Research school methods seminar Genomics and Transcriptomics
|
|
|
- Angel Barker
- 6 years ago
- Views:
Transcription
1 Research school methods seminar Genomics and Transcriptomics Stephan Klee
2 2
3 3
4 4
5 5
6 Genetics, Genomics what are we talking about? Genetics and Genomics Study of genes Role of genes in inheritence Study of single genes and their effects/resulting disease Although both look from different angles, both need to be considered to fully understand the whole picture Study of all of a person s genes and the interplay of the genes Role of interaction of genes with each other and the environment (nongenetic factors) Study of complex diseases such as heart and lung diseases, diabetes and cancer Offers new options to personlized medicine (influence of risk factors) 6
7 How similar are we to. Humans are 99.5 to 99.9% similar to each other (not relatives!) 7
8 8
9 Genomics - What we will deal with in this presentation 1. Methods of sequencing (from the beginnings to next generation to next next generation sequencing) 2. Applications (what can we do with the sequencing tools we ve seen?) 3. (Analyzing your data) ask your bioinformatician of choice :D 9
10 Genomics the 3 Generations of sequencing First generation: Chain-Termination Sequencing (Sanger sequencing) Shotgun sequencing Second generation (next generation sequencing): Roche 454 Sequencing (GS Junior System/ GS FLX+ System) Applied Biosystems SOLID (5500 System/ 5500xl System) Solexa Illumina (HiSeq System/ Genome analyzer Iix/ MySeq) Pacific Biosciences (PacBio RS) Third generation (next next generation sequencing): Oxford Nanopore Technologies (GridION System/ MinION) Helicos (Genetic Analysis System) 10
11 Genomics the 3 Generations of sequencing 11
12 Genomics methods of the first generation - chain termination sequencing (Sanger sequencing) - 1) Denaturation of the dsdna to ssdna 2) Requires initial primer 3) 4 seperate reaction mixes (only differences is different ddntp either A, C, G or T) 4) Dideoxynucleotides lead to early chain termination 5) Seperation on a polyacrylamide gel (each reaction different lane) Gels were radioactively labeled Feasable technique for read length of 100 to 1000bp advances: radioactively labeled ddntps were exchanged for flourescent ddntps (capillary based sequencing) 12
13 Genomics methods of the first generation - shotgun sequencing - Relies on Sanger sequencing, however is capable of sequencing genomes High throughput sequencing technique that can collect a large amount of data at a fast rate. Works by partially digesting a genome or big strand of DNA into small overlapping fragments These small fragments are sequenced and fragments that overlap are matched together 13
14 Genomics methods of the second generation - Roche 454 sequencing - Oldest of the NGS technologies Current: `GS FLX Titanium` since late 2008 Technology is canceled has wide spread user base and niche applications FAST sequencing (<6h per run) Read-length bp (modal length ~700bp) 14
15 Genomics methods of the second generation - Roche 454 sequencing - Fragmentation of DNA ( bp) and adapter ligation (red + green) Deposition in microreactors together with a bead sporting adapter sequences 15
16 Genomics methods of the second generation - Roche 454 sequencing - Binding of fragment onto bead Replication of fragments in the microreactor (polymerase etc in solution) replicas bind to free bead-adapters Lysis of microreactors and extraction of fragment covered beads 16
17 Genomics methods of the second generation - Roche 454 sequencing - Placement of beads in the PicoTiterPlate Filling of the wells with bound reagents Especially reagents responsible for creating the luminous signals (luciferase) 17
18 Genomics methods of the second generation - Roche 454 sequencing - Washing of the plate/wells with dntps, one at a time Recording of the intensity of the pyrophosphat activity 18
19 Genomics methods of the second generation - Roche 454 sequencing - Image interpretation `Flow chart` Conversion to textual representation of sequence-read per well 19
20 Genomics methods of the second generation - Roche 454 sequencing - Advantages: FAST sequencing (<6h per run) Read-length bp (modal length ~500bp) Throughput: ~ 1 mio reads MBases per run (after quality filtering) Areas of application: Whole genome seq Targeted resequencing Sequencing-based Transcriptome Analysis Metagenomics Disadvantages: Poly-NTP errors are common (require specific errorhandling) Low throughput of Mbases per run More expensive than competitors 20
21 Different Plattforms: Genomics methods of the second generation - Illumina - HiSeq (1000/1500/2000/2500/ X ) MiSeq NextSeq They differ in capabilities and throughput, technology is the same NextSeq: up to 150bp paired end (PE) and 120Gbases / 1.5 days MiSeq: up to 300bp PE and 15Gbases / 3 days HiSeq 2000: up to 125bp PE and 600Gbases / 11 days HiSeq 2500: up to 125bp PE and 1Tbases / 6days HiSeq X: up to 150bp PE and 1.8 Tbases / 3 days 21 Maridis Annu. Rev. Genome. Human Genet. 2008
22 Genomics methods of the second generation - Illumina - Fragmentation of sample + ligation of adapters (2 types) + size selection Binding of fragments onto cell surface + initial replication a) Adapter I; b) Adapter II; c) orig. fragment; d) unbound adapters on surface 22
23 Genomics methods of the second generation - Illumina - Bridge formation and polymerase activity using unlabeled dntps Final double stranded bridge a) Full (surface bound) Adapters; b) incomplete Adapters (from aborted polymerase activity: no more space!) 23
24 Genomics methods of the second generation - Illumina - Denaturing of double stranded bridge a) identical (+/- strand) surface bound copies of DNA-fragment Repetition of bridging, amplification, denaturation until a `forest of fragments exists 24
25 Genomics methods of the second generation - Illumina - Removal of adapter I bound fragments, Addition of ddntp like labeled bases + primer (adapter I): Sequencing of base 1 (Laser excitation, recording of fluorescence activity) 25
26 Genomics methods of the second generation - Illumina - Removal sequence elongation terminator Addition of ddntp like labeled bases Sequencing of base 2 Processing of all recorded images into textual format 26
27 Genomics methods of the second generation - Illumina - Advantages: Low error rate Lowest cost per base Tons of data Disadvantages: Must run at very large scale Short read length (50-150bp) Runs take multiple days High startup costs De Novo sequencing difficult Areas of application: DNA sequencing Gene regulation analysis Sequencing-based Transcriptome analysis SNPs and SVs discovery Cytogenetic Analysis ChIP-sequencing Small RNA discovery analysis 27
28 Genomics methods of the second generation - Pacific Bioscience SMRT sequencing - Real time, bound polymerase chain reaction using labeled dntps Pacific Biosciences SMRT (Single Molecule Real Time) Special labeling: fluorescent is situated at the terminal phosphate Incorporation with DNA polymerase releases the label, leaving a natural DNA strand behind. Generates up to 4TB of raw data (per 30 minutes (!!)) Single-molecule sequencing has been developed to circumvent the 2 main biases of PCRdependent sequencing (like 454, Illumina): 1) PCR introduces an uncontrolled bias in template representation because its efficiencies vary as a function of template properties 2) PCR introduces errors (generating false-positive SNPs) 28
29 Genomics methods of the second generation - Pacific Bioscience SMRT sequencing - 29
30 Genomics methods of the second generation - Pacific Bioscience SMRT sequencing - ZMV zero mode waveguide 30
31 Genomics methods of the second generation - Pacific Bioscience SMRT sequencing - Advantages: Very fast Areas of application: Can deliver really long reads de-novo assembly (mean read-length is >5500bp, longest reads can reach Targeted sequencing 30kb) 1 run is not really expensive (~400$ per run) Disadvantages: Only ~ reads per SMRT Cell Need many runs for higher coverage high startup costs B_cUZ8hSYU 31
