DNA Binding Domains: Structural Motifs. Effector Domain. Zinc Fingers. Zinc Fingers, continued. Zif268
|
|
|
- Berenice Blankenship
- 6 years ago
- Views:
Transcription
1 DNA Binding Domains: Structural Motifs Studies of known transcription factors have found several motifs of protein design to allow sequence-specific binding of DNA. We will cover only three of these motifs: Zinc fingers Homeodomains Leucine Zipper (bzip) & bhlh Effector Domain May work by steric hindrance Usually, a region of protein interacts with RNA pol II or another transcription factor Some TFs have effector domains rich in acidic amino acids (GAL4) Some rich in glutamine (Sp1 two domains of 25% glutamine) Some rich in proline (CTF, nearly 25% pro) Zinc Fingers Repeated elements of a 30-aa sequence containing cyteines, histidines or a combination in an orientation that coordinates a zinc ion in space. Finger-shaped loop of amino acids fits well into the major groove, binding several bp each Zinc Fingers, continued There are usually several of these elements repeated in the structure of the protein. That provides for binding of larger sequence elements over a longer stretch of sequence. Consists of α-helix and β-sheet held together by zinc ion. Many types - includes glucocorticoid receptor Zif268 Immediate-early protein one of the first genes activated upon mitogenic stimulation. Three Zn fingers, each a curled shape coordinated by a Zn ion. The 3 Zn fingers form a C shape that locks onto the DNA binding in the major groove. Target bases all lie on one strand G-rich motif. 1
2 One Zinc Finger of Zif268 3 Zn Fingers in Zif268 coordinately bind DNA Zn ++ α-helix anti-parallel β-strand Location of interactions on DNA Zif268 zinc finger-dna complex with Zn 2+ ions in yellow Yeast GAL4 The GAL4-DNA Complex Controls expression of a group of genes metabolizing galactose in yeast. Each target gene contains a GAL4-binding site (called a UAS G ). Binds to UAS G as a dimer. Requires two domains: DNA Binding domain: two Zn ++ ions coordinating a short α-helical region. Dimerization domain: a parallel coiled coil of two α-helical regions. 2
3 Separation of GAL4 Domains Possible to separate DNA binding domain from transcription effector domain. GAL4 binds UAS G site and stimulates transcription in yeast. LexA is a prokaryotic repressor, binds lexa operator, inhibits transcription. Build chimeric proteins. The Chimera has the head of a lion, body of a goat, tail of a serpent. DNA Binding & Transcription Effector domains can be swapped endogenous GAL4 activates trxn. LexA inhibits trxn, LexA-GAL4 substitutes GAL4 s transcription activator domain & activates Nuclear Receptors Class of Zn finger transcription factors that are also hormone receptors. Regulatory domain = ligand binding domain. Sex hormones, glucocorticoids, Vitamin D, thyroid hormone, retinoic acid. Exist in cytoplasm inactive, bound to hsp90. Some are always nuclear (thyroid hormone receptor) bind same site and repress in absence of ligand. 3
4 Homeodomain (helix-turn-helix) Antennapedia Mutation Two short α-helical regions that fit into the major groove of DNA separated by a nonhelical region that provides a "bend" or turn. 180 bp (60 aa) conserved region (homeobox) Best example - homeodomain proteins involved in differentiation. Engrailed, antennapedia, ultrabithorax Leucine Zipper Two domains - LZ & DBD DNA Binding Domain (DBD) - contains basic amino acids & provides specificity. Leucine Zipper (LZ) - combines two polypeptide chains in correct shape for interacting with major groove. LZ Domain α-helical (about 3.5 aa/turn) leu each 7 amino acids - same face of helix Two subunits interact through LZ domains Subunits may be identical (homodimer) or different (heterodimer) Combinations allow diversity of activity. DBD (Basic Region) LZ LZ DBD (Basic Region) 4
5 Coiled coil interaction Scissors Grip Binding of DNA Both LZ & Basic Region Required for Activity of DNA Binding Domain Without dimerization, basic region cannot interact with major groove. Since dimerization region (LZ) required for DNA binding function, it must be considered part of DNA binding domain, along with basic region. Two Theories of Transcription Initiation Basal transcription factors cause a stepwise build-up of pre-initiation complex on DNA. Basal factors already bound to holoenzyme, bind DNA as a unit. 5
6 Regulation by recruitment of basal factors The herpesvirus factor VP16 was shown to bind TFIID through its acidic effector domain. GAL4 seems to interact with TFIIB. Promoter constructed with Ad E4 sequences UAS G added Attached to beads Add Factors & Wash Add each factor, one at a time. Wash off unbound factor. Add next factor. Did bound first factor enable assembly with second factor? GAL4 stimulates assembly of Basal Complex Presence of GAL4 in first step greatly facilitated assembly of transcription complex. Binding of TFIID, however, was not affected by GAL4 presence. Binding of TFIIB was enhanced by GAL4 presence. Lack of TFIIB limited transcription. TFIIB binding requires TFIID, is enhanced by transcription activator (here using an acidic region). Multiple factors act at a distance in a coordinated fashion Each factor binds its own recognition sequence, which may be far away from start site. Complex forms by protein-protein interactions. Requires bending of dsdna. Four theories TF binds sequence and changes shape of DNA, possibly inducing supercoils. TF binds sequence then slides along DNA to find promoter complex. TF binds sequence, loops intervening sequences to interact directly with trxn complex. TF binds sequence, binds second sequence and tracks along DNA. 6
7 Looping proved by catenane construction Two plasmids linked together by catenation were able to interact to stimulate transcription Interferon-β promoter/enhancer IFNβ induced by viral infection. Four binding sites for HMG I(Y), a factor that bends DNA; required for binding by NFκB and ATF-2. NFκB induced by inflammatory responses, unbends DNA; interacts with many promoters. All factors must bind to form enhanceosome complex requires coordinate regulation of all factors. Interferon Regulatory factor Inserting 6 bp between HMGI sites reduces cooperative binding, but inserting 10 bp did not. Mediators (co-activators) Non-DNA binding proteins that promote or interfere with transcription activation. Example is CBP (CREB-binding protein) CREB (camp response element binding protein), activated by phosphorylation by PKA in response to camp, binds CRE (camp response element). But CREB binds CRE even when not phosphorylated how does it work? Phosphorylated CREB binds CBP, which in turn activates basal transcription complex. 7
