development time series using GeneSifter
 Identification of biological themes in microarray data from a mouse heart development time series using GeneSifter VizX Labs, LLC Seattle, WA 98119 Abstract Oligonucleotide microarrays were used to study
Identification of biological themes in microarray data from a mouse heart development time series using GeneSifter VizX Labs, LLC Seattle, WA 98119 Abstract Oligonucleotide microarrays were used to study
Textbook Reading Guidelines
 Understanding Bioinformatics by Marketa Zvelebil and Jeremy Baum Last updated: May 1, 2009 Textbook Reading Guidelines Preface: Read the whole preface, and especially: For the students with Life Science
Understanding Bioinformatics by Marketa Zvelebil and Jeremy Baum Last updated: May 1, 2009 Textbook Reading Guidelines Preface: Read the whole preface, and especially: For the students with Life Science
Figure S1. MUT-16 localization in L4 hermaphrodite, adult hermaphrodite, and adult male germlines. MUT-16 DAPI. Phillips et al. S-1. male.
 Supplementary Material for Phillips et al. Figure S1. MUT-16 localization in L4 hermaphrodite, adult hermaphrodite, and adult male germlines. Figure S2. Mutator foci and P granules localize independently
Supplementary Material for Phillips et al. Figure S1. MUT-16 localization in L4 hermaphrodite, adult hermaphrodite, and adult male germlines. Figure S2. Mutator foci and P granules localize independently
Supplemental Materials
 Supplemental Materials Supplemental Figure S. Phenotypic assessment of alb4 mutant plants under different stress conditions. (A) High-light stress and drought stress. Wild-type (WT) and alb4 mutant plants
Supplemental Materials Supplemental Figure S. Phenotypic assessment of alb4 mutant plants under different stress conditions. (A) High-light stress and drought stress. Wild-type (WT) and alb4 mutant plants
Fig. S1. Phosphorylation of eif2 by PEK-1 is increased in ire-1 mutants. Fig. S2. ire-1
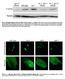 Fig. S1. Phosphorylation of eif2a by PEK-1 is increased in ire-1 mutants. Representative western blot of phosphorylated eif2a and tubulin of day-0 wild-type animals and ire-1 mutants treated with control,
Fig. S1. Phosphorylation of eif2a by PEK-1 is increased in ire-1 mutants. Representative western blot of phosphorylated eif2a and tubulin of day-0 wild-type animals and ire-1 mutants treated with control,
File S1. Program overview and features
 File S1 Program overview and features Query list filtering. Further filtering may be applied through user selected query lists (Figure. 2B, Table S3) that restrict the results and/or report specifically
File S1 Program overview and features Query list filtering. Further filtering may be applied through user selected query lists (Figure. 2B, Table S3) that restrict the results and/or report specifically
Supplemental Data. Zhou et al. (2016). Plant Cell /tpc
 Supplemental Figure 1. Confirmation of mutant mapping results. (A) Complementation assay with stably transformed genomic fragments (ComN-N) (2 kb upstream of TSS and 1.5 kb downstream of TES) and CaMV
Supplemental Figure 1. Confirmation of mutant mapping results. (A) Complementation assay with stably transformed genomic fragments (ComN-N) (2 kb upstream of TSS and 1.5 kb downstream of TES) and CaMV
This place covers: Methods or systems for genetic or protein-related data processing in computational molecular biology.
 G16B BIOINFORMATICS, i.e. INFORMATION AND COMMUNICATION TECHNOLOGY [ICT] SPECIALLY ADAPTED FOR GENETIC OR PROTEIN-RELATED DATA PROCESSING IN COMPUTATIONAL MOLECULAR BIOLOGY Methods or systems for genetic
G16B BIOINFORMATICS, i.e. INFORMATION AND COMMUNICATION TECHNOLOGY [ICT] SPECIALLY ADAPTED FOR GENETIC OR PROTEIN-RELATED DATA PROCESSING IN COMPUTATIONAL MOLECULAR BIOLOGY Methods or systems for genetic
Figure S1. USP-46 is expressed in several tissues including the nervous system
 Supplemental Figure legends Figure S1. USP-46 is expressed in several tissues including the nervous system Transgenic animals expressing a transcriptional reporter (P::GFP) were imaged using epifluorescence
Supplemental Figure legends Figure S1. USP-46 is expressed in several tissues including the nervous system Transgenic animals expressing a transcriptional reporter (P::GFP) were imaged using epifluorescence
Nature Genetics: doi: /ng Supplementary Figure 1. ChIP-seq genome browser views of BRM occupancy at previously identified BRM targets.
