Genome Annotation. Stefan Prost 1. May 27th, States of America. Genome Annotation
|
|
|
- Emory Young
- 6 years ago
- Views:
Transcription
1 Genome Annotation Stefan Prost 1 1 Department of Integrative Biology, University of California, Berkeley, United States of America May 27th, 2015
2 Outline Genome Annotation 1 Repeat Annotation 2
3 Repeat Annotation Repeat Annotation 1 Mask Repeats for 2 Study Repeat Content and Evolution
4 Repeat Annotation Low Complexity Sequences Simple Repeats and Satellites Transposable Elements Class I: Retrotransposons Copy and Paste Class II: DNA Transposons Cut and Paste Hermann et al. 2013
5 Repeat Annotation Class I TEs Long Terminal Repeats (LTR) Endogenous Retrovirus Non-LTR Long Interspersed Nuclear Elements (LINEs) Short Interspersed Nuclear Elements (SINEs) Class II TEs Subclass I (both DNA strands are cleaved) Subclass II (Rolling-circle, self-synthesizing)
6 Repeat Annotation
7 Fishes Repeat Annotation Mammals B Repeat Content (%) Genome Size (Gb)
8 Repeat Annotation Homology RepeatMasker ( De novo RepeatModeler ( WindowMasker (Morgulis et al., 2005) RepeatScout (Price et al., 2005) Piler (Edgar and Myers, 2005) De novo from reads REPdenovo (github) Tedna (Zytnicki et al., 2014) CAVE: De novo tools can wrongly identify highly conserved protein-coding genes
9 Repeat Annotation For 1 RepeatMasker -pa xx -gccalc -nolow -species aves genome.fasta 2 BuildDatabase -name genome genome.fasta.masked 3 RepeatModeler -database genome 4 RepeatMasker -pa xx -gccalc -nolow -lib consensi.fa.classified genome.fasta[.masked] For Repeat Annotation 1 RepeatMasker -pa xx -a -gccalc -species aves genome.fasta 2 RepeatMasker -pa xx -a -gccalc -lib consensi.fa.classified genome.fasta[.masked]
10 Gene Structure (Marina Axelson-Fisk, Springer-Verlag London, 2015)
11 1 Evidence Alignment Protein EST RNA-seq 2 Ab initio Prediction
12 Ab Initio Gene Prediction Do not need evidence data Need to be trained for the organism (codon frequencies, distribution of exon-intron length, etc.) Most find single most likely coding sequence (CDS) Do not report untranslated regions (UTRs) Cannot deal with alternative splicing Accuracy usually 60-70% With training can be up to 100% Need a very good genome assembly
13
14
15 Definition: Sensitivity Sensitivity (SN) is the fraction of the reference feature that is predicted by the gene predictor. SN = TP/(TP + FN) Definition: Specificity Specificity (SP) is the fraction of the prediction overlapping the reference feature. SP = TP/(TP + FP) Definition: Accuracy SN and SP are often combined into a single measure called accuracy (AC)
16 Definition: Annotation Edit Distance Annotation Edit Distance (AED) is calculated in the same manner as SN and SP, but in place of a reference gene model, the coordinates of the union of the aligned evidence are used instead: AED = 1 AC, where AC = (SN + SP)/ indicates perfect agreement with evidence 1... indicates no evidence support for annotation
17
18 Pipelines Maker2 (Holt and Yandell, 2011) Pasa (Haas et al. 2003) Ensembl (Curewen et al., 2004) NCBI (Kitts, 2003) Evidence Mapping BLAST / BLAT Exonerate (Slater and Birey, 2005) Ab Initio Gene Predictors Augustus (Stanke et al., 2006) SNAP (Korf 2004) GeneMark-ES (Zu et al., 2010) Choosers and Combiners JIGSAW (Allen and Salzberg, 2005) GLEAN (Elsik et al., 2007)
19 Visualization / Curation Artemis (Rutherford et al., 2000) Apollo / WebApollo (Lewis et al., 2002) JBROWSE (Skinner at al., 2009) IGV (Robinson et al., 2011)
20 Maker 2
MAKER: An easy to use genome annotation pipeline. Carson Holt Yandell Lab Department of Human Genetics University of Utah
 MAKER: An easy to use genome annotation pipeline Carson Holt Yandell Lab Department of Human Genetics University of Utah Introduction to Genome Annotation What annotations are Importance of genome annotations
MAKER: An easy to use genome annotation pipeline Carson Holt Yandell Lab Department of Human Genetics University of Utah Introduction to Genome Annotation What annotations are Importance of genome annotations
Genome annotation. Erwin Datema (2011) Sandra Smit (2012, 2013)
 Genome annotation Erwin Datema (2011) Sandra Smit (2012, 2013) Genome annotation AGACAAAGATCCGCTAAATTAAATCTGGACTTCACATATTGAAGTGATATCACACGTTTCTCTAAT AATCTCCTCACAATATTATGTTTGGGATGAACTTGTCGTGATTTGCCATTGTAGCAATCACTTGAA
Genome annotation Erwin Datema (2011) Sandra Smit (2012, 2013) Genome annotation AGACAAAGATCCGCTAAATTAAATCTGGACTTCACATATTGAAGTGATATCACACGTTTCTCTAAT AATCTCCTCACAATATTATGTTTGGGATGAACTTGTCGTGATTTGCCATTGTAGCAATCACTTGAA
GENOME ANNOTATION INTRODUCTION TO CONCEPTS AND METHODS. Olivier GARSMEUR & Stéphanie SIDIBE-BOCS
 GENOME ANNOTATION INTRODUCTION TO CONCEPTS AND METHODS Olivier GARSMEUR & Stéphanie SIDIBE-BOCS Introduction two main concepts: Identify the different elements of the genome, (location and stucture) :
GENOME ANNOTATION INTRODUCTION TO CONCEPTS AND METHODS Olivier GARSMEUR & Stéphanie SIDIBE-BOCS Introduction two main concepts: Identify the different elements of the genome, (location and stucture) :
Gene Finding Genome Annotation
 Gene Finding Genome Annotation Gene finding is a cornerstone of genomic analysis Genome content and organization Differential expression analysis Epigenomics Population biology & evolution Medical genomics
Gene Finding Genome Annotation Gene finding is a cornerstone of genomic analysis Genome content and organization Differential expression analysis Epigenomics Population biology & evolution Medical genomics
UCSC Genome Browser. Introduction to ab initio and evidence-based gene finding
 UCSC Genome Browser Introduction to ab initio and evidence-based gene finding Wilson Leung 06/2006 Outline Introduction to annotation ab initio gene finding Basics of the UCSC Browser Evidence-based gene
UCSC Genome Browser Introduction to ab initio and evidence-based gene finding Wilson Leung 06/2006 Outline Introduction to annotation ab initio gene finding Basics of the UCSC Browser Evidence-based gene
Outline. Introduction to ab initio and evidence-based gene finding. Prokaryotic gene predictions
 Outline Introduction to ab initio and evidence-based gene finding Overview of computational gene predictions Different types of eukaryotic gene predictors Common types of gene prediction errors Wilson
Outline Introduction to ab initio and evidence-based gene finding Overview of computational gene predictions Different types of eukaryotic gene predictors Common types of gene prediction errors Wilson
