Next Generation Sequencing Technologies
|
|
|
- Asher Whitehead
- 6 years ago
- Views:
Transcription
1 Next Generation Sequencing Technologies
2 What is first generation? Sanger Sequencing
3
4 DNA Polymerase
5 Base-adding reaction +H +
6 Pros and Cons of Sanger Sequencing Polymerase errors average out Long sequences (~450 bp) Can only do 1 sequence at a time Need a lot of DNA to start with Expensive: 2 /base
7 To solve these cons what do we need? Cheaper Multiplex different samples Smaller starting amount How might you do this? What do you need to be able to do?
8 Design a Sequencer For 1 minute, write down all the things you would need to do to be able to sequence DNA in a multiplexed way. Turn to your neighbors (1-2 people) For 1 minute, discuss the things you would need to do to be able to sequence DNA in a multiplexed way. Be prepared to tell the class what you think you need and why
9 What you need to do multiplexed sequencing Ability to separate individual DNA pieces Ability to observe the sequence of each separated piece individually High sensitivity (as compared to Sanger sequencing)
10 What s different Sequence many sequences at once Technology is paired with DNA sequence agnostic primers Faster than SS Shorter than SS
11 How do we sequence things we don t know the sequence of?
12 Adapt sequences with known sequences This can be done with Sanger too, but need to PCR after this to get enough DNA Mardis, ER; Ann Rev Genom & Hum Gen
13 What you need to do multiplexed sequencing Ability to separate individual DNA pieces Ability to observe the sequence of each separated piece individually High sensitivity (as compared to Sanger sequencing)
14 Emulsion PCR onto beads (454, Ion Torrent)
15 Mardis, E. R. (2013). Next-Generation Sequencing Platforms. Annual Review of Analytical Chemistry, 6(1), doi: /annurevanchem Flow Cell: Bind directly to chip, make bridges (Illumina)
16 What you need to do multiplexed sequencing Ability to separate individual DNA pieces Ability to observe the sequence of each separated piece individually High sensitivity (as compared to Sanger sequencing)
17
18 454:2005 Imaging and light based
19 Illumina: 2006 Expose to all 4 bases Add 1 at a time, 3 OH is reversibly blocked Monitor fluorescence Mardis, E. R. (2013). Next-Generation Sequencing Platforms. Annual Review of Analytical Chemistry, 6(1), doi: / annurev-anchem
20 As of 2010, all were imaging based Why might this be problematic? How else might you follow sequencing?
21 Ion Torrent: 2010 On Chips Most accurate ph meter in the world
22 Ion Torrent Expose to single base type at a time Add as many as possible Monitor change in ph
23 Getting DNA onto beads P A P A P P A
24 Getting DNA onto beads P A
25 P Getting DNA onto beads A P P A P A Which strand do we keep when we make this singlestranded for sequencing? A
26 A P Keep sequence that is complementary to the sequence we read from sequencer A 3 5 P P A P A
27
28
29 Actual Sequence on Bead: GTAACTGTCAAACG What happens on Ion Torrent? Cycle through the following bases: T G A C C ATTGACAGTTTGC GTAACTGTCAAACG T G A C TT G A C T G A C T G A C TTT G A C
30 What does the cyclical process mean for our sequencing? On your sheet, figure out how far each of the sequences will get in 5 cycles
31 Histogram of read lengths
32 What is the sequence of the following DNA? TGAC TCTGGTGA
33 Bead with 2 DNAs T T C G G A A ATCTTAGGTA What happens? 2x as many bases as expected
34 Errors Homopolymers AAAAAAA Polymerase adds all at once System becomes saturated How many are there really of a particular base?
35 And Now for Something Completely Different Single Molecule Sequencing
36 Pacific Biosciences: Single Molecule Sequencing (SMRT) Benjamin A Flusberg, Dale R Webster, Jessica H Lee, Kevin J Travers, Eric C Olivares, Tyson A Clark, Jonas Korlach & Stephen W Turner Nature Methods 7, (2010) Published online: 9 May 2010, doi: /nmeth
37 Pacific Biosciences Can get VERY long sequences 5,000-8,000 bases, on average 30,000 bases sometimes 99.99% accurate for each base No averaging, so can find rare SNPs No amplification needed before sequencing, so less bias Differences in rates of addition allow one to measure epigenetic variations Fewer total sequences so generally end up with fewer total bases Much more expensive than the other techniques
38 Oxford Nanopore Technologies: In beta-testing Similar potential benefits as SMRT technology, but without drawbacks of polymerase and use of imaging technologies
39 Oxford Nanopore Technologies
40
41 Pros and Cons of NGS Fast Cheap (<1 /Mbase) Lots of data Fewer reads of each base are combined, so less accurate overall Short reads (getting longer, up to ~400 bases now)
42 Activity On table: fill in what you think are the pros and cons of each technology we discussed ~ 2 min Discuss with your neighbors what you each put, generate a consensus list to share with the class ~ 4 min
43 Questions?
Overview of Next Generation Sequencing technologies. Céline Keime
 Overview of Next Generation Sequencing technologies Céline Keime keime@igbmc.fr Next Generation Sequencing < Second generation sequencing < General principle < Sequencing by synthesis - Illumina < Sequencing
Overview of Next Generation Sequencing technologies Céline Keime keime@igbmc.fr Next Generation Sequencing < Second generation sequencing < General principle < Sequencing by synthesis - Illumina < Sequencing
Aaron Liston, Oregon State University Botany 2012 Intro to Next Generation Sequencing Workshop
 Output (bp) Aaron Liston, Oregon State University Growth in Next-Gen Sequencing Capacity 3.5E+11 2002 2004 2006 2008 2010 3.0E+11 2.5E+11 2.0E+11 1.5E+11 1.0E+11 Adapted from Mardis, 2011, Nature 5.0E+10
Output (bp) Aaron Liston, Oregon State University Growth in Next-Gen Sequencing Capacity 3.5E+11 2002 2004 2006 2008 2010 3.0E+11 2.5E+11 2.0E+11 1.5E+11 1.0E+11 Adapted from Mardis, 2011, Nature 5.0E+10
Next Generation Sequencing Lecture Saarbrücken, 19. March Sequencing Platforms
 Next Generation Sequencing Lecture Saarbrücken, 19. March 2012 Sequencing Platforms Contents Introduction Sequencing Workflow Platforms Roche 454 ABI SOLiD Illumina Genome Anlayzer / HiSeq Problems Quality
Next Generation Sequencing Lecture Saarbrücken, 19. March 2012 Sequencing Platforms Contents Introduction Sequencing Workflow Platforms Roche 454 ABI SOLiD Illumina Genome Anlayzer / HiSeq Problems Quality
Next Generation Sequencing. Simon Rasmussen Assistant Professor Center for Biological Sequence analysis Technical University of Denmark
 Next eneration Sequencing Simon Rasmussen Assistant Professor enter for Biological Sequence analysis Technical University of Denmark DNA Sequencing DNA sequencing Reading the order of bases in DNA fragments