 Supplementary Figure 1 ChIP-seq genome browser views of BRM occupancy at previously identified BRM targets. Gene structures are shown underneath each panel. Supplementary Figure 2 pref6::ref6-gfp complements
Supplementary Figure 1 ChIP-seq genome browser views of BRM occupancy at previously identified BRM targets. Gene structures are shown underneath each panel. Supplementary Figure 2 pref6::ref6-gfp complements
Computational Genomics
 Computational Genomics Introduction to cell biology, genomics, development, and probability Eric Xing Lecture 1a, January 18, 2007 Reading: Chap. 1, DTM book Introduction to cell biology, functional genomics,
Computational Genomics Introduction to cell biology, genomics, development, and probability Eric Xing Lecture 1a, January 18, 2007 Reading: Chap. 1, DTM book Introduction to cell biology, functional genomics,
Supplemental Data. Sethi et al. (2014). Plant Cell /tpc
 Supplemental Data Supplemental Figure 1. MYC2 Binds to the E-box but not the E1-box of the MPK6 Promoter. (A) E1-box and E-box (wild type) containing MPK6 promoter fragment. The region shown in red denotes
Supplemental Data Supplemental Figure 1. MYC2 Binds to the E-box but not the E1-box of the MPK6 Promoter. (A) E1-box and E-box (wild type) containing MPK6 promoter fragment. The region shown in red denotes
Oligonucleotides used to amplify VvLOXA and VvLOXO for cloning into pentr TEV/D-TOPO vectors
 10.1071/FP09271_AC CSIRO 2010 Accessory Publication: Functional Plant Biology, 2010, 37(8), 767 784. Table S1. Oligonucleotide primers used to obtain full length coding sequence of berry expressed LOXs
10.1071/FP09271_AC CSIRO 2010 Accessory Publication: Functional Plant Biology, 2010, 37(8), 767 784. Table S1. Oligonucleotide primers used to obtain full length coding sequence of berry expressed LOXs
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature11070 Supplementary Figure 1 Purification of FLAG-tagged proteins. a, Purification of FLAG-RNF12 by FLAG-affinity from nuclear extracts of wild-type (WT) and two FLAG- RNF12 transgenic
doi:10.1038/nature11070 Supplementary Figure 1 Purification of FLAG-tagged proteins. a, Purification of FLAG-RNF12 by FLAG-affinity from nuclear extracts of wild-type (WT) and two FLAG- RNF12 transgenic
Biotechnology Explorer
 Biotechnology Explorer C. elegans Behavior Kit Bioinformatics Supplement explorer.bio-rad.com Catalog #166-5120EDU This kit contains temperature-sensitive reagents. Open immediately and see individual
Biotechnology Explorer C. elegans Behavior Kit Bioinformatics Supplement explorer.bio-rad.com Catalog #166-5120EDU This kit contains temperature-sensitive reagents. Open immediately and see individual
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature09937 a Name Position Primersets 1a 1b 2 3 4 b2 Phenotype Genotype b Primerset 1a D T C R I E 10000 8000 6000 5000 4000 3000 2500 2000 1500 1000 800 Donor (D)
SUPPLEMENTARY INFORMATION doi:10.1038/nature09937 a Name Position Primersets 1a 1b 2 3 4 b2 Phenotype Genotype b Primerset 1a D T C R I E 10000 8000 6000 5000 4000 3000 2500 2000 1500 1000 800 Donor (D)
Supplementary Information to: Genome-wide Real-time in vivo Transcriptional Dynamics During Plasmodium falciparum. Blood-stage Development
 Supplementary Information to: Genome-wide Real-time in vivo Transcriptional Dynamics During Plasmodium falciparum Blood-stage Development Painter et al. 1 of 8 Supplementary Figure 1: Supplementary Figure
Supplementary Information to: Genome-wide Real-time in vivo Transcriptional Dynamics During Plasmodium falciparum Blood-stage Development Painter et al. 1 of 8 Supplementary Figure 1: Supplementary Figure
7.012 Problem Set 5. Question 1
 Name Section 7.012 Problem Set 5 Question 1 While studying the problem of infertility, you attempt to isolate a hypothetical rabbit gene that accounts for the prolific reproduction of rabbits. After much
Name Section 7.012 Problem Set 5 Question 1 While studying the problem of infertility, you attempt to isolate a hypothetical rabbit gene that accounts for the prolific reproduction of rabbits. After much
amplification of the 5 flanking region of CAT1 with a tail for the geneticin resistance gene cassette fusion CAT1-3F
 Supplementary materials Table S1. Primer list. Primer Sequence a (5-3 ) Description CAT1-5F CAT1-5R ATACGGATAAGGAAGCGATAGCAGCA gcacaggtacacttgtttagagagcgatccggattttaagtgaacg amplification of the 5 flanking
Supplementary materials Table S1. Primer list. Primer Sequence a (5-3 ) Description CAT1-5F CAT1-5R ATACGGATAAGGAAGCGATAGCAGCA gcacaggtacacttgtttagagagcgatccggattttaagtgaacg amplification of the 5 flanking
Supplemental Figure 1
 Supplemental Figure 1 A gta2-1 gta2-2 1kb AT4G08350 B Col-0 gta2-1 LP+RP+LBb1.3 C Col-0 gta2-2 LP+RP+LB1 D Col-0 gta2-2 gta2-1 GTA2 TUBLIN E Col0 gta2-1 gta2-2 Supplemental Figure 1. Phenotypic analysis
Supplemental Figure 1 A gta2-1 gta2-2 1kb AT4G08350 B Col-0 gta2-1 LP+RP+LBb1.3 C Col-0 gta2-2 LP+RP+LB1 D Col-0 gta2-2 gta2-1 GTA2 TUBLIN E Col0 gta2-1 gta2-2 Supplemental Figure 1. Phenotypic analysis
Mos1 insertion. MosTIC protocol-11/2006. repair template - Mos1 transposase expression - Mos1 excision - DSB formation. homolog arm.
