NGS: Digital RNAseq & Library Prep Seminar. Next-Generation Sequencing Lunch & Learn
|
|
|
- Christal Boyd
- 6 years ago
- Views:
Transcription
1 NGS: Digital RNAseq & Library Prep Seminar Next-Generation Sequencing Lunch & Learn Samuel Rulli, Ph. D Global Product Manager QIAseq Targeted RNA Panels 1
2 Targeted sequencing with UMIs QIAseq mirnaseq Kit UMIs Gel free library prep Complete mirnome Dec mirna DNA Biomarkers QIAseq Targeted DNA Panels Unique molecular index Mutation /SNP analysis CNV Insertions / deletions 2ng Fresh DNA mrna/ lncrna QIAseq Targeted RNA Panel UMIs Gene expression 100pg / 10 cells All kits utilize SPE (single primer extension) Fusion QIAseq Targeted RNAscan Panels Unique molecular index Known fusion genes (validation) Unknown partners (discovery) 2
3 QIAseq Library Products for Illumina Sequencers Nucleic acid extraction NGS Library preparation Sequencing & QIAGEN Bioinformatics Whole Genome Sequencing (WGS) Hybrid Capture Sequencign QIAseq FX DNA Library Kit FX fragmentation step eliminates shearing 2.5 hour workflow from gdna to library Tunable enzymatic fragmentation for any size probes with higher genomic complexity than Nextera DNA Amplicon/ PCR sequencing QIAseq 1-Step Amplicon Library Kit 1-tube, room temperature library prep cfdna for Liquid Biopsy or NIPT QIAseq cfdna Complete Solution QIAamp Circulating Nucleic Acid Isolation Optimized protocol specific for cfdna Low Input ChIP-seq, cfdna, LCM QIAseq Ultralow Input Library Kit Ultra-efficient chemistry for pg+ DNA Barcoded adapters for 96-plex sequencing Single Cell Genomics QIAseq Single Cell FX Lib Kits PCR-free MDA Amplification Sorted single cells or 6pg DNA input 3
4 QIAseq Library Products for Illumina Sequencers Nucleic acid extraction NGS Library preparation Sequencing & QIAGEN Bioinformatics Whole Genome Sequencing (WGS) Hybrid Capture Sequencign QIAseq FX DNA Library Kit FX fragmentation step eliminates shearing 2.5 hour workflow from gdna to library Tunable enzymatic fragmentation for any size probes with higher genomic complexity than Nextera DNA Amplicon/ PCR sequencing QIAseq 1-Step Amplicon Library Kit 1-tube, room temperature library prep cfdna for Liquid Biopsy or NIPT QIAseq cfdna Complete Solution QIAamp Circulating Nucleic Acid Isolation Optimized protocol specific for cfdna Low Input ChIP-seq, cfdna, LCM QIAseq Ultralow Input Library Kit Ultra-efficient chemistry for pg+ DNA Barcoded adapters for 96-plex sequencing Single Cell Genomics QIAseq Single Cell FX Lib Kits PCR-free MDA Amplification Sorted single cells or 6pg DNA input 4
5 Today s Agenda Targeted Sequencing 1. Principles of unique molecular indexes (UMIs) 2. Single primer extension (SPE) versus PCR for library construction 3. UMI and SPE in action gene expression analysis 4. DNA variant analysis and novel gene fusions discovery with UMIs & SPE 5. Single Cell Capture & NGS applications with QIAscout and QIAseq Single Cell FX kits 5
6 Section 1: Principal of unique molecular Index (UMIs) PCR duplicates and amplification bias are major issues in current RNAseq workflows, as they result in biased and inaccurate gene expression profiles Gene A Sample 1 Ratio of original state of genes 4 Ratio of genes based on reads [reads (ratio)] 12 (2) Gene A Sample (1) mrna cdna Raw reads 6
7 Section 1: Principal of unique molecular Index (UMIs) Targeted RNAseq still is a Read based approach to understanding gene expression. How do we go from Reads to counting transcripts? 7
8 Section 1: Principal of unique molecular Index (UMIs) Targeted RNAseq still is a Read based approach to understanding gene expression. How do we go from Reads to counting transcripts? 8
9 Section 1: Principal of unique molecular Index (UMIs) Molecular indexes allow the counting of original transcript levels instead of PCR duplicates, thereby enabling digital sequencing and resulting in unbiased and accurate gene expression profiles. Tag each gene with UMIs Count unique molecules, not reads Gene A Sample 1 Ratio of original state of genes 4 Ratio of genes based on UMIs 4 Ratio of genes based on reads [reads (ratio)] 12 (2) Gene A Sample (1) mrna cdna UMI reads 9
10 Section 1: Principal of unique molecular Index (UMIs) Each capture event is archived with a UMI 12 random bases 16.7 million indexes Strategy for gene expression uses UMI-gene specific primer Strategy for DNA variant and fusion gene is slightly different, but same principle 10
11 Today s Agenda Targeted Sequencing 1. Principles of unique molecular indexes (UMIs) 2. Single primer extension (SPE) versus PCR for library construction 3. UMI and SPE in action gene expression analysis 4. DNA variant analysis and novel gene fusions discovery with UMIs & SPE 5. Summary/Questions? 11
12 Section 2: Single Primer Extension Single Primer Extension 5 3 DNA or cdna 2 Primer Amplification PCR amplification PCR amplification captures information
13 Inefficient sequencing of GC-rich regions by PCR amplification Leads to lack of comprehensive coverage in those regions Normal GC content High GC content PCR Amplification using suboptimal chemistry under uniform conditions Coverage Regions with normal GC content are covered efficiently Regions with High GC content are not covered efficiently 13
14 Complete sequencing of GC-rich regions by SPE Leads to lack of comprehensive coverage in those regions Normal GC content High GC content PCR Amplification following SPE under uniform conditions Coverage Regions with normal GC content are covered efficiently Regions with High GC content are covered as well 14
15 SPE coverage of GC-rich regions CEBPA GC content Coverage CCND1 GC content Coverage 15
16 Section 2: Single Primer Extension Advantages of SPE: 1.) Only need 1 region for primer design unlocks entire transcriptome, genome and fusion genes - 50% primers lowers cost & allows for greater content during multiplexing 2.) Able to adopt to G/C rich and hard to PCR regions - sequence everything 3.) Uniform reaction - uniform library construction uniform sequencing better results 4.) strategy works very well on FFPE, fragmented and low quality samples Disadvantages: 1.) Extra step in library construction - may add 1 hour to total workflow 16
17 Today s Agenda Targeted Sequencing 1. Principles of unique molecular indexes (UMIs) 2. Single primer extension (SPE) versus PCR for library construction 3. UMI and SPE in action gene expression analysis 4. DNA variant analysis and novel gene fusions discovery with UMIs & SPE 5. Summary/Questions? 17
18 Section 3: UMI and SPE in Action Gene expression example Small Molecules Signal Transduction Application cells Experiment is to identify novel compounds that modulate known signal transduction pathways treated cells RNA 18
19 Section 3: UMI and SPE in Action Gene expression example Small Molecules Signal Transduction Application cells treated cells Cells are treated with different chemical inhibitors. Isolate RNA from cells Build library with QIAseq Targeted RNA Panels Human Signal Transduction 421 targets/ 10ng total RNA RNA 19
20 Section 3: UMI and SPE in Action Gene expression example 6 hours GSP1, GSP2 never see other, thereby minimizing primer dimers 20
21 Section 3: UMI and SPE in Action Gene expression example leave no scientist behind. 21
22 Section 3: UMI and SPE in Action Gene expression example Included in Panel Kit Library Quant Index Kit Kit Included in cloud 22
23 Section 3: UMI and SPE in Action Gene expression example Included in Panel Kit Library Quant Index Kit Kit CLC Biomedical workbench with MT plugin 23
24 Section 3: UMI and SPE in Action Gene expression example Requirement Species coverage Biological replicates Short reads for FFPE, and Exosomal RNA QIAseq Targeted RNA Panels Human Catalog, Extended, Virtual, Custom panels Mouse + Rat Custom Essential for robustness of experimental design (and statistics!) Average amplicon 97 bps ; range bases Depth of sequencing High enough to infer accurate statistics determined by UMI- ~2-5 reads per UMI is enough. Stranded library prep Not required, amplicons do not overlap lncrna Type of reads (paired or Unpaired?) mrna and lncrnas Not necessary, 150 base single reads more than enough for accurate data QIAseq was designed against database containing lncrna and mrna. Assay are specific for lncrna or mrna. Currently 54,881 genes from Ensembl version 81 24
25 Section 3: UMI and SPE in Action Gene expression example Free circulating nucleic acids RNA and DNA from dead cells shed into the bloodstream, can contain cancer-related mutations. Exosomes Tiny microvesicles found in body fluids that transport RNA between cells. Circulating tumor cells Tumor cells shed from a tumor into the bloodstream carrying genetic information. Tissue samples Fresh tissue or archived FFPE samples Qiagen comprehensive sample isolation portfolio compatible with QIAseq RNA Compatible with 100pg (10 cells) to 25ng RNA 25
26 Section 3: UMI and SPE in Action Gene expression example 26
27 Section 3: UMI and SPE in Action Gene expression example Flexible experiment design for any Catalog Panel options: Comprehensive Panels (available for 12, 96 or 384 samples) Cancer Transcriptome (395) Inflammation & Immunity Transcriptome (475) Signal Transduction PathwayFinder (406) Stem Cell & Differentiation Markers (293) Molecular Toxicology Transcriptome (370) Angiogenesis & Endothelial Cell Biology (340) Apoptosis & Cell Death (264) ECM & Adhesion Molecules (421) 27
28 Section 3: UMI and SPE in Action Gene expression example Flexible experiment design for any Extended Panels Add 25 of your favorite targets (mrnas or lncrnas) to QIAGEN s comprehensive panel lnc13 GAS5 MEG3 PTCSC3 TERC ADAMT S9 GAS6- AS1 MIR31 HG CAHM GNAS MIR7-3HG ZFAS1 DLEU2 LINC PTCSC1 NEAT NAMA DLX6 GACA T1 LINC What is the role of tumor suppressor lncrnas? Find out! 28
29 Section 3: UMI and SPE in Action Gene expression example Flexible experiment design for any Catalog Panel options: QIAseq Targeted RNA Virtual Panels (available for 12, 96 or 384 samples) Each Panel contains 84 genes + controls and house keeping genes. Choose from over 180 panels! Pathways Diseases mirna Targets 29
30 QIAGEN Automating Sample disruption Purification RNA Quality Control qpcr setup NGS library quantifiaction Real time PCR + HRM High-throughput PCR Fragment Analysis Bench top assay setup Mediumthroughput Pyrosequencing Hybrid capture Low-throughput Integrated assay setup GeneReader NGS 30
31 QIAGEN Automating Purification RNA Quality Control Cells in 96 well plates RNA Integrity (96 samples done automatically while at lunch) RNA isolation of 96 samples RNA quantification (16 samples/ 90 secs) 31
32 QIAGEN Automating Purification RNA Quality Control Assay setup Detection and analysis Cells in 96 well plates RNA Integrity (96 samples done automatically while at lunch) Library quantification Library Integrity RNA isolation of 96 samples Library quantification Setup by qpcr RNA quantification (16 samples/ 90 secs) 32