32 Third generation sequencing - Oxford Nanopore MinION & GridION - Sequencing without fluorescent labels, without fragmenting the DNA Pipetting ions and the entire DNA through a small nanopore located in a synthetic polymer-membrane using a voltage difference A nanopore is the only possibility for current to cross the membrane only small sample volume is required 32
33 Third generation sequencing - Oxford Nanopore MinION & GridION - The inside of the nanopore is engineered for enhanced sensing For each triplet of nucleotides, a characteristic electrical signal caused by the ion-flow is detected The current change can be directly measured Signal for each triplet (overlapping!) is recorded 33
34 Advantages: Third generation sequencing - Oxford Nanopore MinION & GridION - Minimal sample preparation no requirement for polymerase or ligase potential of very long read-lenghts it might well achieve the $1000 per mammalian genome goal the instrument is inexpensive Challenges/Disadavantages: slowing down DNA translocation improving signal/noise ratio Potentially high error rate 34
35 Applications DNA/RNA sequencing can be used for a variety of applications, including: Genome sequencing - De novo sequencing of genomes - resequencing of genomes Detection of variants (SNPs) and mutations exome sequencing Confirmation of clone constructs Detection of methylation events Gene expression studies (transcriptomics) - Whole transcriptome - RNA seq/ small RNA Chip-Seq/PAR-CLIP 35
36 Applications - ChIP sequencing - ChIP-Seq is short for 'chromatin immunoprecipitationsequencing Used to determine the influences of chromatin-associated proteins and transcription factors on the actual transcription. DNA is usually wound up around 'chromatin' Unwinding is necessary for transcription (accessibility) Carried out/aided by transcription-factors and associated proteins How do these work? Where do they bind? => ChIP-Seq tries to deliver the answers. 36
37 Applications - ChIP sequencing - 'Cross-linking' or 'binding' of proteins to DNA 37
38 Applications - ChIP sequencing - Lysate the cell containing the cross-linked proteins 38
39 Applications - ChIP sequencing - Pulldown (magnetic beads) Wash off Undo cross-linking Sequence the DNA (Illumina, SMRT) Rough area (region bp) where protein bound to /interacted with DNA 39
40 Applications - gene expression studies transcriptomics - Transcriptome - set of all mrnas present in certain cell, tissue, organ, - mrna level results from intensity of transcription and mrna stability Transcriptomics - analysis of differences in expression of gene populations under different conditions (treatment, development, disease) - also called expression profiling 40
41 Applications - gene expression studies transcriptomics - Different types of RNA mrna Coding RNA t-rna rrna hnrna sirna mirna Coding RNA Approximately to genes are active in every single cell Different abundance comparing the single mrnas low abundant mrnas about 1 copy per cell highly abundant mrnas more than copies per cells 41
42 Applications - gene expression studies transcriptomics - Why analyzing the transcriptome? How to analyze the transcriptome? allows to analyze expression changes in: using different methods depending on Different cell types the different source of sample/question In different conditions of the to answer: environment/development Microarrays (Affymetrix, Illumina allows to compare healthy vs. Beadchip technology) diseased state RNA-seq (454, Illumina) identification of new genes Real-time quantitative PCR serial analysis of gene expression (SAGE) massively parallel signature sequencing (MPSS) 42
43 Applications - gene expression studies transcriptomics - Workflow of a DNA microarray 43
44 Applications - gene expression studies transcriptomics - Result of a microarray: the heatmap 1 column= 1 sample 1 row = 1 gene Color key: indicating th relative expression 44
45 Applications - gene expression studies transcriptomics - Advantages of microarrays Advantages of RNA-seq affordable fast technique no specific equipment required high resolution Disadvantages of microarrays cross-hybrydisation possible sequences of the targeted genes need to be known material intensive (solutions, RNA) variation among laboratories high-throughput high coverage of mrna compared to microarrays new unidentified exons can be detected can handle all RNA isoforms Disadvantages of RNA-seq high costs computational complexities sample preparation can potentially induce bias (temperature-increased shredding of mrna) 45
46 Thank you! 46
High Throughput Sequencing Technologies. J Fass UCD Genome Center Bioinformatics Core Monday June 16, 2014
 High Throughput Sequencing Technologies J Fass UCD Genome Center Bioinformatics Core Monday June 16, 2014 Sequencing Explosion www.genome.gov/sequencingcosts http://t.co/ka5cvghdqo Sequencing Explosion
High Throughput Sequencing Technologies J Fass UCD Genome Center Bioinformatics Core Monday June 16, 2014 Sequencing Explosion www.genome.gov/sequencingcosts http://t.co/ka5cvghdqo Sequencing Explosion
Overview of Next Generation Sequencing technologies. Céline Keime
 Overview of Next Generation Sequencing technologies Céline Keime keime@igbmc.fr Next Generation Sequencing < Second generation sequencing < General principle < Sequencing by synthesis - Illumina < Sequencing
Overview of Next Generation Sequencing technologies Céline Keime keime@igbmc.fr Next Generation Sequencing < Second generation sequencing < General principle < Sequencing by synthesis - Illumina < Sequencing
High Throughput Sequencing Technologies. J Fass UCD Genome Center Bioinformatics Core Monday September 15, 2014
 High Throughput Sequencing Technologies J Fass UCD Genome Center Bioinformatics Core Monday September 15, 2014 Sequencing Explosion www.genome.gov/sequencingcosts http://t.co/ka5cvghdqo Sequencing Explosion
High Throughput Sequencing Technologies J Fass UCD Genome Center Bioinformatics Core Monday September 15, 2014 Sequencing Explosion www.genome.gov/sequencingcosts http://t.co/ka5cvghdqo Sequencing Explosion
High Throughput Sequencing Technologies. J Fass UCD Genome Center Bioinformatics Core Tuesday December 16, 2014
 High Throughput Sequencing Technologies J Fass UCD Genome Center Bioinformatics Core Tuesday December 16, 2014 Sequencing Explosion www.genome.gov/sequencingcosts http://t.co/ka5cvghdqo Sequencing Explosion
High Throughput Sequencing Technologies J Fass UCD Genome Center Bioinformatics Core Tuesday December 16, 2014 Sequencing Explosion www.genome.gov/sequencingcosts http://t.co/ka5cvghdqo Sequencing Explosion
Third Generation Sequencing
 Third Generation Sequencing By Mohammad Hasan Samiee Aref Medical Genetics Laboratory of Dr. Zeinali History of DNA sequencing 1953 : Discovery of DNA structure by Watson and Crick 1973 : First sequence
Third Generation Sequencing By Mohammad Hasan Samiee Aref Medical Genetics Laboratory of Dr. Zeinali History of DNA sequencing 1953 : Discovery of DNA structure by Watson and Crick 1973 : First sequence
Next Generation Sequencing. Jeroen Van Houdt - Leuven 13/10/2017
 Next Generation Sequencing Jeroen Van Houdt - Leuven 13/10/2017 Landmarks in DNA sequencing 1953 Discovery of DNA double helix structure 1977 A Maxam and W Gilbert "DNA seq by chemical degradation" F Sanger"DNA
Next Generation Sequencing Jeroen Van Houdt - Leuven 13/10/2017 Landmarks in DNA sequencing 1953 Discovery of DNA double helix structure 1977 A Maxam and W Gilbert "DNA seq by chemical degradation" F Sanger"DNA
Next Generation Sequencing (NGS)
 Next Generation Sequencing (NGS) Fernando Alvarez Sección Biomatemática, Facultad de Ciencias, UdelaR 1 Uruguay Montevide o 3 TANGO World Champ 1930 1950 (Maraca 4 Next Generation Sequencing module Next
Next Generation Sequencing (NGS) Fernando Alvarez Sección Biomatemática, Facultad de Ciencias, UdelaR 1 Uruguay Montevide o 3 TANGO World Champ 1930 1950 (Maraca 4 Next Generation Sequencing module Next
Deep Sequencing technologies
 Deep Sequencing technologies Gabriela Salinas 30 October 2017 Transcriptome and Genome Analysis Laboratory http://www.uni-bc.gwdg.de/index.php?id=709 Microarray and Deep-Sequencing Core Facility University