 MosTIC (Mos1 excision induced Transgene Instructed gene Conversion) Valérie Robert (vrobert@biologie.ens.fr) and Jean-Louis Bessereau (jlbesse@biologie.ens.fr) (November 2006) Introduction: MosTIC (Robert
MosTIC (Mos1 excision induced Transgene Instructed gene Conversion) Valérie Robert (vrobert@biologie.ens.fr) and Jean-Louis Bessereau (jlbesse@biologie.ens.fr) (November 2006) Introduction: MosTIC (Robert
Supplemental Material
 Supplemental Material 1 Figure S1. Phylogenetic analysis of Cep72 and Lrrc36, comparative localization of Cep72 and Lrrc36 and Cep72 antibody characterization (A) Phylogenetic alignment of Cep72 and Lrrc36
Supplemental Material 1 Figure S1. Phylogenetic analysis of Cep72 and Lrrc36, comparative localization of Cep72 and Lrrc36 and Cep72 antibody characterization (A) Phylogenetic alignment of Cep72 and Lrrc36
GREG GIBSON SPENCER V. MUSE
 A Primer of Genome Science ience THIRD EDITION TAGCACCTAGAATCATGGAGAGATAATTCGGTGAGAATTAAATGGAGAGTTGCATAGAGAACTGCGAACTG GREG GIBSON SPENCER V. MUSE North Carolina State University Sinauer Associates, Inc.
A Primer of Genome Science ience THIRD EDITION TAGCACCTAGAATCATGGAGAGATAATTCGGTGAGAATTAAATGGAGAGTTGCATAGAGAACTGCGAACTG GREG GIBSON SPENCER V. MUSE North Carolina State University Sinauer Associates, Inc.
Supplementary Figure 1. Homozygous rag2 E450fs mutants are healthy and viable similar to wild-type and heterozygous siblings.
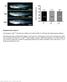 Supplementary Figure 1 Homozygous rag2 E450fs mutants are healthy and viable similar to wild-type and heterozygous siblings. (left) Representative bright-field images of wild type (wt), heterozygous (het)
Supplementary Figure 1 Homozygous rag2 E450fs mutants are healthy and viable similar to wild-type and heterozygous siblings. (left) Representative bright-field images of wild type (wt), heterozygous (het)
Genome research in eukaryotes
 Functional Genomics Genome and EST sequencing can tell us how many POTENTIAL genes are present in the genome Proteomics can tell us about proteins and their interactions The goal of functional genomics
Functional Genomics Genome and EST sequencing can tell us how many POTENTIAL genes are present in the genome Proteomics can tell us about proteins and their interactions The goal of functional genomics
Supplementary Tables and Figures
 Supplementary Tables and Figures The autophagy-related genes BbATG1 and BbATG8 have different functions in differentiation, stress resistance and virulence of mycopathogen Beauveria bassiana Sheng-Hua
Supplementary Tables and Figures The autophagy-related genes BbATG1 and BbATG8 have different functions in differentiation, stress resistance and virulence of mycopathogen Beauveria bassiana Sheng-Hua
Supplemental Data. Osakabe et al. (2013). Plant Cell /tpc
 Supplemental Figure 1. Phylogenetic analysis of KUP in various species. The amino acid sequences of KUPs from green algae and land plants were identified with a BLAST search and aligned using ClustalW
Supplemental Figure 1. Phylogenetic analysis of KUP in various species. The amino acid sequences of KUPs from green algae and land plants were identified with a BLAST search and aligned using ClustalW
Nature Methods: doi: /nmeth Supplementary Figure 1. Construction of a sensitive TetR mediated auxotrophic off-switch.
 Supplementary Figure 1 Construction of a sensitive TetR mediated auxotrophic off-switch. A Production of the Tet repressor in yeast when conjugated to either the LexA4 or LexA8 promoter DNA binding sequences.
Supplementary Figure 1 Construction of a sensitive TetR mediated auxotrophic off-switch. A Production of the Tet repressor in yeast when conjugated to either the LexA4 or LexA8 promoter DNA binding sequences.
BIOINFORMATICS Introduction
 BIOINFORMATICS Introduction Mark Gerstein, Yale University bioinfo.mbb.yale.edu/mbb452a 1 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu What is Bioinformatics? (Molecular) Bio -informatics One idea
BIOINFORMATICS Introduction Mark Gerstein, Yale University bioinfo.mbb.yale.edu/mbb452a 1 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu What is Bioinformatics? (Molecular) Bio -informatics One idea
Supplementary Data Supplementary Figures
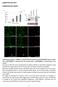 Supplementary Data Supplementary Figures Supplementary Figure 1. Pi04314 is expressed during infection, each GFP-Pi04314 fusion is stable and myr GFP-Pi04314 is removed from the nucleus while NLS GFP-Pi04314
Supplementary Data Supplementary Figures Supplementary Figure 1. Pi04314 is expressed during infection, each GFP-Pi04314 fusion is stable and myr GFP-Pi04314 is removed from the nucleus while NLS GFP-Pi04314
Computational Biology I
 Computational Biology I Microarray data acquisition Gene clustering Practical Microarray Data Acquisition H. Yang From Sample to Target cdna Sample Centrifugation (Buffer) Cell pellets lyse cells (TRIzol)
Computational Biology I Microarray data acquisition Gene clustering Practical Microarray Data Acquisition H. Yang From Sample to Target cdna Sample Centrifugation (Buffer) Cell pellets lyse cells (TRIzol)
A Naturally Occurring Epiallele associates with Leaf Senescence and Local Climate Adaptation in Arabidopsis accessions He et al.
