Eukaryotic Transcription
|
|
|
- Beatrice Knight
- 6 years ago
- Views:
Transcription
1 Eukaryotic Transcription I. Differences between eukaryotic versus prokaryotic transcription. II. (core vs holoenzyme): RNA polymerase II - Promotor elements. - General Pol II transcription factors (GTF). - TBP and TAFs. - Mediator / SRB complex. - Swi/Snf and SAGA complex. III. Regulation of RNA pol II transcription. - Transcription initiation (yeast Gal4 system) - Activators, repressor, enhancer and silencer. - Transcription elongation (HIV Tat-tar system). IV: Chromatin remodeling during transcription. - Role of Swi/Snf complex and TAFs.
2 I. Eukaryotes vs prokaryotes transcription. Eukaryotes Most mrnas are monocistronic (generally one gene for one protein). Transcription and translation of most eukaryotes are spatially separated. Multiple RNA polyermases. Transcription factor directly binds to the DNA template, but not to the RNA pol. Requires ATP for initiation. Nucleosome / chromatin. Prokaryotes Polycistronic mrna. Coupling transcription and translation. Single RNA polyermase RNA pol holoenzyme itself, by associated sigma factor. No ATP Naked DNA I. Eukaryotes vs prokaryotes transcription.
3 RNA polymerase II is capable of transcribing DNA templates, but it will not begin transcription at the correct site. Hence it is competent for elongation but not initiation.?????? Hypothesis: There must be a cis-acting DNA sequence and trans-acting factor that recruits or direct RNA Pol II at the site of transcription initiation. enhancer upstream regulatory elements Start of Transcription core (-200) (-45) (+40) core promoter Core Promoter : Minimal region required for basal transcription. Regulatory Elements: Upstream regulatory sequence (Proximal element) - usually within a few hundred bases of the promoter. Enhancer sequence (Distal element) that functions at a distance (even downstream or within the transcription unit) and orientation independent manner.
4 Identifying regulatory elements: - Systematically mutate DNA and connect to a convenient reporter gene. luciferase chloramphenical acetyl transferase (CAT) ß-galactosidase. - Transfect DNA back into a suitable eucaryotic cell. - Measure reporter gene activity. Caveats: Cell lines are usually immortal so they carry mutations in cell cycle control. Response may be cell-type specific - specific factors or combination of factors may be absent. Chromatin structure may be abnormal. If DNA has randomly inserted into the genome, neighboring sequences can affect transcription. enhancer upstream regulatory elements Start of Transcription core (-200) (-45) (+40) core promoter Start of Transcription (-37 to -32) (-31 to -26) (-2 to +4) (+28 to +32) BRE TATTA Int DPE They are essential for the assembly of the general transcription factors and Pol II complex and the initiation of transcription, and therefore are required for directing where transcription starts. The Inr element encompasses the transcription start site. Found in both TATA-containing as well as TATA-less core promoters. DPE is found most commonly in TATA-less promoters. TFIIB is able to bind directly to the BRE in a sequence-specific manner.
5 enhancer upstream regulatory elements Start of Transcription core (-200) (-45) (+40) core promoter Start of Transcription (-37 to -32) (-31 to -26) (-2 to +4) (+28 to +32) BRE TATTA Int DPE In promoters with TATA box -initiation occurs ~35 bp downstream. Often more than one start site Start site variability greater in S. cerevisiae (40-100bp)
6 enhancer upstream regulatory elements Start of Transcription core (-200) (-45) (+40) Proximal element: (upstream regulatory sequence ): core promoter Relatively GC-rich. Allows binding of Sp1 and related transcription factors (activator or repressor. Often present as multiple copies and can act synergistically. Upstream regulatory sequence (Proximal element):
7 enhancer upstream regulatory elements Start of Transcription core (-200) (-45) (+40) core promoter Distal element: (enhancer / silencer): Facilitate assembly of pre-initiation complex, directly or indirectly. Enhancer: function as binding sites for specific transcription activator (often found in developmentally regulated genes). Function in either orientation at variable distance (0.5 kb - 10 kb). In mammalian system can be positioned downstream of TATA. Silencer also binds to distal element and function as transcriptional repressor. Three distinct DNA-dependent RNA polymerases in the nucleus: RNA pol I, II, and III. Two additional RNA polymerases in organelles: one in mitochondria and one in chloroplasts. Three nuclear RNA Pols were first identified as distinct polymerases because they elute from columns at different salt concentrations and can be classified by the sensitivity to alpha-aminitin (chemical inhibitor of Pol II transcripton).
8
9 Pol I Approx % of total RNA synthesized the precursors for the 28S, 18S, and 5.8S ribosomal RNAs. Regulation is minimal and synthesize stable RNA. Pol II (10-15% of total) typically synthesized mrnas that are translated into proteins. Also some snrnp RNAs involved in mrna splicing. Regulation is diverse and transcripts are less stable. Pol III (5-10% of total) synthesize small RNAs, such as trnas, 5S RNA, some of the other spliceosome RNAs, such as U6 and H1. Regulation is minimal and synthesize stable RNA. None of the subunits appreas to be related to the bacterial sigma-subunit
10 Bushnell, Cramer, Kornberg, Proc Natl Acad Sci USA 99 (2002) p.1218
11 Tandem repeat heptad sequence: ( Tyr Ser Pro Thr Ser Pro Ser ) n Mouse (n = 52) Budding / Fission yeast (n = 26 / 29) Plasmodium (n = 15)
12 CTD of RNA polymerase II Two forms of RNA Pol II: Pol II A: not phosphorylated and preferentially enters the pre-initiation complex. pol IIO: physophorylated at Ser residue and is found in elongating complex. Conversion of IIA to IIO occurs shortly after the transition from initiation to elongation (promoter clearance. RNA pol II lacking the CTD is able to initate transcription in vitro but not from TATA-less promoter. Hypothesis: There must be a cis-acting DNA sequence and trans-acting factor that recruits or direct RNA Pol II at the site of transcription initiation. Approach:??? Biochemically isolate factors that are required for promotor specific transcription (mammalian and viral system). Genetically identify mutants and suppressor genes involved in transcription (yeast and flys).
13 Assay system to isolate transcription factor biochemically. Assay system to isolate transcription factor. G-less Cassette p(c2at) AdML G-less Cassette pml(c2at) - DNA template - RNA polymerase (purified separately) - NTP (ATP, UTP, 32P-CTP and GTP*) - TFIIs (nuclear extract or column fractions) (1) p(c2at) - GTP (2) pml(c2at) - GTP (3) p(c2at) (4) pml(c2at) (5) p(c2at) + RNaseT1 + 2 O-mGTP (6) pml(c2at) + RNaseT1 + 2 O-mGTP Accurate and specific transcription from promoter containing template. Specific transcription can be quantitate by direct measurement of total RNA synthesis.
14 Assay system to isolate transcription factor. Assembly of GTF at the promoter in vitro: Gel mobility shift assay.
15 Assembly of GTF at the promoter in vitro: Gel mobility shift assay. TFIIA Assembly of GTF at the promoter in vitro: Gel mobility shift assay. Purified RNA Pol II, TFII D, A, B, or E do not bind stably to promoter when added individually (lane 1-5) When all the individual components were added (RNA Pol II, TFII D, A, B, ande)formed6complex(lane6) TFIIA is not necessary, but TFIID is absolutely required for complex formation (lane 7, 8 & 12). Lack of TFIIB or RNA Pol II abolished formation of slow migrating complex (lane 9 & 10). This suggest that TFIIB may involved in recruitment of pol II. Note that addition of TFIIB can form complex3intheabsenceofpolii. Omission of TFIIE prevented formation of complex 6 & 7 (compare lanes and lane 6-8).