Deep Sequencing technologies Gabriela Salinas 30 October 2017 Transcriptome and Genome Analysis Laboratory http://www.uni-bc.gwdg.de/index.php?id=709 Microarray and Deep-Sequencing Core Facility University
Aaron Liston, Oregon State University Botany 2012 Intro to Next Generation Sequencing Workshop
 Output (bp) Aaron Liston, Oregon State University Growth in Next-Gen Sequencing Capacity 3.5E+11 2002 2004 2006 2008 2010 3.0E+11 2.5E+11 2.0E+11 1.5E+11 1.0E+11 Adapted from Mardis, 2011, Nature 5.0E+10
Output (bp) Aaron Liston, Oregon State University Growth in Next-Gen Sequencing Capacity 3.5E+11 2002 2004 2006 2008 2010 3.0E+11 2.5E+11 2.0E+11 1.5E+11 1.0E+11 Adapted from Mardis, 2011, Nature 5.0E+10
Matthew Tinning Australian Genome Research Facility. July 2012
 Next-Generation Sequencing: an overview of technologies and applications Matthew Tinning Australian Genome Research Facility July 2012 History of Sequencing Where have we been? 1869 Discovery of DNA 1909
Next-Generation Sequencing: an overview of technologies and applications Matthew Tinning Australian Genome Research Facility July 2012 History of Sequencing Where have we been? 1869 Discovery of DNA 1909
The Journey of DNA Sequencing. Chromosomes. What is a genome? Genome size. H. Sunny Sun
 The Journey of DNA Sequencing H. Sunny Sun What is a genome? Genome is the total genetic complement of a living organism. The nuclear genome comprises approximately 3.2 * 10 9 nucleotides of DNA, divided
The Journey of DNA Sequencing H. Sunny Sun What is a genome? Genome is the total genetic complement of a living organism. The nuclear genome comprises approximately 3.2 * 10 9 nucleotides of DNA, divided
High Throughput Sequencing Technologies. UCD Genome Center Bioinformatics Core Monday 15 June 2015
 High Throughput Sequencing Technologies UCD Genome Center Bioinformatics Core Monday 15 June 2015 Sequencing Explosion www.genome.gov/sequencingcosts http://t.co/ka5cvghdqo Sequencing Explosion 2011 PacBio
High Throughput Sequencing Technologies UCD Genome Center Bioinformatics Core Monday 15 June 2015 Sequencing Explosion www.genome.gov/sequencingcosts http://t.co/ka5cvghdqo Sequencing Explosion 2011 PacBio
Sequencing technologies. Jose Blanca COMAV institute bioinf.comav.upv.es
 Sequencing technologies Jose Blanca COMAV institute bioinf.comav.upv.es Outline Sequencing technologies: Sanger 2nd generation sequencing: 3er generation sequencing: 454 Illumina SOLiD Ion Torrent PacBio
Sequencing technologies Jose Blanca COMAV institute bioinf.comav.upv.es Outline Sequencing technologies: Sanger 2nd generation sequencing: 3er generation sequencing: 454 Illumina SOLiD Ion Torrent PacBio
DNA-Sequencing. Technologies & Devices. Matthias Platzer. Genome Analysis Leibniz Institute on Aging - Fritz Lipmann Institute (FLI)
 DNA-Sequencing Technologies & Devices Matthias Platzer Genome Analysis Leibniz Institute on Aging - Fritz Lipmann Institute (FLI) Genome analysis DNA sequencing platforms ABI 3730xl 4/2004 & 6/2006 1 Mb/day,
DNA-Sequencing Technologies & Devices Matthias Platzer Genome Analysis Leibniz Institute on Aging - Fritz Lipmann Institute (FLI) Genome analysis DNA sequencing platforms ABI 3730xl 4/2004 & 6/2006 1 Mb/day,
DNA-Sequencing. Technologies & Devices. Matthias Platzer. Genome Analysis Leibniz Institute on Aging - Fritz Lipmann Institute (FLI)
 DNA-Sequencing Technologies & Devices Matthias Platzer Genome Analysis Leibniz Institute on Aging - Fritz Lipmann Institute (FLI) Genome analysis DNA sequencing platforms ABI 3730xl 4/2004 & 6/2006 1 Mb/day,
DNA-Sequencing Technologies & Devices Matthias Platzer Genome Analysis Leibniz Institute on Aging - Fritz Lipmann Institute (FLI) Genome analysis DNA sequencing platforms ABI 3730xl 4/2004 & 6/2006 1 Mb/day,
Next Generation Sequencing. Tobias Österlund
 Next Generation Sequencing Tobias Österlund tobiaso@chalmers.se NGS part of the course Week 4 Friday 13/2 15.15-17.00 NGS lecture 1: Introduction to NGS, alignment, assembly Week 6 Thursday 26/2 08.00-09.45
Next Generation Sequencing Tobias Österlund tobiaso@chalmers.se NGS part of the course Week 4 Friday 13/2 15.15-17.00 NGS lecture 1: Introduction to NGS, alignment, assembly Week 6 Thursday 26/2 08.00-09.45
Next generation sequencing techniques" Toma Tebaldi Centre for Integrative Biology University of Trento
 Next generation sequencing techniques" Toma Tebaldi Centre for Integrative Biology University of Trento Mattarello September 28, 2009 Sequencing Fundamental task in modern biology read the information
Next generation sequencing techniques" Toma Tebaldi Centre for Integrative Biology University of Trento Mattarello September 28, 2009 Sequencing Fundamental task in modern biology read the information
DNA-Sequencing. Technologies & Devices
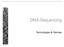 DNA-Sequencing Technologies & Devices Genome analysis DNA sequencing platforms ABI 3730xl 4/2004 & 6/2006 1 Mb/day, 850 nt reads 2 Mb/day, 550 nt reads Roche/454 GS FLX 12/2006 800 Mb/23h, 800 nt reads
DNA-Sequencing Technologies & Devices Genome analysis DNA sequencing platforms ABI 3730xl 4/2004 & 6/2006 1 Mb/day, 850 nt reads 2 Mb/day, 550 nt reads Roche/454 GS FLX 12/2006 800 Mb/23h, 800 nt reads
Next-generation sequencing Technology Overview
 Next-generation sequencing Technology Overview UQ Winter School 2018 Christopher Noune, PhD AGRF Melbourne christopher.noune@agrf.org.au What is NGS? Ion Torrent PGM (Thermo-Fisher) MiSeq (Illumina) High-Throughput
Next-generation sequencing Technology Overview UQ Winter School 2018 Christopher Noune, PhD AGRF Melbourne christopher.noune@agrf.org.au What is NGS? Ion Torrent PGM (Thermo-Fisher) MiSeq (Illumina) High-Throughput
Chapter 7. DNA Microarrays
 Bioinformatics III Structural Bioinformatics and Genome Analysis Chapter 7. DNA Microarrays 7.9 Next Generation Sequencing 454 Sequencing Solexa Illumina Solid TM System Sequencing Process of determining
Bioinformatics III Structural Bioinformatics and Genome Analysis Chapter 7. DNA Microarrays 7.9 Next Generation Sequencing 454 Sequencing Solexa Illumina Solid TM System Sequencing Process of determining
Sequencing technologies. Jose Blanca COMAV institute bioinf.comav.upv.es
 Sequencing technologies Jose Blanca COMAV institute bioinf.comav.upv.es Outline Sequencing technologies: Sanger 2nd generation sequencing: 3er generation sequencing: 454 Illumina SOLiD Ion Torrent PacBio
Sequencing technologies Jose Blanca COMAV institute bioinf.comav.upv.es Outline Sequencing technologies: Sanger 2nd generation sequencing: 3er generation sequencing: 454 Illumina SOLiD Ion Torrent PacBio
Introduction to Next Generation Sequencing (NGS)
 Introduction to Next eneration Sequencing (NS) Simon Rasmussen Assistant Professor enter for Biological Sequence analysis Technical University of Denmark 2012 Today 9.00-9.45: Introduction to NS, How it
Introduction to Next eneration Sequencing (NS) Simon Rasmussen Assistant Professor enter for Biological Sequence analysis Technical University of Denmark 2012 Today 9.00-9.45: Introduction to NS, How it
DNA-Sequencing. Technologies & Devices
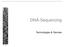 DNA-Sequencing Technologies & Devices Genome analysis DNA sequencing platforms ABI 3730xl 4/2004 & 6/2006 1 Mb/day, 850 nt reads 2 Mb/day, 550 nt reads Roche/454 GS FLX 12/2006 800 Mb/23h, 800 nt reads
DNA-Sequencing Technologies & Devices Genome analysis DNA sequencing platforms ABI 3730xl 4/2004 & 6/2006 1 Mb/day, 850 nt reads 2 Mb/day, 550 nt reads Roche/454 GS FLX 12/2006 800 Mb/23h, 800 nt reads
Contact us for more information and a quotation
 GenePool Information Sheet #1 Installed Sequencing Technologies in the GenePool The GenePool offers sequencing service on three platforms: Sanger (dideoxy) sequencing on ABI 3730 instruments Illumina SOLEXA
GenePool Information Sheet #1 Installed Sequencing Technologies in the GenePool The GenePool offers sequencing service on three platforms: Sanger (dideoxy) sequencing on ABI 3730 instruments Illumina SOLEXA
Next Generation Sequencing Lecture Saarbrücken, 19. March Sequencing Platforms
