Protocol for tissue-specific gene disruption in zebrafish
|
|
|
- Elwin Adrian Thomas
- 6 years ago
- Views:
Transcription
1 Protocol for tissue-specific gene disruption in zebrafish Overview This protocol describes a method to inactivate genes in zebrafish in a tissue-specific manner. It can be used to analyze mosaic loss-of-function phenotypes in F0 embryos or to generate stable tissue-specific knock-out lines. It takes advantage of the Tol2 transposase technology to integrate in the fish genome a vector expressing a guide RNA (grna) from a ubiquitous zebrafish U6 promoter and Cas9 under the control of a tissue-specific promoter. This protocol comprises 5 steps: 1) the identification of efficient CRISPR target sequences in the gene of interest; 2) the annealing of genespecific oligonucleotides; 3) the construction of the tissue-specific CRISPR vector; 4) the injection of the CRISPR construct in zebrafish embryos; 5) the phenotypic analysis and generation of stable lines (Fig. P1). Identification of efficient CRISPR target sequences in the gene of interest (Fig. P2) Annealing of gene-specific oligonucleotides Construction of the tissuespecific CRISPR vector (2 options, see Fig. P3) U6:gRNA Prom:Cas9 Injection of the CRISPR construct in zebrafish embryos Phenotype analysis Generation of stable line Figure P1 Workflow of tissue-specific gene disruption
2 Reagents - Destination vector for Gateway (e.g. pdesttol2 cmlc2:gfp, U6:gRNA or pdesttol2pa2 U6:gRNA, available through Addgene) - Middle entry vector for Gateway containing Cas9 (e.g. pme-cas9 or pme- Cas9-T2A-GFP, available through Addgene) - 5 entry vector for Gateway containing the tissue-specific promoter of interest - 3 entry vector for Gateway containing a polya sequence (p3e-polya, vector #302 from the Tol2Kit) - Gene-specific oligonucleotides (see below) - BseRI enzyme (New England Biolabs) - Gateway LR clonase II (Invitrogen) - T4 DNA ligase (New England Biolabs) - Gel extraction kit or PCR-purification kit (Qiagen) Procedure 1. Identification of efficient CRISPR target sequences in the gene(s) of interest Pick several (3 to 6) CRISPR target sequences in each gene of interest using available tools (Hsu et al., 2013; Montague et al., 2014) (Fig. P2). Produce the grnas in vitro as per usual procedures (Gagnon et al., 2014; Hwang et al., 2013). Inject the grnas along with Cas9 protein or mrna into one-cell stage embryos of a WT strain. Extract DNA from injected embryos at 24 or 48 hpf by the HotSHOT method (Meeker et al., 2007) and assess mutation rates at target loci by sequencing or enzymatic assays (e.g. T7E1 assay, Surveyor assay) (Gagnon et al., 2014; Kim et al., 2009). Proceed with the target sequences that have shown effective targeting (we usually only keep target sequences for which the mutation rates exceed 10%). Note: alternatively, it is possible to test the efficiency of target sequences directly in the context of the Tol2 vector by cloning various target sequences in the U6:gRNA cassette of a vector expressing Cas9 under the control of a ubiquitous promoter.
3 Pick CRISPR target sequences in gene of interest (3-6 per gene) Produce grnas in vitro Inject grnas with Cas9 mrna or protein in zebrafish embryos grna Cas9 Evaluate targeting at the CRISPR site by sequencing or mutagenesis assays GAATCACGGGCAGGGAAAGAT T GAATCAC GGGAAAGATT GAA AGATT GAATCACGG AAAGATT GAATCA GGGAAAGATT GAATCAC AGGGAAAGATT GAATCACGGG GAAAGATT GAATCACGGGC GAATCACGGG Identify efficient target sequences Figure P2 Identification of efficient CRISPR target sequences 2. Annealing of gene-specific oligonucleotides Order 22-mer, unmodified oligonucleotides as follows: Forward: target sequence (20 bases)-gt Reverse: reverse complement of target sequence-ga Note: the target sequence must start with a G. If not, replace the first base by a G. This will introduce a mismatch but most mismatches at the 5 end of the target sequence are well tolerated. Example: for CRISPR target sequence GGTGGGAGAGTGGATGGCTG, order GGTGGGAGAGTGGATGGCTGGT (forward) and CAGCCATCCACTCTCCCACCGA (reverse). Anneal the two oligos in thermocycler (5 min. at 95 C, -1 C/min. down to 20 C).
4 3. Construction of the tissue-specific CRISPR vector There are two options to create a tissue-specific CRISPR vector. The order in which you perform the cloning steps depends on the option you choose. Option 1: Clone the gene-specific seed sequence into the destination vector and then perform the Gateway reaction with the tissue-specific promoter and Cas9 (Fig. P3). This option will be used preferentially to inactivate the same gene or a few genes in several different tissues (e.g. using various tissue-specific promoters with few different grnas) or if the tissue-specific promoter of interest contains one or several BseRI restriction sites. Option 2: Perform the Gateway reaction first to assemble a vector for tissue-specific expression of Cas9 and then clone the gene-specific seed sequence into the U6:gRNA cassette (Fig. P4). This option is better suited for targeting many different genes in the same tissue (e.g. using a single tissue-specific promoter with many different grnas). If you wish to use option 2 with a tissue-specific promoter containing one or several BseRI sites, you may consider modifying them by site-directed mutagenesis on the 5 entry vector for Gateway.