16 Assembly of GTF at the promoter in vitro:dnase 1 Footprint. Coding Strand Non-Coding Strand Complex2:TFIIA,-D. - Protection at -42 to Complex3:TFIIA,-D,-B. - Similar to complex 2 pattern with additional protection -10 to +10 region. Complex 4 and 5: TFFIIA, -D, -B, pol II. - Same upstream boundary (-42) but cleavage downstream was strongly suppressed (up to +20). Complex 6 and 7: TFIIA, -D, -B, pol II and - E. - Enhance cleavage downstream of the promoter (to about +30 on coding strand). (1 subunit) TFIIE TFIIF TFIIH TBP (2 subunits) (9 subunits) (1 subunit) (2 subunits) TAFs (12 subunits)
17 TFIIE: TFIIEiscomposedtof2subunits(56and 34 kda). TFIIF binds prior to TFIIE (lane 4). Requires both TFIIE subunit to form a complex (lane 5-8) - negative result!. Recombinant TFIIE was capable of forming a complex similar to purified TFIIE (lane 4 and 10). Omission of TFIID prevented formation of complex. TFIIH The only GTF with known enzymatic activity: DNA-dependent ATPase ATP-dependent helicase CTD kinase MostcomplexGTFwithatotalmassof 500 kda (~ size of RNA pol II). Converts RNA pol IIA to pol II O (phosphorylation) at CTD Ser 5. TFIIH alters the mobility of the PIC complex in the presence of ATP
18 TFIIH (helicase / ATPase activity appears to act at the promoter clearance step, a transition stage between initiation and elongation. General Transcription Factor Factor Subunits Function TFIID TBP 1 Core promoter recognition (TATA); Recruit TFIIB. TAFs 12 Core promoter recognition (non-tata); positive and negative regulatory function. TFIIA 3 Stabilize TBP binding: stabilization of TAF- DNA interaction; anti-repressor functions. TFIIB 1 pol II-TFIIF recruitment; start-site selection by pol II. TFIIF(w/Pol II) 2 Promoter targeting of pol II; destabilization of non-specific pol II-DNA interactions. TFIIE 2 TFIIH recruitment; modulation of TFIIH helicase, ATPase and kinase activity; enhancement of promoter melting(?). TFIIH 9 Promoter clearance by CTD kinase activity. Promoter melting by helicase activity(?).
19 TATA binding protein: TBP : TATA binding protein. The C-terminal 180 amino acid of TBP is the TATA-binding domain. TBP binds and grossly distorts the minor groove of TATA box
Eukaryotic & Prokaryotic Transcription. RNA polymerases
 Eukaryotic & Prokaryotic Transcription RNA polymerases RNA Polymerases A. E. coli RNA polymerase 1. core enzyme = ββ'(α)2 has catalytic activity but cannot recognize start site of transcription ~500,000
Eukaryotic & Prokaryotic Transcription RNA polymerases RNA Polymerases A. E. coli RNA polymerase 1. core enzyme = ββ'(α)2 has catalytic activity but cannot recognize start site of transcription ~500,000
Chromatographic Separation of the three forms of RNA Polymerase II.
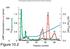 Chromatographic Separation of the three forms of RNA Polymerase II. α-amanitin α-amanitin bound to Pol II Function of the three enzymes. Yeast Pol II. RNA Polymerase Subunit Structures 10-7 Subunit structure.
Chromatographic Separation of the three forms of RNA Polymerase II. α-amanitin α-amanitin bound to Pol II Function of the three enzymes. Yeast Pol II. RNA Polymerase Subunit Structures 10-7 Subunit structure.
30 Gene expression: Transcription
 30 Gene expression: Transcription Gene structure. o Exons coding region of DNA. o Introns non-coding region of DNA. o Introns are interspersed between exons of a single gene. o Promoter region helps enzymes
30 Gene expression: Transcription Gene structure. o Exons coding region of DNA. o Introns non-coding region of DNA. o Introns are interspersed between exons of a single gene. o Promoter region helps enzymes
Mechanisms of Transcription. School of Life Science Shandong University
 Mechanisms of Transcription School of Life Science Shandong University Ch 12: Mechanisms of Transcription 1. RNA polymerase and the transcription cycle 2. The transcription cycle in bacteria 3. Transcription
Mechanisms of Transcription School of Life Science Shandong University Ch 12: Mechanisms of Transcription 1. RNA polymerase and the transcription cycle 2. The transcription cycle in bacteria 3. Transcription
Chapter 3. DNA, RNA, and Protein Synthesis
 Chapter 3. DNA, RNA, and Protein Synthesis 4. Transcription Gene Expression Regulatory region (promoter) 5 flanking region Upstream region Coding region 3 flanking region Downstream region Transcription
Chapter 3. DNA, RNA, and Protein Synthesis 4. Transcription Gene Expression Regulatory region (promoter) 5 flanking region Upstream region Coding region 3 flanking region Downstream region Transcription
The RNA Polymerase II General Transcription Machinery Prof. Michael Hampsey
 The RNA Polymerase II General Transcription Machinery Michael Hampsey, PhD Robert Wood Johnson Medical School Piscataway, New Jersey 1 Central Dogma of Molecular Biology DNA Stores genetic information
The RNA Polymerase II General Transcription Machinery Michael Hampsey, PhD Robert Wood Johnson Medical School Piscataway, New Jersey 1 Central Dogma of Molecular Biology DNA Stores genetic information
Transcription Eukaryotic Cells
 Transcription Eukaryotic Cells Packet #20 1 Introduction Transcription is the process in which genetic information, stored in a strand of DNA (gene), is copied into a strand of RNA. Protein-encoding genes
Transcription Eukaryotic Cells Packet #20 1 Introduction Transcription is the process in which genetic information, stored in a strand of DNA (gene), is copied into a strand of RNA. Protein-encoding genes
TRANSCRIPTION AND PROCESSING OF RNA
 TRANSCRIPTION AND PROCESSING OF RNA 1. The steps of gene expression. 2. General characterization of transcription: steps, components of transcription apparatus. 3. Transcription of eukaryotic structural
TRANSCRIPTION AND PROCESSING OF RNA 1. The steps of gene expression. 2. General characterization of transcription: steps, components of transcription apparatus. 3. Transcription of eukaryotic structural
SIBC504: TRANSCRIPTION & RNA PROCESSING Assistant Professor Dr. Chatchawan Srisawat
 SIBC504: TRANSCRIPTION & RNA PROCESSING Assistant Professor Dr. Chatchawan Srisawat TRANSCRIPTION: AN OVERVIEW Transcription: the synthesis of a single-stranded RNA from a doublestranded DNA template.
SIBC504: TRANSCRIPTION & RNA PROCESSING Assistant Professor Dr. Chatchawan Srisawat TRANSCRIPTION: AN OVERVIEW Transcription: the synthesis of a single-stranded RNA from a doublestranded DNA template.
