Data and Metadata Models Recommendations Version 1.2 Developed by the IHEC Metadata Standards Workgroup
|
|
|
- Roberta McDaniel
- 6 years ago
- Views:
Transcription
1 Data and Metadata Models Recommendations Version 1.2 Developed by the IHEC Metadata Standards Workgroup 1. Introduction The data produced by IHEC is illustrated in Figure 1. Figure 1. The space of epigenomic variation. The two-dimensional plane in Figure 1 corresponding to Assay and Cell Type combinations is displayed on the Epigenome Atlas portal page ( in the form of an interactive grid, as illustrated in Figure 2. Each cell in the grid in Figure 2 corresponds to the same Cell Type Assay combination. Multiple sequencing Runs are combined by Library ID into the same technical replicate and, at the next higher level by Donor ID into a biological replicate. Counts within each grid cell indicate the combined number of technical and biological replicates. By selecting individual cells in the grid, the user may navigate along the Genome coordinate axis by genomic element, gene, pathway and by other means using tools provided on the Epigenome Atlas portal page. The grid in Figure 2 is constructed automatically from the submitted metadata. The metadata may be viewed by clicking on column headers, row headers or individual cells within the grid. 1
2 Figure 2. Human Epigenome Atlas portal (at ). 2
3 2. Data and Metadata Model The data / metadata model in Fig. 3 is used to capture the data from mapping centers and generate data and metadata for submission to the GEO/SRA archives at the NCBI and organize the Human Epigenome Atlas portal. The arrows indicate references ( foreign key references). Specifically, the arrows indicate that a sequencing experiment corresponds to a specific Run, an event which denotes production of a specific unit of data (such as the production of DNA sequencing reads from a single lane of an Illumina sequencer or hybridization of a sample to a specific arrays) in the context of a specific Experiment on a specific Sample in the context of a specific Study. The data and metadata models are based on the SRA XML Schema Version 1.2. This schema was co-developed and is shared by the EBI and NCBI archives, facilitating exchange of data and international collaboration. The SRA XML Schema Version 1.2 allows extensions with data fields that may be required to capture relevant data for specific applications such as epigenomic assays. The NIH Roadmap Epigenomics Initiative has extended the schema with specific data fields for Study, Sample, Experiment, Run and various Analysis Levels and types, as described in the following section. 3
4 3. Metadata Elements Extending SRA XML Schema 1.2 The core SRA XML elements are augmented by additional attributes defined for purposes of the NIH Roadmap Epigenomics as described in this section. The same attribute may be used multiple times in a single XML record. This may be most useful for supplying URIs to multiple ontologies or for supplying multiple references to a single ontology such as in the case of DISEASE_ONTOLOGY_URI. Documentation for the core SRA XML elements is here: The SRA XML schemas are here: Level 0 Data SAMPLES Cell Line MOLECULE - The type of molecule that was extracted from the biological material. Include one of the following: total RNA, polya RNA, cytoplasmic RNA, nuclear RNA, genomic DNA, protein, or other. SAMPLE_ONTOLOGY_URI - links to sample ontology information. DISEASE_ONTOLOGY_URI - links to disease ontology information. DISEASE: Free form field for more specific disease information BIOMATERIAL_PROVIDER - The name of the company, laboratory or person that provided the biological material. BIOMATERIAL_TYPE: Cell Line LINE The name of the cell line. LINEAGE The developmental lineage to which the cell line belongs. DIFFERENTIATION_STAGE - The stage in cell differentiation to which the cell line belongs. DIFFERENTIATION_METHOD The protocol used to differentiation the cell line. 4
5 PASSAGE The number of times the cell line has been re-plated and allowed to grow back to confluency or to some maximum density if using suspension cultures. MEDIUM The medium in which the cell line has been grown. SEX: "Male", "Female" or "Unknown" BATCH The batch from which the cell line is derived. Primarily applicable to initial H1 cell line batches. NA if not applicable. Primary Cell MOLECULE - The type of molecule that was extracted from the biological material. Include one of the following: total RNA, polya RNA, cytoplasmic RNA, nuclear RNA, genomic DNA, protein, or other. SAMPLE_ONTOLOGY_URI - links to sample ontology information. DISEASE_ONTOLOGY_URI - links to disease ontology information. DISEASE: Free form field for more specific disease information BIOMATERIAL_PROVIDER - The name of the company, laboratory or person that provided the biological material. BIOMATERIAL_TYPE: Primary Cell CELL_TYPE The type of cell. MARKERS Markers used to isolate and identify the cell type. DONOR_ID - An identifying designation for the donor that provided the primary cell. DONOR_AGE - The age of the donor that provided the primary cell. NA if not available. DONOR_HEALTH_STATUS - The health status of the donor that provided the primary cell. NA if not available. DONOR_SEX: "Male", "Female" or "Unknown" DONOR_ETHNICITY - The ethnicity of the donor that provided the primary cell. NA if not available. PASSAGE_IF_EXPANDED If the primary cell has been expanded, the number 5
6 of times the primary cell has been re-plated and allowed to grow back to confluency or to some maximum density if using suspension cultures. NA if no expansion. Primary Cell Culture MOLECULE- The type of molecule that was extracted from the biological material. Include one of the following: total RNA, polya RNA, cytoplasmic RNA, nuclear RNA, genomic DNA, protein, or other. SAMPLE_ONTOLOGY_URI - links to sample ontology information. DISEASE_ONTOLOGY_URI - links to disease ontology information. DISEASE: Free form field for more specific disease information. BIOMATERIAL_PROVIDER - The name of the company, laboratory or person that provided the biological material. BIOMATERIAL_TYPE: Primary Cell Culture CELL_TYPE The type of cell. MARKERS Markers used to isolate and identify the cell type. CULTURE_CONDITIONS The conditions under which the primary cell was cultured. DONOR_ID - An identifying designation for the donor that provided the primary cell. DONOR_AGE - The age of the donor that provided the primary cell. NA if not available. DONOR_HEALTH_STATUS - The health status of the donor that provided the primary cell. NA if not available. DONOR_SEX: "Male", "Female" or "Unknown" DONOR_ETHNICITY - The ethnicity of the donor that provided the primary cell. NA if not available. PASSAGE_IF_EXPANDED If the primary cell culture has been expanded, the number of times the cell culture has been re-plated and allowed to grow back to confluency or to some maximum density if using suspension cultures. NA if no expansion. 6
7 Primary Tissue MOLECULE - The type of molecule that was extracted from the biological material. Include one of the following: total RNA, polya RNA, cytoplasmic RNA, nuclear RNA, genomic DNA, protein, or other. SAMPLE_ONTOLOGY_URI - links to sample ontology information. DISEASE_ONTOLOGY_URI - links to disease ontology information. DISEASE: Free form field for more specific disease information. BIOMATERIAL_PROVIDER - The name of the company, laboratory or person that provided the biological material. BIOMATERIAL_TYPE: Primary Tissue TISSUE_TYPE The type of tissue. TISSUE_DEPOT Details about the anatomical location from which the primary tissue was collected. COLLECTION_METHOD The protocol for collecting the primary tissue. DONOR_ID - An identifying designation for the donor that provided the primary tissue. DONOR_AGE - The age of the donor that provided the primary tissue. NA if not available. DONOR_HEALTH_STATUS - The health status of the donor that provided the primary tissue. NA if not available. DONOR_SEX: "Male", "Female" or "Unknown" DONOR_ETHNICITY - The ethnicity of the donor that provided the primary tissue. NA if not available. EXPERIMENTS Chromatin Accessibility EXPERIMENT_TYPE: Chromatin Accessibility EXPERIMENT_ONTOLOGY_URI links to experiment ontology information. EXTRACTION_PROTOCOL - The protocol used to isolate the extract material. 7