Next eneration Sequencing Simon Rasmussen Assistant Professor enter for Biological Sequence analysis Technical University of Denmark DNA Sequencing DNA sequencing Reading the order of bases in DNA fragments
Next generation sequencing techniques" Toma Tebaldi Centre for Integrative Biology University of Trento
 Next generation sequencing techniques" Toma Tebaldi Centre for Integrative Biology University of Trento Mattarello September 28, 2009 Sequencing Fundamental task in modern biology read the information
Next generation sequencing techniques" Toma Tebaldi Centre for Integrative Biology University of Trento Mattarello September 28, 2009 Sequencing Fundamental task in modern biology read the information
Introduction to Next Generation Sequencing (NGS)
 Introduction to Next eneration Sequencing (NS) Simon Rasmussen Assistant Professor enter for Biological Sequence analysis Technical University of Denmark 2012 Today 9.00-9.45: Introduction to NS, How it
Introduction to Next eneration Sequencing (NS) Simon Rasmussen Assistant Professor enter for Biological Sequence analysis Technical University of Denmark 2012 Today 9.00-9.45: Introduction to NS, How it
Sequencing technologies. Jose Blanca COMAV institute bioinf.comav.upv.es
 Sequencing technologies Jose Blanca COMAV institute bioinf.comav.upv.es Outline Sequencing technologies: Sanger 2nd generation sequencing: 3er generation sequencing: 454 Illumina SOLiD Ion Torrent PacBio
Sequencing technologies Jose Blanca COMAV institute bioinf.comav.upv.es Outline Sequencing technologies: Sanger 2nd generation sequencing: 3er generation sequencing: 454 Illumina SOLiD Ion Torrent PacBio
Sequencing technologies. Jose Blanca COMAV institute bioinf.comav.upv.es
 Sequencing technologies Jose Blanca COMAV institute bioinf.comav.upv.es Outline Sequencing technologies: Sanger 2nd generation sequencing: 3er generation sequencing: 454 Illumina SOLiD Ion Torrent PacBio
Sequencing technologies Jose Blanca COMAV institute bioinf.comav.upv.es Outline Sequencing technologies: Sanger 2nd generation sequencing: 3er generation sequencing: 454 Illumina SOLiD Ion Torrent PacBio
Sequencing technologies. Jose Blanca COMAV institute bioinf.comav.upv.es
 Sequencing technologies Jose Blanca COMAV institute bioinf.comav.upv.es Outline Sequencing technologies: Sanger 2nd generation sequencing: 3er generation sequencing: 454 Illumina SOLiD Ion Torrent PacBio
Sequencing technologies Jose Blanca COMAV institute bioinf.comav.upv.es Outline Sequencing technologies: Sanger 2nd generation sequencing: 3er generation sequencing: 454 Illumina SOLiD Ion Torrent PacBio
High Throughput Sequencing Technologies. J Fass UCD Genome Center Bioinformatics Core Monday June 16, 2014
 High Throughput Sequencing Technologies J Fass UCD Genome Center Bioinformatics Core Monday June 16, 2014 Sequencing Explosion www.genome.gov/sequencingcosts http://t.co/ka5cvghdqo Sequencing Explosion
High Throughput Sequencing Technologies J Fass UCD Genome Center Bioinformatics Core Monday June 16, 2014 Sequencing Explosion www.genome.gov/sequencingcosts http://t.co/ka5cvghdqo Sequencing Explosion
Functional Genomics Research Stream. Research Meetings: November 2 & 3, 2009 Next Generation Sequencing
 Functional Genomics Research Stream Research Meetings: November 2 & 3, 2009 Next Generation Sequencing Current Issues Research Meetings: Meet with me this Thursday or Friday. (bring laboratory notebook
Functional Genomics Research Stream Research Meetings: November 2 & 3, 2009 Next Generation Sequencing Current Issues Research Meetings: Meet with me this Thursday or Friday. (bring laboratory notebook
Ultrasequencing: methods and applications of the new generation sequencing platforms
 Ultrasequencing: methods and applications of the new generation sequencing platforms Nuria Tubío Santamaría Course: Genomics Universitat Autònoma de Barcelona 1 Introduction Clasical methods of sequencing:
Ultrasequencing: methods and applications of the new generation sequencing platforms Nuria Tubío Santamaría Course: Genomics Universitat Autònoma de Barcelona 1 Introduction Clasical methods of sequencing:
Sequencing techniques
 Sequencing techniques Workshop on Whole Genome Sequencing and Analysis, 2-4 Oct. 2017 Learning objective: After this lecture, you should be able to account for different techniques for whole genome sequencing
Sequencing techniques Workshop on Whole Genome Sequencing and Analysis, 2-4 Oct. 2017 Learning objective: After this lecture, you should be able to account for different techniques for whole genome sequencing
DNA-Sequencing. Technologies & Devices. Matthias Platzer. Genome Analysis Leibniz Institute on Aging - Fritz Lipmann Institute (FLI)
 DNA-Sequencing Technologies & Devices Matthias Platzer Genome Analysis Leibniz Institute on Aging - Fritz Lipmann Institute (FLI) Genome analysis DNA sequencing platforms ABI 3730xl 4/2004 & 6/2006 1 Mb/day,
DNA-Sequencing Technologies & Devices Matthias Platzer Genome Analysis Leibniz Institute on Aging - Fritz Lipmann Institute (FLI) Genome analysis DNA sequencing platforms ABI 3730xl 4/2004 & 6/2006 1 Mb/day,
DNA-Sequencing. Technologies & Devices. Matthias Platzer. Genome Analysis Leibniz Institute on Aging - Fritz Lipmann Institute (FLI)
 DNA-Sequencing Technologies & Devices Matthias Platzer Genome Analysis Leibniz Institute on Aging - Fritz Lipmann Institute (FLI) Genome analysis DNA sequencing platforms ABI 3730xl 4/2004 & 6/2006 1 Mb/day,
DNA-Sequencing Technologies & Devices Matthias Platzer Genome Analysis Leibniz Institute on Aging - Fritz Lipmann Institute (FLI) Genome analysis DNA sequencing platforms ABI 3730xl 4/2004 & 6/2006 1 Mb/day,
Genome Sequencing. I: Methods. MMG 835, SPRING 2016 Eukaryotic Molecular Genetics. George I. Mias
 Genome Sequencing I: Methods MMG 835, SPRING 2016 Eukaryotic Molecular Genetics George I. Mias Department of Biochemistry and Molecular Biology gmias@msu.edu Sequencing Methods Cost of Sequencing Wetterstrand
Genome Sequencing I: Methods MMG 835, SPRING 2016 Eukaryotic Molecular Genetics George I. Mias Department of Biochemistry and Molecular Biology gmias@msu.edu Sequencing Methods Cost of Sequencing Wetterstrand
Next Gen Sequencing. Expansion of sequencing technology. Contents
 Next Gen Sequencing Contents 1 Expansion of sequencing technology 2 The Next Generation of Sequencing: High-Throughput Technologies 3 High Throughput Sequencing Applied to Genome Sequencing (TEDed CC BY-NC-ND
