Nature Methods: doi: /nmeth Supplementary Figure 1. Retention of RNA with LabelX.
|
|
|
- Eleanor Fleming
- 6 years ago
- Views:
Transcription
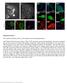
1 Supplementary Figure 1 Retention of RNA with LabelX. (a) Epi-fluorescence image of single molecule FISH (smfish) against GAPDH on HeLa cells expanded without LabelX treatment. (b) Epi-fluorescence image of smfish performed against GAPDH on expanded HeLa cells treated with LabelX. Images are maximum intensity projections of 3-D stacks. Nuclei stained with DAPI (shown in blue). Scale bars: 20 μm (post-expanded units).
2 Supplementary Figure 2 Effect of LabelX on fluorescent in-situ hybridization. To access the effect of LabelX on fluorescent in situ hybridization, fixed HeLa cells were stained with smfish probe-sets, followed by DNAse I treatment to remove the staining. The cells were then treated with LabelX and stained again with the same smfish probesets. (a) UBC staining before LabelX treatment and (b) UBC staining after probe removal and LabelX treatment. (c) EEF2 staining before LabelX treatment. (d) EEF2 staining after probe removal and LabelX treatment. (e) Comparison of smfish spots counted for individual cells before LabelX, and after probe removal and application of LabelX. The number of RNA molecules detected in a given cell was quantified using an automated spot counting algorithm (n=7 cells for each bar). Plotted are mean + standard error; no significant difference in spot counts before vs after LabelX (p > 0.5 for before vs. after for UBC, p > 0.5 for before vs. after for EEF2; t- test, unpaired, two-tailed). Images in a-d are maximum intensity projections of 3-D stacks; scale bars: 10 μm (pre-expanded units).
3 Supplementary Figure 3 High efficiency covalent anchoring of RNA to the ExM polymer gel. Different RNA species spanning 3 orders of magnitude in abundance were detected via single molecule RNA fluorescent in situ hybridization (FISH) in HeLa cells before and after ExM with LabelX treatment (shown in Fig. 1e). (a) Ratio of FISH spots detected after expansion to spots detected before expansion for single cells. Representative before vs. after ExFISH images shown: (b,c) TFRC; (d,e) GAPDH; (f,g) ACTB. Scale bars, 10 μm (pre-expanded units) in b, d, f; c, e, g, expanded physical size 21 μm (imaged in PBS).
4 Supplementary Figure 4 LabelX does not impede nuclear expansion. (a) Pre-expansion widefield image of a cultured HeLa cell stained with DAPI to visualize the nucleus (top panel) and smfish probes against ACTB (bottom panel). (b) Post-expansion widefield image of the same cell as in (a). (c) Pre-expansion widefield image of LabelX treated Thy1-YFP brain slice (Right panel, YFP protein) stained with DAPI (Left panel) (MIP, 4 μm z-depth). (d) Post-expansion image of the same region as in (c) (MIP, 12 μm). (e) Ratio of the expansion factor of cell bodies for individual cells to the expansion factor of their respective nuclei. smfish stain is used to outline the boundaries of the cell bodies of cultured cells while the endogenous YFP protein is used to demarcate the cell bodies of neurons in Thy1-YFP brain slices. Plotted are mean ± standard error. The ratio for both cultured cells and brain slices did not significantly deviate from one (p >0.05 for both, 1-sample t-test; n = 6, cultured HeLa cells; n = 7, cells in 1 brain slice). Scale bars, 10 μm.
5 Supplementary Figure 5 Isotropy of ExFISH. (a) Representative FISH image of TOP2A in a single HeLa cell before expansion (MIP of cell thickness). (b) ExFISH image of cell in (a) taken with the same optical parameters. (c) Merged image of (a) and (b) (red and green for before and after expansion respectively); distance measurements between pairs of mrna spots before (L, red line) and after (L, green line; note that these lines overlap nearly completely) expansion were used to quantify expansion isotropy. (d) Mean of the absolute value of the measurement error (i.e., L-L ) plotted against measurement length (L) for all pairs of mrna spots (mean ± standard deviation, N = 4 samples, 6.8 x 10 5 measurements). Scale bars: white, 10 µm pre-expansion units; blue, white scale bar divided by expansion factor. Orange line indicates diffraction limit of the microscope used (see Methods for details).
6 Supplementary Figure 6 Serially hybridized and multiplexed ExFISH. (a) Five consecutive widefield fluorescence images (top to bottom, then left to right) of GAPDH, applied to the cell of Fig. 2a. (b) Widefield fluorescence images showing ExFISH with serially delivered probes against six RNA targets (right to left, then top to bottom: NEAT1, EEF2, ACTB, UBC, GAPDH, and USF2) in a cultured HeLa cell (raw images of composite shown in Fig. 2e). Scale bars: 20 μm in expanded units.
7 Supplementary Figure 7 Schematic for HCR-mediated signal amplification. FISH probes bearing HCR initiators are hybridized to a target mrna. During amplification, metastable DNA hairpins bearing fluorophores assemble into polymer chains onto the initiators, thus amplifying signal downstream of the FISH probe hybridization event.
8 Supplementary Figure 8 HCR Amplification False Positives. (a) Widefield image of a LabelX treated Thy1-YFP brain slice (YFP protein, green) stained with probes against YFP (red) and Gad1 (magenta) followed by HCR amplification. Probes against YFP transcripts were amplified with the B1 amplifier set (see Methods) while probes against Gad1 transcripts were amplified with the B2 amplifier set (MIP, 59 μm). (b) Widefield image of LabelX treated Thy1-YFP brain slice (YFP protein, green) treated with the same HCR amplifiers as in (a) (namely B1 (red) and B2 (magenta)) without the addition of probes (MIP, 50 μm). (c) HCR spots detected per volume of expanded sample. Analysis was performed on samples which were either treated or not treated with FISH probes followed by HCR amplification. An automated spot counting algorithm (as used in Fig. 1) was used to count HCR spots. The endogenous YFP protein was used to delineate regions used for the analysis. Plotted are mean ± standard error. HCR spot counts are significantly different in the presence of probes than without probes (p <0.05 for both B1 and B2 amplifier sets, Welch s t-test; n=4 fields of view each). Scale bars: 50 μm.
9 Supplementary Figure 9 Lightsheet microscopy of ExFISH. (a) Volume rendering of Thy1-YFP (green) brain tissue acquired by lightsheet microscopy with HCR-ExFISH targeting YFP (red) and Gad1 (blue) mrna. (b) A maximum intensity projection (~8 µm in Z) of a small subsection of the volume, showing the high resolution of imaging and single molecule localization of imaging expanded specimens with lightsheet imaging (scale bar: 10 µm, in pre-expansion units, expansion factor, 3 ). (c) Zoom in of the volume rendering in (a) (scale bar: 20 µm, in pre-expansion units, 3 ).
10 Supplementary Figure 10 Two-color co-localization of FISH probes with HCR amplification in expanded Thy1-YFP brain slices. (a) Schematic showing two color amplification of the same target. A transcript of interest is targeted by probes against alternating parts of the sequence, and bearing two different HCR initiators, allowing for amplification in two colors. (b) Confocal image showing FISH staining with HCR amplification against the Camk2a transcript in two colors (red and blue; YFP fluorescence shown in green). (c) The result of an automated two-color spot co-localization analysis performed on the data set shown in (b). Each purple spot represents a positive co-localization identified by the algorithm and overlaid on the confocal image of YFP. Zoom in of dendrites showing two color FISH staining with HCR amplification against Camk2a (d,e) and Dlg4 (f,g) transcripts. Top row shows the raw two color staining data corresponding to the bottom row showing co-localized spots identified by the automated algorithm (replicated from Fig. 3j-k for convenience). Scale bars: (b,c) 10 μm (3 ); (d-g) 2 μm (3 ). (b-g) are MIP of ~1.6 μm thickness in unexpanded coordinates.