ab initio and Evidence-Based Gene Finding
 ab initio and Evidence-Based Gene Finding A basic introduction to annotation Outline What is annotation? ab initio gene finding Genome databases on the web Basics of the UCSC browser Evidence-based gene
ab initio and Evidence-Based Gene Finding A basic introduction to annotation Outline What is annotation? ab initio gene finding Genome databases on the web Basics of the UCSC browser Evidence-based gene
Wheat Genome Structural Annotation Using a Modular and Evidence-combined Annotation Pipeline
 Wheat Genome Structural Annotation Using a Modular and Evidence-combined Annotation Pipeline Xi Wang Bioinformatics Scientist Computational Life Science Page 1 Bayer 4:3 Template 2010 March 2016 17/01/2017
Wheat Genome Structural Annotation Using a Modular and Evidence-combined Annotation Pipeline Xi Wang Bioinformatics Scientist Computational Life Science Page 1 Bayer 4:3 Template 2010 March 2016 17/01/2017
Collect, analyze and synthesize. Annotation. Annotation for D. virilis. Evidence Based Annotation. GEP goals: Evidence for Gene Models 08/22/2017
 Annotation Annotation for D. virilis Chris Shaffer July 2012 l Big Picture of annotation and then one practical example l This technique may not be the best with other projects (e.g. corn, bacteria) l
Annotation Annotation for D. virilis Chris Shaffer July 2012 l Big Picture of annotation and then one practical example l This technique may not be the best with other projects (e.g. corn, bacteria) l
Collect, analyze and synthesize. Annotation. Annotation for D. virilis. GEP goals: Evidence Based Annotation. Evidence for Gene Models 12/26/2018
 Annotation Annotation for D. virilis Chris Shaffer July 2012 l Big Picture of annotation and then one practical example l This technique may not be the best with other projects (e.g. corn, bacteria) l
Annotation Annotation for D. virilis Chris Shaffer July 2012 l Big Picture of annotation and then one practical example l This technique may not be the best with other projects (e.g. corn, bacteria) l
MAKER: An easy-to-use annotation pipeline designed for emerging model organism genomes
 Resource MAKER: An easy-to-use annotation pipeline designed for emerging model organism genomes Brandi L. Cantarel, 1 Ian Korf, 2 Sofia M.C. Robb, 3 Genis Parra, 2 Eric Ross, 4 Barry Moore, 1 Carson Holt,
Resource MAKER: An easy-to-use annotation pipeline designed for emerging model organism genomes Brandi L. Cantarel, 1 Ian Korf, 2 Sofia M.C. Robb, 3 Genis Parra, 2 Eric Ross, 4 Barry Moore, 1 Carson Holt,
Chimp Chunk 3-14 Annotation by Matthew Kwong, Ruth Howe, and Hao Yang
 Chimp Chunk 3-14 Annotation by Matthew Kwong, Ruth Howe, and Hao Yang Ruth Howe Bio 434W April 1, 2010 INTRODUCTION De novo annotation is the process by which a finished genomic sequence is searched for
Chimp Chunk 3-14 Annotation by Matthew Kwong, Ruth Howe, and Hao Yang Ruth Howe Bio 434W April 1, 2010 INTRODUCTION De novo annotation is the process by which a finished genomic sequence is searched for
HUMAN GENOME BIOINFORMATICS. Tore Samuelsson, Dec 2009
 HUMAN GENOME BIOINFORMATICS Tore Samuelsson, Dec 2009 The sequenced (gray filled) and unsequenced (white) portions of the human genome. Peter F.R. Little Genome Res. 2005; 15: 1759-1766 Human genome organisation
HUMAN GENOME BIOINFORMATICS Tore Samuelsson, Dec 2009 The sequenced (gray filled) and unsequenced (white) portions of the human genome. Peter F.R. Little Genome Res. 2005; 15: 1759-1766 Human genome organisation
Question 2: There are 5 retroelements (2 LINEs and 3 LTRs), 6 unclassified elements (XDMR and XDMR_DM), and 7 satellite sequences.
 Bio4342 Exercise 1 Answers: Detecting and Interpreting Genetic Homology (Answers prepared by Wilson Leung) Question 1: Low complexity DNA can be described as sequences that consist primarily of one or
Bio4342 Exercise 1 Answers: Detecting and Interpreting Genetic Homology (Answers prepared by Wilson Leung) Question 1: Low complexity DNA can be described as sequences that consist primarily of one or
Gene Prediction. Mario Stanke. Institut für Mikrobiologie und Genetik Abteilung Bioinformatik. Gene Prediction p.
 Gene Prediction Mario Stanke mstanke@gwdg.de Institut für Mikrobiologie und Genetik Abteilung Bioinformatik Gene Prediction p.1/23 Why Predict Genes with a Computer? tons of data 39/250 eukaryotic/prokaryotic
Gene Prediction Mario Stanke mstanke@gwdg.de Institut für Mikrobiologie und Genetik Abteilung Bioinformatik Gene Prediction p.1/23 Why Predict Genes with a Computer? tons of data 39/250 eukaryotic/prokaryotic
The Ensembl Database. Dott.ssa Inga Prokopenko. Corso di Genomica
 The Ensembl Database Dott.ssa Inga Prokopenko Corso di Genomica 1 www.ensembl.org Lecture 7.1 2 What is Ensembl? Public annotation of mammalian and other genomes Open source software Relational database
The Ensembl Database Dott.ssa Inga Prokopenko Corso di Genomica 1 www.ensembl.org Lecture 7.1 2 What is Ensembl? Public annotation of mammalian and other genomes Open source software Relational database
Chimp Sequence Annotation: Region 2_3
 Chimp Sequence Annotation: Region 2_3 Jeff Howenstein March 30, 2007 BIO434W Genomics 1 Introduction We received region 2_3 of the ChimpChunk sequence, and the first step we performed was to run RepeatMasker
Chimp Sequence Annotation: Region 2_3 Jeff Howenstein March 30, 2007 BIO434W Genomics 1 Introduction We received region 2_3 of the ChimpChunk sequence, and the first step we performed was to run RepeatMasker
GENOME ANNOTATION INTRODUCTION TO CONCEPTS AND METHODS. Olivier GARSMEUR. Training course in Bioinformatics applied to Musa genome November 2013
 GENOME ANNOTATION INTRODUCTION TO CONCEPTS AND METHODS Olivier GARSMEUR Training course in Bioinformatics applied to Musa genome 18-22 November 2013 Introduction two main concepts: Identify the different
GENOME ANNOTATION INTRODUCTION TO CONCEPTS AND METHODS Olivier GARSMEUR Training course in Bioinformatics applied to Musa genome 18-22 November 2013 Introduction two main concepts: Identify the different
The genome of Fraxinus excelsior (European Ash)
 The genome of Fraxinus excelsior (European Ash) Elizabeth Sollars, Laura Kelly, Bernardo Clavijo, David Swarbreck, Jasmin Zohren, David Boshier, Jo Clark, Anika Joecker, Sarah Ayling, Mario Caccamo, Richard
The genome of Fraxinus excelsior (European Ash) Elizabeth Sollars, Laura Kelly, Bernardo Clavijo, David Swarbreck, Jasmin Zohren, David Boshier, Jo Clark, Anika Joecker, Sarah Ayling, Mario Caccamo, Richard