Next Gen Sequencing Contents 1 Expansion of sequencing technology 2 The Next Generation of Sequencing: High-Throughput Technologies 3 High Throughput Sequencing Applied to Genome Sequencing (TEDed CC BY-NC-ND
2 nd Genera-on ( NextGen ) Sequencing Technologies
 2 nd Genera-on ( NextGen ) Sequencing Technologies Jay Shendure Read Length is Not As Important For Resequencing % of Paired K-mers with Uniquely Assignable Location 100% 90% 80% 70% 60% 50% 40% 30% 20%
2 nd Genera-on ( NextGen ) Sequencing Technologies Jay Shendure Read Length is Not As Important For Resequencing % of Paired K-mers with Uniquely Assignable Location 100% 90% 80% 70% 60% 50% 40% 30% 20%
Human genome sequence
 NGS: the basics Human genome sequence June 26th 2000: official announcement of the completion of the draft of the human genome sequence (truly finished in 2004) Francis Collins Craig Venter HGP: 3 billion
NGS: the basics Human genome sequence June 26th 2000: official announcement of the completion of the draft of the human genome sequence (truly finished in 2004) Francis Collins Craig Venter HGP: 3 billion
Next Generation Sequencing. Jeroen Van Houdt - Leuven 13/10/2017
 Next Generation Sequencing Jeroen Van Houdt - Leuven 13/10/2017 Landmarks in DNA sequencing 1953 Discovery of DNA double helix structure 1977 A Maxam and W Gilbert "DNA seq by chemical degradation" F Sanger"DNA
Next Generation Sequencing Jeroen Van Houdt - Leuven 13/10/2017 Landmarks in DNA sequencing 1953 Discovery of DNA double helix structure 1977 A Maxam and W Gilbert "DNA seq by chemical degradation" F Sanger"DNA
Sequencing technologies
 Sequencing technologies part of High-Throughput Analyzes of Genome Sequenzes Computational EvoDevo University of Leipzig Leipzig, WS 2014/15 Sanger Sequencing (Chain Termination Method) Sequencing of one
Sequencing technologies part of High-Throughput Analyzes of Genome Sequenzes Computational EvoDevo University of Leipzig Leipzig, WS 2014/15 Sanger Sequencing (Chain Termination Method) Sequencing of one
Introductie en Toepassingen van Next-Generation Sequencing in de Klinische Virologie. Sander van Boheemen Medical Microbiology
 Introductie en Toepassingen van Next-Generation Sequencing in de Klinische Virologie Sander van Boheemen Medical Microbiology Next-generation sequencing Next-generation sequencing (NGS), also known as
Introductie en Toepassingen van Next-Generation Sequencing in de Klinische Virologie Sander van Boheemen Medical Microbiology Next-generation sequencing Next-generation sequencing (NGS), also known as
High Throughput Sequencing Technologies. J Fass UCD Genome Center Bioinformatics Core Monday September 15, 2014
 High Throughput Sequencing Technologies J Fass UCD Genome Center Bioinformatics Core Monday September 15, 2014 Sequencing Explosion www.genome.gov/sequencingcosts http://t.co/ka5cvghdqo Sequencing Explosion
High Throughput Sequencing Technologies J Fass UCD Genome Center Bioinformatics Core Monday September 15, 2014 Sequencing Explosion www.genome.gov/sequencingcosts http://t.co/ka5cvghdqo Sequencing Explosion
DNA Sequencing by Ion Torrent. Marc Lavergne CHEM 4590
 DNA Sequencing by Ion Torrent Marc Lavergne CHEM 4590 OVERVIEW History DNA Synthesis and First-Gen Sequencing Technology Sequencing Signal Detection Advantages/Disadvantages Applications Current Research
DNA Sequencing by Ion Torrent Marc Lavergne CHEM 4590 OVERVIEW History DNA Synthesis and First-Gen Sequencing Technology Sequencing Signal Detection Advantages/Disadvantages Applications Current Research
Welcome to the NGS webinar series
 Welcome to the NGS webinar series Webinar 1 NGS: Introduction to technology, and applications NGS Technology Webinar 2 Targeted NGS for Cancer Research NGS in cancer Webinar 3 NGS: Data analysis for genetic
Welcome to the NGS webinar series Webinar 1 NGS: Introduction to technology, and applications NGS Technology Webinar 2 Targeted NGS for Cancer Research NGS in cancer Webinar 3 NGS: Data analysis for genetic
Next Generation Sequencing. Josef K Vogt Slides by: Simon Rasmussen
 Next eneration Sequencing Josef K Vogt Slides by: Simon Rasmussen 2017 Second generation sequencing 454 Illumina 10 6-10 9 90% market share SOLiD Ion Torrent (PM) Library preparation 1.reate library molecules
Next eneration Sequencing Josef K Vogt Slides by: Simon Rasmussen 2017 Second generation sequencing 454 Illumina 10 6-10 9 90% market share SOLiD Ion Torrent (PM) Library preparation 1.reate library molecules
HLA-Typing Strategies
 HLA-Typing Strategies Cologne, 13.5.2017 Joannis Mytilineos MD, PhD Department of Transplantation Immunology Institute for Clinical Transfusion Medicine and Immunogenetics German Red Cross Blood Transfusion
HLA-Typing Strategies Cologne, 13.5.2017 Joannis Mytilineos MD, PhD Department of Transplantation Immunology Institute for Clinical Transfusion Medicine and Immunogenetics German Red Cross Blood Transfusion
Outline General NGS background and terms 11/14/2016 CONFLICT OF INTEREST. HLA region targeted enrichment. NGS library preparation methodologies
 Eric T. Weimer, PhD, D(ABMLI) Assistant Professor, Pathology & Laboratory Medicine, UNC School of Medicine Director, Molecular Immunology Associate Director, Clinical Flow Cytometry, HLA, and Immunology
Eric T. Weimer, PhD, D(ABMLI) Assistant Professor, Pathology & Laboratory Medicine, UNC School of Medicine Director, Molecular Immunology Associate Director, Clinical Flow Cytometry, HLA, and Immunology
Principles of Sequencing and Pla3orms
 Principles of Sequencing and Pla3orms 6/4/2018 RCPA Workshop Ms Leah Roberts PhD candidate University of Queensland TradiMonal diagnosmcs Standardised, established methods and infrastructure, reasonably
Principles of Sequencing and Pla3orms 6/4/2018 RCPA Workshop Ms Leah Roberts PhD candidate University of Queensland TradiMonal diagnosmcs Standardised, established methods and infrastructure, reasonably
DNA-Sequencing. Technologies & Devices
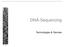 DNA-Sequencing Technologies & Devices Genome analysis DNA sequencing platforms ABI 3730xl 4/2004 & 6/2006 1 Mb/day, 850 nt reads 2 Mb/day, 550 nt reads Roche/454 GS FLX 12/2006 800 Mb/23h, 800 nt reads
DNA-Sequencing Technologies & Devices Genome analysis DNA sequencing platforms ABI 3730xl 4/2004 & 6/2006 1 Mb/day, 850 nt reads 2 Mb/day, 550 nt reads Roche/454 GS FLX 12/2006 800 Mb/23h, 800 nt reads
Research school methods seminar Genomics and Transcriptomics
 Research school methods seminar Genomics and Transcriptomics Stephan Klee 19.11.2014 2 3 4 5 Genetics, Genomics what are we talking about? Genetics and Genomics Study of genes Role of genes in inheritence