DNA Transcription. Dr Aliwaini
 DNA Transcription 1 DNA Transcription-Introduction The synthesis of an RNA molecule from DNA is called Transcription. All eukaryotic cells have five major classes of RNA: ribosomal RNA (rrna), messenger
DNA Transcription 1 DNA Transcription-Introduction The synthesis of an RNA molecule from DNA is called Transcription. All eukaryotic cells have five major classes of RNA: ribosomal RNA (rrna), messenger
Biochemistry Eukaryotic Transcription
 1 Description of Module Subject Name Paper Name Module Name/Title Dr. Vijaya Khader Dr. MC Varadaraj 2 1. Objectives 1. Understand and have an overview of eucaryotic transcriptional regulation. 2. Explain
1 Description of Module Subject Name Paper Name Module Name/Title Dr. Vijaya Khader Dr. MC Varadaraj 2 1. Objectives 1. Understand and have an overview of eucaryotic transcriptional regulation. 2. Explain
Differential Gene Expression
 Biology 4361 Developmental Biology Differential Gene Expression September 28, 2006 Chromatin Structure ~140 bp ~60 bp Transcriptional Regulation: 1. Packing prevents access CH 3 2. Acetylation ( C O )
Biology 4361 Developmental Biology Differential Gene Expression September 28, 2006 Chromatin Structure ~140 bp ~60 bp Transcriptional Regulation: 1. Packing prevents access CH 3 2. Acetylation ( C O )
Lecture 21 Regulation of transcription
 Lecture 21 Regulation of transcription Key learning goals: Understand what a promoter sequence is Understand the role of sigma factors in prokaryotic RNAP initiation Understand what an open complex is,
Lecture 21 Regulation of transcription Key learning goals: Understand what a promoter sequence is Understand the role of sigma factors in prokaryotic RNAP initiation Understand what an open complex is,
Chapter 17 Lecture. Concepts of Genetics. Tenth Edition. Regulation of Gene Expression in Eukaryotes
 Chapter 17 Lecture Concepts of Genetics Tenth Edition Regulation of Gene Expression in Eukaryotes Chapter Contents 17.1 Eukaryotic Gene Regulation Can Occur at Any of the Steps Leading from DNA to Protein
Chapter 17 Lecture Concepts of Genetics Tenth Edition Regulation of Gene Expression in Eukaryotes Chapter Contents 17.1 Eukaryotic Gene Regulation Can Occur at Any of the Steps Leading from DNA to Protein
GENETICS - CLUTCH CH.10 TRANSCRIPTION.
 !! www.clutchprep.com CONCEPT: OVERVIEW OF TRANSCRIPTION Transcription is the process of using DNA as a template to RNA RNA polymerase is the enzyme that transcribes DNA - There are many different types
!! www.clutchprep.com CONCEPT: OVERVIEW OF TRANSCRIPTION Transcription is the process of using DNA as a template to RNA RNA polymerase is the enzyme that transcribes DNA - There are many different types
Transcription Regulation And Gene Expression in Eukaryotes FS 2016 Graduate Course G2 P Matthias and RG Clerc Pharmazentrum Hörsaal 2 16h15-18h00
 Transcription Regulation And Gene Expression in Eukaryotes FS 2016 Graduate Course G2 P Matthias and RG Clerc Pharmazentrum Hörsaal 2 16h15-18h00 The general problem RG Clerc March 2, 2016 RNA Transcription
Transcription Regulation And Gene Expression in Eukaryotes FS 2016 Graduate Course G2 P Matthias and RG Clerc Pharmazentrum Hörsaal 2 16h15-18h00 The general problem RG Clerc March 2, 2016 RNA Transcription
Transcription. Manzur Ali PP, DBT,M.E.S College,Marampally
 Transcription Manzur Ali PP, DBT,M.E.S College,Marampally manzursir@gmail.com RNA transcription is actively regulated Not all DNA is transcribed in a given cell (less than 50% even in prokaryotes) For
Transcription Manzur Ali PP, DBT,M.E.S College,Marampally manzursir@gmail.com RNA transcription is actively regulated Not all DNA is transcribed in a given cell (less than 50% even in prokaryotes) For
Differential Gene Expression
 Biology 4361 Developmental Biology Differential Gene Expression June 19, 2008 Differential Gene Expression Overview Chromatin structure Gene anatomy RNA processing and protein production Initiating transcription:
Biology 4361 Developmental Biology Differential Gene Expression June 19, 2008 Differential Gene Expression Overview Chromatin structure Gene anatomy RNA processing and protein production Initiating transcription:
DNA Prokaryote Transcription Steps (updated February 2013)
 URS AACTGT ATATTA - 35-10 transcription Pribnow Box discriminator +1 AGGAGGT TTA TCCTCCA ATT Gene C TGA TAG ACT ATC rho or GC hairpin loop transcription termination DNA Prokaryote Transcription Steps (updated
URS AACTGT ATATTA - 35-10 transcription Pribnow Box discriminator +1 AGGAGGT TTA TCCTCCA ATT Gene C TGA TAG ACT ATC rho or GC hairpin loop transcription termination DNA Prokaryote Transcription Steps (updated
M1 - Biochemistry. Nucleic Acid Structure II/Transcription I
 M1 - Biochemistry Nucleic Acid Structure II/Transcription I PH Ratz, PhD (Resources: Lehninger et al., 5th ed., Chapters 8, 24 & 26) 1 Nucleic Acid Structure II/Transcription I Learning Objectives: 1.
M1 - Biochemistry Nucleic Acid Structure II/Transcription I PH Ratz, PhD (Resources: Lehninger et al., 5th ed., Chapters 8, 24 & 26) 1 Nucleic Acid Structure II/Transcription I Learning Objectives: 1.
Molecular Biology (BIOL 4320) Exam #1 March 12, 2002
 Molecular Biology (BIOL 4320) Exam #1 March 12, 2002 Name KEY SS# This exam is worth a total of 100 points. The number of points each question is worth is shown in parentheses after the question number.
Molecular Biology (BIOL 4320) Exam #1 March 12, 2002 Name KEY SS# This exam is worth a total of 100 points. The number of points each question is worth is shown in parentheses after the question number.
We can now identify three major pathways of information flow in the cell (in replication, information passes from one DNA molecule to other DNA
 1 We can now identify three major pathways of information flow in the cell (in replication, information passes from one DNA molecule to other DNA molecules; in transcription, information passes from DNA
1 We can now identify three major pathways of information flow in the cell (in replication, information passes from one DNA molecule to other DNA molecules; in transcription, information passes from DNA
Computational Biology I LSM5191 (2003/4)
 Computational Biology I LSM5191 (2003/4) Aylwin Ng, D.Phil Lecture Notes: Transcriptome: Molecular Biology of Gene Expression I Flow of information: DNA to polypeptide DNA Start Exon1 Intron Exon2 Termination
Computational Biology I LSM5191 (2003/4) Aylwin Ng, D.Phil Lecture Notes: Transcriptome: Molecular Biology of Gene Expression I Flow of information: DNA to polypeptide DNA Start Exon1 Intron Exon2 Termination
Complex Transcription Machinery
 Complex Transcription Machinery Subunits of the Basal Txn Apparatus Stepwise Assembly of the Pre-initiation Complex Interplay of Activators, Co-regulators and RNA Polymerase at the Promoter Divide and
Complex Transcription Machinery Subunits of the Basal Txn Apparatus Stepwise Assembly of the Pre-initiation Complex Interplay of Activators, Co-regulators and RNA Polymerase at the Promoter Divide and
Transcription in Eukaryotes
 Transcription in Eukaryotes Biology I Hayder A Giha Transcription Transcription is a DNA-directed synthesis of RNA, which is the first step in gene expression. Gene expression, is transformation of the
Transcription in Eukaryotes Biology I Hayder A Giha Transcription Transcription is a DNA-directed synthesis of RNA, which is the first step in gene expression. Gene expression, is transformation of the
However, only a fraction of these genes are transcribed in an individual cell at any given time.