5 grna grna U6:gRNA BseRI digestion seed ligation U6:gRNA gene + 5 Entry Promoter U6:gRNA gene Prom:Cas9 Gateway reaction Middle Entry Cas9 3 Entry PolyA Figure P3 Construction of a tissue-specific CRISPR vector, option 1 Option 1 - Digest the destination vector of your choice with BseRI enzyme. - Purify on 1% agarose gel or column. - Ligate the gene-specific seed sequence into BseRI-digested destination vector (1:20 vector:insert molar ratio). - Transform chemically competent ccdb-survival bacteria, plate on LB Agar and select with ampicillin + chloramphenicol. - Extract plasmid DNA from 2-3 distinct colonies and sequence using the following primer: CCTCACACAAACTCTGGATT. - Perform the Gateway reaction (Hartley et al., 2000) with the destination vector, a 5 entry vector containing a tissue-specific promoter of interest, a middle entry vector containing zebrafish codon-optimized Cas9 and p3e-polya, according to manufacturer s protocol. - Transform chemically competent bacteria, plate on LB Agar and select with ampicillin. - Extract plasmid DNA from 2-3 colonies and check correct recombination by digestion or sequencing.
6 grna 5 Entry Promoter grna U6:gRNA + Middle Entry Cas9 Gateway reaction U6:gRNA Prom:Cas9 3 Entry PolyA BseRI digestion grna U6:gRNA gene Prom:Cas9 seed ligation Figure P4 Construction of a tissue-specific CRISPR vector, option 2 Option 2 - Perform the Gateway reaction (Hartley et al., 2000) with the destination vector of your choice, a 5 entry vector containing a tissue-specific promoter of interest, a middle entry vector containing zebrafish codon-optimized Cas9 and p3e-polya, according to manufacturer s protocol. - Transform chemically competent bacteria, plate on LB Agar and select with ampicillin. - Extract plasmid DNA from 2-3 colonies and check correct recombination by digestion or sequencing. - Digest the resulting CRISPR vector with BseRI enzyme. - Purify on 1% agarose gel or column. - Ligate the gene-specific seed sequence into BseRI-digested destination vector (1:20 vector:insert molar ratio). - Transform chemically competent bacteria, plate on LB Agar and select with ampicillin. - Extract plasmid DNA from 2-3 distinct colonies and sequence using the following primer: CCTCACACAAACTCTGGATT.
7 4. Injection of the CRISPR construct in zebrafish embryos Mix and inject pg of CRISPR vector and 20 pg of Tol2 mrna into one-cell stage embryos (Kawakami et al., 2004). For reliable phenotypic analyses, we recommend injecting >50 embryos per construct. Vectors expressing grnas targeting an irrelevant gene or driving Cas9 expression in a different tissue can be used as negative controls. Allow injected embryos to develop at 28.5 C. 5. Phenotypic analysis and generation of stable lines Sort injected F0 embryos based on the expression of a transgenesis marker present either in the destination vector (e.g. cmlc2:gfp) or in the middle entry part of the vector (e.g. T2A-GFP). Only positive embryos should be considered for further analysis. For the generation of stable lines, raise positive F0 fish to adulthood. Back-cross them to the strain used for injection and sort positive F1 embryos (according to the expression of the transgenesis marker). F1 embryos can be analyzed phenotypically or raised to adulthood. Note: depending on the transgenesis marker used, it may be possible to evaluate the level of mosaicism in injected embryos. In that case, injected embryos could be further sorted according to their level of mosaicism. References Gagnon, J.A., Valen, E., Thyme, S.B., Huang, P., Ahkmetova, L., Pauli, A., Montague, T.G., Zimmerman, S., Richter, C., and Schier, A.F. (2014). Efficient mutagenesis by Cas9 protein-mediated oligonucleotide insertion and large-scale assessment of single-guide RNAs. PLoS One 9, e Hartley, J.L., Temple, G.F., and Brasch, M.A. (2000). DNA cloning using in vitro sitespecific recombination. Genome Res 10, Hsu, P.D., Scott, D.A., Weinstein, J.A., Ran, F.A., Konermann, S., Agarwala, V., Li, Y., Fine, E.J., Wu, X., Shalem, O., et al. (2013). DNA targeting specificity of RNAguided Cas9 nucleases. Nat Biotechnol 31,