33 QIAseq Targeted RNA Application Data Small Molecules Signal Transduction Application cells HEK293T Cells were treated with 90 different chemical inhibitors. The 421 Signal Transduction Gene QIAseq Panel was interrogated. treated cells RNA In one day we went from total RNA to sequence ready libraries for 96 samples. The final libraries were quantified, normalized, and pooled. Prior to loading onto a NextSeq, the denatured libraries were diluted to the appropriate input concentration to obtain to generate suitable clusters on the NextSeq. Indexed libraries The parameters of the NextSeq sequencing run were; single 151 bp read, with a Custom Sequencing Primer (included in kit). Normalized, pooled libraries 33
34 Primary Data Analysis for QIAseq Targeted RNA Sequencing QIAseq Targeted RNA Data Analysis automated workflow Read Mapping Primer Trimming UMI Count Read Mapping Identify the possible position of the read within the reference genome Align the read sequence to reference sequences Primer Trimming Remove the primer sequences from the reads Go get coffee UMI Counting 34
35 Small Molecule Application Data Primary Data Analysis for QIAseq Targeted RNA Sequencing 35
36 Small Molecule Application Data Primary Data Analysis for QIAseq Targeted RNA Sequencing QIAseq RNA Quantification - Read Details: Unique Captures per Target Gene Count Differential gene expression inter- and intra-samples 36
37 QIAseq secondary data analysis setup Analysis: What kinds of things get flagged? Low tag #, high gdna, poor normalizer performance 37
38 Secondary Data Analysis for QIAseq Targeted RNA Sequencing Scatter plot and clustergram (HDAC Sample compared to Control) 38
39 Secondary Data Analysis for QIAseq Targeted RNA Sequencing Changes in gene expression due to chemical perturbation and quantified by QIAseq RNA NGS were characterized. 39
40 Ingenuity IPA analysis HDAC Mechanistic Network in HEK293T Cells Treated with Trichostatin A HDAC is predicted to be inhibited by Trichostatin A and drives a mechanistic network with 18 other regulators. Cell cycle NHR, proliferation Transcriptional activator 40
41 Where can you run this? QIAseq sample multiplexing guidelines on NGS platforms 41
42 Unparalleled efficiency and flexibility vs PCR An example: 96 samples, 421 genes Parameter QIAseq targeted RNA panels RT-PCR Material required One pool of primers well plates Run time 14 hours for NextSeq run 310 hours (2 hours per plate) Hands-on time 3 hours (for 96 samples) 105 hours (one hour per plate) Sample 10 ng each sample 4000 ng each sample 42
43 Today s Agenda Targeted Sequencing 1. Principles of unique molecular indexes (UMIs) 2. Single primer extension (SPE) versus PCR for library construction 3. UMI and SPE in action gene expression analysis 4. DNA variant analysis and novel gene fusions discovery with UMIs & SPE 5. Summary/Questions? 43
44 Section 4: DNA variant analysis and novel gene fusions discovery QIAseq mirnaseq Kit UMIs Gel free library prep Complete mirnome Dec mirna DNA Biomarkers QIAseq Targeted DNA Panels Unique molecular index Mutation /SNP analysis CNV Insertions / deletions 2ng Fresh DNA mrna/ lncrna QIAseq Targeted RNA Panel UMIs Gene expression 100pg / 10 cells Fusion QIAseq Targeted RNAscan Panels Unique molecular index Known fusion genes (validation) Unknown partners (discovery) 44
45 Section 4: DNA variant analysis and novel gene fusions discovery DNA QIAseq Targeted DNA Panels Unique molecular index Mutation /SNP analysis CNV Insertions / deletions 2ng Fresh DNA mirna Biomarkers mrna/ lncrna Fusion 45
46 Section 4: DNA variant analysis and novel gene fusions discovery PCR and sequencing errors (artifacts) limit variant calling accuracy Traditional targeted DNA sequencing EGFR exon 21 * A mutation is seen in 1 out of 5 reads that map to EGFR exon 21. Cannot accurately tell whether the mutation is: 1. A PCR or sequencing error (artifact) (False positive), or 2. A true low-frequency mutations Variant calling based on non-unique reads does not reflect the mutational status of original DNA molecules Applies to a wide range of panels 46
47 Section 4: DNA variant analysis and novel gene fusions discovery Count and analyze single original molecules (not total reads) = digital sequencing Traditional targeted DNA sequencing EGFR exon 21 * A mutation is seen in 1 out of 5 reads that map to EGFR exon 21. Cannot accurately tell whether the mutation is: 1. A PCR or sequencing error (artifact) (False positive), or 2. A true low-frequency mutations Digital sequencing with UMIs * * * False variant is present in some fragments carrying the same UMI Traditional Targeted DNA Seq True variant is present in all fragments carrying the same UMI 47
48 Section 4: DNA variant analysis and novel gene fusions discovery Count and analyze single original molecules (not total reads) = digital sequencing Traditional targeted DNA sequencing EGFR exon 21 * A mutation is seen in 1 out of 5 reads that map to EGFR exon 21. Cannot accurately tell whether the mutation is: 1. A PCR or sequencing error (artifact) (False positive), or 2. A true low-frequency mutations Add UMIs before any amplification Digital sequencing with UMIs UMI * * False variant is present in some fragments carrying the same UMI True variant is present in all fragments carrying the same UMI 48
49 Section 4: DNA variant analysis and novel gene fusions discovery QIAseq Targeted DNA Panel Workflow 49
50 QIAseq targeted DNA panels List of panels Panel Variant (Cat) number Number of genes Number of primers Type of coverage Breast cancer panel DHS-001Z Colorectal cancer panel DHS-002Z Myeloid Neoplasms panel DHS-003Z Lung cancer panel DHS-005Z Actionable solid tumor panel DHS-101Z BRCA1 and BRCA2 panel DHS-102Z BRCA1 and BRCA2 Plus panel DHS-103Z Pharmacogenomics panel DHS-104Z Mitochondria panel DHS-105Z Chromosome M Inherited diseases panel DHS-3011Z Comprehensive cancer panel DHS-3501Z Types of coverage: 1. Exonic regions of genes plus 10 bases to cover intron/exon junctions 2. Mix of type of coverage 1 (for tumor suppressor genes) and HotSpots for Oncogenes 3. SNPs 4. Full chromosome 50
51 QIAseq targeted DNA panels List of panels Panel Variant (Cat) number Panel size (bases) Specificity (reads with primers) (%) Uniformity (0.2x mean basemt) (%) Breast cancer panel DHS-001Z 370, Colorectal cancer panel DHS-002Z 215, Myeloid Neoplasms panel DHS-003Z 436, Lung cancer panel DHS-005Z 318, Actionable solid tumor panel DHS-101Z 15, BRCA1 and BRCA2 panel DHS-102Z 16, BRCA1 and BRCA2 Plus panel DHS-103Z 25, Pharmacogenomics panel DHS-104Z 3, Mitochondria panel DHS-105Z 16, Inherited diseases panel DHS-3011Z 838, Comprehensive cancer panel DHS-3501Z 836, Uniformity and Specificity are defined based on NA12878 tests 51
52 Section 4: DNA variant analysis and novel gene fusions discovery DNA mirna Biomarkers mrna/ lncrna Fusion QIAseq Targeted RNAscan Panels Unique molecular index Known fusion genes (validation) Unknown partners (discovery) 52
53 Section 4: DNA variant analysis and novel gene fusions discovery QIASeq Targeted RNAscan is a RNA target enrichment method that allows verification of known fusions and discovery of novel fusions with next-generation sequencing (NGS). RPS6KB1-VMP1 ARFGEF2-SULF2
54 Section 4: DNA variant analysis and novel gene fusions discovery 54
55 QIAseq targeted RNAscan panels List of panels Panel Variant (Cat) number Number of primers Type of coverage Hematology panel FHS-001Z 156 Breakpoint Solid Tumor panel FHS-002Z 101 Breakpoint Lung cancer panel FHS-003Z 137 Breakpoint Oncology panel FHS-3001Z 950 Breakpoint
56 Summary: Biomarkers come in many flavors. Gene Expression Indels mirna expression Biomarkers Mutations Fusions Copy number variants
57 Summary: NGS can be used for all of them NGS Gene Expression Indels mirna expression Biomarkers Mutations Fusions Copy number variants 57
58 QIAseq solutions to detect all Biomarkers using NGS Gene Expression QIAseq targeted RNA panels QIAseq targeted DNA panels Indels mirna expression QIAseq mirna sequencing system Biomarkers QIAseq targeted DNA panels Mutations Copy number variants Fusions QIAseq targeted RNAscan panels QIAseq targeted DNA panels 58
59 QIAseq Library Products for Illumina Sequencers Nucleic acid extraction NGS Library preparation Sequencing & QIAGEN Bioinformatics Whole Genome Sequencing (WGS) Hybrid Capture Sequencign QIAseq FX DNA Library Kit FX fragmentation step eliminates shearing 2.5 hour workflow from gdna to library Tunable enzymatic fragmentation for any size probes with higher genomic complexity than Nextera DNA Amplicon/ PCR sequencing QIAseq 1-Step Amplicon Library Kit 1-tube, room temperature library prep cfdna for Liquid Biopsy or NIPT QIAseq cfdna Complete Solution QIAamp Circulating Nucleic Acid Isolation Optimized protocol specific for cfdna Low Input ChIP-seq, cfdna, LCM QIAseq Ultralow Input Library Kit Ultra-efficient chemistry for pg+ DNA Barcoded adapters for 96-plex sequencing Single Cell Genomics QIAseq Single Cell FX Lib Kits) PCR-free MDA Amplification Sorted single cells or 6pg DNA input 59
60 QIAscout Release Device: Interface with Microscope Objective on revolving turret 4 adapters cover a diameter range from mm Objective magnifications include 4x, 5x, and 10x Dimensions based on popular objectives from 4 major players Release device controlled by button on controller box 60
61 Interface with microscope
62 QIAscout raft Can be used as standard cell culture dish Microrafts are magnetic, dislodgeable and self-sealing Each array contains 12,000 microrafts Cells settle according to Poisson distribution 6000 cells means a 30% chance of having a singe cell in a microraft 62
63 Cells can be imaged directly on the array by brightfield or fluorescence CellTracker Green Hoechst 63
64 Numbered microrafts make it easy to isolate cells Numbered microrafts on an unused Array 2. Array with seeded cells in medium 3. Microraft dislodged with needle 3. 64
65 Introducing QIAscout 65
66 Qiagen s Single Cell Portfolio: The cutting edge of genomics Highest accuracy Superior coverage High diversity, high quality libraries Streamlined workflows Data analyis & visualization Biological interpretation WGA or WTA Library preparation Sequencing & QIAGEN Bioinformatics Single-cell whole genome sequencing REPLI-g Single Cell FX DNA Library Kit Solution for variant calling, ideal for applications where SNV & CNV is equally important Single-cell gene expression analysis by RNAseq REPLI-g Single Cell FX RNA Library Kit Streamlined workflow with reliable detection of even low-abundance transcripts Single-cell targeted sequencing REPLI-g Single Cell DNA or WTA Kit QIAseq Targeted DNA\RNA Panels High quality targeted sequencing from a single cell for DNA variants or RNA expression 66