 All cells in an organism contain the same set of genes. However, only a fraction of these genes are transcribed in an individual cell at any given time. It is the pattern of gene expression that determines
All cells in an organism contain the same set of genes. However, only a fraction of these genes are transcribed in an individual cell at any given time. It is the pattern of gene expression that determines
Chapter 11. Transcription. The biochemistry and molecular biology department of CMU
 Chapter 11 Transcription The biochemistry and molecular biology department of CMU Transcription The synthesis of RNA molecules using DNA strands as the templates so that the genetic information can be
Chapter 11 Transcription The biochemistry and molecular biology department of CMU Transcription The synthesis of RNA molecules using DNA strands as the templates so that the genetic information can be
DNA Evolution of knowledge about gene. Contains information about RNAs and proteins. Polynucleotide chains; Double stranded molecule;
 Evolution of knowledge about gene G. Mendel Hereditary factors W.Johannsen, 1909 G.W.Beadle, E.L.Tatum, 1945 Ingram, 1957 Actual concepts The gene hereditary unit located in chromosomes Hypotheses One
Evolution of knowledge about gene G. Mendel Hereditary factors W.Johannsen, 1909 G.W.Beadle, E.L.Tatum, 1945 Ingram, 1957 Actual concepts The gene hereditary unit located in chromosomes Hypotheses One
Resources. This lecture Campbell and Farrell's Biochemistry, Chapter 11
 Transcription Resources This lecture Campbell and Farrell's Biochemistry, Chapter 11 2 Definition of a gene The entire nucleic acid sequence that is necessary for the synthesis of a functional polypeptide
Transcription Resources This lecture Campbell and Farrell's Biochemistry, Chapter 11 2 Definition of a gene The entire nucleic acid sequence that is necessary for the synthesis of a functional polypeptide
Differential Gene Expression
 Biology 4361 - Developmental Biology Differential Gene Expression June 18, 2009 Differential Gene Expression Overview Chromatin structure Gene anatomy RNA processing and protein production Initiating transcription:
Biology 4361 - Developmental Biology Differential Gene Expression June 18, 2009 Differential Gene Expression Overview Chromatin structure Gene anatomy RNA processing and protein production Initiating transcription:
Initiation and termination of transcription, Post transcription modification of the RNA. Mitesh Shrestha
 Initiation and termination of transcription, Post transcription modification of the RNA Mitesh Shrestha Transcription: overview In prokaryotes transcription and translation are coupled. Proteins are synthesized
Initiation and termination of transcription, Post transcription modification of the RNA Mitesh Shrestha Transcription: overview In prokaryotes transcription and translation are coupled. Proteins are synthesized
Francisco Navarro. Departamento de Biología Experimental Área de Genética Universidad de Jaén
 Regulación de la expresión génica Francisco Navarro Departamento de Biología Experimental Área de Genética Universidad de Jaén Gene expression From genes to proteins Gene expression From genes to proteins
Regulación de la expresión génica Francisco Navarro Departamento de Biología Experimental Área de Genética Universidad de Jaén Gene expression From genes to proteins Gene expression From genes to proteins
Differential Gene Expression
 Developmental Biology Biology 4361 Differential Gene Expression October 13, 2005 core transcription initiation site 5 promoter 3 TATAT +1 upstream downstream Basal transcription factors (eukaryotes) TFIID
Developmental Biology Biology 4361 Differential Gene Expression October 13, 2005 core transcription initiation site 5 promoter 3 TATAT +1 upstream downstream Basal transcription factors (eukaryotes) TFIID
BS 50 Genetics and Genomics Week of Oct 24
 BS 50 Genetics and Genomics Week of Oct 24 Additional Practice Problems for Section Question 1: The following table contains a list of statements that apply to replication, transcription, both, or neither.
BS 50 Genetics and Genomics Week of Oct 24 Additional Practice Problems for Section Question 1: The following table contains a list of statements that apply to replication, transcription, both, or neither.
Transcription & post transcriptional modification
 Transcription & post transcriptional modification Transcription The synthesis of RNA molecules using DNA strands as the templates so that the genetic information can be transferred from DNA to RNA Similarity
Transcription & post transcriptional modification Transcription The synthesis of RNA molecules using DNA strands as the templates so that the genetic information can be transferred from DNA to RNA Similarity
Chapter 24: Promoters and Enhancers
 Chapter 24: Promoters and Enhancers A typical gene transcribed by RNA polymerase II has a promoter that usually extends upstream from the site where transcription is initiated the (#1) of transcription
Chapter 24: Promoters and Enhancers A typical gene transcribed by RNA polymerase II has a promoter that usually extends upstream from the site where transcription is initiated the (#1) of transcription
CLASS 3.5: 03/29/07 EUKARYOTIC TRANSCRIPTION I: PROMOTERS AND ENHANCERS
 CLASS 3.5: 03/29/07 EUKARYOTIC TRANSCRIPTION I: PROMOTERS AND ENHANCERS A. Promoters and Polymerases (RNA pols): 1. General characteristics - Initiation of transcription requires a. Transcription factors
CLASS 3.5: 03/29/07 EUKARYOTIC TRANSCRIPTION I: PROMOTERS AND ENHANCERS A. Promoters and Polymerases (RNA pols): 1. General characteristics - Initiation of transcription requires a. Transcription factors
Expression of the genome. Books: 1. Molecular biology of the gene: Watson et al 2. Genetics: Peter J. Russell
 Expression of the genome Books: 1. Molecular biology of the gene: Watson et al 2. Genetics: Peter J. Russell 1 Transcription 1. Francis Crick (1956) named the flow of information from DNA RNA protein the
Expression of the genome Books: 1. Molecular biology of the gene: Watson et al 2. Genetics: Peter J. Russell 1 Transcription 1. Francis Crick (1956) named the flow of information from DNA RNA protein the
Structure/function relationship in DNA-binding proteins
 PHRM 836 September 22, 2015 Structure/function relationship in DNA-binding proteins Devlin Chapter 8.8-9 u General description of transcription factors (TFs) u Sequence-specific interactions between DNA
PHRM 836 September 22, 2015 Structure/function relationship in DNA-binding proteins Devlin Chapter 8.8-9 u General description of transcription factors (TFs) u Sequence-specific interactions between DNA
Lecture 11. Initiation of RNA Pol II transcription. Transcription Initiation Complex
 Lecture 11 *Eukaryotic Transcription Gene Organization RNA Processing 5 cap 3 polyadenylation splicing Translation Initiation of RNA Pol II transcription Consensus sequence of promoter TATA Transcription
Lecture 11 *Eukaryotic Transcription Gene Organization RNA Processing 5 cap 3 polyadenylation splicing Translation Initiation of RNA Pol II transcription Consensus sequence of promoter TATA Transcription
Practice Exam A. Briefly describe how IL-25 treatment might be able to help this responder subgroup of liver cancer patients.