8 Hwang, W.Y., Fu, Y., Reyon, D., Maeder, M.L., Tsai, S.Q., Sander, J.D., Peterson, R.T., Yeh, J.R., and Joung, J.K. (2013). Efficient genome editing in zebrafish using a CRISPR-Cas system. Nat Biotechnol 31, Kawakami, K., Takeda, H., Kawakami, N., Kobayashi, M., Matsuda, N., and Mishina, M. (2004). A transposon-mediated gene trap approach identifies developmentally regulated genes in zebrafish. Dev Cell 7, Kim, H.J., Lee, H.J., Kim, H., Cho, S.W., and Kim, J.S. (2009). Targeted genome editing in human cells with zinc finger nucleases constructed via modular assembly. Genome Res 19, Meeker, N.D., Hutchinson, S.A., Ho, L., and Trede, N.S. (2007). Method for isolation of PCR-ready genomic DNA from zebrafish tissues. Biotechniques 43, 610, 612, 614. Montague, T.G., Cruz, J.M., Gagnon, J.A., Church, G.M., and Valen, E. (2014). CHOPCHOP: a CRISPR/Cas9 and TALEN web tool for genome editing. Nucleic Acids Res 42, W
A CRISPR/Cas9 Vector System for Tissue-Specific Gene Disruption in Zebrafish
 Developmental Cell Supplemental Information A CRISPR/Cas9 Vector System for Tissue-Specific Gene Disruption in Zebrafish Julien Ablain, Ellen M. Durand, Song Yang, Yi Zhou, and Leonard I. Zon % larvae
Developmental Cell Supplemental Information A CRISPR/Cas9 Vector System for Tissue-Specific Gene Disruption in Zebrafish Julien Ablain, Ellen M. Durand, Song Yang, Yi Zhou, and Leonard I. Zon % larvae
Supplementary Methods pcfd5 cloning protocol
 Supplementary Methods cloning protocol vermilion trna grna trna grna U6:3 Terminator AmpR attb is a vector for expressing one or multiple trna-flanked Cas9 grnas under the control of the strong, ubiquitous
Supplementary Methods cloning protocol vermilion trna grna trna grna U6:3 Terminator AmpR attb is a vector for expressing one or multiple trna-flanked Cas9 grnas under the control of the strong, ubiquitous
Supplementary Materials
 Supplementary Materials Supplementary Methods Supplementary Discussion Supplementary Figure 1 Calculated frequencies of embryo cells bearing bi-allelic alterations. Targeted indel mutations induced by
Supplementary Materials Supplementary Methods Supplementary Discussion Supplementary Figure 1 Calculated frequencies of embryo cells bearing bi-allelic alterations. Targeted indel mutations induced by
Protocol for Genome Editing via the RNA-guided Cas9 Nuclease in. Zebrafish Embryos 1
 Protocol for Genome Editing via the RNA-guided Cas9 Nuclease in Zebrafish Embryos 1 1. In vitro synthesis of capped Cas9 mrna The full length of humanized Cas9 cdnas with double NLS were cloned into pxt7
Protocol for Genome Editing via the RNA-guided Cas9 Nuclease in Zebrafish Embryos 1 1. In vitro synthesis of capped Cas9 mrna The full length of humanized Cas9 cdnas with double NLS were cloned into pxt7
CRISPR RNA-guided activation of endogenous human genes
 CRISPR RNA-guided activation of endogenous human genes Morgan L Maeder, Samantha J Linder, Vincent M Cascio, Yanfang Fu, Quan H Ho, J Keith Joung Supplementary Figure 1 Comparison of VEGF activation induced
CRISPR RNA-guided activation of endogenous human genes Morgan L Maeder, Samantha J Linder, Vincent M Cascio, Yanfang Fu, Quan H Ho, J Keith Joung Supplementary Figure 1 Comparison of VEGF activation induced
Farnham Lab Protocol for CRISPR/Cas 9- mediated enhancer deletion using a puromycin selection vector Created by Yu (Phoebe) Guo
 Farnham Lab Protocol for CRISPR/Cas 9- mediated enhancer deletion using a puromycin selection vector Created by Yu (Phoebe) Guo 20151024 Part A. Design grna oligos 1. Design grnas Go to Optimized CRISPR
Farnham Lab Protocol for CRISPR/Cas 9- mediated enhancer deletion using a puromycin selection vector Created by Yu (Phoebe) Guo 20151024 Part A. Design grna oligos 1. Design grnas Go to Optimized CRISPR
Protocols for cloning SEC-based repair templates using SapTrap assembly
 Protocols for cloning SEC-based repair templates using SapTrap assembly Written by Dan Dickinson (ddickins@live.unc.edu) and last updated July 2016. Overview SapTrap (Schwartz and Jorgensen, 2016) is a
Protocols for cloning SEC-based repair templates using SapTrap assembly Written by Dan Dickinson (ddickins@live.unc.edu) and last updated July 2016. Overview SapTrap (Schwartz and Jorgensen, 2016) is a
Supplementary Figure 1. Diagram for CATCHA construct. Nature Biotechnology doi: /nbt.3444
 Supplementary Figure 1 Diagram for CATCHA construct. Supplementary Figure 2 Representative view of ebony (left) and non-ebony (right) F2 flies from experiments described in Fig. 1c. F0 #1 F0 #2 F0 #3 F0
Supplementary Figure 1 Diagram for CATCHA construct. Supplementary Figure 2 Representative view of ebony (left) and non-ebony (right) F2 flies from experiments described in Fig. 1c. F0 #1 F0 #2 F0 #3 F0
Protocols for cell lines using CRISPR/CAS