11 Supplementary Figure 11 HCR reversal via toe-hold mediated strand displacement. (a) Schematic for HCR amplification and reversal. HCR amplification is initiated with custom-made HCR hairpins bearing toe-holds for toe-hold mediated strand displacement. After amplification, the addition of a disassembling strand initiates the disassembly of the HCR polymers via strand displacement. (b) ExFISH-treated Thy1-YFP brain slice (YFP in blue) is shown stained with YFP FISH probes bearing HCR initiators and amplified with custom made HCR hairpins bearing toe-holds for strand displacement (green dots). The different panels show the state of HCR reversal at different times after the addition of strands to initiate the disassembly of the HCR polymers. Scale bars: 20 μm (in post-expansion units).
12 Supplementary Figure 12 Dependence of RNA FISH spot intensity on degree of expansion and concentration of LabelX. HeLa cells, treated with LabelX diluted to different final concentrations of Label-IT Amine concentration, were expanded and stained with a probe-set against GAPDH. After staining, the gelled samples were expanded in 1 PBS (~2 expansion ratio) and water (~4 expansion ratio) and the spot intensity for the different samples was quantified. Plotted are mean + standard error; N = 6 cells.
13 Supplemental Table 1. List of reagents and suppliers Chemical Supplies Chemical Name Supplier Part Number ExM Gel or Preparation Hybridization Buffer Sodium Acrylate (purity note:*) Sigma Acrylamide Sigma A9099 N,N -Methylenebisacrylamide Sigma M7279 Ammonium Persulfate Sigma A3678 N,N,N,N -Tetramethylethylenediamine Sigma T7024 VA-044 Wako Hydroxy-TEMPO Sigma Dextran Sulfate Sigma D g SSC Thermo Fisher AM9765 Formamide Thermo Fisher AM9342 Paraformaldehyde Electron Microscopy Sciences Fixation and Permeabilization Protein Digestion Tissue-prep Buffered 10% Formalin Electron Microscopy Sciences Triton X-100 Sigma Ethyl Alcohol Sigma E7023 Glycine Sigma x PBS Thermo Fisher AM9624 Proteinase K New England Biolabs P8107S Ethylenediaminetetraacetic acid Sigma EDS Sodium Chloride Sigma S9888 Tris-HCl Life Technologies AM9855 HCR Amplification Amplification Buffer Molecular Instruments N/A Tween 20 Sigma P1379 LabelX Preparation Label-IT Amine Mirus Bio MIR 3900 Acryloyl-X, SE Thermo Fisher A20770 LabelX Treatment MOPS Sigma M G Reembeded Gels Staining DNAse I Sigma Bind-silane Treatment Bind-Silane Sigma GE * check for yellow color upon resuspension: that indicates poor quality; solution should be clear (see
14 Decades (Transcript Abundance) Mean (Ratio of # spots detected in individual cells after ExM, to # spots detected before ExM) Standard Deviation Sample size (n) p-value 10s s s Supplementary Table 2. Statistical Analysis of RNA FISH spots detected in individual cells before and after ExFISH. For RNA molecules detected before vs after expansion, spots were counted by an automated algorithm. The ratio of the number of spots after ExM to spots counted before ExM was determined for each cell. Spot counts were grouped into decades based on the pre-expansion spot count. The table shows the results of a one-sample T-test performed on the ratio of spots counts for each decade to determine significant deviation from the expected mean ratio value of one.
15 Target Total Spot Count (Averaged Across Both Red and Blue Channels) Co-localized Spots Co-localization % Hybridization Efficiency Volume analyzed Density (Colocalized Puncta per (µm 3 in unexpanded coordinates) µm 3 ) ActB Dlg Camk2a Dlg4 Missense mcherry Supplementary Table 3. Analysis of two-color colocalization of FISH probes with HCR amplification in expanded slices.
16 Supplementary Video 1. Volume rendering of Thy1-YFP (green) brain tissue acquired by lightsheet microscopy with HCR-ExFISH targeting YFP (red) and Gad1 (blue) mrna. Movie of volume in Supp. Fig. 9.
Improvements on expansion microscopy: protein-retention expansion microscopy (proexm) and expansion FISH (exfish)
 Improvements on expansion microscopy: protein-retention expansion microscopy (proexm) and expansion FISH (exfish) Technical Journal Club 19.07.2016. Orsolya Török Microscopy-history μικρός, mikrós, "small"
Improvements on expansion microscopy: protein-retention expansion microscopy (proexm) and expansion FISH (exfish) Technical Journal Club 19.07.2016. Orsolya Török Microscopy-history μικρός, mikrós, "small"
Supplementary Figure 1
 Supplementary Figure 1 (A) Schematic of sequential hybridization and barcoding. (B) Schematic of the FISH images of the cell. In each round of hybridization, the same spots are detected, but the dye associated
Supplementary Figure 1 (A) Schematic of sequential hybridization and barcoding. (B) Schematic of the FISH images of the cell. In each round of hybridization, the same spots are detected, but the dye associated
Nanoscale Imaging of RNA with Expansion Microscopy
 Nanoscale Imaging of RNA with Expansion Microscopy The Harvard community has made this article openly available. Please share how this access benefits you. Your story matters Citation Chen, F., A. T. Wassie,
Nanoscale Imaging of RNA with Expansion Microscopy The Harvard community has made this article openly available. Please share how this access benefits you. Your story matters Citation Chen, F., A. T. Wassie,
Multiplexed imaging using same species primary antibodies with signal amplification
 Multiplexed imaging using same species primary antibodies with signal amplification Yu Wang 1,2, Wenxin Xie 1,2, Richie E. Kohman 1 and George M. Church 1,2 1. Wyss Institute for Biologically Inspired
Multiplexed imaging using same species primary antibodies with signal amplification Yu Wang 1,2, Wenxin Xie 1,2, Richie E. Kohman 1 and George M. Church 1,2 1. Wyss Institute for Biologically Inspired
Nature Biotechnology: doi: /nbt Supplementary Figure 1. Mass spectrometry characterization of epoxide reactions.
 Supplementary Figure 1 Mass spectrometry characterization of epoxide reactions. (a) Degree of amine reactivity of bovine serum albumin (BSA) with epoxide molecules having different numbers of epoxide groups:
Supplementary Figure 1 Mass spectrometry characterization of epoxide reactions. (a) Degree of amine reactivity of bovine serum albumin (BSA) with epoxide molecules having different numbers of epoxide groups:
Technical Review. Real time PCR
 Technical Review Real time PCR Normal PCR: Analyze with agarose gel Normal PCR vs Real time PCR Real-time PCR, also known as quantitative PCR (qpcr) or kinetic PCR Key feature: Used to amplify and simultaneously
Technical Review Real time PCR Normal PCR: Analyze with agarose gel Normal PCR vs Real time PCR Real-time PCR, also known as quantitative PCR (qpcr) or kinetic PCR Key feature: Used to amplify and simultaneously
Supplementary Information
 Supplementary Information Live imaging reveals the dynamics and regulation of mitochondrial nucleoids during the cell cycle in Fucci2-HeLa cells Taeko Sasaki 1, Yoshikatsu Sato 2, Tetsuya Higashiyama 1,2,
Supplementary Information Live imaging reveals the dynamics and regulation of mitochondrial nucleoids during the cell cycle in Fucci2-HeLa cells Taeko Sasaki 1, Yoshikatsu Sato 2, Tetsuya Higashiyama 1,2,
A Supersandwich Fluorescence in Situ Hybridization (SFISH) Strategy. for Highly Sensitive and Selective mrna Imaging in Tumor Cells
 Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2015 Electronic Supplementary Information (ESI) A Supersandwich Fluorescence in Situ Hybridization (SFISH)
Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2015 Electronic Supplementary Information (ESI) A Supersandwich Fluorescence in Situ Hybridization (SFISH)
Post-expansion antibody delivery, after epitope-preserving homogenization.