 Next Generation Sequencing Lecture Saarbrücken, 19. March 2012 Sequencing Platforms Contents Introduction Sequencing Workflow Platforms Roche 454 ABI SOLiD Illumina Genome Anlayzer / HiSeq Problems Quality
Next Generation Sequencing Lecture Saarbrücken, 19. March 2012 Sequencing Platforms Contents Introduction Sequencing Workflow Platforms Roche 454 ABI SOLiD Illumina Genome Anlayzer / HiSeq Problems Quality
Sequencing technologies. Jose Blanca COMAV institute bioinf.comav.upv.es
 Sequencing technologies Jose Blanca COMAV institute bioinf.comav.upv.es Outline Sequencing technologies: Sanger 2nd generation sequencing: 3er generation sequencing: 454 Illumina SOLiD Ion Torrent PacBio
Sequencing technologies Jose Blanca COMAV institute bioinf.comav.upv.es Outline Sequencing technologies: Sanger 2nd generation sequencing: 3er generation sequencing: 454 Illumina SOLiD Ion Torrent PacBio
Wheat CAP Gene Expression with RNA-Seq
 Wheat CAP Gene Expression with RNA-Seq July 9 th -13 th, 2018 Overview of the workshop, Alina Akhunova http://www.ksre.k-state.edu/igenomics/workshops/ RNA-Seq Workshop Activities Lectures Laboratory Molecular
Wheat CAP Gene Expression with RNA-Seq July 9 th -13 th, 2018 Overview of the workshop, Alina Akhunova http://www.ksre.k-state.edu/igenomics/workshops/ RNA-Seq Workshop Activities Lectures Laboratory Molecular
High Throughput Sequencing the Multi-Tool of Life Sciences. Lutz Froenicke DNA Technologies and Expression Analysis Cores UCD Genome Center
 High Throughput Sequencing the Multi-Tool of Life Sciences Lutz Froenicke DNA Technologies and Expression Analysis Cores UCD Genome Center Complementary Approaches Illumina Still-imaging of clusters (~1000
High Throughput Sequencing the Multi-Tool of Life Sciences Lutz Froenicke DNA Technologies and Expression Analysis Cores UCD Genome Center Complementary Approaches Illumina Still-imaging of clusters (~1000
Bioinformatics Advice on Experimental Design
 Bioinformatics Advice on Experimental Design Where do I start? Please refer to the following guide to better plan your experiments for good statistical analysis, best suited for your research needs. Statistics
Bioinformatics Advice on Experimental Design Where do I start? Please refer to the following guide to better plan your experiments for good statistical analysis, best suited for your research needs. Statistics
Next Gen Sequencing. Expansion of sequencing technology. Contents
 Next Gen Sequencing Contents 1 Expansion of sequencing technology 2 The Next Generation of Sequencing: High-Throughput Technologies 3 High Throughput Sequencing Applied to Genome Sequencing (TEDed CC BY-NC-ND
Next Gen Sequencing Contents 1 Expansion of sequencing technology 2 The Next Generation of Sequencing: High-Throughput Technologies 3 High Throughput Sequencing Applied to Genome Sequencing (TEDed CC BY-NC-ND
INTRODUCCIÓ A LES TECNOLOGIES DE 'NEXT GENERATION SEQUENCING'
 INTRODUCCIÓ A LES TECNOLOGIES DE 'NEXT GENERATION SEQUENCING' Bioinformàtica per a la Recerca Biomèdica Ricardo Gonzalo Sanz ricardo.gonzalo@vhir.org 14/12/2016 1. Introduction to NGS 2. First Generation
INTRODUCCIÓ A LES TECNOLOGIES DE 'NEXT GENERATION SEQUENCING' Bioinformàtica per a la Recerca Biomèdica Ricardo Gonzalo Sanz ricardo.gonzalo@vhir.org 14/12/2016 1. Introduction to NGS 2. First Generation
Sequencing techniques and applications
 I519 Introduction to Bioinformatics Sequencing techniques and applications Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Contents Sequencing techniques Sanger sequencing Next generation
I519 Introduction to Bioinformatics Sequencing techniques and applications Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Contents Sequencing techniques Sanger sequencing Next generation
Sequencing technologies
 Sequencing technologies part of High-Throughput Analyzes of Genome Sequenzes Computational EvoDevo University of Leipzig Leipzig, WS 2014/15 Sanger Sequencing (Chain Termination Method) Sequencing of one
Sequencing technologies part of High-Throughput Analyzes of Genome Sequenzes Computational EvoDevo University of Leipzig Leipzig, WS 2014/15 Sanger Sequencing (Chain Termination Method) Sequencing of one
High Throughput Sequencing the Multi-Tool of Life Sciences. Lutz Froenicke DNA Technologies and Expression Analysis Cores UCD Genome Center
 High Throughput Sequencing the Multi-Tool of Life Sciences Lutz Froenicke DNA Technologies and Expression Analysis Cores UCD Genome Center DNA Technologies & Expression Analysis Cores HT Sequencing (Illumina
High Throughput Sequencing the Multi-Tool of Life Sciences Lutz Froenicke DNA Technologies and Expression Analysis Cores UCD Genome Center DNA Technologies & Expression Analysis Cores HT Sequencing (Illumina
Human genome sequence
 NGS: the basics Human genome sequence June 26th 2000: official announcement of the completion of the draft of the human genome sequence (truly finished in 2004) Francis Collins Craig Venter HGP: 3 billion
NGS: the basics Human genome sequence June 26th 2000: official announcement of the completion of the draft of the human genome sequence (truly finished in 2004) Francis Collins Craig Venter HGP: 3 billion
The Expanded Illumina Sequencing Portfolio New Sample Prep Solutions and Workflow
 The Expanded Illumina Sequencing Portfolio New Sample Prep Solutions and Workflow Marcus Hausch, Ph.D. 2010 Illumina, Inc. All rights reserved. Illumina, illuminadx, Solexa, Making Sense Out of Life, Oligator,
The Expanded Illumina Sequencing Portfolio New Sample Prep Solutions and Workflow Marcus Hausch, Ph.D. 2010 Illumina, Inc. All rights reserved. Illumina, illuminadx, Solexa, Making Sense Out of Life, Oligator,
Next-Generation Sequencing. Technologies
 Next-Generation Next-Generation Sequencing Technologies Sequencing Technologies Nicholas E. Navin, Ph.D. MD Anderson Cancer Center Dept. Genetics Dept. Bioinformatics Introduction to Bioinformatics GS011062
Next-Generation Next-Generation Sequencing Technologies Sequencing Technologies Nicholas E. Navin, Ph.D. MD Anderson Cancer Center Dept. Genetics Dept. Bioinformatics Introduction to Bioinformatics GS011062
Gene Expression Technology
 Gene Expression Technology Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Gene expression Gene expression is the process by which information from a gene
Gene Expression Technology Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Gene expression Gene expression is the process by which information from a gene
Genome Resequencing. Rearrangements. SNPs, Indels CNVs. De novo genome Sequencing. Metagenomics. Exome Sequencing. RNA-seq Gene Expression
 Genome Resequencing De novo genome Sequencing SNPs, Indels CNVs Rearrangements Metagenomics RNA-seq Gene Expression Splice Isoform Abundance High Throughput Short Read Sequencing: Illumina Exome Sequencing
Genome Resequencing De novo genome Sequencing SNPs, Indels CNVs Rearrangements Metagenomics RNA-seq Gene Expression Splice Isoform Abundance High Throughput Short Read Sequencing: Illumina Exome Sequencing
Wet-lab Considerations for Illumina data analysis
 Wet-lab Considerations for Illumina data analysis Based on a presentation by Henriette O Geen Lutz Froenicke DNA Technologies and Expression Analysis Cores UCD Genome Center Complementary Approaches Illumina
Wet-lab Considerations for Illumina data analysis Based on a presentation by Henriette O Geen Lutz Froenicke DNA Technologies and Expression Analysis Cores UCD Genome Center Complementary Approaches Illumina
NGS technologies: a user s guide. Karim Gharbi & Mark Blaxter
 NGS technologies: a user s guide Karim Gharbi & Mark Blaxter genepool-manager@ed.ac.uk Natural history of sequencing 2 Brief history of sequencing 100s bp throughput 100 Gb 1977 1986 1995 1999 2005 2007
NGS technologies: a user s guide Karim Gharbi & Mark Blaxter genepool-manager@ed.ac.uk Natural history of sequencing 2 Brief history of sequencing 100s bp throughput 100 Gb 1977 1986 1995 1999 2005 2007
2/5/16. Honeypot Ants. DNA sequencing, Transcriptomics and Genomics. Gene sequence changes? And/or gene expression changes?