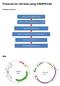 Protocols for cell lines using CRISPR/CAS Procedure overview Map Preparation of CRISPR/CAS plasmids Expression vectors for guide RNA (U6-gRNA) and Cas9 gene (CMV-p-Cas9) are ampicillin-resist ant and stable
Protocols for cell lines using CRISPR/CAS Procedure overview Map Preparation of CRISPR/CAS plasmids Expression vectors for guide RNA (U6-gRNA) and Cas9 gene (CMV-p-Cas9) are ampicillin-resist ant and stable
HE Swift Cloning Kit
 HE Swift Cloning Kit For high-efficient cloning of PCR products either blunt or sticky-end Kit Contents Contents VTT-BB05 phe Vector (35 ng/µl) 20 µl T4 DNA Ligase (3 U/µl) 20 µl 2 Reaction Buffer 100
HE Swift Cloning Kit For high-efficient cloning of PCR products either blunt or sticky-end Kit Contents Contents VTT-BB05 phe Vector (35 ng/µl) 20 µl T4 DNA Ligase (3 U/µl) 20 µl 2 Reaction Buffer 100
Paquet et al Supplementary Information
 Paquet et al. 2009 Supplementary Information Paquet et al. 2009 Supplementary Information Supplementary Figure 1 Integration of Gateway cloning technology can further improve the applicability of the transgenesis
Paquet et al. 2009 Supplementary Information Paquet et al. 2009 Supplementary Information Supplementary Figure 1 Integration of Gateway cloning technology can further improve the applicability of the transgenesis
Creating pentr vectors by BP reaction
 Creating pentr vectors by BP reaction Tanya Lepikhova and Rafael Martinez 15072011 Overview: 1. Design primers to add the attb sites to gene of interest 2. Perform PCR with a high fidelity DNA polymerase
Creating pentr vectors by BP reaction Tanya Lepikhova and Rafael Martinez 15072011 Overview: 1. Design primers to add the attb sites to gene of interest 2. Perform PCR with a high fidelity DNA polymerase
Data Sheet Quick PCR Cloning Kit
 Data Sheet Quick PCR Cloning Kit 6044 Cornerstone Ct. West, Ste. E DESCRIPTION: The Quick PCR Cloning Kit is a simple and highly efficient method to insert any gene or DNA fragment into a vector, without
Data Sheet Quick PCR Cloning Kit 6044 Cornerstone Ct. West, Ste. E DESCRIPTION: The Quick PCR Cloning Kit is a simple and highly efficient method to insert any gene or DNA fragment into a vector, without
Supplementary Information
 Supplementary Information Vidigal and Ventura a wt locus 5 region 3 region CCTCTGCCACTGCGAGGGCGTCCAATGGTGCTTG(...)AACAGGTGGAATATCCCTACTCTA predicted deletion clone 1 clone 2 clone 3 CCTCTGCCACTGCGAGGGCGTC-AGGTGGAATATCCCTACTCTA
Supplementary Information Vidigal and Ventura a wt locus 5 region 3 region CCTCTGCCACTGCGAGGGCGTCCAATGGTGCTTG(...)AACAGGTGGAATATCCCTACTCTA predicted deletion clone 1 clone 2 clone 3 CCTCTGCCACTGCGAGGGCGTC-AGGTGGAATATCCCTACTCTA
Polymerase Chain Reaction
 Polymerase Chain Reaction Amplify your insert or verify its presence 3H Taq platinum PCR mix primers Ultrapure Water PCR tubes PCR machine A. Insert amplification For insert amplification, use the Taq
Polymerase Chain Reaction Amplify your insert or verify its presence 3H Taq platinum PCR mix primers Ultrapure Water PCR tubes PCR machine A. Insert amplification For insert amplification, use the Taq
Conversion of plasmids into Gateway compatible cloning
 Conversion of plasmids into Gateway compatible cloning Rafael Martinez 14072011 Overview: 1. Select the right Gateway cassette (A, B or C). 2. Design primers to amplify the right Gateway cassette from
Conversion of plasmids into Gateway compatible cloning Rafael Martinez 14072011 Overview: 1. Select the right Gateway cassette (A, B or C). 2. Design primers to amplify the right Gateway cassette from
BP-reaction (recombination of PCR products into entry vector) Created on: 06/01/10 Version: 1.1 No.: 1 Page 1 of 6
 Created on: 06/01/10 Version: 1.1 No.: 1 Page 1 of 6 1. Background This reaction generates bacteria that contain recombinant Gateway-compatible entry plasmid. The PCRamplified ORFs (SOPs ORF-PCR 1 (two
Created on: 06/01/10 Version: 1.1 No.: 1 Page 1 of 6 1. Background This reaction generates bacteria that contain recombinant Gateway-compatible entry plasmid. The PCRamplified ORFs (SOPs ORF-PCR 1 (two
SUPPLEMENT MATERIALS FOR CURTIN,
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 SUPPLEMENT MATERIALS FOR CURTIN, et al. Validating genome-wide association candidates: Selecting, testing, and characterizing
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 SUPPLEMENT MATERIALS FOR CURTIN, et al. Validating genome-wide association candidates: Selecting, testing, and characterizing
Multiplex CRISPR/Cas9 Assembly System Kit protocol (Yamamoto lab)