Research school methods seminar Genomics and Transcriptomics Stephan Klee 19.11.2014 2 3 4 5 Genetics, Genomics what are we talking about? Genetics and Genomics Study of genes Role of genes in inheritence
Next-generation sequencing Technology Overview
 Next-generation sequencing Technology Overview UQ Winter School 2018 Christopher Noune, PhD AGRF Melbourne christopher.noune@agrf.org.au What is NGS? Ion Torrent PGM (Thermo-Fisher) MiSeq (Illumina) High-Throughput
Next-generation sequencing Technology Overview UQ Winter School 2018 Christopher Noune, PhD AGRF Melbourne christopher.noune@agrf.org.au What is NGS? Ion Torrent PGM (Thermo-Fisher) MiSeq (Illumina) High-Throughput
A Crash Course in NGS for GI Pathologists. Sandra O Toole
 A Crash Course in NGS for GI Pathologists Sandra O Toole The Sanger Technique First generation sequencing Uses dideoxynucleotides (dideoxyadenine, dideoxyguanine, etc) These are molecules that resemble
A Crash Course in NGS for GI Pathologists Sandra O Toole The Sanger Technique First generation sequencing Uses dideoxynucleotides (dideoxyadenine, dideoxyguanine, etc) These are molecules that resemble
Third Generation Sequencing
 Third Generation Sequencing By Mohammad Hasan Samiee Aref Medical Genetics Laboratory of Dr. Zeinali History of DNA sequencing 1953 : Discovery of DNA structure by Watson and Crick 1973 : First sequence
Third Generation Sequencing By Mohammad Hasan Samiee Aref Medical Genetics Laboratory of Dr. Zeinali History of DNA sequencing 1953 : Discovery of DNA structure by Watson and Crick 1973 : First sequence
DNA-Sequencing. Technologies & Devices
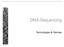 DNA-Sequencing Technologies & Devices Genome analysis DNA sequencing platforms ABI 3730xl 4/2004 & 6/2006 1 Mb/day, 850 nt reads 2 Mb/day, 550 nt reads Roche/454 GS FLX 12/2006 800 Mb/23h, 800 nt reads
DNA-Sequencing Technologies & Devices Genome analysis DNA sequencing platforms ABI 3730xl 4/2004 & 6/2006 1 Mb/day, 850 nt reads 2 Mb/day, 550 nt reads Roche/454 GS FLX 12/2006 800 Mb/23h, 800 nt reads
The Journey of DNA Sequencing. Chromosomes. What is a genome? Genome size. H. Sunny Sun
 The Journey of DNA Sequencing H. Sunny Sun What is a genome? Genome is the total genetic complement of a living organism. The nuclear genome comprises approximately 3.2 * 10 9 nucleotides of DNA, divided
The Journey of DNA Sequencing H. Sunny Sun What is a genome? Genome is the total genetic complement of a living organism. The nuclear genome comprises approximately 3.2 * 10 9 nucleotides of DNA, divided
Phenotype analysis: biological-biochemical analysis. Genotype analysis: molecular and physical analysis
 1 Genetic Analysis Phenotype analysis: biological-biochemical analysis Behaviour under specific environmental conditions Behaviour of specific genetic configurations Behaviour of progeny in crosses - Genotype
1 Genetic Analysis Phenotype analysis: biological-biochemical analysis Behaviour under specific environmental conditions Behaviour of specific genetic configurations Behaviour of progeny in crosses - Genotype
NEXT GENERATION SEQUENCING. Farhat Habib
 NEXT GENERATION SEQUENCING HISTORY HISTORY Sanger Dominant for last ~30 years 1000bp longest read Based on primers so not good for repetitive or SNPs sites HISTORY Sanger Dominant for last ~30 years 1000bp
NEXT GENERATION SEQUENCING HISTORY HISTORY Sanger Dominant for last ~30 years 1000bp longest read Based on primers so not good for repetitive or SNPs sites HISTORY Sanger Dominant for last ~30 years 1000bp
Genome 373: High- Throughput DNA Sequencing. Doug Fowler
 Genome 373: High- Throughput DNA Sequencing Doug Fowler Tasks give ML unity We learned about three tasks that are commonly encountered in ML Models/Algorithms Give ML Diversity Classification Regression
Genome 373: High- Throughput DNA Sequencing Doug Fowler Tasks give ML unity We learned about three tasks that are commonly encountered in ML Models/Algorithms Give ML Diversity Classification Regression
Biochemistry 412. New Strategies, Technologies, & Applications For DNA Sequencing. 12 February 2008
 Biochemistry 412 New Strategies, Technologies, & Applications For DNA Sequencing 12 February 2008 Note: Scale is wrong!! (at least for sequences) 10 6 In 1980, the sequencing cost per finished bp $1.00
Biochemistry 412 New Strategies, Technologies, & Applications For DNA Sequencing 12 February 2008 Note: Scale is wrong!! (at least for sequences) 10 6 In 1980, the sequencing cost per finished bp $1.00
Introduction to NGS. Josef K Vogt Slides by: Simon Rasmussen Next Generation Sequencing Analysis
 Introduction to NGS Josef K Vogt Slides by: Simon Rasmussen 2017 Life science data deluge Massive unstructured data from several areas DNA, patient journals, proteomics, imaging,... Impacts Industry, Environment,
Introduction to NGS Josef K Vogt Slides by: Simon Rasmussen 2017 Life science data deluge Massive unstructured data from several areas DNA, patient journals, proteomics, imaging,... Impacts Industry, Environment,
Phenotype analysis: biological-biochemical analysis. Genotype analysis: molecular and physical analysis
 1 Genetic Analysis Phenotype analysis: biological-biochemical analysis Behaviour under specific environmental conditions Behaviour of specific genetic configurations Behaviour of progeny in crosses - Genotype
1 Genetic Analysis Phenotype analysis: biological-biochemical analysis Behaviour under specific environmental conditions Behaviour of specific genetic configurations Behaviour of progeny in crosses - Genotype
Deep Sequencing technologies
 Deep Sequencing technologies Gabriela Salinas 30 October 2017 Transcriptome and Genome Analysis Laboratory http://www.uni-bc.gwdg.de/index.php?id=709 Microarray and Deep-Sequencing Core Facility University
Deep Sequencing technologies Gabriela Salinas 30 October 2017 Transcriptome and Genome Analysis Laboratory http://www.uni-bc.gwdg.de/index.php?id=709 Microarray and Deep-Sequencing Core Facility University
Chapter 7. DNA Microarrays
 Bioinformatics III Structural Bioinformatics and Genome Analysis Chapter 7. DNA Microarrays 7.9 Next Generation Sequencing 454 Sequencing Solexa Illumina Solid TM System Sequencing Process of determining
Bioinformatics III Structural Bioinformatics and Genome Analysis Chapter 7. DNA Microarrays 7.9 Next Generation Sequencing 454 Sequencing Solexa Illumina Solid TM System Sequencing Process of determining
13-2 Manipulating DNA Slide 1 of 32
 1 of 32 The Tools of Molecular Biology The Tools of Molecular Biology How do scientists make changes to DNA? Scientists use their knowledge of the structure of DNA and its chemical properties to study