67 QIAGEN s REPLI-g Technology: use MDA, not PCR! MDA: Multiple Displacement Amplification Alkaline denaturation Hexamer random primers Phi29 polymerase strand displacement (30 C) Displaced strand becomes a template for replication Gentle denaturation results in intact DNA-template Even priming events covering the whole genome immediate amplification across all regions Modified high-fidelity Phi29 polymerase (SensiPhi) o High enzyme processivity molecular weight product ( kb) o Proofreading activity 1000-fold higher accuracy than normal PCR-enzyme Multiple displacement events efficient amplification of DNA 67
68 MDA amplifies through gdna secondary structure regions Denatured gdna has a complex secondary structure Consists of regions of ssdna and dsdna that can form complicated hairpins and loops QIAGEN s MDA enzyme handles complex DNA structures generating extremely long amplicons 68
69
70 QIAseq Library Products for Illumina Sequencers Products for Illumina Sequencers: Nucleic acid extraction NGS Library preparation Sequencing & QIAGEN Bioinformatics Whole Genome Sequencing (WGS) Hybrid Capture Sequencign QIAseq FX DNA Library Kit FX fragmentation step eliminates shearing 2.5 hour workflow from gdna to library Tunable enzymatic fragmentation for any size probes with higher genomic complexity than Nextera DNA Amplicon/ PCR sequencing QIAseq 1-Step Amplicon Library Kit 1-tube, room temperature library prep cfdna for Liquid Biopsy or NIPT QIAseq cfdna Complete Solution QIAamp Circulating Nucleic Acid Isolation Optimized protocol specific for cfdna Low Input ChIP-seq, cfdna, LCM QIAseq Ultralow Input Library Kit Ultra-efficient chemistry for pg+ DNA Barcoded adapters for 96-plex sequencing Single Cell Genomics (QIAseq Single Cell FX Lib Kits) PCR-free MDA Amplification Sorted single cells or 6pg DNA input 70
71 QIAseq FX: Gold-Standard WGS libraries in just 2.5 hours Purified gdna 1ng 1µg 1000bp QIAseq FX 2.5 hour workflow: - gdna Fragmentation - Barcoded Adapter Ligation - Library Amplification 250bp 450bp Library QC Hybrid Capture (Optional) Target Enrichment (IDT xgen, Agilent SureSelect) Illumina HiSeq Compatible with Hybrid Capture downstream target enrichment Flexible to generate any size sample DNA fragment 71
72 QIAseq FX the ideal balance of speed & high data quality Illumina Truseq Nano Fragmentase + NEB Ultra Illumina Nextera QIASeq FX Total /Hands-on 5 hr/ 1hr 3.5 hr/45 min 1.5 hr/ 15min 2.5 hr /20min Pros Cons Good sequencing performance Relies on Covaris; Long Protocol Fully Enzymatic Fast Workflow Library Complexity Application Flexibility Known Bias Poor Reproducibility Loss of Complexity Precise input requirements Illumina-specific kit Fragmentation (Covaris) Fragmentase Enzyme Tagmentation Fragmentation End Repair A-Tailing End Repair Cleanup Cleanup Adapter Ligation Cleanup End Repair A-Addition Library Amp Cleanup A-Additon Adapter Ligation Library Amp Adapter Ligation Cleanup Cleanup Library Amp Library Amp 72
73 Superior coverage distribution & gold-standard % Duplication Fraction of target genome Coverage Distribution Coverage depth (X) QIAseq FX (100 ng) Supplier N Enzymatic (100 ng) Covaris + Standard LP (100 ng) Tagmentation Supplier I (1 ng) Duplication Rate, 1ng Input QIAseq FX (1ng and 100ng) Supplier N Enzymatic (1ng) Covaris + Standard LP (1 ng) Tagmentation Supplier I (1 ng) 73
74 QIAseq FX: Gold-Standard WGS libraries in just 2.5 hours For high throughput Illumina NGS facilities or small labs sequencing the whole genome (or human whole exome) of any organism, including metagenomics Need even genomic coverage to minimize the total sequencing and avoid dropout regions without sacrificing a fast workflow that s easy to scale QIAseq FX provides gold-standard WGS libraries in just 2.5 hours Competitor products: Covaris Fragmentation + Any Kit: Slow, extra tube transfers, expensive Illumina Nextera: Strong sequence bias, very expensive Illumina Nextera XT: Strong sequence bias, only for small genomes Kapa HyperPlus: No adapters provided, customer must source separately QIAseq FX DNA Library Kit (24), with 24-plex adapter plate QIAseq FX DNA Library Kit (96), with 96-plex adapter plate 74
75 QIAseq Library Products for Illumina Sequencers Products for Illumina Sequencers: Nucleic acid extraction NGS Library preparation Sequencing & QIAGEN Bioinformatics Whole Genome Sequencing (WGS) Hybrid Capture Sequencign QIAseq FX DNA Library Kit FX fragmentation step eliminates shearing 2.5 hour workflow from gdna to library Tunable enzymatic fragmentation for any size probes with higher genomic complexity than Nextera DNA Amplicon/ PCR sequencing QIAseq 1-Step Amplicon Library Kit 1-tube, room temperature library prep cfdna for Liquid Biopsy or NIPT QIAseq cfdna Complete Solution QIAamp Circulating Nucleic Acid Isolation Optimized protocol specific for cfdna Low Input ChIP-seq, cfdna, LCM QIAseq Ultralow Input Library Kit Ultra-efficient chemistry for pg+ DNA Barcoded adapters for 96-plex sequencing Single Cell Genomics (QIAseq Single Cell FX Lib Kits) PCR-free MDA Amplification Sorted single cells or 6pg DNA input 75
76 QIAseq 1-Step Amplicon: Fast Library Prep for PCR Products 30 minute single-tube room-temperature library prep for amplicon targeted re-sequencing Purified PCR Products (1-500 ng) GeneRead V2 Targeted Panels Other Panels (ie. AmpliSeq) Other PCR reactions Single-tube room temperature library prep (30 min) Proprietary single-tube technology: Only 1 pipetting step Fastest Possible Library Prep: 30 minutes! Compatible with any PCR amplicons and gene panels from QIAGEN or other manufacturers library amplification (optional) (45 min) 76
77 QIAseq 1-Step Amplicon: Fast Library Prep for PCR Products For translational research, biomarker discovery and applied testing using a targeted DNA panel (QIAGEN, 3 rd party, or home brew) on Illumina sequencers QIAseq 1-Step Amplicon provides the fastest, easiest workflow from PCR product to sequencer-ready libraries Competitor products workflows 2.5 hours (Kapa) to 6 hours (Illumina TruSeq) Illumina TruSeq Nano: 5-6 hours QIAseq 1-Step Amplicon advantages: 30 minute PCR-free workflow with a single room temp reaction to set up 96-plex adapters included in 96 reaction kit Compatible with ANY multiplexed PCR or gene panels QIAseq 1-Step Amplicon Library Kit (12), no adapters QIAseq 1-Step Amplicon Library Kit (96), with adapters 77
78 QIAseq Library Products for Illumina Sequencers Products for Illumina Sequencers: Nucleic acid extraction NGS Library preparation Sequencing & QIAGEN Bioinformatics Whole Genome Sequencing (WGS) Hybrid Capture Sequencign QIAseq FX DNA Library Kit FX fragmentation step eliminates shearing 2.5 hour workflow from gdna to library Tunable enzymatic fragmentation for any size probes with higher genomic complexity than Nextera DNA Amplicon/ PCR sequencing QIAseq 1-Step Amplicon Library Kit 1-tube, room temperature library prep cfdna for Liquid Biopsy or NIPT QIAseq cfdna Complete Solution QIAamp Circulating Nucleic Acid Isolation Optimized protocol specific for cfdna Low Input ChIP-seq, cfdna, LCM QIAseq Ultralow Input Library Kit Ultra-efficient chemistry for pg+ DNA Barcoded adapters for 96-plex sequencing Single Cell Genomics (QIAseq Single Cell FX Lib Kits) PCR-free MDA Amplification Sorted single cells or 6pg DNA input 78
79 QIAseq Ultralow Input: Maximize Sub-Nanogram Samples Fragmented gdna 10pg 100ng End-polishing Barcoded Adapter Ligation ULTRA-EFFICIENT library prep for maximum library with minimum PCR SUPERIOR DATA including low % adapter and even G/C coverage WHOLE GENOME library complexity from subnanogram input IDEAL for oncology LCM, Liquid Biopsy cfdna, ChIP-Seq, and low input whole genome sequencing Reaction Cleanup Library Amplification (Optional for >10ng input) 79
80 The highest library yield and quality from as little as pg DNA Comparison of Input Requirements ULTRA-EFFICIENT chemistry balances yield with cycles of PCR SUPERIOR % adapter and G/C coverage WHOLE GENOME library complexity IDEAL for cfdna, ChIP-Seq, and low input whole genome sequencing 80
81 QIAseq Ultralow Input: Maximize Sub-Nanogram Samples Launch Date: July 2016 For Illumina sequencing of fragmented DNA, especially limited DNA sample types. Liquid Biopsy cfdna, Ancient DNA, Laser Capture Microdissection, FFPE Sequencing, ChIP-Seq The best possible yield and genome coverage from as little as pg of DNA The best option for low input when REPLI-g pre-amplification is not possible Compatible with Covaris mechanical shearing (up to 100ng DNA) Competitor products: Illumina TruSeq Nano (100ng): Slow, extra tube transfers, expensive Kapa HyperPrep (1ng): No adapters provided, must source separately NEB Ultra II (500pg): 24-plex, tube-format adapters offered separately QIAseq Ultralow Input Library Kit (12), without adapters QIAseq Ultralow Input Library Kit (96), with 96 plex adapters 81
82 QIAGEN Recommended Workflows for Exome Hybrid Capture Exome Sequencing from Human gdna Exome Sequencing from circulating cfdna Exome Sequencing from gdna using Covaris Sample Type Human gdna Circulating cell-free DNA Human gdna Fragmentation Library Prep QIAseq FX DNA Library Kit Cat no QIAseq FX DNA Library Kit Cat no None cfdna is naturally fragmented QIAseq Ultralow Input DNA Library Kit Cat no Covaris QIAseq Ultralow Input DNA Library Kit Cat no Exome Capture Hybrid Capture Probes such as: IDT xgen Exome Research Panel v1.0 Protocol: Hybridization capture of DNA libraries using xgen Lockdown Probes and Reagents Amp GeneRead DNA I Amp Kit (100) Cat no QIAseq FX IDT xgen Exome Hybrid Capture (5.5 hours) GeneRead I Amp Kit QIAseq Library Quant QC QIAseq Ultralow Input IDT xgen Exome Hybrid Capture (5.5 hours) GeneRead I Amp Kit QIAseq Library Quant QC Fragmentation (Covaris) QIAseq Ultralow Input IDT xgen Exome Hybrid Capture (5.5 hours) GeneRead I Amp Kit QIAseq Library Quant QC 82
83 QIAseq Library Products for Illumina Sequencers Products for Illumina Sequencers: Nucleic acid extraction NGS Library preparation Sequencing & QIAGEN Bioinformatics Whole Genome Sequencing (WGS) Hybrid Capture Sequencign QIAseq FX DNA Library Kit FX fragmentation step eliminates shearing 2.5 hour workflow from gdna to library Tunable enzymatic fragmentation for any size probes with higher genomic complexity than Nextera DNA Amplicon/ PCR sequencing QIAseq 1-Step Amplicon Library Kit 1-tube, room temperature library prep cfdna for Liquid Biopsy or NIPT QIAseq cfdna Complete Solution QIAamp Circulating Nucleic Acid Isolation Optimized protocol specific for cfdna Low Input ChIP-seq, cfdna, LCM QIAseq Ultralow Input Library Kit Ultra-efficient chemistry for pg+ DNA Barcoded adapters for 96-plex sequencing Single Cell Genomics (QIAseq Single Cell FX Lib Kits) PCR-free MDA Amplification Sorted single cells or 6pg DNA input 83
