Impact of Retinoic acid induced-1 (Rai1) on Regulators of Metabolism and Adipogenesis
|
|
|
- Lynette Black
- 6 years ago
- Views:
Transcription
1 Impact of Retinoic acid induced-1 (Rai1) on Regulators of Metabolism and Adipogenesis The mammalian system undergoes ~24 hour cycles known as circadian rhythms that temporally orchestrate metabolism, behavior, and physiology through the manipulation of a complex transcriptional circuitry. In mammals, a central master clock located in the SCN of the hypothalamus generates signals that control peripheral clocks in other tissues of the body to regulate body temperature, hormone product, sleeping and feeding patterns, and other biological activities. Given the complexity of this regulator (Fig 1), it is not surprising that differential expression of clock components may have widespread effects on downstream genes, notably those that regulate energy metabolism, an idea supported by the observation that Clock gene mutant mice develop obesity and metabolic syndrome (Turek et al. 2005). A recent study in the Elsea lab demonstrated that alteration of transcription factor Retinoic acid induced-1 (RAI1) impacts the expression of CLOCK, leading to a wide number of abnormalities (Williams et al. 2012). Haploinsufficiency of RAI1 is the primary cause of Smith-Magenis syndrome (OMIM ), a complex congenital disease characterized by neurobehavioral complications, craniofacial abnormalities, obesity, and an inverted circadian rhythm (Elsea & Williams 2011). A large portion of SMS patients exhibit aberrant fat distribution leading to truncal obesity. Mouse models of SMS also display obesity along with high levels of leptin and corticosterone (Fig 2). Moreover, Rai1-Tg (over expressing Rai1) mice are underweight, indicating the importance of Rai1 dosage (Burns et al. 2010). Interestingly, SMS patients do not have an increased risk for metabolic syndromes, including insulin resistance Fig 1. The transcriptional circuitry that integrators circadian oscillator and metabolism (Froy, 2011) Fig 2. (A) Wild type (normal littermates) and Rai1 +/- (heterozygous) mice of same age are shown. Older Rai1 +/- animals may grow to >70 g. (B) Shown are mouse weights at 5-30 weeks of age. Older Rai1 +/- animals may grow to >70 g (Burns et al. 2010). 1
2 and diabetes, suggesting a more complicated mechanism of pathogenesis. Despite preliminary data regarding Rai1 dosage on levels of several circadian and metabolic genes, the etiology of the metabolic complications observed in persons with SMS remains largely unexplored. Rai1 regulates the core components of the circadian clock, but how does Rai1dosage impact its downstream transcriptional circuitry involved in metabolic regulation? In the present work, we propose to investigate the expressional dynamics of key metabolic genes in Rai1 haploinsufficient and over-expressing mouse metabolic tissues, using wet-lab and computational approaches. We will also conduct a histological analysis of adipose and liver tissue to investigate lipid accumulation (Fig 3). The data generated from the following experimental plan will establish a foundation for designing future studies regarding potential treatments targeting affected pathways to systemically alter circadian regulators and facilitate healthy metabolism in persons with SMS. Fig 3: A summary of the work plan to investigate our research question regarding Rai1 and its impact on metabolic processes. The plan is designed to reach three main aims that will be reached through the shown work flow. 2
3 Aim 1: Determining the expression levels of key metabolic genes (6 weeks) The transcription network at the intersection between the central circadian regulator and the peripheral metabolic tissues is complex, and to begin understanding the role of Rai1 dosage in metabolic regulation, we must first determine its impact on the key elements of this network. Therefore, we propose to assess the expression levels of several key metabolic regulator as well as adipocyte-specific genes in Rai1+/- and Rai1-Tg mice tissue (Table 1). White adipose tissue (WAT), muscle, and hypothalamic tissues, key metabolic sites in humans and mice, will be utilized for this assessment. Tissues will be collected from 3-5 mice for each genotype Table 1: Key metabolic regulator and adiposity genes for this investigation. Expression levels of these genes will be assessed in Rai1 dosage altered tissues from mice. in accordance with standard protocol and frozen at -80 C. In order to implement this project, I will obtain training and approval to work on Dr. Elsea s IACUC protocol (AM10101) prior to summer. The expression levels of these genes will be determined using quantitative real-time PCR. Total cellular RNA will be extracted from all three tissues using RNeasy kit (Qiagen protocol), and then reverse transcribed to obtain cdna. Left and right primers for each gene in Table 1 will be designed using Taqman assays. qpcr results will be analyzed using the software accompanying the qpcr machine. This study will result in graphical data that will quantify the expression each chosen gene relative to expression in WT mouse tissues. Given the regulatory role of Rai1 on the Clock gene, our current prediction is that many of these downstream metabolic and adiposity genes will also show aberrant expression levels. These data will allow us to do more complicated analysis and focus future studies on manipulating these genes in an effort to alter the dynamics of the affected pathways. Aim 2: Assessing the characteristics of Rai1 altered adipose tissue (2-3 weeks) In addition to investigating the molecular changes that occur due to altered Rai1 dosage, we will conduct a phenotypic analysis of the adipose tissue from Rai1 heterozygous and transgenic mice. Currently published work shows that adipocytes in Rai1 haploinsufficient tissue show consistent size and seem to be full relative to WT fat cells (Burns et al. 2010). Interestingly, an increased number of evenly shaped fat cells are 3
4 seen in a given section of tissue rather than adipocytes enlargement, which is normally observed due to excess adiposity. This perhaps indicates increased lipid accumulation, and can be easily investigated using an Oil Red O (ORO) staining protocol that shows lipid localization in tissues. Frozen or fixed adipose tissue will be sent to VCU Pathology lab for sectioning and will be subsequently stained using a standard protocol. Fat will be seen as red droplets in the tissue, with a blue nuclear counter stain. This visualization will help us evaluate the lipid accumulation in Rai1 deficient or over-expressing adipose tissue, and based on results, various other staining might also be conducted to further characterize the cell count and morphology in these tissues. Aim 3: Identifying a consensus DNA binding site sequence for Rai1 (All summer) Alongside investigations using wet-lab techniques, it will be essential to approach our research question from an in silico perspective through analysis of data available regarding Rai1 transcriptional activity at the genome-wide scale. What promoters does Rai1 protein bind to? Which of these genes are involved in metabolism? The answers can be explored through computational analysis of Chromatin Immunoprecipitation with microarray (ChIP-Chip) data sets that are currently available to the Elsea lab. Briefly, this data contains information regarding Rai1 protein interaction with functional elements in the mouse genome, and has DNA sequence of sites that Rai1 protein binds to in an in vivo environment. Determination of a consensus binding site sequence for Rai1 using these sequence fragments will allow us to search promoters of metabolic genes in other genomes and identify the direct targets of Rai1 transcriptional activity. This can be done by generating sequence alignments using ClustalW2, and visualizing them using a Jalview platform. It will be of interest to see whether some of the candidate genes from this analysis overlap with the differentially expressed genes we identify through q-pcr. In addition, we will generate a list of all metabolic genes that are reported in the ChIP-chip data sets and classify them based on their role in various metabolic processes. Comparison with a previous microarray data set, which lists genes that are differentially expressed when Rai1 is down regulated in mouse hypothalamus, will be crucial during this analysis, and we expect to find Rai1 associated genes that are common to both data sets. 4
