Annotating 7G24-63 Justin Richner May 4, Figure 1: Map of my sequence
|
|
|
- Norma Johnston
- 6 years ago
- Views:
Transcription
1 Annotating 7G24-63 Justin Richner May 4, 2005 Zfh2 exons Thd1 exons Pur-alpha exons 0 40 kb 8 = 1 kb = LINE, Penelope = DNA/Transib, Transib1 = DINE = Novel Repeat = LTR/PAO, Diver2 I = LTR/Gypsy, Invader = Transposon, Tel1 = DNA, DNAREP1 DM Figure 1: Map of my sequence I was given 80,940 bases of sequence to annotate from the Drosophila virilis dot chromosome. This consisted of two approximately 40 kb fosmids joined together; 7G24 and 63. Fosmid 7G24 comprises bases 1 to 39,070. Fosmid 63 was annotated last year (Figure 1), and three genes were found; zfh2, thd1, and pur-alpha. I also found and annotated the same three genes. Zfh2 is zinc finger homeodomain protein 2, a probable transcription factor that is required for wing development. Zfh2 stretches from to and contains nine exons. Thd1 is mismatch dependent uracil/thymine DNA glycosylase, which removes mismatched uracil or thymine in double stranded DNA. Thd1 stretches from to and contains five exons. Pur-alpha is purine-rich binding protein-α, which is a single stranded DNA binding protein thought to be involved in DNA replication. Pur-alpha begins at and extends past the end of my sequence. Two of the Pur-alpha exons are within my sequence. The entire sequence contains 32 repeated segments, one of which is a novel repeat, and five of which are DINES. The protein Zfh2 is conserved across species in the zinc finge binding domain. No conserved non-genic regions were found. This segment of the dot chromosome has high synteny with the fourth chromosome of D. melanogaster. Figure 2: Gene map from last year s submitted paper
2 2 Genes: I first tried to identify genes using the Twinscan output on the Goose server within the UCSC genome browser format (Figure 3). The first gene predicted (chr6001.1) is the tel1 gene, a protein involved in transposable elements. I will look at this gene more closely in the Repeat section. Figure 3: UCSC output on goose server The next predicted feature I analyzed was chr Twinscan predicts this to be a single exon feature, but Genescan and mrna data suggests that there are multiple exons. When Blast was performed against the nr database, the feature shows very good homology to the Zfh2 protein. But, the Zfh2 protein was much longer than the predicted one exon gene from Twinscan. I did a Blast search with the next predicted feature, chr and again found high homology to Zfh2. I decided that these were most likely the exons for this same gene and attempted to find the rest of the exons. At this point, I did not know how to use Ensembl or FlyBase, so to look for the exons, I blasted my entire repeat masked sequence to the nr database, and looked for the exons using herne on the Blast output file. The results were not expected. I had the first four exons transcribed in the forward direction from around to bases (Figure 4), and the last five exons transcribed in the reverse direction from the very end of my sequence to about bases (Figure 5). Figure 4: Two of the exons for Zfh2 transcribed in the forward direction. Figure 5: Three of the exons for Zfh2 transcribed in the reverse direction. I realized that my sequence was not assembled correctly, and XAAA63 should have been orientated in the opposite direction before it was joined with 7G24. Chris corrected my sequence but could not put the corrected sequence into the UCSC output on the Goose server. All of the numbers in the second half of my sequence were incorrect
3 3 when looking at data on the UCSC output, and I continually had to do Blast2 alignments in order to find the proper numbers. Also, the Twinscan output was wrong for Zfh2. After performing a Blast search with the corrected sequence file, I looked at the hits to Zfh2. With an e-value score of 0.0, predicted exons for nearly all of the amino acids, no stop codons within the predicted exons, and last years data, I concluded that zfh2 is a real gene. I than begin searching for exons. The first exon predicted by Twinscan was much shorter than the first exon in D. melanogaster, obtained from the Ensembl database. However, I noticed that the exon could extend for quite some distance in the +2 frame without encountering a stop codon as shown by the green arrow in Figure 6. I hypothesized that the exon actually continued through the first three exons predicted by Genescan, as shown in Figure 6. Figure 6: UCSC output of first exon of zfh2 I performed a Blast2 alignment against my hypothesized exon and the D. melanogaster first exon, and obtained a good match (Figure 7). I hypothesize that this region, from to 24577, is the first exon of zfh2. Figure 7: D. melanogaster Vs. predicted zfh2 first exon Figure 8: Blast2 of D. melanogaster 2 nd exon with my sequence. At this point I realized two things; Twinscan and Genscan are not reliable, and the method used to find the first exon was highly inefficient. I began to search for exons
4 4 much more quickly by performing Blast2 with the D, melanogaster exons from Ensembl and my entire sequence (Figure 8). Later, I came back to exon 1 and examined intron/exon boundaries to determine the exact stop site of this exon. The beginning of exon 1 was moved farther back to bases because of mrna data, Figure 9, and now the exon has a 5 un-translated region. The end of exon 1 had to be moved forward a couple of bases to because all introns begin with the base GT, see Figure 10. Figure 9: Beginning of exon 1; Red arrow = old boundary; Green arrow = new boundary Figure 10: End of exon 1 Exons 2, 3, and 4 were found without much difficulty. When searching for exon 5, only half of the exon predicted by D. melanogaster matched with my sequence. I joined exons 5 and 6 of D. melanogaster and performed a Blast2 alignment with my sequence and found a complete exon encompassing both predicted exons without any internal stop codons (Figure 11). I hypothesize that exons 5 and 6 from melanogaster have combined to form one exon in virilis.
5 5 Figure 11: Exons 5 and 6 of D. melanogaster aligned with my sequence Exons 6, 7, 8, and 9 were all pretty straight forward and matched the exons from D. melanogaster. Because exon 9 is the last exon in the ORF, it ends with a stop codon. I was unable to find any 3 un-translated region for zfh2. Table 1 shows all the identified exons for Zfh2. Table 1: Zfh2 exons; Capital letters are exons Exon Start base Sequence End base Sequence Length (bases) tgctaacgacggct GTGCTCGgtaagttc tttgttacagctgcg GGCAGgtacgtttt ccgttccaggccaa CTGAAGgtatgtc aatttcagatcca AGCTTgtcgatct gcagtcccccca ACCCAGgtaagtcg tagcaacaatt GAAGgtaccacgtcga atattcaaacagggttg TACAAgtaagtcaa gggctttcacaggtttgg TCACCGgtaagaatt cgtaaaacaagacacg GACTAAacgaaatt 89
6 6 To ensure the accuracy of the predicted exons, I joined all of the exons into one file forming the DNA sequence of the protein. Using the translate tool on Expassy, I translated the protein s DNA sequence. If the intron/exon boundaries are incorrect, than the translated protein will be full of stop codons, as occurred on the initial attempt with Zfh2 (Figure 12). Figure 12: Translated Zfh2 with predicted exons I made the intron boundaries incorrect between the 5 th and 6 th exons, which caused a frame shift. Between exons, the annotator has to be sure to keep in the same frame. When comparing Figure 13 to Figure 12, it becomes apparent that I was in the 3 frame instead of the desired 1 frame. This problem resulted from the end of exon 5 where I was off by just one base, Figure 14. Figure 13: Frame shift in exon 6 Figure 14: Wrong exon boundary at the end of exon 5 After fixing this, I recompiled the exons together and translated the sequence. The result was exactly what I wanted (Figure 15). I confirmed that this was the correct sequence by blasting the translated amino acid sequence against Zfh2 and got a nearly perfect alignment. Figure 15: Zfh2 with correct exons
7 7 The next feature I analyzed was Twinscan output chr When I performed a Blast against the nr database with this feature, a hit to CG1981 appeared with an evalue of e^-100. Flybase showed this gene to be thd1. I assumed this gene to be real because it was annotated last year, and when I ran blast with my entire sequence against the nr database, I matched this gene with multiple exons and no internal stop codons. Thd1 clearly contains more exons than just the one predicted by Twinscan. When attempting to find the first exon, I could not match the first 144 amino acids of the protein, even with a high e-value and the filter turned off (Figure 16). Because I could not find the start site by using Blast, I used the first methionine that was upstream of the area that matched in Figure 16. Fortunately, the methionine was about 140 amino acids away. Figure 16: Blast2 with D. melanogaster exon 1 and my sequence When looking at the first exon. I noticed that the score gets better and better the more you use the raw sequence instead of filtered data. In Figure 17 all panels show the output from the same Blast2 as in Figure 16. The top panel shows the score using my sequence after Repeat Masker was run and turning on the filter from the Blast2 website. The middle panel shows the same reaction but with the filter turned off. The bottom panel shows the same reaction but the filter off, and using my unmasked sequence. The rest of the exons were not difficult to find for Thd1, and Table 2 shows all of the exons. I compiled the exons as before and attempted to translate the predicted sequence of thd1. The first attempt failed, but after making adjustments to account for the gene going in the opposite direction, I was successful (Figure 18).