 Practice Exam 2007 1. A special JAK-STAT signaling system (JAK5-STAT5) was recently identified in which a gene called TS5 becomes selectively transcribed and expressed in the liver upon induction by a
Practice Exam 2007 1. A special JAK-STAT signaling system (JAK5-STAT5) was recently identified in which a gene called TS5 becomes selectively transcribed and expressed in the liver upon induction by a
DNA makes RNA makes Proteins. The Central Dogma
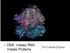 DNA makes RNA makes Proteins The Central Dogma TRANSCRIPTION DNA RNA transcript RNA polymerase RNA PROCESSING Exon RNA transcript (pre-mrna) Intron Aminoacyl-tRNA synthetase NUCLEUS CYTOPLASM FORMATION
DNA makes RNA makes Proteins The Central Dogma TRANSCRIPTION DNA RNA transcript RNA polymerase RNA PROCESSING Exon RNA transcript (pre-mrna) Intron Aminoacyl-tRNA synthetase NUCLEUS CYTOPLASM FORMATION
Make the protein through the genetic dogma process.
 Make the protein through the genetic dogma process. Coding Strand 5 AGCAATCATGGATTGGGTACATTTGTAACTGT 3 Template Strand mrna Protein Complete the table. DNA strand DNA s strand G mrna A C U G T A T Amino
Make the protein through the genetic dogma process. Coding Strand 5 AGCAATCATGGATTGGGTACATTTGTAACTGT 3 Template Strand mrna Protein Complete the table. DNA strand DNA s strand G mrna A C U G T A T Amino
Transcription. By : Lucia Dhiantika Witasari M.Biotech., Apt
 Transcription By : Lucia Dhiantika Witasari M.Biotech., Apt REGULATION OF GENE EXPRESSION 11/26/2010 2 RNA Messenger RNAs (mrnas) encode the amino acid sequence of one or more polypeptides specified by
Transcription By : Lucia Dhiantika Witasari M.Biotech., Apt REGULATION OF GENE EXPRESSION 11/26/2010 2 RNA Messenger RNAs (mrnas) encode the amino acid sequence of one or more polypeptides specified by
Synthetic cells: do bacteria need all its genes? No.
 NO NEED TO REFER TO THE SLIDES. بسم هللا الرحمن الرحيم Do we need all the non coding regions of the DNA? Two weeks ago, they discovered that the genome of a plant is very small (recall that plant genome
NO NEED TO REFER TO THE SLIDES. بسم هللا الرحمن الرحيم Do we need all the non coding regions of the DNA? Two weeks ago, they discovered that the genome of a plant is very small (recall that plant genome
Bis2A 12.2 Eukaryotic Transcription
 OpenStax-CNX module: m56061 1 Bis2A 12.2 Eukaryotic Transcription Mitch Singer Based on Eukaryotic Transcription by OpenStax College This work is produced by OpenStax-CNX and licensed under the Creative
OpenStax-CNX module: m56061 1 Bis2A 12.2 Eukaryotic Transcription Mitch Singer Based on Eukaryotic Transcription by OpenStax College This work is produced by OpenStax-CNX and licensed under the Creative
The discovery of the role of RNA RNA structure, synthesis and function
 Central Dogma The discovery of the role of RNA RNA structure, synthesis and function! Fundamental observations in genetics!! Genes are located in nuclei (in eukaryotes)!! Polypeptides are synthesised in
Central Dogma The discovery of the role of RNA RNA structure, synthesis and function! Fundamental observations in genetics!! Genes are located in nuclei (in eukaryotes)!! Polypeptides are synthesised in
Nucleotide Entry Port. Scaffold Subunits. Polymerase Activity β Sliding Clamp. Clamp Loader. Promoter Recognition
 Nucleotide Entry ort α (2) Scaffold Subunits olymerase Activity Sliding Clamp σ Clamp Loader romoter Recognition -35-10 NNAAA AA T A TTTTNNAAAANNN TT T N N17 N6 α α α α α α α α α α α α α α +1 α α α α Subunit
Nucleotide Entry ort α (2) Scaffold Subunits olymerase Activity Sliding Clamp σ Clamp Loader romoter Recognition -35-10 NNAAA AA T A TTTTNNAAAANNN TT T N N17 N6 α α α α α α α α α α α α α α +1 α α α α Subunit
BIO 311C Spring Lecture 36 Wednesday 28 Apr.
 BIO 311C Spring 2010 1 Lecture 36 Wednesday 28 Apr. Synthesis of a Polypeptide Chain 5 direction of ribosome movement along the mrna 3 ribosome mrna NH 2 polypeptide chain direction of mrna movement through
BIO 311C Spring 2010 1 Lecture 36 Wednesday 28 Apr. Synthesis of a Polypeptide Chain 5 direction of ribosome movement along the mrna 3 ribosome mrna NH 2 polypeptide chain direction of mrna movement through
Enhancers. Activators and repressors of transcription
 Enhancers Can be >50 kb away from the gene they regulate. Can be upstream from a promoter, downstream from a promoter, within an intron, or even downstream of the final exon of a gene. Are often cell type
Enhancers Can be >50 kb away from the gene they regulate. Can be upstream from a promoter, downstream from a promoter, within an intron, or even downstream of the final exon of a gene. Are often cell type
Transcription. The sugar molecule found in RNA is ribose, rather than the deoxyribose found in DNA.
 Transcription RNA (ribonucleic acid) is a key intermediary between a DNA sequence and a polypeptide. RNA is an informational polynucleotide similar to DNA, but it differs from DNA in three ways: RNA generally
Transcription RNA (ribonucleic acid) is a key intermediary between a DNA sequence and a polypeptide. RNA is an informational polynucleotide similar to DNA, but it differs from DNA in three ways: RNA generally
DNA Transcription. Visualizing Transcription. The Transcription Process
 DNA Transcription By: Suzanne Clancy, Ph.D. 2008 Nature Education Citation: Clancy, S. (2008) DNA transcription. Nature Education 1(1) If DNA is a book, then how is it read? Learn more about the DNA transcription
DNA Transcription By: Suzanne Clancy, Ph.D. 2008 Nature Education Citation: Clancy, S. (2008) DNA transcription. Nature Education 1(1) If DNA is a book, then how is it read? Learn more about the DNA transcription
(c) 2014 Dr. Alice Heicklen & Dr. Deborah Mowshowitz, Columbia University, New York, NY. Last update 02/26/ :57 PM
 C2006/F2402 '14 OUTLINE OF LECTURE #11 (c) 2014 Dr. Alice Heicklen & Dr. Deborah Mowshowitz, Columbia University, New York, NY. Last update 02/26/2014 12:57 PM Handouts: 10C -- Typical Eukaryotic Gene,
C2006/F2402 '14 OUTLINE OF LECTURE #11 (c) 2014 Dr. Alice Heicklen & Dr. Deborah Mowshowitz, Columbia University, New York, NY. Last update 02/26/2014 12:57 PM Handouts: 10C -- Typical Eukaryotic Gene,
Gene Expression: Transcription, Translation, RNAs and the Genetic Code
 Lecture 28-29 Gene Expression: Transcription, Translation, RNAs and the Genetic Code Central dogma of molecular biology During transcription, the information in a DNA sequence (a gene) is copied into a
Lecture 28-29 Gene Expression: Transcription, Translation, RNAs and the Genetic Code Central dogma of molecular biology During transcription, the information in a DNA sequence (a gene) is copied into a
Branches of Genetics
 Branches of Genetics 1. Transmission genetics Classical genetics or Mendelian genetics 2. Molecular genetics chromosomes, DNA, regulation of gene expression recombinant DNA, biotechnology, bioinformatics,