 Supplementary Figure 1 Post-expansion antibody delivery, after epitope-preserving homogenization. (a, b) Wide-field fluorescence images of Thy1-YFP-expressing mouse brain hemisphere slice before expansion
Supplementary Figure 1 Post-expansion antibody delivery, after epitope-preserving homogenization. (a, b) Wide-field fluorescence images of Thy1-YFP-expressing mouse brain hemisphere slice before expansion
LysoTracker Red DND-99 (Invitrogen) was used as a marker of lysosome or acidic
 information MATERIAL AND METHODS Lysosome staining LysoTracker Red DND-99 (Invitrogen) was used as a marker of lysosome or acidic compartments, according to the manufacturer s protocol. Plasmid independent
information MATERIAL AND METHODS Lysosome staining LysoTracker Red DND-99 (Invitrogen) was used as a marker of lysosome or acidic compartments, according to the manufacturer s protocol. Plasmid independent
SUPPLEMENTARY INFORMATION. Transcriptional output transiently spikes upon mitotic exit
 SUPPLEMENTARY INFORMATION Transcriptional output transiently spikes upon mitotic exit Viola Vaňková Hausnerová 1, 2, Christian Lanctôt 1* 1 BIOCEV and Department of Cell Biology, Faculty of Science, Charles
SUPPLEMENTARY INFORMATION Transcriptional output transiently spikes upon mitotic exit Viola Vaňková Hausnerová 1, 2, Christian Lanctôt 1* 1 BIOCEV and Department of Cell Biology, Faculty of Science, Charles
Stellaris FISH Probes Protocols and Storage
 Stellaris FISH Probes Protocols and Storage Catalog No. SMF-2035-1 Product Name Stellaris FISH Probes, Human MALAT1 with Quasar 570 Dye Product Description Product consists of Quasar 570-labeled oligos
Stellaris FISH Probes Protocols and Storage Catalog No. SMF-2035-1 Product Name Stellaris FISH Probes, Human MALAT1 with Quasar 570 Dye Product Description Product consists of Quasar 570-labeled oligos
TR DISTRIBUTION STATEMENT A: Approved for public release; distribution is unlimited.
 Chapter 14. Intracellular detection of viral transcription and replication using RNA FISH i. Summary/Abstract Many hemorrhagic fever viruses require BSL-3 or 4 laboratory containment for study. The necessary
Chapter 14. Intracellular detection of viral transcription and replication using RNA FISH i. Summary/Abstract Many hemorrhagic fever viruses require BSL-3 or 4 laboratory containment for study. The necessary
Gene Expression Technology
 Gene Expression Technology Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Gene expression Gene expression is the process by which information from a gene
Gene Expression Technology Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Gene expression Gene expression is the process by which information from a gene
INTRODUCTION TO REVERSE TRANSCRIPTION PCR (RT-PCR) ABCF 2016 BecA-ILRI Hub, Nairobi 21 st September 2016 Roger Pelle Principal Scientist
 INTRODUCTION TO REVERSE TRANSCRIPTION PCR (RT-PCR) ABCF 2016 BecA-ILRI Hub, Nairobi 21 st September 2016 Roger Pelle Principal Scientist Objective of PCR To provide a solution to one of the most pressing
INTRODUCTION TO REVERSE TRANSCRIPTION PCR (RT-PCR) ABCF 2016 BecA-ILRI Hub, Nairobi 21 st September 2016 Roger Pelle Principal Scientist Objective of PCR To provide a solution to one of the most pressing
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/NCHEM.1805 Visualization and Selective Chemical Targeting of RNA G-quadruplex Structures in the Cytoplasm of Human Cells Giulia Biffi 1, Marco Di Antonio 2, David Tannahill 1 and Shankar Balasubramanian
DOI: 10.1038/NCHEM.1805 Visualization and Selective Chemical Targeting of RNA G-quadruplex Structures in the Cytoplasm of Human Cells Giulia Biffi 1, Marco Di Antonio 2, David Tannahill 1 and Shankar Balasubramanian
Supplementary Figure 1: (a) 3D illustration of the mobile phone microscopy attachment, containing a 3D movable sample holder (red piece for z
 Supplementary Figure 1: (a) 3D illustration of the mobile phone microscopy attachment, containing a 3D movable sample holder (red piece for z movement and blue piece for x-y movement). Illumination sources
Supplementary Figure 1: (a) 3D illustration of the mobile phone microscopy attachment, containing a 3D movable sample holder (red piece for z movement and blue piece for x-y movement). Illumination sources
DNA Microarray Technology
 2 DNA Microarray Technology 2.1 Overview DNA microarrays are assays for quantifying the types and amounts of mrna transcripts present in a collection of cells. The number of mrna molecules derived from
2 DNA Microarray Technology 2.1 Overview DNA microarrays are assays for quantifying the types and amounts of mrna transcripts present in a collection of cells. The number of mrna molecules derived from
Multiplexed imaging of highdensity. MERFISH and expansion microscopy
 Multiplexed imaging of highdensity libraries of RNAs with MERFISH and expansion microscopy The Harvard community has made this article openly available. Please share how this access benefits you. Your
Multiplexed imaging of highdensity libraries of RNAs with MERFISH and expansion microscopy The Harvard community has made this article openly available. Please share how this access benefits you. Your
Confocal immunofluorescence microscopy
 Confocal immunofluorescence microscopy HL-6 and cells were cultured and cytospun onto glass slides. The cells were double immunofluorescence stained for Mt NPM1 and fibrillarin (nucleolar marker). Briefly,
Confocal immunofluorescence microscopy HL-6 and cells were cultured and cytospun onto glass slides. The cells were double immunofluorescence stained for Mt NPM1 and fibrillarin (nucleolar marker). Briefly,
Nature Methods: doi: /nmeth.4396
 Supplementary Figure 1 Comparison of technical replicate consistency between and across the standard ATAC-seq method, DNase-seq, and Omni-ATAC. (a) Heatmap-based representation of ATAC-seq quality control
Supplementary Figure 1 Comparison of technical replicate consistency between and across the standard ATAC-seq method, DNase-seq, and Omni-ATAC. (a) Heatmap-based representation of ATAC-seq quality control
Roche Molecular Biochemicals Technical Note No. LC 10/2000
 Roche Molecular Biochemicals Technical Note No. LC 10/2000 LightCycler Overview of LightCycler Quantification Methods 1. General Introduction Introduction Content Definitions This Technical Note will introduce
Roche Molecular Biochemicals Technical Note No. LC 10/2000 LightCycler Overview of LightCycler Quantification Methods 1. General Introduction Introduction Content Definitions This Technical Note will introduce
Flow Cytometry - The Essentials
 Flow Cytometry - The Essentials Pocket Guide to Flow Cytometry: 1. Know your Cytometer 2. Understanding Fluorescence and Fluorophores 3. Gating Process 4. Controls 5. Optimization 6. Panel Building 7.
Flow Cytometry - The Essentials Pocket Guide to Flow Cytometry: 1. Know your Cytometer 2. Understanding Fluorescence and Fluorophores 3. Gating Process 4. Controls 5. Optimization 6. Panel Building 7.