Gene Prediction in Eukaryotes
 Gene Prediction in Eukaryotes Jan-Jaap Wesselink Biomol Informatics, S.L. jjw@biomol-informatics.com June 2010/Madrid jjw@biomol-informatics.com (BI) Gene Prediction June 2010/Madrid 1 / 34 Outline 1 Gene
Gene Prediction in Eukaryotes Jan-Jaap Wesselink Biomol Informatics, S.L. jjw@biomol-informatics.com June 2010/Madrid jjw@biomol-informatics.com (BI) Gene Prediction June 2010/Madrid 1 / 34 Outline 1 Gene
Gene Annotation Project. Group 1. Tyler Tiede Yanzhu Ji Jenae Skelton
 Gene Annotation Project Group 1 Tyler Tiede Yanzhu Ji Jenae Skelton Outline Tools Overview of 150kb region Overview of annotation process Characterization of 5 putative gene regions Analysis of masked
Gene Annotation Project Group 1 Tyler Tiede Yanzhu Ji Jenae Skelton Outline Tools Overview of 150kb region Overview of annotation process Characterization of 5 putative gene regions Analysis of masked
EvidentialGene: Perfect Genes Constructed from Gigabases of RNA Gilbert, Don Indiana University, Biology, Bloomington, IN 47405;
 EvidentialGene: Perfect Genes Constructed from Gigabases of RNA Gilbert, Don Indiana University, Biology, Bloomington, IN 47405; gilbertd@indiana.edu Gene construction, not prediction. The decade of gene
EvidentialGene: Perfect Genes Constructed from Gigabases of RNA Gilbert, Don Indiana University, Biology, Bloomington, IN 47405; gilbertd@indiana.edu Gene construction, not prediction. The decade of gene
Agenda. Web Databases for Drosophila. Gene annotation workflow. GEP Drosophila annotation projects 01/01/2018. Annotation adding labels to a sequence
 Agenda GEP annotation project overview Web Databases for Drosophila An introduction to web tools, databases and NCBI BLAST Web databases for Drosophila annotation UCSC Genome Browser NCBI / BLAST FlyBase
Agenda GEP annotation project overview Web Databases for Drosophila An introduction to web tools, databases and NCBI BLAST Web databases for Drosophila annotation UCSC Genome Browser NCBI / BLAST FlyBase
Genomic region (ENCODE) Gene definitions
 DNA From genes to proteins Bioinformatics Methods RNA PROMOTER ELEMENTS TRANSCRIPTION Iosif Vaisman mrna SPLICE SITES SPLICING Email: ivaisman@gmu.edu START CODON STOP CODON TRANSLATION PROTEIN From genes
DNA From genes to proteins Bioinformatics Methods RNA PROMOTER ELEMENTS TRANSCRIPTION Iosif Vaisman mrna SPLICE SITES SPLICING Email: ivaisman@gmu.edu START CODON STOP CODON TRANSLATION PROTEIN From genes
Agenda. Annotation of Drosophila. Muller element nomenclature. Annotation: Adding labels to a sequence. GEP Drosophila annotation projects 01/03/2018
 Agenda Annotation of Drosophila January 2018 Overview of the GEP annotation project GEP annotation strategy Types of evidence Analysis tools Web databases Annotation of a single isoform (walkthrough) Wilson
Agenda Annotation of Drosophila January 2018 Overview of the GEP annotation project GEP annotation strategy Types of evidence Analysis tools Web databases Annotation of a single isoform (walkthrough) Wilson
Selective constraints on noncoding DNA of mammals. Peter Keightley Institute of Evolutionary Biology University of Edinburgh
 Selective constraints on noncoding DNA of mammals Peter Keightley Institute of Evolutionary Biology University of Edinburgh Most mammalian noncoding DNA evolves rapidly Homo-Pan Divergence (%) 1.5 1.25
Selective constraints on noncoding DNA of mammals Peter Keightley Institute of Evolutionary Biology University of Edinburgh Most mammalian noncoding DNA evolves rapidly Homo-Pan Divergence (%) 1.5 1.25
Bacterial Genome Annotation
 Bacterial Genome Annotation Bacterial Genome Annotation For an annotation you want to predict from the sequence, all of... protein-coding genes their stop-start the resulting protein the function the control
Bacterial Genome Annotation Bacterial Genome Annotation For an annotation you want to predict from the sequence, all of... protein-coding genes their stop-start the resulting protein the function the control
Biol 478/595 Intro to Bioinformatics
 Biol 478/595 Intro to Bioinformatics September M 1 Labor Day 4 W 3 MG Database Searching Ch. 6 5 F 5 MG Database Searching Hw1 6 M 8 MG Scoring Matrices Ch 3 and Ch 4 7 W 10 MG Pairwise Alignment 8 F 12
Biol 478/595 Intro to Bioinformatics September M 1 Labor Day 4 W 3 MG Database Searching Ch. 6 5 F 5 MG Database Searching Hw1 6 M 8 MG Scoring Matrices Ch 3 and Ch 4 7 W 10 MG Pairwise Alignment 8 F 12
GenBank Growth. In 2003 ~ 31 million sequences ~ 37 billion base pairs
 Gene Finding GenBank Growth GenBank Growth In 2003 ~ 31 million sequences ~ 37 billion base pairs GenBank: Exponential Growth Growth of GenBank in billions of base pairs from release 3 in April of 1994
Gene Finding GenBank Growth GenBank Growth In 2003 ~ 31 million sequences ~ 37 billion base pairs GenBank: Exponential Growth Growth of GenBank in billions of base pairs from release 3 in April of 1994
A transposable elements annotation pipeline and expression analysis reveal potentially active elements in the microalga Tisochrysis lutea
 Additional file 1: A transposable elements annotation pipeline and expression analysis reveal potentially active elements in the microalga Tisochrysis lutea Jérémy Berthelier 1 *, Nathalie Casse 2, Nicolas
Additional file 1: A transposable elements annotation pipeline and expression analysis reveal potentially active elements in the microalga Tisochrysis lutea Jérémy Berthelier 1 *, Nathalie Casse 2, Nicolas
user s guide Question 1
 Question 1 How does one find a gene of interest and determine that gene s structure? Once the gene has been located on the map, how does one easily examine other genes in that same region? doi:10.1038/ng966
Question 1 How does one find a gene of interest and determine that gene s structure? Once the gene has been located on the map, how does one easily examine other genes in that same region? doi:10.1038/ng966
CSE 549: RNA-Seq aided gene finding
 CSE 549: RNA-Seq aided gene finding Finding Genes We ll break gene finding methods into 3 main categories. ab initio latin from the beginning w/o experimental evidence comparative make use of knowledge
CSE 549: RNA-Seq aided gene finding Finding Genes We ll break gene finding methods into 3 main categories. ab initio latin from the beginning w/o experimental evidence comparative make use of knowledge
Identifying Genes and Pseudogenes in a Chimpanzee Sequence Adapted from Chimp BAC analysis: TWINSCAN and UCSC Browser by Dr. M.