Branches of Genetics 1. Transmission genetics Classical genetics or Mendelian genetics 2. Molecular genetics chromosomes, DNA, regulation of gene expression recombinant DNA, biotechnology, bioinformatics,
Proofreading, post-replication modification of DNA. Mitesh Shrestha
 Proofreading, post-replication modification of DNA Mitesh Shrestha Proofreading During DNA replication (copying), most DNA polymerases can check their work with each base that they add. This process is
Proofreading, post-replication modification of DNA Mitesh Shrestha Proofreading During DNA replication (copying), most DNA polymerases can check their work with each base that they add. This process is
Chapter 9-II - Transcriptional Control of Gene Expression
 Chapter 9-II - Transcriptional Control of Gene Expression Transcriptional Control of Gene Expression 9.3 RNA Polymerase II Promoters and General Transcription Factors Three types of promoter sequences
Chapter 9-II - Transcriptional Control of Gene Expression Transcriptional Control of Gene Expression 9.3 RNA Polymerase II Promoters and General Transcription Factors Three types of promoter sequences
Molecular Cell Biology - Problem Drill 08: Transcription, Translation and the Genetic Code
 Molecular Cell Biology - Problem Drill 08: Transcription, Translation and the Genetic Code Question No. 1 of 10 1. Which of the following statements about how genes function is correct? Question #1 (A)
Molecular Cell Biology - Problem Drill 08: Transcription, Translation and the Genetic Code Question No. 1 of 10 1. Which of the following statements about how genes function is correct? Question #1 (A)
Transcriptional Regulation in Eukaryotes
 Transcriptional Regulation in Eukaryotes Concepts, Strategies, and Techniques Michael Carey Stephen T. Smale COLD SPRING HARBOR LABORATORY PRESS NEW YORK 2000 Cold Spring Harbor Laboratory Press, 0-87969-537-4
Transcriptional Regulation in Eukaryotes Concepts, Strategies, and Techniques Michael Carey Stephen T. Smale COLD SPRING HARBOR LABORATORY PRESS NEW YORK 2000 Cold Spring Harbor Laboratory Press, 0-87969-537-4
RNA Expression of the information in a gene generally involves production of an RNA molecule transcribed from a DNA template. RNA differs from DNA
 RNA Expression of the information in a gene generally involves production of an RNA molecule transcribed from a DNA template. RNA differs from DNA that it has a hydroxyl group at the 2 position of the
RNA Expression of the information in a gene generally involves production of an RNA molecule transcribed from a DNA template. RNA differs from DNA that it has a hydroxyl group at the 2 position of the
CH 17 :From Gene to Protein
 CH 17 :From Gene to Protein Defining a gene gene gene Defining a gene is problematic because one gene can code for several protein products, some genes code only for RNA, two genes can overlap, and there
CH 17 :From Gene to Protein Defining a gene gene gene Defining a gene is problematic because one gene can code for several protein products, some genes code only for RNA, two genes can overlap, and there
The Nature of Genes. The Nature of Genes. The Nature of Genes. The Nature of Genes. The Nature of Genes. The Genetic Code. Genes and How They Work
 Genes and How They Work Chapter 15 Early ideas to explain how genes work came from studying human diseases. Archibald Garrod studied alkaptonuria, 1902 Garrod recognized that the disease is inherited via
Genes and How They Work Chapter 15 Early ideas to explain how genes work came from studying human diseases. Archibald Garrod studied alkaptonuria, 1902 Garrod recognized that the disease is inherited via
Transcription in Prokaryotes. Jörg Bungert, PhD Phone:
 Transcription in Prokaryotes Jörg Bungert, PhD Phone: 352-273-8098 Email: jbungert@ufl.edu Objectives Understand the basic mechanism of transcription. Know the function of promoter elements and associating
Transcription in Prokaryotes Jörg Bungert, PhD Phone: 352-273-8098 Email: jbungert@ufl.edu Objectives Understand the basic mechanism of transcription. Know the function of promoter elements and associating
Chapter 8 Lecture Outline. Transcription, Translation, and Bioinformatics
 Chapter 8 Lecture Outline Transcription, Translation, and Bioinformatics Replication, Transcription, Translation n Repetitive processes Build polymers of nucleotides or amino acids n All have 3 major steps
Chapter 8 Lecture Outline Transcription, Translation, and Bioinformatics Replication, Transcription, Translation n Repetitive processes Build polymers of nucleotides or amino acids n All have 3 major steps
Gene function at the level of traits Gene function at the molecular level
 Gene expression Gene function at the level of traits Gene function at the molecular level Two levels tied together since the molecular level affects the structure and function of cells which determines
Gene expression Gene function at the level of traits Gene function at the molecular level Two levels tied together since the molecular level affects the structure and function of cells which determines
Biotechnology Unit 3: DNA to Proteins. From DNA to RNA
 From DNA to RNA Biotechnology Unit 3: DNA to Proteins I. After the discovery of the structure of DNA, the major question remaining was how does the stored in the 4 letter code of DNA direct the and of
From DNA to RNA Biotechnology Unit 3: DNA to Proteins I. After the discovery of the structure of DNA, the major question remaining was how does the stored in the 4 letter code of DNA direct the and of
Chapter 3 Gene Function. Transcription Prokaroyotes Eukaryotes Transcript processing Proteins Translation Genetic nomenclature
 Chapter 3 Gene Function Transcription Prokaroyotes Eukaryotes Transcript processing Proteins Translation Genetic nomenclature Transcription RNA composition ATP, GTP, UTP, CTP are substrates for RNA polymerase.
Chapter 3 Gene Function Transcription Prokaroyotes Eukaryotes Transcript processing Proteins Translation Genetic nomenclature Transcription RNA composition ATP, GTP, UTP, CTP are substrates for RNA polymerase.
Genes and How They Work. Chapter 15
 Genes and How They Work Chapter 15 The Nature of Genes They proposed the one gene one enzyme hypothesis. Today we know this as the one gene one polypeptide hypothesis. 2 The Nature of Genes The central
Genes and How They Work Chapter 15 The Nature of Genes They proposed the one gene one enzyme hypothesis. Today we know this as the one gene one polypeptide hypothesis. 2 The Nature of Genes The central
Classes of eukaryotic cellular RNAs
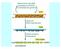 Classes of eukaryotic cellular RNAs ribosomal RNA (rrna) 18S (small subunit) 28S (large subunit) 5.8S (large subunit) 5S (large subunit) transfer RNA (trna) messenger RNA (mrna) heterogeneous nuclear RNA
Classes of eukaryotic cellular RNAs ribosomal RNA (rrna) 18S (small subunit) 28S (large subunit) 5.8S (large subunit) 5S (large subunit) transfer RNA (trna) messenger RNA (mrna) heterogeneous nuclear RNA
CELL BIOLOGY - CLUTCH CH. 7 - GENE EXPRESSION.