5 Work Plan and Future Studies Assuming hours in the lab, a tentative work plan for each week is shown in Table 2. A mid-term project report regarding the data obtained from this ten week research investigation will be submitted August 1, 2012 and the research project results will be presented at the VCU Undergraduate Symposium/Poster Day in Spring This study, however, will not conclude with these three aims. Based on the preliminary data that will be obtained this summer, we will be able to determine additional lines of study. Integration of expression data will help determine if certain metabolic pathways are affected. If so, we can design studies that alter mouse feeding schedule, etc, to see whether we can manipulate the circadian-metabolic oscillator and therefore the expression of downstream genes that innervate the affected metabolic pathways. Aim* Methodology Week 1: Design primers for target genes: Taqman Assays Week 2: Collect tissues from transgenic mice Week 3: Collect tissues from wild type mice Week 4: 1 RNA Extraction from tissues: preparation of cdna Week 5: qpcr and expression data analysis Week 6: Result replication and Analysis (Starting preparation of mid-term report) Week 7: Adipose tissue sectioning at the VCU Pathology Dept. Week 8: 2 Oil Red O staining protocol Week 9: Analysis and imaging of stained tissues Week 10: Submit mid-term report on August 1st. *Aim 3 (ChIP-Chip data analysis) is not included as it will be conducted all summer. Table 2: A detailed 10-week work plan for implementation of the proposed work. References: 1. Chen J, Xu H, Aronow BJ, Jegga AG Improved human disease candidate gene prioritization using mouse phenotype. BMC Bioinformatics 8: Burns, Brooke, et al. "Rai1 haploinsufficiency causes reduced Bdnf expression resulting in hyperphagia, obesity and altered fat distribution in mice and humans with no evidence of metabolic syndrome." Human Molecular Genetics (2010): Froy O. Metabolism and circadian rhythms: implications for obesity. Endocr Rev 2011; 31: Nadler, S T, et al. "The expression of adipogenic genes is decreased in obesity and diabetes mellitus." Proceedings of the National Academy of Sciences (2000): Turek, Fred W, et al. "Obesity and metabolic syndrome in circadian Clock mutant mice." Science (2005): Williams SR, Zies, D, Mullegama SV, Grotewiel MS, Elsea SH Smith-Magenis syndrome results in disruption of CLOCK gene transcription and reveals an integral role for RAI1 in the maintenance of the circadian rhythmicity. (Manuscript Submitted). 5
6th Form Open Day 15th July 2015
 6th Form Open Day 15 th July 2015 DNA is like a computer program but far, far more advanced than any software ever created. Bill Gates Welcome to the Open Day event at the Institute of Genetic Medicine
6th Form Open Day 15 th July 2015 DNA is like a computer program but far, far more advanced than any software ever created. Bill Gates Welcome to the Open Day event at the Institute of Genetic Medicine
How to deal with the microarray results.
 How to deal with the microarray results. Britt Gabrielsson PhD RCEM, Div of metabolism and cardiovascular research Department of Medicine The Sahlgrenska Academy at Göteborg University and then we will
How to deal with the microarray results. Britt Gabrielsson PhD RCEM, Div of metabolism and cardiovascular research Department of Medicine The Sahlgrenska Academy at Göteborg University and then we will
Supplementary Methods
 Supplementary Methods Reverse transcribed Quantitative PCR. Total RNA was isolated from bone marrow derived macrophages using RNeasy Mini Kit (Qiagen), DNase-treated (Promega RQ1), and reverse transcribed
Supplementary Methods Reverse transcribed Quantitative PCR. Total RNA was isolated from bone marrow derived macrophages using RNeasy Mini Kit (Qiagen), DNase-treated (Promega RQ1), and reverse transcribed
GENETICS - CLUTCH CH.15 GENOMES AND GENOMICS.
 !! www.clutchprep.com CONCEPT: OVERVIEW OF GENOMICS Genomics is the study of genomes in their entirety Bioinformatics is the analysis of the information content of genomes - Genes, regulatory sequences,
!! www.clutchprep.com CONCEPT: OVERVIEW OF GENOMICS Genomics is the study of genomes in their entirety Bioinformatics is the analysis of the information content of genomes - Genes, regulatory sequences,
C57BL/6N Female. C57BL/6N Male. Prl2 WT Allele. Prl2 Targeted Allele **** **** WT Het KO PRL bp. 230 bp CNX. C57BL/6N Male 30.
 A Prl Allele Prl Targeted Allele 631 bp 1 3 4 5 6 bp 1 SA β-geo pa 3 4 5 6 Het B Het 631 bp bp PRL CNX C 1 C57BL/6N Male (14) Het (7) (1) 1 C57BL/6N Female (19) Het (31) (1) % survival 5 % survival 5 ****
A Prl Allele Prl Targeted Allele 631 bp 1 3 4 5 6 bp 1 SA β-geo pa 3 4 5 6 Het B Het 631 bp bp PRL CNX C 1 C57BL/6N Male (14) Het (7) (1) 1 C57BL/6N Female (19) Het (31) (1) % survival 5 % survival 5 ****
DNA Microarray Technology
 CHAPTER 1 DNA Microarray Technology All living organisms are composed of cells. As a functional unit, each cell can make copies of itself, and this process depends on a proper replication of the genetic
CHAPTER 1 DNA Microarray Technology All living organisms are composed of cells. As a functional unit, each cell can make copies of itself, and this process depends on a proper replication of the genetic
Combining Techniques to Answer Molecular Questions
 Combining Techniques to Answer Molecular Questions UNIT FM02 How to cite this article: Curr. Protoc. Essential Lab. Tech. 9:FM02.1-FM02.5. doi: 10.1002/9780470089941.etfm02s9 INTRODUCTION This manual is
Combining Techniques to Answer Molecular Questions UNIT FM02 How to cite this article: Curr. Protoc. Essential Lab. Tech. 9:FM02.1-FM02.5. doi: 10.1002/9780470089941.etfm02s9 INTRODUCTION This manual is
Figure S1. Figure S2. Figure S3 HB Anti-FSP27 (COOH-terminal peptide) Ab. Anti-GST-FSP27(45-127) Ab.
 / 36B4 mrna ratio Figure S1 * 2. 1.6 1.2.8 *.4 control TNFα BRL49653 Figure S2 Su bw AT p iw Anti- (COOH-terminal peptide) Ab Blot : Anti-GST-(45-127) Ab β-actin Figure S3 HB2 HW AT BA T Figure S4 A TAG
/ 36B4 mrna ratio Figure S1 * 2. 1.6 1.2.8 *.4 control TNFα BRL49653 Figure S2 Su bw AT p iw Anti- (COOH-terminal peptide) Ab Blot : Anti-GST-(45-127) Ab β-actin Figure S3 HB2 HW AT BA T Figure S4 A TAG
Chapter 15 Gene Technologies and Human Applications
 Chapter Outline Chapter 15 Gene Technologies and Human Applications Section 1: The Human Genome KEY IDEAS > Why is the Human Genome Project so important? > How do genomics and gene technologies affect
Chapter Outline Chapter 15 Gene Technologies and Human Applications Section 1: The Human Genome KEY IDEAS > Why is the Human Genome Project so important? > How do genomics and gene technologies affect
QPCR ASSAYS FOR MIRNA EXPRESSION PROFILING
 TECH NOTE 4320 Forest Park Ave Suite 303 Saint Louis, MO 63108 +1 (314) 833-9764 mirna qpcr ASSAYS - powered by NAWGEN Our mirna qpcr Assays were developed by mirna experts at Nawgen to improve upon previously
TECH NOTE 4320 Forest Park Ave Suite 303 Saint Louis, MO 63108 +1 (314) 833-9764 mirna qpcr ASSAYS - powered by NAWGEN Our mirna qpcr Assays were developed by mirna experts at Nawgen to improve upon previously
3.1.4 DNA Microarray Technology
 3.1.4 DNA Microarray Technology Scientists have discovered that one of the differences between healthy and cancer is which genes are turned on in each. Scientists can compare the gene expression patterns
3.1.4 DNA Microarray Technology Scientists have discovered that one of the differences between healthy and cancer is which genes are turned on in each. Scientists can compare the gene expression patterns
Gene expression analysis. Biosciences 741: Genomics Fall, 2013 Week 5. Gene expression analysis
 Gene expression analysis Biosciences 741: Genomics Fall, 2013 Week 5 Gene expression analysis From EST clusters to spotted cdna microarrays Long vs. short oligonucleotide microarrays vs. RT-PCR Methods
Gene expression analysis Biosciences 741: Genomics Fall, 2013 Week 5 Gene expression analysis From EST clusters to spotted cdna microarrays Long vs. short oligonucleotide microarrays vs. RT-PCR Methods
Chapter 20 Recombinant DNA Technology. Copyright 2009 Pearson Education, Inc.