8 8 Figure 17: Progression of score when decreasing filtering Exon Start base Sequence End base Sequence Length (bases) aggcacgaagatggc AAGGTTgtgagtaacgtat atattattgcagaacac ACAATGgtgagttcctat atcttgaaacagcggcgg TTATAgtgagttgtaaa aaaaaccctgcaggtcgg ATACTgtaagcatattt aatttcagtatatct TCTGAtggcagcagcag 2556 Table 2: Thd1 exons Figure 18: Thd1 translated
9 9 The next feature to investigate was chr , a predicted single exon gene. I performed blast on this feature, searched for EST data, cdna data, CDS data, and mrna data and found no hits to the region around or including this feature. This suggests a false hit by Twinscan. Chr was the next feature predicted by Twinscan. This feature, like chr , had no hits to any actual data. After this, I completely gave up on Twinscan and used the Blast file, with my sequence and the nr database, to see that there was only one other hit with a good evalue score; the gene CG1507, Pur-alpha (Figure19). This protein has several different splicing patterns according to Ensembl. Figure 19: Herne view of Blast output with my sequence and nr database zoomed in at the end I could not locate the first exon for this gene, so I used the mrna data available (Figure 20). The gene starts at around in the figure and is in the 3 frame. The blue area is where my sequence and exon 2 of D. melanogaster aligned. I hypothesize that the first exon is that shown by the mrna data in Figure 20 and the area prior to the Methionine is 5 un-translated region. Figure 20: Pur-alpha exon 1 Exon 2 was found using Ensembl and mrna data. The rest of pur-alpha extends past my sequence. Table 3 shows the exon information. I compiled the exons, transcribed them, and got the desired translation.
10 10 Exon Start base Sequence End base Sequence Length (bases) tcttttattttcaga GGTATgttataaaaaaa cagccgtcagtgcag GGCCGAGgtaaatata 106 Table 3: Pur-alpha exons Conserved Non-Genic Regions: I searched for, but could not find, any CNG regions. Repeats: The large table below contains all the repeats in my sequence. The black entries are the repeats found by Repeat Masker. All of the red entries indicate repeats found upon further analysis. Repetitive features from this table make up 16.9% of my sequence. Repeat Masker ran with out the no low option found 74 additional regions of low complexity or simple repeats. Repeat ID# Position on Sequence Repeat Family Repeat LINE PENELOPE LINE PENELOPE LINE PENELOPE Novel??? Probably end of Penelope LINE PENELOPE DNA DNAREP1 DM DINE LTR/Pao DIVER2 I LTR/Pao BATUMI I Transposon Tel LINE PENELOPE LINE PENELOPE DINE DNA/Transib TRANSIB Novel??? Probably end of Transib LINE PENELOPE LINE PENELOPE LINE PENELOPE DINE LINE PENELOPE DINE LINE PENELOPE LINE PENELOPE Novel??? Probably joins entries 22 and LINE PENELOPE DINE Novel LTR/Gypsy INVADER3 I LTR/Gypsy INVADER2 I DNA DNAREP1 DM DNA DNAREP1 DM LINE PENELOPE LINE PENELOPE
11 11 When searching for proteins through the Twinscan output, the first feature analyzed hit perfectly to tel1 when run on Blast against the nr database. Tel1 is a protein involved in transposable elements. Tel1 lifts a region out of a DNA sequence and places it elsewhere. Tel1 is adjacent to repeat #8 on the table, and possibly lifts this section out of the DNA sequence. Tel1 is not a novel repeat and should have been recognized by repeat masker. Tel1 is on the table of repeating elements under entry #10. I found five DINE s in my sequence by performing a Blast2 alignment with my sequence and the generic DINE sequence supplied by Libby. After the initial matches, I performed a Blast2 with the suspected DINE regions and the known DINE sequences from different sources. The suspected DINE s had significant matches to all of the different types of DINE s in the exact same areas. The characteristic common to all DINE s is two highly conserved regions of DNA separated by a non-conserved region, as is shown in Figure 23. Figure 21: DINE with two section of conserved sequence To find novel repeats, or repeats not known by Repeat Masker, I performed a BlastN operation with my sequence against the rest of the dot chromosome of D. virillis, and found four potential novel repeats. Three of the potential novel repeats were very close to either end of repeats found in Repeat Masker, and are probably extensions of the known repeats. Repeat Masker often will not recognize the end of a repeat within a sequence due to the program s method of scoring. The other novel repeat had no matches to any known protein, and I hypothesize this to be truly novel. Interestingly, this novel repeat is found within an intron of Thd1. The four potential novel repeats are found on the table under entry # s, 4, 15, 24, and 27, with #27 being the truly novel repeat. ClustalW: For the Clustal analysis, I compared Zfh2 with different zinc finger proteins from a wide-range of species. Organisms and the proteins that I used include; Zfh2 from D. melanogaster, Zinc finger homeodomain 4 from Homo sapiens, Zinc finger homeodomain from Caenorhabditis elegans, and the Homeobox protein from Arabidopsis thaliana. The Clustal analysis with all of the species did not show any conservation except in a small area, and this was not good conservation. I hypothesized that conservation would be more evident without A. thaliana because of the great evolutionary distance between any of the other species. I ran another Clustal analysis without A. thaliana and
12 12 found a much higher conserved sequence in the same region that showed little conservation before (Figure 22). The conserved sequence represents the Zinc finger domain. This domain is conserved across animal species, but it appears not to be conserved in plants. Figure 24: Clustal without A. thaliana Synteny: My sequence has high synteny to the D. melanogaster dot chromosome, in that all the genes are in the same order and orientation. Figure 25 shows the region on the dot chromosome of D. melanogaster, and Figure 26 shows my region with just the genes. Figure 25: Ensembl map of region on 4 th chromosome of melanogaster Figure 26: Map with just my genes
13 13 In my sequence, about 17.5 kilobases separate the first translated exons of Thd and Pur-alpha, compared to 4 kilobases in D. melanogaster. This is a very large difference and is unexpected considering that D. virilis is more genetically dense than D. melanogaster in the dot chromosome. There is a large repeat section in my sequence that could account for some of the space difference. Between the last translated exons of Thd1 and Zfh2, both D. virilis and D. melanogaster contains about 8.5 kilobases of sequence. The region before Zfh2 does not contain any known genetic features for more than 30 kilobases in both species. Both These regions show high synteny between D. virilis and D. melanogaster. The region in front of Zfh2 is hypothesized to contain an important element of Zfh2, be it a 5 un-translated region or a promoter. When a P-element is inserted into this empty region, the fly does not survive. Unfortunately, I did not have enough time to analyze this section of sequence.
Annotating Fosmid 14p24 of D. Virilis chromosome 4
 Lo 1 Annotating Fosmid 14p24 of D. Virilis chromosome 4 Lo, Louis April 20, 2006 Annotation Report Introduction In the first half of Research Explorations in Genomics I finished a 38kb fragment of chromosome
Lo 1 Annotating Fosmid 14p24 of D. Virilis chromosome 4 Lo, Louis April 20, 2006 Annotation Report Introduction In the first half of Research Explorations in Genomics I finished a 38kb fragment of chromosome
Annotation of contig27 in the Muller F Element of D. elegans. Contig27 is a 60,000 bp region located in the Muller F element of the D. elegans.