 A Naturally Occurring Epiallele associates with Leaf Senescence and Local Climate Adaptation in Arabidopsis accessions He et al. Supplementary Notes Origin of NMR19 elements Because there are two copies
A Naturally Occurring Epiallele associates with Leaf Senescence and Local Climate Adaptation in Arabidopsis accessions He et al. Supplementary Notes Origin of NMR19 elements Because there are two copies
Targeted modification of gene function exploiting homology directed repair of TALENmediated double strand breaks in barley
 Targeted modification of gene function exploiting homology directed repair of TALENmediated double strand breaks in barley Nagaveni Budhagatapalli a, Twan Rutten b, Maia Gurushidze a, Jochen Kumlehn a,
Targeted modification of gene function exploiting homology directed repair of TALENmediated double strand breaks in barley Nagaveni Budhagatapalli a, Twan Rutten b, Maia Gurushidze a, Jochen Kumlehn a,
TIGR THE INSTITUTE FOR GENOMIC RESEARCH
 Introduction to Genome Annotation: Overview of What You Will Learn This Week C. Robin Buell May 21, 2007 Types of Annotation Structural Annotation: Defining genes, boundaries, sequence motifs e.g. ORF,
Introduction to Genome Annotation: Overview of What You Will Learn This Week C. Robin Buell May 21, 2007 Types of Annotation Structural Annotation: Defining genes, boundaries, sequence motifs e.g. ORF,
On the adequate use of microarray data
 On the adequate use of microarray data About semi-quantitative analysis Andreas Hoppe, Charité Universtitätsmedizin Berlin Computational systems biochemistry group Contents Introduction Gene array accuracy
On the adequate use of microarray data About semi-quantitative analysis Andreas Hoppe, Charité Universtitätsmedizin Berlin Computational systems biochemistry group Contents Introduction Gene array accuracy
The title page. Institution: College of Horticulture, South China Agricultural University
 The title page The manuscript title: Overexpression of the Eggplant (Solanum melongena) NAC Family Transcription Factor, SmNAC, Suppresses Resistance to Bacterial Wilt. The author list: 1 Cao Bihao: Corresponding
The title page The manuscript title: Overexpression of the Eggplant (Solanum melongena) NAC Family Transcription Factor, SmNAC, Suppresses Resistance to Bacterial Wilt. The author list: 1 Cao Bihao: Corresponding
Supplementary Figure 1. Isolation of GFPHigh cells.
 Supplementary Figure 1. Isolation of GFP High cells. (A) Schematic diagram of cell isolation based on Wnt signaling activity. Colorectal cancer (CRC) cell lines were stably transduced with lentivirus encoding
Supplementary Figure 1. Isolation of GFP High cells. (A) Schematic diagram of cell isolation based on Wnt signaling activity. Colorectal cancer (CRC) cell lines were stably transduced with lentivirus encoding
A Repressor Complex Governs the Integration of
 Developmental Cell 15 Supplemental Data A Repressor Complex Governs the Integration of Flowering Signals in Arabidopsis Dan Li, Chang Liu, Lisha Shen, Yang Wu, Hongyan Chen, Masumi Robertson, Chris A.
Developmental Cell 15 Supplemental Data A Repressor Complex Governs the Integration of Flowering Signals in Arabidopsis Dan Li, Chang Liu, Lisha Shen, Yang Wu, Hongyan Chen, Masumi Robertson, Chris A.
Computational methods in bioinformatics: Lecture 1
 Computational methods in bioinformatics: Lecture 1 Graham J.L. Kemp 2 November 2015 What is biology? Ecosystem Rain forest, desert, fresh water lake, digestive tract of an animal Community All species
Computational methods in bioinformatics: Lecture 1 Graham J.L. Kemp 2 November 2015 What is biology? Ecosystem Rain forest, desert, fresh water lake, digestive tract of an animal Community All species
To investigate the heredity of the WFP gene, we selected plants that were homozygous
 Supplementary information Supplementary Note ST-12 WFP allele is semi-dominant To investigate the heredity of the WFP gene, we selected plants that were homozygous for chromosome 1 of Nipponbare and heterozygous
Supplementary information Supplementary Note ST-12 WFP allele is semi-dominant To investigate the heredity of the WFP gene, we selected plants that were homozygous for chromosome 1 of Nipponbare and heterozygous
Supplemental Figure 1 HDA18 has an HDAC domain and therefore has concentration dependent and TSA inhibited histone deacetylase activity.
 Supplemental Figure 1 HDA18 has an HDAC domain and therefore has concentration dependent and TSA inhibited histone deacetylase activity. (A) Amino acid alignment of HDA5, HDA15 and HDA18. The blue line
Supplemental Figure 1 HDA18 has an HDAC domain and therefore has concentration dependent and TSA inhibited histone deacetylase activity. (A) Amino acid alignment of HDA5, HDA15 and HDA18. The blue line
NGS Approaches to Epigenomics
 I519 Introduction to Bioinformatics, 2013 NGS Approaches to Epigenomics Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Contents Background: chromatin structure & DNA methylation Epigenomic
I519 Introduction to Bioinformatics, 2013 NGS Approaches to Epigenomics Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Contents Background: chromatin structure & DNA methylation Epigenomic
Nature Genetics: doi: /ng Supplementary Figure 1
 Supplementary Figure 1 Ihh interacts preferentially with its upstream neighboring gene Nhej1. Genes are indicated by gray lines, and Ihh and Nhej1 are highlighted in blue. 4C seq performed in E14.5 limbs
Supplementary Figure 1 Ihh interacts preferentially with its upstream neighboring gene Nhej1. Genes are indicated by gray lines, and Ihh and Nhej1 are highlighted in blue. 4C seq performed in E14.5 limbs
Supplemental Figure 1.
 Supplemental Data. Charron et al. Dynamic landscapes of four histone modifications during de-etiolation in Arabidopsis. Plant Cell (2009). 10.1105/tpc.109.066845 Supplemental Figure 1. Immunodetection
Supplemental Data. Charron et al. Dynamic landscapes of four histone modifications during de-etiolation in Arabidopsis. Plant Cell (2009). 10.1105/tpc.109.066845 Supplemental Figure 1. Immunodetection
embryos. Asterisk represents loss of or reduced expression. Brackets represent
 Supplemental Figures Supplemental Figure 1. tfec expression is highly enriched in tail endothelial cells (A- B) ISH of tfec at 15 and 16hpf in WT embryos. (C- D) ISH of tfec at 36 and 38hpf in WT embryos.