 Multiplex CRISPR/Cas9 Assembly System Kit protocol (Yamamoto lab) Ver. 1.1 July 2014 Tetsushi Sakuma, Ph. D. E-mail: tetsushi-sakuma@hiroshima-u.ac.jp Takashi Yamamoto Lab. Department of Mathematical and
Multiplex CRISPR/Cas9 Assembly System Kit protocol (Yamamoto lab) Ver. 1.1 July 2014 Tetsushi Sakuma, Ph. D. E-mail: tetsushi-sakuma@hiroshima-u.ac.jp Takashi Yamamoto Lab. Department of Mathematical and
peco TM -T7-nGST, Eco cloning Kit User Manual (Patent pending)
 peco TM -T7-nGST, Eco cloning Kit User Manual (Patent pending) Cloning PCR products for E Coli expression of N-term GST-tagged protein Cat# Contents Amounts Application IC-1004 peco-t7-ngst vector built-in
peco TM -T7-nGST, Eco cloning Kit User Manual (Patent pending) Cloning PCR products for E Coli expression of N-term GST-tagged protein Cat# Contents Amounts Application IC-1004 peco-t7-ngst vector built-in
Gateway Cloning Protocol (Clough Lab Edition) This document is a modification of the Gateway cloning protocol developed by Manju in Chris Taylor's lab
 Gateway Cloning Protocol (Clough Lab Edition) This document is a modification of the Gateway cloning protocol developed by Manju in Chris Taylor's lab With the Gateway cloning system, a PCR fragment is
Gateway Cloning Protocol (Clough Lab Edition) This document is a modification of the Gateway cloning protocol developed by Manju in Chris Taylor's lab With the Gateway cloning system, a PCR fragment is
Simple protocol for gene editing using GenCrisprTM Cas9 nuclease
 Simple protocol for gene editing using GenCrisprTM Cas9 nuclease Contents Protocol Step 1: Choose the target DNA sequence Step 2: Design sgrna Step 3: Preparation for sgrna 3.1 In vitro transcription of
Simple protocol for gene editing using GenCrisprTM Cas9 nuclease Contents Protocol Step 1: Choose the target DNA sequence Step 2: Design sgrna Step 3: Preparation for sgrna 3.1 In vitro transcription of
Protocols for cloning SEC-based repair templates using SapTrap assembly
 Protocols for cloning SEC-based repair templates using SapTrap assembly Written by Dan Dickinson (daniel.dickinson@austin.utexas.edu) and last updated January 2018. Overview SapTrap (Schwartz and Jorgensen,
Protocols for cloning SEC-based repair templates using SapTrap assembly Written by Dan Dickinson (daniel.dickinson@austin.utexas.edu) and last updated January 2018. Overview SapTrap (Schwartz and Jorgensen,
Application of Molecular Biology tools for cloning of a foreign gene
 IFM/Kemi Linköpings Universitet September 2013/LGM Labmanual Project course Application of Molecular Biology tools for cloning of a foreign gene Table of contents Introduction... 3 Amplification of a gene
IFM/Kemi Linköpings Universitet September 2013/LGM Labmanual Project course Application of Molecular Biology tools for cloning of a foreign gene Table of contents Introduction... 3 Amplification of a gene
Multiplex CRISPR/Cas9 Assembly System Kit protocol (Yamamoto lab)
 Multiplex CRISPR/Cas9 Assembly System Kit protocol (Yamamoto lab) Ver. 2.0 May 2015 Tetsushi Sakuma, Ph. D. E-mail: tetsushi-sakuma@hiroshima-u.ac.jp Takashi Yamamoto Lab. Department of Mathematical and
Multiplex CRISPR/Cas9 Assembly System Kit protocol (Yamamoto lab) Ver. 2.0 May 2015 Tetsushi Sakuma, Ph. D. E-mail: tetsushi-sakuma@hiroshima-u.ac.jp Takashi Yamamoto Lab. Department of Mathematical and
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION Supplementary figure 1: List of primers/oligonucleotides used in this study. 1 Supplementary figure 2: Sequences and mirna-targets of i) mcherry expresses in transgenic fish used
SUPPLEMENTARY INFORMATION Supplementary figure 1: List of primers/oligonucleotides used in this study. 1 Supplementary figure 2: Sequences and mirna-targets of i) mcherry expresses in transgenic fish used
CELLTECHGEN For Research Only. Construction of all-in-one vector for Lenti-virus system (Example: Lenti-EF1 -Cas9-EGFP-U6 sgrna vector)
 Construction of all-in-one vector for Lenti-virus system (Example: Lenti-EF1 -Cas9-EGFP-U6 sgrna vector) Catalog number: CTG-CAS9-18 Introduction The vector Lenti-EF1 -Cas9-EGFP-U6 sgrna is designed for
Construction of all-in-one vector for Lenti-virus system (Example: Lenti-EF1 -Cas9-EGFP-U6 sgrna vector) Catalog number: CTG-CAS9-18 Introduction The vector Lenti-EF1 -Cas9-EGFP-U6 sgrna is designed for
Contents... vii. List of Figures... xii. List of Tables... xiv. Abbreviatons... xv. Summary... xvii. 1. Introduction In vitro evolution...