 2/5/16 DNA sequencing, Transcriptomics and Genomics Honeypot Ants "nequacatl" BY2208, Mani Lecture 3 Gene sequence changes? And/or gene expression changes? gene expression differences DNA sequencing, Transcriptomics
2/5/16 DNA sequencing, Transcriptomics and Genomics Honeypot Ants "nequacatl" BY2208, Mani Lecture 3 Gene sequence changes? And/or gene expression changes? gene expression differences DNA sequencing, Transcriptomics
Genome Sequencing. I: Methods. MMG 835, SPRING 2016 Eukaryotic Molecular Genetics. George I. Mias
 Genome Sequencing I: Methods MMG 835, SPRING 2016 Eukaryotic Molecular Genetics George I. Mias Department of Biochemistry and Molecular Biology gmias@msu.edu Sequencing Methods Cost of Sequencing Wetterstrand
Genome Sequencing I: Methods MMG 835, SPRING 2016 Eukaryotic Molecular Genetics George I. Mias Department of Biochemistry and Molecular Biology gmias@msu.edu Sequencing Methods Cost of Sequencing Wetterstrand
1. Introduction Gene regulation Genomics and genome analyses
 1. Introduction Gene regulation Genomics and genome analyses 2. Gene regulation tools and methods Regulatory sequences and motif discovery TF binding sites Databases 3. Technologies Microarrays Deep sequencing
1. Introduction Gene regulation Genomics and genome analyses 2. Gene regulation tools and methods Regulatory sequences and motif discovery TF binding sites Databases 3. Technologies Microarrays Deep sequencing
The New Genome Analyzer IIx Delivering more data, faster, and easier than ever before. Jeremy Preston, PhD Marketing Manager, Sequencing
 The New Genome Analyzer IIx Delivering more data, faster, and easier than ever before Jeremy Preston, PhD Marketing Manager, Sequencing Illumina Genome Analyzer: a Paradigm Shift 2000x gain in efficiency
The New Genome Analyzer IIx Delivering more data, faster, and easier than ever before Jeremy Preston, PhD Marketing Manager, Sequencing Illumina Genome Analyzer: a Paradigm Shift 2000x gain in efficiency
Concepts and methods in sequencing and genome assembly
 BCM-2002 Concepts and methods in sequencing and genome assembly http://megasun.bch.umontreal.ca/papers/bcm-2002/sequencing-bcm2002-nov2015.pdf B. Franz LANG, Département de Biochimie Bureau: H307-15 Courrier
BCM-2002 Concepts and methods in sequencing and genome assembly http://megasun.bch.umontreal.ca/papers/bcm-2002/sequencing-bcm2002-nov2015.pdf B. Franz LANG, Département de Biochimie Bureau: H307-15 Courrier
Genome 373: High- Throughput DNA Sequencing. Doug Fowler
 Genome 373: High- Throughput DNA Sequencing Doug Fowler Tasks give ML unity We learned about three tasks that are commonly encountered in ML Models/Algorithms Give ML Diversity Classification Regression
Genome 373: High- Throughput DNA Sequencing Doug Fowler Tasks give ML unity We learned about three tasks that are commonly encountered in ML Models/Algorithms Give ML Diversity Classification Regression
 www.illumina.com/hiseq www.illumina.com FOR RESEARCH USE ONLY 2012 2014 Illumina, Inc. All rights reserved. Illumina, BaseSpace, cbot, CSPro, Genetic Energy, HiSeq, Nextera, TruSeq, the pumpkin orange
www.illumina.com/hiseq www.illumina.com FOR RESEARCH USE ONLY 2012 2014 Illumina, Inc. All rights reserved. Illumina, BaseSpace, cbot, CSPro, Genetic Energy, HiSeq, Nextera, TruSeq, the pumpkin orange
Ultrasequencing: methods and applications of the new generation sequencing platforms
 Ultrasequencing: methods and applications of the new generation sequencing platforms Nuria Tubío Santamaría Course: Genomics Universitat Autònoma de Barcelona 1 Introduction Clasical methods of sequencing:
Ultrasequencing: methods and applications of the new generation sequencing platforms Nuria Tubío Santamaría Course: Genomics Universitat Autònoma de Barcelona 1 Introduction Clasical methods of sequencing:
Welcome to the NGS webinar series
 Welcome to the NGS webinar series Webinar 1 NGS: Introduction to technology, and applications NGS Technology Webinar 2 Targeted NGS for Cancer Research NGS in cancer Webinar 3 NGS: Data analysis for genetic
Welcome to the NGS webinar series Webinar 1 NGS: Introduction to technology, and applications NGS Technology Webinar 2 Targeted NGS for Cancer Research NGS in cancer Webinar 3 NGS: Data analysis for genetic
Sequence Assembly and Next Generation Sequencing Informatics CBPS7711
 Sequence Assembly and Next Generation Sequencing Informatics CBPS7711 Oct 5, 2010 Sonia Leach, PhD Assistant Professor National Jewish Health sonia.leach@gmail.com Next Generation Sequencing Increasing
Sequence Assembly and Next Generation Sequencing Informatics CBPS7711 Oct 5, 2010 Sonia Leach, PhD Assistant Professor National Jewish Health sonia.leach@gmail.com Next Generation Sequencing Increasing
DNA-Sequenzierung. Technologien & Geräte
 DNA-Sequenzierung Technologien & Geräte Genome analysis DNA sequencing platforms ABI 3730xl 4/2004 & 6/2006 1 Mb/day, 850 nt reads 2 Mb/day, 550 nt reads Roche/454 GS FLX 12/2006 400 Mb/7h, 350 nt reads
DNA-Sequenzierung Technologien & Geräte Genome analysis DNA sequencing platforms ABI 3730xl 4/2004 & 6/2006 1 Mb/day, 850 nt reads 2 Mb/day, 550 nt reads Roche/454 GS FLX 12/2006 400 Mb/7h, 350 nt reads
CM581A2: NEXT GENERATION SEQUENCING PLATFORMS AND LIBRARY GENERATION
 CM581A2: NEXT GENERATION SEQUENCING PLATFORMS AND LIBRARY GENERATION Fall 2015 Instructors: Coordinator: Carol Wilusz, Associate Professor MIP, CMB Instructor: Dan Sloan, Assistant Professor, Biology,
CM581A2: NEXT GENERATION SEQUENCING PLATFORMS AND LIBRARY GENERATION Fall 2015 Instructors: Coordinator: Carol Wilusz, Associate Professor MIP, CMB Instructor: Dan Sloan, Assistant Professor, Biology,
Biochemistry 412. New Strategies, Technologies, & Applications For DNA Sequencing. 12 February 2008
 Biochemistry 412 New Strategies, Technologies, & Applications For DNA Sequencing 12 February 2008 Note: Scale is wrong!! (at least for sequences) 10 6 In 1980, the sequencing cost per finished bp $1.00
Biochemistry 412 New Strategies, Technologies, & Applications For DNA Sequencing 12 February 2008 Note: Scale is wrong!! (at least for sequences) 10 6 In 1980, the sequencing cost per finished bp $1.00
Phenotype analysis: biological-biochemical analysis. Genotype analysis: molecular and physical analysis
 1 Genetic Analysis Phenotype analysis: biological-biochemical analysis Behaviour under specific environmental conditions Behaviour of specific genetic configurations Behaviour of progeny in crosses - Genotype
1 Genetic Analysis Phenotype analysis: biological-biochemical analysis Behaviour under specific environmental conditions Behaviour of specific genetic configurations Behaviour of progeny in crosses - Genotype
How much sequencing do I need? Emily Crisovan Genomics Core
 How much sequencing do I need? Emily Crisovan Genomics Core How much sequencing? Three questions: 1. How much sequence is required for good experimental design? 2. What type of sequencing run is best?