8 DNASE_PROTOCOL The protocol used for DNAse treatment. WGBS (NOTE: this is a new name to be used instead of Bisulfite-Seq ) EXPERIMENT_TYPE: DNA Methylation EXTRACTION_PROTOCOL - The protocol used to isolate the extract material. EXTRACTION_PROTOCOL_TYPE_OF_SONICATOR - The type of sonicator used for extraction. EXTRACTION_PROTOCOL_SONICATION_CYCLES - The number of sonication cycles used for extraction. DNA_PREPARATION_INITIAL_DNA_QNTY The initial DNA quantity used in preparation. DNA_PREPARATION_FRAGMENT_SIZE_RANGE The DNA fragment size range used in preparation. DNA_PREPARATION_ADAPTOR_SEQUENCE The sequence of the adaptor used in preparation. DNA_PREPARATION_ADAPTOR_LIGATION_PROTOCOL The protocol used for adaptor ligation. DNA_PREPARATION_POST-LIGATION_FRAGMENT_SIZE_SELECTION The fragment size selection after adaptor ligation. BISULFITE_CONVERSION_PROTOCOL The bisulfite conversion protocol. BISULFITE_CONVERSION_PERCENT The bisulfite conversion percent and how it was determined. LIBRARY_GENERATION_PCR_TEMPLATE_CONC The PCR template concentration for library generation. LIBRARY_GENERATION_PCR_POLYMERASE_TYPE The PCR polymerase used for library generation LIBRARY_GENERATION_PCR_THERMOCYCLING_PROGRAM The thermocycling program used for library generation. LIBRARY_GENERATION_PCR_NUMBER_CYCLES The number of PCR cycles used for library generation. 8
9 LIBRARY_GENERATION_PCR_F_PRIMER_SEQUENCE The sequence of the PCR forward primer used for library generation. LIBRARY_GENERATION_PCR_R_PRIMER_SEQUENCE The sequence of the PCR reverse primer used for library generation. LIBRARY_GENERATION_PCR_PRIMER_CONC The concentration of the PCR primers used for library generation. LIBRARY_GENERATION_PCR_PRODUCT_ISOLATION_PROTOCOL The protocol for isolating PCR products used for library generation. MeDIP-Seq EXPERIMENT_TYPE: DNA Methylation EXPERIMENT_ONTOLOGY_URI - links to experiment ontology information. EXTRACTION_PROTOCOL - The protocol used to isolate the extract material. EXTRACTION_PROTOCOL_TYPE_OF_SONICATOR - The type of sonicator used for extraction. EXTRACTION_PROTOCOL_SONICATION_CYCLES - The number of sonication cycles used for extraction. MeDIP_PROTOCOL The MeDIP protocol used. MeDIP_PROTOCOL_DNA_AMOUNT The amount of DNA used in the MeDIP protocol. MeDIP_PROTOCOL_BEAD_TYPE The type of bead used in the MeDIP protocol. MeDIP_PROTOCOL_BEAD_AMOUNT The amount of beads used in the MeDIP protocol. MeDIP_PROTOCOL_ANTIBODY_AMOUNT The amount of antibody used in the MeDIP protocol. MeDIP_ANTIBODY The specific antibody used in the MeDIP protocol. MeDIP_ANTIBODY_PROVIDER - The name of the company, laboratory or person that provided the antibody. MeDIP_ANTIBODY_CATALOG The catalog from which the antibody was purchased. 9
10 MeDIP_ANTIBODY_LOT The lot identifier of the antibody. MRE-Seq EXPERIMENT_TYPE: DNA Methylation EXPERIMENT_ONTOLOGY_URI elements that contain links to experiment ontology information. MRE_PROTOCOL The MRE protocol. MRE_PROTOCOL_CHROMATIN_AMOUNT The amount of chromatin used in the MRE protocol. MRE_PROTOCOL_RESTRICTION_ENZYME The restriction enzyme(s) used in the MRE protocol. MRE_PROTOCOL_SIZE_FRACTION The size of the fragments selected in the MRE protocol. Chip-Seq Input EXPERIMENT_TYPE: ChIP-Seq Input EXPERIMENT_ONTOLOGY_URIS - container element for EXPERIMENT_ONTOLOGY_URI elements that contain links to sample ontology information. EXTRACTION_PROTOCOL - The protocol used to isolate the extract material. EXTRACTION_PROTOCOL_TYPE_OF_SONICATOR - The type of sonicator used for extraction. EXTRACTION_PROTOCOL_SONICATION_CYCLES - The number of sonication cycles used for extraction. CHIP_PROTOCOL: Input CHIP_PROTOCOL_CHROMATIN_AMOUNT The amount of chromatin used in the ChIP protocol. Chip-Seq EXPERIMENT_TYPE: 'Histone H3K4me1','Histone H3K4me3','Histone H3K9me3','Histone H3K9ac','Histone H3K27me3', 'Histone H3K36me3', etc. EXPERIMENT_ONTOLOGY_URI - links to experiment ontology information. 10
11 EXTRACTION_PROTOCOL - The protocol used to isolate the extract material. EXTRACTION_PROTOCOL_TYPE_OF_SONICATOR - The type of sonicator used for extraction. EXTRACTION_PROTOCOL_SONICATION_CYCLES - The number of sonication cycles used for extraction. CHIP_PROTOCOL The ChIP protocol used. CHIP_PROTOCOL_CHROMATIN_AMOUNT - The amount of chromatin used in the ChIP protocol. CHIP_PROTOCOL_BEAD_TYPE - The type of bead used in the ChIP protocol. CHIP_PROTOCOL_BEAD_AMOUNT - The amount of beads used in the ChIP protocol. CHIP_PROTOCOL_ANTIBODY_AMOUNT The amount of antibody used in the ChIP protocol. CHIP_ANTIBODY - The specific antibody used in the ChIP protocol. CHIP_ANTIBODY_PROVIDER - The name of the company, laboratory or person that provided the antibody. CHIP_ANTIBODY_CATALOG The catalog from which the antibody was purchased. CHIP_ANTIBODY_LOT The lot identifier of the antibody. CHIP_PROTOCOL_CROSSLINK_TIME - The timespan in which the chromatin is crosslinked LIBRARY_GENERATION_FRAGMENT_SIZE_RANGE The fragment size range of the preparation. mrna-seq EXPERIMENT_TYPE: mrna-seq EXPERIMENT_ONTOLOGY_URI - links to experiment ontology information. EXTRACTION_PROTOCOL - The protocol used to isolate the extract material. EXTRACTION_PROTOCOL_MRNA_ENRICHMENT The mrna enrichment method used in the extraction protocol. 11
12 EXTRACTION_PROTOCOL_FRAGMENTATION The fragmentation method used in the extraction protocol. MRNA_PREPARATION_FRAGMENT_SIZE_RANGE The mrna fragment size range of the preparation. RNA_PREPARATION_5'_RNA_ADAPTER_SEQUENCE The sequence of the 5 RNA adapter used in preparation. RNA_PREPARATION_3'_RNA_ADAPTER_SEQUENCE - The sequence of the 3 RNA adapter used in preparation. RNA_PREPARATION_REVERSE_TRANSCRIPTION_PRIMER_SEQUENCE The sequence of the primer for reverse transcription used in preparation. RNA_PREPARATION_5'_DEPHOSPHORYLATION The protocol for 5 dephosphorylation used in preparation. RNA_PREPARATION_5'_PHOSPHORYLATION The protocol for 5 phosphorylation used in preparation. RNA_PREPARATION_3'_RNA ADAPTER_LIGATION_PROTOCOL The protocol for 3 adapter ligation used in preparation. RNA_PREPARATION_5'_RNA_ADAPTER_LIGATION_PROTOCOL - The protocol for 5 adapter ligation used in preparation. LIBRARY_GENERATION_PCR_TEMPLATE_CONC The PCR template concentration for library generation. LIBRARY_GENERATION_PCR_POLYMERASE_TYPE The PCR polymerase used for library generation LIBRARY_GENERATION_PCR_THERMOCYCLING_PROGRAM The thermocycling program used for library generation. LIBRARY_GENERATION_PCR_NUMBER_CYCLES The number of PCR cycles used for library generation. LIBRARY_GENERATION_PCR_F_PRIMER_SEQUENCE The sequence of the PCR forward primer used for library generation. LIBRARY_GENERATION_PCR_R_PRIMER_SEQUENCE The sequence of the PCR reverse primer used for library generation. 12
13 LIBRARY_GENERATION_PCR_PRIMER_CONC The concentration of the PCR primers used for library generation. LIBRARY_GENERATION_PCR_PRODUCT_ISOLATION_PROTOCOL The protocol for isolating PCR products used for library generation. TEMPLATE_TYPE - mrna or cdna - The type of template. AMPLIFIED - True or False - Is the sample amplified? PREPARATION_INITIAL_MRNA_QNTY -The initial mrna quantity used in preparation. PREPARATION_REVERSE_TRANSCRIPTION_PROTOCOL - The protocol for reverse transcription used in preparation. PREPARATION_PCR_NUMBER_CYCLES - The number of PCR cycles used to amplify. LIBRARY_GENERATION_PROTOCOL - The protocol used to generate the library. LIBRARY_GENERATION_FRAGMENTATION - The fragmentation method used in the library protocol. LIBRARY_GENERATION_FRAGMENT_SIZE_RANGE The fragment size range of the preparation. LIBRARY_GENERATION_3'_ADAPTER_SEQUENCE The sequence of the 3' adapter used for library generation. LIBRARY_GENERATION_5'_ADAPTER_SEQUENCE The sequence of the 5' adapter used for library generation. smrna-seq EXPERIMENT_TYPE:smRNA-Seq EXPERIMENT_ONTOLOGY_URI - links to experiment ontology information. EXTRACTION_PROTOCOL - The protocol used to isolate the extract material. EXTRACTION_PROTOCOL_SMRNA_ENRICHMENT - The smrna enrichment method used in the extraction protocol. SMRNA_PREPARATION_INITIAL_SMRNA_QNTY - The initial smrna quantity 13