 vii Contents Contents... vii List of Figures... xii List of Tables... xiv Abbreviatons... xv Summary... xvii 1. Introduction...1 1.1 In vitro evolution... 1 1.2 Phage Display Technology... 3 1.3 Cell surface
vii Contents Contents... vii List of Figures... xii List of Tables... xiv Abbreviatons... xv Summary... xvii 1. Introduction...1 1.1 In vitro evolution... 1 1.2 Phage Display Technology... 3 1.3 Cell surface
Supplemental Protocols Gagnon et al., 2014
 Last updated April 15th 2014 There may be a more up-to-date version of this protocol on our website http://labs.mcb.harvard.edu/schier/index.html Protocol Page Flowchart for Cas9-mediated mutagenesis 1
Last updated April 15th 2014 There may be a more up-to-date version of this protocol on our website http://labs.mcb.harvard.edu/schier/index.html Protocol Page Flowchart for Cas9-mediated mutagenesis 1
CRISPR Ribonucleoprotein (RNP) User Manual
 CRISPR Ribonucleoprotein (RNP) User Manual Description This user manual describes how to use GenScript s CRISPR Ribonucleoprotein (RNP) products for targeted genome editing. The RNP system is comprised
CRISPR Ribonucleoprotein (RNP) User Manual Description This user manual describes how to use GenScript s CRISPR Ribonucleoprotein (RNP) products for targeted genome editing. The RNP system is comprised
CRISPR/Cas9 Genome Editing: Transfection Methods
 CRISPR/ Genome Editing: Transfection Methods For over 20 years Mirus Bio has developed and manufactured high performance transfection products and technologies. That expertise is now being applied to the
CRISPR/ Genome Editing: Transfection Methods For over 20 years Mirus Bio has developed and manufactured high performance transfection products and technologies. That expertise is now being applied to the
Molecular Genetics Techniques. BIT 220 Chapter 20
 Molecular Genetics Techniques BIT 220 Chapter 20 What is Cloning? Recombinant DNA technologies 1. Producing Recombinant DNA molecule Incorporate gene of interest into plasmid (cloning vector) 2. Recombinant
Molecular Genetics Techniques BIT 220 Chapter 20 What is Cloning? Recombinant DNA technologies 1. Producing Recombinant DNA molecule Incorporate gene of interest into plasmid (cloning vector) 2. Recombinant
Targeted Protein Degradation
 The PITCh dtag vectors for rapid protein degradation of endogenous targets The dissection of complex biological systems requires target-specific control of protein function or abundance. Genetic perturbations
The PITCh dtag vectors for rapid protein degradation of endogenous targets The dissection of complex biological systems requires target-specific control of protein function or abundance. Genetic perturbations
Supplementary Information
 Supplementary Information This Supplementary Information file contains the following contents: Supplementary Methods, Supplementary Discussion, Supplementary Figures (S1~S9), Supplementary Tables (1~3)
Supplementary Information This Supplementary Information file contains the following contents: Supplementary Methods, Supplementary Discussion, Supplementary Figures (S1~S9), Supplementary Tables (1~3)
Targeted Gene Mutation in Rice Using a CRISPR-Cas9 System Kabin Xie 1, Bastian Minkenberg 2 and Yinong Yang 2*
 Targeted Gene Mutation in Rice Using a CRISPR-Cas9 System Kabin Xie 1, Bastian Minkenberg 2 and Yinong Yang 2* 1 Department of Plant Pathology and Environmental Microbiology, Pennsylvania State University,
Targeted Gene Mutation in Rice Using a CRISPR-Cas9 System Kabin Xie 1, Bastian Minkenberg 2 and Yinong Yang 2* 1 Department of Plant Pathology and Environmental Microbiology, Pennsylvania State University,
Efficient In Vivo Genome Editing Using RNA-Guided Nucleases
 Efficient In Vivo Genome Editing Using RNA-Guided Nucleases The Harvard community has made this article openly available. Please share how this access benefits you. Your story matters Citation Hwang, Woong
Efficient In Vivo Genome Editing Using RNA-Guided Nucleases The Harvard community has made this article openly available. Please share how this access benefits you. Your story matters Citation Hwang, Woong
Hetero-Stagger PCR Cloning Kit
 Product Name: Code No: Size: DynaExpress Hetero-Stagger PCR Cloning Kit DS150 20 reactions Kit Components: Box 1 (-20 ) phst-1 Vector, linearized Annealing Buffer Ligase Mixture phst Forward Sequence Primer
Product Name: Code No: Size: DynaExpress Hetero-Stagger PCR Cloning Kit DS150 20 reactions Kit Components: Box 1 (-20 ) phst-1 Vector, linearized Annealing Buffer Ligase Mixture phst Forward Sequence Primer
Supplementary Materials for
 www.sciencemag.org/cgi/content/full/science.aaa5945/dc1 Supplementary Materials for The mutagenic chain reaction: A method for converting heterozygous to homozygous mutations Valentino M. Gantz* and Ethan
www.sciencemag.org/cgi/content/full/science.aaa5945/dc1 Supplementary Materials for The mutagenic chain reaction: A method for converting heterozygous to homozygous mutations Valentino M. Gantz* and Ethan
Cat # Box1 Box2. DH5a Competent E. coli cells CCK-20 (20 rxns) 40 µl 40 µl 50 µl x 20 tubes. Choo-Choo Cloning TM Enzyme Mix
 Molecular Cloning Laboratories User Manual Version 3.3 Product name: Choo-Choo Cloning Kits Cat #: CCK-10, CCK-20, CCK-096, CCK-384 Description: Choo-Choo Cloning is a highly efficient directional PCR
Molecular Cloning Laboratories User Manual Version 3.3 Product name: Choo-Choo Cloning Kits Cat #: CCK-10, CCK-20, CCK-096, CCK-384 Description: Choo-Choo Cloning is a highly efficient directional PCR
CRISPR Genome Editing Embryo Microinjection Service Catalog
 CRISPR Genome Editing Embryo Microinjection Service Catalog CRISPR Genome Editing Services (Drosophila) Embryo Microinjection Services (Drosophila & Mosquito) ellgenetics.com +8862-2651-1809 +8862-2782-9911
CRISPR Genome Editing Embryo Microinjection Service Catalog CRISPR Genome Editing Services (Drosophila) Embryo Microinjection Services (Drosophila & Mosquito) ellgenetics.com +8862-2651-1809 +8862-2782-9911
NZYGene Synthesis kit
 Kit components Component Concentration Amount NZYGene Synthesis kit Catalogue number: MB33901, 10 reactions GS DNA Polymerase 1U/ μl 30 μl Reaction Buffer for GS DNA Polymerase 10 150 μl dntp mix 2 mm
Kit components Component Concentration Amount NZYGene Synthesis kit Catalogue number: MB33901, 10 reactions GS DNA Polymerase 1U/ μl 30 μl Reaction Buffer for GS DNA Polymerase 10 150 μl dntp mix 2 mm
In order to make our construct pah12 (Mlra) and pah05 (venus) biobrick compatitable PCR was conducted. Following PCR mixes were prepared
 BioBrick cloning Project: BioBricks Authors: Antti Koistinen Dates: 2016-09-15 to 2016-10-06 THURSDAY, 9/15 In order to make our construct pah12 (Mlra) and pah05 (venus) biobrick compatitable PCR was conducted.