8 CBP also found in MAP Kinase Cascade Addition of growth factors (mitogens) stimulates activition of mitogen-activated protein kinase (MAPK) by phosphorylation. Phospho-MAPK enters nucleus, phosphorylates factors like Sap-1a, jun. These TFs use CBP to mediate activation of promoters. CBP is also a histone acetyl transferase enzyme, can loosen histones on DNA. Convergence of multiple pathways on a common intermediate forces interaction of pathways. Integration of Signaling Cascades Mitogen acts at extracellular receptor to stimulate receptor dimerization (activation) & activation of tyrosine kinase. Receptor monomers phosphorylate each other. Phosphotyrosines recognized by GRB2, an adaptor. GRB2 now binds the ras exchanger Sos. Sos replaces GDP bound to ras with GTP. Ras delivers Raf to cell membrane, where it phosporylates MAPKK. MAPKK phosphorylates MAPK, which translocates to nucleus. MAPK phosporylates jun. Jun associates with fos to make AP-1. AP-1 binds an AP-1 site and stimulates transcription through interaction with CBP. Why a cascade? Each step amplifies the signal! Signal transduction linked to transcription factors Each TF regulated by some signaling pathway using some mechanism (phosphorylation, ligand binding, release from cytoplasm ) TF bind sites on DNA (enhancers). Multiple TF/enhancer complexes interact by looping DNA. Multiple regulatory complexes coordinate binding and/or activation activities to integrate signaling. Some activating domains increase association of TFIIB, others TFIID. Affect efficiency of transcription initiation! 8
Enhancers. Activators and repressors of transcription
 Enhancers Can be >50 kb away from the gene they regulate. Can be upstream from a promoter, downstream from a promoter, within an intron, or even downstream of the final exon of a gene. Are often cell type
Enhancers Can be >50 kb away from the gene they regulate. Can be upstream from a promoter, downstream from a promoter, within an intron, or even downstream of the final exon of a gene. Are often cell type
Eukaryotic transcription (II)
 Eukaryotic transcription (II) Transcription factors Prokaryote: Sigma factors they are similar to general transcription factor of eukaryotes, they help bring RNA pol to the promoter Transcription factors
Eukaryotic transcription (II) Transcription factors Prokaryote: Sigma factors they are similar to general transcription factor of eukaryotes, they help bring RNA pol to the promoter Transcription factors
Chapter 25: Regulating Eukaryotic Transcription The Ligand Responsive Activators
 Chapter 25: Regulating Eukaryotic Transcription The Ligand Responsive Activators At least 5 potential gene expression control points Superfamily of Gene Regulators Activation of gene structure Initiation
Chapter 25: Regulating Eukaryotic Transcription The Ligand Responsive Activators At least 5 potential gene expression control points Superfamily of Gene Regulators Activation of gene structure Initiation
Structure/function relationship in DNA-binding proteins
 PHRM 836 September 22, 2015 Structure/function relationship in DNA-binding proteins Devlin Chapter 8.8-9 u General description of transcription factors (TFs) u Sequence-specific interactions between DNA
PHRM 836 September 22, 2015 Structure/function relationship in DNA-binding proteins Devlin Chapter 8.8-9 u General description of transcription factors (TFs) u Sequence-specific interactions between DNA
Molecular Biology (BIOL 4320) Exam #1 March 12, 2002
 Molecular Biology (BIOL 4320) Exam #1 March 12, 2002 Name KEY SS# This exam is worth a total of 100 points. The number of points each question is worth is shown in parentheses after the question number.
Molecular Biology (BIOL 4320) Exam #1 March 12, 2002 Name KEY SS# This exam is worth a total of 100 points. The number of points each question is worth is shown in parentheses after the question number.
Transcription Activators in Eukaryotes. Chapter 12
 Transcription Activators in Eukaryotes Chapter 12 Techniques Run-off transcription Primer extension Quantitative S1 mapping Modular Protein domain is an independingly folding region. DA binding domain
Transcription Activators in Eukaryotes Chapter 12 Techniques Run-off transcription Primer extension Quantitative S1 mapping Modular Protein domain is an independingly folding region. DA binding domain
Eukaryotic Gene Expression Prof. P. N. Rangarajan Department of Biochemistry Indian Institute of Science, Bangalore
 Eukaryotic Gene Expression Prof. P. N. Rangarajan Department of Biochemistry Indian Institute of Science, Bangalore Lecture No. # 06 Eukaryotic Transcription Factors: Transcription Activation Domains In
Eukaryotic Gene Expression Prof. P. N. Rangarajan Department of Biochemistry Indian Institute of Science, Bangalore Lecture No. # 06 Eukaryotic Transcription Factors: Transcription Activation Domains In
Gene regulation V Biochemistry 302. March 6, 2006
 Gene regulation V Biochemistry 302 March 6, 2006 Common structural motifs associated with transcriptional regulatory proteins Helix-turn-helix Prokaryotic repressors and activators Eukaryotic homeodomain
Gene regulation V Biochemistry 302 March 6, 2006 Common structural motifs associated with transcriptional regulatory proteins Helix-turn-helix Prokaryotic repressors and activators Eukaryotic homeodomain
Gene expression DNA RNA. Protein DNA. Replication. Initiation Elongation Processing Export. DNA RNA Protein. Transcription. Degradation.
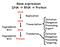 Gene expression DNA RNA Protein DNA DNA Degradation RNA Degradation Protein Replication Transcription Translation Initiation Elongation Processing Export Initiation Elongation Processing Targeting Chapter
Gene expression DNA RNA Protein DNA DNA Degradation RNA Degradation Protein Replication Transcription Translation Initiation Elongation Processing Export Initiation Elongation Processing Targeting Chapter
Regulation of gene expression. (Lehninger pg )
 Regulation of gene expression (Lehninger pg. 1072-1085) Today s lecture Gene expression Constitutive, inducible, repressible genes Specificity factors, activators, repressors Negative and positive gene
Regulation of gene expression (Lehninger pg. 1072-1085) Today s lecture Gene expression Constitutive, inducible, repressible genes Specificity factors, activators, repressors Negative and positive gene
Another groups asks with what does Gal4- VP16 interact?
 Another groups asks with what does Gal4- VP16 interact? DNA attached to agarose beads Now add components in steps Ask if the recruitment of a component requires that Gal4-VP16 be present. Primer extension
Another groups asks with what does Gal4- VP16 interact? DNA attached to agarose beads Now add components in steps Ask if the recruitment of a component requires that Gal4-VP16 be present. Primer extension
Overlap with exam 1 begins
 Exam 2 begins here Overlap with exam 1 begins Another groups asks with what does Gal4- VP16 interact? DNA attached to agarose beads Now add components in steps Ask if the recruitment of a component requires
Exam 2 begins here Overlap with exam 1 begins Another groups asks with what does Gal4- VP16 interact? DNA attached to agarose beads Now add components in steps Ask if the recruitment of a component requires
Synthetic cells: do bacteria need all its genes? No.