 Identifying Genes and Pseudogenes in a Chimpanzee Sequence Adapted from Chimp BAC analysis: TWINSCAN and UCSC Browser by Dr. M. Brent Prerequisites: A Simple Introduction to NCBI BLAST Resources: The GENSCAN
Identifying Genes and Pseudogenes in a Chimpanzee Sequence Adapted from Chimp BAC analysis: TWINSCAN and UCSC Browser by Dr. M. Brent Prerequisites: A Simple Introduction to NCBI BLAST Resources: The GENSCAN
Genome Annotation. What Does Annotation Describe??? Genome duplications Genes Mobile genetic elements Small repeats Genetic diversity
 Genome Annotation Genome Sequencing Costliest aspect of sequencing the genome o But Devoid of content Genome must be annotated o Annotation definition Analyzing the raw sequence of a genome and describing
Genome Annotation Genome Sequencing Costliest aspect of sequencing the genome o But Devoid of content Genome must be annotated o Annotation definition Analyzing the raw sequence of a genome and describing
GENOME ANNOTATION INTRODUCTION TO CONCEPTS AND METHODS
 GENOME ANNOTATION INTRODUCTION TO CONCEPTS AND METHODS garsmeur@2016 Introduc=on two main concepts: Iden=fy the different elements of the genome, (loca=on and stucture) : structural annota=on AIribute
GENOME ANNOTATION INTRODUCTION TO CONCEPTS AND METHODS garsmeur@2016 Introduc=on two main concepts: Iden=fy the different elements of the genome, (loca=on and stucture) : structural annota=on AIribute
BIO4342 Lab Exercise: Detecting and Interpreting Genetic Homology
 BIO4342 Lab Exercise: Detecting and Interpreting Genetic Homology Jeremy Buhler March 15, 2004 In this lab, we ll annotate an interesting piece of the D. melanogaster genome. Along the way, you ll get
BIO4342 Lab Exercise: Detecting and Interpreting Genetic Homology Jeremy Buhler March 15, 2004 In this lab, we ll annotate an interesting piece of the D. melanogaster genome. Along the way, you ll get
From assembled genome to annotated genome
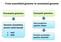 From assembled genome to annotated genome Procaryotic genomes Eucaryotic genomes Genome annotation servers (web based) 1. RAST 2. NCBI Gene prediction pipeline: Maker Function annotation pipeline: Blast2GO
From assembled genome to annotated genome Procaryotic genomes Eucaryotic genomes Genome annotation servers (web based) 1. RAST 2. NCBI Gene prediction pipeline: Maker Function annotation pipeline: Blast2GO
What the Genome of Raffaelea lauricola Can Tell Us About Laurel Wilt
 What the Genome of Raffaelea lauricola Can Tell Us About Laurel Wilt Laurel Wilt Summit November 3-4, 2016 Dr. Jeffrey Rollins Associate Professor Plant Pathology Department University of Florida Gainesville,
What the Genome of Raffaelea lauricola Can Tell Us About Laurel Wilt Laurel Wilt Summit November 3-4, 2016 Dr. Jeffrey Rollins Associate Professor Plant Pathology Department University of Florida Gainesville,
Proteogenomics Workflow for Neoantigen Discovery
 Proteogenomics Workflow for Neoantigen Discovery XIE Lu, Shanghai Center for Bioinformation Technology August 29-31, 2018. The 16th KJC Bioinformatics Symposium, Hayama, Japan Proteogenomics 目录 CONTENTS
Proteogenomics Workflow for Neoantigen Discovery XIE Lu, Shanghai Center for Bioinformation Technology August 29-31, 2018. The 16th KJC Bioinformatics Symposium, Hayama, Japan Proteogenomics 目录 CONTENTS
Genome annotation & EST
 Genome annotation & EST What is genome annotation? The process of taking the raw DNA sequence produced by the genome sequence projects and adding the layers of analysis and interpretation necessary
Genome annotation & EST What is genome annotation? The process of taking the raw DNA sequence produced by the genome sequence projects and adding the layers of analysis and interpretation necessary
Methods and Algorithms for Gene Prediction
 Methods and Algorithms for Gene Prediction Chaochun Wei 韦朝春 Sc.D. ccwei@sjtu.edu.cn http://cbb.sjtu.edu.cn/~ccwei Shanghai Jiao Tong University Shanghai Center for Bioinformation Technology 5/12/2011 K-J-C
Methods and Algorithms for Gene Prediction Chaochun Wei 韦朝春 Sc.D. ccwei@sjtu.edu.cn http://cbb.sjtu.edu.cn/~ccwei Shanghai Jiao Tong University Shanghai Center for Bioinformation Technology 5/12/2011 K-J-C
Outline. Gene Finding Questions. Recap: Prokaryotic gene finding Eukaryotic gene finding The human gene complement Regulation
 Tues, Nov 29: Gene Finding 1 Online FCE s: Thru Dec 12 Thurs, Dec 1: Gene Finding 2 Tues, Dec 6: PS5 due Project presentations 1 (see course web site for schedule) Thurs, Dec 8 Final papers due Project
Tues, Nov 29: Gene Finding 1 Online FCE s: Thru Dec 12 Thurs, Dec 1: Gene Finding 2 Tues, Dec 6: PS5 due Project presentations 1 (see course web site for schedule) Thurs, Dec 8 Final papers due Project
Sequence Based Function Annotation
 Sequence Based Function Annotation Qi Sun Bioinformatics Facility Biotechnology Resource Center Cornell University Sequence Based Function Annotation 1. Given a sequence, how to predict its biological
Sequence Based Function Annotation Qi Sun Bioinformatics Facility Biotechnology Resource Center Cornell University Sequence Based Function Annotation 1. Given a sequence, how to predict its biological
Annotation of contig27 in the Muller F Element of D. elegans. Contig27 is a 60,000 bp region located in the Muller F element of the D. elegans.