BioBrick cloning Project: BioBricks Authors: Antti Koistinen Dates: 2016-09-15 to 2016-10-06 THURSDAY, 9/15 In order to make our construct pah12 (Mlra) and pah05 (venus) biobrick compatitable PCR was conducted.
3 Designing Primers for Site-Directed Mutagenesis
 3 Designing Primers for Site-Directed Mutagenesis 3.1 Learning Objectives During the next two labs you will learn the basics of site-directed mutagenesis: you will design primers for the mutants you designed
3 Designing Primers for Site-Directed Mutagenesis 3.1 Learning Objectives During the next two labs you will learn the basics of site-directed mutagenesis: you will design primers for the mutants you designed
Designing CRISPR mediated Gene disruptions with gblocks Gene Fragments
 Designing CRISPR mediated Gene disruptions with gblocks Gene Fragments Adam Clore, PhD Manager, Synthetic Biology Design Integrated DNA technology Typical CRISPR timeline in Mammalian cell lines Design
Designing CRISPR mediated Gene disruptions with gblocks Gene Fragments Adam Clore, PhD Manager, Synthetic Biology Design Integrated DNA technology Typical CRISPR timeline in Mammalian cell lines Design
Protocol for cloning SEC-based repair templates using Gibson assembly and ccdb negative selection
 Protocol for cloning SEC-based repair templates using Gibson assembly and ccdb negative selection Written by Dan Dickinson (daniel.dickinson@austin.utexas.edu) and last updated January 2018. A version
Protocol for cloning SEC-based repair templates using Gibson assembly and ccdb negative selection Written by Dan Dickinson (daniel.dickinson@austin.utexas.edu) and last updated January 2018. A version
PrecisionX Multiplex grna Cloning Kit. Cat. # CAS9-GRNA-KIT. User Manual
 PrecisionX Multiplex grna Cloning Kit Store at -20 C upon receipt A limited-use label license covers this product. By use of this product, you accept the terms and conditions outlined in the Licensing
PrecisionX Multiplex grna Cloning Kit Store at -20 C upon receipt A limited-use label license covers this product. By use of this product, you accept the terms and conditions outlined in the Licensing
Programmable Sequence-Specific Transcriptional Regulation of Mammalian Genome Using Designer TAL Effectors
 Supplementary Information Programmable Sequence-Specific Transcriptional Regulation of Mammalian Genome Using Designer TAL Effectors Feng Zhang 1,2,3,5,7 *,±, Le Cong 2,3,4 *, Simona Lodato 5,6, Sriram
Supplementary Information Programmable Sequence-Specific Transcriptional Regulation of Mammalian Genome Using Designer TAL Effectors Feng Zhang 1,2,3,5,7 *,±, Le Cong 2,3,4 *, Simona Lodato 5,6, Sriram
Optimized, chemically-modified crrna:tracrrna complexes for CRISPR gene editing
 Optimized, chemically-modified crrna:tracrrna complexes for CRISPR gene editing Mark Behlke MD, PhD Chief Scientific Officer February 24, 2016 1 Implementing CRISPR/Cas9 gene editing 2 To focus on RNA
Optimized, chemically-modified crrna:tracrrna complexes for CRISPR gene editing Mark Behlke MD, PhD Chief Scientific Officer February 24, 2016 1 Implementing CRISPR/Cas9 gene editing 2 To focus on RNA
Figure 1. Map of cloning vector pgem T-Easy (bacterial plasmid DNA)
 Texas A&M University-Corpus Christi CHEM4402 Biochemistry II Laboratory Laboratory 6: Ligation & Bacterial Transformation (Bring your text and laptop to class if you wish to work on your assignment during
Texas A&M University-Corpus Christi CHEM4402 Biochemistry II Laboratory Laboratory 6: Ligation & Bacterial Transformation (Bring your text and laptop to class if you wish to work on your assignment during
Ligation Independent Cloning (LIC) Procedure
 Ligation Independent Cloning (LIC) Procedure Ligation Independent Cloning (LIC) LIC cloning allows insertion of DNA fragments without using restriction enzymes into specific vectors containing engineered
Ligation Independent Cloning (LIC) Procedure Ligation Independent Cloning (LIC) LIC cloning allows insertion of DNA fragments without using restriction enzymes into specific vectors containing engineered
pdsipher and pdsipher -GFP shrna Vector User s Guide
 pdsipher and pdsipher -GFP shrna Vector User s Guide NOTE: PLEASE READ THE ENTIRE PROTOCOL CAREFULLY BEFORE USE Page 1. Introduction... 1 2. Vector Overview... 1 3. Vector Maps 2 4. Materials Provided...