 NO NEED TO REFER TO THE SLIDES. بسم هللا الرحمن الرحيم Do we need all the non coding regions of the DNA? Two weeks ago, they discovered that the genome of a plant is very small (recall that plant genome
NO NEED TO REFER TO THE SLIDES. بسم هللا الرحمن الرحيم Do we need all the non coding regions of the DNA? Two weeks ago, they discovered that the genome of a plant is very small (recall that plant genome
Transcription factors
 Atlas of Genetics and Cytogenetics in Oncology and Haematology Transcription factors I Introduction * II Initiation of transcription III Transcription factors family pdf version I Introduction III.1 Helix-Turn-Helix
Atlas of Genetics and Cytogenetics in Oncology and Haematology Transcription factors I Introduction * II Initiation of transcription III Transcription factors family pdf version I Introduction III.1 Helix-Turn-Helix
Complex Transcription Machinery
 Complex Transcription Machinery Subunits of the Basal Txn Apparatus Stepwise Assembly of the Pre-initiation Complex Interplay of Activators, Co-regulators and RNA Polymerase at the Promoter Divide and
Complex Transcription Machinery Subunits of the Basal Txn Apparatus Stepwise Assembly of the Pre-initiation Complex Interplay of Activators, Co-regulators and RNA Polymerase at the Promoter Divide and
Chapter 14 Regulation of Transcription
 Chapter 14 Regulation of Transcription Cis-acting sequences Distance-independent cis-acting elements Dissecting regulatory elements Transcription factors Overview transcriptional regulation Transcription
Chapter 14 Regulation of Transcription Cis-acting sequences Distance-independent cis-acting elements Dissecting regulatory elements Transcription factors Overview transcriptional regulation Transcription
CELL BIOLOGY - CLUTCH CH. 7 - GENE EXPRESSION.
 !! www.clutchprep.com CONCEPT: CONTROL OF GENE EXPRESSION BASICS Gene expression is the process through which cells selectively to express some genes and not others Every cell in an organism is a clone
!! www.clutchprep.com CONCEPT: CONTROL OF GENE EXPRESSION BASICS Gene expression is the process through which cells selectively to express some genes and not others Every cell in an organism is a clone
Differential Gene Expression
 Developmental Biology Biology 4361 Differential Gene Expression October 13, 2005 core transcription initiation site 5 promoter 3 TATAT +1 upstream downstream Basal transcription factors (eukaryotes) TFIID
Developmental Biology Biology 4361 Differential Gene Expression October 13, 2005 core transcription initiation site 5 promoter 3 TATAT +1 upstream downstream Basal transcription factors (eukaryotes) TFIID
CHAPTER 18 LECTURE NOTES: CONTROL OF GENE EXPRESSION PART B: CONTROL IN EUKARYOTES
 CHAPTER 18 LECTURE NOTES: CONTROL OF GENE EXPRESSION PART B: CONTROL IN EUKARYOTES I. Introduction A. No operon structures in eukaryotes B. Regulation of gene expression is frequently tissue specific.
CHAPTER 18 LECTURE NOTES: CONTROL OF GENE EXPRESSION PART B: CONTROL IN EUKARYOTES I. Introduction A. No operon structures in eukaryotes B. Regulation of gene expression is frequently tissue specific.
17.5 Eukaryotic Transcription Initiation Is Regulated by Transcription Factors That Bind to Cis-Acting Sites
 17.5 Eukaryotic Transcription Initiation Is Regulated by Transcription Factors That Bind to Cis-Acting Sites 1 Section 17.5 Transcription regulatory proteins, transcription factors, target cis-acting sites
17.5 Eukaryotic Transcription Initiation Is Regulated by Transcription Factors That Bind to Cis-Acting Sites 1 Section 17.5 Transcription regulatory proteins, transcription factors, target cis-acting sites
- all cells express large number of the same genes - housekeeping or common genes - many cells also express cell-type-specific genes
 Developmental Biology - Biology 4361 Lecture 8 - Differential Gene Expression October 13, 2005 The principle of genomic equivalence states that all cells in a developing organism have the same genetic
Developmental Biology - Biology 4361 Lecture 8 - Differential Gene Expression October 13, 2005 The principle of genomic equivalence states that all cells in a developing organism have the same genetic
Mechanism of action-1
 Mechanism of action-1 receptors: mediators of hormone action, membrane associated vs. intracellular receptors: measurements of receptor - ligand interaction, mechanism surface-receptors: kinases, phosphatases,
Mechanism of action-1 receptors: mediators of hormone action, membrane associated vs. intracellular receptors: measurements of receptor - ligand interaction, mechanism surface-receptors: kinases, phosphatases,
Gene Expression and Regulation - 1
 Gene Expression and Regulation - 1 We have been discussing the molecular structure of DNA and its function in DNA replication and in transcription. Earlier we discussed how genes interact in transmission
Gene Expression and Regulation - 1 We have been discussing the molecular structure of DNA and its function in DNA replication and in transcription. Earlier we discussed how genes interact in transmission
DNA & DNA : Protein Interactions BIBC 100
 DNA & DNA : Protein Interactions BIBC 100 Sequence = Information Alphabet = language L,I,F,E LIFE DNA = DNA code A, T, C, G CAC=Histidine CAG=Glutamine GGG=Glycine Protein = Protein code 20 a.a. LIVE EVIL
DNA & DNA : Protein Interactions BIBC 100 Sequence = Information Alphabet = language L,I,F,E LIFE DNA = DNA code A, T, C, G CAC=Histidine CAG=Glutamine GGG=Glycine Protein = Protein code 20 a.a. LIVE EVIL
Differential Gene Expression
 Biology 4361 Developmental Biology Differential Gene Expression September 28, 2006 Chromatin Structure ~140 bp ~60 bp Transcriptional Regulation: 1. Packing prevents access CH 3 2. Acetylation ( C O )
Biology 4361 Developmental Biology Differential Gene Expression September 28, 2006 Chromatin Structure ~140 bp ~60 bp Transcriptional Regulation: 1. Packing prevents access CH 3 2. Acetylation ( C O )
Chapter 8 DNA Recognition in Prokaryotes by Helix-Turn-Helix Motifs
 Chapter 8 DNA Recognition in Prokaryotes by Helix-Turn-Helix Motifs 1. Helix-turn-helix proteins 2. Zinc finger proteins 3. Leucine zipper proteins 4. Beta-scaffold factors 5. Others λ-repressor AND CRO
Chapter 8 DNA Recognition in Prokaryotes by Helix-Turn-Helix Motifs 1. Helix-turn-helix proteins 2. Zinc finger proteins 3. Leucine zipper proteins 4. Beta-scaffold factors 5. Others λ-repressor AND CRO
Resources. This lecture Campbell and Farrell's Biochemistry, Chapter 11
 Transcription Resources This lecture Campbell and Farrell's Biochemistry, Chapter 11 2 Definition of a gene The entire nucleic acid sequence that is necessary for the synthesis of a functional polypeptide
Transcription Resources This lecture Campbell and Farrell's Biochemistry, Chapter 11 2 Definition of a gene The entire nucleic acid sequence that is necessary for the synthesis of a functional polypeptide
Chapter 9-II - Transcriptional Control of Gene Expression
 Chapter 9-II - Transcriptional Control of Gene Expression Transcriptional Control of Gene Expression 9.3 RNA Polymerase II Promoters and General Transcription Factors Three types of promoter sequences
Chapter 9-II - Transcriptional Control of Gene Expression Transcriptional Control of Gene Expression 9.3 RNA Polymerase II Promoters and General Transcription Factors Three types of promoter sequences
Mechanism of action-1
 Mechanism of action-1 receptors: mediators of hormone action, membrane associated vs. intracellular receptors: measurements of receptor - ligand interactions, regulation mechanisms surface - receptors:
Mechanism of action-1 receptors: mediators of hormone action, membrane associated vs. intracellular receptors: measurements of receptor - ligand interactions, regulation mechanisms surface - receptors:
Chem 465 Biochemistry II
 Chem 465 Biochemistry II Name: 2 points Multiple choice (4 points apiece): 1. Which of the following is not true of trna molecules? A) The 3'-terminal sequence is -CCA. B) Their anticodons are complementary
Chem 465 Biochemistry II Name: 2 points Multiple choice (4 points apiece): 1. Which of the following is not true of trna molecules? A) The 3'-terminal sequence is -CCA. B) Their anticodons are complementary
Differential Gene Expression
 Biology 4361 Developmental Biology Differential Gene Expression June 19, 2008 Differential Gene Expression Overview Chromatin structure Gene anatomy RNA processing and protein production Initiating transcription:
Biology 4361 Developmental Biology Differential Gene Expression June 19, 2008 Differential Gene Expression Overview Chromatin structure Gene anatomy RNA processing and protein production Initiating transcription:
Control of Gene Expression
 Control of Gene Expression 1 How Gene Regulation Works 2 Control of Gene Expression Controlling gene expression is often accomplished by controlling transcription initiation Regulatory proteins bind to
Control of Gene Expression 1 How Gene Regulation Works 2 Control of Gene Expression Controlling gene expression is often accomplished by controlling transcription initiation Regulatory proteins bind to
M1 - Biochemistry. Nucleic Acid Structure II/Transcription I
 M1 - Biochemistry Nucleic Acid Structure II/Transcription I PH Ratz, PhD (Resources: Lehninger et al., 5th ed., Chapters 8, 24 & 26) 1 Nucleic Acid Structure II/Transcription I Learning Objectives: 1.