84 QIAGEN s REPLI-g Technology: use MDA, not PCR! MDA: Multiple Displacement Amplification Alkaline denaturation Hexamer random primers Phi29 polymerase strand displacement (30 C) Displaced strand becomes a template for replication Gentle denaturation results in intact DNA-template Even priming events covering the whole genome immediate amplification across all regions Modified high-fidelity Phi29 polymerase (SensiPhi) o High enzyme processivity molecular weight product ( kb) o Proofreading activity 1000-fold higher accuracy than normal PCR-enzyme Multiple displacement events efficient amplification of DNA 84
85 MDA amplifies through gdna secondary structure regions Denatured gdna has a complex secondary structure Consists of regions of ssdna and dsdna that can form complicated hairpins and loops QIAGEN s MDA enzyme handles complex DNA structures generating extremely long amplicons 85
86 Qiagen s Single Cell Portfolio: The cutting edge of genomics Highest accuracy Superior coverage High diversity, high quality libraries Streamlined workflows Data analyis & visualization Biological interpretation WGA or WTA Library preparation Sequencing & QIAGEN Bioinformatics Single-cell whole genome sequencing REPLI-g Single Cell DNA Library Kit (48) Cat no Q3 2016: QIAseq FX Single Cell DNA Kit Superior solution for variant calling, ideal for applications where SNV & CNV is equally important Single-cell gene expression analysis by RNAseq REPLI-g Single Cell RNA Library Kit(24) Cat no Q32016: QIAseq FX Single Cell RNA Kit Streamlined workflow with reliable detection of even low-abundance transcripts REPLI-g Single Cell Kit Single-cell targeted sequencing & QIAseq Targeted DNA Panels High quality targeted sequencing from a single cell for DNA variants 86
87 QIAseq Library Products for Illumina Sequencers Products for Illumina Sequencers: Nucleic acid extraction NGS Library preparation Sequencing & QIAGEN Bioinformatics Whole Genome Sequencing (WGS) Hybrid Capture Sequencign QIAseq FX DNA Library Kit FX fragmentation step eliminates shearing 2.5 hour workflow from gdna to library Tunable enzymatic fragmentation for any size probes with higher genomic complexity than Nextera DNA Amplicon/ PCR sequencing QIAseq 1-Step Amplicon Library Kit 1-tube, room temperature library prep cfdna for Liquid Biopsy or NIPT QIAseq cfdna Complete Solution QIAamp Circulating Nucleic Acid Isolation Optimized protocol specific for cfdna Low Input ChIP-seq, cfdna, LCM QIAseq Ultralow Input Library Kit Ultra-efficient chemistry for pg+ DNA Barcoded adapters for 96-plex sequencing Single Cell Genomics (QIAseq Single Cell FX Lib Kits) PCR-free MDA Amplification Sorted single cells or 6pg DNA input 87
88 QIAseq ccf DNA Workflow Designed for any NGS-based cfdna research Up to 5 ml plasma can be used for cfdna extraction using QIAamp Mini Columns. 24 samples are processed in less than 2 h ng of cfdna can be directly used from up to 56.5 µl eluant volume. Quantifcation of cfdna prior to library prep is not necessary. Library prep includes end-polishing and adapter ligation and takes < 1 h. Optional HiFi amplification can be performed for < 10 ng cfdna before samples undergo sequencing. 88
89 QIAseq ccf DNA Workflow 89
90 QIAseq sequencing summary Unique molecular indexes remove bias due to technique and give improved data Applications for DNA variants, Gene expression, Fusion gene, mirna Complete workflow from start to data analysis On-Line based platform CLC-Bio solution (gene expression, more to follow) Leverage Qiagen content know-how for NGS Disease and pathway specific collections Extended panels where you add your targetes to our Panels Customizable solutions Library Preparation kits and applications Circulating cell-free DNA Single Cell Genomics (DNA/RNA) Genomic DNA (WGS/HC) enzymatic fragmentation in 1 tube Ultra-low input sequencing (10pg) 1-step amplicon sequencing 90
QIAGEN s NGS Solutions for Biomarkers NGS & Bioinformatics team QIAGEN (Suzhou) Translational Medicine Co.,Ltd
 QIAGEN s NGS Solutions for Biomarkers NGS & Bioinformatics team QIAGEN (Suzhou) Translational Medicine Co.,Ltd 1 Our current NGS & Bioinformatics Platform 2 Our NGS workflow and applications 3 QIAGEN s
QIAGEN s NGS Solutions for Biomarkers NGS & Bioinformatics team QIAGEN (Suzhou) Translational Medicine Co.,Ltd 1 Our current NGS & Bioinformatics Platform 2 Our NGS workflow and applications 3 QIAGEN s
Sample to Insight. Dr. Bhagyashree S. Birla NGS Field Application Scientist
 Dr. Bhagyashree S. Birla NGS Field Application Scientist bhagyashree.birla@qiagen.com NGS spans a broad range of applications DNA Applications Human ID Liquid biopsy Biomarker discovery Inherited and somatic
Dr. Bhagyashree S. Birla NGS Field Application Scientist bhagyashree.birla@qiagen.com NGS spans a broad range of applications DNA Applications Human ID Liquid biopsy Biomarker discovery Inherited and somatic
Exploring new frontiers with next-generation sequencing
 Spotlight: NGS Exploring new frontiers with next-generation sequencing Pushing the limits of discovery Next-generation sequencing (NGS) is being utilized for numerous new and exciting applications, such
Spotlight: NGS Exploring new frontiers with next-generation sequencing Pushing the limits of discovery Next-generation sequencing (NGS) is being utilized for numerous new and exciting applications, such
Digital DNA/RNA sequencing enables highly accurate and sensitive biomarker detection and quantification
 Digital DNA/RNA sequencing enables highly accurate and sensitive biomarker detection and quantification Erwin Chen ( 陳立德 ) Technical Product Specialist QIAGEN Taiwan Precision medicine: Right drug, right
Digital DNA/RNA sequencing enables highly accurate and sensitive biomarker detection and quantification Erwin Chen ( 陳立德 ) Technical Product Specialist QIAGEN Taiwan Precision medicine: Right drug, right
Development of quantitative targeted RNA-seq methodology for use in differential gene expression
 Development of quantitative targeted RNA-seq methodology for use in differential gene expression Dr. Jens Winter, Market Development Group Biological Biological Research Content EMEA QIAGEN Universal Workflows
Development of quantitative targeted RNA-seq methodology for use in differential gene expression Dr. Jens Winter, Market Development Group Biological Biological Research Content EMEA QIAGEN Universal Workflows
Introducing QIAseq. Accelerate your NGS performance through Sample to Insight solutions. Sample to Insight
 Introducing QIAseq Accelerate your NGS performance through Sample to Insight solutions Sample to Insight From Sample to Insight let QIAGEN enhance your NGS-based research High-throughput next-generation
Introducing QIAseq Accelerate your NGS performance through Sample to Insight solutions Sample to Insight From Sample to Insight let QIAGEN enhance your NGS-based research High-throughput next-generation
Welcome to the NGS webinar series
 Welcome to the NGS webinar series Webinar 1 NGS: Introduction to technology, and applications NGS Technology Webinar 2 Targeted NGS for Cancer Research NGS in cancer Webinar 3 NGS: Data analysis for genetic
Welcome to the NGS webinar series Webinar 1 NGS: Introduction to technology, and applications NGS Technology Webinar 2 Targeted NGS for Cancer Research NGS in cancer Webinar 3 NGS: Data analysis for genetic
Accelerate your NGS performance through Sample to Insight solutions
 Accelerate your NGS performance through Sample to Insight solutions Complete your NGS workflow and save up to 40% Sample to Insight From Sample to Insight let QIAGEN enhance your NGS-based research High-throughput
Accelerate your NGS performance through Sample to Insight solutions Complete your NGS workflow and save up to 40% Sample to Insight From Sample to Insight let QIAGEN enhance your NGS-based research High-throughput
QIAseq SPE technology for Illumina : Redefining amplicon sequencing
 Application Note QIAseq SPE technology for Illumina : Redefining amplicon sequencing Amplicon-based enrichment and sequencing takes advantage of PCR workflows to turn amplicons that represent regions of
Application Note QIAseq SPE technology for Illumina : Redefining amplicon sequencing Amplicon-based enrichment and sequencing takes advantage of PCR workflows to turn amplicons that represent regions of
ACCEL-NGS 2S DNA LIBRARY KITS
 ACCEL-NGS 2S DNA LIBRARY KITS Accel-NGS 2S DNA Library Kits produce high quality libraries with an all-inclusive, easy-to-use format. The kits contain all reagents necessary to build high complexity libraries
ACCEL-NGS 2S DNA LIBRARY KITS Accel-NGS 2S DNA Library Kits produce high quality libraries with an all-inclusive, easy-to-use format. The kits contain all reagents necessary to build high complexity libraries
QIAseq mirna Library Kit The next-generation in mirna sequencing products
 QIAseq mirna Library Kit The next-generation in mirna sequencing products What challenges are there with mirna Sequencing? Sample input is one of many challenges facing mirna sequencing reactions. Kits
QIAseq mirna Library Kit The next-generation in mirna sequencing products What challenges are there with mirna Sequencing? Sample input is one of many challenges facing mirna sequencing reactions. Kits
SureSelect XT HS. Target Enrichment
 SureSelect XT HS Target Enrichment What Is It? SureSelect XT HS joins the SureSelect library preparation reagent family as Agilent s highest sensitivity hybrid capture-based library prep and target enrichment
SureSelect XT HS Target Enrichment What Is It? SureSelect XT HS joins the SureSelect library preparation reagent family as Agilent s highest sensitivity hybrid capture-based library prep and target enrichment
Integrated NGS Sample Preparation Solutions for Limiting Amounts of RNA and DNA. March 2, Steven R. Kain, Ph.D. ABRF 2013
 Integrated NGS Sample Preparation Solutions for Limiting Amounts of RNA and DNA March 2, 2013 Steven R. Kain, Ph.D. ABRF 2013 NuGEN s Core Technologies Selective Sequence Priming Nucleic Acid Amplification
Integrated NGS Sample Preparation Solutions for Limiting Amounts of RNA and DNA March 2, 2013 Steven R. Kain, Ph.D. ABRF 2013 NuGEN s Core Technologies Selective Sequence Priming Nucleic Acid Amplification
Deep Sequencing technologies
 Deep Sequencing technologies Gabriela Salinas 30 October 2017 Transcriptome and Genome Analysis Laboratory http://www.uni-bc.gwdg.de/index.php?id=709 Microarray and Deep-Sequencing Core Facility University
Deep Sequencing technologies Gabriela Salinas 30 October 2017 Transcriptome and Genome Analysis Laboratory http://www.uni-bc.gwdg.de/index.php?id=709 Microarray and Deep-Sequencing Core Facility University
Ion S5 and Ion S5 XL Systems
 Ion S5 and Ion S5 XL Systems Targeted sequencing has never been simpler Explore the Ion S5 and Ion S5 XL Systems Adopting next-generation sequencing (NGS) in your lab is now simpler than ever The Ion S5
Ion S5 and Ion S5 XL Systems Targeted sequencing has never been simpler Explore the Ion S5 and Ion S5 XL Systems Adopting next-generation sequencing (NGS) in your lab is now simpler than ever The Ion S5
HaloPlex HS. Get to Know Your DNA. Every Single Fragment. Kevin Poon, Ph.D.