 David Wang Bio 434W 4/27/15 Annotation of contig27 in the Muller F Element of D. elegans Abstract Contig27 is a 60,000 bp region located in the Muller F element of the D. elegans. Genscan predicted six
David Wang Bio 434W 4/27/15 Annotation of contig27 in the Muller F Element of D. elegans Abstract Contig27 is a 60,000 bp region located in the Muller F element of the D. elegans. Genscan predicted six
Draft 3 Annotation of DGA06H06, Contig 1 Jeannette Wong Bio4342W 27 April 2009
 Page 1 Draft 3 Annotation of DGA06H06, Contig 1 Jeannette Wong Bio4342W 27 April 2009 Page 2 Introduction: Annotation is the process of analyzing the genomic sequence of an organism. Besides identifying
Page 1 Draft 3 Annotation of DGA06H06, Contig 1 Jeannette Wong Bio4342W 27 April 2009 Page 2 Introduction: Annotation is the process of analyzing the genomic sequence of an organism. Besides identifying
Collect, analyze and synthesize. Annotation. Annotation for D. virilis. Evidence Based Annotation. GEP goals: Evidence for Gene Models 08/22/2017
 Annotation Annotation for D. virilis Chris Shaffer July 2012 l Big Picture of annotation and then one practical example l This technique may not be the best with other projects (e.g. corn, bacteria) l
Annotation Annotation for D. virilis Chris Shaffer July 2012 l Big Picture of annotation and then one practical example l This technique may not be the best with other projects (e.g. corn, bacteria) l
Collect, analyze and synthesize. Annotation. Annotation for D. virilis. GEP goals: Evidence Based Annotation. Evidence for Gene Models 12/26/2018
 Annotation Annotation for D. virilis Chris Shaffer July 2012 l Big Picture of annotation and then one practical example l This technique may not be the best with other projects (e.g. corn, bacteria) l
Annotation Annotation for D. virilis Chris Shaffer July 2012 l Big Picture of annotation and then one practical example l This technique may not be the best with other projects (e.g. corn, bacteria) l
Aaditya Khatri. Abstract
 Abstract In this project, Chimp-chunk 2-7 was annotated. Chimp-chunk 2-7 is an 80 kb region on chromosome 5 of the chimpanzee genome. Analysis with the Mapviewer function using the NCBI non-redundant database
Abstract In this project, Chimp-chunk 2-7 was annotated. Chimp-chunk 2-7 is an 80 kb region on chromosome 5 of the chimpanzee genome. Analysis with the Mapviewer function using the NCBI non-redundant database
Lab Week 9 - A Sample Annotation Problem (adapted by Chris Shaffer from a worksheet by Varun Sundaram, WU-STL, Class of 2009)
 Lab Week 9 - A Sample Annotation Problem (adapted by Chris Shaffer from a worksheet by Varun Sundaram, WU-STL, Class of 2009) Prerequisites: BLAST Exercise: An In-Depth Introduction to NCBI BLAST Familiarity
Lab Week 9 - A Sample Annotation Problem (adapted by Chris Shaffer from a worksheet by Varun Sundaram, WU-STL, Class of 2009) Prerequisites: BLAST Exercise: An In-Depth Introduction to NCBI BLAST Familiarity
Chimp Sequence Annotation: Region 2_3
 Chimp Sequence Annotation: Region 2_3 Jeff Howenstein March 30, 2007 BIO434W Genomics 1 Introduction We received region 2_3 of the ChimpChunk sequence, and the first step we performed was to run RepeatMasker
Chimp Sequence Annotation: Region 2_3 Jeff Howenstein March 30, 2007 BIO434W Genomics 1 Introduction We received region 2_3 of the ChimpChunk sequence, and the first step we performed was to run RepeatMasker
Annotation of Drosophila erecta Contig 14. Kimberly Chau Dr. Laura Hoopes. Pomona College 24 February 2009
 Annotation of Drosophila erecta Contig 14 Kimberly Chau Dr. Laura Hoopes Pomona College 24 February 2009 1 Table of Contents I. Overview A. Introduction..1 B. Final Gene Model.....1 II. Genes A. Initial
Annotation of Drosophila erecta Contig 14 Kimberly Chau Dr. Laura Hoopes Pomona College 24 February 2009 1 Table of Contents I. Overview A. Introduction..1 B. Final Gene Model.....1 II. Genes A. Initial
Genomic Annotation Lab Exercise By Jacob Jipp and Marian Kaehler Luther College, Department of Biology Genomics Education Partnership 2010
 Genomic Annotation Lab Exercise By Jacob Jipp and Marian Kaehler Luther College, Department of Biology Genomics Education Partnership 2010 Genomics is a new and expanding field with an increasing impact
Genomic Annotation Lab Exercise By Jacob Jipp and Marian Kaehler Luther College, Department of Biology Genomics Education Partnership 2010 Genomics is a new and expanding field with an increasing impact
ab initio and Evidence-Based Gene Finding
 ab initio and Evidence-Based Gene Finding A basic introduction to annotation Outline What is annotation? ab initio gene finding Genome databases on the web Basics of the UCSC browser Evidence-based gene
ab initio and Evidence-Based Gene Finding A basic introduction to annotation Outline What is annotation? ab initio gene finding Genome databases on the web Basics of the UCSC browser Evidence-based gene
MODULE 5: TRANSLATION
 MODULE 5: TRANSLATION Lesson Plan: CARINA ENDRES HOWELL, LEOCADIA PALIULIS Title Translation Objectives Determine the codons for specific amino acids and identify reading frames by looking at the Base
MODULE 5: TRANSLATION Lesson Plan: CARINA ENDRES HOWELL, LEOCADIA PALIULIS Title Translation Objectives Determine the codons for specific amino acids and identify reading frames by looking at the Base
UCSC Genome Browser. Introduction to ab initio and evidence-based gene finding
 UCSC Genome Browser Introduction to ab initio and evidence-based gene finding Wilson Leung 06/2006 Outline Introduction to annotation ab initio gene finding Basics of the UCSC Browser Evidence-based gene
UCSC Genome Browser Introduction to ab initio and evidence-based gene finding Wilson Leung 06/2006 Outline Introduction to annotation ab initio gene finding Basics of the UCSC Browser Evidence-based gene
Annotating the D. virilis Fourth Chromosome: Fosmid 99M21
 Sonal Singhal 3 May 2006 Bio 4342W Annotating the D. virilis Fourth Chromosome: Fosmid 99M21 Abstract In this project, I annotated a chunk of the D. virilis fourth chromosome (fosmid 99M21) by considering
Sonal Singhal 3 May 2006 Bio 4342W Annotating the D. virilis Fourth Chromosome: Fosmid 99M21 Abstract In this project, I annotated a chunk of the D. virilis fourth chromosome (fosmid 99M21) by considering
Computational gene finding
 Computational gene finding Devika Subramanian Comp 470 Outline (3 lectures) Lec 1 Lec 2 Lec 3 The biological context Markov models and Hidden Markov models Ab-initio methods for gene finding Comparative
Computational gene finding Devika Subramanian Comp 470 Outline (3 lectures) Lec 1 Lec 2 Lec 3 The biological context Markov models and Hidden Markov models Ab-initio methods for gene finding Comparative
Annotation Practice Activity [Based on materials from the GEP Summer 2010 Workshop] Special thanks to Chris Shaffer for document review Parts A-G
![Annotation Practice Activity [Based on materials from the GEP Summer 2010 Workshop] Special thanks to Chris Shaffer for document review Parts A-G Annotation Practice Activity [Based on materials from the GEP Summer 2010 Workshop] Special thanks to Chris Shaffer for document review Parts A-G](/thumbs/95/123686269.jpg) Annotation Practice Activity [Based on materials from the GEP Summer 2010 Workshop] Special thanks to Chris Shaffer for document review Parts A-G Introduction: A genome is the total genetic content of
Annotation Practice Activity [Based on materials from the GEP Summer 2010 Workshop] Special thanks to Chris Shaffer for document review Parts A-G Introduction: A genome is the total genetic content of
BIO4342 Lab Exercise: Detecting and Interpreting Genetic Homology
 BIO4342 Lab Exercise: Detecting and Interpreting Genetic Homology Jeremy Buhler March 15, 2004 In this lab, we ll annotate an interesting piece of the D. melanogaster genome. Along the way, you ll get
BIO4342 Lab Exercise: Detecting and Interpreting Genetic Homology Jeremy Buhler March 15, 2004 In this lab, we ll annotate an interesting piece of the D. melanogaster genome. Along the way, you ll get
Outline. Annotation of Drosophila Primer. Gene structure nomenclature. Muller element nomenclature. GEP Drosophila annotation projects 01/04/2018
 Outline Overview of the GEP annotation projects Annotation of Drosophila Primer January 2018 GEP annotation workflow Practice applying the GEP annotation strategy Wilson Leung and Chris Shaffer AAACAACAATCATAAATAGAGGAAGTTTTCGGAATATACGATAAGTGAAATATCGTTCT
Outline Overview of the GEP annotation projects Annotation of Drosophila Primer January 2018 GEP annotation workflow Practice applying the GEP annotation strategy Wilson Leung and Chris Shaffer AAACAACAATCATAAATAGAGGAAGTTTTCGGAATATACGATAAGTGAAATATCGTTCT
Identifying Genes and Pseudogenes in a Chimpanzee Sequence Adapted from Chimp BAC analysis: TWINSCAN and UCSC Browser by Dr. M.