 !! www.clutchprep.com CONCEPT: CONTROL OF GENE EXPRESSION BASICS Gene expression is the process through which cells selectively to express some genes and not others Every cell in an organism is a clone
!! www.clutchprep.com CONCEPT: CONTROL OF GENE EXPRESSION BASICS Gene expression is the process through which cells selectively to express some genes and not others Every cell in an organism is a clone
Eukaryotic Gene Expression John O. Thomas
 Eukaryotic Gene Expression John O. Thomas I) RNA polymerases A) There are four RNA polymerases in human cells. 1) RNA polymerase I, located in the nucleolar region of the nucleus, is responsible for the
Eukaryotic Gene Expression John O. Thomas I) RNA polymerases A) There are four RNA polymerases in human cells. 1) RNA polymerase I, located in the nucleolar region of the nucleus, is responsible for the
SCBC203 Gene Expression. Assoc. Prof. Rutaiwan Tohtong Department of Biochemistry Faculty of Science PR318
 SCBC203 Gene Expression Assoc. Prof. Rutaiwan Tohtong Department of Biochemistry Faculty of Science PR318 Rutaiwan.toh@mahidol.ac.th 1 Gene Expression Gene expression is a process where by the genetic
SCBC203 Gene Expression Assoc. Prof. Rutaiwan Tohtong Department of Biochemistry Faculty of Science PR318 Rutaiwan.toh@mahidol.ac.th 1 Gene Expression Gene expression is a process where by the genetic
Control of Gene Expression
 Control of Gene Expression 1 How Gene Regulation Works 2 Control of Gene Expression Controlling gene expression is often accomplished by controlling transcription initiation Regulatory proteins bind to
Control of Gene Expression 1 How Gene Regulation Works 2 Control of Gene Expression Controlling gene expression is often accomplished by controlling transcription initiation Regulatory proteins bind to
Transcription and Post Transcript Modification
 Transcription and Post Transcript Modification You Should Be Able To 1. Describe transcription. 2. Compare and contrast eukaryotic + prokaryotic transcription. 3. Explain mrna processing in eukaryotes.
Transcription and Post Transcript Modification You Should Be Able To 1. Describe transcription. 2. Compare and contrast eukaryotic + prokaryotic transcription. 3. Explain mrna processing in eukaryotes.
RNA synthesis/transcription I Biochemistry 302. February 6, 2004 Bob Kelm
 RNA synthesis/transcription I Biochemistry 302 February 6, 2004 Bob Kelm Overview of RNA classes Messenger RNA (mrna) Encodes protein Relatively short half-life ( 3 min in E. coli, 30 min in eukaryotic
RNA synthesis/transcription I Biochemistry 302 February 6, 2004 Bob Kelm Overview of RNA classes Messenger RNA (mrna) Encodes protein Relatively short half-life ( 3 min in E. coli, 30 min in eukaryotic
Transcription. DNA to RNA
 Transcription from DNA to RNA The Central Dogma of Molecular Biology replication DNA RNA Protein transcription translation Why call it transcription and translation? transcription is such a direct copy
Transcription from DNA to RNA The Central Dogma of Molecular Biology replication DNA RNA Protein transcription translation Why call it transcription and translation? transcription is such a direct copy
5. Which of the following enzymes catalyze the attachment of an amino acid to trna in the formation of aminoacyl trna?
 Sample Examination Questions for Exam 3 Material Biology 3300 / Dr. Jerald Hendrix Warning! These questions are posted solely to provide examples of past test questions. There is no guarantee that any
Sample Examination Questions for Exam 3 Material Biology 3300 / Dr. Jerald Hendrix Warning! These questions are posted solely to provide examples of past test questions. There is no guarantee that any
Bi 8 Lecture 10. Ellen Rothenberg 4 February 2016
 Bi 8 Lecture 10 Bacterial regulation, II Ellen Rothenberg 4 February 2016 Not all bacterial promoters use the same σ factors, and this provides added regulation capability Most sigma factors are related
Bi 8 Lecture 10 Bacterial regulation, II Ellen Rothenberg 4 February 2016 Not all bacterial promoters use the same σ factors, and this provides added regulation capability Most sigma factors are related
Molecular Genetics of the RNA Polymerase II General Transcriptional Machinery
 MICROBIOLOGY AND MOLECULAR BIOLOGY REVIEWS, June 1998, p. 465 503 Vol. 62, No. 2 1092-2172/98/$04.00 0 Copyright 1998, American Society for Microbiology Molecular Genetics of the RNA Polymerase II General
MICROBIOLOGY AND MOLECULAR BIOLOGY REVIEWS, June 1998, p. 465 503 Vol. 62, No. 2 1092-2172/98/$04.00 0 Copyright 1998, American Society for Microbiology Molecular Genetics of the RNA Polymerase II General
Section C: The Control of Gene Expression
 Section C: The Control of Gene Expression 1. Each cell of a multicellular eukaryote expresses only a small fraction of its genes 2. The control of gene expression can occur at any step in the pathway from
Section C: The Control of Gene Expression 1. Each cell of a multicellular eukaryote expresses only a small fraction of its genes 2. The control of gene expression can occur at any step in the pathway from
BEADLE & TATUM EXPERIMENT
 FROM DNA TO PROTEINS: gene expression Chapter 14 LECTURE OBJECTIVES What Is the Evidence that Genes Code for Proteins? How Does Information Flow from Genes to Proteins? How Is the Information Content in
FROM DNA TO PROTEINS: gene expression Chapter 14 LECTURE OBJECTIVES What Is the Evidence that Genes Code for Proteins? How Does Information Flow from Genes to Proteins? How Is the Information Content in
Chapter 18: Regulation of Gene Expression. 1. Gene Regulation in Bacteria 2. Gene Regulation in Eukaryotes 3. Gene Regulation & Cancer
 Chapter 18: Regulation of Gene Expression 1. Gene Regulation in Bacteria 2. Gene Regulation in Eukaryotes 3. Gene Regulation & Cancer Gene Regulation Gene regulation refers to all aspects of controlling
Chapter 18: Regulation of Gene Expression 1. Gene Regulation in Bacteria 2. Gene Regulation in Eukaryotes 3. Gene Regulation & Cancer Gene Regulation Gene regulation refers to all aspects of controlling
Ch. 10 From DNA to Protein. AP Biology
 Ch. 10 From DNA to Protein Protein Synthesis Metabolism and Gene Expression n Inheritance of metabolic diseases suggests that genes coded for enzymes n Diseases (phenotypes) caused by non-functional gene
Ch. 10 From DNA to Protein Protein Synthesis Metabolism and Gene Expression n Inheritance of metabolic diseases suggests that genes coded for enzymes n Diseases (phenotypes) caused by non-functional gene
Analyzed Fungi Neurospora crassa mutants. Mutants were UNABLE to grow without Arginine (an amino acid) Other biochemical experiments indicated:
 From Gene to Protein Beadle and Tatum Analyzed Fungi Neurospora crassa mutants Mutants were UNABLE to grow without Arginine (an amino acid) Other biochemical experiments indicated: Precursor Ornithine
From Gene to Protein Beadle and Tatum Analyzed Fungi Neurospora crassa mutants Mutants were UNABLE to grow without Arginine (an amino acid) Other biochemical experiments indicated: Precursor Ornithine
Chapter 14 Regulation of Transcription
 Chapter 14 Regulation of Transcription Cis-acting sequences Distance-independent cis-acting elements Dissecting regulatory elements Transcription factors Overview transcriptional regulation Transcription