Supplemental Figures Supplemental Figure 1. tfec expression is highly enriched in tail endothelial cells (A- B) ISH of tfec at 15 and 16hpf in WT embryos. (C- D) ISH of tfec at 36 and 38hpf in WT embryos.
Supplemental Data. Guo et al. (2015). Plant Cell /tpc
 Supplemental Figure 1. The Mutant exb1-d Displayed Pleiotropic Phenotypes and Produced Branches in the Axils of Cotyledons. (A) Branches were developed in exb1-d but not in wild-type plants. (B) and (C)
Supplemental Figure 1. The Mutant exb1-d Displayed Pleiotropic Phenotypes and Produced Branches in the Axils of Cotyledons. (A) Branches were developed in exb1-d but not in wild-type plants. (B) and (C)
LysoTracker Red DND-99 (Invitrogen) was used as a marker of lysosome or acidic
 information MATERIAL AND METHODS Lysosome staining LysoTracker Red DND-99 (Invitrogen) was used as a marker of lysosome or acidic compartments, according to the manufacturer s protocol. Plasmid independent
information MATERIAL AND METHODS Lysosome staining LysoTracker Red DND-99 (Invitrogen) was used as a marker of lysosome or acidic compartments, according to the manufacturer s protocol. Plasmid independent
SAS6-like protein in Plasmodium indicates that conoid-associated apical complex proteins persist in invasive stages within the mosquito vector
 SAS6-like protein in Plasmodium indicates that conoid-associated apical complex proteins persist in invasive stages within the mosquito vector Richard J. Wall 1,#, Magali Roques 1,+, Nicholas J. Katris
SAS6-like protein in Plasmodium indicates that conoid-associated apical complex proteins persist in invasive stages within the mosquito vector Richard J. Wall 1,#, Magali Roques 1,+, Nicholas J. Katris
Supplemental Data. Li et al. (2015). Plant Cell /tpc
 Supplemental Data Supplemental Figure 1: Characterization of asr3 T-DNA knockout lines and complementation transgenic lines. (A) The scheme of At2G33550 (ASR3) with gray boxes indicating exons and dash
Supplemental Data Supplemental Figure 1: Characterization of asr3 T-DNA knockout lines and complementation transgenic lines. (A) The scheme of At2G33550 (ASR3) with gray boxes indicating exons and dash
Supplementary Fig. 1. Schematic structure of TRAIP and RAP80. The prey line below TRAIP indicates bait and the two lines above RAP80 highlight the
 Supplementary Fig. 1. Schematic structure of TRAIP and RAP80. The prey line below TRAIP indicates bait and the two lines above RAP80 highlight the prey clones identified in the yeast two hybrid screen.
Supplementary Fig. 1. Schematic structure of TRAIP and RAP80. The prey line below TRAIP indicates bait and the two lines above RAP80 highlight the prey clones identified in the yeast two hybrid screen.
Nature Methods: doi: /nmeth Supplementary Figure 1
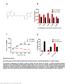 Supplementary Figure 1 Enrichment with anti-biotin antibody significantly increases detection of biotinylated peptides in complex samples. (A) Structure of NHS-biotin (B) Results of spike-in studies. Bar
Supplementary Figure 1 Enrichment with anti-biotin antibody significantly increases detection of biotinylated peptides in complex samples. (A) Structure of NHS-biotin (B) Results of spike-in studies. Bar
Supplemental Data. Wu et al. (2). Plant Cell..5/tpc RGLG Hormonal treatment H2O B RGLG µm ABA µm ACC µm GA Time (hours) µm µm MJ µm IA
 Supplemental Data. Wu et al. (2). Plant Cell..5/tpc..4. A B Supplemental Figure. Immunoblot analysis verifies the expression of the AD-PP2C and BD-RGLG proteins in the Y2H assay. Total proteins were extracted
Supplemental Data. Wu et al. (2). Plant Cell..5/tpc..4. A B Supplemental Figure. Immunoblot analysis verifies the expression of the AD-PP2C and BD-RGLG proteins in the Y2H assay. Total proteins were extracted
Supplemental Table 1. List of PCR primers
 Supplemental Table 1. List of PCR primers Primer Sequences (5 to 3 ) 1 TATCCATGGCGCCGGCGGCGAGGGCGGAG 2 TATAAGCTTCTTGGCGTGTCCAGCCCACGGGGCGTAGAACTCGAC 3 ATAAAGCTTGCTCCAGAGTATGAGAAAGCT 4 ATACTCGAGGAGCTCATCCTTGAGAGGCTC
Supplemental Table 1. List of PCR primers Primer Sequences (5 to 3 ) 1 TATCCATGGCGCCGGCGGCGAGGGCGGAG 2 TATAAGCTTCTTGGCGTGTCCAGCCCACGGGGCGTAGAACTCGAC 3 ATAAAGCTTGCTCCAGAGTATGAGAAAGCT 4 ATACTCGAGGAGCTCATCCTTGAGAGGCTC
Bioinformatics for Cell Biologists
 Bioinformatics for Cell Biologists 15 19 March 2010 Developmental Biology and Regnerative Medicine (DBRM) Schedule Monday, March 15 09.00 11.00 Introduction to course and Bioinformatics (L1) D224 Helena
Bioinformatics for Cell Biologists 15 19 March 2010 Developmental Biology and Regnerative Medicine (DBRM) Schedule Monday, March 15 09.00 11.00 Introduction to course and Bioinformatics (L1) D224 Helena
Supplemental Figure 1
 Supplemental Figure 1 A LK sls1 lks1-2 F 1 sls1 B LK sls1 lks1-2 F 1 lks1-2 sls1 F 1 lks1-2 sls1 F 2 Col lks1-2 Col+LKS1 Col Col+LKS1 Supplemental Figure 1. Genetic analysis of sls1 mutant. (A) and (B)
Supplemental Figure 1 A LK sls1 lks1-2 F 1 sls1 B LK sls1 lks1-2 F 1 lks1-2 sls1 F 1 lks1-2 sls1 F 2 Col lks1-2 Col+LKS1 Col Col+LKS1 Supplemental Figure 1. Genetic analysis of sls1 mutant. (A) and (B)
Introduction to Bioinformatics and Gene Expression Technology
 Vocabulary Introduction to Bioinformatics and Gene Expression Technology Utah State University Spring 2014 STAT 5570: Statistical Bioinformatics Notes 1.1 Gene: Genetics: Genome: Genomics: hereditary DNA
Vocabulary Introduction to Bioinformatics and Gene Expression Technology Utah State University Spring 2014 STAT 5570: Statistical Bioinformatics Notes 1.1 Gene: Genetics: Genome: Genomics: hereditary DNA
Revised: RG-RV2 by Fukuhara et al.