14 used in preparation. RNA_PREPARATION_5'_RNA_ADAPTER_SEQUENCE The sequence of the 5 RNA adapter used in preparation. RNA_PREPARATION_3'_RNA_ADAPTER_SEQUENCE - The sequence of the 3 RNA adapter used in preparation. RNA_PREPARATION_REVERSE_TRANSCRIPTION_PRIMER_SEQUENCE The sequence of the primer for reverse transcription used in preparation. RNA_PREPARATION_3'_RNA ADAPTER_LIGATION_PROTOCOL The protocol for 3 adapter ligation used in preparation. RNA_PREPARATION_5'_RNA_ADAPTER_LIGATION_PROTOCOL - The protocol for 5 adapter ligation used in preparation. RNA_PREPARATION_REVERSE_TRANSCRIPTION_PROTOCOL - The protocol for reverse transcription used in preparation. LIBRARY_GENERATION_PCR_TEMPLATE_CONC The PCR template concentration for library generation. LIBRARY_GENERATION_PCR_POLYMERASE_TYPE The PCR polymerase used for library generation LIBRARY_GENERATION_PCR_THERMOCYCLING_PROGRAM The thermocycling program used for library generation. LIBRARY_GENERATION_PCR_NUMBER_CYCLES The number of PCR cycles used for library generation. LIBRARY_GENERATION_PCR_F_PRIMER_SEQUENCE The sequence of the PCR forward primer used for library generation. LIBRARY_GENERATION_PCR_R_PRIMER_SEQUENCE The sequence of the PCR reverse primer used for library generation. LIBRARY_GENERATION_PCR_PRIMER_CONC The concentration of the PCR primers used for library generation. LIBRARY_GENERATION_PCR_PRODUCT_ISOLATION_PROTOCOL The protocol for isolating PCR products used for library generation. Level 1 Data (SRA: ANALYSIS_TYPE -REFERENCE_ALIGNMENT) 14
15 DATA_ANALYSIS_LEVEL - 1 EXPERIMENT_TYPE - The type of experiment (Chromatin Accessibility, Bisulfite-Seq, MeDIP-Seq, MRE-Seq, ChIP-Seq, mrna-seq, smrna-seq). EXPERIMENT_ONTOLOGY_URI - links to experiment ontology information. GENOME_ASSEMBLY The genome assembly to which the reads are mapped. SOFTWARE The name of the software used for mapping. SOFTWARE_VERSION The version of the software used for mapping. SOFTWARE_COMMAND_LINE - The command line used to run the software. ANALYSIS_PROTOCOL - Description of how the analysis was performed. MAXIMUM_ALIGNMENT_LENGTH The maximum read alignment length supported by the software. If the software aligns the entire read use Read Length. MISMATCHES_ALLOWED The number of mismatches allowed in an alignment. ALIGNMENTS_ALLOWED The number of locations to which a read is allowed to align. TREATMENT_OF_MULTIPLE_ALIGNMENTS How reads aligning to multiple locations are treated. TREATMENT_OF_IDENTICAL_ALIGNMENTS_OF_MULTIPLE_READS How multiple reads aligning to the same location are treated. This applies to clonal duplicate removal. ALIGNMENT_POSTPROCESSING Any postprocessing applied to the alignments. NUMBER_OF_MAPPED_READS The number of mapped reads. Quality Control Quality control related analysis attributes. These are dependent on the type of experiment. Level 2 Data (SRA: ANALYSIS_TYPE - ABUNDANCE_MEASUREMENT) DATA_ANALYSIS_LEVEL: 2 15
16 EXPERIMENT_TYPE - The type of experiment (Chromatin Accessibility, Bisulfite-Seq, MeDIP-Seq, MRE-Seq, ChIP-Seq, mrna-seq, smrna-seq). EXPERIMENT_ONTOLOGY_URI - links to experiment ontology information. GENOME_ASSEMBLY The genome assembly to which the reads are mapped. SOFTWARE The name of the software used for determining signal (read density). SOFTWARE_VERSION The version of the software used for determining signal (read density). SOFTWARE_COMMAND_LINE - The command line used to run the software. ANALYSIS_PROTOCOL - Description of how the analysis was performed. READ_EXTENSION If read mappings are extended before determining signal (read density), the length to which the read is extended in bp. NA if not applicable. GENOMIC_WINDOW The bp size of the window in which the signal (read density) is calculated. TREATMENT_OF_REGIONS_PRONE_TO_MULTIPLE_ALIGNMENTS Any treatment of regions, such as repeats, which are prone to multiple alignments. NA if not applicable. NUMBER_OF_MAPPED_READS The number of mapped reads. Quality Control Quality control related analysis attributes. These are dependent on the type of experiment. 4. Accepted Ontologies Sample Ontologies Cell Lines - Experimental Factor Ontology (EFO) Primary Cells - Cell Ontology (CL) Primary Tissue - Uberon Disease Ontologies Disease - NCI Metathesaurus Experiment Ontologies Assays and Platforms - Ontology for Biomedical Investigations (OBI) 16
Assay Standards Working Group Nov 2012 Assay Standards Working Group Recommendations, November 2012
 Assay Standards Working Group Recommendations, November 2012 Contents Assay Standards Working Group Recommendations, August 2012... 1 Contents... 1 Introduction... 2 1: Reference Epigenome Criteria...
Assay Standards Working Group Recommendations, November 2012 Contents Assay Standards Working Group Recommendations, August 2012... 1 Contents... 1 Introduction... 2 1: Reference Epigenome Criteria...
Deep Sequencing technologies
 Deep Sequencing technologies Gabriela Salinas 30 October 2017 Transcriptome and Genome Analysis Laboratory http://www.uni-bc.gwdg.de/index.php?id=709 Microarray and Deep-Sequencing Core Facility University
Deep Sequencing technologies Gabriela Salinas 30 October 2017 Transcriptome and Genome Analysis Laboratory http://www.uni-bc.gwdg.de/index.php?id=709 Microarray and Deep-Sequencing Core Facility University
NGS Approaches to Epigenomics
 I519 Introduction to Bioinformatics, 2013 NGS Approaches to Epigenomics Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Contents Background: chromatin structure & DNA methylation Epigenomic
I519 Introduction to Bioinformatics, 2013 NGS Approaches to Epigenomics Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Contents Background: chromatin structure & DNA methylation Epigenomic
Applied Bioinformatics - Lecture 16: Transcriptomics
 Applied Bioinformatics - Lecture 16: Transcriptomics David Hendrix Oregon State University Feb 15th 2016 Transcriptomics High-throughput Sequencing (deep sequencing) High-throughput sequencing (also
Applied Bioinformatics - Lecture 16: Transcriptomics David Hendrix Oregon State University Feb 15th 2016 Transcriptomics High-throughput Sequencing (deep sequencing) High-throughput sequencing (also
Next-generation sequencing technologies
 Next-generation sequencing technologies Illumina: Summary https://www.youtube.com/watch?v=fcd6b5hraz8 Illumina platforms: Benchtop sequencers https://www.illumina.com/systems/sequencing-platforms.html
Next-generation sequencing technologies Illumina: Summary https://www.youtube.com/watch?v=fcd6b5hraz8 Illumina platforms: Benchtop sequencers https://www.illumina.com/systems/sequencing-platforms.html
Gene expression analysis. Biosciences 741: Genomics Fall, 2013 Week 5. Gene expression analysis
 Gene expression analysis Biosciences 741: Genomics Fall, 2013 Week 5 Gene expression analysis From EST clusters to spotted cdna microarrays Long vs. short oligonucleotide microarrays vs. RT-PCR Methods
Gene expression analysis Biosciences 741: Genomics Fall, 2013 Week 5 Gene expression analysis From EST clusters to spotted cdna microarrays Long vs. short oligonucleotide microarrays vs. RT-PCR Methods
Next-generation sequencing technologies
 Next-generation sequencing technologies NGS applications Illumina sequencing workflow Overview Sequencing by ligation Short-read NGS Sequencing by synthesis Illumina NGS Single-molecule approach Long-read
Next-generation sequencing technologies NGS applications Illumina sequencing workflow Overview Sequencing by ligation Short-read NGS Sequencing by synthesis Illumina NGS Single-molecule approach Long-read
Gene Expression Technology
 Gene Expression Technology Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Gene expression Gene expression is the process by which information from a gene
Gene Expression Technology Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Gene expression Gene expression is the process by which information from a gene
M Keramatipour 2. M Keramatipour 1. M Keramatipour 4. M Keramatipour 3. M Keramatipour 5. M Keramatipour
 Molecular Cloning Methods Mohammad Keramatipour MD, PhD keramatipour@tums.ac.ir Outline DNA recombinant technology DNA cloning co Cell based PCR PCR-based Some application of DNA cloning Genomic libraries
Molecular Cloning Methods Mohammad Keramatipour MD, PhD keramatipour@tums.ac.ir Outline DNA recombinant technology DNA cloning co Cell based PCR PCR-based Some application of DNA cloning Genomic libraries
Subtractive hybridization is a powerful technique particularly well-suited for the identification of target cdnas that correspond to rare
 Subtractive hybridization is a powerful technique particularly well-suited for the identification of target cdnas that correspond to rare transcripts, that are expressed in one population but not in the