 David Wang Bio 434W 4/27/15 Annotation of contig27 in the Muller F Element of D. elegans Abstract Contig27 is a 60,000 bp region located in the Muller F element of the D. elegans. Genscan predicted six
David Wang Bio 434W 4/27/15 Annotation of contig27 in the Muller F Element of D. elegans Abstract Contig27 is a 60,000 bp region located in the Muller F element of the D. elegans. Genscan predicted six
Gene Prediction 10/21/05
 Gene Prediction 1/21/5 1/21/5 Gene Prediction Announcements Eam 2 - net Friday Posted online: Eam 2 Study Guide 544 Reading Assignment (2 papers) (formerly Gene Prediction - ) 1/21/5 D Dobbs ISU - BCB
Gene Prediction 1/21/5 1/21/5 Gene Prediction Announcements Eam 2 - net Friday Posted online: Eam 2 Study Guide 544 Reading Assignment (2 papers) (formerly Gene Prediction - ) 1/21/5 D Dobbs ISU - BCB
9/19/13. cdna libraries, EST clusters, gene prediction and functional annotation. Biosciences 741: Genomics Fall, 2013 Week 3
 cdna libraries, EST clusters, gene prediction and functional annotation Biosciences 741: Genomics Fall, 2013 Week 3 1 2 3 4 5 6 Figure 2.14 Relationship between gene structure, cdna, and EST sequences
cdna libraries, EST clusters, gene prediction and functional annotation Biosciences 741: Genomics Fall, 2013 Week 3 1 2 3 4 5 6 Figure 2.14 Relationship between gene structure, cdna, and EST sequences
Transcriptome Assembly, Functional Annotation (and a few other related thoughts)
 Transcriptome Assembly, Functional Annotation (and a few other related thoughts) Monica Britton, Ph.D. Sr. Bioinformatics Analyst June 23, 2017 Differential Gene Expression Generalized Workflow File Types
Transcriptome Assembly, Functional Annotation (and a few other related thoughts) Monica Britton, Ph.D. Sr. Bioinformatics Analyst June 23, 2017 Differential Gene Expression Generalized Workflow File Types
Genomes summary. Bacterial genome sizes
 Genomes summary 1. >930 bacterial genomes sequenced. 2. Circular. Genes densely packed. 3. 2-10 Mbases, 470-7,000 genes 4. Genomes of >200 eukaryotes (45 higher ) sequenced. 5. Linear chromosomes 6. On
Genomes summary 1. >930 bacterial genomes sequenced. 2. Circular. Genes densely packed. 3. 2-10 Mbases, 470-7,000 genes 4. Genomes of >200 eukaryotes (45 higher ) sequenced. 5. Linear chromosomes 6. On
Perfect~ Arthropod Genes Constructed from Gigabases of RNA
 Perfect~ Arthropod Genes Constructed from Gigabases of RNA Don Gilbert Biology Dept., Indiana University gilbertd@indiana.edu May/June 2012 Perfect Arthropod Genes Gen 2 genome informatics Gene prediction
Perfect~ Arthropod Genes Constructed from Gigabases of RNA Don Gilbert Biology Dept., Indiana University gilbertd@indiana.edu May/June 2012 Perfect Arthropod Genes Gen 2 genome informatics Gene prediction
Chapter 4 METHOD FOR PREDICTING GENES IN EUKARYOTIC GENOMES
 qene Prediction Chapter 4 METHOD FOR PREDICTING GENES IN EUKARYOTIC GENOMES 4.1 INTRODUCTION Evaluation of seven ab initio methods that were evaluated on a non-homologous mammalian data set (Rogic et al.
qene Prediction Chapter 4 METHOD FOR PREDICTING GENES IN EUKARYOTIC GENOMES 4.1 INTRODUCTION Evaluation of seven ab initio methods that were evaluated on a non-homologous mammalian data set (Rogic et al.
Training materials.
 Training materials Ensembl training materials are protected by a CC BY license http://creativecommons.org/licenses/by/4.0/ If you wish to re-use these materials, please credit Ensembl for their creation
Training materials Ensembl training materials are protected by a CC BY license http://creativecommons.org/licenses/by/4.0/ If you wish to re-use these materials, please credit Ensembl for their creation
Sequence Analysis. II: Sequence Patterns and Matrices. George Bell, Ph.D. WIBR Bioinformatics and Research Computing
 Sequence Analysis II: Sequence Patterns and Matrices George Bell, Ph.D. WIBR Bioinformatics and Research Computing Sequence Patterns and Matrices Multiple sequence alignments Sequence patterns Sequence
Sequence Analysis II: Sequence Patterns and Matrices George Bell, Ph.D. WIBR Bioinformatics and Research Computing Sequence Patterns and Matrices Multiple sequence alignments Sequence patterns Sequence
Small Exon Finder User Guide
 Small Exon Finder User Guide Author Wilson Leung wleung@wustl.edu Document History Initial Draft 01/09/2011 First Revision 08/03/2014 Current Version 12/29/2015 Table of Contents Author... 1 Document History...
Small Exon Finder User Guide Author Wilson Leung wleung@wustl.edu Document History Initial Draft 01/09/2011 First Revision 08/03/2014 Current Version 12/29/2015 Table of Contents Author... 1 Document History...
Improved Splice Site Detection in Genie
 Improved Splice Site Detection in Genie Martin Reese Informatics Group Human Genome Center Lawrence Berkeley National Laboratory MGReese@lbl.gov http://www-hgc.lbl.gov/inf Santa Fe, 1/23/97 Database Homologies
Improved Splice Site Detection in Genie Martin Reese Informatics Group Human Genome Center Lawrence Berkeley National Laboratory MGReese@lbl.gov http://www-hgc.lbl.gov/inf Santa Fe, 1/23/97 Database Homologies
Bioinformatika a výpočetní biologie. KFC/BIN VI. Geny predikce a ontologie
 Bioinformatika a výpočetní biologie KFC/BIN VI. Geny predikce a ontologie RNDr. Karel Berka, Ph.D. Univerzita Palackého v Olomouci Predikce genů gene is "a locatable region of genomic sequence, corresponding
Bioinformatika a výpočetní biologie KFC/BIN VI. Geny predikce a ontologie RNDr. Karel Berka, Ph.D. Univerzita Palackého v Olomouci Predikce genů gene is "a locatable region of genomic sequence, corresponding
Sequencing the genomes of Nicotiana sylvestris and Nicotiana tomentosiformis Nicolas Sierro
 Sequencing the genomes of Nicotiana sylvestris and Nicotiana tomentosiformis Nicolas Sierro Philip Morris International R&D, Philip Morris Products S.A., Neuchatel, Switzerland Introduction Nicotiana sylvestris
Sequencing the genomes of Nicotiana sylvestris and Nicotiana tomentosiformis Nicolas Sierro Philip Morris International R&D, Philip Morris Products S.A., Neuchatel, Switzerland Introduction Nicotiana sylvestris
Bioinformatics for Proteomics. Ann Loraine
 Bioinformatics for Proteomics Ann Loraine aloraine@uab.edu What is bioinformatics? The science of collecting, processing, organizing, storing, analyzing, and mining biological information, especially data