M1 - Biochemistry Nucleic Acid Structure II/Transcription I PH Ratz, PhD (Resources: Lehninger et al., 5th ed., Chapters 8, 24 & 26) 1 Nucleic Acid Structure II/Transcription I Learning Objectives: 1.
Eukaryotic Gene Expression John O. Thomas
 Eukaryotic Gene Expression John O. Thomas I) RNA polymerases A) There are four RNA polymerases in human cells. 1) RNA polymerase I, located in the nucleolar region of the nucleus, is responsible for the
Eukaryotic Gene Expression John O. Thomas I) RNA polymerases A) There are four RNA polymerases in human cells. 1) RNA polymerase I, located in the nucleolar region of the nucleus, is responsible for the
Transcription in Prokaryotes. Jörg Bungert, PhD Phone:
 Transcription in Prokaryotes Jörg Bungert, PhD Phone: 352-273-8098 Email: jbungert@ufl.edu Objectives Understand the basic mechanism of transcription. Know the function of promoter elements and associating
Transcription in Prokaryotes Jörg Bungert, PhD Phone: 352-273-8098 Email: jbungert@ufl.edu Objectives Understand the basic mechanism of transcription. Know the function of promoter elements and associating
Transcription Regulation And Gene Expression in Eukaryotes FS 2016 Graduate Course G2 P Matthias and RG Clerc Pharmazentrum Hörsaal 2 16h15-18h00
 Transcription Regulation And Gene Expression in Eukaryotes FS 2016 Graduate Course G2 P Matthias and RG Clerc Pharmazentrum Hörsaal 2 16h15-18h00 MODULAR STRUCTURE OF UPSTREAM TRANSCRIPTION FACTORS Experimental
Transcription Regulation And Gene Expression in Eukaryotes FS 2016 Graduate Course G2 P Matthias and RG Clerc Pharmazentrum Hörsaal 2 16h15-18h00 MODULAR STRUCTURE OF UPSTREAM TRANSCRIPTION FACTORS Experimental
Transcription factors
 Transcription factors R. de Martin Department of Vascular Biology and Thrombosis Research Medical University of Vienna K63 A20, TTP, XIAP IκBα K48 IL-6, Il-8, MCP-1, ICAM1, SELE, ciap1/2, A20, IκBα Adapted
Transcription factors R. de Martin Department of Vascular Biology and Thrombosis Research Medical University of Vienna K63 A20, TTP, XIAP IκBα K48 IL-6, Il-8, MCP-1, ICAM1, SELE, ciap1/2, A20, IκBα Adapted
Eukaryotic & Prokaryotic Transcription. RNA polymerases
 Eukaryotic & Prokaryotic Transcription RNA polymerases RNA Polymerases A. E. coli RNA polymerase 1. core enzyme = ββ'(α)2 has catalytic activity but cannot recognize start site of transcription ~500,000
Eukaryotic & Prokaryotic Transcription RNA polymerases RNA Polymerases A. E. coli RNA polymerase 1. core enzyme = ββ'(α)2 has catalytic activity but cannot recognize start site of transcription ~500,000
Part IV => DNA and RNA. 4.7 Gene Regulation 4.7a Chromatin Remodeling 4.7b Transcriptional Control
 Part IV => DNA and RNA 4.7 Gene Regulation 4.7a Chromatin Remodeling 4.7b Transcriptional Control Overview of Gene regulation - The expression of the so-called housekeeping genes (eg basal transcription
Part IV => DNA and RNA 4.7 Gene Regulation 4.7a Chromatin Remodeling 4.7b Transcriptional Control Overview of Gene regulation - The expression of the so-called housekeeping genes (eg basal transcription
Eukaryotic Transcription
 Eukaryotic Transcription I. Differences between eukaryotic versus prokaryotic transcription. II. (core vs holoenzyme): RNA polymerase II - Promotor elements. - General Pol II transcription factors (GTF).
Eukaryotic Transcription I. Differences between eukaryotic versus prokaryotic transcription. II. (core vs holoenzyme): RNA polymerase II - Promotor elements. - General Pol II transcription factors (GTF).