 Identifying Genes and Pseudogenes in a Chimpanzee Sequence Adapted from Chimp BAC analysis: TWINSCAN and UCSC Browser by Dr. M. Brent Prerequisites: A Simple Introduction to NCBI BLAST Resources: The GENSCAN
Identifying Genes and Pseudogenes in a Chimpanzee Sequence Adapted from Chimp BAC analysis: TWINSCAN and UCSC Browser by Dr. M. Brent Prerequisites: A Simple Introduction to NCBI BLAST Resources: The GENSCAN
TIGR THE INSTITUTE FOR GENOMIC RESEARCH
 Introduction to Genome Annotation: Overview of What You Will Learn This Week C. Robin Buell May 21, 2007 Types of Annotation Structural Annotation: Defining genes, boundaries, sequence motifs e.g. ORF,
Introduction to Genome Annotation: Overview of What You Will Learn This Week C. Robin Buell May 21, 2007 Types of Annotation Structural Annotation: Defining genes, boundaries, sequence motifs e.g. ORF,
Transcription Start Sites Project Report
 Transcription Start Sites Project Report Student name: Student email: Faculty advisor: College/university: Project details Project name: Project species: Date of submission: Number of genes in project:
Transcription Start Sites Project Report Student name: Student email: Faculty advisor: College/university: Project details Project name: Project species: Date of submission: Number of genes in project:
user s guide Question 1
 Question 1 How does one find a gene of interest and determine that gene s structure? Once the gene has been located on the map, how does one easily examine other genes in that same region? doi:10.1038/ng966
Question 1 How does one find a gene of interest and determine that gene s structure? Once the gene has been located on the map, how does one easily examine other genes in that same region? doi:10.1038/ng966
Annotation Walkthrough Workshop BIO 173/273 Genomics and Bioinformatics Spring 2013 Developed by Justin R. DiAngelo at Hofstra University
 Annotation Walkthrough Workshop NAME: BIO 173/273 Genomics and Bioinformatics Spring 2013 Developed by Justin R. DiAngelo at Hofstra University A Simple Annotation Exercise Adapted from: Alexis Nagengast,
Annotation Walkthrough Workshop NAME: BIO 173/273 Genomics and Bioinformatics Spring 2013 Developed by Justin R. DiAngelo at Hofstra University A Simple Annotation Exercise Adapted from: Alexis Nagengast,
Gene Annotation Project. Group 1. Tyler Tiede Yanzhu Ji Jenae Skelton
 Gene Annotation Project Group 1 Tyler Tiede Yanzhu Ji Jenae Skelton Outline Tools Overview of 150kb region Overview of annotation process Characterization of 5 putative gene regions Analysis of masked
Gene Annotation Project Group 1 Tyler Tiede Yanzhu Ji Jenae Skelton Outline Tools Overview of 150kb region Overview of annotation process Characterization of 5 putative gene regions Analysis of masked
Question 2: There are 5 retroelements (2 LINEs and 3 LTRs), 6 unclassified elements (XDMR and XDMR_DM), and 7 satellite sequences.
 Bio4342 Exercise 1 Answers: Detecting and Interpreting Genetic Homology (Answers prepared by Wilson Leung) Question 1: Low complexity DNA can be described as sequences that consist primarily of one or
Bio4342 Exercise 1 Answers: Detecting and Interpreting Genetic Homology (Answers prepared by Wilson Leung) Question 1: Low complexity DNA can be described as sequences that consist primarily of one or
Computational gene finding
 Computational gene finding Devika Subramanian Comp 470 Outline (3 lectures) Lec 1 Lec 2 Lec 3 The biological context Markov models and Hidden Markov models Ab-initio methods for gene finding Comparative
Computational gene finding Devika Subramanian Comp 470 Outline (3 lectures) Lec 1 Lec 2 Lec 3 The biological context Markov models and Hidden Markov models Ab-initio methods for gene finding Comparative
Finishing of Fosmid 1042D14. Project 1042D14 is a roughly 40 kb segment of Drosophila ananassae
 Schefkind 1 Adam Schefkind Bio 434W 03/08/2014 Finishing of Fosmid 1042D14 Abstract Project 1042D14 is a roughly 40 kb segment of Drosophila ananassae genomic DNA. Through a comprehensive analysis of forward-
Schefkind 1 Adam Schefkind Bio 434W 03/08/2014 Finishing of Fosmid 1042D14 Abstract Project 1042D14 is a roughly 40 kb segment of Drosophila ananassae genomic DNA. Through a comprehensive analysis of forward-
Genomes: What we know and what we don t know
 Genomes: What we know and what we don t know Complete draft sequence 2001 October 15, 2007 Dr. Stefan Maas, BioS Lehigh U. What we know Raw genome data The range of genome sizes in the animal & plant kingdoms!
Genomes: What we know and what we don t know Complete draft sequence 2001 October 15, 2007 Dr. Stefan Maas, BioS Lehigh U. What we know Raw genome data The range of genome sizes in the animal & plant kingdoms!
Genome annotation. Erwin Datema (2011) Sandra Smit (2012, 2013)
 Genome annotation Erwin Datema (2011) Sandra Smit (2012, 2013) Genome annotation AGACAAAGATCCGCTAAATTAAATCTGGACTTCACATATTGAAGTGATATCACACGTTTCTCTAAT AATCTCCTCACAATATTATGTTTGGGATGAACTTGTCGTGATTTGCCATTGTAGCAATCACTTGAA
Genome annotation Erwin Datema (2011) Sandra Smit (2012, 2013) Genome annotation AGACAAAGATCCGCTAAATTAAATCTGGACTTCACATATTGAAGTGATATCACACGTTTCTCTAAT AATCTCCTCACAATATTATGTTTGGGATGAACTTGTCGTGATTTGCCATTGTAGCAATCACTTGAA
MODULE 1: INTRODUCTION TO THE GENOME BROWSER: WHAT IS A GENE?
 MODULE 1: INTRODUCTION TO THE GENOME BROWSER: WHAT IS A GENE? Lesson Plan: Title Introduction to the Genome Browser: what is a gene? JOYCE STAMM Objectives Demonstrate basic skills in using the UCSC Genome
MODULE 1: INTRODUCTION TO THE GENOME BROWSER: WHAT IS A GENE? Lesson Plan: Title Introduction to the Genome Browser: what is a gene? JOYCE STAMM Objectives Demonstrate basic skills in using the UCSC Genome
Agenda. Annotation of Drosophila. Muller element nomenclature. Annotation: Adding labels to a sequence. GEP Drosophila annotation projects 01/03/2018
 Agenda Annotation of Drosophila January 2018 Overview of the GEP annotation project GEP annotation strategy Types of evidence Analysis tools Web databases Annotation of a single isoform (walkthrough) Wilson
Agenda Annotation of Drosophila January 2018 Overview of the GEP annotation project GEP annotation strategy Types of evidence Analysis tools Web databases Annotation of a single isoform (walkthrough) Wilson
Annotation of Contig8 Sakura Oyama Dr. Elgin, Dr. Shaffer, Dr. Bednarski Bio 434W May 2, 2016
 Annotation of Contig8 Sakura Oyama Dr. Elgin, Dr. Shaffer, Dr. Bednarski Bio 434W May 2, 2016 Abstract Contig8, a 45 kb region of the fourth chromosome of Drosophila ficusphila, was annotated using the
Annotation of Contig8 Sakura Oyama Dr. Elgin, Dr. Shaffer, Dr. Bednarski Bio 434W May 2, 2016 Abstract Contig8, a 45 kb region of the fourth chromosome of Drosophila ficusphila, was annotated using the
Annotation of contig62 from Drosophila elegans Dot Chromosome
 Abstract: Annotation of contig62 from Drosophila elegans Dot Chromosome 1 Maxwell Wang The goal of this project is to annotate the Drosophila elegans Dot chromosome contig62. Contig62 is a 32,259 bp contig
Abstract: Annotation of contig62 from Drosophila elegans Dot Chromosome 1 Maxwell Wang The goal of this project is to annotate the Drosophila elegans Dot chromosome contig62. Contig62 is a 32,259 bp contig
Drosophila ficusphila F element
 5/2/2016 CONTIG52 Drosophila ficusphila F element Vahag Kechejian BIO434W Abstract Contig52 is a 35,000 bp region located on the F element of Drosophila ficusphila. Genscan predicts six features in the
5/2/2016 CONTIG52 Drosophila ficusphila F element Vahag Kechejian BIO434W Abstract Contig52 is a 35,000 bp region located on the F element of Drosophila ficusphila. Genscan predicts six features in the
Lecture 7 Motif Databases and Gene Finding
 Introduction to Bioinformatics for Medical Research Gideon Greenspan gdg@cs.technion.ac.il Lecture 7 Motif Databases and Gene Finding Motif Databases & Gene Finding Motifs Recap Motif Databases TRANSFAC
Introduction to Bioinformatics for Medical Research Gideon Greenspan gdg@cs.technion.ac.il Lecture 7 Motif Databases and Gene Finding Motif Databases & Gene Finding Motifs Recap Motif Databases TRANSFAC
Small Exon Finder User Guide
 Small Exon Finder User Guide Author Wilson Leung wleung@wustl.edu Document History Initial Draft 01/09/2011 First Revision 08/03/2014 Current Version 12/29/2015 Table of Contents Author... 1 Document History...