BIO 315 Lab Exam I. Section #: Name:
 Section #: Name: Also provide this information on the computer grid sheet given to you. (Section # in special code box) BIO 315 Lab Exam I 1. In labeling the parts of a standard compound light microscope
Section #: Name: Also provide this information on the computer grid sheet given to you. (Section # in special code box) BIO 315 Lab Exam I 1. In labeling the parts of a standard compound light microscope
Rapid Method for the Purification of Total RNA from Formalin- Fixed Paraffin-Embedded (FFPE) Tissue Samples
 Application Note 17 RNA Sample Preparation Rapid Method for the Purification of Total RNA from Formalin- Fixed Paraffin-Embedded (FFPE) Tissue Samples M. Melmogy 1, V. Misic 1, B. Lam, PhD 1, C. Dobbin,
Application Note 17 RNA Sample Preparation Rapid Method for the Purification of Total RNA from Formalin- Fixed Paraffin-Embedded (FFPE) Tissue Samples M. Melmogy 1, V. Misic 1, B. Lam, PhD 1, C. Dobbin,
Supporting Information
 Electronic Supplementary Material (ESI) for Chemical Science. This journal is The Royal Society of Chemistry 2018 Supporting Information An enzyme-free molecular catalytic device: dynamically self-assembled
Electronic Supplementary Material (ESI) for Chemical Science. This journal is The Royal Society of Chemistry 2018 Supporting Information An enzyme-free molecular catalytic device: dynamically self-assembled
Stellaris RNA FISH Protocol for Adherent Cells in 96 Well Glass Bottom Plates
 Stellaris RNA FISH Protocol for Adherent Cells in 96 Well Glass Bottom Plates This protocol is specifically designed for high throughput applications of Stellaris in 96 well glass bottom plates. General
Stellaris RNA FISH Protocol for Adherent Cells in 96 Well Glass Bottom Plates This protocol is specifically designed for high throughput applications of Stellaris in 96 well glass bottom plates. General
Stellaris RNA FISH Protocol for D. Melanogaster Wing Imaginal Discs
 Stellaris RNA FISH Protocol for D. Melanogaster Wing Imaginal Discs General Protocol & Storage Product Description A set of Stellaris RNA FISH Probes is comprised of up to 48 singly labeled oligonucleotides
Stellaris RNA FISH Protocol for D. Melanogaster Wing Imaginal Discs General Protocol & Storage Product Description A set of Stellaris RNA FISH Probes is comprised of up to 48 singly labeled oligonucleotides
Supplementary Material for
 www.sciencemag.org/cgi/content/full/science.aaa6090/dc1 Supplementary Material for Spatially resolved, highly multiplexed RNA profiling in single cells Kok Hao Chen, Alistair N. Boettiger, Jeffrey R. Moffitt,
www.sciencemag.org/cgi/content/full/science.aaa6090/dc1 Supplementary Material for Spatially resolved, highly multiplexed RNA profiling in single cells Kok Hao Chen, Alistair N. Boettiger, Jeffrey R. Moffitt,
BIO 315 Lab Exam I. Section #: Name:
 Section #: Name: Also provide this information on the computer grid sheet given to you. (Section # in special code box) BIO 315 Lab Exam I 1. In labeling the parts of a standard compound light microscope
Section #: Name: Also provide this information on the computer grid sheet given to you. (Section # in special code box) BIO 315 Lab Exam I 1. In labeling the parts of a standard compound light microscope
Vital Reagents, Essential Products for Life Science Research
 Vital Reagents, Essential Products for Life Science Research Bringing Chemistry to Life Find the Perfect Reagents for your Life Science Application Fisher BioReagents offer over 1,000 products dedicated
Vital Reagents, Essential Products for Life Science Research Bringing Chemistry to Life Find the Perfect Reagents for your Life Science Application Fisher BioReagents offer over 1,000 products dedicated
Assay Design Considerations, Optimization and Validation
 Assay Design Considerations, Optimization and Validation Ray Meng, Ph.D. International Field Applications Specialist Gene Expression Division Bio-Rad Laboratories, Inc. Assay Design Considerations Experiment
Assay Design Considerations, Optimization and Validation Ray Meng, Ph.D. International Field Applications Specialist Gene Expression Division Bio-Rad Laboratories, Inc. Assay Design Considerations Experiment
3.1.4 DNA Microarray Technology
 3.1.4 DNA Microarray Technology Scientists have discovered that one of the differences between healthy and cancer is which genes are turned on in each. Scientists can compare the gene expression patterns
3.1.4 DNA Microarray Technology Scientists have discovered that one of the differences between healthy and cancer is which genes are turned on in each. Scientists can compare the gene expression patterns
THE BASICS OF IMMUNOHISTOCHEMISTRY
 THE BASICS OF IMMUNOHISTOCHEMISTRY Introduction Immunohistochemistry (IHC) identifies specific tissue components by means of a specific antigen/antibody reaction tagged with a visible label. IHC makes
THE BASICS OF IMMUNOHISTOCHEMISTRY Introduction Immunohistochemistry (IHC) identifies specific tissue components by means of a specific antigen/antibody reaction tagged with a visible label. IHC makes
Introduction to Microarray Analysis
 Introduction to Microarray Analysis Methods Course: Gene Expression Data Analysis -Day One Rainer Spang Microarrays Highly parallel measurement devices for gene expression levels 1. How does the microarray
Introduction to Microarray Analysis Methods Course: Gene Expression Data Analysis -Day One Rainer Spang Microarrays Highly parallel measurement devices for gene expression levels 1. How does the microarray
Supporting Information. ribosome-mrna interactions in single cells. Kelly S. Burke, Katie A. Antilla, and David A. Tirrell*
 Supporting Information A fluorescence in situ hybridization method to quantify mrna translation by visualizing ribosome-mrna interactions in single cells. Kelly S. Burke, Katie A. Antilla, and David A.
Supporting Information A fluorescence in situ hybridization method to quantify mrna translation by visualizing ribosome-mrna interactions in single cells. Kelly S. Burke, Katie A. Antilla, and David A.
Stellaris RNA FISH Protocol for Simultaneous IF + FISH in Adherent Cells
 Stellaris RNA FISH Protocol for Simultaneous IF + FISH in Adherent Cells General Protocol & Storage Product Description A set of Stellaris RNA FISH Probes is comprised of up to 48 singly labeled oligonucleotides
Stellaris RNA FISH Protocol for Simultaneous IF + FISH in Adherent Cells General Protocol & Storage Product Description A set of Stellaris RNA FISH Probes is comprised of up to 48 singly labeled oligonucleotides
Supplementary Figure 1. NORAD expression in mouse (A) and dog (B). The black boxes indicate the position of the regions alignable to the 12 repeat