Subtractive hybridization is a powerful technique particularly well-suited for the identification of target cdnas that correspond to rare transcripts, that are expressed in one population but not in the
02 Agenda Item 03 Agenda Item
 01 Agenda Item 02 Agenda Item 03 Agenda Item SOLiD 3 System: Applications Overview April 12th, 2010 Jennifer Stover Field Application Specialist - SOLiD Applications Workflow for SOLiD Application Application
01 Agenda Item 02 Agenda Item 03 Agenda Item SOLiD 3 System: Applications Overview April 12th, 2010 Jennifer Stover Field Application Specialist - SOLiD Applications Workflow for SOLiD Application Application
2/10/17. Contents. Applications of HMMs in Epigenomics
 2/10/17 I529: Machine Learning in Bioinformatics (Spring 2017) Contents Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2017 Background:
2/10/17 I529: Machine Learning in Bioinformatics (Spring 2017) Contents Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2017 Background:
RNA-Sequencing analysis
 RNA-Sequencing analysis Markus Kreuz 25. 04. 2012 Institut für Medizinische Informatik, Statistik und Epidemiologie Content: Biological background Overview transcriptomics RNA-Seq RNA-Seq technology Challenges
RNA-Sequencing analysis Markus Kreuz 25. 04. 2012 Institut für Medizinische Informatik, Statistik und Epidemiologie Content: Biological background Overview transcriptomics RNA-Seq RNA-Seq technology Challenges
Molecular Cell Biology - Problem Drill 11: Recombinant DNA
 Molecular Cell Biology - Problem Drill 11: Recombinant DNA Question No. 1 of 10 1. Which of the following statements about the sources of DNA used for molecular cloning is correct? Question #1 (A) cdna
Molecular Cell Biology - Problem Drill 11: Recombinant DNA Question No. 1 of 10 1. Which of the following statements about the sources of DNA used for molecular cloning is correct? Question #1 (A) cdna
Reading Lecture 8: Lecture 9: Lecture 8. DNA Libraries. Definition Types Construction
 Lecture 8 Reading Lecture 8: 96-110 Lecture 9: 111-120 DNA Libraries Definition Types Construction 142 DNA Libraries A DNA library is a collection of clones of genomic fragments or cdnas from a certain
Lecture 8 Reading Lecture 8: 96-110 Lecture 9: 111-120 DNA Libraries Definition Types Construction 142 DNA Libraries A DNA library is a collection of clones of genomic fragments or cdnas from a certain
Introduction to genome biology
 Introduction to genome biology Lisa Stubbs Deep transcritpomes for traditional model species from ENCODE (and modencode) Deep RNA-seq and chromatin analysis on 147 human cell types, as well as tissues,
Introduction to genome biology Lisa Stubbs Deep transcritpomes for traditional model species from ENCODE (and modencode) Deep RNA-seq and chromatin analysis on 147 human cell types, as well as tissues,
Bi 8 Lecture 5. Ellen Rothenberg 19 January 2016
 Bi 8 Lecture 5 MORE ON HOW WE KNOW WHAT WE KNOW and intro to the protein code Ellen Rothenberg 19 January 2016 SIZE AND PURIFICATION BY SYNTHESIS: BASIS OF EARLY SEQUENCING complex mixture of aborted DNA
Bi 8 Lecture 5 MORE ON HOW WE KNOW WHAT WE KNOW and intro to the protein code Ellen Rothenberg 19 January 2016 SIZE AND PURIFICATION BY SYNTHESIS: BASIS OF EARLY SEQUENCING complex mixture of aborted DNA
Integrated NGS Sample Preparation Solutions for Limiting Amounts of RNA and DNA. March 2, Steven R. Kain, Ph.D. ABRF 2013
 Integrated NGS Sample Preparation Solutions for Limiting Amounts of RNA and DNA March 2, 2013 Steven R. Kain, Ph.D. ABRF 2013 NuGEN s Core Technologies Selective Sequence Priming Nucleic Acid Amplification
Integrated NGS Sample Preparation Solutions for Limiting Amounts of RNA and DNA March 2, 2013 Steven R. Kain, Ph.D. ABRF 2013 NuGEN s Core Technologies Selective Sequence Priming Nucleic Acid Amplification
TECH NOTE Ligation-Free ChIP-Seq Library Preparation
 TECH NOTE Ligation-Free ChIP-Seq Library Preparation The DNA SMART ChIP-Seq Kit Ligation-free template switching technology: Minimize sample handling in a single-tube workflow >> Simplified protocol with
TECH NOTE Ligation-Free ChIP-Seq Library Preparation The DNA SMART ChIP-Seq Kit Ligation-free template switching technology: Minimize sample handling in a single-tube workflow >> Simplified protocol with
2/19/13. Contents. Applications of HMMs in Epigenomics
 2/19/13 I529: Machine Learning in Bioinformatics (Spring 2013) Contents Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2013 Background:
2/19/13 I529: Machine Learning in Bioinformatics (Spring 2013) Contents Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2013 Background:
ChIP-seq/Functional Genomics/Epigenomics. CBSU/3CPG/CVG Next-Gen Sequencing Workshop. Josh Waterfall. March 31, 2010
 ChIP-seq/Functional Genomics/Epigenomics CBSU/3CPG/CVG Next-Gen Sequencing Workshop Josh Waterfall March 31, 2010 Outline Introduction to ChIP-seq Control data sets Peak/enriched region identification
ChIP-seq/Functional Genomics/Epigenomics CBSU/3CPG/CVG Next-Gen Sequencing Workshop Josh Waterfall March 31, 2010 Outline Introduction to ChIP-seq Control data sets Peak/enriched region identification
Applications of HMMs in Epigenomics
 I529: Machine Learning in Bioinformatics (Spring 2013) Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2013 Contents Background:
I529: Machine Learning in Bioinformatics (Spring 2013) Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2013 Contents Background:
RNA Sequencing. Next gen insight into transcriptomes , Elio Schijlen
 RNA Sequencing Next gen insight into transcriptomes 05-06-2013, Elio Schijlen Transcriptome complete set of transcripts in a cell, and their quantity, for a specific developmental stage or physiological
RNA Sequencing Next gen insight into transcriptomes 05-06-2013, Elio Schijlen Transcriptome complete set of transcripts in a cell, and their quantity, for a specific developmental stage or physiological
ENCODE and modencode Guidelines For Experiments Generating Chromatin and Transcription Factor Data Version 1.0, April 09, 2010
 ENCODE and modencode Guidelines For Experiments Generating Chromatin and Transcription Factor Data Version 1.0, April 09, 2010 Every data producer aims to generate high- quality data sets. To help achieve
ENCODE and modencode Guidelines For Experiments Generating Chromatin and Transcription Factor Data Version 1.0, April 09, 2010 Every data producer aims to generate high- quality data sets. To help achieve
APPLICATION NOTE. Abstract. Introduction
 From minuscule amounts to magnificent results: reliable ChIP-seq data from 1, cells with the True MicroChIP and the MicroPlex Library Preparation kits Abstract Diagenode has developed groundbreaking solutions
From minuscule amounts to magnificent results: reliable ChIP-seq data from 1, cells with the True MicroChIP and the MicroPlex Library Preparation kits Abstract Diagenode has developed groundbreaking solutions
Design. Construction. Characterization
 Design Construction Characterization DNA mrna (messenger) A C C transcription translation C A C protein His A T G C T A C G Plasmids replicon copy number incompatibility selection marker origin of replication
Design Construction Characterization DNA mrna (messenger) A C C transcription translation C A C protein His A T G C T A C G Plasmids replicon copy number incompatibility selection marker origin of replication
Novel methods for RNA and DNA- Seq analysis using SMART Technology. Andrew Farmer, D. Phil. Vice President, R&D Clontech Laboratories, Inc.
 Novel methods for RNA and DNA- Seq analysis using SMART Technology Andrew Farmer, D. Phil. Vice President, R&D Clontech Laboratories, Inc. Agenda Enabling Single Cell RNA-Seq using SMART Technology SMART
Novel methods for RNA and DNA- Seq analysis using SMART Technology Andrew Farmer, D. Phil. Vice President, R&D Clontech Laboratories, Inc. Agenda Enabling Single Cell RNA-Seq using SMART Technology SMART
Genome 373: High- Throughput DNA Sequencing. Doug Fowler
 Genome 373: High- Throughput DNA Sequencing Doug Fowler Tasks give ML unity We learned about three tasks that are commonly encountered in ML Models/Algorithms Give ML Diversity Classification Regression
Genome 373: High- Throughput DNA Sequencing Doug Fowler Tasks give ML unity We learned about three tasks that are commonly encountered in ML Models/Algorithms Give ML Diversity Classification Regression
Chapter 20 Recombinant DNA Technology. Copyright 2009 Pearson Education, Inc.