How much sequencing do I need? Emily Crisovan Genomics Core How much sequencing? Three questions: 1. How much sequence is required for good experimental design? 2. What type of sequencing run is best?
Introduction to Bioinformatics and Gene Expression Technologies
 Introduction to Bioinformatics and Gene Expression Technologies Utah State University Fall 2017 Statistical Bioinformatics (Biomedical Big Data) Notes 1 1 Vocabulary Gene: hereditary DNA sequence at a
Introduction to Bioinformatics and Gene Expression Technologies Utah State University Fall 2017 Statistical Bioinformatics (Biomedical Big Data) Notes 1 1 Vocabulary Gene: hereditary DNA sequence at a
Introduction to Bioinformatics and Gene Expression Technologies
 Vocabulary Introduction to Bioinformatics and Gene Expression Technologies Utah State University Fall 2017 Statistical Bioinformatics (Biomedical Big Data) Notes 1 Gene: Genetics: Genome: Genomics: hereditary
Vocabulary Introduction to Bioinformatics and Gene Expression Technologies Utah State University Fall 2017 Statistical Bioinformatics (Biomedical Big Data) Notes 1 Gene: Genetics: Genome: Genomics: hereditary
How much sequencing do I need? Emily Crisovan Genomics Core September 26, 2018
 How much sequencing do I need? Emily Crisovan Genomics Core September 26, 2018 How much sequencing? Three questions: 1. How much sequence is required for good experimental design? 2. What type of sequencing
How much sequencing do I need? Emily Crisovan Genomics Core September 26, 2018 How much sequencing? Three questions: 1. How much sequence is required for good experimental design? 2. What type of sequencing
Functional Genomics Research Stream. Research Meetings: November 2 & 3, 2009 Next Generation Sequencing
 Functional Genomics Research Stream Research Meetings: November 2 & 3, 2009 Next Generation Sequencing Current Issues Research Meetings: Meet with me this Thursday or Friday. (bring laboratory notebook
Functional Genomics Research Stream Research Meetings: November 2 & 3, 2009 Next Generation Sequencing Current Issues Research Meetings: Meet with me this Thursday or Friday. (bring laboratory notebook
High throughput DNA Sequencing. An Equal Opportunity University!
 High throughput DNA Sequencing An Equal Opportunity University! irst Generation DNA sequencing utilize chain terminator technologies (adaptation of Sanger sequencing) Adapt fluorescence chemistry, high-resolution
High throughput DNA Sequencing An Equal Opportunity University! irst Generation DNA sequencing utilize chain terminator technologies (adaptation of Sanger sequencing) Adapt fluorescence chemistry, high-resolution
Genetics and Genomics in Medicine Chapter 3. Questions & Answers
 Genetics and Genomics in Medicine Chapter 3 Multiple Choice Questions Questions & Answers Question 3.1 Which of the following statements, if any, is false? a) Amplifying DNA means making many identical
Genetics and Genomics in Medicine Chapter 3 Multiple Choice Questions Questions & Answers Question 3.1 Which of the following statements, if any, is false? a) Amplifying DNA means making many identical
Phenotype analysis: biological-biochemical analysis. Genotype analysis: molecular and physical analysis
 1 Genetic Analysis Phenotype analysis: biological-biochemical analysis Behaviour under specific environmental conditions Behaviour of specific genetic configurations Behaviour of progeny in crosses - Genotype
1 Genetic Analysis Phenotype analysis: biological-biochemical analysis Behaviour under specific environmental conditions Behaviour of specific genetic configurations Behaviour of progeny in crosses - Genotype
Understanding the science and technology of whole genome sequencing
 Understanding the science and technology of whole genome sequencing Dag Undlien Department of Medical Genetics Oslo University Hospital University of Oslo and The Norwegian Sequencing Centre d.e.undlien@medisin.uio.no
Understanding the science and technology of whole genome sequencing Dag Undlien Department of Medical Genetics Oslo University Hospital University of Oslo and The Norwegian Sequencing Centre d.e.undlien@medisin.uio.no
Using New ThiNGS on Small Things. Shane Byrne
 Using New ThiNGS on Small Things Shane Byrne Next Generation Sequencing New Things Small Things NGS Next Generation Sequencing = 2 nd generation of sequencing 454 GS FLX, SOLiD, GAIIx, HiSeq, MiSeq, Ion
Using New ThiNGS on Small Things Shane Byrne Next Generation Sequencing New Things Small Things NGS Next Generation Sequencing = 2 nd generation of sequencing 454 GS FLX, SOLiD, GAIIx, HiSeq, MiSeq, Ion
resequencing storage SNP ncrna metagenomics private trio de novo exome ncrna RNA DNA bioinformatics RNA-seq comparative genomics
 RNA Sequencing T TM variation genetics validation SNP ncrna metagenomics private trio de novo exome mendelian ChIP-seq RNA DNA bioinformatics custom target high-throughput resequencing storage ncrna comparative
RNA Sequencing T TM variation genetics validation SNP ncrna metagenomics private trio de novo exome mendelian ChIP-seq RNA DNA bioinformatics custom target high-throughput resequencing storage ncrna comparative
The Genome Analysis Centre. Building Excellence in Genomics and Computa5onal Bioscience
 Building Excellence in Genomics and Computa5onal Bioscience Resequencing approaches Sarah Ayling Crop Genomics and Diversity sarah.ayling@tgac.ac.uk Why re- sequence plants? To iden
Building Excellence in Genomics and Computa5onal Bioscience Resequencing approaches Sarah Ayling Crop Genomics and Diversity sarah.ayling@tgac.ac.uk Why re- sequence plants? To iden
Basics of RNA-Seq. (With a Focus on Application to Single Cell RNA-Seq) Michael Kelly, PhD Team Lead, NCI Single Cell Analysis Facility
 2018 ABRF Meeting Satellite Workshop 4 Bridging the Gap: Isolation to Translation (Single Cell RNA-Seq) Sunday, April 22 Basics of RNA-Seq (With a Focus on Application to Single Cell RNA-Seq) Michael Kelly,
2018 ABRF Meeting Satellite Workshop 4 Bridging the Gap: Isolation to Translation (Single Cell RNA-Seq) Sunday, April 22 Basics of RNA-Seq (With a Focus on Application to Single Cell RNA-Seq) Michael Kelly,
Lecture 8: Sequencing and SNP. Sept 15, 2006
 Lecture 8: Sequencing and SNP Sept 15, 2006 Announcements Random questioning during literature discussion sessions starts next week for real! Schedule changes Moved QTL lecture up Removed landscape genetics
Lecture 8: Sequencing and SNP Sept 15, 2006 Announcements Random questioning during literature discussion sessions starts next week for real! Schedule changes Moved QTL lecture up Removed landscape genetics
RNA Sequencing. Next gen insight into transcriptomes , Elio Schijlen
 RNA Sequencing Next gen insight into transcriptomes 05-06-2013, Elio Schijlen Transcriptome complete set of transcripts in a cell, and their quantity, for a specific developmental stage or physiological
RNA Sequencing Next gen insight into transcriptomes 05-06-2013, Elio Schijlen Transcriptome complete set of transcripts in a cell, and their quantity, for a specific developmental stage or physiological