 Supplementary Figure 1. NORAD expression in mouse (A) and dog (B). The black boxes indicate the position of the regions alignable to the 12 repeat units in the human genome. Annotated transposable elements
Supplementary Figure 1. NORAD expression in mouse (A) and dog (B). The black boxes indicate the position of the regions alignable to the 12 repeat units in the human genome. Annotated transposable elements
in-situ PCR Presented for: Presented by: Date:
 in-situ PCR Presented for: Presented by: Date: 2 in situ Hybridization - Definition in situ PCR is a method in which the polymerase chain reaction actually takes place in the cell on a slide, and the product
in-situ PCR Presented for: Presented by: Date: 2 in situ Hybridization - Definition in situ PCR is a method in which the polymerase chain reaction actually takes place in the cell on a slide, and the product
Methods of Biomaterials Testing Lesson 3-5. Biochemical Methods - Molecular Biology -
 Methods of Biomaterials Testing Lesson 3-5 Biochemical Methods - Molecular Biology - Chromosomes in the Cell Nucleus DNA in the Chromosome Deoxyribonucleic Acid (DNA) DNA has double-helix structure The
Methods of Biomaterials Testing Lesson 3-5 Biochemical Methods - Molecular Biology - Chromosomes in the Cell Nucleus DNA in the Chromosome Deoxyribonucleic Acid (DNA) DNA has double-helix structure The
Single-Molecule Imaging Reveals Dynamics of CREB Transcription Factor Bound to Its Target Sequence
 Supplementary Information Single-Molecule Imaging Reveals Dynamics of CREB Transcription Factor Bound to Its Target Sequence Noriyuki Sugo, Masatoshi Morimatsu, Yoshiyuki Arai, Yoshinori Kousoku, Aya Ohkuni,
Supplementary Information Single-Molecule Imaging Reveals Dynamics of CREB Transcription Factor Bound to Its Target Sequence Noriyuki Sugo, Masatoshi Morimatsu, Yoshiyuki Arai, Yoshinori Kousoku, Aya Ohkuni,
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/10/496/eaam6291/dc1 Supplementary Materials for Regulation of autophagy, NF-κB signaling, and cell viability by mir-124 in KRAS mutant mesenchymal-like NSCLC cells
www.sciencesignaling.org/cgi/content/full/10/496/eaam6291/dc1 Supplementary Materials for Regulation of autophagy, NF-κB signaling, and cell viability by mir-124 in KRAS mutant mesenchymal-like NSCLC cells
Detection of Histone Modifications at Specific Gene Loci in Single Cells in Histological Sections
 Detection of Histone Modifications at Specific Gene Loci in Single Cells in Histological Sections Delphine Gomez; Laura L Shankman; Anh T Nguyen; Gary K Owens. Supplementary Figures 1-13 Supplementary
Detection of Histone Modifications at Specific Gene Loci in Single Cells in Histological Sections Delphine Gomez; Laura L Shankman; Anh T Nguyen; Gary K Owens. Supplementary Figures 1-13 Supplementary
Selected Techniques Part I
 1 Selected Techniques Part I Gel Electrophoresis Can be both qualitative and quantitative Qualitative About what size is the fragment? How many fragments are present? Is there in insert or not? Quantitative
1 Selected Techniques Part I Gel Electrophoresis Can be both qualitative and quantitative Qualitative About what size is the fragment? How many fragments are present? Is there in insert or not? Quantitative
Chapter 6 - Molecular Genetic Techniques
 Chapter 6 - Molecular Genetic Techniques Two objects of molecular & genetic technologies For analysis For generation Molecular genetic technologies! For analysis DNA gel electrophoresis Southern blotting
Chapter 6 - Molecular Genetic Techniques Two objects of molecular & genetic technologies For analysis For generation Molecular genetic technologies! For analysis DNA gel electrophoresis Southern blotting
ProductInformation TECHNICAL BULLETIN MULTIPLE TISSUE NORTHERN BLOT, MOUSE. Product No. BLOT-2 Technical Bulletin No. MB-865 February 2000
 MULTIPLE TISSUE NORTHERN BLOT, MOUSE Product No. BLOT-2 Technical Bulletin No. MB-865 February 2000 ProductInformation TECHNICAL BULLETIN Product Description Sigma s Mouse Multiple Tissue Northern Blot
MULTIPLE TISSUE NORTHERN BLOT, MOUSE Product No. BLOT-2 Technical Bulletin No. MB-865 February 2000 ProductInformation TECHNICAL BULLETIN Product Description Sigma s Mouse Multiple Tissue Northern Blot
Rapid Method for the Purification of Total RNA from Formalin-Fixed Paraffin-Embedded (FFPE) Tissue Samples
 Application Note 17 RNA Sample Preparation Rapid Method for the Purification of Total RNA from Formalin-Fixed Paraffin-Embedded (FFPE) Tissue Samples M. Melmogy 1, V. Misic 1, B. Lam, PhD 1, C. Dobbin,
Application Note 17 RNA Sample Preparation Rapid Method for the Purification of Total RNA from Formalin-Fixed Paraffin-Embedded (FFPE) Tissue Samples M. Melmogy 1, V. Misic 1, B. Lam, PhD 1, C. Dobbin,
Localization Microscopy
 Localization Microscopy Theory, Sample Prep & Practical Considerations Patrina Pellett & Ann McEvoy Applications Scientist GE Healthcare, Cell Technologies May 27 th, 2015 Localization Microscopy Talk
Localization Microscopy Theory, Sample Prep & Practical Considerations Patrina Pellett & Ann McEvoy Applications Scientist GE Healthcare, Cell Technologies May 27 th, 2015 Localization Microscopy Talk
Supporting Information for
 Supporting Information for Building Electromagnetic Hot Spots in Living Cells via Target-Triggered Nanoparticle Dimerization Wen Zhou, 1,2 Qiang Li, 1 Huiqiao Liu, 1 Jie Yang, 1 Dingbin Liu 1,2 * 1. College
Supporting Information for Building Electromagnetic Hot Spots in Living Cells via Target-Triggered Nanoparticle Dimerization Wen Zhou, 1,2 Qiang Li, 1 Huiqiao Liu, 1 Jie Yang, 1 Dingbin Liu 1,2 * 1. College
Multiplexed 3D FRET imaging in deep tissue of live embryos Ming Zhao, Xiaoyang Wan, Yu Li, Weibin Zhou and Leilei Peng
 Scientific Reports Multiplexed 3D FRET imaging in deep tissue of live embryos Ming Zhao, Xiaoyang Wan, Yu Li, Weibin Zhou and Leilei Peng 1 Supplementary figures and notes Supplementary Figure S1 Volumetric
Scientific Reports Multiplexed 3D FRET imaging in deep tissue of live embryos Ming Zhao, Xiaoyang Wan, Yu Li, Weibin Zhou and Leilei Peng 1 Supplementary figures and notes Supplementary Figure S1 Volumetric
Introduction to Fluorescent In Situ Hybridization (FISH)
 Robert Driscoll, M.F.S. Heather Cunningham, M.S. Introduction to Fluorescent In Situ Hybridization (FISH) Fluorescent In Situ Hybridization (FISH) FISH is a cytogenetic technique used to detect the presence
Robert Driscoll, M.F.S. Heather Cunningham, M.S. Introduction to Fluorescent In Situ Hybridization (FISH) Fluorescent In Situ Hybridization (FISH) FISH is a cytogenetic technique used to detect the presence
!! PLEASE READ BEFORE USE!!