 HaloPlex HS Get to Know Your DNA. Every Single Fragment. Kevin Poon, Ph.D. Sr. Global Product Manager Diagnostics & Genomics Group Agilent Technologies For Research Use Only. Not for Use in Diagnostic
HaloPlex HS Get to Know Your DNA. Every Single Fragment. Kevin Poon, Ph.D. Sr. Global Product Manager Diagnostics & Genomics Group Agilent Technologies For Research Use Only. Not for Use in Diagnostic
Ion S5 and Ion S5 XL Systems
 Ion S5 and Ion S5 XL Systems Targeted sequencing has never been simpler Explore the Ion S5 and Ion S5 XL Systems Adopting next-generation sequencing (NGS) in your lab is now simpler than ever The Ion S5
Ion S5 and Ion S5 XL Systems Targeted sequencing has never been simpler Explore the Ion S5 and Ion S5 XL Systems Adopting next-generation sequencing (NGS) in your lab is now simpler than ever The Ion S5
RAPID, ROBUST & RELIABLE
 Roche Sample Prep Solutions for RNA-Seq Sequence what matters RAPID, ROBUST & RELIABLE Sample P le Samp Quant ifi /QC tion ca As the first step in the NGS workflow continuum, sample prep holds the key
Roche Sample Prep Solutions for RNA-Seq Sequence what matters RAPID, ROBUST & RELIABLE Sample P le Samp Quant ifi /QC tion ca As the first step in the NGS workflow continuum, sample prep holds the key
High-quality stranded RNA-seq libraries from single cells using the SMART-Seq Stranded Kit Product highlights:
 TECH NOTE High-quality stranded RNA-seq libraries from single cells using the SMART-Seq Stranded Kit Product highlights: Simple workflow starts directly from 1 1,000 cells or 10 pg 10 ng total RNA to generate
TECH NOTE High-quality stranded RNA-seq libraries from single cells using the SMART-Seq Stranded Kit Product highlights: Simple workflow starts directly from 1 1,000 cells or 10 pg 10 ng total RNA to generate
Your Best Data: Teaming QIAGEN Chemistry & Bioinformatics to Drive Samples to Insight
 Your Best Data: Teaming QIAGEN Chemistry & Bioinformatics to Drive Samples to Insight Aysel Heckel Director Clinical Solutions Sales Dr. Anne Arens Field Application Scientist Course on Variant Detection
Your Best Data: Teaming QIAGEN Chemistry & Bioinformatics to Drive Samples to Insight Aysel Heckel Director Clinical Solutions Sales Dr. Anne Arens Field Application Scientist Course on Variant Detection
Surely Better Target Enrichment from Sample to Sequencer
 sureselect TARGET ENRICHMENT solutions Surely Better Target Enrichment from Sample to Sequencer Agilent s market leading SureSelect platform provides a complete portfolio of catalog to custom products,
sureselect TARGET ENRICHMENT solutions Surely Better Target Enrichment from Sample to Sequencer Agilent s market leading SureSelect platform provides a complete portfolio of catalog to custom products,
Incorporating Molecular ID Technology. Accel-NGS 2S MID Indexing Kits
 Incorporating Molecular ID Technology Accel-NGS 2S MID Indexing Kits Molecular Identifiers (MIDs) MIDs are indices used to label unique library molecules MIDs can assess duplicate molecules in sequencing
Incorporating Molecular ID Technology Accel-NGS 2S MID Indexing Kits Molecular Identifiers (MIDs) MIDs are indices used to label unique library molecules MIDs can assess duplicate molecules in sequencing
DNA concentration and purity were initially measured by NanoDrop 2000 and verified on Qubit 2.0 Fluorometer.
 DNA Preparation and QC Extraction DNA was extracted from whole blood or flash frozen post-mortem tissue using a DNA mini kit (QIAmp #51104 and QIAmp#51404, respectively) following the manufacturer s recommendations.
DNA Preparation and QC Extraction DNA was extracted from whole blood or flash frozen post-mortem tissue using a DNA mini kit (QIAmp #51104 and QIAmp#51404, respectively) following the manufacturer s recommendations.
Novel methods for RNA and DNA- Seq analysis using SMART Technology. Andrew Farmer, D. Phil. Vice President, R&D Clontech Laboratories, Inc.
 Novel methods for RNA and DNA- Seq analysis using SMART Technology Andrew Farmer, D. Phil. Vice President, R&D Clontech Laboratories, Inc. Agenda Enabling Single Cell RNA-Seq using SMART Technology SMART
Novel methods for RNA and DNA- Seq analysis using SMART Technology Andrew Farmer, D. Phil. Vice President, R&D Clontech Laboratories, Inc. Agenda Enabling Single Cell RNA-Seq using SMART Technology SMART
Lab methods: Exome / Genome. Ewart de Bruijn
 Lab methods: Exome / Genome 27 06 2013 Ewart de Bruijn Library prep is only a small part of the complete DNA analysis workflow DNA isolation library prep enrichment flowchip prep sequencing bioinformatics
Lab methods: Exome / Genome 27 06 2013 Ewart de Bruijn Library prep is only a small part of the complete DNA analysis workflow DNA isolation library prep enrichment flowchip prep sequencing bioinformatics
SURESELECTXT LOW INPUT TARGET ENRICHMENT
 SURESELECTXT LOW INPUT TARGET ENRICHMENT Low Input FFPE Optimized Streamlined Workflow SureSelect XT Low Input What is it? SureSelect XT Low Input is a low-input, FFPE-optimized library preparation kit.
SURESELECTXT LOW INPUT TARGET ENRICHMENT Low Input FFPE Optimized Streamlined Workflow SureSelect XT Low Input What is it? SureSelect XT Low Input is a low-input, FFPE-optimized library preparation kit.
Illumina s Suite of Targeted Resequencing Solutions
 Illumina s Suite of Targeted Resequencing Solutions Colin Baron Sr. Product Manager Sequencing Applications 2011 Illumina, Inc. All rights reserved. Illumina, illuminadx, Solexa, Making Sense Out of Life,
Illumina s Suite of Targeted Resequencing Solutions Colin Baron Sr. Product Manager Sequencing Applications 2011 Illumina, Inc. All rights reserved. Illumina, illuminadx, Solexa, Making Sense Out of Life,
RIPTIDE HIGH THROUGHPUT RAPID LIBRARY PREP (HT-RLP)
 Application Note: RIPTIDE HIGH THROUGHPUT RAPID LIBRARY PREP (HT-RLP) Introduction: Innovations in DNA sequencing during the 21st century have revolutionized our ability to obtain nucleotide information
Application Note: RIPTIDE HIGH THROUGHPUT RAPID LIBRARY PREP (HT-RLP) Introduction: Innovations in DNA sequencing during the 21st century have revolutionized our ability to obtain nucleotide information
Next Generation Sequencing. Target Enrichment
 Next Generation Sequencing Target Enrichment Next Generation Sequencing Your Partner in Every Step from Sample to Data NGS: Revolutionizing Genetic Analysis with Single-Molecule Resolution Next generation
Next Generation Sequencing Target Enrichment Next Generation Sequencing Your Partner in Every Step from Sample to Data NGS: Revolutionizing Genetic Analysis with Single-Molecule Resolution Next generation
High Throughput Sequencing the Multi-Tool of Life Sciences. Lutz Froenicke DNA Technologies and Expression Analysis Cores UCD Genome Center
 High Throughput Sequencing the Multi-Tool of Life Sciences Lutz Froenicke DNA Technologies and Expression Analysis Cores UCD Genome Center Complementary Approaches Illumina Still-imaging of clusters (~1000
High Throughput Sequencing the Multi-Tool of Life Sciences Lutz Froenicke DNA Technologies and Expression Analysis Cores UCD Genome Center Complementary Approaches Illumina Still-imaging of clusters (~1000
High-yield, Scalable Library Preparation with the NEBNext Ultra II FS DNA Library Prep Kit
 be INSPIRED drive DISCOVERY stay GENUINE TECHNICAL NOTE High-yield, Scalable Library Preparation with the NEBNext Ultra II FS DNA Library Prep Kit Improving performance, ease of use and reliability of
be INSPIRED drive DISCOVERY stay GENUINE TECHNICAL NOTE High-yield, Scalable Library Preparation with the NEBNext Ultra II FS DNA Library Prep Kit Improving performance, ease of use and reliability of
Agilent NGS Solutions : Addressing Today s Challenges
 Agilent NGS Solutions : Addressing Today s Challenges Charmian Cher, Ph.D Director, Global Marketing Programs 1 10 years of Next-Gen Sequencing 2003 Completion of the Human Genome Project 2004 Pyrosequencing
Agilent NGS Solutions : Addressing Today s Challenges Charmian Cher, Ph.D Director, Global Marketing Programs 1 10 years of Next-Gen Sequencing 2003 Completion of the Human Genome Project 2004 Pyrosequencing
Cancer Genetics Solutions
 Cancer Genetics Solutions Cancer Genetics Solutions Pushing the Boundaries in Cancer Genetics Cancer is a formidable foe that presents significant challenges. The complexity of this disease can be daunting
Cancer Genetics Solutions Cancer Genetics Solutions Pushing the Boundaries in Cancer Genetics Cancer is a formidable foe that presents significant challenges. The complexity of this disease can be daunting
Surely Better Target Enrichment from Sample to Sequencer and Analysis
 sureselect TARGET ENRIChment solutions Surely Better Target Enrichment from Sample to Sequencer and Analysis Agilent s market leading SureSelect platform provides a complete portfolio of catalog to custom
sureselect TARGET ENRIChment solutions Surely Better Target Enrichment from Sample to Sequencer and Analysis Agilent s market leading SureSelect platform provides a complete portfolio of catalog to custom
Single Cell Genomics
 Single Cell Genomics Application Cost Platform/Protocol Note 3 mrna-seq Cell capture/rt/library prep $1,990/Sample 10x Genomics Chromium 500-10,000 cells/sample 5 V(D)J + mrna Cell capture/rt/library prep
Single Cell Genomics Application Cost Platform/Protocol Note 3 mrna-seq Cell capture/rt/library prep $1,990/Sample 10x Genomics Chromium 500-10,000 cells/sample 5 V(D)J + mrna Cell capture/rt/library prep
QIAGEN s Preanalytic NGS Solutions
 QIAGEN s Preanalytic NGS Solutions For outstanding DNAseq results Choose QIAGEN for every step of your NGS workflow Sample & Assay Technologies 1. Next-generation sequencing (NGS) Purification and Sample
QIAGEN s Preanalytic NGS Solutions For outstanding DNAseq results Choose QIAGEN for every step of your NGS workflow Sample & Assay Technologies 1. Next-generation sequencing (NGS) Purification and Sample
Gene Regulation Solutions. Microarrays and Next-Generation Sequencing
 Gene Regulation Solutions Microarrays and Next-Generation Sequencing Gene Regulation Solutions The Microarrays Advantage Microarrays Lead the Industry in: Comprehensive Content SurePrint G3 Human Gene
Gene Regulation Solutions Microarrays and Next-Generation Sequencing Gene Regulation Solutions The Microarrays Advantage Microarrays Lead the Industry in: Comprehensive Content SurePrint G3 Human Gene
Single Cell Genomics
 Single Cell Genomics Application Cost Platform/Protoc ol Note Single cell 3 mrna-seq cell lysis/rt/library prep $2460/Sample 10X Genomics Chromium 500-10,000 cells/sample Single cell 5 V(D)J mrna-seq cell
Single Cell Genomics Application Cost Platform/Protoc ol Note Single cell 3 mrna-seq cell lysis/rt/library prep $2460/Sample 10X Genomics Chromium 500-10,000 cells/sample Single cell 5 V(D)J mrna-seq cell
G E N OM I C S S E RV I C ES
 GENOMICS SERVICES ABOUT T H E N E W YOR K G E NOM E C E N T E R NYGC is an independent non-profit implementing advanced genomic research to improve diagnosis and treatment of serious diseases. Through
GENOMICS SERVICES ABOUT T H E N E W YOR K G E NOM E C E N T E R NYGC is an independent non-profit implementing advanced genomic research to improve diagnosis and treatment of serious diseases. Through
Get to Know Your DNA. Every Single Fragment.