Section C: The Control of Gene Expression
 Section C: The Control of Gene Expression 1. Each cell of a multicellular eukaryote expresses only a small fraction of its genes 2. The control of gene expression can occur at any step in the pathway from
Section C: The Control of Gene Expression 1. Each cell of a multicellular eukaryote expresses only a small fraction of its genes 2. The control of gene expression can occur at any step in the pathway from
Gene Regulation in Eukaryotes
 Gene Regulation in Eukaryotes The latest estimates are that a human cell, a eukaryotic cell, contains 20,000 25,000 genes. Some of these are expressed in all cells all the time. These so-called housekeeping
Gene Regulation in Eukaryotes The latest estimates are that a human cell, a eukaryotic cell, contains 20,000 25,000 genes. Some of these are expressed in all cells all the time. These so-called housekeeping
30 Gene expression: Transcription
 30 Gene expression: Transcription Gene structure. o Exons coding region of DNA. o Introns non-coding region of DNA. o Introns are interspersed between exons of a single gene. o Promoter region helps enzymes
30 Gene expression: Transcription Gene structure. o Exons coding region of DNA. o Introns non-coding region of DNA. o Introns are interspersed between exons of a single gene. o Promoter region helps enzymes
Chapter 18: Regulation of Gene Expression. 1. Gene Regulation in Bacteria 2. Gene Regulation in Eukaryotes 3. Gene Regulation & Cancer
 Chapter 18: Regulation of Gene Expression 1. Gene Regulation in Bacteria 2. Gene Regulation in Eukaryotes 3. Gene Regulation & Cancer Gene Regulation Gene regulation refers to all aspects of controlling
Chapter 18: Regulation of Gene Expression 1. Gene Regulation in Bacteria 2. Gene Regulation in Eukaryotes 3. Gene Regulation & Cancer Gene Regulation Gene regulation refers to all aspects of controlling
Computational Biology I LSM5191
 Computational Biology I LSM5191 Aylwin Ng, D.Phil Lecture 4 Notes: Gene Regulation & Control Do all cells in an individual have the same DNA content? PART I: CONTROL OF GENE EXPRESSION REGULATION OF GENE
Computational Biology I LSM5191 Aylwin Ng, D.Phil Lecture 4 Notes: Gene Regulation & Control Do all cells in an individual have the same DNA content? PART I: CONTROL OF GENE EXPRESSION REGULATION OF GENE
Spring 2006 Biochemistry 302 Exam 2
 Name Spring 2006 Biochemistry 302 Exam 2 Directions: This exam has 45 questions/problems totaling 110 points. Check to make sure you have seven pages. Some questions have multiple parts so read each one
Name Spring 2006 Biochemistry 302 Exam 2 Directions: This exam has 45 questions/problems totaling 110 points. Check to make sure you have seven pages. Some questions have multiple parts so read each one
7.06 Problem Set #3, 2006
 7.06 Problem Set #3, 2006 1. You are studying the EGF/Ras/MAPK pathway in cultured cells. When the pathway is activated, cells are signaled to proliferate. You generate various mutants described below.
7.06 Problem Set #3, 2006 1. You are studying the EGF/Ras/MAPK pathway in cultured cells. When the pathway is activated, cells are signaled to proliferate. You generate various mutants described below.
Protein Synthesis Notes
 Protein Synthesis Notes Protein Synthesis: Overview Transcription: synthesis of mrna under the direction of DNA. Translation: actual synthesis of a polypeptide under the direction of mrna. Transcription
Protein Synthesis Notes Protein Synthesis: Overview Transcription: synthesis of mrna under the direction of DNA. Translation: actual synthesis of a polypeptide under the direction of mrna. Transcription
GENE EXPRESSSION. Promoter sequence where RNA polymerase binds. Operator sequence that acts as a switch (yellow) OPERON
 GENE EXPRESSSION 1 GENE REGULATION IN PROKARYOTES Bacteria can turn genes on or off depending on their environment Prokaryotes have operons clusters of related genes and regulatory sequences Promoter sequence
GENE EXPRESSSION 1 GENE REGULATION IN PROKARYOTES Bacteria can turn genes on or off depending on their environment Prokaryotes have operons clusters of related genes and regulatory sequences Promoter sequence
Computational Biology I LSM5191 (2003/4)
 Computational Biology I LSM5191 (2003/4) Aylwin Ng, D.Phil Lecture Notes: Transcriptome: Molecular Biology of Gene Expression I Flow of information: DNA to polypeptide DNA Start Exon1 Intron Exon2 Termination
Computational Biology I LSM5191 (2003/4) Aylwin Ng, D.Phil Lecture Notes: Transcriptome: Molecular Biology of Gene Expression I Flow of information: DNA to polypeptide DNA Start Exon1 Intron Exon2 Termination
BIOLOGY. Chapter 16 GenesExpression
 BIOLOGY Chapter 16 GenesExpression CAMPBELL BIOLOGY TENTH EDITION Reece Urry Cain Wasserman Minorsky Jackson 18 Gene Expression 2014 Pearson Education, Inc. Figure 16.1 Differential Gene Expression results
BIOLOGY Chapter 16 GenesExpression CAMPBELL BIOLOGY TENTH EDITION Reece Urry Cain Wasserman Minorsky Jackson 18 Gene Expression 2014 Pearson Education, Inc. Figure 16.1 Differential Gene Expression results
Chapter 17 Lecture. Concepts of Genetics. Tenth Edition. Regulation of Gene Expression in Eukaryotes
 Chapter 17 Lecture Concepts of Genetics Tenth Edition Regulation of Gene Expression in Eukaryotes Chapter Contents 17.1 Eukaryotic Gene Regulation Can Occur at Any of the Steps Leading from DNA to Protein
Chapter 17 Lecture Concepts of Genetics Tenth Edition Regulation of Gene Expression in Eukaryotes Chapter Contents 17.1 Eukaryotic Gene Regulation Can Occur at Any of the Steps Leading from DNA to Protein
Contact information Blaine Bartholomew Office: Neckers Bldg., Rm. 211 Phone:
 Contact information Blaine Bartholomew Office: Neckers Bldg., Rm. 211 Phone: 453-6437 Email: bbartholomew@siumed.edu Class notes http://www.siumed.edu/~bbartholomew/hbtxn.html Other references Principles
Contact information Blaine Bartholomew Office: Neckers Bldg., Rm. 211 Phone: 453-6437 Email: bbartholomew@siumed.edu Class notes http://www.siumed.edu/~bbartholomew/hbtxn.html Other references Principles
Hmwk # 8 : DNA-Binding Proteins : Part II
 The purpose of this exercise is : Hmwk # 8 : DNA-Binding Proteins : Part II 1). to examine the case of a tandem head-to-tail homodimer binding to DNA 2). to view a Zn finger motif 3). to consider the case
The purpose of this exercise is : Hmwk # 8 : DNA-Binding Proteins : Part II 1). to examine the case of a tandem head-to-tail homodimer binding to DNA 2). to view a Zn finger motif 3). to consider the case
DNA Transcription. Dr Aliwaini
 DNA Transcription 1 DNA Transcription-Introduction The synthesis of an RNA molecule from DNA is called Transcription. All eukaryotic cells have five major classes of RNA: ribosomal RNA (rrna), messenger
DNA Transcription 1 DNA Transcription-Introduction The synthesis of an RNA molecule from DNA is called Transcription. All eukaryotic cells have five major classes of RNA: ribosomal RNA (rrna), messenger
Control of Eukaryotic Genes. AP Biology
 Control of Eukaryotic Genes The BIG Questions How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different, specialized functions? Evolution
Control of Eukaryotic Genes The BIG Questions How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different, specialized functions? Evolution
The Little Things About the Little Things Inside of Us The Eukaryotic Genome and Its Expression
 The Little Things About the Little Things Inside of Us The Eukaryotic Genome and Its Expression What Are the Characteristics of the Eukaryotic Genome? Key differences between eukaryotic and prokaryotic
The Little Things About the Little Things Inside of Us The Eukaryotic Genome and Its Expression What Are the Characteristics of the Eukaryotic Genome? Key differences between eukaryotic and prokaryotic
Chromatographic Separation of the three forms of RNA Polymerase II.