1 of 32 The Tools of Molecular Biology The Tools of Molecular Biology How do scientists make changes to DNA? Scientists use their knowledge of the structure of DNA and its chemical properties to study
1. Introduction Gene regulation Genomics and genome analyses
 1. Introduction Gene regulation Genomics and genome analyses 2. Gene regulation tools and methods Regulatory sequences and motif discovery TF binding sites Databases 3. Technologies Microarrays Deep sequencing
1. Introduction Gene regulation Genomics and genome analyses 2. Gene regulation tools and methods Regulatory sequences and motif discovery TF binding sites Databases 3. Technologies Microarrays Deep sequencing
Data Basics. Josef K Vogt Slides by: Simon Rasmussen Next Generation Sequencing Analysis
 Data Basics Josef K Vogt Slides by: Simon Rasmussen 2017 Generalized NGS analysis Sample prep & Sequencing Data size Main data reductive steps SNPs, genes, regions Application Assembly: Compare Raw Pre-
Data Basics Josef K Vogt Slides by: Simon Rasmussen 2017 Generalized NGS analysis Sample prep & Sequencing Data size Main data reductive steps SNPs, genes, regions Application Assembly: Compare Raw Pre-
Next Generation Sequencing (NGS)
 Next Generation Sequencing (NGS) Fernando Alvarez Sección Biomatemática, Facultad de Ciencias, UdelaR 1 Uruguay Montevide o 3 TANGO World Champ 1930 1950 (Maraca 4 Next Generation Sequencing module Next
Next Generation Sequencing (NGS) Fernando Alvarez Sección Biomatemática, Facultad de Ciencias, UdelaR 1 Uruguay Montevide o 3 TANGO World Champ 1930 1950 (Maraca 4 Next Generation Sequencing module Next
NGS technologies: a user s guide. Karim Gharbi & Mark Blaxter
 NGS technologies: a user s guide Karim Gharbi & Mark Blaxter genepool-manager@ed.ac.uk Natural history of sequencing 2 Brief history of sequencing 100s bp throughput 100 Gb 1977 1986 1995 1999 2005 2007
NGS technologies: a user s guide Karim Gharbi & Mark Blaxter genepool-manager@ed.ac.uk Natural history of sequencing 2 Brief history of sequencing 100s bp throughput 100 Gb 1977 1986 1995 1999 2005 2007
DNA-Sequenzierung. Technologien & Geräte
 DNA-Sequenzierung Technologien & Geräte Genome analysis DNA sequencing platforms ABI 3730xl 4/2004 & 6/2006 1 Mb/day, 850 nt reads 2 Mb/day, 550 nt reads Roche/454 GS FLX 12/2006 400 Mb/7h, 350 nt reads
DNA-Sequenzierung Technologien & Geräte Genome analysis DNA sequencing platforms ABI 3730xl 4/2004 & 6/2006 1 Mb/day, 850 nt reads 2 Mb/day, 550 nt reads Roche/454 GS FLX 12/2006 400 Mb/7h, 350 nt reads
Ultrasequencing: Methods and Applications of the New Generation Sequencing Platforms
 Ultrasequencing: Methods and Applications of the New Generation Sequencing Platforms Laura Moya Andérico Master in Advanced Genetics Genomics Class December 16 th, 2015 Brief Overview First-generation
Ultrasequencing: Methods and Applications of the New Generation Sequencing Platforms Laura Moya Andérico Master in Advanced Genetics Genomics Class December 16 th, 2015 Brief Overview First-generation
Human Genome Sequencing Over the Decades The capacity to sequence all 3.2 billion bases of the human genome (at 30X coverage) has increased
 Human Genome Sequencing Over the Decades The capacity to sequence all 3.2 billion bases of the human genome (at 30X coverage) has increased exponentially since the 1990s. In 2005, with the introduction
Human Genome Sequencing Over the Decades The capacity to sequence all 3.2 billion bases of the human genome (at 30X coverage) has increased exponentially since the 1990s. In 2005, with the introduction
CM581A2: NEXT GENERATION SEQUENCING PLATFORMS AND LIBRARY GENERATION
 CM581A2: NEXT GENERATION SEQUENCING PLATFORMS AND LIBRARY GENERATION Fall 2015 Instructors: Coordinator: Carol Wilusz, Associate Professor MIP, CMB Instructor: Dan Sloan, Assistant Professor, Biology,
CM581A2: NEXT GENERATION SEQUENCING PLATFORMS AND LIBRARY GENERATION Fall 2015 Instructors: Coordinator: Carol Wilusz, Associate Professor MIP, CMB Instructor: Dan Sloan, Assistant Professor, Biology,
NGS technologies approaches, applications and challenges!
 www.supagro.fr NGS technologies approaches, applications and challenges! Jean-François Martin Centre de Biologie pour la Gestion des Populations Centre international d études supérieures en sciences agronomiques
www.supagro.fr NGS technologies approaches, applications and challenges! Jean-François Martin Centre de Biologie pour la Gestion des Populations Centre international d études supérieures en sciences agronomiques
DNA Sequencing and Assembly
 DNA Sequencing and Assembly CS 262 Lecture Notes, Winter 2016 February 2nd, 2016 Scribe: Mark Berger Abstract In this lecture, we survey a variety of different sequencing technologies, including their
DNA Sequencing and Assembly CS 262 Lecture Notes, Winter 2016 February 2nd, 2016 Scribe: Mark Berger Abstract In this lecture, we survey a variety of different sequencing technologies, including their
High Throughput Sequencing Technologies. J Fass UCD Genome Center Bioinformatics Core Tuesday December 16, 2014
 High Throughput Sequencing Technologies J Fass UCD Genome Center Bioinformatics Core Tuesday December 16, 2014 Sequencing Explosion www.genome.gov/sequencingcosts http://t.co/ka5cvghdqo Sequencing Explosion
High Throughput Sequencing Technologies J Fass UCD Genome Center Bioinformatics Core Tuesday December 16, 2014 Sequencing Explosion www.genome.gov/sequencingcosts http://t.co/ka5cvghdqo Sequencing Explosion
Outline. General principles of clonal sequencing Analysis principles Applications CNV analysis Genome architecture
 The use of new sequencing technologies for genome analysis Chris Mattocks National Genetics Reference Laboratory (Wessex) NGRL (Wessex) 2008 Outline General principles of clonal sequencing Analysis principles
The use of new sequencing technologies for genome analysis Chris Mattocks National Genetics Reference Laboratory (Wessex) NGRL (Wessex) 2008 Outline General principles of clonal sequencing Analysis principles
Next-Generation Sequencing. Technologies
 Next-Generation Next-Generation Sequencing Technologies Sequencing Technologies Nicholas E. Navin, Ph.D. MD Anderson Cancer Center Dept. Genetics Dept. Bioinformatics Introduction to Bioinformatics GS011062
Next-Generation Next-Generation Sequencing Technologies Sequencing Technologies Nicholas E. Navin, Ph.D. MD Anderson Cancer Center Dept. Genetics Dept. Bioinformatics Introduction to Bioinformatics GS011062
Introduction to NGS. Simon Rasmussen Associate Professor DTU Bioinformatics Technical University of Denmark 2018
 Introduction to NGS Simon Rasmussen Associate Professor DTU Bioinformatics Technical University of Denmark 2018 Life science data deluge Massive unstructured data from several areas DNA, patient journals,