Small Exon Finder User Guide Author Wilson Leung wleung@wustl.edu Document History Initial Draft 01/09/2011 First Revision 08/03/2014 Current Version 12/29/2015 Table of Contents Author... 1 Document History...
BME 110 Midterm Examination
 BME 110 Midterm Examination May 10, 2011 Name: (please print) Directions: Please circle one answer for each question, unless the question specifies "circle all correct answers". You can use any resource
BME 110 Midterm Examination May 10, 2011 Name: (please print) Directions: Please circle one answer for each question, unless the question specifies "circle all correct answers". You can use any resource
BCHM 6280 Tutorial: Gene specific information using NCBI, Ensembl and genome viewers
 BCHM 6280 Tutorial: Gene specific information using NCBI, Ensembl and genome viewers Web resources: NCBI database: http://www.ncbi.nlm.nih.gov/ Ensembl database: http://useast.ensembl.org/index.html UCSC
BCHM 6280 Tutorial: Gene specific information using NCBI, Ensembl and genome viewers Web resources: NCBI database: http://www.ncbi.nlm.nih.gov/ Ensembl database: http://useast.ensembl.org/index.html UCSC
Gene Identification in silico
 Gene Identification in silico Nita Parekh, IIIT Hyderabad Presented at National Seminar on Bioinformatics and Functional Genomics, at Bioinformatics centre, Pondicherry University, Feb 15 17, 2006. Introduction
Gene Identification in silico Nita Parekh, IIIT Hyderabad Presented at National Seminar on Bioinformatics and Functional Genomics, at Bioinformatics centre, Pondicherry University, Feb 15 17, 2006. Introduction
Genome annotation & EST
 Genome annotation & EST What is genome annotation? The process of taking the raw DNA sequence produced by the genome sequence projects and adding the layers of analysis and interpretation necessary
Genome annotation & EST What is genome annotation? The process of taking the raw DNA sequence produced by the genome sequence projects and adding the layers of analysis and interpretation necessary
 Genome Annotation Genome annotation What is the function of each part of the genome? Where are the genes? What is the mrna sequence (transcription, splicing) What is the protein sequence? What does
Genome Annotation Genome annotation What is the function of each part of the genome? Where are the genes? What is the mrna sequence (transcription, splicing) What is the protein sequence? What does
Sections 12.3, 13.1, 13.2
 Sections 12.3, 13.1, 13.2 Background: Watson & Crick recognized that base pairing in the double helix allows DNA to be copied, or replicated Each strand in the double helix has all the information to remake
Sections 12.3, 13.1, 13.2 Background: Watson & Crick recognized that base pairing in the double helix allows DNA to be copied, or replicated Each strand in the double helix has all the information to remake
GENETICS - CLUTCH CH.15 GENOMES AND GENOMICS.
 !! www.clutchprep.com CONCEPT: OVERVIEW OF GENOMICS Genomics is the study of genomes in their entirety Bioinformatics is the analysis of the information content of genomes - Genes, regulatory sequences,
!! www.clutchprep.com CONCEPT: OVERVIEW OF GENOMICS Genomics is the study of genomes in their entirety Bioinformatics is the analysis of the information content of genomes - Genes, regulatory sequences,
Computational gene finding. Devika Subramanian Comp 470
 Computational gene finding Devika Subramanian Comp 470 Outline (3 lectures) The biological context Lec 1 Lec 2 Lec 3 Markov models and Hidden Markov models Ab-initio methods for gene finding Comparative
Computational gene finding Devika Subramanian Comp 470 Outline (3 lectures) The biological context Lec 1 Lec 2 Lec 3 Markov models and Hidden Markov models Ab-initio methods for gene finding Comparative
Data Retrieval from GenBank
 Data Retrieval from GenBank Peter J. Myler Bioinformatics of Intracellular Pathogens JNU, Feb 7-0, 2009 http://www.ncbi.nlm.nih.gov (January, 2007) http://ncbi.nlm.nih.gov/sitemap/resourceguide.html Accessing
Data Retrieval from GenBank Peter J. Myler Bioinformatics of Intracellular Pathogens JNU, Feb 7-0, 2009 http://www.ncbi.nlm.nih.gov (January, 2007) http://ncbi.nlm.nih.gov/sitemap/resourceguide.html Accessing
Annotation of a Drosophila Gene
 Annotation of a Drosophila Gene Wilson Leung Last Update: 12/30/2018 Prerequisites Lecture: Annotation of Drosophila Lecture: RNA-Seq Primer BLAST Walkthrough: An Introduction to NCBI BLAST Resources FlyBase:
Annotation of a Drosophila Gene Wilson Leung Last Update: 12/30/2018 Prerequisites Lecture: Annotation of Drosophila Lecture: RNA-Seq Primer BLAST Walkthrough: An Introduction to NCBI BLAST Resources FlyBase:
Bacterial Genome Annotation
 Bacterial Genome Annotation Bacterial Genome Annotation For an annotation you want to predict from the sequence, all of... protein-coding genes their stop-start the resulting protein the function the control
Bacterial Genome Annotation Bacterial Genome Annotation For an annotation you want to predict from the sequence, all of... protein-coding genes their stop-start the resulting protein the function the control
Complete draft sequence 2001
 Genomes: What we know and what we don t know Complete draft sequence 2001 November11, 2009 Dr. Stefan Maas, BioS Lehigh U. What we know Raw genome data The range of genome sizes in the animal & plant kingdoms
Genomes: What we know and what we don t know Complete draft sequence 2001 November11, 2009 Dr. Stefan Maas, BioS Lehigh U. What we know Raw genome data The range of genome sizes in the animal & plant kingdoms
Week 1 BCHM 6280 Tutorial: Gene specific information using NCBI, Ensembl and genome viewers
 Week 1 BCHM 6280 Tutorial: Gene specific information using NCBI, Ensembl and genome viewers Web resources: NCBI database: http://www.ncbi.nlm.nih.gov/ Ensembl database: http://useast.ensembl.org/index.html
Week 1 BCHM 6280 Tutorial: Gene specific information using NCBI, Ensembl and genome viewers Web resources: NCBI database: http://www.ncbi.nlm.nih.gov/ Ensembl database: http://useast.ensembl.org/index.html
Agenda. Web Databases for Drosophila. Gene annotation workflow. GEP Drosophila annotation projects 01/01/2018. Annotation adding labels to a sequence
 Agenda GEP annotation project overview Web Databases for Drosophila An introduction to web tools, databases and NCBI BLAST Web databases for Drosophila annotation UCSC Genome Browser NCBI / BLAST FlyBase
Agenda GEP annotation project overview Web Databases for Drosophila An introduction to web tools, databases and NCBI BLAST Web databases for Drosophila annotation UCSC Genome Browser NCBI / BLAST FlyBase
The common structure of a DNA nucleotide. Hewitt
 GENETICS Unless otherwise noted* the artwork and photographs in this slide show are original and by Burt Carter. Permission is granted to use them for non-commercial, non-profit educational purposes provided
GENETICS Unless otherwise noted* the artwork and photographs in this slide show are original and by Burt Carter. Permission is granted to use them for non-commercial, non-profit educational purposes provided
Files for this Tutorial: All files needed for this tutorial are compressed into a single archive: [BLAST_Intro.tar.gz]
![Files for this Tutorial: All files needed for this tutorial are compressed into a single archive: [BLAST_Intro.tar.gz] Files for this Tutorial: All files needed for this tutorial are compressed into a single archive: [BLAST_Intro.tar.gz]](/thumbs/75/71877651.jpg) BLAST Exercise: Detecting and Interpreting Genetic Homology Adapted by W. Leung and SCR Elgin from Detecting and Interpreting Genetic Homology by Dr. J. Buhler Prequisites: None Resources: The BLAST web
BLAST Exercise: Detecting and Interpreting Genetic Homology Adapted by W. Leung and SCR Elgin from Detecting and Interpreting Genetic Homology by Dr. J. Buhler Prequisites: None Resources: The BLAST web
MATH 5610, Computational Biology
 MATH 5610, Computational Biology Lecture 2 Intro to Molecular Biology (cont) Stephen Billups University of Colorado at Denver MATH 5610, Computational Biology p.1/24 Announcements Error on syllabus Class