 Chapter 20 Recombinant DNA Technology Copyright 2009 Pearson Education, Inc. 20.1 Recombinant DNA Technology Began with Two Key Tools: Restriction Enzymes and DNA Cloning Vectors Recombinant DNA refers
Chapter 20 Recombinant DNA Technology Copyright 2009 Pearson Education, Inc. 20.1 Recombinant DNA Technology Began with Two Key Tools: Restriction Enzymes and DNA Cloning Vectors Recombinant DNA refers
Functional Genomics Overview RORY STARK PRINCIPAL BIOINFORMATICS ANALYST CRUK CAMBRIDGE INSTITUTE 18 SEPTEMBER 2017
 Functional Genomics Overview RORY STARK PRINCIPAL BIOINFORMATICS ANALYST CRUK CAMBRIDGE INSTITUTE 18 SEPTEMBER 2017 Agenda What is Functional Genomics? RNA Transcription/Gene Expression Measuring Gene
Functional Genomics Overview RORY STARK PRINCIPAL BIOINFORMATICS ANALYST CRUK CAMBRIDGE INSTITUTE 18 SEPTEMBER 2017 Agenda What is Functional Genomics? RNA Transcription/Gene Expression Measuring Gene
Introduction to genome biology
 Introduction to genome biology Lisa Stubbs We ve found most genes; but what about the rest of the genome? Genome size* 12 Mb 95 Mb 170 Mb 1500 Mb 2700 Mb 3200 Mb #coding genes ~7000 ~20000 ~14000 ~26000
Introduction to genome biology Lisa Stubbs We ve found most genes; but what about the rest of the genome? Genome size* 12 Mb 95 Mb 170 Mb 1500 Mb 2700 Mb 3200 Mb #coding genes ~7000 ~20000 ~14000 ~26000
ChIP DNA Clean & Concentrator Catalog Nos. D5201 & D5205
 Page 0 INSTRUCTION MANUAL ChIP DNA Clean & Concentrator Catalog Nos. D5201 & D5205 Highlights Quick (2 minute) recovery of ultra-pure DNA from chromatin immunoprecipitation (ChIP) assays, cell lysates,
Page 0 INSTRUCTION MANUAL ChIP DNA Clean & Concentrator Catalog Nos. D5201 & D5205 Highlights Quick (2 minute) recovery of ultra-pure DNA from chromatin immunoprecipitation (ChIP) assays, cell lysates,
Applications of ChIP. November 05, David Grotsky, PhD Scientific Support Specialist - Epigenetics
 Applications of ChIP November 05, 2014 David Grotsky, PhD Scientific Support Specialist - Epigenetics Overview Introduction to chromatin and ChIP The histone code hypothesis What we learn from ChIP-on-chip
Applications of ChIP November 05, 2014 David Grotsky, PhD Scientific Support Specialist - Epigenetics Overview Introduction to chromatin and ChIP The histone code hypothesis What we learn from ChIP-on-chip
454 Sample Prep / Workflow at the BioMedical Genomics Center (BMGC) University of Minnesota. Sushmita Singh
 454 Sample Prep / Workflow at the BioMedical Genomics Center (BMGC) University of Minnesota Sushmita Singh 1 Consultation 2 Sample Prep (client) 3 Sample Prep (Core) Sequencing 4 Data QC 1 Consultation
454 Sample Prep / Workflow at the BioMedical Genomics Center (BMGC) University of Minnesota Sushmita Singh 1 Consultation 2 Sample Prep (client) 3 Sample Prep (Core) Sequencing 4 Data QC 1 Consultation
SOLiD Total RNA-Seq Kit SOLiD RNA Barcoding Kit
 SOLiD Total RNA-Seq Kit SOLiD RNA Barcoding Kit Agenda SOLiD Total RNAseq Kit Overview Kit Configurations Barcoding Kit Introduction New Small RNA and WT Workflow Small RNA Workflow Step-by-step Workflow
SOLiD Total RNA-Seq Kit SOLiD RNA Barcoding Kit Agenda SOLiD Total RNAseq Kit Overview Kit Configurations Barcoding Kit Introduction New Small RNA and WT Workflow Small RNA Workflow Step-by-step Workflow
EPIGENTEK. EpiQuik Chromatin Immunoprecipitation Kit. Base Catalog # P-2002 PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE
 EpiQuik Chromatin Immunoprecipitation Kit Base Catalog # P-2002 PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE The EpiQuik Chromatin Immunoprecipitation Kit is suitable for combining the specificity of
EpiQuik Chromatin Immunoprecipitation Kit Base Catalog # P-2002 PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE The EpiQuik Chromatin Immunoprecipitation Kit is suitable for combining the specificity of
EPIGENTEK. EpiNext DNA Library Preparation Kit (Illumina) Base Catalog # P-1051 PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE
 EpiNext DNA Library Preparation Kit (Illumina) Base Catalog # PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE Uses: The EpiNext DNA Library Preparation Kit (Illumina) is suitable for preparing a DNA library
EpiNext DNA Library Preparation Kit (Illumina) Base Catalog # PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE Uses: The EpiNext DNA Library Preparation Kit (Illumina) is suitable for preparing a DNA library
The Journey of DNA Sequencing. Chromosomes. What is a genome? Genome size. H. Sunny Sun
 The Journey of DNA Sequencing H. Sunny Sun What is a genome? Genome is the total genetic complement of a living organism. The nuclear genome comprises approximately 3.2 * 10 9 nucleotides of DNA, divided
The Journey of DNA Sequencing H. Sunny Sun What is a genome? Genome is the total genetic complement of a living organism. The nuclear genome comprises approximately 3.2 * 10 9 nucleotides of DNA, divided
Bootcamp: Molecular Biology Techniques and Interpretation
 Bootcamp: Molecular Biology Techniques and Interpretation Bi8 Winter 2016 Today s outline Detecting and quantifying nucleic acids and proteins: Basic nucleic acid properties Hybridization PCR and Designing
Bootcamp: Molecular Biology Techniques and Interpretation Bi8 Winter 2016 Today s outline Detecting and quantifying nucleic acids and proteins: Basic nucleic acid properties Hybridization PCR and Designing
Welcome to the NGS webinar series
 Welcome to the NGS webinar series Webinar 1 NGS: Introduction to technology, and applications NGS Technology Webinar 2 Targeted NGS for Cancer Research NGS in cancer Webinar 3 NGS: Data analysis for genetic
Welcome to the NGS webinar series Webinar 1 NGS: Introduction to technology, and applications NGS Technology Webinar 2 Targeted NGS for Cancer Research NGS in cancer Webinar 3 NGS: Data analysis for genetic
Complete Sample to Analysis Solutions for DNA Methylation Discovery using Next Generation Sequencing
 Complete Sample to Analysis Solutions for DNA Methylation Discovery using Next Generation Sequencing SureSelect Human/Mouse Methyl-Seq Kyeong Jeong PhD February 5, 2013 CAG EMEAI DGG/GSD/GFO Agilent Restricted
Complete Sample to Analysis Solutions for DNA Methylation Discovery using Next Generation Sequencing SureSelect Human/Mouse Methyl-Seq Kyeong Jeong PhD February 5, 2013 CAG EMEAI DGG/GSD/GFO Agilent Restricted
Incorporating Molecular ID Technology. Accel-NGS 2S MID Indexing Kits
 Incorporating Molecular ID Technology Accel-NGS 2S MID Indexing Kits Molecular Identifiers (MIDs) MIDs are indices used to label unique library molecules MIDs can assess duplicate molecules in sequencing
Incorporating Molecular ID Technology Accel-NGS 2S MID Indexing Kits Molecular Identifiers (MIDs) MIDs are indices used to label unique library molecules MIDs can assess duplicate molecules in sequencing
Functional Genomics Research Stream. Research Meeting: November 8, 2011 cdna Library Construction for RNA-Seq
 Functional Genomics Research Stream Research Meeting: November 8, 2011 cdna Library Construction for RNA-Seq Lab Issues Don t leave out boxes of water + ethidium bromide Minimize tip boxes in phenol waste
Functional Genomics Research Stream Research Meeting: November 8, 2011 cdna Library Construction for RNA-Seq Lab Issues Don t leave out boxes of water + ethidium bromide Minimize tip boxes in phenol waste
Non-Organic-Based Isolation of Mammalian microrna using Norgen s microrna Purification Kit
 Application Note 13 RNA Sample Preparation Non-Organic-Based Isolation of Mammalian microrna using Norgen s microrna Purification Kit B. Lam, PhD 1, P. Roberts, MSc 1 Y. Haj-Ahmad, M.Sc., Ph.D 1,2 1 Norgen
Application Note 13 RNA Sample Preparation Non-Organic-Based Isolation of Mammalian microrna using Norgen s microrna Purification Kit B. Lam, PhD 1, P. Roberts, MSc 1 Y. Haj-Ahmad, M.Sc., Ph.D 1,2 1 Norgen
ChampionChIP Quick, High Throughput Chromatin Immunoprecipitation Assay System
 ChampionChIP Quick, High Throughput Chromatin Immunoprecipitation Assay System Liyan Pang, Ph.D. Application Scientist 1 Topics to be Covered Introduction What is ChIP-qPCR? Challenges Facing Biological
ChampionChIP Quick, High Throughput Chromatin Immunoprecipitation Assay System Liyan Pang, Ph.D. Application Scientist 1 Topics to be Covered Introduction What is ChIP-qPCR? Challenges Facing Biological
The Biotechnology Toolbox
 Chapter 15 The Biotechnology Toolbox Cutting and Pasting DNA Cutting DNA Restriction endonuclease or restriction enzymes Cellular protection mechanism for infected foreign DNA Recognition and cutting specific
Chapter 15 The Biotechnology Toolbox Cutting and Pasting DNA Cutting DNA Restriction endonuclease or restriction enzymes Cellular protection mechanism for infected foreign DNA Recognition and cutting specific
Impact of gdna Integrity on the Outcome of DNA Methylation Studies
 Impact of gdna Integrity on the Outcome of DNA Methylation Studies Application Note Nucleic Acid Analysis Authors Emily Putnam, Keith Booher, and Xueguang Sun Zymo Research Corporation, Irvine, CA, USA
Impact of gdna Integrity on the Outcome of DNA Methylation Studies Application Note Nucleic Acid Analysis Authors Emily Putnam, Keith Booher, and Xueguang Sun Zymo Research Corporation, Irvine, CA, USA
Chapter 6 - Molecular Genetic Techniques
 Chapter 6 - Molecular Genetic Techniques Two objects of molecular & genetic technologies For analysis For generation Molecular genetic technologies! For analysis DNA gel electrophoresis Southern blotting
Chapter 6 - Molecular Genetic Techniques Two objects of molecular & genetic technologies For analysis For generation Molecular genetic technologies! For analysis DNA gel electrophoresis Southern blotting
NEXTFLEX ChIP-Seq Kit (For Illumina Platforms) Catalog #NOVA (Kit contains 8 reactions) Bioo Scientific Corp V15.