Chapter 14 Regulation of Transcription Cis-acting sequences Distance-independent cis-acting elements Dissecting regulatory elements Transcription factors Overview transcriptional regulation Transcription
DNA Bending and Wrapping around RNA Polymerase: a Revolutionary Model Describing Transcriptional Mechanisms
 MICROBIOLOGY AND MOLECULAR BIOLOGY REVIEWS, June 1999, p. 457 478 Vol. 63, No. 2 1092-2172/99/$04.00 0 Copyright 1999, American Society for Microbiology. All Rights Reserved. DNA Bending and Wrapping around
MICROBIOLOGY AND MOLECULAR BIOLOGY REVIEWS, June 1999, p. 457 478 Vol. 63, No. 2 1092-2172/99/$04.00 0 Copyright 1999, American Society for Microbiology. All Rights Reserved. DNA Bending and Wrapping around
RNA Polymerase II The Transcription Machine
 RNA Polymerase II The Transcription Machine Nobel Prize in Chemistry 2006 Jiyoti Verma 1, Aruna Naorem 2, Anand Kumar 3, Manimala Sen 4 and Parag Sadhale 5 It was 2005 December. The distinguished speaker
RNA Polymerase II The Transcription Machine Nobel Prize in Chemistry 2006 Jiyoti Verma 1, Aruna Naorem 2, Anand Kumar 3, Manimala Sen 4 and Parag Sadhale 5 It was 2005 December. The distinguished speaker
Student name ID # Second Mid Term Exam, Biology 2020, Spring 2002 Scores Total
 Second Mid Term Exam, Biology 2020, Spring 2002 Scores 1. 2. 3. 4. 5. 6. 7. 8. 9. 10. 11. 12. 13. 14. 15. 16. 17. 18. 19. 20. 21. Total 1 1. Matching (7 pts). Each answer is used exactly once Helicase
Second Mid Term Exam, Biology 2020, Spring 2002 Scores 1. 2. 3. 4. 5. 6. 7. 8. 9. 10. 11. 12. 13. 14. 15. 16. 17. 18. 19. 20. 21. Total 1 1. Matching (7 pts). Each answer is used exactly once Helicase
CONTENTS. LIST OF FIGURES... i. LIST OF TABLES... iii. SUMMARY OF THE WORK DONE... vi
 CONTENTS SECTION ~ LIST OF FIGURES... i LIST OF TABLES... iii LIST OF ABBREVIATIONS USED... ~... iv SUMMARY OF THE WORK DONE... vi 1. INTROD UCTION... :.................................... 1 1.1. AN OVERVIEW
CONTENTS SECTION ~ LIST OF FIGURES... i LIST OF TABLES... iii LIST OF ABBREVIATIONS USED... ~... iv SUMMARY OF THE WORK DONE... vi 1. INTROD UCTION... :.................................... 1 1.1. AN OVERVIEW
MOLECULAR GENETICS PROTEIN SYNTHESIS. Molecular Genetics Activity #2 page 1
 AP BIOLOGY MOLECULAR GENETICS ACTIVITY #2 NAME DATE HOUR PROTEIN SYNTHESIS Molecular Genetics Activity #2 page 1 GENETIC CODE PROTEIN SYNTHESIS OVERVIEW Molecular Genetics Activity #2 page 2 PROTEIN SYNTHESIS
AP BIOLOGY MOLECULAR GENETICS ACTIVITY #2 NAME DATE HOUR PROTEIN SYNTHESIS Molecular Genetics Activity #2 page 1 GENETIC CODE PROTEIN SYNTHESIS OVERVIEW Molecular Genetics Activity #2 page 2 PROTEIN SYNTHESIS
17.5 Eukaryotic Transcription Initiation Is Regulated by Transcription Factors That Bind to Cis-Acting Sites
 17.5 Eukaryotic Transcription Initiation Is Regulated by Transcription Factors That Bind to Cis-Acting Sites 1 Section 17.5 Transcription regulatory proteins, transcription factors, target cis-acting sites
17.5 Eukaryotic Transcription Initiation Is Regulated by Transcription Factors That Bind to Cis-Acting Sites 1 Section 17.5 Transcription regulatory proteins, transcription factors, target cis-acting sites
Lecture Summary: Regulation of transcription. General mechanisms-what are the major regulatory points?
 BCH 401G Lecture 37 Andres Lecture Summary: Regulation of transcription. General mechanisms-what are the major regulatory points? RNA processing: Capping, polyadenylation, splicing. Why process mammalian
BCH 401G Lecture 37 Andres Lecture Summary: Regulation of transcription. General mechanisms-what are the major regulatory points? RNA processing: Capping, polyadenylation, splicing. Why process mammalian
AP Biology. The BIG Questions. Chapter 19. Prokaryote vs. eukaryote genome. Prokaryote vs. eukaryote genome. Why turn genes on & off?
 The BIG Questions Chapter 19. Control of Eukaryotic Genome How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different, specialized functions?
The BIG Questions Chapter 19. Control of Eukaryotic Genome How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different, specialized functions?
Chapter 2. An Introduction to Genes and Genomes
 PowerPoint Lectures for Introduction to Biotechnology, Second Edition William J.Thieman and Michael A.Palladino Chapter 2 An Introduction to Genes and Genomes Lectures by Lara Dowland Chapter Contents
PowerPoint Lectures for Introduction to Biotechnology, Second Edition William J.Thieman and Michael A.Palladino Chapter 2 An Introduction to Genes and Genomes Lectures by Lara Dowland Chapter Contents
Chromatin and Transcription
 Chromatin and Transcription Chromatin Structure Chromatin Represses Transcription Nucleosome Positioning Histone Acetylation Chromatin Remodeling Histone Methylation CHIP Analysis Chromatin and Elongation
Chromatin and Transcription Chromatin Structure Chromatin Represses Transcription Nucleosome Positioning Histone Acetylation Chromatin Remodeling Histone Methylation CHIP Analysis Chromatin and Elongation
Control of Eukaryotic Gene Expression (Learning Objectives)
 Control of Eukaryotic Gene Expression (Learning Objectives) 1. Compare and contrast chromatin and chromosome: composition, proteins involved and level of packing. Explain the structure and function of
Control of Eukaryotic Gene Expression (Learning Objectives) 1. Compare and contrast chromatin and chromosome: composition, proteins involved and level of packing. Explain the structure and function of
Biochemistry 302. Exam 2. March 10, Answer Key
 1 Biochemistry 302 Exam 2 March 10, 2004 Answer Key 2 Biochemistry 302, Spring 2004 Exam 2 (100 points) Name I. Short answer 1. Identify the 5 end, 3 end, amino acid acceptor nucleoside, and bases comprising
1 Biochemistry 302 Exam 2 March 10, 2004 Answer Key 2 Biochemistry 302, Spring 2004 Exam 2 (100 points) Name I. Short answer 1. Identify the 5 end, 3 end, amino acid acceptor nucleoside, and bases comprising
Transcription is the first stage of gene expression
 Transcription is the first stage of gene expression RNA synthesis is catalyzed by RNA polymerase, which pries the DNA strands apart and hooks together the RNA nucleotides The RNA is complementary to the
Transcription is the first stage of gene expression RNA synthesis is catalyzed by RNA polymerase, which pries the DNA strands apart and hooks together the RNA nucleotides The RNA is complementary to the
Lecture for Wednesday. Dr. Prince BIOL 1408
 Lecture for Wednesday Dr. Prince BIOL 1408 THE FLOW OF GENETIC INFORMATION FROM DNA TO RNA TO PROTEIN Copyright 2009 Pearson Education, Inc. Genes are expressed as proteins A gene is a segment of DNA that
Lecture for Wednesday Dr. Prince BIOL 1408 THE FLOW OF GENETIC INFORMATION FROM DNA TO RNA TO PROTEIN Copyright 2009 Pearson Education, Inc. Genes are expressed as proteins A gene is a segment of DNA that
Biological information flow
 BCMB 3100 Chapters 36-38 Transcription & RNA Processing Definition of gene RNA Polymerase Gene coding vs template strand Promoter Transcription in E. coli Transcription factors mrna processing Biological
BCMB 3100 Chapters 36-38 Transcription & RNA Processing Definition of gene RNA Polymerase Gene coding vs template strand Promoter Transcription in E. coli Transcription factors mrna processing Biological