MATH 5610, Computational Biology Lecture 2 Intro to Molecular Biology (cont) Stephen Billups University of Colorado at Denver MATH 5610, Computational Biology p.1/24 Announcements Error on syllabus Class
Finishing of DELE Drosophila elegans has been sequenced using Roche 454 pyrosequencing and Illumina
 Sarah Swiezy Dr. Elgin, Dr. Shaffer Bio 434W 27 February 2015 Finishing of DELE8596009 Abstract Drosophila elegans has been sequenced using Roche 454 pyrosequencing and Illumina technology. DELE8596009,
Sarah Swiezy Dr. Elgin, Dr. Shaffer Bio 434W 27 February 2015 Finishing of DELE8596009 Abstract Drosophila elegans has been sequenced using Roche 454 pyrosequencing and Illumina technology. DELE8596009,
Hands-On Four Investigating Inherited Diseases
 Hands-On Four Investigating Inherited Diseases The purpose of these exercises is to introduce bioinformatics databases and tools. We investigate an important human gene and see how mutations give rise
Hands-On Four Investigating Inherited Diseases The purpose of these exercises is to introduce bioinformatics databases and tools. We investigate an important human gene and see how mutations give rise
I. Gene Expression Figure 1: Central Dogma of Molecular Biology
 I. Gene Expression Figure 1: Central Dogma of Molecular Biology Central Dogma: Gene Expression: RNA Structure RNA nucleotides contain the pentose sugar Ribose instead of deoxyribose. Contain the bases
I. Gene Expression Figure 1: Central Dogma of Molecular Biology Central Dogma: Gene Expression: RNA Structure RNA nucleotides contain the pentose sugar Ribose instead of deoxyribose. Contain the bases
CS313 Exercise 1 Cover Page Fall 2017
 CS313 Exercise 1 Cover Page Fall 2017 Due by the start of class on Monday, September 18, 2017. Name(s): In the TIME column, please estimate the time you spent on the parts of this exercise. Please try
CS313 Exercise 1 Cover Page Fall 2017 Due by the start of class on Monday, September 18, 2017. Name(s): In the TIME column, please estimate the time you spent on the parts of this exercise. Please try
A Guide to Consed Michelle Itano, Carolyn Cain, Tien Chusak, Justin Richner, and SCR Elgin.
 1 A Guide to Consed Michelle Itano, Carolyn Cain, Tien Chusak, Justin Richner, and SCR Elgin. Main Window Figure 1. The Main Window is the starting point when Consed is opened. From here, you can access
1 A Guide to Consed Michelle Itano, Carolyn Cain, Tien Chusak, Justin Richner, and SCR Elgin. Main Window Figure 1. The Main Window is the starting point when Consed is opened. From here, you can access
Chimp BAC analysis: Adapted by Wilson Leung and Sarah C.R. Elgin from Chimp BAC analysis: TWINSCAN and UCSC Browser by Dr. Michael R.
 Chimp BAC analysis: Adapted by Wilson Leung and Sarah C.R. Elgin from Chimp BAC analysis: TWINSCAN and UCSC Browser by Dr. Michael R. Brent Prerequisites: BLAST exercise: Detecting and Interpreting Genetic
Chimp BAC analysis: Adapted by Wilson Leung and Sarah C.R. Elgin from Chimp BAC analysis: TWINSCAN and UCSC Browser by Dr. Michael R. Brent Prerequisites: BLAST exercise: Detecting and Interpreting Genetic
Chimp Chunk 3-14 Annotation by Matthew Kwong, Ruth Howe, and Hao Yang
 Chimp Chunk 3-14 Annotation by Matthew Kwong, Ruth Howe, and Hao Yang Ruth Howe Bio 434W April 1, 2010 INTRODUCTION De novo annotation is the process by which a finished genomic sequence is searched for
Chimp Chunk 3-14 Annotation by Matthew Kwong, Ruth Howe, and Hao Yang Ruth Howe Bio 434W April 1, 2010 INTRODUCTION De novo annotation is the process by which a finished genomic sequence is searched for
How does the human genome stack up? Genomic Size. Genome Size. Number of Genes. Eukaryotic genomes are generally larger.
 How does the human genome stack up? Organism Human (Homo sapiens) Laboratory mouse (M. musculus) Mustard weed (A. thaliana) Roundworm (C. elegans) Fruit fly (D. melanogaster) Yeast (S. cerevisiae) Bacterium
How does the human genome stack up? Organism Human (Homo sapiens) Laboratory mouse (M. musculus) Mustard weed (A. thaliana) Roundworm (C. elegans) Fruit fly (D. melanogaster) Yeast (S. cerevisiae) Bacterium
COMPUTER RESOURCES II:
 COMPUTER RESOURCES II: Using the computer to analyze data, using the internet, and accessing online databases Bio 210, Fall 2006 Linda S. Huang, Ph.D. University of Massachusetts Boston In the first computer
COMPUTER RESOURCES II: Using the computer to analyze data, using the internet, and accessing online databases Bio 210, Fall 2006 Linda S. Huang, Ph.D. University of Massachusetts Boston In the first computer
Outline. Gene Finding Questions. Recap: Prokaryotic gene finding Eukaryotic gene finding The human gene complement Regulation
 Tues, Nov 29: Gene Finding 1 Online FCE s: Thru Dec 12 Thurs, Dec 1: Gene Finding 2 Tues, Dec 6: PS5 due Project presentations 1 (see course web site for schedule) Thurs, Dec 8 Final papers due Project
Tues, Nov 29: Gene Finding 1 Online FCE s: Thru Dec 12 Thurs, Dec 1: Gene Finding 2 Tues, Dec 6: PS5 due Project presentations 1 (see course web site for schedule) Thurs, Dec 8 Final papers due Project
Biotechnology Unit 3: DNA to Proteins. From DNA to RNA
 From DNA to RNA Biotechnology Unit 3: DNA to Proteins I. After the discovery of the structure of DNA, the major question remaining was how does the stored in the 4 letter code of DNA direct the and of
From DNA to RNA Biotechnology Unit 3: DNA to Proteins I. After the discovery of the structure of DNA, the major question remaining was how does the stored in the 4 letter code of DNA direct the and of
Applications of HMMs in Computational Biology. BMI/CS Colin Dewey
 Applications of HMMs in Computational Biology BMI/CS 576 www.biostat.wisc.edu/bmi576.html Colin Dewey cdewey@biostat.wisc.edu Fall 2008 The Gene Finding Task Given: an uncharacterized DNA sequence Do:
Applications of HMMs in Computational Biology BMI/CS 576 www.biostat.wisc.edu/bmi576.html Colin Dewey cdewey@biostat.wisc.edu Fall 2008 The Gene Finding Task Given: an uncharacterized DNA sequence Do:
Sequence Based Function Annotation
 Sequence Based Function Annotation Qi Sun Bioinformatics Facility Biotechnology Resource Center Cornell University Sequence Based Function Annotation 1. Given a sequence, how to predict its biological
Sequence Based Function Annotation Qi Sun Bioinformatics Facility Biotechnology Resource Center Cornell University Sequence Based Function Annotation 1. Given a sequence, how to predict its biological
BIOINFORMATICS TO ANALYZE AND COMPARE GENOMES
 BIOINFORMATICS TO ANALYZE AND COMPARE GENOMES We sequenced and assembled a genome, but this is only a long stretch of ATCG What should we do now? 1. find genes What are the starting and end points for
BIOINFORMATICS TO ANALYZE AND COMPARE GENOMES We sequenced and assembled a genome, but this is only a long stretch of ATCG What should we do now? 1. find genes What are the starting and end points for
Transcription and Translation. DANILO V. ROGAYAN JR. Faculty, Department of Natural Sciences
 Transcription and Translation DANILO V. ROGAYAN JR. Faculty, Department of Natural Sciences Protein Structure Made up of amino acids Polypeptide- string of amino acids 20 amino acids are arranged in different
Transcription and Translation DANILO V. ROGAYAN JR. Faculty, Department of Natural Sciences Protein Structure Made up of amino acids Polypeptide- string of amino acids 20 amino acids are arranged in different
From DNA to Protein: Genotype to Phenotype
 12 From DNA to Protein: Genotype to Phenotype 12.1 What Is the Evidence that Genes Code for Proteins? The gene-enzyme relationship is one-gene, one-polypeptide relationship. Example: In hemoglobin, each
12 From DNA to Protein: Genotype to Phenotype 12.1 What Is the Evidence that Genes Code for Proteins? The gene-enzyme relationship is one-gene, one-polypeptide relationship. Example: In hemoglobin, each
Section 10.3 Outline 10.3 How Is the Base Sequence of a Messenger RNA Molecule Translated into Protein?