Bioinformatics for Proteomics Ann Loraine aloraine@uab.edu What is bioinformatics? The science of collecting, processing, organizing, storing, analyzing, and mining biological information, especially data
Using the Genome Browser: A Practical Guide. Travis Saari
 Using the Genome Browser: A Practical Guide Travis Saari What is it for? Problem: Bioinformatics programs produce an overwhelming amount of data Difficult to understand anything from the raw data Data
Using the Genome Browser: A Practical Guide Travis Saari What is it for? Problem: Bioinformatics programs produce an overwhelming amount of data Difficult to understand anything from the raw data Data
Outline. Annotation of Drosophila Primer. Gene structure nomenclature. Muller element nomenclature. GEP Drosophila annotation projects 01/04/2018
 Outline Overview of the GEP annotation projects Annotation of Drosophila Primer January 2018 GEP annotation workflow Practice applying the GEP annotation strategy Wilson Leung and Chris Shaffer AAACAACAATCATAAATAGAGGAAGTTTTCGGAATATACGATAAGTGAAATATCGTTCT
Outline Overview of the GEP annotation projects Annotation of Drosophila Primer January 2018 GEP annotation workflow Practice applying the GEP annotation strategy Wilson Leung and Chris Shaffer AAACAACAATCATAAATAGAGGAAGTTTTCGGAATATACGATAAGTGAAATATCGTTCT
Annotation of Drosophila erecta Contig 14. Kimberly Chau Dr. Laura Hoopes. Pomona College 24 February 2009
 Annotation of Drosophila erecta Contig 14 Kimberly Chau Dr. Laura Hoopes Pomona College 24 February 2009 1 Table of Contents I. Overview A. Introduction..1 B. Final Gene Model.....1 II. Genes A. Initial
Annotation of Drosophila erecta Contig 14 Kimberly Chau Dr. Laura Hoopes Pomona College 24 February 2009 1 Table of Contents I. Overview A. Introduction..1 B. Final Gene Model.....1 II. Genes A. Initial
Ensembl workshop. Thomas Randall, PhD bioinformatics.unc.edu. handouts, papers, datasets
 Ensembl workshop Thomas Randall, PhD tarandal@email.unc.edu bioinformatics.unc.edu www.unc.edu/~tarandal/ensembl handouts, papers, datasets Ensembl is a joint project between EMBL - EBI and the Sanger
Ensembl workshop Thomas Randall, PhD tarandal@email.unc.edu bioinformatics.unc.edu www.unc.edu/~tarandal/ensembl handouts, papers, datasets Ensembl is a joint project between EMBL - EBI and the Sanger
Ensembl gene annotation project
 Ensembl gene annotation project Sus scrofa (Pig) Raw Computes Stage: Searching for sequence patterns, aligning proteins and cdnas to the genome. The annotation process of the Sscrofa10.2 assembly began
Ensembl gene annotation project Sus scrofa (Pig) Raw Computes Stage: Searching for sequence patterns, aligning proteins and cdnas to the genome. The annotation process of the Sscrofa10.2 assembly began
Annotation of contig62 from Drosophila elegans Dot Chromosome
 Abstract: Annotation of contig62 from Drosophila elegans Dot Chromosome 1 Maxwell Wang The goal of this project is to annotate the Drosophila elegans Dot chromosome contig62. Contig62 is a 32,259 bp contig
Abstract: Annotation of contig62 from Drosophila elegans Dot Chromosome 1 Maxwell Wang The goal of this project is to annotate the Drosophila elegans Dot chromosome contig62. Contig62 is a 32,259 bp contig
Annotating the D. virilis Fourth Chromosome: Fosmid 99M21
 Sonal Singhal 3 May 2006 Bio 4342W Annotating the D. virilis Fourth Chromosome: Fosmid 99M21 Abstract In this project, I annotated a chunk of the D. virilis fourth chromosome (fosmid 99M21) by considering
Sonal Singhal 3 May 2006 Bio 4342W Annotating the D. virilis Fourth Chromosome: Fosmid 99M21 Abstract In this project, I annotated a chunk of the D. virilis fourth chromosome (fosmid 99M21) by considering
Aaditya Khatri. Abstract
 Abstract In this project, Chimp-chunk 2-7 was annotated. Chimp-chunk 2-7 is an 80 kb region on chromosome 5 of the chimpanzee genome. Analysis with the Mapviewer function using the NCBI non-redundant database
Abstract In this project, Chimp-chunk 2-7 was annotated. Chimp-chunk 2-7 is an 80 kb region on chromosome 5 of the chimpanzee genome. Analysis with the Mapviewer function using the NCBI non-redundant database
BIO 4342 Lecture on Repeats
 BIO 4342 Lecture on Repeats Jeremy Buhler June 14, 2006 1 How RepeatMasker Works Running RepeatMasker is the most common first step in annotating genomic DNA sequences. What exactly does it do? Given a
BIO 4342 Lecture on Repeats Jeremy Buhler June 14, 2006 1 How RepeatMasker Works Running RepeatMasker is the most common first step in annotating genomic DNA sequences. What exactly does it do? Given a
Reading Lecture 8: Lecture 9: Lecture 8. DNA Libraries. Definition Types Construction
 Lecture 8 Reading Lecture 8: 96-110 Lecture 9: 111-120 DNA Libraries Definition Types Construction 142 DNA Libraries A DNA library is a collection of clones of genomic fragments or cdnas from a certain
Lecture 8 Reading Lecture 8: 96-110 Lecture 9: 111-120 DNA Libraries Definition Types Construction 142 DNA Libraries A DNA library is a collection of clones of genomic fragments or cdnas from a certain
Annotation Walkthrough Workshop BIO 173/273 Genomics and Bioinformatics Spring 2013 Developed by Justin R. DiAngelo at Hofstra University
 Annotation Walkthrough Workshop NAME: BIO 173/273 Genomics and Bioinformatics Spring 2013 Developed by Justin R. DiAngelo at Hofstra University A Simple Annotation Exercise Adapted from: Alexis Nagengast,
Annotation Walkthrough Workshop NAME: BIO 173/273 Genomics and Bioinformatics Spring 2013 Developed by Justin R. DiAngelo at Hofstra University A Simple Annotation Exercise Adapted from: Alexis Nagengast,
Annotating 7G24-63 Justin Richner May 4, Figure 1: Map of my sequence
 Annotating 7G24-63 Justin Richner May 4, 2005 Zfh2 exons Thd1 exons Pur-alpha exons 0 40 kb 8 = 1 kb = LINE, Penelope = DNA/Transib, Transib1 = DINE = Novel Repeat = LTR/PAO, Diver2 I = LTR/Gypsy, Invader
Annotating 7G24-63 Justin Richner May 4, 2005 Zfh2 exons Thd1 exons Pur-alpha exons 0 40 kb 8 = 1 kb = LINE, Penelope = DNA/Transib, Transib1 = DINE = Novel Repeat = LTR/PAO, Diver2 I = LTR/Gypsy, Invader
Applications of HMMs in Computational Biology. BMI/CS Colin Dewey
 Applications of HMMs in Computational Biology BMI/CS 576 www.biostat.wisc.edu/bmi576.html Colin Dewey cdewey@biostat.wisc.edu Fall 2008 The Gene Finding Task Given: an uncharacterized DNA sequence Do:
Applications of HMMs in Computational Biology BMI/CS 576 www.biostat.wisc.edu/bmi576.html Colin Dewey cdewey@biostat.wisc.edu Fall 2008 The Gene Finding Task Given: an uncharacterized DNA sequence Do:
CSE 527 Computational Biology Autumn Lectures ~14-15 Gene Prediction
 CSE 527 Computational Biology Autumn 2004 Lectures ~14-15 Gene Prediction Some References A great online bib http://www.nslij-genetics.org/gene/ A good intro survey JM Claverie (1997) "Computational methods
CSE 527 Computational Biology Autumn 2004 Lectures ~14-15 Gene Prediction Some References A great online bib http://www.nslij-genetics.org/gene/ A good intro survey JM Claverie (1997) "Computational methods
Sequence Alignments. Week 3
 Sequence Alignments Week 3 Independent Project Gene Due: 9/25 (Monday--must be submitted by email) Rough Draft Due: 11/13 (hard copy due at the beginning of class, and emailed to me) Final Version Due:
Sequence Alignments Week 3 Independent Project Gene Due: 9/25 (Monday--must be submitted by email) Rough Draft Due: 11/13 (hard copy due at the beginning of class, and emailed to me) Final Version Due:
Biotechnology Project Lab
 Only for teaching purposes - not for reproduction or sale Advanced Cell Biology & Biotechnology Biotechnology Project Lab Giovanna Gambarotta COMPETENCES THAT YOU WILL ACQUIRE - compare DNA sequences -
Only for teaching purposes - not for reproduction or sale Advanced Cell Biology & Biotechnology Biotechnology Project Lab Giovanna Gambarotta COMPETENCES THAT YOU WILL ACQUIRE - compare DNA sequences -
Gene Prediction Final Presentation
 Gene Prediction Final Presentation Final Proposed Pipeline Assembled Genome Protein - coding Gene Prediction Ab Initio Prodigal Glimmer GeneMarkS RNA Gene Prediction ncrna Specific trnascanse (trna) RNAmmer
Gene Prediction Final Presentation Final Proposed Pipeline Assembled Genome Protein - coding Gene Prediction Ab Initio Prodigal Glimmer GeneMarkS RNA Gene Prediction ncrna Specific trnascanse (trna) RNAmmer
Introduction to Bioinformatics
 Introduction to Bioinformatics Bas E. Dutilh Systems Biology: Bioinformatic Data Analysis Utrecht University, February 8 th 2016 Info and documentation http://tbb.bio.uu.nl/bda/ http://www.google.com/
Introduction to Bioinformatics Bas E. Dutilh Systems Biology: Bioinformatic Data Analysis Utrecht University, February 8 th 2016 Info and documentation http://tbb.bio.uu.nl/bda/ http://www.google.com/
Annotating Fosmid 14p24 of D. Virilis chromosome 4
 Lo 1 Annotating Fosmid 14p24 of D. Virilis chromosome 4 Lo, Louis April 20, 2006 Annotation Report Introduction In the first half of Research Explorations in Genomics I finished a 38kb fragment of chromosome
Lo 1 Annotating Fosmid 14p24 of D. Virilis chromosome 4 Lo, Louis April 20, 2006 Annotation Report Introduction In the first half of Research Explorations in Genomics I finished a 38kb fragment of chromosome
AN OVERVIEW OF GENE STRUCTURE & FUNCTION PREDICTION. Marcus Chibucos, Ph.D. University of Maryland School of Medicine June 2013
 AN OVERVIEW OF GENE STRUCTURE & FUNCTION PREDICTION Marcus Chibucos, Ph.D. University of Maryland School of Medicine June 2013 Overview & goals Understand 1. How we predict presence & structure of coding
AN OVERVIEW OF GENE STRUCTURE & FUNCTION PREDICTION Marcus Chibucos, Ph.D. University of Maryland School of Medicine June 2013 Overview & goals Understand 1. How we predict presence & structure of coding
Computational Biology I LSM5191 (2003/4)
 Computational Biology I LSM5191 (2003/4) Aylwin Ng, D.Phil. Lecture Notes: Features of the Human Genome Reading List International Human Genome Sequencing Consortium (2001). Initial sequencing and analysis
Computational Biology I LSM5191 (2003/4) Aylwin Ng, D.Phil. Lecture Notes: Features of the Human Genome Reading List International Human Genome Sequencing Consortium (2001). Initial sequencing and analysis
Red: an intelligent, rapid, accurate tool for detecting repeats de-novo on the genomic scale
 Girgis BMC Bioinformatics (2015) 16:227 DOI 10.1186/s12859-015-0654-5 SOFTWARE Open Access Red: an intelligent, rapid, accurate tool for detecting repeats de-novo on the genomic scale Hani Z. Girgis 1,2
Girgis BMC Bioinformatics (2015) 16:227 DOI 10.1186/s12859-015-0654-5 SOFTWARE Open Access Red: an intelligent, rapid, accurate tool for detecting repeats de-novo on the genomic scale Hani Z. Girgis 1,2
RNA-Seq with the Tuxedo Suite
 RNA-Seq with the Tuxedo Suite Monica Britton, Ph.D. Sr. Bioinformatics Analyst September 2015 Workshop The Basic Tuxedo Suite References Trapnell C, et al. 2009 TopHat: discovering splice junctions with
RNA-Seq with the Tuxedo Suite Monica Britton, Ph.D. Sr. Bioinformatics Analyst September 2015 Workshop The Basic Tuxedo Suite References Trapnell C, et al. 2009 TopHat: discovering splice junctions with
The i5k a pan-arthropoda Genome Database. Chris Childers and Monica Poelchau USDA-ARS, National Agricultural Library
 The i5k Workspace@NAL: a pan-arthropoda Genome Database Chris Childers and Monica Poelchau USDA-ARS, National Agricultural Library Outline Background and overview Why join the i5k Workspace? What do we
The i5k Workspace@NAL: a pan-arthropoda Genome Database Chris Childers and Monica Poelchau USDA-ARS, National Agricultural Library Outline Background and overview Why join the i5k Workspace? What do we
Drosophila ficusphila F element
 5/2/2016 CONTIG52 Drosophila ficusphila F element Vahag Kechejian BIO434W Abstract Contig52 is a 35,000 bp region located on the F element of Drosophila ficusphila. Genscan predicts six features in the
5/2/2016 CONTIG52 Drosophila ficusphila F element Vahag Kechejian BIO434W Abstract Contig52 is a 35,000 bp region located on the F element of Drosophila ficusphila. Genscan predicts six features in the
Transcription and Post Transcript Modification
 Transcription and Post Transcript Modification You Should Be Able To 1. Describe transcription. 2. Compare and contrast eukaryotic + prokaryotic transcription. 3. Explain mrna processing in eukaryotes.
Transcription and Post Transcript Modification You Should Be Able To 1. Describe transcription. 2. Compare and contrast eukaryotic + prokaryotic transcription. 3. Explain mrna processing in eukaryotes.