 Chapter 20 Recombinant DNA Technology Copyright 2009 Pearson Education, Inc. 20.1 Recombinant DNA Technology Began with Two Key Tools: Restriction Enzymes and DNA Cloning Vectors Recombinant DNA refers
Chapter 20 Recombinant DNA Technology Copyright 2009 Pearson Education, Inc. 20.1 Recombinant DNA Technology Began with Two Key Tools: Restriction Enzymes and DNA Cloning Vectors Recombinant DNA refers
CircadiOmics: Integrating Circadian Genomics, Transcriptomics, Proteomics, and Metabolomics
 CircadiOmics: Integrating Circadian Genomics, Transcriptomics, Proteomics, and Metabolomics Vishal R Patel, Kristin Eckel-Mahan, Paolo Sassone-Corsi, Pierre Baldi Supplementary Figure 1: CircadiOmics Webserver
CircadiOmics: Integrating Circadian Genomics, Transcriptomics, Proteomics, and Metabolomics Vishal R Patel, Kristin Eckel-Mahan, Paolo Sassone-Corsi, Pierre Baldi Supplementary Figure 1: CircadiOmics Webserver
Introduction to Microarray Analysis
 Introduction to Microarray Analysis Methods Course: Gene Expression Data Analysis -Day One Rainer Spang Microarrays Highly parallel measurement devices for gene expression levels 1. How does the microarray
Introduction to Microarray Analysis Methods Course: Gene Expression Data Analysis -Day One Rainer Spang Microarrays Highly parallel measurement devices for gene expression levels 1. How does the microarray
GREG GIBSON SPENCER V. MUSE
 A Primer of Genome Science ience THIRD EDITION TAGCACCTAGAATCATGGAGAGATAATTCGGTGAGAATTAAATGGAGAGTTGCATAGAGAACTGCGAACTG GREG GIBSON SPENCER V. MUSE North Carolina State University Sinauer Associates, Inc.
A Primer of Genome Science ience THIRD EDITION TAGCACCTAGAATCATGGAGAGATAATTCGGTGAGAATTAAATGGAGAGTTGCATAGAGAACTGCGAACTG GREG GIBSON SPENCER V. MUSE North Carolina State University Sinauer Associates, Inc.
ChampionChIP Quick, High Throughput Chromatin Immunoprecipitation Assay System
 ChampionChIP Quick, High Throughput Chromatin Immunoprecipitation Assay System Liyan Pang, Ph.D. Application Scientist 1 Topics to be Covered Introduction What is ChIP-qPCR? Challenges Facing Biological
ChampionChIP Quick, High Throughput Chromatin Immunoprecipitation Assay System Liyan Pang, Ph.D. Application Scientist 1 Topics to be Covered Introduction What is ChIP-qPCR? Challenges Facing Biological
EPIGENTEK. EpiQuik Tissue Chromatin Immunoprecipitation Kit. Base Catalog # P-2003 PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE
 EpiQuik Tissue Chromatin Immunoprecipitation Kit Base Catalog # P-2003 PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE The EpiQuik Tissue Chromatin Immunoprecipitation Kit is suitable for combining the specificity
EpiQuik Tissue Chromatin Immunoprecipitation Kit Base Catalog # P-2003 PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE The EpiQuik Tissue Chromatin Immunoprecipitation Kit is suitable for combining the specificity
Non-Organic-Based Isolation of Mammalian microrna using Norgen s microrna Purification Kit
 Application Note 13 RNA Sample Preparation Non-Organic-Based Isolation of Mammalian microrna using Norgen s microrna Purification Kit B. Lam, PhD 1, P. Roberts, MSc 1 Y. Haj-Ahmad, M.Sc., Ph.D 1,2 1 Norgen
Application Note 13 RNA Sample Preparation Non-Organic-Based Isolation of Mammalian microrna using Norgen s microrna Purification Kit B. Lam, PhD 1, P. Roberts, MSc 1 Y. Haj-Ahmad, M.Sc., Ph.D 1,2 1 Norgen
Human skin punch biopsies were obtained under informed consent from normal healthy
 SUPPLEMENTAL METHODS Acquisition of human skin specimens. Human skin punch biopsies were obtained under informed consent from normal healthy volunteers (n = 30) and psoriasis patients (n = 45) under a
SUPPLEMENTAL METHODS Acquisition of human skin specimens. Human skin punch biopsies were obtained under informed consent from normal healthy volunteers (n = 30) and psoriasis patients (n = 45) under a
Practice Exam A. Briefly describe how IL-25 treatment might be able to help this responder subgroup of liver cancer patients.
 Practice Exam 2007 1. A special JAK-STAT signaling system (JAK5-STAT5) was recently identified in which a gene called TS5 becomes selectively transcribed and expressed in the liver upon induction by a
Practice Exam 2007 1. A special JAK-STAT signaling system (JAK5-STAT5) was recently identified in which a gene called TS5 becomes selectively transcribed and expressed in the liver upon induction by a
Cellular Reprogramming and Controllability of Complex Systems
 Cellular Reprogramming and Controllability of Complex Systems University of Maryland and ISR April 25, 2016 Roger Brockett John A. Paulson School of Engineering and Applied Science Harvard University 1
Cellular Reprogramming and Controllability of Complex Systems University of Maryland and ISR April 25, 2016 Roger Brockett John A. Paulson School of Engineering and Applied Science Harvard University 1
EPIGENTEK. EpiQuik Chromatin Immunoprecipitation Kit. Base Catalog # P-2002 PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE
 EpiQuik Chromatin Immunoprecipitation Kit Base Catalog # P-2002 PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE The EpiQuik Chromatin Immunoprecipitation Kit is suitable for combining the specificity of
EpiQuik Chromatin Immunoprecipitation Kit Base Catalog # P-2002 PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE The EpiQuik Chromatin Immunoprecipitation Kit is suitable for combining the specificity of
Applied Bioinformatics - Lecture 16: Transcriptomics
 Applied Bioinformatics - Lecture 16: Transcriptomics David Hendrix Oregon State University Feb 15th 2016 Transcriptomics High-throughput Sequencing (deep sequencing) High-throughput sequencing (also
Applied Bioinformatics - Lecture 16: Transcriptomics David Hendrix Oregon State University Feb 15th 2016 Transcriptomics High-throughput Sequencing (deep sequencing) High-throughput sequencing (also
Supplementary Figure 1.
 Supplementary Figure 1. (A) UCSC Genome Browser view of region immediately downstream of the NEUROG3 start codon. All candidate grna target sites which meet the G(N 19 )NGG constraint are aligned to illustrate
Supplementary Figure 1. (A) UCSC Genome Browser view of region immediately downstream of the NEUROG3 start codon. All candidate grna target sites which meet the G(N 19 )NGG constraint are aligned to illustrate
Thyroid hormone transcriptome and cerebellum development.
 Thyroid hormone transcriptome and cerebellum development. 1) Developmental biology: General concepts 2) Genomic data: mouse and human genomes global view 3) A biological problem: mouse cerebellum and thyroid
Thyroid hormone transcriptome and cerebellum development. 1) Developmental biology: General concepts 2) Genomic data: mouse and human genomes global view 3) A biological problem: mouse cerebellum and thyroid
SUPPLEMENTARY DATA. Supplementary Table 1. Sequences of qpcr primers.