 Supplemental Figure 1 The generation of Spns2 conditional knockout mice. (A) Schematic representation of the wild type Spns2 locus (Spns2 + ), the targeted allele, the floxed allele (Spns2 f ) and the
Supplemental Figure 1 The generation of Spns2 conditional knockout mice. (A) Schematic representation of the wild type Spns2 locus (Spns2 + ), the targeted allele, the floxed allele (Spns2 f ) and the
GENETICS - CLUTCH CH.15 GENOMES AND GENOMICS.
 !! www.clutchprep.com CONCEPT: OVERVIEW OF GENOMICS Genomics is the study of genomes in their entirety Bioinformatics is the analysis of the information content of genomes - Genes, regulatory sequences,
!! www.clutchprep.com CONCEPT: OVERVIEW OF GENOMICS Genomics is the study of genomes in their entirety Bioinformatics is the analysis of the information content of genomes - Genes, regulatory sequences,
Supplemental Data. Wu and Xue (2010). Plant Cell /tpc
 Supplemental Data. Wu and Xue (21). Plant Cell 1.115/tpc.11.75564 A P1-S P1-A P2-S P2-A P3-S P3-A P4-S P4-A B Relative expression Relative expression C 5. 4.5 4. 3.5 3. 2.5 2. 1.5 1..5 5. 4.5 4. 3.5 3.
Supplemental Data. Wu and Xue (21). Plant Cell 1.115/tpc.11.75564 A P1-S P1-A P2-S P2-A P3-S P3-A P4-S P4-A B Relative expression Relative expression C 5. 4.5 4. 3.5 3. 2.5 2. 1.5 1..5 5. 4.5 4. 3.5 3.
Aminoacid change in chromophore. PIN3::PIN3-GFP GFP S65 (no change) (5.9) (Kneen et al., 1998)
 Supplemental Table. Table S1. Fluorophore characteristics of used fluorescent proteins, Related to Figure 2 and 3. Transgenic line Fluprescent marker Aminoacid change in chromophore pk(a) Ref. PIN3::PIN3-GFP
Supplemental Table. Table S1. Fluorophore characteristics of used fluorescent proteins, Related to Figure 2 and 3. Transgenic line Fluprescent marker Aminoacid change in chromophore pk(a) Ref. PIN3::PIN3-GFP
Perspectives on the Priorities for Bioinformatics Education in the 21 st Century
 Perspectives on the Priorities for Bioinformatics Education in the 21 st Century Oyekanmi Nash, PhD Associate Professor & Director Genetics, Genomics & Bioinformatics National Biotechnology Development
Perspectives on the Priorities for Bioinformatics Education in the 21 st Century Oyekanmi Nash, PhD Associate Professor & Director Genetics, Genomics & Bioinformatics National Biotechnology Development
Bioinformatics & Protein Structural Analysis. Bioinformatics & Protein Structural Analysis. Learning Objective. Proteomics
 The molecular structures of proteins are complex and can be defined at various levels. These structures can also be predicted from their amino-acid sequences. Protein structure prediction is one of the
The molecular structures of proteins are complex and can be defined at various levels. These structures can also be predicted from their amino-acid sequences. Protein structure prediction is one of the
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/5/244/ra72/dc1 Supplementary Materials for An Interaction Between BZR1 and DELLAs Mediates Direct Signaling Crosstalk Between Brassinosteroids and Gibberellins
www.sciencesignaling.org/cgi/content/full/5/244/ra72/dc1 Supplementary Materials for An Interaction Between BZR1 and DELLAs Mediates Direct Signaling Crosstalk Between Brassinosteroids and Gibberellins
Xu et al., Supplementary Figures 1-7
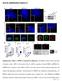 Xu et al., Supplementary Figures 1-7 Supplementary Figure 1. PIPKI is required for ciliogenesis. (a) PIPKI localizes at the basal body of primary cilium. RPE-1 cells treated with two sirnas targeting to
Xu et al., Supplementary Figures 1-7 Supplementary Figure 1. PIPKI is required for ciliogenesis. (a) PIPKI localizes at the basal body of primary cilium. RPE-1 cells treated with two sirnas targeting to
Chapter 20 Recombinant DNA Technology. Copyright 2009 Pearson Education, Inc.