 In situ Proximity Ligation Assay protocols!! PLEASE READ BEFORE USE!! The test protocol is a guideline, user need to determine their optimal experimental condition for best performance. The following protocol
In situ Proximity Ligation Assay protocols!! PLEASE READ BEFORE USE!! The test protocol is a guideline, user need to determine their optimal experimental condition for best performance. The following protocol
Propidium Iodide. Catalog Number: Structure: Molecular Formula: C 27H 34I 2N 4. Molecular Weight: CAS #
 Catalog Number: 195458 Propidium Iodide Structure: Molecular Formula: C 27H 34I 2N 4 Molecular Weight: 668.45 CAS # 25535-16-4 Physical Description: Dark red crystals Description: Reagent used for the
Catalog Number: 195458 Propidium Iodide Structure: Molecular Formula: C 27H 34I 2N 4 Molecular Weight: 668.45 CAS # 25535-16-4 Physical Description: Dark red crystals Description: Reagent used for the
hnrnp D/AUF1 Rabbit IgG hnrnp M
 Mouse IgG Goat IgG Rabbit IgG Mouse IgG hnrnp F Goat IgG Mouse IgG Kb 6 4 3 2 15 5 Supplementary Figure S1. In vivo binding of TERRA-bound RBPs to target RNAs. Immunoprecipitation (IP) assay using 3 mg
Mouse IgG Goat IgG Rabbit IgG Mouse IgG hnrnp F Goat IgG Mouse IgG Kb 6 4 3 2 15 5 Supplementary Figure S1. In vivo binding of TERRA-bound RBPs to target RNAs. Immunoprecipitation (IP) assay using 3 mg
Figure S6. Detection of anti-gfp antibodies in anti-dna and normal plasma without competition DNA--9
 Supplementary Information Ultrasensitive antibody detection by agglutination-pcr (ADAP) Cheng-ting Tsai 1 *, Peter V. Robinson 1 *, Carole A. Spencer 2 and Carolyn R. Bertozzi 3,4ǂ Department of 1 Chemistry,
Supplementary Information Ultrasensitive antibody detection by agglutination-pcr (ADAP) Cheng-ting Tsai 1 *, Peter V. Robinson 1 *, Carole A. Spencer 2 and Carolyn R. Bertozzi 3,4ǂ Department of 1 Chemistry,
Immunofluorescence Confocal Microscopy of 3D Cultures Grown on Alvetex
 Immunofluorescence Confocal Microscopy of 3D Cultures Grown on Alvetex 1.0. Introduction Immunofluorescence uses the recognition of cellular targets by fluorescent dyes or antigen-specific antibodies coupled
Immunofluorescence Confocal Microscopy of 3D Cultures Grown on Alvetex 1.0. Introduction Immunofluorescence uses the recognition of cellular targets by fluorescent dyes or antigen-specific antibodies coupled
Supplementary Figure 1 Strategy for parallel detection of DHSs and adjacent nucleosomes
 Supplementary Figure 1 Strategy for parallel detection of DHSs and adjacent nucleosomes DNase I cleavage DNase I DNase I digestion Sucrose gradient enrichment Small Large F1 F2...... F9 F1 F1 F2 F3 F4
Supplementary Figure 1 Strategy for parallel detection of DHSs and adjacent nucleosomes DNase I cleavage DNase I DNase I digestion Sucrose gradient enrichment Small Large F1 F2...... F9 F1 F1 F2 F3 F4
Roche Molecular Biochemicals Technical Note No. LC 12/2000
 Roche Molecular Biochemicals Technical Note No. LC 12/2000 LightCycler Absolute Quantification with External Standards and an Internal Control 1. General Introduction Purpose of this Note Overview of Method
Roche Molecular Biochemicals Technical Note No. LC 12/2000 LightCycler Absolute Quantification with External Standards and an Internal Control 1. General Introduction Purpose of this Note Overview of Method
SUPPORTING INFORMATION. A cleavage-responsive stem-loop hairpin for assaying guide RNA activity
 SUPPORTING INFORMATION A cleavage-responsive stem-loop hairpin for assaying guide RNA activity Tara R. deboer 1, Noreen Wauford 1, Jing-Yi Chung, Miguel Salvador Torres Perez, and Niren Murthy* University
SUPPORTING INFORMATION A cleavage-responsive stem-loop hairpin for assaying guide RNA activity Tara R. deboer 1, Noreen Wauford 1, Jing-Yi Chung, Miguel Salvador Torres Perez, and Niren Murthy* University
Supplemental Figure 1 Human REEP family of proteins can be divided into two distinct subfamilies. Residues (single letter amino acid code) identical
 Supplemental Figure Human REEP family of proteins can be divided into two distinct subfamilies. Residues (single letter amino acid code) identical in all six REEPs are highlighted in green. Additional
Supplemental Figure Human REEP family of proteins can be divided into two distinct subfamilies. Residues (single letter amino acid code) identical in all six REEPs are highlighted in green. Additional
PCNA appears in two populations of slow and fast diffusion with a constant ratio throughout S-phase in replicating mammalian cells
 PCNA appears in two populations of slow and fast diffusion with a constant ratio throughout S-phase in replicating mammalian cells Patrick J. M. Zessin 1, Anje Sporbert 2, *, Mike Heilemann 1, * 1 Institute
PCNA appears in two populations of slow and fast diffusion with a constant ratio throughout S-phase in replicating mammalian cells Patrick J. M. Zessin 1, Anje Sporbert 2, *, Mike Heilemann 1, * 1 Institute
Supplemental Information. Dynamic Organization of Chromatin Domains. Revealed by Super-Resolution Live-Cell Imaging
 Molecular Cell, Volume 67 Supplemental Information Dynamic Organization of Chromatin Domains Revealed by Super-Resolution Live-Cell Imaging Tadasu Nozaki, Ryosuke Imai, Mai Tanbo, Ryosuke Nagashima, Sachiko
Molecular Cell, Volume 67 Supplemental Information Dynamic Organization of Chromatin Domains Revealed by Super-Resolution Live-Cell Imaging Tadasu Nozaki, Ryosuke Imai, Mai Tanbo, Ryosuke Nagashima, Sachiko
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Endogenous gene tagging to study subcellular localization and chromatin binding. a, b, Schematic of experimental set-up to endogenously tag RNAi factors using the CRISPR Cas9 technology,
Supplementary Figure 1 Endogenous gene tagging to study subcellular localization and chromatin binding. a, b, Schematic of experimental set-up to endogenously tag RNAi factors using the CRISPR Cas9 technology,
Introduction to N-STORM
 Introduction to N-STORM Dan Metcalf Advanced Imaging Manager Outline Introduction Principles of STORM Applications N-STORM overview Biological Scale Mitochondrion Microtubule Amino Acid 1Å Kinesin 1nm
Introduction to N-STORM Dan Metcalf Advanced Imaging Manager Outline Introduction Principles of STORM Applications N-STORM overview Biological Scale Mitochondrion Microtubule Amino Acid 1Å Kinesin 1nm
T H E J O U R N A L O F C E L L B I O L O G Y
 Supplemental material Wang et al., http://www.jcb.org/cgi/content/full/jcb.201405026/dc1 T H E J O U R N A L O F C E L L B I O L O G Y Figure S1. Generation and characterization of unc-40 alleles. (A and
Supplemental material Wang et al., http://www.jcb.org/cgi/content/full/jcb.201405026/dc1 T H E J O U R N A L O F C E L L B I O L O G Y Figure S1. Generation and characterization of unc-40 alleles. (A and
MEFs were treated with the indicated concentrations of LLOMe for three hours, washed
 Supplementary Materials and Methods Cell Fractionation MEFs were treated with the indicated concentrations of LLOMe for three hours, washed with ice-cold PBS, collected by centrifugation, and then homogenized
Supplementary Materials and Methods Cell Fractionation MEFs were treated with the indicated concentrations of LLOMe for three hours, washed with ice-cold PBS, collected by centrifugation, and then homogenized
Comparative Genomic Hybridization
 Comparative Genomic Hybridization Srikesh G. Arunajadai Division of Biostatistics University of California Berkeley PH 296 Presentation Fall 2002 December 9 th 2002 OUTLINE CGH Introduction Methodology,
Comparative Genomic Hybridization Srikesh G. Arunajadai Division of Biostatistics University of California Berkeley PH 296 Presentation Fall 2002 December 9 th 2002 OUTLINE CGH Introduction Methodology,
DENAzist Asia's High Quality Kits and Molecular Biology Products Price list_version Cat. No. Name Application Price (Rial)
 RNA Isolation kits S1010 Total RNA isolation kit (25) A unique combination of solutions for extraction of RNA from cells, tissues, 1,190,000 S10101 Total RNA isolation kit (50) plants, bacteria, yeasts.
RNA Isolation kits S1010 Total RNA isolation kit (25) A unique combination of solutions for extraction of RNA from cells, tissues, 1,190,000 S10101 Total RNA isolation kit (50) plants, bacteria, yeasts.