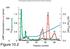 Chromatographic Separation of the three forms of RNA Polymerase II. α-amanitin α-amanitin bound to Pol II Function of the three enzymes. Yeast Pol II. RNA Polymerase Subunit Structures 10-7 Subunit structure.
Chromatographic Separation of the three forms of RNA Polymerase II. α-amanitin α-amanitin bound to Pol II Function of the three enzymes. Yeast Pol II. RNA Polymerase Subunit Structures 10-7 Subunit structure.
Gene Circuits -2. Shu-Ping Lin, Ph.D. Institute of Biomedical Engineering
 Gene Circuits -2 Shu-Ping Lin, Ph.D. Institute of Biomedical Engineering E-mail: splin@dragon.nchu.edu.tw Website: http://web.nchu.edu.tw/pweb/users/splin/ Genes Flow of information in gene expression
Gene Circuits -2 Shu-Ping Lin, Ph.D. Institute of Biomedical Engineering E-mail: splin@dragon.nchu.edu.tw Website: http://web.nchu.edu.tw/pweb/users/splin/ Genes Flow of information in gene expression
Differential Gene Expression
 Biology 4361 - Developmental Biology Differential Gene Expression June 18, 2009 Differential Gene Expression Overview Chromatin structure Gene anatomy RNA processing and protein production Initiating transcription:
Biology 4361 - Developmental Biology Differential Gene Expression June 18, 2009 Differential Gene Expression Overview Chromatin structure Gene anatomy RNA processing and protein production Initiating transcription:
Transcription. Manzur Ali PP, DBT,M.E.S College,Marampally
 Transcription Manzur Ali PP, DBT,M.E.S College,Marampally manzursir@gmail.com RNA transcription is actively regulated Not all DNA is transcribed in a given cell (less than 50% even in prokaryotes) For
Transcription Manzur Ali PP, DBT,M.E.S College,Marampally manzursir@gmail.com RNA transcription is actively regulated Not all DNA is transcribed in a given cell (less than 50% even in prokaryotes) For
GENETICS - CLUTCH CH.10 TRANSCRIPTION.
 !! www.clutchprep.com CONCEPT: OVERVIEW OF TRANSCRIPTION Transcription is the process of using DNA as a template to RNA RNA polymerase is the enzyme that transcribes DNA - There are many different types
!! www.clutchprep.com CONCEPT: OVERVIEW OF TRANSCRIPTION Transcription is the process of using DNA as a template to RNA RNA polymerase is the enzyme that transcribes DNA - There are many different types
Chapter 24: Promoters and Enhancers
 Chapter 24: Promoters and Enhancers A typical gene transcribed by RNA polymerase II has a promoter that usually extends upstream from the site where transcription is initiated the (#1) of transcription
Chapter 24: Promoters and Enhancers A typical gene transcribed by RNA polymerase II has a promoter that usually extends upstream from the site where transcription is initiated the (#1) of transcription
The Stringent Response
 The Stringent Response When amino acids are limiting a response is triggered to shut down a wide range of biosynthetic processes This process is called the Stringent Response It results in the synthesis
The Stringent Response When amino acids are limiting a response is triggered to shut down a wide range of biosynthetic processes This process is called the Stringent Response It results in the synthesis
Plant Molecular and Cellular Biology Lecture 9: Nuclear Genome Organization: Chromosome Structure, Chromatin, DNA Packaging, Mitosis Gary Peter
 Plant Molecular and Cellular Biology Lecture 9: Nuclear Genome Organization: Chromosome Structure, Chromatin, DNA Packaging, Mitosis Gary Peter 9/16/2008 1 Learning Objectives 1. List and explain how DNA
Plant Molecular and Cellular Biology Lecture 9: Nuclear Genome Organization: Chromosome Structure, Chromatin, DNA Packaging, Mitosis Gary Peter 9/16/2008 1 Learning Objectives 1. List and explain how DNA
M I C R O B I O L O G Y WITH DISEASES BY TAXONOMY, THIRD EDITION
 M I C R O B I O L O G Y WITH DISEASES BY TAXONOMY, THIRD EDITION Chapter 7 Microbial Genetics Lecture prepared by Mindy Miller-Kittrell, University of Tennessee, Knoxville The Structure and Replication
M I C R O B I O L O G Y WITH DISEASES BY TAXONOMY, THIRD EDITION Chapter 7 Microbial Genetics Lecture prepared by Mindy Miller-Kittrell, University of Tennessee, Knoxville The Structure and Replication
The RNA Polymerase II General Transcription Machinery Prof. Michael Hampsey
 The RNA Polymerase II General Transcription Machinery Michael Hampsey, PhD Robert Wood Johnson Medical School Piscataway, New Jersey 1 Central Dogma of Molecular Biology DNA Stores genetic information
The RNA Polymerase II General Transcription Machinery Michael Hampsey, PhD Robert Wood Johnson Medical School Piscataway, New Jersey 1 Central Dogma of Molecular Biology DNA Stores genetic information
Mechanisms of Transcription. School of Life Science Shandong University
 Mechanisms of Transcription School of Life Science Shandong University Ch 12: Mechanisms of Transcription 1. RNA polymerase and the transcription cycle 2. The transcription cycle in bacteria 3. Transcription
Mechanisms of Transcription School of Life Science Shandong University Ch 12: Mechanisms of Transcription 1. RNA polymerase and the transcription cycle 2. The transcription cycle in bacteria 3. Transcription
Fig Ch 17: From Gene to Protein
 Fig. 17-1 Ch 17: From Gene to Protein Basic Principles of Transcription and Translation RNA is the intermediate between genes and the proteins for which they code Transcription is the synthesis of RNA
Fig. 17-1 Ch 17: From Gene to Protein Basic Principles of Transcription and Translation RNA is the intermediate between genes and the proteins for which they code Transcription is the synthesis of RNA
Control of Eukaryotic Genes
 Control of Eukaryotic Genes 2007-2008 The BIG Questions How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different, specialized functions?
Control of Eukaryotic Genes 2007-2008 The BIG Questions How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different, specialized functions?
However, only a fraction of these genes are transcribed in an individual cell at any given time.