 NEXTFLEX ChIP-Seq Kit (For Illumina Platforms) Catalog #NOVA-5143-01 (Kit contains 8 reactions) Bioo Scientific Corp. 2015-2018 V15.07 This product is for research use only. Not for use in diagnostic procedures.
NEXTFLEX ChIP-Seq Kit (For Illumina Platforms) Catalog #NOVA-5143-01 (Kit contains 8 reactions) Bioo Scientific Corp. 2015-2018 V15.07 This product is for research use only. Not for use in diagnostic procedures.
Green Center Computational Core ChIP- Seq Pipeline, Just a Click Away
 Green Center Computational Core ChIP- Seq Pipeline, Just a Click Away Venkat Malladi Computational Biologist Computational Core Cecil H. and Ida Green Center for Reproductive Biology Science Introduc
Green Center Computational Core ChIP- Seq Pipeline, Just a Click Away Venkat Malladi Computational Biologist Computational Core Cecil H. and Ida Green Center for Reproductive Biology Science Introduc
Mammalian Chromatin Extraction Kit
 Mammalian Chromatin Extraction Kit Catalog Number KA3925 100 extractions Version: 01 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Intended Use... 3 Background... 3
Mammalian Chromatin Extraction Kit Catalog Number KA3925 100 extractions Version: 01 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Intended Use... 3 Background... 3
Chapter 8: Recombinant DNA. Ways this technology touches us. Overview. Genetic Engineering
 Chapter 8 Recombinant DNA and Genetic Engineering Genetic manipulation Ways this technology touches us Criminal justice The Justice Project, started by law students to advocate for DNA testing of Death
Chapter 8 Recombinant DNA and Genetic Engineering Genetic manipulation Ways this technology touches us Criminal justice The Justice Project, started by law students to advocate for DNA testing of Death
Supplemental Figure 1.
 Supplemental Data. Charron et al. Dynamic landscapes of four histone modifications during de-etiolation in Arabidopsis. Plant Cell (2009). 10.1105/tpc.109.066845 Supplemental Figure 1. Immunodetection
Supplemental Data. Charron et al. Dynamic landscapes of four histone modifications during de-etiolation in Arabidopsis. Plant Cell (2009). 10.1105/tpc.109.066845 Supplemental Figure 1. Immunodetection
Applications of Next Generation Sequencing in Metagenomics Studies
 Applications of Next Generation Sequencing in Metagenomics Studies Francesca Rizzo, PhD Genomix4life Laboratory of Molecular Medicine and Genomics Department of Medicine and Surgery University of Salerno
Applications of Next Generation Sequencing in Metagenomics Studies Francesca Rizzo, PhD Genomix4life Laboratory of Molecular Medicine and Genomics Department of Medicine and Surgery University of Salerno
DNA METHYLATION NH 2 H 3. Solutions using bisulfite conversion and immunocapture Ideal for NGS, Sanger sequencing, Pyrosequencing, and qpcr
 NH 2 DNA METHYLATION H 3 C Solutions using bisulfite conversion and immunocapture Ideal for NGS, Sanger sequencing, Pyrosequencing, and qpcr NH 2 N N H PAGE 3 Understanding DNA Methylation DNA methylation
NH 2 DNA METHYLATION H 3 C Solutions using bisulfite conversion and immunocapture Ideal for NGS, Sanger sequencing, Pyrosequencing, and qpcr NH 2 N N H PAGE 3 Understanding DNA Methylation DNA methylation
Development of quantitative targeted RNA-seq methodology for use in differential gene expression
 Development of quantitative targeted RNA-seq methodology for use in differential gene expression Dr. Jens Winter, Market Development Group Biological Biological Research Content EMEA QIAGEN Universal Workflows
Development of quantitative targeted RNA-seq methodology for use in differential gene expression Dr. Jens Winter, Market Development Group Biological Biological Research Content EMEA QIAGEN Universal Workflows
Basics of RNA-Seq. (With a Focus on Application to Single Cell RNA-Seq) Michael Kelly, PhD Team Lead, NCI Single Cell Analysis Facility
 2018 ABRF Meeting Satellite Workshop 4 Bridging the Gap: Isolation to Translation (Single Cell RNA-Seq) Sunday, April 22 Basics of RNA-Seq (With a Focus on Application to Single Cell RNA-Seq) Michael Kelly,
2018 ABRF Meeting Satellite Workshop 4 Bridging the Gap: Isolation to Translation (Single Cell RNA-Seq) Sunday, April 22 Basics of RNA-Seq (With a Focus on Application to Single Cell RNA-Seq) Michael Kelly,
DNA:CHROMATIN INTERACTIONS
 DNA:CHROMATIN INTERACTIONS Exploring transcription factor binding and the epigenomic landscape Chris Seward Introductions Cell and Developmental Biology PhD Candidate in Dr. Lisa Stubbs Laboratory Currently
DNA:CHROMATIN INTERACTIONS Exploring transcription factor binding and the epigenomic landscape Chris Seward Introductions Cell and Developmental Biology PhD Candidate in Dr. Lisa Stubbs Laboratory Currently
PLNT2530 (2018) Unit 6b Sequence Libraries
 PLNT2530 (2018) Unit 6b Sequence Libraries Molecular Biotechnology (Ch 4) Analysis of Genes and Genomes (Ch 5) Unless otherwise cited or referenced, all content of this presenataion is licensed under the
PLNT2530 (2018) Unit 6b Sequence Libraries Molecular Biotechnology (Ch 4) Analysis of Genes and Genomes (Ch 5) Unless otherwise cited or referenced, all content of this presenataion is licensed under the
Molecular Biology and Functional Genomic Core Facility
 Molecular Biology and Functional Genomic Core Facility General Presentation Dr Odile Neyret Core Manager Myriam Rondeau Research Assistant Agnès Dumont Research Assistant Institut de recherche clinique
Molecular Biology and Functional Genomic Core Facility General Presentation Dr Odile Neyret Core Manager Myriam Rondeau Research Assistant Agnès Dumont Research Assistant Institut de recherche clinique
GATCGTGCACGATCTCGGCAATTCGGGATGCCGGCTCGTCACCGGTCGCT
 Problem. (pts) A. (5pts) Your colleague professor Eugene Mathew Lateed generated a genome-wide DNA methylation map for normal colon cells using MRE-seq and MeDIP-seq. In an intergenic region, he found
Problem. (pts) A. (5pts) Your colleague professor Eugene Mathew Lateed generated a genome-wide DNA methylation map for normal colon cells using MRE-seq and MeDIP-seq. In an intergenic region, he found
G E N OM I C S S E RV I C ES
 GENOMICS SERVICES ABOUT T H E N E W YOR K G E NOM E C E N T E R NYGC is an independent non-profit implementing advanced genomic research to improve diagnosis and treatment of serious diseases. Through
GENOMICS SERVICES ABOUT T H E N E W YOR K G E NOM E C E N T E R NYGC is an independent non-profit implementing advanced genomic research to improve diagnosis and treatment of serious diseases. Through
Increased transcription detection with the NEBNext Single Cell/Low Input RNA Library Prep Kit
 be INSPIRED drive DISCOVERY stay GENUINE TECHNICAL NOTE Increased transcription detection with the NEBNext Single Cell/Low Input RNA Library Prep Kit Highly sensitive, robust generation of high quality
be INSPIRED drive DISCOVERY stay GENUINE TECHNICAL NOTE Increased transcription detection with the NEBNext Single Cell/Low Input RNA Library Prep Kit Highly sensitive, robust generation of high quality
DNA METHYLATION RESEARCH TOOLS
 SeqCap Epi Enrichment System Revolutionize your epigenomic research DNA METHYLATION RESEARCH TOOLS Methylated DNA The SeqCap Epi System is a set of target enrichment tools for DNA methylation assessment
SeqCap Epi Enrichment System Revolutionize your epigenomic research DNA METHYLATION RESEARCH TOOLS Methylated DNA The SeqCap Epi System is a set of target enrichment tools for DNA methylation assessment
Lieberman-Aiden et al. (2009) Comprehensive Mapping of Long-Range Interactions Reveals Folding Principles of the Human Genome. Science 326:
 Lieberman-Aiden et al. (2009) Comprehensive Mapping of Long-Range Interactions Reveals Folding Principles of the Human Genome. Science 326: 289-293. : Understanding the 3D conformation of the genome can
Lieberman-Aiden et al. (2009) Comprehensive Mapping of Long-Range Interactions Reveals Folding Principles of the Human Genome. Science 326: 289-293. : Understanding the 3D conformation of the genome can
Gene Regulation Solutions. Microarrays and Next-Generation Sequencing
 Gene Regulation Solutions Microarrays and Next-Generation Sequencing Gene Regulation Solutions The Microarrays Advantage Microarrays Lead the Industry in: Comprehensive Content SurePrint G3 Human Gene
Gene Regulation Solutions Microarrays and Next-Generation Sequencing Gene Regulation Solutions The Microarrays Advantage Microarrays Lead the Industry in: Comprehensive Content SurePrint G3 Human Gene
MeDIP-seq library construction protocol v2 Costello Lab June Notes:
 MeDIP-seq library construction protocol v2 Costello Lab June 2010 Notes: A. For all Qiagen gel extraction steps (Qiaquick and MinElute), melt gel slice at 37º C instead of 50º (see Quail et al 2008 Nature
MeDIP-seq library construction protocol v2 Costello Lab June 2010 Notes: A. For all Qiagen gel extraction steps (Qiaquick and MinElute), melt gel slice at 37º C instead of 50º (see Quail et al 2008 Nature
Introduction Bioo Scientific
 Next Generation Sequencing Catalog 2014-2015 Introduction Bioo Scientific Bioo Scientific is a global life science company headquartered in Austin, TX, committed to providing innovative products and superior
Next Generation Sequencing Catalog 2014-2015 Introduction Bioo Scientific Bioo Scientific is a global life science company headquartered in Austin, TX, committed to providing innovative products and superior
Genomic resources. for non-model systems
 Genomic resources for non-model systems 1 Genomic resources Whole genome sequencing reference genome sequence comparisons across species identify signatures of natural selection population-level resequencing
Genomic resources for non-model systems 1 Genomic resources Whole genome sequencing reference genome sequence comparisons across species identify signatures of natural selection population-level resequencing
Go to Bottom Left click WashU Epigenome Browser. Click
 Now you are going to look at the Human Epigenome Browswer. It has a more sophisticated but weirder interface than the UCSC Genome Browser. All the data that you will view as tracks is in reality just files
Now you are going to look at the Human Epigenome Browswer. It has a more sophisticated but weirder interface than the UCSC Genome Browser. All the data that you will view as tracks is in reality just files
Contents... vii. List of Figures... xii. List of Tables... xiv. Abbreviatons... xv. Summary... xvii. 1. Introduction In vitro evolution...