Transcriptomics analysis with RNA seq: an overview Frederik Coppens
 Transcriptomics analysis with RNA seq: an overview Frederik Coppens Platforms Applications Analysis Quantification RNA content Platforms Platforms Short (few hundred bases) Long reads (multiple kilobases)
Transcriptomics analysis with RNA seq: an overview Frederik Coppens Platforms Applications Analysis Quantification RNA content Platforms Platforms Short (few hundred bases) Long reads (multiple kilobases)
High throughput sequencing technologies
 High throughput sequencing technologies and NGS applications Mei-yeh Lu 呂美曄 High Throughput Sequencing Core Manager g g p q g g Academia Sinica 6/30/2011 Outlines Evolution of sequencing technologies Sanger
High throughput sequencing technologies and NGS applications Mei-yeh Lu 呂美曄 High Throughput Sequencing Core Manager g g p q g g Academia Sinica 6/30/2011 Outlines Evolution of sequencing technologies Sanger
CSC Assignment1SequencingReview- 1109_Su N_NEXT_GENERATION_SEQUENCING.docx By Anonymous. Similarity Index
 Page 1 of 6 Document Viewer TurnitinUK Originality Report Processed on: 05-Dec-20 10:49 AM GMT ID: 13 Word Count: 1587 Submitted: 1 CSC8313-201 - Assignment1SequencingReview- 1109_Su N_NEXT_GENERATION_SEQUENCING.docx
Page 1 of 6 Document Viewer TurnitinUK Originality Report Processed on: 05-Dec-20 10:49 AM GMT ID: 13 Word Count: 1587 Submitted: 1 CSC8313-201 - Assignment1SequencingReview- 1109_Su N_NEXT_GENERATION_SEQUENCING.docx
DNA Sequencing. Happiness Kumburu BSU- workshop Nov, 2016
 DNA Sequencing Happiness Kumburu BSU- workshop Nov, 2016 OUT LINE History of DNA sequencing Purpose of DNA sequencing DNA Sequencing Methods Advantages and Disadvantages References DNA SEQUENCING DNA sequencing-the
DNA Sequencing Happiness Kumburu BSU- workshop Nov, 2016 OUT LINE History of DNA sequencing Purpose of DNA sequencing DNA Sequencing Methods Advantages and Disadvantages References DNA SEQUENCING DNA sequencing-the
Applications of Next Generation Sequencing in Metagenomics Studies
 Applications of Next Generation Sequencing in Metagenomics Studies Francesca Rizzo, PhD Genomix4life Laboratory of Molecular Medicine and Genomics Department of Medicine and Surgery University of Salerno
Applications of Next Generation Sequencing in Metagenomics Studies Francesca Rizzo, PhD Genomix4life Laboratory of Molecular Medicine and Genomics Department of Medicine and Surgery University of Salerno
Application of NGS (nextgeneration. for studying RNA regulation. Sung Wook Chi. Sungkyunkwan University (SKKU) Samsung Medical Center (SMC)
 Application of NGS (nextgeneration sequencing) for studying RNA regulation Samsung Advanced Institute of Heath Sciences and Technology (SAIHST) Sungkyunkwan University (SKKU) Samsung Research Institute
Application of NGS (nextgeneration sequencing) for studying RNA regulation Samsung Advanced Institute of Heath Sciences and Technology (SAIHST) Sungkyunkwan University (SKKU) Samsung Research Institute
Outline. General principles of clonal sequencing Analysis principles Applications CNV analysis Genome architecture
 The use of new sequencing technologies for genome analysis Chris Mattocks National Genetics Reference Laboratory (Wessex) NGRL (Wessex) 2008 Outline General principles of clonal sequencing Analysis principles
The use of new sequencing technologies for genome analysis Chris Mattocks National Genetics Reference Laboratory (Wessex) NGRL (Wessex) 2008 Outline General principles of clonal sequencing Analysis principles
RNA-Seq data analysis course September 7-9, 2015
 RNA-Seq data analysis course September 7-9, 2015 Peter-Bram t Hoen (LUMC) Jan Oosting (LUMC) Celia van Gelder, Jacintha Valk (BioSB) Anita Remmelzwaal (LUMC) Expression profiling DNA mrna protein Comprehensive
RNA-Seq data analysis course September 7-9, 2015 Peter-Bram t Hoen (LUMC) Jan Oosting (LUMC) Celia van Gelder, Jacintha Valk (BioSB) Anita Remmelzwaal (LUMC) Expression profiling DNA mrna protein Comprehensive
DNA and genome sequencing. Matthew Hudson Dept of Crop Sciences University of Illinois
 DNA and genome sequencing Matthew Hudson Dept of Crop Sciences University of Illinois Genome projects 2,424 ongoing genome projects 696 for eukaryotes 520 completed genomes 47 from eukaryotes Almost every
DNA and genome sequencing Matthew Hudson Dept of Crop Sciences University of Illinois Genome projects 2,424 ongoing genome projects 696 for eukaryotes 520 completed genomes 47 from eukaryotes Almost every
Next Generation Sequencing: An Overview
 Next Generation Sequencing: An Overview Cavan Reilly November 13, 2017 Table of contents Next generation sequencing NGS and microarrays Study design Quality assessment Burrows Wheeler transform Next generation
Next Generation Sequencing: An Overview Cavan Reilly November 13, 2017 Table of contents Next generation sequencing NGS and microarrays Study design Quality assessment Burrows Wheeler transform Next generation
NEXT GENERATION SEQUENCING. Farhat Habib
 NEXT GENERATION SEQUENCING HISTORY HISTORY Sanger Dominant for last ~30 years 1000bp longest read Based on primers so not good for repetitive or SNPs sites HISTORY Sanger Dominant for last ~30 years 1000bp
NEXT GENERATION SEQUENCING HISTORY HISTORY Sanger Dominant for last ~30 years 1000bp longest read Based on primers so not good for repetitive or SNPs sites HISTORY Sanger Dominant for last ~30 years 1000bp
Galaxy Workshop
 Galaxy Workshop 1-8-13 Intros: Tom Bair thomas-bair@uiowa.edu Ann Black-Ziegelbein annblack@eng.uiowa.edu Srinivas Maddhi srinivas-maddhi@uiowa.edu What is galaxy good for Access to resources Documentation
Galaxy Workshop 1-8-13 Intros: Tom Bair thomas-bair@uiowa.edu Ann Black-Ziegelbein annblack@eng.uiowa.edu Srinivas Maddhi srinivas-maddhi@uiowa.edu What is galaxy good for Access to resources Documentation
Design. Construction. Characterization
 Design Construction Characterization DNA mrna (messenger) A C C transcription translation C A C protein His A T G C T A C G Plasmids replicon copy number incompatibility selection marker origin of replication
Design Construction Characterization DNA mrna (messenger) A C C transcription translation C A C protein His A T G C T A C G Plasmids replicon copy number incompatibility selection marker origin of replication
TREE CODE PRODUCT BROCHURE
 TREE CODE PRODUCT BROCHURE Single Molecule, Real-Time (SMRT) Sequencing technology offers: Long read sequencing ~10 Gb with 20 kb average read lengths for WGS ~20 Gb with 40 kb average read length for
TREE CODE PRODUCT BROCHURE Single Molecule, Real-Time (SMRT) Sequencing technology offers: Long read sequencing ~10 Gb with 20 kb average read lengths for WGS ~20 Gb with 40 kb average read length for
FGCZ NEWSLETTER FALL Next Generation Sequencing at the Functional Genomics Center Zurich
 FGCZ NEWSLETTER FALL 2011 newsletter Technologies, Applications, and Access to Support Next Generation Sequencing at the Functional Genomics Center Zurich OVERVIEW 1 NGS AT THE FGCZ Technologies and organization
FGCZ NEWSLETTER FALL 2011 newsletter Technologies, Applications, and Access to Support Next Generation Sequencing at the Functional Genomics Center Zurich OVERVIEW 1 NGS AT THE FGCZ Technologies and organization