 All cells in an organism contain the same set of genes. However, only a fraction of these genes are transcribed in an individual cell at any given time. It is the pattern of gene expression that determines
All cells in an organism contain the same set of genes. However, only a fraction of these genes are transcribed in an individual cell at any given time. It is the pattern of gene expression that determines
GENES AND CHROMOSOMES V. Lecture 7. Biology Department Concordia University. Dr. S. Azam BIOL 266/
 1 GENES AND CHROMOSOMES V Lecture 7 BIOL 266/4 2014-15 Dr. S. Azam Biology Department Concordia University 2 CELL NUCLEUS AND THE CONTROL OF GENE EXPRESSION An Overview of Gene Regulation in Eukaryotes
1 GENES AND CHROMOSOMES V Lecture 7 BIOL 266/4 2014-15 Dr. S. Azam Biology Department Concordia University 2 CELL NUCLEUS AND THE CONTROL OF GENE EXPRESSION An Overview of Gene Regulation in Eukaryotes
RNA does not adopt the classic B-DNA helix conformation when it forms a self-complementary double helix
 Reason: RNA has ribose sugar ring, with a hydroxyl group (OH) If RNA in B-from conformation there would be unfavorable steric contact between the hydroxyl group, base, and phosphate backbone. RNA structure
Reason: RNA has ribose sugar ring, with a hydroxyl group (OH) If RNA in B-from conformation there would be unfavorable steric contact between the hydroxyl group, base, and phosphate backbone. RNA structure
Control of Eukaryotic Genes. AP Biology
 Control of Eukaryotic Genes The BIG Questions How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different, specialized functions? Evolution
Control of Eukaryotic Genes The BIG Questions How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different, specialized functions? Evolution
Unit 7. Genetic Regulation, Development, and Biotechnology. AP Biology
 Unit 7 Genetic Regulation, Development, and Biotechnology The BIG Questions How are genes turned on & off in eukaryotes and prokaryotes? How do cells with the same genes differentiate to perform completely
Unit 7 Genetic Regulation, Development, and Biotechnology The BIG Questions How are genes turned on & off in eukaryotes and prokaryotes? How do cells with the same genes differentiate to perform completely
Biol 3301 Genetics Exam #2A October 26, 2004
 Biol 3301 Genetics Exam #2A October 26, 2004 This exam consists of 40 multiple choice questions worth 2.5 points each, for a total of 100 points. Good luck. Name SS# 1. Which of the following statements
Biol 3301 Genetics Exam #2A October 26, 2004 This exam consists of 40 multiple choice questions worth 2.5 points each, for a total of 100 points. Good luck. Name SS# 1. Which of the following statements
Transcription is the first stage of gene expression
 Transcription is the first stage of gene expression RNA synthesis is catalyzed by RNA polymerase, which pries the DNA strands apart and hooks together the RNA nucleotides The RNA is complementary to the
Transcription is the first stage of gene expression RNA synthesis is catalyzed by RNA polymerase, which pries the DNA strands apart and hooks together the RNA nucleotides The RNA is complementary to the
Translation Mechanisms
 Translation Mechanisms Biology I Hayder A. Giha Translation The translation is the process of protein synthesis, where information in nucleotides sequences of a mrna is translated into amino acids sequence
Translation Mechanisms Biology I Hayder A. Giha Translation The translation is the process of protein synthesis, where information in nucleotides sequences of a mrna is translated into amino acids sequence
SIBC504: TRANSCRIPTION & RNA PROCESSING Assistant Professor Dr. Chatchawan Srisawat
 SIBC504: TRANSCRIPTION & RNA PROCESSING Assistant Professor Dr. Chatchawan Srisawat TRANSCRIPTION: AN OVERVIEW Transcription: the synthesis of a single-stranded RNA from a doublestranded DNA template.
SIBC504: TRANSCRIPTION & RNA PROCESSING Assistant Professor Dr. Chatchawan Srisawat TRANSCRIPTION: AN OVERVIEW Transcription: the synthesis of a single-stranded RNA from a doublestranded DNA template.
BCH 4054 Fall 2000 Chapter 31 Lecture Notes
 BCH 4054 Fall 2000 Chapter 31 Lecture Notes 1 Chapter 31 Transcription and Regulation of Gene Expression 2 Messenger RNA Central Dogma (Francis Crick, 1958) DNA RNA Protein (Fig 31.1) Jacob-Monod Hypothesis:
BCH 4054 Fall 2000 Chapter 31 Lecture Notes 1 Chapter 31 Transcription and Regulation of Gene Expression 2 Messenger RNA Central Dogma (Francis Crick, 1958) DNA RNA Protein (Fig 31.1) Jacob-Monod Hypothesis:
PART FOUR - ANSWERS PART FOUR: GENE REGULATION ANSWERS. Answers to questions from Chapter 15 on Positive and negative control of the lac operon
 PART FOUR: GENE REGULATION ANSWERS Answers to questions from Chapter 15 on Positive and negative control of the lac operon 15.1 laci, lacz, lacy, laca 15.2 The lac operon is negatively regulated by a repressor,
PART FOUR: GENE REGULATION ANSWERS Answers to questions from Chapter 15 on Positive and negative control of the lac operon 15.1 laci, lacz, lacy, laca 15.2 The lac operon is negatively regulated by a repressor,
Einführung in die Genetik
 Einführung in die Genetik Prof. Dr. Kay Schneitz (EBio Pflanzen) http://plantdev.bio.wzw.tum.de schneitz@wzw.tum.de Prof. Dr. Claus Schwechheimer (PlaSysBiol) http://wzw.tum.de/sysbiol claus.schwechheimer@wzw.tum.de
Einführung in die Genetik Prof. Dr. Kay Schneitz (EBio Pflanzen) http://plantdev.bio.wzw.tum.de schneitz@wzw.tum.de Prof. Dr. Claus Schwechheimer (PlaSysBiol) http://wzw.tum.de/sysbiol claus.schwechheimer@wzw.tum.de
Chapter 8 Lecture Outline. Transcription, Translation, and Bioinformatics
 Chapter 8 Lecture Outline Transcription, Translation, and Bioinformatics Replication, Transcription, Translation n Repetitive processes Build polymers of nucleotides or amino acids n All have 3 major steps
Chapter 8 Lecture Outline Transcription, Translation, and Bioinformatics Replication, Transcription, Translation n Repetitive processes Build polymers of nucleotides or amino acids n All have 3 major steps
Translation. Protein Synthesis
 Protein Structure Translation Protein Synthesis Size and Shape Comparison of Proteins Levels of Protein Structure 1 o 2 o 3 o 4 o Amino Acids Peptide Bonds Proteins are formed by creating peptide bonds
Protein Structure Translation Protein Synthesis Size and Shape Comparison of Proteins Levels of Protein Structure 1 o 2 o 3 o 4 o Amino Acids Peptide Bonds Proteins are formed by creating peptide bonds
Cell Nucleus. Chen Li. Department of Cellular and Genetic Medicine
 Cell Nucleus Chen Li Department of Cellular and Genetic Medicine 13 223 chenli2008@fudan.edu.cn Outline A. Historical background B. Structure of the nucleus: nuclear pore complex (NPC), lamina, nucleolus,
Cell Nucleus Chen Li Department of Cellular and Genetic Medicine 13 223 chenli2008@fudan.edu.cn Outline A. Historical background B. Structure of the nucleus: nuclear pore complex (NPC), lamina, nucleolus,
TRANSCRIPTION AND PROCESSING OF RNA
 TRANSCRIPTION AND PROCESSING OF RNA 1. The steps of gene expression. 2. General characterization of transcription: steps, components of transcription apparatus. 3. Transcription of eukaryotic structural
TRANSCRIPTION AND PROCESSING OF RNA 1. The steps of gene expression. 2. General characterization of transcription: steps, components of transcription apparatus. 3. Transcription of eukaryotic structural
Transcription in Eukaryotes
 Transcription in Eukaryotes Biology I Hayder A Giha Transcription Transcription is a DNA-directed synthesis of RNA, which is the first step in gene expression. Gene expression, is transformation of the
Transcription in Eukaryotes Biology I Hayder A Giha Transcription Transcription is a DNA-directed synthesis of RNA, which is the first step in gene expression. Gene expression, is transformation of the
of signal transduction pathways?