 Chapter 20 Recombinant DNA Technology Copyright 2009 Pearson Education, Inc. 20.1 Recombinant DNA Technology Began with Two Key Tools: Restriction Enzymes and DNA Cloning Vectors Recombinant DNA refers
Chapter 20 Recombinant DNA Technology Copyright 2009 Pearson Education, Inc. 20.1 Recombinant DNA Technology Began with Two Key Tools: Restriction Enzymes and DNA Cloning Vectors Recombinant DNA refers
0.5% Sucrose. Supplemental Data. Chao et al. (2011). Plant Cell /tpc Col-0 tsc10a-2. Root length (mm)
 A 0% 0.5% 1% Col-0 tsc10a-2 B Root length (mm) 35 30 25 20 15 10 5 0 0% 0.50% 1% 2% 4% Sucrose concentration 2% 4% Sucrose Col-0 tsc10a-2 Supplemental Figure 1. The effect of sucrose on root growth of
A 0% 0.5% 1% Col-0 tsc10a-2 B Root length (mm) 35 30 25 20 15 10 5 0 0% 0.50% 1% 2% 4% Sucrose concentration 2% 4% Sucrose Col-0 tsc10a-2 Supplemental Figure 1. The effect of sucrose on root growth of
Supplementary information
 Supplementary information Supplementary figures Figure S1 Level of mycdet1 protein in DET1 OE-1, OE-2 and OE-3 transgenic lines. Total protein extract from wild type Col0, det1-1 mutant and DET1 OE lines
Supplementary information Supplementary figures Figure S1 Level of mycdet1 protein in DET1 OE-1, OE-2 and OE-3 transgenic lines. Total protein extract from wild type Col0, det1-1 mutant and DET1 OE lines
Supplementary Information
 Supplementary Information MED18 interaction with distinct transcription factors regulates plant immunity, flowering time and responses to hormones Supplementary Figure 1. Diagram showing T-DNA insertion
Supplementary Information MED18 interaction with distinct transcription factors regulates plant immunity, flowering time and responses to hormones Supplementary Figure 1. Diagram showing T-DNA insertion
Identification and characterization of a pathogenicity-related gene VdCYP1 from Verticillium dahliae
 Identification and characterization of a pathogenicity-related gene VdCYP1 from Verticillium dahliae Dan-Dan Zhang*, Xin-Yan Wang*, Jie-Yin Chen*, Zhi-Qiang Kong, Yue-Jing Gui, Nan-Yang Li, Yu-Ming Bao,
Identification and characterization of a pathogenicity-related gene VdCYP1 from Verticillium dahliae Dan-Dan Zhang*, Xin-Yan Wang*, Jie-Yin Chen*, Zhi-Qiang Kong, Yue-Jing Gui, Nan-Yang Li, Yu-Ming Bao,
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION Supplementary figures Supplementary Figure 1: Suv39h1, but not Suv39h2, promotes HP1α sumoylation in vivo. In vivo HP1α sumoylation assay. Top: experimental scheme. Middle: we
SUPPLEMENTARY INFORMATION Supplementary figures Supplementary Figure 1: Suv39h1, but not Suv39h2, promotes HP1α sumoylation in vivo. In vivo HP1α sumoylation assay. Top: experimental scheme. Middle: we
Mannen et al., http :// /cgi /content /full /jcb /DC1
 Supplemental material JCB Mannen et al., http ://www.jcb.org /cgi /content /full /jcb.201601024 /DC1 THE JOURNAL OF CELL BIOLOGY Figure S1. Characterization of SNB components. (A) SNB localization of Venus-tagged
Supplemental material JCB Mannen et al., http ://www.jcb.org /cgi /content /full /jcb.201601024 /DC1 THE JOURNAL OF CELL BIOLOGY Figure S1. Characterization of SNB components. (A) SNB localization of Venus-tagged
Nature Biotechnology: doi: /nbt.4166
 Supplementary Figure 1 Validation of correct targeting at targeted locus. (a) by immunofluorescence staining of 2C-HR-CRISPR microinjected embryos cultured to the blastocyst stage. Embryos were stained
Supplementary Figure 1 Validation of correct targeting at targeted locus. (a) by immunofluorescence staining of 2C-HR-CRISPR microinjected embryos cultured to the blastocyst stage. Embryos were stained
indicated numbers of pups at day of life (DOL) 10, or embryonic day (ED) B. Male mice of
 SUPPLEMENTRY FIGURE LEGENDS Figure S1. USP44 loss leads to chromosome missegregation.. Genotypes obtained from the indicated numbers of pups at day of life (DOL) 10, or embryonic day (ED) 13.5.. Male mice
SUPPLEMENTRY FIGURE LEGENDS Figure S1. USP44 loss leads to chromosome missegregation.. Genotypes obtained from the indicated numbers of pups at day of life (DOL) 10, or embryonic day (ED) 13.5.. Male mice
MARINE BIOINFORMATICS & NANOBIOTECHNOLOGY - PBBT305
 MARINE BIOINFORMATICS & NANOBIOTECHNOLOGY - PBBT305 UNIT-1 MARINE GENOMICS AND PROTEOMICS 1. Define genomics? 2. Scope and functional genomics? 3. What is Genetics? 4. Define functional genomics? 5. What
MARINE BIOINFORMATICS & NANOBIOTECHNOLOGY - PBBT305 UNIT-1 MARINE GENOMICS AND PROTEOMICS 1. Define genomics? 2. Scope and functional genomics? 3. What is Genetics? 4. Define functional genomics? 5. What
Supplementary Information
 1 Supplementary Information 2 3 4 5 6 Characterization and expression of the cytochrome P450 gene family in diamondback moth, Plutella xylostella (L.) Liying Yu 1,2,3, Weiqi Tang 1,2, Weiyi He 1,3, Xiaoli
1 Supplementary Information 2 3 4 5 6 Characterization and expression of the cytochrome P450 gene family in diamondback moth, Plutella xylostella (L.) Liying Yu 1,2,3, Weiqi Tang 1,2, Weiyi He 1,3, Xiaoli
Genome annotation & EST
 Genome annotation & EST What is genome annotation? The process of taking the raw DNA sequence produced by the genome sequence projects and adding the layers of analysis and interpretation necessary
Genome annotation & EST What is genome annotation? The process of taking the raw DNA sequence produced by the genome sequence projects and adding the layers of analysis and interpretation necessary
Supplemental Figure 1 A
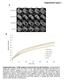 Supplemental Figure A prebleach postbleach 2 min 6 min 3 min mh2a.-gfp mh2a.2-gfp mh2a2-gfp GFP-H2A..9 Relative Intensity.8.7.6.5 mh2a. GFP n=8.4 mh2a.2 GFP n=4.3 mh2a2 GFP n=2.2 GFP H2A n=24. GFP n=7.