 Supplementary Table 1. Sequences of qpcr primers. Lpl Fatp1 (Slc27a1) Fatp4 (Slc27a4) Cd36 Acsl1 Gpam Agpat2 Dgat1 Dgat2 Plin1 Atgl Hsl Mgll Lpin1 Got2 Cav1 Cav2 Fitm2 Nr1h2 Nr1h3 Acsl4 Acsl5 Agpat6 Agpat9
Supplementary Table 1. Sequences of qpcr primers. Lpl Fatp1 (Slc27a1) Fatp4 (Slc27a4) Cd36 Acsl1 Gpam Agpat2 Dgat1 Dgat2 Plin1 Atgl Hsl Mgll Lpin1 Got2 Cav1 Cav2 Fitm2 Nr1h2 Nr1h3 Acsl4 Acsl5 Agpat6 Agpat9
Integrated Omics Study Delineates the Dynamics of Lipid Droplets in Rhodococcus Opacus PD630
 School of Natural Sciences and Mathematics 2013-10-22 Integrated Omics Study Delineates the Dynamics of Lipid Droplets in Rhodococcus Opacus PD630 UTD AUTHOR(S): Michael Qiwei Zhang 2013 The Authors This
School of Natural Sciences and Mathematics 2013-10-22 Integrated Omics Study Delineates the Dynamics of Lipid Droplets in Rhodococcus Opacus PD630 UTD AUTHOR(S): Michael Qiwei Zhang 2013 The Authors This
Functional Genomics Overview RORY STARK PRINCIPAL BIOINFORMATICS ANALYST CRUK CAMBRIDGE INSTITUTE 18 SEPTEMBER 2017
 Functional Genomics Overview RORY STARK PRINCIPAL BIOINFORMATICS ANALYST CRUK CAMBRIDGE INSTITUTE 18 SEPTEMBER 2017 Agenda What is Functional Genomics? RNA Transcription/Gene Expression Measuring Gene
Functional Genomics Overview RORY STARK PRINCIPAL BIOINFORMATICS ANALYST CRUK CAMBRIDGE INSTITUTE 18 SEPTEMBER 2017 Agenda What is Functional Genomics? RNA Transcription/Gene Expression Measuring Gene
Nature Genetics: doi: /ng Supplementary Figure 1. ChIP-seq genome browser views of BRM occupancy at previously identified BRM targets.
 Supplementary Figure 1 ChIP-seq genome browser views of BRM occupancy at previously identified BRM targets. Gene structures are shown underneath each panel. Supplementary Figure 2 pref6::ref6-gfp complements
Supplementary Figure 1 ChIP-seq genome browser views of BRM occupancy at previously identified BRM targets. Gene structures are shown underneath each panel. Supplementary Figure 2 pref6::ref6-gfp complements
Biology 4361 Developmental Biology Lecture 4. The Genetic Core of Development
 Biology 4361 Developmental Biology Lecture 4. The Genetic Core of Development The only way to get from genotype to phenotype is through developmental processes. - Remember the analogy that the zygote contains
Biology 4361 Developmental Biology Lecture 4. The Genetic Core of Development The only way to get from genotype to phenotype is through developmental processes. - Remember the analogy that the zygote contains
Nature Genetics: doi: /ng Supplementary Figure 1
 Supplementary Figure 1 Ihh interacts preferentially with its upstream neighboring gene Nhej1. Genes are indicated by gray lines, and Ihh and Nhej1 are highlighted in blue. 4C seq performed in E14.5 limbs
Supplementary Figure 1 Ihh interacts preferentially with its upstream neighboring gene Nhej1. Genes are indicated by gray lines, and Ihh and Nhej1 are highlighted in blue. 4C seq performed in E14.5 limbs
Efficient Method for Isolation of High Quality Concentrated Cellular RNA with Extremely Low Levels of Genomic DNA Contamination Application
 Efficient Method for Isolation of High Quality Concentrated Cellular RNA with Extremely Low Levels of Genomic DNA Contamination Application Gene Expression Authors Ilgar Abbaszade, Claudia Robbins, John
Efficient Method for Isolation of High Quality Concentrated Cellular RNA with Extremely Low Levels of Genomic DNA Contamination Application Gene Expression Authors Ilgar Abbaszade, Claudia Robbins, John
Expressed genes profiling (Microarrays) Overview Of Gene Expression Control Profiling Of Expressed Genes
 Expressed genes profiling (Microarrays) Overview Of Gene Expression Control Profiling Of Expressed Genes Genes can be regulated at many levels Usually, gene regulation, are referring to transcriptional
Expressed genes profiling (Microarrays) Overview Of Gene Expression Control Profiling Of Expressed Genes Genes can be regulated at many levels Usually, gene regulation, are referring to transcriptional
CAP BIOINFORMATICS Su-Shing Chen CISE. 10/5/2005 Su-Shing Chen, CISE 1
 CAP 5510-9 BIOINFORMATICS Su-Shing Chen CISE 10/5/2005 Su-Shing Chen, CISE 1 Basic BioTech Processes Hybridization PCR Southern blotting (spot or stain) 10/5/2005 Su-Shing Chen, CISE 2 10/5/2005 Su-Shing
CAP 5510-9 BIOINFORMATICS Su-Shing Chen CISE 10/5/2005 Su-Shing Chen, CISE 1 Basic BioTech Processes Hybridization PCR Southern blotting (spot or stain) 10/5/2005 Su-Shing Chen, CISE 2 10/5/2005 Su-Shing
From Genotype to Phenotype
 From Genotype to Phenotype Johanna Vilkki Green technology, Natural Resources Institute Finland Systems biology Genome Transcriptome genes mrna Genotyping methodology SNP TOOLS, WG SEQUENCING Functional
From Genotype to Phenotype Johanna Vilkki Green technology, Natural Resources Institute Finland Systems biology Genome Transcriptome genes mrna Genotyping methodology SNP TOOLS, WG SEQUENCING Functional
Nature Biotechnology: doi: /nbt Supplementary Figure 1
 Supplementary Figure 1 Schematic and results of screening the combinatorial antibody library for Sox2 replacement activity. A single batch of MEFs were plated and transduced with doxycycline inducible
Supplementary Figure 1 Schematic and results of screening the combinatorial antibody library for Sox2 replacement activity. A single batch of MEFs were plated and transduced with doxycycline inducible
Chapter 10 Genetic Engineering: A Revolution in Molecular Biology
 Chapter 10 Genetic Engineering: A Revolution in Molecular Biology Genetic Engineering Direct, deliberate modification of an organism s genome bioengineering Biotechnology use of an organism s biochemical
Chapter 10 Genetic Engineering: A Revolution in Molecular Biology Genetic Engineering Direct, deliberate modification of an organism s genome bioengineering Biotechnology use of an organism s biochemical
Outline and learning objectives. From Proteomics to Systems Biology. Integration of omics - information
 From to Systems Biology Outline and learning objectives Omics science provides global analysis tools to study entire systems How to obtain omics - What can we learn Limitations Integration of omics - In-class
From to Systems Biology Outline and learning objectives Omics science provides global analysis tools to study entire systems How to obtain omics - What can we learn Limitations Integration of omics - In-class
Illumina Genome Analyzer. Progenika Experience. - Susana Catarino -
 Illumina Genome Analyzer Progenika Experience - Susana Catarino - Who are we? 2000 PROGENIKA BIOPHARMA Development, production and commercialization of new genomic tools for diagnosis, prognosis and drug-response
Illumina Genome Analyzer Progenika Experience - Susana Catarino - Who are we? 2000 PROGENIKA BIOPHARMA Development, production and commercialization of new genomic tools for diagnosis, prognosis and drug-response