 CELL BIOLOGY Signal Transduction + How do you study activation of signal transduction pathways? + IDENTIFICATION OF PATHWAY-SPECIFIC TRANSCRIPTION ACTIVATION + EASIER, FASTER THAN BLOTTING OR GEL-SHIFT
CELL BIOLOGY Signal Transduction + How do you study activation of signal transduction pathways? + IDENTIFICATION OF PATHWAY-SPECIFIC TRANSCRIPTION ACTIVATION + EASIER, FASTER THAN BLOTTING OR GEL-SHIFT
Transcription Eukaryotic Cells
 Transcription Eukaryotic Cells Packet #20 1 Introduction Transcription is the process in which genetic information, stored in a strand of DNA (gene), is copied into a strand of RNA. Protein-encoding genes
Transcription Eukaryotic Cells Packet #20 1 Introduction Transcription is the process in which genetic information, stored in a strand of DNA (gene), is copied into a strand of RNA. Protein-encoding genes
GENE REGULATION slide shows by Kim Foglia modified Slides with blue edges are Kim s
 GENE REGULATION slide shows by Kim Foglia modified Slides with blue edges are Kim s 2007-2008 Bacterial metabolism Bacteria need to respond quickly to changes in their environment STOP GO if they have
GENE REGULATION slide shows by Kim Foglia modified Slides with blue edges are Kim s 2007-2008 Bacterial metabolism Bacteria need to respond quickly to changes in their environment STOP GO if they have
Division Ave. High School AP Biology
 Control of Eukaryotic Genes 2007-2008 The BIG Questions n How are genes turned on & off in eukaryotes? n How do cells with the same genes differentiate to perform completely different, specialized functions?
Control of Eukaryotic Genes 2007-2008 The BIG Questions n How are genes turned on & off in eukaryotes? n How do cells with the same genes differentiate to perform completely different, specialized functions?
Chromatin and Transcription
 Chromatin and Transcription Chromatin Structure Chromatin Represses Transcription Nucleosome Positioning Histone Acetylation Chromatin Remodeling Histone Methylation CHIP Analysis Chromatin and Elongation
Chromatin and Transcription Chromatin Structure Chromatin Represses Transcription Nucleosome Positioning Histone Acetylation Chromatin Remodeling Histone Methylation CHIP Analysis Chromatin and Elongation
Bi 8 Lecture 7. Ellen Rothenberg 26 January Reading: Ch. 3, pp ; panel 3-1
 Bi 8 Lecture 7 PROTEIN STRUCTURE, Functional analysis, and evolution Ellen Rothenberg 26 January 2016 Reading: Ch. 3, pp. 109-134; panel 3-1 (end with free amine) aromatic, hydrophobic small, hydrophilic
Bi 8 Lecture 7 PROTEIN STRUCTURE, Functional analysis, and evolution Ellen Rothenberg 26 January 2016 Reading: Ch. 3, pp. 109-134; panel 3-1 (end with free amine) aromatic, hydrophobic small, hydrophilic
Chromatin. Structure and modification of chromatin. Chromatin domains
 Chromatin Structure and modification of chromatin Chromatin domains 2 DNA consensus 5 3 3 DNA DNA 4 RNA 5 ss RNA forms secondary structures with ds hairpins ds forms 6 of nucleic acids Form coiling bp/turn
Chromatin Structure and modification of chromatin Chromatin domains 2 DNA consensus 5 3 3 DNA DNA 4 RNA 5 ss RNA forms secondary structures with ds hairpins ds forms 6 of nucleic acids Form coiling bp/turn
Protein Structure. Protein Structure Tertiary & Quaternary
 Lecture 4 Protein Structure Protein Structure Tertiary & Quaternary Dr. Sameh Sarray Hlaoui Primary structure: The linear sequence of amino acids held together by peptide bonds. Secondary structure: The
Lecture 4 Protein Structure Protein Structure Tertiary & Quaternary Dr. Sameh Sarray Hlaoui Primary structure: The linear sequence of amino acids held together by peptide bonds. Secondary structure: The
Gene Regulation in Eukaryotes. Dr. Syahril Abdullah Medical Genetics Laboratory
 Gene Regulation in Eukaryotes Dr. Syahril Abdullah Medical Genetics Laboratory syahril@medic.upm.edu.my Lecture Outline 1. The Genome 2. Overview of Gene Control 3. Cellular Differentiation in Higher Eukaryotes
Gene Regulation in Eukaryotes Dr. Syahril Abdullah Medical Genetics Laboratory syahril@medic.upm.edu.my Lecture Outline 1. The Genome 2. Overview of Gene Control 3. Cellular Differentiation in Higher Eukaryotes
Post-translational modification
 Protein expression Western blotting, is a widely used and accepted technique to detect levels of protein expression in a cell or tissue extract. This technique measures protein levels in a biological sample
Protein expression Western blotting, is a widely used and accepted technique to detect levels of protein expression in a cell or tissue extract. This technique measures protein levels in a biological sample
Einführung in die Genetik
 Einführung in die Genetik Prof. Dr. Kay Schneitz (EBio Pflanzen) http://plantdev.bio.wzw.tum.de schneitz@wzw.tum.de Twitter: @PlantDevTUM, #genetiktum FB: Plant Development TUM Prof. Dr. Claus Schwechheimer
Einführung in die Genetik Prof. Dr. Kay Schneitz (EBio Pflanzen) http://plantdev.bio.wzw.tum.de schneitz@wzw.tum.de Twitter: @PlantDevTUM, #genetiktum FB: Plant Development TUM Prof. Dr. Claus Schwechheimer
Biochemistry 674, Fall, 1995: Nucleic Acids Exam II: November 16, 1995 Your Name Here: PCR A C
 CHM 674 Exam II, 1995 1 iochemistry 674, Fall, 1995: Nucleic cids Prof. Jason Kahn Exam II: November 16, 1995 Your Name Here: This exam has five questions worth 20 points each. nswer all five. You do not
CHM 674 Exam II, 1995 1 iochemistry 674, Fall, 1995: Nucleic cids Prof. Jason Kahn Exam II: November 16, 1995 Your Name Here: This exam has five questions worth 20 points each. nswer all five. You do not
Control of Eukaryotic Genes. AP Biology
 Control of Eukaryotic Genes The BIG Questions How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different, specialized functions? Evolution
Control of Eukaryotic Genes The BIG Questions How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different, specialized functions? Evolution