Supplemental Figure A prebleach postbleach 2 min 6 min 3 min mh2a.-gfp mh2a.2-gfp mh2a2-gfp GFP-H2A..9 Relative Intensity.8.7.6.5 mh2a. GFP n=8.4 mh2a.2 GFP n=4.3 mh2a2 GFP n=2.2 GFP H2A n=24. GFP n=7.
Genome manipulation by homologous recombination in Drosophila Xiaolin Bi and Yikang S. Rong Date received (in revised form): 9th May 2003
 Xiaolin Bi is a post doctoral research fellow at the Laboratory of Molecular Cell Biology, National Cancer Institute, National Institutes of Health, Bethesda, Maryland, USA. Yikang S. Rong is the principal
Xiaolin Bi is a post doctoral research fellow at the Laboratory of Molecular Cell Biology, National Cancer Institute, National Institutes of Health, Bethesda, Maryland, USA. Yikang S. Rong is the principal
Supplemental Data. Jing et al. (2013). Plant Cell /tpc
 Supplemental Figure 1. Characterization of epp1 Mutants. (A) Cotyledon angles of 5-d-old Col wild-type (gray bars) and epp1-1 (black bars) seedlings under red (R), far-red (FR) and blue (BL) light conditions,
Supplemental Figure 1. Characterization of epp1 Mutants. (A) Cotyledon angles of 5-d-old Col wild-type (gray bars) and epp1-1 (black bars) seedlings under red (R), far-red (FR) and blue (BL) light conditions,
Annotation. (Chapter 8)
 Annotation (Chapter 8) Genome annotation Genome annotation is the process of attaching biological information to sequences: identify elements on the genome attach biological information to elements store
Annotation (Chapter 8) Genome annotation Genome annotation is the process of attaching biological information to sequences: identify elements on the genome attach biological information to elements store
This practical aims to walk you through the process of text searching DNA and protein databases for sequence entries.
 PRACTICAL 1: BLAST and Sequence Alignment The EBI and NCBI websites, two of the most widely used life science web portals are introduced along with some of the principal databases: the NCBI Protein database,
PRACTICAL 1: BLAST and Sequence Alignment The EBI and NCBI websites, two of the most widely used life science web portals are introduced along with some of the principal databases: the NCBI Protein database,
CS 262 Lecture 14 Notes Human Genome Diversity, Coalescence and Haplotypes
 CS 262 Lecture 14 Notes Human Genome Diversity, Coalescence and Haplotypes Coalescence Scribe: Alex Wells 2/18/16 Whenever you observe two sequences that are similar, there is actually a single individual
CS 262 Lecture 14 Notes Human Genome Diversity, Coalescence and Haplotypes Coalescence Scribe: Alex Wells 2/18/16 Whenever you observe two sequences that are similar, there is actually a single individual
Description of Supplementary Files. File name: Supplementary Information Description: Supplementary figures and supplementary tables.
 Description of Supplementary Files File name: Supplementary Information Description: Supplementary figures and supplementary tables. File name: Supplementary Data 1 Description: Differential expression
Description of Supplementary Files File name: Supplementary Information Description: Supplementary figures and supplementary tables. File name: Supplementary Data 1 Description: Differential expression
Supplemental Data. Hu et al. Plant Cell (2017) /tpc
 1 2 3 4 Supplemental Figure 1. DNA gel blot analysis of homozygous transgenic plants. (Supports Figure 1.) 5 6 7 8 Rice genomic DNA was digested with the restriction enzymes EcoRⅠ and BamHⅠ. Lanes in the
1 2 3 4 Supplemental Figure 1. DNA gel blot analysis of homozygous transgenic plants. (Supports Figure 1.) 5 6 7 8 Rice genomic DNA was digested with the restriction enzymes EcoRⅠ and BamHⅠ. Lanes in the
Supplemental Data. Cui et al. (2012). Plant Cell /tpc a b c d. Stem UBC32 ACTIN
 A Root Stem Leaf Flower Silique Senescence leaf B a b c d UBC32 ACTIN C * Supplemental Figure 1. Expression Pattern and Protein Sequence of UBC32 Homologues in Yeast, Human, and Arabidopsis. (A) Expression
A Root Stem Leaf Flower Silique Senescence leaf B a b c d UBC32 ACTIN C * Supplemental Figure 1. Expression Pattern and Protein Sequence of UBC32 Homologues in Yeast, Human, and Arabidopsis. (A) Expression