Corn ChIPs and RNA-seq: Researchers Dip into Advanced Tools and Resources to Examine bzip Transcription Factor Function in the Maize Endosperm
 Plant Cell Advance Publication. Published on November 21, 2018, doi:10.1105/tpc.18.00882 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 41
Plant Cell Advance Publication. Published on November 21, 2018, doi:10.1105/tpc.18.00882 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 41
Pioneering Clinical Omics
 Pioneering Clinical Omics Clinical Genomics Strand NGS An analysis tool for data generated by cutting-edge Next Generation Sequencing(NGS) instruments. Strand NGS enables read alignment and analysis of
Pioneering Clinical Omics Clinical Genomics Strand NGS An analysis tool for data generated by cutting-edge Next Generation Sequencing(NGS) instruments. Strand NGS enables read alignment and analysis of
Name AP Biology Mrs. Laux Take home test #11 on Chapters 14, 15, and 17 DUE: MONDAY, DECEMBER 21, 2009
 MULTIPLE CHOICE QUESTIONS 1. Inducible genes are usually actively transcribed when: A. the molecule degraded by the enzyme(s) is present in the cell. B. repressor molecules bind to the promoter. C. lactose
MULTIPLE CHOICE QUESTIONS 1. Inducible genes are usually actively transcribed when: A. the molecule degraded by the enzyme(s) is present in the cell. B. repressor molecules bind to the promoter. C. lactose
Gene Expression Technology
 Gene Expression Technology Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Gene expression Gene expression is the process by which information from a gene
Gene Expression Technology Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Gene expression Gene expression is the process by which information from a gene
From Proteomics to Systems Biology. Integration of omics - information
 From Proteomics to Systems Biology Integration of omics - information Outline and learning objectives Omics science provides global analysis tools to study entire systems How to obtain omics - data What
From Proteomics to Systems Biology Integration of omics - information Outline and learning objectives Omics science provides global analysis tools to study entire systems How to obtain omics - data What
Intracellular receptors specify complex patterns of gene expression that are cell and gene
 SUPPLEMENTAL RESULTS AND DISCUSSION Some HPr-1AR ARE-containing Genes Are Unresponsive to Androgen Intracellular receptors specify complex patterns of gene expression that are cell and gene specific. For
SUPPLEMENTAL RESULTS AND DISCUSSION Some HPr-1AR ARE-containing Genes Are Unresponsive to Androgen Intracellular receptors specify complex patterns of gene expression that are cell and gene specific. For
Department of Genetics
 Department of Genetics Program Specific Outcomes (PSO) Program- MSc. Genetics (404) The course prepares students for pursuing further research and teaching. Key features: Students have high success rate
Department of Genetics Program Specific Outcomes (PSO) Program- MSc. Genetics (404) The course prepares students for pursuing further research and teaching. Key features: Students have high success rate
Biology 644: Bioinformatics
 Processes Activation Repression Initiation Elongation.... Processes Splicing Editing Degradation Translation.... Transcription Translation DNA Regulators DNA-Binding Transcription Factors Chromatin Remodelers....
Processes Activation Repression Initiation Elongation.... Processes Splicing Editing Degradation Translation.... Transcription Translation DNA Regulators DNA-Binding Transcription Factors Chromatin Remodelers....
ChIP-seq data analysis with Chipster. Eija Korpelainen CSC IT Center for Science, Finland
 ChIP-seq data analysis with Chipster Eija Korpelainen CSC IT Center for Science, Finland chipster@csc.fi What will I learn? Short introduction to ChIP-seq Analyzing ChIP-seq data Central concepts Analysis
ChIP-seq data analysis with Chipster Eija Korpelainen CSC IT Center for Science, Finland chipster@csc.fi What will I learn? Short introduction to ChIP-seq Analyzing ChIP-seq data Central concepts Analysis
A Genome wide RNAi Screen. in Human Cells. Eric E. Zhang,1,2,5 Andrew C. Liu,1,3,5 Tsuyoshi Hirota,1,2,5 Loren J. Miraglia,1 Genevieve Welch,1
 A Genome wide RNAi Screen for Modifiers of the Circadian Clock in Human Cells Eric E. Zhang,1,2,5 Andrew C. Liu,1,3,5 Tsuyoshi Hirota,1,2,5 Loren J. Miraglia,1 Genevieve Welch,1 Pagkapol Y. Pongsawakul,2
A Genome wide RNAi Screen for Modifiers of the Circadian Clock in Human Cells Eric E. Zhang,1,2,5 Andrew C. Liu,1,3,5 Tsuyoshi Hirota,1,2,5 Loren J. Miraglia,1 Genevieve Welch,1 Pagkapol Y. Pongsawakul,2
NGS to address ncrna and viruses
 NGS to address ncrna and viruses Introduction & TRON Next generation sequencing transcriptomics ncrnas vrna June 30, 2010 John Castle Institute for Translational Oncology and Immunology (TRON) Mainz, Germany
NGS to address ncrna and viruses Introduction & TRON Next generation sequencing transcriptomics ncrnas vrna June 30, 2010 John Castle Institute for Translational Oncology and Immunology (TRON) Mainz, Germany
Honours and PostDoctorate Research Projects. Lions Eye Institute, Perth
 Honours and PostDoctorate Research Projects Available from 1 st Semester, 2018 Lions Eye Institute, Perth 1 About the Lions Eye Institute. The Lions Eye Institute is a not-for-profit centre of excellence
Honours and PostDoctorate Research Projects Available from 1 st Semester, 2018 Lions Eye Institute, Perth 1 About the Lions Eye Institute. The Lions Eye Institute is a not-for-profit centre of excellence
THE ANALYSIS OF CHROMATIN CONDENSATION STATE AND TRANSCRIPTIONAL ACTIVITY USING DNA MICROARRAYS 1. INTRODUCTION
 JOURNAL OF MEDICAL INFORMATICS & TECHNOLOGIES Vol.6/2003, ISSN 1642-6037 Piotr WIDŁAK *, Krzysztof FUJAREWICZ ** DNA microarrays, chromatin, transcription THE ANALYSIS OF CHROMATIN CONDENSATION STATE AND
JOURNAL OF MEDICAL INFORMATICS & TECHNOLOGIES Vol.6/2003, ISSN 1642-6037 Piotr WIDŁAK *, Krzysztof FUJAREWICZ ** DNA microarrays, chromatin, transcription THE ANALYSIS OF CHROMATIN CONDENSATION STATE AND
Tissue Acetyl-Histone H4 ChIP Kit
 Tissue Acetyl-Histone H4 ChIP Kit Catalog Number KA0672 48 assays Version: 03 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Intended Use... 3 Background... 3 Principle
Tissue Acetyl-Histone H4 ChIP Kit Catalog Number KA0672 48 assays Version: 03 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Intended Use... 3 Background... 3 Principle
1. why study multiple traits together?
 Multiple Traits & Microarrays 1. why study multiple traits together? 2-10 diabetes case study 2. design issues 11-13 selective phenotyping 3. why are traits correlated? 14-17 close linkage or pleiotropy?
Multiple Traits & Microarrays 1. why study multiple traits together? 2-10 diabetes case study 2. design issues 11-13 selective phenotyping 3. why are traits correlated? 14-17 close linkage or pleiotropy?