Please purchase PDFcamp Printer on to remove this watermark. DNA microarray
 DNA microarray Example of an approximately 40,000 probe spotted oligo microarray with enlarged inset to show detail. A DNA microarray is a multiplex technology used in molecular biology. It consists of
DNA microarray Example of an approximately 40,000 probe spotted oligo microarray with enlarged inset to show detail. A DNA microarray is a multiplex technology used in molecular biology. It consists of
Real-Time PCR Principles and Applications
 Real-Time PCR Principles and Applications Dr Esam Ibraheem Azhar (BSc, MSc, Ph.D Molecular Medical Virology) Asst. Prof. Medical Laboratory Technology Department Objectives Real-Time PCR Principles and
Real-Time PCR Principles and Applications Dr Esam Ibraheem Azhar (BSc, MSc, Ph.D Molecular Medical Virology) Asst. Prof. Medical Laboratory Technology Department Objectives Real-Time PCR Principles and
Intercalation-based single-molecule fluorescence assay to study DNA supercoil
 Intercalation-based single-molecule fluorescence assay to study DNA supercoil dynamics Mahipal Ganji, Sung Hyun Kim, Jaco van der Torre, Elio Abbondanzieri*, Cees Dekker* Supplementary information Supplementary
Intercalation-based single-molecule fluorescence assay to study DNA supercoil dynamics Mahipal Ganji, Sung Hyun Kim, Jaco van der Torre, Elio Abbondanzieri*, Cees Dekker* Supplementary information Supplementary
Visualizing mechanical tension across membrane receptors with a fluorescent sensor
 Nature Methods Visualizing mechanical tension across membrane receptors with a fluorescent sensor Daniel R. Stabley, Carol Jurchenko, Stephen S. Marshall, Khalid S. Salaita Supplementary Figure 1 Fabrication
Nature Methods Visualizing mechanical tension across membrane receptors with a fluorescent sensor Daniel R. Stabley, Carol Jurchenko, Stephen S. Marshall, Khalid S. Salaita Supplementary Figure 1 Fabrication
SUPPLEMENTARY INFORMATION
 DOI:.38/ncb327 a b Sequence coverage (%) 4 3 2 IP: -GFP isoform IP: GFP IP: -GFP IP: GFP Sequence coverage (%) 4 3 2 IP: -GFP IP: GFP 33 52 58 isoform 2 33 49 47 IP: Control IP: Peptide Sequence Start
DOI:.38/ncb327 a b Sequence coverage (%) 4 3 2 IP: -GFP isoform IP: GFP IP: -GFP IP: GFP Sequence coverage (%) 4 3 2 IP: -GFP IP: GFP 33 52 58 isoform 2 33 49 47 IP: Control IP: Peptide Sequence Start
Supporting Information
 Supporting Information Deng et al. 10.1073/pnas.1515692112 SI Materials and Methods FPLC. All fusion proteins were expressed and purified through a three-step FPLC purification protocol, as described (20),
Supporting Information Deng et al. 10.1073/pnas.1515692112 SI Materials and Methods FPLC. All fusion proteins were expressed and purified through a three-step FPLC purification protocol, as described (20),
Supplementary Figure 1. Supporting data for Figure 1
 Supplementary Figure 1 Supporting data for Figure 1 (A) Schematic of BLI assay used to measure dissociation. 5 monobiotinylated substrate DNA (identical to λ1, Figure 2) is associated with streptavidin-coated
Supplementary Figure 1 Supporting data for Figure 1 (A) Schematic of BLI assay used to measure dissociation. 5 monobiotinylated substrate DNA (identical to λ1, Figure 2) is associated with streptavidin-coated
Multiple choice questions (numbers in brackets indicate the number of correct answers)
 1 Multiple choice questions (numbers in brackets indicate the number of correct answers) February 1, 2013 1. Ribose is found in Nucleic acids Proteins Lipids RNA DNA (2) 2. Most RNA in cells is transfer
1 Multiple choice questions (numbers in brackets indicate the number of correct answers) February 1, 2013 1. Ribose is found in Nucleic acids Proteins Lipids RNA DNA (2) 2. Most RNA in cells is transfer
Validation of Hits from an sirna Library Screen using Real-Time qpcr. Gregory L. Shipley The University of Texas Health Science Center- Houston
 Validation of Hits from an sirna Library Screen using Real-Time qpcr Gregory L. Shipley The University of Texas Health Science Center- Houston sirnas and Gene Silencing Small interfering RNAs or sirnas
Validation of Hits from an sirna Library Screen using Real-Time qpcr Gregory L. Shipley The University of Texas Health Science Center- Houston sirnas and Gene Silencing Small interfering RNAs or sirnas
Supplemental Table/Figure Legends
 MiR-26a is required for skeletal muscle differentiation and regeneration in mice Bijan K. Dey, Jeffrey Gagan, Zhen Yan #, Anindya Dutta Supplemental Table/Figure Legends Suppl. Table 1: Effect of overexpression
MiR-26a is required for skeletal muscle differentiation and regeneration in mice Bijan K. Dey, Jeffrey Gagan, Zhen Yan #, Anindya Dutta Supplemental Table/Figure Legends Suppl. Table 1: Effect of overexpression
DAPI mrna intron. Chromosome 19 UBA2 EIF3K
 Supplementary Figure DAPI mrna intron a b c d p-arm Chromosome 9 q-arm.79 3.98.2.76 2.79 7.97 8.68 23. 3.92 39. 39.9.33.3 2.36.39.7 7.28 2.69.7 6.6 e intron mrna Chr. 9 Supplementary Fig.. Targeting 2
Supplementary Figure DAPI mrna intron a b c d p-arm Chromosome 9 q-arm.79 3.98.2.76 2.79 7.97 8.68 23. 3.92 39. 39.9.33.3 2.36.39.7 7.28 2.69.7 6.6 e intron mrna Chr. 9 Supplementary Fig.. Targeting 2
RNA oligonucleotides and 2 -O-methylated oligonucleotides were synthesized by. 5 AGACACAAACACCAUUGUCACACUCCACAGC; Rand-2 OMe,
 Materials and methods Oligonucleotides and DNA constructs RNA oligonucleotides and 2 -O-methylated oligonucleotides were synthesized by Dharmacon Inc. (Lafayette, CO). The sequences were: 122-2 OMe, 5
Materials and methods Oligonucleotides and DNA constructs RNA oligonucleotides and 2 -O-methylated oligonucleotides were synthesized by Dharmacon Inc. (Lafayette, CO). The sequences were: 122-2 OMe, 5
Supplementary Data. Flvcr1a TCTAAGGCCCAGTAGGACCC GGCCTCAACTGCCTGGGAGC AGAGGGCAACCTCGGTGTCC
 Supplementary Data Supplementary Materials and Methods Measurement of reactive oxygen species accumulation in fresh intestinal rings Accumulation of reactive oxygen species in fresh intestinal rings was
Supplementary Data Supplementary Materials and Methods Measurement of reactive oxygen species accumulation in fresh intestinal rings Accumulation of reactive oxygen species in fresh intestinal rings was
Figure S1 Proteasome inhibition leads to formation of aggregates in human cells and tissues. (a)
 SUPPLEMENTARY MATERIAL Figure S1 Proteasome inhibition leads to formation of aggregates in human cells and tissues. (a) Flow cytometry. Cells were treated with MG132 for the indicated times. Cells were
SUPPLEMENTARY MATERIAL Figure S1 Proteasome inhibition leads to formation of aggregates in human cells and tissues. (a) Flow cytometry. Cells were treated with MG132 for the indicated times. Cells were
Stellaris RNA FISH Protocol for C. elegans
 Stellaris RNA FISH Protocol for C. elegans General Protocol & Storage Product Description A set of Stellaris RNA FISH Probes is comprised of up to 48 singly labeled oligonucleotides designed to selectively
Stellaris RNA FISH Protocol for C. elegans General Protocol & Storage Product Description A set of Stellaris RNA FISH Probes is comprised of up to 48 singly labeled oligonucleotides designed to selectively
Protein Electrophoresis EZ-Run Protein Gel Solution EZ-Run Protein Standards EZ-Run Gel Staining Solution Traditional SDS-PAGE Reagents
 Protein Electrophoresis EZ-Run Protein Gel Solution EZ-Run Protein Standards EZ-Run Gel Staining Solution Traditional SDS-PAGE Reagents Reliability. Purity. Certainty. Introduction Sodium dodecyl sulfate
Protein Electrophoresis EZ-Run Protein Gel Solution EZ-Run Protein Standards EZ-Run Gel Staining Solution Traditional SDS-PAGE Reagents Reliability. Purity. Certainty. Introduction Sodium dodecyl sulfate