 vii Contents Contents... vii List of Figures... xii List of Tables... xiv Abbreviatons... xv Summary... xvii 1. Introduction...1 1.1 In vitro evolution... 1 1.2 Phage Display Technology... 3 1.3 Cell surface
vii Contents Contents... vii List of Figures... xii List of Tables... xiv Abbreviatons... xv Summary... xvii 1. Introduction...1 1.1 In vitro evolution... 1 1.2 Phage Display Technology... 3 1.3 Cell surface
Lecture 7. Next-generation sequencing technologies
 Lecture 7 Next-generation sequencing technologies Next-generation sequencing technologies General principles of short-read NGS Construct a library of fragments Generate clonal template populations Massively
Lecture 7 Next-generation sequencing technologies Next-generation sequencing technologies General principles of short-read NGS Construct a library of fragments Generate clonal template populations Massively
Matthew Tinning Australian Genome Research Facility. July 2012
 Next-Generation Sequencing: an overview of technologies and applications Matthew Tinning Australian Genome Research Facility July 2012 History of Sequencing Where have we been? 1869 Discovery of DNA 1909
Next-Generation Sequencing: an overview of technologies and applications Matthew Tinning Australian Genome Research Facility July 2012 History of Sequencing Where have we been? 1869 Discovery of DNA 1909
Chapter 17 Lecture. Concepts of Genetics. Tenth Edition. Regulation of Gene Expression in Eukaryotes
 Chapter 17 Lecture Concepts of Genetics Tenth Edition Regulation of Gene Expression in Eukaryotes Chapter Contents 17.1 Eukaryotic Gene Regulation Can Occur at Any of the Steps Leading from DNA to Protein
Chapter 17 Lecture Concepts of Genetics Tenth Edition Regulation of Gene Expression in Eukaryotes Chapter Contents 17.1 Eukaryotic Gene Regulation Can Occur at Any of the Steps Leading from DNA to Protein
GENETICS EXAM 3 FALL a) is a technique that allows you to separate nucleic acids (DNA or RNA) by size.
 Student Name: All questions are worth 5 pts. each. GENETICS EXAM 3 FALL 2004 1. a) is a technique that allows you to separate nucleic acids (DNA or RNA) by size. b) Name one of the materials (of the two
Student Name: All questions are worth 5 pts. each. GENETICS EXAM 3 FALL 2004 1. a) is a technique that allows you to separate nucleic acids (DNA or RNA) by size. b) Name one of the materials (of the two
Microarrays: since we use probes we obviously must know the sequences we are looking at!
 These background are needed: 1. - Basic Molecular Biology & Genetics DNA replication Transcription Post-transcriptional RNA processing Translation Post-translational protein modification Gene expression
These background are needed: 1. - Basic Molecular Biology & Genetics DNA replication Transcription Post-transcriptional RNA processing Translation Post-translational protein modification Gene expression
Recent technology allow production of microarrays composed of 70-mers (essentially a hybrid of the two techniques)
 Microarrays and Transcript Profiling Gene expression patterns are traditionally studied using Northern blots (DNA-RNA hybridization assays). This approach involves separation of total or polya + RNA on
Microarrays and Transcript Profiling Gene expression patterns are traditionally studied using Northern blots (DNA-RNA hybridization assays). This approach involves separation of total or polya + RNA on
Supplemental Figure 1 A
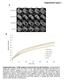 Supplemental Figure A prebleach postbleach 2 min 6 min 3 min mh2a.-gfp mh2a.2-gfp mh2a2-gfp GFP-H2A..9 Relative Intensity.8.7.6.5 mh2a. GFP n=8.4 mh2a.2 GFP n=4.3 mh2a2 GFP n=2.2 GFP H2A n=24. GFP n=7.
Supplemental Figure A prebleach postbleach 2 min 6 min 3 min mh2a.-gfp mh2a.2-gfp mh2a2-gfp GFP-H2A..9 Relative Intensity.8.7.6.5 mh2a. GFP n=8.4 mh2a.2 GFP n=4.3 mh2a2 GFP n=2.2 GFP H2A n=24. GFP n=7.
EpiNext 5-mC RNA Bisulfite-Seq Easy Kit (Illumina)
 EpiNext 5-mC RNA Bisulfite-Seq Easy Kit (Illumina) Base Catalog # PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE Uses: The EpiNext 5-mC RNA Bisulfite-Seq Easy Kit (Illumina) is designed for easily carrying
EpiNext 5-mC RNA Bisulfite-Seq Easy Kit (Illumina) Base Catalog # PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE Uses: The EpiNext 5-mC RNA Bisulfite-Seq Easy Kit (Illumina) is designed for easily carrying
NEXTFLEX Rapid Directional RNA-Seq Kit (For Illumina Platforms) Catalog #NOVA (Kit contains 8 reactions)
 NEXTFLEX Rapid Directional RNA-Seq Kit (For Illumina Platforms) Catalog #NOVA-5138-07 (Kit contains 8 reactions) Bioo Scientific Corp. 2014-2018 V18.07 This product is for research use only. Not for use
NEXTFLEX Rapid Directional RNA-Seq Kit (For Illumina Platforms) Catalog #NOVA-5138-07 (Kit contains 8 reactions) Bioo Scientific Corp. 2014-2018 V18.07 This product is for research use only. Not for use
Whole Transcriptome Analysis of Illumina RNA- Seq Data. Ryan Peters Field Application Specialist
 Whole Transcriptome Analysis of Illumina RNA- Seq Data Ryan Peters Field Application Specialist Partek GS in your NGS Pipeline Your Start-to-Finish Solution for Analysis of Next Generation Sequencing Data
Whole Transcriptome Analysis of Illumina RNA- Seq Data Ryan Peters Field Application Specialist Partek GS in your NGS Pipeline Your Start-to-Finish Solution for Analysis of Next Generation Sequencing Data
Rapid Method for the Purification of Total RNA from Formalin- Fixed Paraffin-Embedded (FFPE) Tissue Samples
 Application Note 17 RNA Sample Preparation Rapid Method for the Purification of Total RNA from Formalin- Fixed Paraffin-Embedded (FFPE) Tissue Samples M. Melmogy 1, V. Misic 1, B. Lam, PhD 1, C. Dobbin,
Application Note 17 RNA Sample Preparation Rapid Method for the Purification of Total RNA from Formalin- Fixed Paraffin-Embedded (FFPE) Tissue Samples M. Melmogy 1, V. Misic 1, B. Lam, PhD 1, C. Dobbin,
INTRODUCTION TO REVERSE TRANSCRIPTION PCR (RT-PCR) ABCF 2016 BecA-ILRI Hub, Nairobi 21 st September 2016 Roger Pelle Principal Scientist
 INTRODUCTION TO REVERSE TRANSCRIPTION PCR (RT-PCR) ABCF 2016 BecA-ILRI Hub, Nairobi 21 st September 2016 Roger Pelle Principal Scientist Objective of PCR To provide a solution to one of the most pressing
INTRODUCTION TO REVERSE TRANSCRIPTION PCR (RT-PCR) ABCF 2016 BecA-ILRI Hub, Nairobi 21 st September 2016 Roger Pelle Principal Scientist Objective of PCR To provide a solution to one of the most pressing
ChIP-seq data analysis with Chipster. Eija Korpelainen CSC IT Center for Science, Finland
 ChIP-seq data analysis with Chipster Eija Korpelainen CSC IT Center for Science, Finland chipster@csc.fi What will I learn? Short introduction to ChIP-seq Analyzing ChIP-seq data Central concepts Analysis
ChIP-seq data analysis with Chipster Eija Korpelainen CSC IT Center for Science, Finland chipster@csc.fi What will I learn? Short introduction to ChIP-seq Analyzing ChIP-seq data Central concepts Analysis
A more efficient, sensitive and robust method of chromatin immunoprecipitation (ChIP)
 A more efficient, sensitive and robust method of chromatin immunoprecipitation (ChIP) ADVANCEMENTS IN EPIGENETICS Introducing ChIP and Chromatrap Chromatrap is a more efficient, sensitive and robust method
A more efficient, sensitive and robust method of chromatin immunoprecipitation (ChIP) ADVANCEMENTS IN EPIGENETICS Introducing ChIP and Chromatrap Chromatrap is a more efficient, sensitive and robust method
Recitation CHAPTER 9 DNA Technologies
 Recitation CHAPTER 9 DNA Technologies DNA Cloning: General Scheme A cloning vector and eukaryotic chromosomes are separately cleaved with the same restriction endonuclease. (A single chromosome is shown
Recitation CHAPTER 9 DNA Technologies DNA Cloning: General Scheme A cloning vector and eukaryotic chromosomes are separately cleaved with the same restriction endonuclease. (A single chromosome is shown
HMP Data Set Documentation
 HMP Data Set Documentation Introduction This document provides detail about files available via the DACC website. The goal of the HMP consortium is to make the metagenomics sequence data generated by the
HMP Data Set Documentation Introduction This document provides detail about files available via the DACC website. The goal of the HMP consortium is to make the metagenomics sequence data generated by the
DNA. bioinformatics. epigenetics methylation structural variation. custom. assembly. gene. tumor-normal. mendelian. BS-seq. prediction.