1. Introduction. 1 Page 1
 1. INTRODUCTION Osteoblast differentiation is the key component of the bone formation, growth and remodeling. During this incompletely understood process a subpopulation of multipotential mesenchymal stem
1. INTRODUCTION Osteoblast differentiation is the key component of the bone formation, growth and remodeling. During this incompletely understood process a subpopulation of multipotential mesenchymal stem
Submitting Samples to the KOMP Analysis Pipeline Date: 8/12/2017
 Submitting Samples to the KOMP Analysis Pipeline Date: 8/12/2017 We want to encourage investigators who have developed or studied mice that have abnormalities in their skeletal health to submit these animals
Submitting Samples to the KOMP Analysis Pipeline Date: 8/12/2017 We want to encourage investigators who have developed or studied mice that have abnormalities in their skeletal health to submit these animals
FY10 INVITATION PROPOSAL SUMMARY PAGE
 FY10 INVITATION PROPOSAL SUMMARY PAGE Project Title: Pilot Sequence: High Density DNA Sequencing to Detect Population Structure of Pacific Herring (Project Number: 10100165) Project Period: October 1,
FY10 INVITATION PROPOSAL SUMMARY PAGE Project Title: Pilot Sequence: High Density DNA Sequencing to Detect Population Structure of Pacific Herring (Project Number: 10100165) Project Period: October 1,
Supplementary Table 1. Primers used to construct full-length or various truncated mutants of ISG12b2.
 Supplementary Table 1. Primers used to construct full-length or various truncated mutants of ISG12b2. Construct name ISG12b2 (No tag) HA-ISG12b2 (N-HA) ISG12b2-HA (C-HA; FL-HA) 94-283-HA (FL-GFP) 93-GFP
Supplementary Table 1. Primers used to construct full-length or various truncated mutants of ISG12b2. Construct name ISG12b2 (No tag) HA-ISG12b2 (N-HA) ISG12b2-HA (C-HA; FL-HA) 94-283-HA (FL-GFP) 93-GFP
Supplemental Information
 Supplemental Information Itemized List Materials and Methods, Related to Supplemental Figures S5A-C and S6. Supplemental Figure S1, Related to Figures 1 and 2. Supplemental Figure S2, Related to Figure
Supplemental Information Itemized List Materials and Methods, Related to Supplemental Figures S5A-C and S6. Supplemental Figure S1, Related to Figures 1 and 2. Supplemental Figure S2, Related to Figure
The Power to Cure: Therapeutic Innovation in Academia
 The Power to Cure: Therapeutic Innovation in Academia Leslie Molony, Ph.D. November 7, 2011 Sanford-Burnham Medical Research Institute: a Non Profit Basic Research Institute Santa Barbara Est. 2006 La
The Power to Cure: Therapeutic Innovation in Academia Leslie Molony, Ph.D. November 7, 2011 Sanford-Burnham Medical Research Institute: a Non Profit Basic Research Institute Santa Barbara Est. 2006 La
The RRPA knock-in allele was generated by homologous recombination in TC1 ES cells.
 Supplemental Materials Materials & Methods Generation of RRPA and RAPA Knock-in Mice The RRPA knock-in allele was generated by homologous recombination in TC1 ES cells. Targeted ES clones in which the
Supplemental Materials Materials & Methods Generation of RRPA and RAPA Knock-in Mice The RRPA knock-in allele was generated by homologous recombination in TC1 ES cells. Targeted ES clones in which the
Supplementary Figure 1. TSA (10 nmol/l), non-class-selective HDAC inhibitor, potentiates
 Supplementary Figure 1. TSA (10 nmol/l), non-class-selective HDAC inhibitor, potentiates vascular calcification (VC). (a) Von Kossa staining shows that TSA potentiated the Pi-induced VC. Scale bar, 100
Supplementary Figure 1. TSA (10 nmol/l), non-class-selective HDAC inhibitor, potentiates vascular calcification (VC). (a) Von Kossa staining shows that TSA potentiated the Pi-induced VC. Scale bar, 100
At E17.5, the embryos were rinsed in phosphate-buffered saline (PBS) and immersed in
 Supplementary Materials and Methods Barrier function assays At E17.5, the embryos were rinsed in phosphate-buffered saline (PBS) and immersed in acidic X-gal mix (100 mm phosphate buffer at ph4.3, 3 mm
Supplementary Materials and Methods Barrier function assays At E17.5, the embryos were rinsed in phosphate-buffered saline (PBS) and immersed in acidic X-gal mix (100 mm phosphate buffer at ph4.3, 3 mm
Nature Methods: doi: /nmeth Supplementary Figure 1
 Supplementary Figure 1 Workflow for multimodal analysis using sc-gem on a programmable microfluidic device (Fluidigm). 1) Cells are captured and lysed, 2) RNA from lysed single cell is reverse-transcribed
Supplementary Figure 1 Workflow for multimodal analysis using sc-gem on a programmable microfluidic device (Fluidigm). 1) Cells are captured and lysed, 2) RNA from lysed single cell is reverse-transcribed
GENE EXPRESSION REAGENTS MARKETS (SAMPLE COPY, NOT FOR RESALE)
 TriMark Publications April 2007 Volume: TMRGER07-0401 GENE EXPRESSION REAGENTS MARKETS (SAMPLE COPY, NOT FOR RESALE) Trends, Industry Participants, Product Overviews and Market Drivers TABLE OF CONTENTS
TriMark Publications April 2007 Volume: TMRGER07-0401 GENE EXPRESSION REAGENTS MARKETS (SAMPLE COPY, NOT FOR RESALE) Trends, Industry Participants, Product Overviews and Market Drivers TABLE OF CONTENTS
Applicazioni biotecnologiche
 Applicazioni biotecnologiche Analisi forense Sintesi di proteine ricombinanti Restriction Fragment Length Polymorphism (RFLP) Polymorphism (more fully genetic polymorphism) refers to the simultaneous occurrence
Applicazioni biotecnologiche Analisi forense Sintesi di proteine ricombinanti Restriction Fragment Length Polymorphism (RFLP) Polymorphism (more fully genetic polymorphism) refers to the simultaneous occurrence
Introduction to Bioinformatics and Gene Expression Technologies
 Introduction to Bioinformatics and Gene Expression Technologies Utah State University Fall 2017 Statistical Bioinformatics (Biomedical Big Data) Notes 1 1 Vocabulary Gene: hereditary DNA sequence at a
Introduction to Bioinformatics and Gene Expression Technologies Utah State University Fall 2017 Statistical Bioinformatics (Biomedical Big Data) Notes 1 1 Vocabulary Gene: hereditary DNA sequence at a
Introduction to Bioinformatics and Gene Expression Technologies
 Vocabulary Introduction to Bioinformatics and Gene Expression Technologies Utah State University Fall 2017 Statistical Bioinformatics (Biomedical Big Data) Notes 1 Gene: Genetics: Genome: Genomics: hereditary
Vocabulary Introduction to Bioinformatics and Gene Expression Technologies Utah State University Fall 2017 Statistical Bioinformatics (Biomedical Big Data) Notes 1 Gene: Genetics: Genome: Genomics: hereditary
This place covers: Methods or systems for genetic or protein-related data processing in computational molecular biology.
 G16B BIOINFORMATICS, i.e. INFORMATION AND COMMUNICATION TECHNOLOGY [ICT] SPECIALLY ADAPTED FOR GENETIC OR PROTEIN-RELATED DATA PROCESSING IN COMPUTATIONAL MOLECULAR BIOLOGY Methods or systems for genetic
G16B BIOINFORMATICS, i.e. INFORMATION AND COMMUNICATION TECHNOLOGY [ICT] SPECIALLY ADAPTED FOR GENETIC OR PROTEIN-RELATED DATA PROCESSING IN COMPUTATIONAL MOLECULAR BIOLOGY Methods or systems for genetic
SUPPLEMENTARY INFORMATION
 Ca 2+ /calmodulin Regulates Salicylic Acid-mediated Plant Immunity Liqun Du, Gul S. Ali, Kayla A. Simons, Jingguo Hou, Tianbao Yang, A.S.N. Reddy and B. W. Poovaiah * *To whom correspondence should be
Ca 2+ /calmodulin Regulates Salicylic Acid-mediated Plant Immunity Liqun Du, Gul S. Ali, Kayla A. Simons, Jingguo Hou, Tianbao Yang, A.S.N. Reddy and B. W. Poovaiah * *To whom correspondence should be
CHAPTER 21 LECTURE SLIDES
 CHAPTER 21 LECTURE SLIDES Prepared by Brenda Leady University of Toledo To run the animations you must be in Slideshow View. Use the buttons on the animation to play, pause, and turn audio/text on or off.