Microarray. Key components Array Probes Detection system. Normalisation. Data-analysis - ratio generation
 Microarray Key components Array Probes Detection system Normalisation Data-analysis - ratio generation MICROARRAY Measures Gene Expression Global - Genome wide scale Why Measure Gene Expression? What information
Microarray Key components Array Probes Detection system Normalisation Data-analysis - ratio generation MICROARRAY Measures Gene Expression Global - Genome wide scale Why Measure Gene Expression? What information
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10016 Supplementary discussion on binding site density for protein complexes on the surface: The density of biotin sites on the chip is ~10 3 biotin-peg per µm 2. The biotin sites are
doi:10.1038/nature10016 Supplementary discussion on binding site density for protein complexes on the surface: The density of biotin sites on the chip is ~10 3 biotin-peg per µm 2. The biotin sites are
Supplementary Figure 1
 Supplementary Figure 1 CITE-seq library preparation. (a) Illustration of the DNA-barcoded antibodies used in CITE-seq. (b) Antibody-oligonucleotide complexes appear as a high-molecular-weight smear when
Supplementary Figure 1 CITE-seq library preparation. (a) Illustration of the DNA-barcoded antibodies used in CITE-seq. (b) Antibody-oligonucleotide complexes appear as a high-molecular-weight smear when
7/24/2012. DNA Probes. Hybridization and Probes. CLS 420 Immunology & Molecular Diagnostics. Target Sequences. Target Sequences. Nucleic Acid Probes
 Hybridization and Probes CLS 420 Immunology & Molecular Diagnostics Molecular Diagnostics Techniques: Hybridization and Probes Nucleic acid probes: A short, known sequence of DNA or RNA Used to detect
Hybridization and Probes CLS 420 Immunology & Molecular Diagnostics Molecular Diagnostics Techniques: Hybridization and Probes Nucleic acid probes: A short, known sequence of DNA or RNA Used to detect
catalytic hairpin DNA assembly for dual-signal amplification toward homogenous analysis of protein and
 Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2015 Supporting Information Programmable Mg 2+ -dependent DNAzyme switch by the catalytic hairpin DNA
Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2015 Supporting Information Programmable Mg 2+ -dependent DNAzyme switch by the catalytic hairpin DNA
Technical Manual No Version
 TUNEL Apoptosis Detection Kit Cat. No. L00301 (For Cryopreserved Tissue Sections, FITC-labled POD) Technical Manual No. 0269 Version 01132011 I Description. 1 II Key Features.... 1 III Kit Contents.. 1
TUNEL Apoptosis Detection Kit Cat. No. L00301 (For Cryopreserved Tissue Sections, FITC-labled POD) Technical Manual No. 0269 Version 01132011 I Description. 1 II Key Features.... 1 III Kit Contents.. 1
ViewRNA ISH Cell Assays. Visualize RNA with single-molecule sensitivity and single-cell resolution
 ViewRNA ISH Cell Assays Visualize RNA with single-molecule sensitivity and single-cell resolution ViewRNA ISH Cell Assays Analyze sample heterogeneity Study noncoding RNAs, including mirna and lncrna,
ViewRNA ISH Cell Assays Visualize RNA with single-molecule sensitivity and single-cell resolution ViewRNA ISH Cell Assays Analyze sample heterogeneity Study noncoding RNAs, including mirna and lncrna,
Quantitative Real Time PCR USING SYBR GREEN
 Quantitative Real Time PCR USING SYBR GREEN SYBR Green SYBR Green is a cyanine dye that binds to double stranded DNA. When it is bound to D.S. DNA it has a much greater fluorescence than when bound to
Quantitative Real Time PCR USING SYBR GREEN SYBR Green SYBR Green is a cyanine dye that binds to double stranded DNA. When it is bound to D.S. DNA it has a much greater fluorescence than when bound to
ApoTrack Cytochrome c Apoptosis ICC Antibody Kit: 2 color immunocytochemistry of cytochrome c and mitochondria.
 PROTOCOL ApoTrack Cytochrome c Apoptosis ICC Antibody Kit 1850 Millrace Drive, Suite 3A Eugene, Oregon 97403 MSA07 Rev.1 DESCRIPTION ApoTrack Cytochrome c Apoptosis ICC Antibody Kit: 2 color immunocytochemistry
PROTOCOL ApoTrack Cytochrome c Apoptosis ICC Antibody Kit 1850 Millrace Drive, Suite 3A Eugene, Oregon 97403 MSA07 Rev.1 DESCRIPTION ApoTrack Cytochrome c Apoptosis ICC Antibody Kit: 2 color immunocytochemistry
Coleman et al., Supplementary Figure 1
 Coleman et al., Supplementary Figure 1 BrdU Merge G1 Early S Mid S Supplementary Figure 1. Sequential destruction of CRL4 Cdt2 targets during the G1/S transition. HCT116 cells were synchronized by sequential
Coleman et al., Supplementary Figure 1 BrdU Merge G1 Early S Mid S Supplementary Figure 1. Sequential destruction of CRL4 Cdt2 targets during the G1/S transition. HCT116 cells were synchronized by sequential
Nature Immunology: doi: /ni Supplementary Figure 1. Construction of lnckdm2b RFP reporter and lnckdm2b-knockout mice.
 Supplementary Figure 1 Construction of lnckdm2b RFP reporter and lnckdm2b-knockout mice. (a) ILC3s (Lin CD45 lo CD90 hi ) were isolated from small intestines, followed by immunofluorescence staining. LncKdm2b
Supplementary Figure 1 Construction of lnckdm2b RFP reporter and lnckdm2b-knockout mice. (a) ILC3s (Lin CD45 lo CD90 hi ) were isolated from small intestines, followed by immunofluorescence staining. LncKdm2b
Real Time PCR. Advanced Biotechnology Lab I Florida Atlantic University April 2, 2008
 Real Time PCR Advanced Biotechnology Lab I Florida Atlantic University April 2, 2008 Introduction We wish to compare the expression levels of our gene under study (Drosophila MsrA) for two different treatment
Real Time PCR Advanced Biotechnology Lab I Florida Atlantic University April 2, 2008 Introduction We wish to compare the expression levels of our gene under study (Drosophila MsrA) for two different treatment
T H E J O U R N A L O F C E L L B I O L O G Y
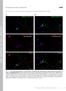 T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Kanaani et al., http://www.jcb.org/cgi/content/full/jcb.200912101/dc1 Figure S1. The K2 rabbit polyclonal antibody is specific for GAD67,
T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Kanaani et al., http://www.jcb.org/cgi/content/full/jcb.200912101/dc1 Figure S1. The K2 rabbit polyclonal antibody is specific for GAD67,
Figure S1. Related to Figure 1, to show 3-D images of nerve/vessel patterning.
 Supplemental Information Inventory of Supplemental Information Figure S1. Related to Figure 1, to show 3-D images of nerve/vessel patterning. Figure S2. Related to Figure 4, to validate Npn-1 signaling
Supplemental Information Inventory of Supplemental Information Figure S1. Related to Figure 1, to show 3-D images of nerve/vessel patterning. Figure S2. Related to Figure 4, to validate Npn-1 signaling
Increased transcription detection with the NEBNext Single Cell/Low Input RNA Library Prep Kit
 be INSPIRED drive DISCOVERY stay GENUINE TECHNICAL NOTE Increased transcription detection with the NEBNext Single Cell/Low Input RNA Library Prep Kit Highly sensitive, robust generation of high quality
be INSPIRED drive DISCOVERY stay GENUINE TECHNICAL NOTE Increased transcription detection with the NEBNext Single Cell/Low Input RNA Library Prep Kit Highly sensitive, robust generation of high quality
Sensitivity vs Specificity
 Viral Detection Animal Inoculation Culturing the Virus Definitive Length of time Serology Detecting antibodies to the infectious agent Detecting Viral Proteins Western Blot ELISA Detecting the Viral Genome
Viral Detection Animal Inoculation Culturing the Virus Definitive Length of time Serology Detecting antibodies to the infectious agent Detecting Viral Proteins Western Blot ELISA Detecting the Viral Genome