Introduction to NGS Simon Rasmussen Associate Professor DTU Bioinformatics Technical University of Denmark 2018 Life science data deluge Massive unstructured data from several areas DNA, patient journals,
3.1.4 DNA Microarray Technology
 3.1.4 DNA Microarray Technology Scientists have discovered that one of the differences between healthy and cancer is which genes are turned on in each. Scientists can compare the gene expression patterns
3.1.4 DNA Microarray Technology Scientists have discovered that one of the differences between healthy and cancer is which genes are turned on in each. Scientists can compare the gene expression patterns
Targeted Sequencing Using Droplet-Based Microfluidics. Keith Brown Director, Sales
 Targeted Sequencing Using Droplet-Based Microfluidics Keith Brown Director, Sales brownk@raindancetech.com Who we are: is a Provider of Microdroplet-based Solutions The Company s RainStorm TM Technology
Targeted Sequencing Using Droplet-Based Microfluidics Keith Brown Director, Sales brownk@raindancetech.com Who we are: is a Provider of Microdroplet-based Solutions The Company s RainStorm TM Technology
BIOINFORMATICS 1 SEQUENCING TECHNOLOGY. DNA story. DNA story. Sequencing: infancy. Sequencing: beginnings 26/10/16. bioinformatic challenges
 BIOINFORMATICS 1 or why biologists need computers SEQUENCING TECHNOLOGY bioinformatic challenges http://www.bioinformatics.uni-muenster.de/teaching/courses-2012/bioinf1/index.hbi Prof. Dr. Wojciech Makałowski"
BIOINFORMATICS 1 or why biologists need computers SEQUENCING TECHNOLOGY bioinformatic challenges http://www.bioinformatics.uni-muenster.de/teaching/courses-2012/bioinf1/index.hbi Prof. Dr. Wojciech Makałowski"
The Genome Analysis Centre. Building Excellence in Genomics and Computa5onal Bioscience
 Building Excellence in Genomics and Computa5onal Bioscience Resequencing approaches Sarah Ayling Crop Genomics and Diversity sarah.ayling@tgac.ac.uk Why re- sequence plants? To iden
Building Excellence in Genomics and Computa5onal Bioscience Resequencing approaches Sarah Ayling Crop Genomics and Diversity sarah.ayling@tgac.ac.uk Why re- sequence plants? To iden
NEXT-GENERATION SEQUENCING AND BIOINFORMATICS
 NEXT-GENERATION SEQUENCING AND BIOINFORMATICS Moore's law: the number of transistors in a dense integrated circuit doubles every two years Moore's law calculates and predicts the pace of improvement of
NEXT-GENERATION SEQUENCING AND BIOINFORMATICS Moore's law: the number of transistors in a dense integrated circuit doubles every two years Moore's law calculates and predicts the pace of improvement of
Understanding Accuracy in SMRT Sequencing
 Understanding Accuracy in SMRT Sequencing Jonas Korlach, Chief Scientific Officer, Pacific Biosciences Introduction Single Molecule, Real-Time (SMRT ) DNA sequencing achieves highly accurate sequencing
Understanding Accuracy in SMRT Sequencing Jonas Korlach, Chief Scientific Officer, Pacific Biosciences Introduction Single Molecule, Real-Time (SMRT ) DNA sequencing achieves highly accurate sequencing
FUTURE PROSPECTS IN MOLECULAR INFECTIOUS DISEASES DIAGNOSIS
 FUTURE PROSPECTS IN MOLECULAR INFECTIOUS DISEASES DIAGNOSIS Richard L. Hodinka, Ph.D. University of South Carolina School of Medicine Greenville Greenville Health System, Greenville, SC hodinka@greenvillemed.sc.edu
FUTURE PROSPECTS IN MOLECULAR INFECTIOUS DISEASES DIAGNOSIS Richard L. Hodinka, Ph.D. University of South Carolina School of Medicine Greenville Greenville Health System, Greenville, SC hodinka@greenvillemed.sc.edu
Concepts and methods in sequencing and genome assembly
 BCM-2002 Concepts and methods in sequencing and genome assembly http://megasun.bch.umontreal.ca/papers/bcm-2002/sequencing-bcm2002-nov2015.pdf B. Franz LANG, Département de Biochimie Bureau: H307-15 Courrier
BCM-2002 Concepts and methods in sequencing and genome assembly http://megasun.bch.umontreal.ca/papers/bcm-2002/sequencing-bcm2002-nov2015.pdf B. Franz LANG, Département de Biochimie Bureau: H307-15 Courrier
Bio(tech) Interlude. 3 Nobel Prizes: PCR: Kary Mullis, 1993 Electrophoresis: A.W.K. Tiselius, 1948 DNA Sequencing: Frederick Sanger, 1980
 Bio(tech) Interlude 3 Nobel Prizes: PCR: Kary Mullis, 1993 Electrophoresis: A.W.K. Tiselius, 1948 DNA Sequencing: Frederick Sanger, 1980 PCR 1: 25ºC G 2: 95ºC A 3: 60ºC T 5 3 A A G 3 G T C 5 T T T 6: 72ºC
Bio(tech) Interlude 3 Nobel Prizes: PCR: Kary Mullis, 1993 Electrophoresis: A.W.K. Tiselius, 1948 DNA Sequencing: Frederick Sanger, 1980 PCR 1: 25ºC G 2: 95ºC A 3: 60ºC T 5 3 A A G 3 G T C 5 T T T 6: 72ºC
Genome Resequencing. Rearrangements. SNPs, Indels CNVs. De novo genome Sequencing. Metagenomics. Exome Sequencing. RNA-seq Gene Expression
 Genome Resequencing De novo genome Sequencing SNPs, Indels CNVs Rearrangements Metagenomics RNA-seq Gene Expression Splice Isoform Abundance High Throughput Short Read Sequencing: Illumina Exome Sequencing
Genome Resequencing De novo genome Sequencing SNPs, Indels CNVs Rearrangements Metagenomics RNA-seq Gene Expression Splice Isoform Abundance High Throughput Short Read Sequencing: Illumina Exome Sequencing
Get to Know Your DNA. Every Single Fragment.
 HaloPlex HS NGS Target Enrichment System Get to Know Your DNA. Every Single Fragment. High sensitivity detection of rare variants using molecular barcodes How Does Molecular Barcoding Work? HaloPlex HS
HaloPlex HS NGS Target Enrichment System Get to Know Your DNA. Every Single Fragment. High sensitivity detection of rare variants using molecular barcodes How Does Molecular Barcoding Work? HaloPlex HS
NEXT GENERATION SEQUENCING Whole Gene Sequencing
 NEXT GENERATION SEQUENCING Whole Gene Sequencing Ingrid Faé Educational Session 3: Next generation sequencing Stockholm, Friday, June 27 th 2014 Department for Blood Group Serology and Transfusion Medicine
NEXT GENERATION SEQUENCING Whole Gene Sequencing Ingrid Faé Educational Session 3: Next generation sequencing Stockholm, Friday, June 27 th 2014 Department for Blood Group Serology and Transfusion Medicine
Factors affecting PCR
 Lec. 11 Dr. Ahmed K. Ali Factors affecting PCR The sequences of the primers are critical to the success of the experiment, as are the precise temperatures used in the heating and cooling stages of the
Lec. 11 Dr. Ahmed K. Ali Factors affecting PCR The sequences of the primers are critical to the success of the experiment, as are the precise temperatures used in the heating and cooling stages of the
Student Exercise on Polymerase Chain Reaction (PCR) Prepared by the Office of Biotechnology, Iowa State University. Part I. Part III.