Gene & genome organisation. Computational gene identification
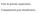 Gene & genome organisation Computational gene identification Eubacterial gene Eukaryotic gene Regulatory elements Promoter Translation start Introns polya signal Transcription stop DNA Transcription
Gene & genome organisation Computational gene identification Eubacterial gene Eukaryotic gene Regulatory elements Promoter Translation start Introns polya signal Transcription stop DNA Transcription
Aligning GENCODE and RefSeq transcripts By EMBL-EBI and NCBI
 Aligning GENCODE and RefSeq transcripts By EMBL-EBI and NCBI Joannella Morales, Ph.D. LRG Project Manager jmorales@ebi.ac.uk contact@lrg-sequence.org https://www.lrg-sequence.org https://www.ensembl.org
Aligning GENCODE and RefSeq transcripts By EMBL-EBI and NCBI Joannella Morales, Ph.D. LRG Project Manager jmorales@ebi.ac.uk contact@lrg-sequence.org https://www.lrg-sequence.org https://www.ensembl.org
Gene-centered resources at NCBI
 COURSE OF BIOINFORMATICS a.a. 2014-2015 Gene-centered resources at NCBI We searched Accession Number: M60495 AT NCBI Nucleotide Gene has been implemented at NCBI to organize information about genes, serving
COURSE OF BIOINFORMATICS a.a. 2014-2015 Gene-centered resources at NCBI We searched Accession Number: M60495 AT NCBI Nucleotide Gene has been implemented at NCBI to organize information about genes, serving
Gene Identification in silico
 Gene Identification in silico Nita Parekh, IIIT Hyderabad Presented at National Seminar on Bioinformatics and Functional Genomics, at Bioinformatics centre, Pondicherry University, Feb 15 17, 2006. Introduction
Gene Identification in silico Nita Parekh, IIIT Hyderabad Presented at National Seminar on Bioinformatics and Functional Genomics, at Bioinformatics centre, Pondicherry University, Feb 15 17, 2006. Introduction
Genscan. The Genscan HMM model Training Genscan Validating Genscan. (c) Devika Subramanian,
 Genscan The Genscan HMM model Training Genscan Validating Genscan (c) Devika Subramanian, 2009 96 Gene structure assumed by Genscan donor site acceptor site (c) Devika Subramanian, 2009 97 A simple model
Genscan The Genscan HMM model Training Genscan Validating Genscan (c) Devika Subramanian, 2009 96 Gene structure assumed by Genscan donor site acceptor site (c) Devika Subramanian, 2009 97 A simple model
MODULE 5: TRANSLATION
 MODULE 5: TRANSLATION Lesson Plan: CARINA ENDRES HOWELL, LEOCADIA PALIULIS Title Translation Objectives Determine the codons for specific amino acids and identify reading frames by looking at the Base
MODULE 5: TRANSLATION Lesson Plan: CARINA ENDRES HOWELL, LEOCADIA PALIULIS Title Translation Objectives Determine the codons for specific amino acids and identify reading frames by looking at the Base
Accurate & Complete Gene Construction with EvidentialGene. eugenes.org/evidentialgene/ 2016 June
 Accurate & Complete Gene Construction with EvidentialGene eugenes.org/evidentialgene/ Don Gilbert 2016 June gilbertd@indiana.edu What is EvidentialGene? Classifier of gene models Class = good, alternate,
Accurate & Complete Gene Construction with EvidentialGene eugenes.org/evidentialgene/ Don Gilbert 2016 June gilbertd@indiana.edu What is EvidentialGene? Classifier of gene models Class = good, alternate,
AC Algorithms for Mining Biological Sequences (COMP 680)
 AC-04-18 Algorithms for Mining Biological Sequences (COMP 680) Instructor: Mathieu Blanchette School of Computer Science and McGill Centre for Bioinformatics, 332 Duff Building McGill University, Montreal,
AC-04-18 Algorithms for Mining Biological Sequences (COMP 680) Instructor: Mathieu Blanchette School of Computer Science and McGill Centre for Bioinformatics, 332 Duff Building McGill University, Montreal,
Eukaryotic Gene Prediction. Wei Zhu May 2007
 Eukaryotic Gene Prediction Wei Zhu May 2007 In nature, nothing is perfect... - Alice Walker Gene Structure What is Gene Prediction? Gene prediction is the problem of parsing a sequence into nonoverlapping
Eukaryotic Gene Prediction Wei Zhu May 2007 In nature, nothing is perfect... - Alice Walker Gene Structure What is Gene Prediction? Gene prediction is the problem of parsing a sequence into nonoverlapping
Introduction to BIOINFORMATICS
 COURSE OF BIOINFORMATICS a.a. 2016-2017 Introduction to BIOINFORMATICS What is Bioinformatics? (I) The sinergy between biology and informatics What is Bioinformatics? (II) From: http://www.bioteach.ubc.ca/bioinfo2010/
COURSE OF BIOINFORMATICS a.a. 2016-2017 Introduction to BIOINFORMATICS What is Bioinformatics? (I) The sinergy between biology and informatics What is Bioinformatics? (II) From: http://www.bioteach.ubc.ca/bioinfo2010/
Analysis of data from high-throughput molecular biology experiments Lecture 6 (F6, RNA-seq ),
 Analysis of data from high-throughput molecular biology experiments Lecture 6 (F6, RNA-seq ), 2012-01-26 What is a gene What is a transcriptome History of gene expression assessment RNA-seq RNA-seq analysis
Analysis of data from high-throughput molecular biology experiments Lecture 6 (F6, RNA-seq ), 2012-01-26 What is a gene What is a transcriptome History of gene expression assessment RNA-seq RNA-seq analysis
Lab Week 9 - A Sample Annotation Problem (adapted by Chris Shaffer from a worksheet by Varun Sundaram, WU-STL, Class of 2009)
 Lab Week 9 - A Sample Annotation Problem (adapted by Chris Shaffer from a worksheet by Varun Sundaram, WU-STL, Class of 2009) Prerequisites: BLAST Exercise: An In-Depth Introduction to NCBI BLAST Familiarity
Lab Week 9 - A Sample Annotation Problem (adapted by Chris Shaffer from a worksheet by Varun Sundaram, WU-STL, Class of 2009) Prerequisites: BLAST Exercise: An In-Depth Introduction to NCBI BLAST Familiarity
Supplementary Table 1. Summary of whole genome shotgun sequence used for genome assembly
 Supplementary Tables Supplementary Table 1. Summary of whole genome shotgun sequence used for genome assembly Library Read length Raw data Filtered data insert size (bp) * Total Sequence depth Total Sequence
Supplementary Tables Supplementary Table 1. Summary of whole genome shotgun sequence used for genome assembly Library Read length Raw data Filtered data insert size (bp) * Total Sequence depth Total Sequence
Leonardo Mariño-Ramírez, PhD NCBI / NLM / NIH. BIOL 7210 A Computational Genomics 2/18/2015
 Leonardo Mariño-Ramírez, PhD NCBI / NLM / NIH BIOL 7210 A Computational Genomics 2/18/2015 The $1,000 genome is here! http://www.illumina.com/systems/hiseq-x-sequencing-system.ilmn Bioinformatics bottleneck
Leonardo Mariño-Ramírez, PhD NCBI / NLM / NIH BIOL 7210 A Computational Genomics 2/18/2015 The $1,000 genome is here! http://www.illumina.com/systems/hiseq-x-sequencing-system.ilmn Bioinformatics bottleneck
Multiple choice questions (numbers in brackets indicate the number of correct answers)
 1 February 15, 2013 Multiple choice questions (numbers in brackets indicate the number of correct answers) 1. Which of the following statements are not true Transcriptomes consist of mrnas Proteomes consist
1 February 15, 2013 Multiple choice questions (numbers in brackets indicate the number of correct answers) 1. Which of the following statements are not true Transcriptomes consist of mrnas Proteomes consist
Guided tour to Ensembl
 Guided tour to Ensembl Introduction Introduction to the Ensembl project Walk-through of the browser Variations and Functional Genomics Comparative Genomics BioMart Ensembl Genome browser http://www.ensembl.org
Guided tour to Ensembl Introduction Introduction to the Ensembl project Walk-through of the browser Variations and Functional Genomics Comparative Genomics BioMart Ensembl Genome browser http://www.ensembl.org
Genome Assembly and Annotation of Isochrysis Galbana
 Genome Assembly and Annotation of Isochrysis Galbana By: Yi Wang Institution: California State University San Marcos Date: May 14, 2014 Abstract Isochrysis Galbana is a species of cocoolithophores, which
Genome Assembly and Annotation of Isochrysis Galbana By: Yi Wang Institution: California State University San Marcos Date: May 14, 2014 Abstract Isochrysis Galbana is a species of cocoolithophores, which