 Section 10.3 Outline 10.3 How Is the Base Sequence of a Messenger RNA Molecule Translated into Protein? Messenger RNA Carries Information for Protein Synthesis from the DNA to Ribosomes Ribosomes Consist
Section 10.3 Outline 10.3 How Is the Base Sequence of a Messenger RNA Molecule Translated into Protein? Messenger RNA Carries Information for Protein Synthesis from the DNA to Ribosomes Ribosomes Consist
Bioinformatics Tools. Stuart M. Brown, Ph.D Dept of Cell Biology NYU School of Medicine
 Bioinformatics Tools Stuart M. Brown, Ph.D Dept of Cell Biology NYU School of Medicine Bioinformatics Tools Stuart M. Brown, Ph.D Dept of Cell Biology NYU School of Medicine Overview This lecture will
Bioinformatics Tools Stuart M. Brown, Ph.D Dept of Cell Biology NYU School of Medicine Bioinformatics Tools Stuart M. Brown, Ph.D Dept of Cell Biology NYU School of Medicine Overview This lecture will
Biology. Biology. Slide 1 of 39. End Show. Copyright Pearson Prentice Hall
 Biology Biology 1 of 39 12-3 RNA and Protein Synthesis 2 of 39 Essential Question What is transcription and translation and how do they take place? 3 of 39 12 3 RNA and Protein Synthesis Genes are coded
Biology Biology 1 of 39 12-3 RNA and Protein Synthesis 2 of 39 Essential Question What is transcription and translation and how do they take place? 3 of 39 12 3 RNA and Protein Synthesis Genes are coded
Biology. Biology. Slide 1 of 39. End Show. Copyright Pearson Prentice Hall
 Biology Biology 1 of 39 12-3 RNA and Protein Synthesis 2 of 39 12 3 RNA and Protein Synthesis Genes are coded DNA instructions that control the production of proteins. Genetic messages can be decoded by
Biology Biology 1 of 39 12-3 RNA and Protein Synthesis 2 of 39 12 3 RNA and Protein Synthesis Genes are coded DNA instructions that control the production of proteins. Genetic messages can be decoded by
PROTEIN SYNTHESIS Flow of Genetic Information The flow of genetic information can be symbolized as: DNA RNA Protein
 PROTEIN SYNTHESIS Flow of Genetic Information The flow of genetic information can be symbolized as: DNA RNA Protein This is also known as: The central dogma of molecular biology Protein Proteins are made
PROTEIN SYNTHESIS Flow of Genetic Information The flow of genetic information can be symbolized as: DNA RNA Protein This is also known as: The central dogma of molecular biology Protein Proteins are made
Last Update: 12/31/2017. Recommended Background Tutorial: An Introduction to NCBI BLAST
 BLAST Exercise: Detecting and Interpreting Genetic Homology Adapted by T. Cordonnier, C. Shaffer, W. Leung and SCR Elgin from Detecting and Interpreting Genetic Homology by Dr. J. Buhler Recommended Background
BLAST Exercise: Detecting and Interpreting Genetic Homology Adapted by T. Cordonnier, C. Shaffer, W. Leung and SCR Elgin from Detecting and Interpreting Genetic Homology by Dr. J. Buhler Recommended Background
The Flow of Genetic Information
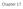 Chapter 17 The Flow of Genetic Information The DNA inherited by an organism leads to specific traits by dictating the synthesis of proteins and of RNA molecules involved in protein synthesis. Proteins
Chapter 17 The Flow of Genetic Information The DNA inherited by an organism leads to specific traits by dictating the synthesis of proteins and of RNA molecules involved in protein synthesis. Proteins
The Ensembl Database. Dott.ssa Inga Prokopenko. Corso di Genomica
 The Ensembl Database Dott.ssa Inga Prokopenko Corso di Genomica 1 www.ensembl.org Lecture 7.1 2 What is Ensembl? Public annotation of mammalian and other genomes Open source software Relational database
The Ensembl Database Dott.ssa Inga Prokopenko Corso di Genomica 1 www.ensembl.org Lecture 7.1 2 What is Ensembl? Public annotation of mammalian and other genomes Open source software Relational database
Unit 1: DNA and the Genome. Sub-Topic (1.3) Gene Expression
 Unit 1: DNA and the Genome Sub-Topic (1.3) Gene Expression Unit 1: DNA and the Genome Sub-Topic (1.3) Gene Expression On completion of this subtopic I will be able to State the meanings of the terms genotype,
Unit 1: DNA and the Genome Sub-Topic (1.3) Gene Expression Unit 1: DNA and the Genome Sub-Topic (1.3) Gene Expression On completion of this subtopic I will be able to State the meanings of the terms genotype,
How to Use This Presentation
 How to Use This Presentation To View the presentation as a slideshow with effects select View on the menu bar and click on Slide Show. To advance through the presentation, click the right-arrow key or
How to Use This Presentation To View the presentation as a slideshow with effects select View on the menu bar and click on Slide Show. To advance through the presentation, click the right-arrow key or
CHapter 14. From DNA to Protein
 CHapter 14 From DNA to Protein How? DNA to RNA to Protein to Trait Types of RNA 1. Messenger RNA: carries protein code or transcript 2. Ribosomal RNA: part of ribosomes 3. Transfer RNA: delivers amino
CHapter 14 From DNA to Protein How? DNA to RNA to Protein to Trait Types of RNA 1. Messenger RNA: carries protein code or transcript 2. Ribosomal RNA: part of ribosomes 3. Transfer RNA: delivers amino
Lecture 2: Biology Basics Continued. Fall 2018 August 23, 2018
 Lecture 2: Biology Basics Continued Fall 2018 August 23, 2018 Genetic Material for Life Central Dogma DNA: The Code of Life The structure and the four genomic letters code for all living organisms Adenine,
Lecture 2: Biology Basics Continued Fall 2018 August 23, 2018 Genetic Material for Life Central Dogma DNA: The Code of Life The structure and the four genomic letters code for all living organisms Adenine,
GenBank Growth. In 2003 ~ 31 million sequences ~ 37 billion base pairs
 Gene Finding GenBank Growth GenBank Growth In 2003 ~ 31 million sequences ~ 37 billion base pairs GenBank: Exponential Growth Growth of GenBank in billions of base pairs from release 3 in April of 1994
Gene Finding GenBank Growth GenBank Growth In 2003 ~ 31 million sequences ~ 37 billion base pairs GenBank: Exponential Growth Growth of GenBank in billions of base pairs from release 3 in April of 1994
Ensembl workshop. Thomas Randall, PhD bioinformatics.unc.edu. handouts, papers, datasets
 Ensembl workshop Thomas Randall, PhD tarandal@email.unc.edu bioinformatics.unc.edu www.unc.edu/~tarandal/ensembl handouts, papers, datasets Ensembl is a joint project between EMBL - EBI and the Sanger
Ensembl workshop Thomas Randall, PhD tarandal@email.unc.edu bioinformatics.unc.edu www.unc.edu/~tarandal/ensembl handouts, papers, datasets Ensembl is a joint project between EMBL - EBI and the Sanger
Genes found in the genome include protein-coding genes and non-coding RNA genes. Which nucleotide is not normally found in non-coding RNA genes?