 Student Exercise on Polymerase Chain Reaction (PCR) Prepared by the Office of Biotechnology, Iowa State University Part I. Segment of interest (02) Original-2 5' 3' 1. The purpose of PCR is to make copies
Student Exercise on Polymerase Chain Reaction (PCR) Prepared by the Office of Biotechnology, Iowa State University Part I. Segment of interest (02) Original-2 5' 3' 1. The purpose of PCR is to make copies
Human Genomics. 1 P a g e
 Human Genomics What were the aims of the human genome project? To identify all the approximately 20,000-25,000 genes in Human DNA. To find where each gene is located To determine the sequences of the 3
Human Genomics What were the aims of the human genome project? To identify all the approximately 20,000-25,000 genes in Human DNA. To find where each gene is located To determine the sequences of the 3
Contact us for more information and a quotation
 GenePool Information Sheet #1 Installed Sequencing Technologies in the GenePool The GenePool offers sequencing service on three platforms: Sanger (dideoxy) sequencing on ABI 3730 instruments Illumina SOLEXA
GenePool Information Sheet #1 Installed Sequencing Technologies in the GenePool The GenePool offers sequencing service on three platforms: Sanger (dideoxy) sequencing on ABI 3730 instruments Illumina SOLEXA
Illumina (Solexa) Throughput: 4 Tbp in one run (5 days) Cheapest sequencing technology. Mismatch errors dominate. Cost: ~$1000 per human genme
 Illumina (Solexa) Current market leader Based on sequencing by synthesis Current read length 100-150bp Paired-end easy, longer matepairs harder Error ~0.1% Mismatch errors dominate Throughput: 4 Tbp in
Illumina (Solexa) Current market leader Based on sequencing by synthesis Current read length 100-150bp Paired-end easy, longer matepairs harder Error ~0.1% Mismatch errors dominate Throughput: 4 Tbp in
BST227 Introduction to Statistical Genetics. Lecture 8: Variant calling from high-throughput sequencing data
 BST227 Introduction to Statistical Genetics Lecture 8: Variant calling from high-throughput sequencing data 1 PC recap typical genome Differs from the reference genome at 4-5 million sites ~85% SNPs ~15%
BST227 Introduction to Statistical Genetics Lecture 8: Variant calling from high-throughput sequencing data 1 PC recap typical genome Differs from the reference genome at 4-5 million sites ~85% SNPs ~15%
Overview of techniques in Molecular Diagnostics ELKE BOONE 23 MAART 2017
 Overview of techniques in Molecular Diagnostics ELKE BOONE 23 MAART 2017 Molecular Diagnostics Molecular Diagnostics? Molecular Diagnostics FISH or CISH techniques FISH or CISH techniques FISH or CISH
Overview of techniques in Molecular Diagnostics ELKE BOONE 23 MAART 2017 Molecular Diagnostics Molecular Diagnostics? Molecular Diagnostics FISH or CISH techniques FISH or CISH techniques FISH or CISH
Matthew Tinning Australian Genome Research Facility. July 2012
 Next-Generation Sequencing: an overview of technologies and applications Matthew Tinning Australian Genome Research Facility July 2012 History of Sequencing Where have we been? 1869 Discovery of DNA 1909
Next-Generation Sequencing: an overview of technologies and applications Matthew Tinning Australian Genome Research Facility July 2012 History of Sequencing Where have we been? 1869 Discovery of DNA 1909
Paper Primer-Matching Activity to Reinforce the Concept of PCR (Polymerase Chain Reaction)
 Paper Primer-Matching Activity to Reinforce the Concept of PCR (Polymerase Chain Reaction) This is a hands-on paper primer matching activity that confirms that students understand the role of primers in
Paper Primer-Matching Activity to Reinforce the Concept of PCR (Polymerase Chain Reaction) This is a hands-on paper primer matching activity that confirms that students understand the role of primers in
15:30-16:15 Sean Prosser New Developments for Natural History Collection Barcoding. DNA Barcoding Natural History Collections
 15:30-16:15 Sean Prosser New Developments for Natural History Collection Barcoding DNA Barcoding Natural History Collections Recap Barcoding Museum Specimens Age Target Amplicons Final Sequence Length
15:30-16:15 Sean Prosser New Developments for Natural History Collection Barcoding DNA Barcoding Natural History Collections Recap Barcoding Museum Specimens Age Target Amplicons Final Sequence Length
QIAseq SPE technology for Illumina : Redefining amplicon sequencing
 Application Note QIAseq SPE technology for Illumina : Redefining amplicon sequencing Amplicon-based enrichment and sequencing takes advantage of PCR workflows to turn amplicons that represent regions of
Application Note QIAseq SPE technology for Illumina : Redefining amplicon sequencing Amplicon-based enrichment and sequencing takes advantage of PCR workflows to turn amplicons that represent regions of
II. Integrative Genomics interactions between molecules and genes
 . Structural Genomics Structure of Genome I. Functional Genomics Expression of Genome a. Transcriptomics b. Proteomics II. Integrative Genomics interactions between molecules and genes Fields of Genomics
. Structural Genomics Structure of Genome I. Functional Genomics Expression of Genome a. Transcriptomics b. Proteomics II. Integrative Genomics interactions between molecules and genes Fields of Genomics
I. Structure of Genome Structural genomics II. Expression of Genome Functional genomics. a. Transcriptomics b. Proteomics
 I. Structure of Genome Structural genomics II. Expression of Genome Functional genomics a. Transcriptomics b. Proteomics Fields of Genomics Structural genomics Functional genomics Transcriptomics Proteomics
I. Structure of Genome Structural genomics II. Expression of Genome Functional genomics a. Transcriptomics b. Proteomics Fields of Genomics Structural genomics Functional genomics Transcriptomics Proteomics
NextGen Sequencing Technologies Sequencing overview
 Outline Conventional NextGen High-throughput sequencing (Next-Gen sequencing) technologies. Illumina sequencing in detail. Quality control. Sequence coverage. Multiplexing. FASTQ files. Shendure and Ji
Outline Conventional NextGen High-throughput sequencing (Next-Gen sequencing) technologies. Illumina sequencing in detail. Quality control. Sequence coverage. Multiplexing. FASTQ files. Shendure and Ji
High Throughput Sequencing Technologies. UCD Genome Center Bioinformatics Core Monday 15 June 2015
 High Throughput Sequencing Technologies UCD Genome Center Bioinformatics Core Monday 15 June 2015 Sequencing Explosion www.genome.gov/sequencingcosts http://t.co/ka5cvghdqo Sequencing Explosion 2011 PacBio
High Throughput Sequencing Technologies UCD Genome Center Bioinformatics Core Monday 15 June 2015 Sequencing Explosion www.genome.gov/sequencingcosts http://t.co/ka5cvghdqo Sequencing Explosion 2011 PacBio
The New Genome Analyzer IIx Delivering more data, faster, and easier than ever before. Jeremy Preston, PhD Marketing Manager, Sequencing
 The New Genome Analyzer IIx Delivering more data, faster, and easier than ever before Jeremy Preston, PhD Marketing Manager, Sequencing Illumina Genome Analyzer: a Paradigm Shift 2000x gain in efficiency
The New Genome Analyzer IIx Delivering more data, faster, and easier than ever before Jeremy Preston, PhD Marketing Manager, Sequencing Illumina Genome Analyzer: a Paradigm Shift 2000x gain in efficiency
High throughput DNA Sequencing. An Equal Opportunity University!