 HaloPlex HS NGS Target Enrichment System Get to Know Your DNA. Every Single Fragment. High sensitivity detection of rare variants using molecular barcodes How Does Molecular Barcoding Work? HaloPlex HS
HaloPlex HS NGS Target Enrichment System Get to Know Your DNA. Every Single Fragment. High sensitivity detection of rare variants using molecular barcodes How Does Molecular Barcoding Work? HaloPlex HS
Increased transcription detection with the NEBNext Single Cell/Low Input RNA Library Prep Kit
 be INSPIRED drive DISCOVERY stay GENUINE TECHNICAL NOTE Increased transcription detection with the NEBNext Single Cell/Low Input RNA Library Prep Kit Highly sensitive, robust generation of high quality
be INSPIRED drive DISCOVERY stay GENUINE TECHNICAL NOTE Increased transcription detection with the NEBNext Single Cell/Low Input RNA Library Prep Kit Highly sensitive, robust generation of high quality
SureSelect Target Enrichment for the Ion Proton TM Next Generation Sequencing System
 SureSelect Target Enrichment for the Ion Proton TM Next Generation Sequencing System Demonstrated performance you can count on Christina Chiu Product Manager, SureSelect Kyeong Jeong Ph.D. R&D Scientist
SureSelect Target Enrichment for the Ion Proton TM Next Generation Sequencing System Demonstrated performance you can count on Christina Chiu Product Manager, SureSelect Kyeong Jeong Ph.D. R&D Scientist
Whole Genome Amplification (WGA): What to Do When You Don t Have Enough Genomic DNA
 Whole Genome Amplification (WGA): What to Do When You Don t Have Enough Genomic DNA Rob Brazas, Ph.D. Senior Product Manager, Lucigen January, 2017 www.lucigen.com Agenda Improving Whole Genome Amplified
Whole Genome Amplification (WGA): What to Do When You Don t Have Enough Genomic DNA Rob Brazas, Ph.D. Senior Product Manager, Lucigen January, 2017 www.lucigen.com Agenda Improving Whole Genome Amplified
Wet-lab Considerations for Illumina data analysis
 Wet-lab Considerations for Illumina data analysis Based on a presentation by Henriette O Geen Lutz Froenicke DNA Technologies and Expression Analysis Cores UCD Genome Center Complementary Approaches Illumina
Wet-lab Considerations for Illumina data analysis Based on a presentation by Henriette O Geen Lutz Froenicke DNA Technologies and Expression Analysis Cores UCD Genome Center Complementary Approaches Illumina
Fully Automated Library Quantification for Illumina Sequencing on the NGS STAR
 Fully Automated Library Quantification for Illumina Sequencing on the NGS STAR Introduction Hamilton Robotics, an industry leader in liquid handling and laboratory automation equipment, has partnered with
Fully Automated Library Quantification for Illumina Sequencing on the NGS STAR Introduction Hamilton Robotics, an industry leader in liquid handling and laboratory automation equipment, has partnered with
Next-generation sequencing technologies
 Next-generation sequencing technologies NGS applications Illumina sequencing workflow Overview Sequencing by ligation Short-read NGS Sequencing by synthesis Illumina NGS Single-molecule approach Long-read
Next-generation sequencing technologies NGS applications Illumina sequencing workflow Overview Sequencing by ligation Short-read NGS Sequencing by synthesis Illumina NGS Single-molecule approach Long-read
ab High Sensitivity DNA Library Preparation Kit (For Illumina )
 ab185905 High Sensitivity DNA Library Preparation Kit (For Illumina ) Instructions for Use For the preparation of a DNA library using sub-nanogram amounts of DNA input for next generation sequencing applications
ab185905 High Sensitivity DNA Library Preparation Kit (For Illumina ) Instructions for Use For the preparation of a DNA library using sub-nanogram amounts of DNA input for next generation sequencing applications
ab High Sensitivity DNA Library Preparation Kit (For Illumina )
 ab185905 High Sensitivity DNA Library Preparation Kit (For Illumina ) Instructions for Use For the preparation of a DNA library using sub-nanogram amounts of DNA input for next generation sequencing applications
ab185905 High Sensitivity DNA Library Preparation Kit (For Illumina ) Instructions for Use For the preparation of a DNA library using sub-nanogram amounts of DNA input for next generation sequencing applications
TECH NOTE Pushing the Limit: A Complete Solution for Generating Stranded RNA Seq Libraries from Picogram Inputs of Total Mammalian RNA
 TECH NOTE Pushing the Limit: A Complete Solution for Generating Stranded RNA Seq Libraries from Picogram Inputs of Total Mammalian RNA Stranded, Illumina ready library construction in
TECH NOTE Pushing the Limit: A Complete Solution for Generating Stranded RNA Seq Libraries from Picogram Inputs of Total Mammalian RNA Stranded, Illumina ready library construction in
High Throughput Sequencing the Multi-Tool of Life Sciences. Lutz Froenicke DNA Technologies and Expression Analysis Cores UCD Genome Center
 High Throughput Sequencing the Multi-Tool of Life Sciences Lutz Froenicke DNA Technologies and Expression Analysis Cores UCD Genome Center DNA Technologies & Expression Analysis Cores HT Sequencing (Illumina
High Throughput Sequencing the Multi-Tool of Life Sciences Lutz Froenicke DNA Technologies and Expression Analysis Cores UCD Genome Center DNA Technologies & Expression Analysis Cores HT Sequencing (Illumina
EPIGENTEK. EpiNext DNA Library Preparation Kit (Illumina) Base Catalog # P-1051 PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE
 EpiNext DNA Library Preparation Kit (Illumina) Base Catalog # PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE Uses: The EpiNext DNA Library Preparation Kit (Illumina) is suitable for preparing a DNA library
EpiNext DNA Library Preparation Kit (Illumina) Base Catalog # PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE Uses: The EpiNext DNA Library Preparation Kit (Illumina) is suitable for preparing a DNA library
GENOMICS WORKFLOW SOLUTIONS THAT GO WHERE THE SCIENCE LEADS. Genomics Solutions Portfolio
 GENOMICS WORKFLOW SOLUTIONS THAT GO WHERE THE SCIENCE LEADS Genomics Solutions Portfolio WORKFLOW SOLUTIONS FROM EXTRACTION TO ANALYSIS Application-based answers for every step of your workflow Scientists
GENOMICS WORKFLOW SOLUTIONS THAT GO WHERE THE SCIENCE LEADS Genomics Solutions Portfolio WORKFLOW SOLUTIONS FROM EXTRACTION TO ANALYSIS Application-based answers for every step of your workflow Scientists
A Genomics (R)evolution: Harnessing the Power of Single Cells
 A Genomics (R)evolution: Harnessing the Power of Single Cells Fundamental Question #1 If Transcriptional Heterogeneity ( Noise ) is so great in single cells What s the Point? Single Cells = True Biology
A Genomics (R)evolution: Harnessing the Power of Single Cells Fundamental Question #1 If Transcriptional Heterogeneity ( Noise ) is so great in single cells What s the Point? Single Cells = True Biology
SOLiD Total RNA-Seq Kit SOLiD RNA Barcoding Kit
 SOLiD Total RNA-Seq Kit SOLiD RNA Barcoding Kit Agenda SOLiD Total RNAseq Kit Overview Kit Configurations Barcoding Kit Introduction New Small RNA and WT Workflow Small RNA Workflow Step-by-step Workflow
SOLiD Total RNA-Seq Kit SOLiD RNA Barcoding Kit Agenda SOLiD Total RNAseq Kit Overview Kit Configurations Barcoding Kit Introduction New Small RNA and WT Workflow Small RNA Workflow Step-by-step Workflow
Maximizing your NGS sequencing with IDT. Adam Chernick, PhD Field Applications Manager, Functional Genomics
 Maximizing your NGS sequencing with IDT Adam Chernick, PhD Field Applications Manager, Functional Genomics 1 Contents Expanding our NGS portfolio what s next? xgen technology and Lockdown probe advantages
Maximizing your NGS sequencing with IDT Adam Chernick, PhD Field Applications Manager, Functional Genomics 1 Contents Expanding our NGS portfolio what s next? xgen technology and Lockdown probe advantages
New Frontiers of Genetic Profiling Achieve Higher Sensitivity and Greater Insights with Molecular Barcodes, Long Read Capture and Optimized Exomes
 New Frontiers of Genetic Profiling Achieve Higher Sensitivity and Greater Insights with Molecular Barcodes, Long Read Capture and Optimized Exomes Jennifer Jones, PhD Senior Field Application Scientist
New Frontiers of Genetic Profiling Achieve Higher Sensitivity and Greater Insights with Molecular Barcodes, Long Read Capture and Optimized Exomes Jennifer Jones, PhD Senior Field Application Scientist
Roche Sequencing Solutions CHANGING SCIENCE CHANGING LIVES
 Roche Sequencing Solutions CHANGING SCIENCE CHANGING LIVES Roche Sequencing Solutions Roche is helping to shape the future of personalized medicine by integrating best-in-class sequencing technologies
Roche Sequencing Solutions CHANGING SCIENCE CHANGING LIVES Roche Sequencing Solutions Roche is helping to shape the future of personalized medicine by integrating best-in-class sequencing technologies
Isolation of total nucleic acids from FFPE tissues using FormaPure DNA
 APPLICATION NOTE Isolation of total nucleic acids from FFPE tissues using FormaPure DNA Jung Hoon Doh, Ph.D. Senior Application Scientist Beckman Coulter Life Sciences, Indianapolis, IN USA Summary Extensive
APPLICATION NOTE Isolation of total nucleic acids from FFPE tissues using FormaPure DNA Jung Hoon Doh, Ph.D. Senior Application Scientist Beckman Coulter Life Sciences, Indianapolis, IN USA Summary Extensive
Accessible answers. Targeted sequencing: accelerating and amplifying answers for oncology research
 Accessible answers Targeted sequencing: accelerating and amplifying answers for oncology research Help advance precision medicine Accelerate results with Ion Torrent NGS Life without cancer. This is our
Accessible answers Targeted sequencing: accelerating and amplifying answers for oncology research Help advance precision medicine Accelerate results with Ion Torrent NGS Life without cancer. This is our
Obtain superior NGS library performance with lower input amounts using the NEBNext Ultra II Directional RNA Library Prep Kit for Illumina