 Midterm Q Genes found in the genome include protein-coding genes and non-coding RNA genes Which nucleotide is not normally found in non-coding RNA genes? G T 3 A 4 C 5 U 00% Midterm Q Which of the following
Midterm Q Genes found in the genome include protein-coding genes and non-coding RNA genes Which nucleotide is not normally found in non-coding RNA genes? G T 3 A 4 C 5 U 00% Midterm Q Which of the following
Tutorial for Stop codon reassignment in the wild
 Tutorial for Stop codon reassignment in the wild Learning Objectives This tutorial has two learning objectives: 1. Finding evidence of stop codon reassignment on DNA fragments. 2. Detecting and confirming
Tutorial for Stop codon reassignment in the wild Learning Objectives This tutorial has two learning objectives: 1. Finding evidence of stop codon reassignment on DNA fragments. 2. Detecting and confirming
Biology Chapter 12 Test: Molecular Genetics
 Class: Date: ID: A Biology Chapter 12 Test: Molecular Genetics True/False Indicate whether the statement is true or false. 1. RNA polymerase has to bind to DMA for an enzyme to be synthesized. 2. The only
Class: Date: ID: A Biology Chapter 12 Test: Molecular Genetics True/False Indicate whether the statement is true or false. 1. RNA polymerase has to bind to DMA for an enzyme to be synthesized. 2. The only
Transcription is the first stage of gene expression
 Transcription is the first stage of gene expression RNA synthesis is catalyzed by RNA polymerase, which pries the DNA strands apart and hooks together the RNA nucleotides The RNA is complementary to the
Transcription is the first stage of gene expression RNA synthesis is catalyzed by RNA polymerase, which pries the DNA strands apart and hooks together the RNA nucleotides The RNA is complementary to the
Biology A: Chapter 9 Annotating Notes Protein Synthesis
 Name: Pd: Biology A: Chapter 9 Annotating Notes Protein Synthesis -As you read your textbook, please fill out these notes. -Read each paragraph state the big/main idea on the left side. -On the right side
Name: Pd: Biology A: Chapter 9 Annotating Notes Protein Synthesis -As you read your textbook, please fill out these notes. -Read each paragraph state the big/main idea on the left side. -On the right side
A tutorial introduction into the MIPS PlantsDB barley&wheat database instances
 transplant 2 nd user training workshop Poznan, Poland, June, 27 th, 2013 A tutorial introduction into the MIPS PlantsDB barley&wheat database instances TUTORIAL ANSWERS Please direct any questions related
transplant 2 nd user training workshop Poznan, Poland, June, 27 th, 2013 A tutorial introduction into the MIPS PlantsDB barley&wheat database instances TUTORIAL ANSWERS Please direct any questions related
Assemblytics: a web analytics tool for the detection of assembly-based variants Maria Nattestad and Michael C. Schatz
 Assemblytics: a web analytics tool for the detection of assembly-based variants Maria Nattestad and Michael C. Schatz Table of Contents Supplementary Note 1: Unique Anchor Filtering Supplementary Figure
Assemblytics: a web analytics tool for the detection of assembly-based variants Maria Nattestad and Michael C. Schatz Table of Contents Supplementary Note 1: Unique Anchor Filtering Supplementary Figure
user s guide Question 3
 Question 3 During a positional cloning project aimed at finding a human disease gene, linkage data have been obtained suggesting that the gene of interest lies between two sequence-tagged site markers.
Question 3 During a positional cloning project aimed at finding a human disease gene, linkage data have been obtained suggesting that the gene of interest lies between two sequence-tagged site markers.
From DNA to Protein: Genotype to Phenotype
 12 From DNA to Protein: Genotype to Phenotype 12.1 What Is the Evidence that Genes Code for Proteins? The gene-enzyme relationship is one-gene, one-polypeptide relationship. Example: In hemoglobin, each
12 From DNA to Protein: Genotype to Phenotype 12.1 What Is the Evidence that Genes Code for Proteins? The gene-enzyme relationship is one-gene, one-polypeptide relationship. Example: In hemoglobin, each
Bio 101 Sample questions: Chapter 10
 Bio 101 Sample questions: Chapter 10 1. Which of the following is NOT needed for DNA replication? A. nucleotides B. ribosomes C. Enzymes (like polymerases) D. DNA E. all of the above are needed 2 The information
Bio 101 Sample questions: Chapter 10 1. Which of the following is NOT needed for DNA replication? A. nucleotides B. ribosomes C. Enzymes (like polymerases) D. DNA E. all of the above are needed 2 The information
Gene Expression Transcription/Translation Protein Synthesis
 Gene Expression Transcription/Translation Protein Synthesis 1. Describe how genetic information is transcribed into sequences of bases in RNA molecules and is finally translated into sequences of amino
Gene Expression Transcription/Translation Protein Synthesis 1. Describe how genetic information is transcribed into sequences of bases in RNA molecules and is finally translated into sequences of amino
Investigating Inherited Diseases
 Investigating Inherited Diseases The purpose of these exercises is to introduce bioinformatics databases and tools. We investigate an important human gene and see how mutations give rise to inherited diseases.
Investigating Inherited Diseases The purpose of these exercises is to introduce bioinformatics databases and tools. We investigate an important human gene and see how mutations give rise to inherited diseases.
Genome 373: Gene Predic/on I. Doug Fowler
 Genome 373: Gene Predic/on I Doug Fowler Outline Review of gene structure Scale of the problem Solu;ons Empirical methods Ab ini&o predic;on What is a gene? A locatable region of genomic sequence, corresponding
Genome 373: Gene Predic/on I Doug Fowler Outline Review of gene structure Scale of the problem Solu;ons Empirical methods Ab ini&o predic;on What is a gene? A locatable region of genomic sequence, corresponding
3. human genomics clone genes associated with genetic disorders. 4. many projects generate ordered clones that cover genome
 Lectures 30 and 31 Genome analysis I. Genome analysis A. two general areas 1. structural 2. functional B. genome projects a status report 1. 1 st sequenced: several viral genomes 2. mitochondria and chloroplasts
Lectures 30 and 31 Genome analysis I. Genome analysis A. two general areas 1. structural 2. functional B. genome projects a status report 1. 1 st sequenced: several viral genomes 2. mitochondria and chloroplasts
Textbook Reading Guidelines
 Understanding Bioinformatics by Marketa Zvelebil and Jeremy Baum Last updated: January 16, 2013 Textbook Reading Guidelines Preface: Read the whole preface, and especially: For the students with Life Science
Understanding Bioinformatics by Marketa Zvelebil and Jeremy Baum Last updated: January 16, 2013 Textbook Reading Guidelines Preface: Read the whole preface, and especially: For the students with Life Science
BIOLOGY. Monday 14 Mar 2016
 BIOLOGY Monday 14 Mar 2016 Entry Task List the terms that were mentioned last week in the video. Translation, Transcription, Messenger RNA (mrna), codon, Ribosomal RNA (rrna), Polypeptide, etc. Agenda
BIOLOGY Monday 14 Mar 2016 Entry Task List the terms that were mentioned last week in the video. Translation, Transcription, Messenger RNA (mrna), codon, Ribosomal RNA (rrna), Polypeptide, etc. Agenda
Biotechnology Explorer
 Biotechnology Explorer C. elegans Behavior Kit Bioinformatics Supplement explorer.bio-rad.com Catalog #166-5120EDU This kit contains temperature-sensitive reagents. Open immediately and see individual
Biotechnology Explorer C. elegans Behavior Kit Bioinformatics Supplement explorer.bio-rad.com Catalog #166-5120EDU This kit contains temperature-sensitive reagents. Open immediately and see individual