CHAPTER 21 LECTURE SLIDES Prepared by Brenda Leady University of Toledo To run the animations you must be in Slideshow View. Use the buttons on the animation to play, pause, and turn audio/text on or off.
MeCP2. MeCP2/α-tubulin. GFP mir1-1 mir132
 Conservation Figure S1. Schematic showing 3 UTR (top; thick black line), mir132 MRE (arrow) and nucleotide sequence conservation (vertical black lines; http://genome.ucsc.edu). a GFP mir1-1 mir132 b GFP
Conservation Figure S1. Schematic showing 3 UTR (top; thick black line), mir132 MRE (arrow) and nucleotide sequence conservation (vertical black lines; http://genome.ucsc.edu). a GFP mir1-1 mir132 b GFP
a rj13 ORNLf CP CONF-~~/!CI~~--The Functional Genomics Initiative at Oak Ridge National Laboratory*
 I I d ORNLf CP-94806 CONF-~~/!CI~~--The Functional Genomics Initiative at Oak Ridge National Laboratory* Dabney Johnson t, Monica Justice t; Ken Beattie t, Michelle Buchanan $, Michael Ramsey 0,Rose Ramsey
I I d ORNLf CP-94806 CONF-~~/!CI~~--The Functional Genomics Initiative at Oak Ridge National Laboratory* Dabney Johnson t, Monica Justice t; Ken Beattie t, Michelle Buchanan $, Michael Ramsey 0,Rose Ramsey
Introduction to Bioinformatics
 Introduction to Bioinformatics If the 19 th century was the century of chemistry and 20 th century was the century of physic, the 21 st century promises to be the century of biology...professor Dr. Satoru
Introduction to Bioinformatics If the 19 th century was the century of chemistry and 20 th century was the century of physic, the 21 st century promises to be the century of biology...professor Dr. Satoru
Supplemental Figure 1. Study flow chart.
 Supplemental Figure 1. Study flow chart. 1 Supplemental Figure 2. Histograms represent fold changes of mir-22 in HL-1 cells. Vehicle-treated (gray) or sildenafil-treated (black b) cells. Results are expressed
Supplemental Figure 1. Study flow chart. 1 Supplemental Figure 2. Histograms represent fold changes of mir-22 in HL-1 cells. Vehicle-treated (gray) or sildenafil-treated (black b) cells. Results are expressed
Methods of Biomaterials Testing Lesson 3-5. Biochemical Methods - Molecular Biology -
 Methods of Biomaterials Testing Lesson 3-5 Biochemical Methods - Molecular Biology - Chromosomes in the Cell Nucleus DNA in the Chromosome Deoxyribonucleic Acid (DNA) DNA has double-helix structure The
Methods of Biomaterials Testing Lesson 3-5 Biochemical Methods - Molecular Biology - Chromosomes in the Cell Nucleus DNA in the Chromosome Deoxyribonucleic Acid (DNA) DNA has double-helix structure The
REPRODUCTIVE HEALTH. Phosphorus. Diagnostics
 14 32 6 32 7 REPRODUCTIVE HEALTH Phosphorus Diagnostics Genomics, Illuminated We focus on your peace of mind. Our expert-guided assay designs, powerful data analytics, and rigorous variant interpretation
14 32 6 32 7 REPRODUCTIVE HEALTH Phosphorus Diagnostics Genomics, Illuminated We focus on your peace of mind. Our expert-guided assay designs, powerful data analytics, and rigorous variant interpretation
Supplemental Data. Na Xu et al. (2016). Plant Cell /tpc
 Supplemental Figure 1. The weak fluorescence phenotype is not caused by the mutation in At3g60240. (A) A mutation mapped to the gene At3g60240. Map-based cloning strategy was used to map the mutated site
Supplemental Figure 1. The weak fluorescence phenotype is not caused by the mutation in At3g60240. (A) A mutation mapped to the gene At3g60240. Map-based cloning strategy was used to map the mutated site
Dingjun Hao, Baorong He, Liang Yan. Hong Hui Hospital, Xi an Jiaotong University College of. Medicine, Xi an, Shaanxi , China
 Xi an Hong Hui Hospital Xi an, Shaanxi, China A single nucleotide polymorphism in the human bone morphogenetic protein-2 gene (109T>G) affects the Smad signaling pathway and the predisposition to ossification
Xi an Hong Hui Hospital Xi an, Shaanxi, China A single nucleotide polymorphism in the human bone morphogenetic protein-2 gene (109T>G) affects the Smad signaling pathway and the predisposition to ossification
Supplementary Data. Supplementary Methods Three-step protocol for spontaneous differentiation of mouse induced pluripotent stem (embryonic stem) cells
 Supplementary Data Supplementary Methods Three-step protocol for spontaneous differentiation of mouse induced pluripotent stem (embryonic stem) cells Mouse induced pluripotent stem cells (ipscs) were cultured
Supplementary Data Supplementary Methods Three-step protocol for spontaneous differentiation of mouse induced pluripotent stem (embryonic stem) cells Mouse induced pluripotent stem cells (ipscs) were cultured
SUPPLEMENTARY INFORMATION
 AS-NMD modulates FLM-dependent thermosensory flowering response in Arabidopsis NATURE PLANTS www.nature.com/natureplants 1 Supplementary Figure 1. Genomic sequence of FLM along with the splice sites. Sequencing
AS-NMD modulates FLM-dependent thermosensory flowering response in Arabidopsis NATURE PLANTS www.nature.com/natureplants 1 Supplementary Figure 1. Genomic sequence of FLM along with the splice sites. Sequencing
BIOLOGY Dr.Locke Lecture# 27 An Introduction to Polymerase Chain Reaction (PCR)
 BIOLOGY 207 - Dr.Locke Lecture# 27 An Introduction to Polymerase Chain Reaction (PCR) Required readings and problems: Reading: Open Genetics, Chapter 8.1 Problems: Chapter 8 Optional Griffiths (2008) 9
BIOLOGY 207 - Dr.Locke Lecture# 27 An Introduction to Polymerase Chain Reaction (PCR) Required readings and problems: Reading: Open Genetics, Chapter 8.1 Problems: Chapter 8 Optional Griffiths (2008) 9
Guided Notes Unit 5: Molecular Genetics
 Name: Date: Block: Chapter 8: From DNA to Protein I. Concept 8.4: Transcription a. Central Dogma of Molecular Biology i. Information flows in one direction: ii. How? Guided Notes Unit 5: Molecular Genetics
Name: Date: Block: Chapter 8: From DNA to Protein I. Concept 8.4: Transcription a. Central Dogma of Molecular Biology i. Information flows in one direction: ii. How? Guided Notes Unit 5: Molecular Genetics
Technical Review. Real time PCR
 Technical Review Real time PCR Normal PCR: Analyze with agarose gel Normal PCR vs Real time PCR Real-time PCR, also known as quantitative PCR (qpcr) or kinetic PCR Key feature: Used to amplify and simultaneously
Technical Review Real time PCR Normal PCR: Analyze with agarose gel Normal PCR vs Real time PCR Real-time PCR, also known as quantitative PCR (qpcr) or kinetic PCR Key feature: Used to amplify and simultaneously
Gene Expression on the Fluidigm BioMark HD
 Gene Expression on the Fluidigm BioMark HD Overview Introduction to Fluidigm James Miller Advantages of the technology Running a Fluidigm gene expression project Paul Lacaze Assay design, chemistry, experimental
Gene Expression on the Fluidigm BioMark HD Overview Introduction to Fluidigm James Miller Advantages of the technology Running a Fluidigm gene expression project Paul Lacaze Assay design, chemistry, experimental
Progress and Future Directions in Integrated Systems Toxicology. Mary McBride Agilent Technologies
 Progress and Future Directions in Integrated Systems Toxicology Mary McBride Agilent Technologies 1 Toxicity testing tools of the late 20 th century Patchwork approach to testing dates back to the 1930
Progress and Future Directions in Integrated Systems Toxicology Mary McBride Agilent Technologies 1 Toxicity testing tools of the late 20 th century Patchwork approach to testing dates back to the 1930
CURRICULUM VITAE. Educational Qualification: DEGREE INSTITUTION YEAR PROGRESS. Bharathidasan University Tamil Nadu
 CURRICULUM VITAE M.BHARATH VIJAYARAGAVAN Masters in Biomedical sciences Nationality: Indian E-mail: bharathragavan@gmail.com Mobile: 91-9911738539 Sex: Male; Age: 23 Objective: Cell Signaling is an important
CURRICULUM VITAE M.BHARATH VIJAYARAGAVAN Masters in Biomedical sciences Nationality: Indian E-mail: bharathragavan@gmail.com Mobile: 91-9911738539 Sex: Male; Age: 23 Objective: Cell Signaling is an important
Novel methods for RNA and DNA- Seq analysis using SMART Technology. Andrew Farmer, D. Phil. Vice President, R&D Clontech Laboratories, Inc.