 be INSPIRED drive DISCOVERY stay GENINE TECHNICAL NOTE Directional rrna depletion Obtain superior NGS library performance with lower input amounts using the NEBNext ltra II Directional RNA Library Prep
be INSPIRED drive DISCOVERY stay GENINE TECHNICAL NOTE Directional rrna depletion Obtain superior NGS library performance with lower input amounts using the NEBNext ltra II Directional RNA Library Prep
Obtain superior NGS library performance with lower input amounts using the NEBNext Ultra II Directional RNA Library Prep Kit for Illumina
 be INSPIRED drive DISCOVERY stay GENINE TECHNICAL NOTE Directional rrna depletion Obtain superior NGS library performance with lower input amounts using the NEBNext ltra II Directional RNA Library Prep
be INSPIRED drive DISCOVERY stay GENINE TECHNICAL NOTE Directional rrna depletion Obtain superior NGS library performance with lower input amounts using the NEBNext ltra II Directional RNA Library Prep
The Expanded Illumina Sequencing Portfolio New Sample Prep Solutions and Workflow
 The Expanded Illumina Sequencing Portfolio New Sample Prep Solutions and Workflow Marcus Hausch, Ph.D. 2010 Illumina, Inc. All rights reserved. Illumina, illuminadx, Solexa, Making Sense Out of Life, Oligator,
The Expanded Illumina Sequencing Portfolio New Sample Prep Solutions and Workflow Marcus Hausch, Ph.D. 2010 Illumina, Inc. All rights reserved. Illumina, illuminadx, Solexa, Making Sense Out of Life, Oligator,
APPLICATION NOTE. Abstract. Introduction
 From minuscule amounts to magnificent results: reliable ChIP-seq data from 1, cells with the True MicroChIP and the MicroPlex Library Preparation kits Abstract Diagenode has developed groundbreaking solutions
From minuscule amounts to magnificent results: reliable ChIP-seq data from 1, cells with the True MicroChIP and the MicroPlex Library Preparation kits Abstract Diagenode has developed groundbreaking solutions
P HENIX. PHENIX PCR Enzyme Guide Tools For Life Science Discovery RESEARCH PRODUCTS
 PHENIX PCR Enzyme Guide PHENIX offers a broad line of premium quality PCR Enzymes. This PCR Enzyme Guide will help simplify your polymerase selection process. Each DNA Polymerase has different characteristics
PHENIX PCR Enzyme Guide PHENIX offers a broad line of premium quality PCR Enzymes. This PCR Enzyme Guide will help simplify your polymerase selection process. Each DNA Polymerase has different characteristics
MULTIPLEXING SIMULTANEOUSLY DETECT MULTIPLE TARGETS IN SINGLE ASSAYS. WhiteSci Whitehead Scientific (Pty) Ltd. Products. Expertise. Support.
 MULTIPLEXING SIMULTANEOUSLY DETECT MULTIPLE TARGETS IN SINGLE ASSAYS WhiteSci Whitehead Scientific (Pty) Ltd Products. Expertise. Support. MULTIPLEXING Allowing researchers to gain more insight into precious
MULTIPLEXING SIMULTANEOUSLY DETECT MULTIPLE TARGETS IN SINGLE ASSAYS WhiteSci Whitehead Scientific (Pty) Ltd Products. Expertise. Support. MULTIPLEXING Allowing researchers to gain more insight into precious
The MiniSeq System. Explore the possibilities. Discover demonstrated NGS workflows for molecular biology applications.
 The MiniSeq System. Explore the possibilities. Discover demonstrated NGS workflows for molecular biology applications. Let your work flow with Illumina NGS. The MiniSeq System delivers powerful and cost-effective
The MiniSeq System. Explore the possibilities. Discover demonstrated NGS workflows for molecular biology applications. Let your work flow with Illumina NGS. The MiniSeq System delivers powerful and cost-effective
SMARTer Ultra Low RNA Kit for Illumina Sequencing Two powerful technologies combine to enable sequencing with ultra-low levels of RNA
 SMARTer Ultra Low RNA Kit for Illumina Sequencing Two powerful technologies combine to enable sequencing with ultra-low levels of RNA The most sensitive cdna synthesis technology, combined with next-generation
SMARTer Ultra Low RNA Kit for Illumina Sequencing Two powerful technologies combine to enable sequencing with ultra-low levels of RNA The most sensitive cdna synthesis technology, combined with next-generation
TECH NOTE Stranded NGS libraries from FFPE samples
 TECH NOTE Stranded NGS libraries from FFPE samples Robust performance with extremely degraded FFPE RNA (DV 200 >25%) Consistent library quality across a range of input amounts (5 ng 50 ng) Compatibility
TECH NOTE Stranded NGS libraries from FFPE samples Robust performance with extremely degraded FFPE RNA (DV 200 >25%) Consistent library quality across a range of input amounts (5 ng 50 ng) Compatibility
Unique, dual-matched adapters mitigate index hopping between NGS samples. Kristina Giorda, PhD
 Unique, dual-matched adapters mitigate index hopping between NGS samples Kristina Giorda, PhD 1 Outline NGS workflow and cross-talk Sources of sample cross-talk and mitigation strategies Adapter recommendations
Unique, dual-matched adapters mitigate index hopping between NGS samples Kristina Giorda, PhD 1 Outline NGS workflow and cross-talk Sources of sample cross-talk and mitigation strategies Adapter recommendations
GENOMICS WORKFLOW SOLUTIONS THAT GO WHERE THE SCIENCE LEADS. Genomics Solutions Portfolio
 GENOMICS WORKFLOW SOLUTIONS THAT GO WHERE THE SCIENCE LEADS Genomics Solutions Portfolio WORKFLOW SOLUTIONS FROM EXTRACTION TO ANALYSIS Application-based answers for every step of your workflow Scientists
GENOMICS WORKFLOW SOLUTIONS THAT GO WHERE THE SCIENCE LEADS Genomics Solutions Portfolio WORKFLOW SOLUTIONS FROM EXTRACTION TO ANALYSIS Application-based answers for every step of your workflow Scientists
Product selection guide Ion GeneStudio S5 Series
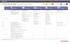 Cancer genomics research Molecular profiling Ion AmpliSeq Comprehensive Cancer Panel Cat. No. 4477685 Ion AmpliSeq Made-to-Order Panels (Customize your own or browse redesigned community panels at ampliseq.com)
Cancer genomics research Molecular profiling Ion AmpliSeq Comprehensive Cancer Panel Cat. No. 4477685 Ion AmpliSeq Made-to-Order Panels (Customize your own or browse redesigned community panels at ampliseq.com)
Automation of xgen hybridization capture on the Sciclone G3 NGS Workstation
 next generation sequencing protocol Automation of xgen hybridization capture on the Sciclone G3 NGS Workstation www.idtdna.com For Research Use Only Version 2 Revision history Document version Date released
next generation sequencing protocol Automation of xgen hybridization capture on the Sciclone G3 NGS Workstation www.idtdna.com For Research Use Only Version 2 Revision history Document version Date released
solid S Y S T E M s e q u e n c i n g See the Difference Discover the Quality Genome
 solid S Y S T E M s e q u e n c i n g See the Difference Discover the Quality Genome See the Difference With a commitment to your peace of mind, Life Technologies provides a portfolio of robust and scalable
solid S Y S T E M s e q u e n c i n g See the Difference Discover the Quality Genome See the Difference With a commitment to your peace of mind, Life Technologies provides a portfolio of robust and scalable
CM581A2: NEXT GENERATION SEQUENCING PLATFORMS AND LIBRARY GENERATION
 CM581A2: NEXT GENERATION SEQUENCING PLATFORMS AND LIBRARY GENERATION Fall 2015 Instructors: Coordinator: Carol Wilusz, Associate Professor MIP, CMB Instructor: Dan Sloan, Assistant Professor, Biology,
CM581A2: NEXT GENERATION SEQUENCING PLATFORMS AND LIBRARY GENERATION Fall 2015 Instructors: Coordinator: Carol Wilusz, Associate Professor MIP, CMB Instructor: Dan Sloan, Assistant Professor, Biology,
DNA Workflow. Marine Biological Laboratory. Mark Bratz Applications Scientist, Promega Corporation. August 2016
 DNA Workflow Marine Biological Laboratory Mark Bratz Applications Scientist, August 2016 Scientific Applications Support Mission Realize customer driven applications of Promega technologies to enhance
DNA Workflow Marine Biological Laboratory Mark Bratz Applications Scientist, August 2016 Scientific Applications Support Mission Realize customer driven applications of Promega technologies to enhance
Rapid, Accurate and Flexible DNA Quantitation Using the QuantiFluor dsdna System on the MANTIS Liquid Handler
 Rapid, Accurate and Flexible DNA Quantitation Using the QuantiFluor dsdna System on the MANTIS Liquid Handler Materials Required MANTIS Instrument (Formulatrix) High Volume Chips (Formulatrix Cat.# MCHS6)
Rapid, Accurate and Flexible DNA Quantitation Using the QuantiFluor dsdna System on the MANTIS Liquid Handler Materials Required MANTIS Instrument (Formulatrix) High Volume Chips (Formulatrix Cat.# MCHS6)
FFPE in your NGS Study
 FFPE in your NGS Study Richard Corbett Canada s Michael Smith Genome Sciences Centre Vancouver, British Columbia Dec 6, 2017 Our mandate is to advance knowledge about cancer and other diseases and to use
FFPE in your NGS Study Richard Corbett Canada s Michael Smith Genome Sciences Centre Vancouver, British Columbia Dec 6, 2017 Our mandate is to advance knowledge about cancer and other diseases and to use
resequencing storage SNP ncrna metagenomics private trio de novo exome ncrna RNA DNA bioinformatics RNA-seq comparative genomics
 RNA Sequencing T TM variation genetics validation SNP ncrna metagenomics private trio de novo exome mendelian ChIP-seq RNA DNA bioinformatics custom target high-throughput resequencing storage ncrna comparative
RNA Sequencing T TM variation genetics validation SNP ncrna metagenomics private trio de novo exome mendelian ChIP-seq RNA DNA bioinformatics custom target high-throughput resequencing storage ncrna comparative
HyperCap, an automatable workflow on the Agilent Bravo B
 Automation Note February 2018 HyperCap, an automatable workflow on the Agilent Bravo B 1. OVERVIEW As the demand for next-generation sequencing (NGS) grows, laboratories must adapt to manage increased
Automation Note February 2018 HyperCap, an automatable workflow on the Agilent Bravo B 1. OVERVIEW As the demand for next-generation sequencing (NGS) grows, laboratories must adapt to manage increased
Reverse Transcription & RT-PCR