 High throughput DNA Sequencing An Equal Opportunity University! irst Generation DNA sequencing utilize chain terminator technologies (adaptation of Sanger sequencing) Adapt fluorescence chemistry, high-resolution
High throughput DNA Sequencing An Equal Opportunity University! irst Generation DNA sequencing utilize chain terminator technologies (adaptation of Sanger sequencing) Adapt fluorescence chemistry, high-resolution
Rapid Transcriptome Characterization for a nonmodel organism using 454 pyrosequencing
 Rapid Transcriptome Characterization for a nonmodel organism using 454 pyrosequencing "#$%&'()*+,"(-*."#$%&/.,"*01*0.,(%-*.&0("2*01*3,$,45,"-*4#66&*71** 3"#)(82,"-*2&9:)($*)1*"(03&"2-*#)66(*.(8$6#*;
Rapid Transcriptome Characterization for a nonmodel organism using 454 pyrosequencing "#$%&'()*+,"(-*."#$%&/.,"*01*0.,(%-*.&0("2*01*3,$,45,"-*4#66&*71** 3"#)(82,"-*2&9:)($*)1*"(03&"2-*#)66(*.(8$6#*;
Sequencing Theory. Brett E. Pickett, Ph.D. J. Craig Venter Institute
 Sequencing Theory Brett E. Pickett, Ph.D. J. Craig Venter Institute Applications of Genomics and Bioinformatics to Infectious Diseases GABRIEL Network Agenda Sequencing Instruments Sanger Illumina Ion
Sequencing Theory Brett E. Pickett, Ph.D. J. Craig Venter Institute Applications of Genomics and Bioinformatics to Infectious Diseases GABRIEL Network Agenda Sequencing Instruments Sanger Illumina Ion
Wheat CAP Gene Expression with RNA-Seq
 Wheat CAP Gene Expression with RNA-Seq July 9 th -13 th, 2018 Overview of the workshop, Alina Akhunova http://www.ksre.k-state.edu/igenomics/workshops/ RNA-Seq Workshop Activities Lectures Laboratory Molecular
Wheat CAP Gene Expression with RNA-Seq July 9 th -13 th, 2018 Overview of the workshop, Alina Akhunova http://www.ksre.k-state.edu/igenomics/workshops/ RNA-Seq Workshop Activities Lectures Laboratory Molecular
CHEM 4420 Exam I Spring 2013 Page 1 of 6
 CHEM 4420 Exam I Spring 2013 Page 1 of 6 Name Use complete sentences when requested. There are 100 possible points on this exam. The multiple choice questions are worth 2 points each. All other questions
CHEM 4420 Exam I Spring 2013 Page 1 of 6 Name Use complete sentences when requested. There are 100 possible points on this exam. The multiple choice questions are worth 2 points each. All other questions
Next-generation sequencing technologies
 Next-generation sequencing technologies NGS applications Illumina sequencing workflow Overview Sequencing by ligation Short-read NGS Sequencing by synthesis Illumina NGS Single-molecule approach Long-read
Next-generation sequencing technologies NGS applications Illumina sequencing workflow Overview Sequencing by ligation Short-read NGS Sequencing by synthesis Illumina NGS Single-molecule approach Long-read
AKESOgen, Inc, 3155 Northwoods Place, NW, Norcross, GA 30071, USA Tel: +1 (770) Page 1
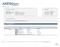 Page 1 HLA Sequencing Based Typing Workflow: A- Sample Collection: iswab-discovery (Mawi DNA Technologies) - Sample was transported by standard US postal services mail, no cold chain of any sort was involved
Page 1 HLA Sequencing Based Typing Workflow: A- Sample Collection: iswab-discovery (Mawi DNA Technologies) - Sample was transported by standard US postal services mail, no cold chain of any sort was involved
TREE CODE PRODUCT BROCHURE
 TREE CODE PRODUCT BROCHURE Single Molecule, Real-Time (SMRT) Sequencing technology offers: Long read sequencing ~10 Gb with 20 kb average read lengths for WGS ~20 Gb with 40 kb average read length for
TREE CODE PRODUCT BROCHURE Single Molecule, Real-Time (SMRT) Sequencing technology offers: Long read sequencing ~10 Gb with 20 kb average read lengths for WGS ~20 Gb with 40 kb average read length for
MHC Region. MHC expression: Class I: All nucleated cells and platelets Class II: Antigen presenting cells
 DNA based HLA typing methods By: Yadollah Shakiba, MD, PhD MHC Region MHC expression: Class I: All nucleated cells and platelets Class II: Antigen presenting cells Nomenclature of HLA Alleles Assigned
DNA based HLA typing methods By: Yadollah Shakiba, MD, PhD MHC Region MHC expression: Class I: All nucleated cells and platelets Class II: Antigen presenting cells Nomenclature of HLA Alleles Assigned
qpcr Quantitative PCR or Real-time PCR Gives a measurement of PCR product at end of each cycle real time
 qpcr qpcr Quantitative PCR or Real-time PCR Gives a measurement of PCR product at end of each cycle real time Differs from endpoint PCR gel on last cycle Used to determines relative amount of template
qpcr qpcr Quantitative PCR or Real-time PCR Gives a measurement of PCR product at end of each cycle real time Differs from endpoint PCR gel on last cycle Used to determines relative amount of template
Next- gen sequencing. STAMPS 2015 Hilary G. Morrison Joe Vineis, Nora Downey, Be>e Hecox- Lea, Kim Finnegan
 Next- gen sequencing STAMPS 2015 Hilary G. Morrison Joe Vineis, Nora Downey, Be>e Hecox- Lea, Kim Finnegan QuesIons What is the difference between standard and next- gen sequencing? How is next- gen sequencing
Next- gen sequencing STAMPS 2015 Hilary G. Morrison Joe Vineis, Nora Downey, Be>e Hecox- Lea, Kim Finnegan QuesIons What is the difference between standard and next- gen sequencing? How is next- gen sequencing
Next Generation Sequencing. Tobias Österlund
 Next Generation Sequencing Tobias Österlund tobiaso@chalmers.se NGS part of the course Week 4 Friday 13/2 15.15-17.00 NGS lecture 1: Introduction to NGS, alignment, assembly Week 6 Thursday 26/2 08.00-09.45
Next Generation Sequencing Tobias Österlund tobiaso@chalmers.se NGS part of the course Week 4 Friday 13/2 15.15-17.00 NGS lecture 1: Introduction to NGS, alignment, assembly Week 6 Thursday 26/2 08.00-09.45
Incorporating Molecular ID Technology. Accel-NGS 2S MID Indexing Kits
 Incorporating Molecular ID Technology Accel-NGS 2S MID Indexing Kits Molecular Identifiers (MIDs) MIDs are indices used to label unique library molecules MIDs can assess duplicate molecules in sequencing
Incorporating Molecular ID Technology Accel-NGS 2S MID Indexing Kits Molecular Identifiers (MIDs) MIDs are indices used to label unique library molecules MIDs can assess duplicate molecules in sequencing