 Epigenomics T TM activation SNP target ncrna validation metagenomics genetics private RRBS-seq de novo trio RIP-seq exome mendelian comparative genomics DNA NGS ChIP-seq bioinformatics assembly tumor-normal
Epigenomics T TM activation SNP target ncrna validation metagenomics genetics private RRBS-seq de novo trio RIP-seq exome mendelian comparative genomics DNA NGS ChIP-seq bioinformatics assembly tumor-normal
SO YOU WANT TO DO A: RNA-SEQ EXPERIMENT MATT SETTLES, PHD UNIVERSITY OF CALIFORNIA, DAVIS
 SO YOU WANT TO DO A: RNA-SEQ EXPERIMENT MATT SETTLES, PHD UNIVERSITY OF CALIFORNIA, DAVIS SETTLES@UCDAVIS.EDU Bioinformatics Core Genome Center UC Davis BIOINFORMATICS.UCDAVIS.EDU DISCLAIMER This talk/workshop
SO YOU WANT TO DO A: RNA-SEQ EXPERIMENT MATT SETTLES, PHD UNIVERSITY OF CALIFORNIA, DAVIS SETTLES@UCDAVIS.EDU Bioinformatics Core Genome Center UC Davis BIOINFORMATICS.UCDAVIS.EDU DISCLAIMER This talk/workshop
ab High Sensitivity DNA Library Preparation Kit (For Illumina )
 ab185905 High Sensitivity DNA Library Preparation Kit (For Illumina ) Instructions for Use For the preparation of a DNA library using sub-nanogram amounts of DNA input for next generation sequencing applications
ab185905 High Sensitivity DNA Library Preparation Kit (For Illumina ) Instructions for Use For the preparation of a DNA library using sub-nanogram amounts of DNA input for next generation sequencing applications
ab High Sensitivity DNA Library Preparation Kit (For Illumina )
 ab185905 High Sensitivity DNA Library Preparation Kit (For Illumina ) Instructions for Use For the preparation of a DNA library using sub-nanogram amounts of DNA input for next generation sequencing applications
ab185905 High Sensitivity DNA Library Preparation Kit (For Illumina ) Instructions for Use For the preparation of a DNA library using sub-nanogram amounts of DNA input for next generation sequencing applications
Overview of Next Generation Sequencing technologies. Céline Keime
 Overview of Next Generation Sequencing technologies Céline Keime keime@igbmc.fr Next Generation Sequencing < Second generation sequencing < General principle < Sequencing by synthesis - Illumina < Sequencing
Overview of Next Generation Sequencing technologies Céline Keime keime@igbmc.fr Next Generation Sequencing < Second generation sequencing < General principle < Sequencing by synthesis - Illumina < Sequencing
A Crash Course in NGS for GI Pathologists. Sandra O Toole
 A Crash Course in NGS for GI Pathologists Sandra O Toole The Sanger Technique First generation sequencing Uses dideoxynucleotides (dideoxyadenine, dideoxyguanine, etc) These are molecules that resemble
A Crash Course in NGS for GI Pathologists Sandra O Toole The Sanger Technique First generation sequencing Uses dideoxynucleotides (dideoxyadenine, dideoxyguanine, etc) These are molecules that resemble
DNA Arrays Affymetrix GeneChip System
 DNA Arrays Affymetrix GeneChip System chip scanner Affymetrix Inc. hybridization Affymetrix Inc. data analysis Affymetrix Inc. mrna 5' 3' TGTGATGGTGGGAATTGGGTCAGAAGGACTGTGGGCGCTGCC... GGAATTGGGTCAGAAGGACTGTGGC
DNA Arrays Affymetrix GeneChip System chip scanner Affymetrix Inc. hybridization Affymetrix Inc. data analysis Affymetrix Inc. mrna 5' 3' TGTGATGGTGGGAATTGGGTCAGAAGGACTGTGGGCGCTGCC... GGAATTGGGTCAGAAGGACTGTGGC
Targeted Sequencing Using Droplet-Based Microfluidics. Keith Brown Director, Sales
 Targeted Sequencing Using Droplet-Based Microfluidics Keith Brown Director, Sales brownk@raindancetech.com Who we are: is a Provider of Microdroplet-based Solutions The Company s RainStorm TM Technology
Targeted Sequencing Using Droplet-Based Microfluidics Keith Brown Director, Sales brownk@raindancetech.com Who we are: is a Provider of Microdroplet-based Solutions The Company s RainStorm TM Technology
Methods of Biomaterials Testing Lesson 3-5. Biochemical Methods - Molecular Biology -
 Methods of Biomaterials Testing Lesson 3-5 Biochemical Methods - Molecular Biology - Chromosomes in the Cell Nucleus DNA in the Chromosome Deoxyribonucleic Acid (DNA) DNA has double-helix structure The
Methods of Biomaterials Testing Lesson 3-5 Biochemical Methods - Molecular Biology - Chromosomes in the Cell Nucleus DNA in the Chromosome Deoxyribonucleic Acid (DNA) DNA has double-helix structure The
Supplementary data. sienigma. F-Enigma F-EnigmaSM. a-p53
 Supplementary data Supplemental Figure 1 A sienigma #2 sienigma sicontrol a-enigma - + ++ - - - - - - + ++ - - - - - - ++ B sienigma F-Enigma F-EnigmaSM a-flag HLK3 cells - - - + ++ + ++ - + - + + - -
Supplementary data Supplemental Figure 1 A sienigma #2 sienigma sicontrol a-enigma - + ++ - - - - - - + ++ - - - - - - ++ B sienigma F-Enigma F-EnigmaSM a-flag HLK3 cells - - - + ++ + ++ - + - + + - -
EPIGENTEK. EpiQuik Methylated DNA Immunoprecipitation Kit. Base Catalog # P-2019 PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE
 EpiQuik Methylated DNA Immunoprecipitation Kit Base Catalog # PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE The EpiQuik MeDIP Kit can be used for immunoprecipitating the methylated DNA from a broad range
EpiQuik Methylated DNA Immunoprecipitation Kit Base Catalog # PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE The EpiQuik MeDIP Kit can be used for immunoprecipitating the methylated DNA from a broad range
Finding Genes with Genomics Technologies
 PLNT2530 Plant Biotechnology (2018) Unit 7 Finding Genes with Genomics Technologies Unless otherwise cited or referenced, all content of this presenataion is licensed under the Creative Commons License
PLNT2530 Plant Biotechnology (2018) Unit 7 Finding Genes with Genomics Technologies Unless otherwise cited or referenced, all content of this presenataion is licensed under the Creative Commons License
Reverse transcription-pcr (rt-pcr) Dr. Hani Alhadrami
 Reverse transcription-pcr (rt-pcr) Dr. Hani Alhadrami hanialhadrami@kau.edu.sa www.hanialhadrami.kau.edu.sa Overview Several techniques are available to detect and analyse RNA. Examples of these techniques
Reverse transcription-pcr (rt-pcr) Dr. Hani Alhadrami hanialhadrami@kau.edu.sa www.hanialhadrami.kau.edu.sa Overview Several techniques are available to detect and analyse RNA. Examples of these techniques