 Creating Gene Expression Solutions Reverse Transcription & RT-PCR Reverse transcription, a process that involves a reverse transcriptase (RTase) which uses RNA as the template to make complementary DNA
Creating Gene Expression Solutions Reverse Transcription & RT-PCR Reverse transcription, a process that involves a reverse transcriptase (RTase) which uses RNA as the template to make complementary DNA
QIAGEN Whole Genome Amplification REPLI-g Eliminating Sample Limitations, Potential Use for Reference Material
 QIAGEN Whole Genome Amplification REPLI-g Eliminating Sample Limitations, Potential Use for Reference Material MDA Technology Protocols Locus Bias Applications Kits & Service Summary QIAGEN REPLI-g WGA
QIAGEN Whole Genome Amplification REPLI-g Eliminating Sample Limitations, Potential Use for Reference Material MDA Technology Protocols Locus Bias Applications Kits & Service Summary QIAGEN REPLI-g WGA
TECHNICAL BULLETIN. SeqPlex DNA Amplification Kit for use with high throughput sequencing technologies. Catalog Number SEQX Storage Temperature 20 C
 SeqPlex DNA Amplification Kit for use with high throughput sequencing technologies Catalog Number SEQX Storage Temperature 20 C TECHNICAL BULLETIN Product Description The SeqPlex DNA Amplification Kit
SeqPlex DNA Amplification Kit for use with high throughput sequencing technologies Catalog Number SEQX Storage Temperature 20 C TECHNICAL BULLETIN Product Description The SeqPlex DNA Amplification Kit
Genome Resequencing. Rearrangements. SNPs, Indels CNVs. De novo genome Sequencing. Metagenomics. Exome Sequencing. RNA-seq Gene Expression
 Genome Resequencing De novo genome Sequencing SNPs, Indels CNVs Rearrangements Metagenomics RNA-seq Gene Expression Splice Isoform Abundance High Throughput Short Read Sequencing: Illumina Exome Sequencing
Genome Resequencing De novo genome Sequencing SNPs, Indels CNVs Rearrangements Metagenomics RNA-seq Gene Expression Splice Isoform Abundance High Throughput Short Read Sequencing: Illumina Exome Sequencing
NGS-based innovations within the Leiden Network
 NGS-based innovations within the Leiden Network A strong bridge between two partners Dr. Mark de Jong 2017-09-29 Design accurate and robust NGS tests and generate data sets essential for Diagnostics &
NGS-based innovations within the Leiden Network A strong bridge between two partners Dr. Mark de Jong 2017-09-29 Design accurate and robust NGS tests and generate data sets essential for Diagnostics &
It s All in the Details (or Small RNA): Simplified and Improved mirna Purification from Tissue
 It s All in the Details (or Small RNA): Simplified and Improved mirna Purification from Tissue Douglas Horejsh January 2015 Discussion Outline Non-coding RNA General Overview mirna What is it? The Future
It s All in the Details (or Small RNA): Simplified and Improved mirna Purification from Tissue Douglas Horejsh January 2015 Discussion Outline Non-coding RNA General Overview mirna What is it? The Future
TaqMan Advanced mirna Assays
 PRODUCT BULLETIN TaqMan Advanced mirna Assays TaqMan Advanced mirna Assays Key features Universal reverse transcription (RT) one RT step for all Applied Biosystems TaqMan Advanced mirna Assays Sensitive
PRODUCT BULLETIN TaqMan Advanced mirna Assays TaqMan Advanced mirna Assays Key features Universal reverse transcription (RT) one RT step for all Applied Biosystems TaqMan Advanced mirna Assays Sensitive
Implementation of Automated Sample Quality Control in Whole Exome Sequencing
 Journal of Life Sciences 11 (2017) 261-268 doi: 10.17265/1934-7391/2017.06.001 D DAVID PUBLISHING Implementation of Automated Sample Quality Control in Whole Exome Sequencing Elisa Viering 1, Jana Molitor
Journal of Life Sciences 11 (2017) 261-268 doi: 10.17265/1934-7391/2017.06.001 D DAVID PUBLISHING Implementation of Automated Sample Quality Control in Whole Exome Sequencing Elisa Viering 1, Jana Molitor
Enterprise Interest I am an employee of ThermoFisher Scientific.
 Enterprise Interest I am an employee of ThermoFisher Scientific. Developing a multiplex next-generation sequencing assay to study highly clonal tumor samples Dumitru Brinza, Ph.D Clinical Next-Generation
Enterprise Interest I am an employee of ThermoFisher Scientific. Developing a multiplex next-generation sequencing assay to study highly clonal tumor samples Dumitru Brinza, Ph.D Clinical Next-Generation
Supplementary Information for:
 Supplementary Information for: A streamlined and high-throughput targeting approach for human germline and cancer genomes using Oligonucleotide-Selective Sequencing Samuel Myllykangas 1, Jason D. Buenrostro
Supplementary Information for: A streamlined and high-throughput targeting approach for human germline and cancer genomes using Oligonucleotide-Selective Sequencing Samuel Myllykangas 1, Jason D. Buenrostro
Functional DNA Quality Analysis Improves the Accuracy of Next Generation Sequencing from Clinical Specimens
 Functional DNA Quality Analysis Improves the Accuracy of Next Generation Sequencing from Clinical Specimens Overview We have developed a novel QC, the SuraSeq DNA Quantitative Functional Index (QFI ).
Functional DNA Quality Analysis Improves the Accuracy of Next Generation Sequencing from Clinical Specimens Overview We have developed a novel QC, the SuraSeq DNA Quantitative Functional Index (QFI ).
sparq HiFi PCR Master Mix
 sparq HiFi PCR Master Mix Cat. No. 95192-050 Size: 50 reactions Store at -25 C to -15 C 95192-250 250 reactions Description The sparq HiFi PCR Master Mix is a high efficiency, high-fidelity, and low bias
sparq HiFi PCR Master Mix Cat. No. 95192-050 Size: 50 reactions Store at -25 C to -15 C 95192-250 250 reactions Description The sparq HiFi PCR Master Mix is a high efficiency, high-fidelity, and low bias
The New Genome Analyzer IIx Delivering more data, faster, and easier than ever before. Jeremy Preston, PhD Marketing Manager, Sequencing
 The New Genome Analyzer IIx Delivering more data, faster, and easier than ever before Jeremy Preston, PhD Marketing Manager, Sequencing Illumina Genome Analyzer: a Paradigm Shift 2000x gain in efficiency
The New Genome Analyzer IIx Delivering more data, faster, and easier than ever before Jeremy Preston, PhD Marketing Manager, Sequencing Illumina Genome Analyzer: a Paradigm Shift 2000x gain in efficiency
Low input RNA-seq library preparation provides higher small non-coding RNA diversity and greatly reduced hands-on time
 TECHNICAL NOTE Low input RNA-seq library preparation provides higher small non-coding RNA diversity and greatly reduced hands-on time INTRODUCTION RNA-seq is a next-generation sequencing technique that
TECHNICAL NOTE Low input RNA-seq library preparation provides higher small non-coding RNA diversity and greatly reduced hands-on time INTRODUCTION RNA-seq is a next-generation sequencing technique that
NEBNext. for Ion Torrent LIBRARY PREPARATION KITS
 NEBNext for Ion Torrent LIBRARY PREPARATION KITS NEBNEXT PRODUCTS FOR ION TORRENT Table of Contents 3 General Introduction 4 5 6 6 7 8 DNA Library Preparation Workflow Product Selection Product Details
NEBNext for Ion Torrent LIBRARY PREPARATION KITS NEBNEXT PRODUCTS FOR ION TORRENT Table of Contents 3 General Introduction 4 5 6 6 7 8 DNA Library Preparation Workflow Product Selection Product Details
Considerations for Illumina library preparation. Henriette O Geen June 20, 2014 UCD Genome Center
 Considerations for Illumina library preparation Henriette O Geen June 20, 2014 UCD Genome Center Diversity of applications De novo genome Sequencing ranscriptome Expression Splice Isoform bundance Genotyping
Considerations for Illumina library preparation Henriette O Geen June 20, 2014 UCD Genome Center Diversity of applications De novo genome Sequencing ranscriptome Expression Splice Isoform bundance Genotyping
APPLICATION NOTE
 APPLICATION NOTE www.swiftbiosci.com Approaching Single-Cell Sequencing by Understanding NGS Library Complexity and Bias Abstract Demands are growing on genomics to deliver higher quality sequencing data
APPLICATION NOTE www.swiftbiosci.com Approaching Single-Cell Sequencing by Understanding NGS Library Complexity and Bias Abstract Demands are growing on genomics to deliver higher quality sequencing data
Access Array BRCA1 / BRCA2 / TP53 Target-Specific Panel Build the highest quality amplicon libraries with qualified assays
 DATA SHEET PN 100-3489 B1 Access Array BRCA1 / BRCA2 / TP53 Target-Specific Panel Build the highest quality amplicon libraries with qualified assays Covers 100% of the exons within the genes Supported
DATA SHEET PN 100-3489 B1 Access Array BRCA1 / BRCA2 / TP53 Target-Specific Panel Build the highest quality amplicon libraries with qualified assays Covers 100% of the exons within the genes Supported
Automation of the IDT xgen Lockdown Panels on the Sciclone G3 NGS Workstation
 A PPLI C A TION NOT E NGS Library Preparation Sciclone G3 NGSx Workstation Automation of the IDT xgen Lockdown Panels on the Sciclone G3 NGS Workstation Introduction Integrated DNA Technologies (IDT) xgen
A PPLI C A TION NOT E NGS Library Preparation Sciclone G3 NGSx Workstation Automation of the IDT xgen Lockdown Panels on the Sciclone G3 NGS Workstation Introduction Integrated DNA Technologies (IDT) xgen
TREE CODE PRODUCT BROCHURE
 TREE CODE PRODUCT BROCHURE Single Molecule, Real-Time (SMRT) Sequencing technology offers: Long read sequencing ~10 Gb with 20 kb average read lengths for WGS ~20 Gb with 40 kb average read length for
TREE CODE PRODUCT BROCHURE Single Molecule, Real-Time (SMRT) Sequencing technology offers: Long read sequencing ~10 Gb with 20 kb average read lengths for WGS ~20 Gb with 40 kb average read length for
Admera Health RUO Services
 Admera Health RUO Services Services 1. RNA-Seq... 2 2. Single-cell RNAseq (scrna-seq)... 3 3. Whole Exome Sequencing (WES)... 4 4. Whole Genome Sequencing (WGS)... 5 5. Whole Genome Bisulfite Sequencing
Admera Health RUO Services Services 1. RNA-Seq... 2 2. Single-cell RNAseq (scrna-seq)... 3 3. Whole Exome Sequencing (WES)... 4 4. Whole Genome Sequencing (WGS)... 5 5. Whole Genome Bisulfite Sequencing