Introduction to NGS. Josef K Vogt Slides by: Simon Rasmussen Next Generation Sequencing Analysis
 Introduction to NGS Josef K Vogt Slides by: Simon Rasmussen 2017 Life science data deluge Massive unstructured data from several areas DNA, patient journals, proteomics, imaging,... Impacts Industry, Environment,
Introduction to NGS Josef K Vogt Slides by: Simon Rasmussen 2017 Life science data deluge Massive unstructured data from several areas DNA, patient journals, proteomics, imaging,... Impacts Industry, Environment,
Next Generation Sequencing. Simon Rasmussen Assistant Professor Center for Biological Sequence analysis Technical University of Denmark
 Next eneration Sequencing Simon Rasmussen Assistant Professor enter for Biological Sequence analysis Technical University of Denmark DNA Sequencing DNA sequencing Reading the order of bases in DNA fragments
Next eneration Sequencing Simon Rasmussen Assistant Professor enter for Biological Sequence analysis Technical University of Denmark DNA Sequencing DNA sequencing Reading the order of bases in DNA fragments
Advanced Technology in Phytoplasma Research
 Advanced Technology in Phytoplasma Research Sequencing and Phylogenetics Wednesday July 8 Pauline Wang pauline.wang@utoronto.ca Lethal Yellowing Disease Phytoplasma Healthy palm Lethal yellowing of palm
Advanced Technology in Phytoplasma Research Sequencing and Phylogenetics Wednesday July 8 Pauline Wang pauline.wang@utoronto.ca Lethal Yellowing Disease Phytoplasma Healthy palm Lethal yellowing of palm
NextGen Sequencing and Target Enrichment
 NextGen Sequencing and Target Enrichment Laurent FARINELLI 7 September 2010 Agilent 3rd Analytic Forum Basel, Switzerland Outline The illumina HiSEQ 2000 system Applications Target enrichment Outlook 7
NextGen Sequencing and Target Enrichment Laurent FARINELLI 7 September 2010 Agilent 3rd Analytic Forum Basel, Switzerland Outline The illumina HiSEQ 2000 system Applications Target enrichment Outlook 7
A Crash Course in NGS for GI Pathologists. Sandra O Toole
 A Crash Course in NGS for GI Pathologists Sandra O Toole The Sanger Technique First generation sequencing Uses dideoxynucleotides (dideoxyadenine, dideoxyguanine, etc) These are molecules that resemble
A Crash Course in NGS for GI Pathologists Sandra O Toole The Sanger Technique First generation sequencing Uses dideoxynucleotides (dideoxyadenine, dideoxyguanine, etc) These are molecules that resemble
Selected Techniques Part I
 1 Selected Techniques Part I Gel Electrophoresis Can be both qualitative and quantitative Qualitative About what size is the fragment? How many fragments are present? Is there in insert or not? Quantitative
1 Selected Techniques Part I Gel Electrophoresis Can be both qualitative and quantitative Qualitative About what size is the fragment? How many fragments are present? Is there in insert or not? Quantitative
NextGen Sequencing Technologies Sequencing overview
 Outline Conventional NextGen High-throughput sequencing (Next-Gen sequencing) technologies. Illumina sequencing in detail. Quality control. Sequence coverage. Multiplexing. FASTQ files. Shendure and Ji
Outline Conventional NextGen High-throughput sequencing (Next-Gen sequencing) technologies. Illumina sequencing in detail. Quality control. Sequence coverage. Multiplexing. FASTQ files. Shendure and Ji
HLA-Typing Strategies
 HLA-Typing Strategies Cologne, 13.5.2017 Joannis Mytilineos MD, PhD Department of Transplantation Immunology Institute for Clinical Transfusion Medicine and Immunogenetics German Red Cross Blood Transfusion
HLA-Typing Strategies Cologne, 13.5.2017 Joannis Mytilineos MD, PhD Department of Transplantation Immunology Institute for Clinical Transfusion Medicine and Immunogenetics German Red Cross Blood Transfusion
DNA Sequencing by Ion Torrent. Marc Lavergne CHEM 4590
 DNA Sequencing by Ion Torrent Marc Lavergne CHEM 4590 OVERVIEW History DNA Synthesis and First-Gen Sequencing Technology Sequencing Signal Detection Advantages/Disadvantages Applications Current Research
DNA Sequencing by Ion Torrent Marc Lavergne CHEM 4590 OVERVIEW History DNA Synthesis and First-Gen Sequencing Technology Sequencing Signal Detection Advantages/Disadvantages Applications Current Research
Ultrasequencing: Methods and Applications of the New Generation Sequencing Platforms
 Ultrasequencing: Methods and Applications of the New Generation Sequencing Platforms Laura Moya Andérico Master in Advanced Genetics Genomics Class December 16 th, 2015 Brief Overview First-generation
Ultrasequencing: Methods and Applications of the New Generation Sequencing Platforms Laura Moya Andérico Master in Advanced Genetics Genomics Class December 16 th, 2015 Brief Overview First-generation
Next-generation sequencing technologies
 Next-generation sequencing technologies NGS applications Illumina sequencing workflow Overview Sequencing by ligation Short-read NGS Sequencing by synthesis Illumina NGS Single-molecule approach Long-read
Next-generation sequencing technologies NGS applications Illumina sequencing workflow Overview Sequencing by ligation Short-read NGS Sequencing by synthesis Illumina NGS Single-molecule approach Long-read
Molecular Biology and Functional Genomic Core Facility
 Molecular Biology and Functional Genomic Core Facility General Presentation Dr Odile Neyret Core Manager Myriam Rondeau Research Assistant Agnès Dumont Research Assistant Institut de recherche clinique
Molecular Biology and Functional Genomic Core Facility General Presentation Dr Odile Neyret Core Manager Myriam Rondeau Research Assistant Agnès Dumont Research Assistant Institut de recherche clinique
you can see that if if you look into the you know the capability kilobases per day, per machine kind of calculation if you do.
 Functional Genomics Professor S Ganesh Department of Biological Sciences & Bioengineering Indian Institute of Technology Kanpur Lecture No 11 DNA Sequencing Methods Part 2 So welcome back to this course
Functional Genomics Professor S Ganesh Department of Biological Sciences & Bioengineering Indian Institute of Technology Kanpur Lecture No 11 DNA Sequencing Methods Part 2 So welcome back to this course
DNA Sequencing and Assembly
 DNA Sequencing and Assembly CS 262 Lecture Notes, Winter 2016 February 2nd, 2016 Scribe: Mark Berger Abstract In this lecture, we survey a variety of different sequencing technologies, including their
DNA Sequencing and Assembly CS 262 Lecture Notes, Winter 2016 February 2nd, 2016 Scribe: Mark Berger Abstract In this lecture, we survey a variety of different sequencing technologies, including their
Principles of Sequencing and Pla3orms
 Principles of Sequencing and Pla3orms 6/4/2018 RCPA Workshop Ms Leah Roberts PhD candidate University of Queensland TradiMonal diagnosmcs Standardised, established methods and infrastructure, reasonably
Principles of Sequencing and Pla3orms 6/4/2018 RCPA Workshop Ms Leah Roberts PhD candidate University of Queensland TradiMonal diagnosmcs Standardised, established methods and infrastructure, reasonably