 Novel methods for RNA and DNA- Seq analysis using SMART Technology Andrew Farmer, D. Phil. Vice President, R&D Clontech Laboratories, Inc. Agenda Enabling Single Cell RNA-Seq using SMART Technology SMART
Novel methods for RNA and DNA- Seq analysis using SMART Technology Andrew Farmer, D. Phil. Vice President, R&D Clontech Laboratories, Inc. Agenda Enabling Single Cell RNA-Seq using SMART Technology SMART
Motivation From Protein to Gene
 MOLECULAR BIOLOGY 2003-4 Topic B Recombinant DNA -principles and tools Construct a library - what for, how Major techniques +principles Bioinformatics - in brief Chapter 7 (MCB) 1 Motivation From Protein
MOLECULAR BIOLOGY 2003-4 Topic B Recombinant DNA -principles and tools Construct a library - what for, how Major techniques +principles Bioinformatics - in brief Chapter 7 (MCB) 1 Motivation From Protein
A Repressor Complex Governs the Integration of
 Developmental Cell 15 Supplemental Data A Repressor Complex Governs the Integration of Flowering Signals in Arabidopsis Dan Li, Chang Liu, Lisha Shen, Yang Wu, Hongyan Chen, Masumi Robertson, Chris A.
Developmental Cell 15 Supplemental Data A Repressor Complex Governs the Integration of Flowering Signals in Arabidopsis Dan Li, Chang Liu, Lisha Shen, Yang Wu, Hongyan Chen, Masumi Robertson, Chris A.
RESEARCH OPPORTUNITY PROGRAM 299Y/399Y PROJECT DESCRIPTIONS SUMMER
 Project Code: MGY 1S RESEARCH OPPORTUNITY PROGRAM 299Y/399Y PROJECT DESCRIPTIONS 2018 2019 SUMMER Name and Title: Department: Amy A. Caudy, Associate Professor Molecular Genetics TITLE OF RESEARCH PROJECT:
Project Code: MGY 1S RESEARCH OPPORTUNITY PROGRAM 299Y/399Y PROJECT DESCRIPTIONS 2018 2019 SUMMER Name and Title: Department: Amy A. Caudy, Associate Professor Molecular Genetics TITLE OF RESEARCH PROJECT:
Multiple Traits & Microarrays
 Multiple Traits & Microarrays 1. why study multiple traits together? 2-10 diabetes case study 2. design issues 11-13 selective phenotyping 3. why are traits correlated? 14-17 close linkage or pleiotropy?
Multiple Traits & Microarrays 1. why study multiple traits together? 2-10 diabetes case study 2. design issues 11-13 selective phenotyping 3. why are traits correlated? 14-17 close linkage or pleiotropy?
Supplementary Figure 1. Homozygous rag2 E450fs mutants are healthy and viable similar to wild-type and heterozygous siblings.
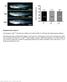 Supplementary Figure 1 Homozygous rag2 E450fs mutants are healthy and viable similar to wild-type and heterozygous siblings. (left) Representative bright-field images of wild type (wt), heterozygous (het)
Supplementary Figure 1 Homozygous rag2 E450fs mutants are healthy and viable similar to wild-type and heterozygous siblings. (left) Representative bright-field images of wild type (wt), heterozygous (het)
Supplementary information. Generation of heterozygous fibrillin-1 mutant cloned pigs from genome-edited foetal fibroblasts.
 Supplementary information Generation of heterozygous fibrillin-1 mutant cloned pigs from genome-edited foetal fibroblasts. Kazuhiro Umeyama 1,2, Kota Watanabe 3, Masahito Watanabe 1,2, Keisuke Horiuchi
Supplementary information Generation of heterozygous fibrillin-1 mutant cloned pigs from genome-edited foetal fibroblasts. Kazuhiro Umeyama 1,2, Kota Watanabe 3, Masahito Watanabe 1,2, Keisuke Horiuchi
Chromatin immunoprecipitation: five steps to great results
 Chromatin immunoprecipitation: five steps to great results Introduction The discovery and use of antibodies in life science research has been critical to many advancements across applications, including
Chromatin immunoprecipitation: five steps to great results Introduction The discovery and use of antibodies in life science research has been critical to many advancements across applications, including
4 Mutant Hunts - To Select or to Screen (Perhaps Even by Brute Force)
 Genetic Techniques for Biological Research Corinne A. Michels Copyright q 2002 John Wiley & Sons, Ltd ISBNs: 0-471-89921-6 (Hardback); 0-470-84662-3 (Electronic) 4 Mutant Hunts - To Select or to Screen
Genetic Techniques for Biological Research Corinne A. Michels Copyright q 2002 John Wiley & Sons, Ltd ISBNs: 0-471-89921-6 (Hardback); 0-470-84662-3 (Electronic) 4 Mutant Hunts - To Select or to Screen
Resources Available through the Albert Einstein College of Medicine Nathan Shock Center
 Resources Available through the Albert Einstein College of Medicine Nathan Shock Center http://www.einstein.yu.edu/centers/aging/ Available Proteostasis-related plasmids Sent as filter spotted DNA - shrna
Resources Available through the Albert Einstein College of Medicine Nathan Shock Center http://www.einstein.yu.edu/centers/aging/ Available Proteostasis-related plasmids Sent as filter spotted DNA - shrna
BIOTECHNOLOGY. Unit 8
 BIOTECHNOLOGY Unit 8 PART 1 BASIC/FUNDAMENTAL SCIENCE VS. APPLIED SCIENCE! Basic/Fundamental Science the development and establishment of information to aid our understanding of the world.! Applied Science
BIOTECHNOLOGY Unit 8 PART 1 BASIC/FUNDAMENTAL SCIENCE VS. APPLIED SCIENCE! Basic/Fundamental Science the development and establishment of information to aid our understanding of the world.! Applied Science
Supplement figure legends
 Supplement figure legends Figure S1. Alignments of PRDM16 amino acid sequences of mouse, human, dog, chicken, and Xenopus. Two putative CtBP-binding motifs are indicated in the boxes. Numbers represent
Supplement figure legends Figure S1. Alignments of PRDM16 amino acid sequences of mouse, human, dog, chicken, and Xenopus. Two putative CtBP-binding motifs are indicated in the boxes. Numbers represent