pdsipher and pdsipher -GFP shrna Vector User s Guide NOTE: PLEASE READ THE ENTIRE PROTOCOL CAREFULLY BEFORE USE Page 1. Introduction... 1 2. Vector Overview... 1 3. Vector Maps 2 4. Materials Provided...
GenBuilder TM Plus Cloning Kit User Manual
 GenBuilder TM Plus Cloning Kit User Manual Cat. No. L00744 Version 11242017 Ⅰ. Introduction... 2 I.1 Product Information... 2 I.2 Kit Contents and Storage... 2 I.3 GenBuilder Cloning Kit Workflow... 2
GenBuilder TM Plus Cloning Kit User Manual Cat. No. L00744 Version 11242017 Ⅰ. Introduction... 2 I.1 Product Information... 2 I.2 Kit Contents and Storage... 2 I.3 GenBuilder Cloning Kit Workflow... 2
XactEdit Cas9 Nuclease with NLS User Manual
 XactEdit Cas9 Nuclease with NLS User Manual An RNA-guided recombinant endonuclease for efficient targeted DNA cleavage Catalog Numbers CE1000-50K, CE1000-50, CE1000-250, CE1001-250, CE1001-1000 Table of
XactEdit Cas9 Nuclease with NLS User Manual An RNA-guided recombinant endonuclease for efficient targeted DNA cleavage Catalog Numbers CE1000-50K, CE1000-50, CE1000-250, CE1001-250, CE1001-1000 Table of
GenBuilder TM Plus Cloning Kit User Manual
 GenBuilder TM Plus Cloning Kit User Manual Cat.no L00744 Version 11242017 Ⅰ. Introduction... 2 I.1 Product Information... 2 I.2 Kit Contents and Storage... 2 I.3 GenBuilder Cloning Kit Workflow... 2 Ⅱ.
GenBuilder TM Plus Cloning Kit User Manual Cat.no L00744 Version 11242017 Ⅰ. Introduction... 2 I.1 Product Information... 2 I.2 Kit Contents and Storage... 2 I.3 GenBuilder Cloning Kit Workflow... 2 Ⅱ.
Genome Engineering with ZFNs, TALENs and CRISPR/Cas9
 Genome Engineering with ZFNs, TALENs and CRISPR/Cas9 Designer Endonucleases ZFNs (zinc finger nucleases), TALENs (transcription activator-like effector nucleases) and CRISPR/Cas9 (clustered regularly interspaced
Genome Engineering with ZFNs, TALENs and CRISPR/Cas9 Designer Endonucleases ZFNs (zinc finger nucleases), TALENs (transcription activator-like effector nucleases) and CRISPR/Cas9 (clustered regularly interspaced
File S1. Detailed protocol for CRISPR/Cas9 mediated genome editing with dual marker cassettes
 File S1 Detailed protocol for CRISPR/Cas9 mediated genome editing with dual marker cassettes A) Preparing sgrna vectors We ve modified the strategy to create new sgrna vectors starting with the klp 12
File S1 Detailed protocol for CRISPR/Cas9 mediated genome editing with dual marker cassettes A) Preparing sgrna vectors We ve modified the strategy to create new sgrna vectors starting with the klp 12
Organization of the libraries. For PCR screening
 Organization of the libraries For PCR screening Recall : Construction of a library (YAC and BAC) Genomic DNA Vector BAC YAC restriction enzyme digestion + PFGE BAC YAC ligation transformation Host-cells
Organization of the libraries For PCR screening Recall : Construction of a library (YAC and BAC) Genomic DNA Vector BAC YAC restriction enzyme digestion + PFGE BAC YAC ligation transformation Host-cells
XXII DNA cloning and sequencing. Outline
 XXII DNA cloning and sequencing 1) Deriving DNA for cloning Outline 2) Vectors; forming recombinant DNA; cloning DNA; and screening for clones containing recombinant DNA [replica plating and autoradiography;
XXII DNA cloning and sequencing 1) Deriving DNA for cloning Outline 2) Vectors; forming recombinant DNA; cloning DNA; and screening for clones containing recombinant DNA [replica plating and autoradiography;
CELLTECHGEN For Research Only. Construction of sgrna expression vector for Lenti-virus system (Example: Lenti-U6 sgrna-ef1 -Puro vector)
 Construction of sgrna expression vector for Lenti-virus system (Example: Lenti-U6 sgrna-ef1 -Puro vector) Catalog number: CTG-CAS9-11 Introduction The vector Lenti-U6 sgrna-ef1 -Puro is designed for expression
Construction of sgrna expression vector for Lenti-virus system (Example: Lenti-U6 sgrna-ef1 -Puro vector) Catalog number: CTG-CAS9-11 Introduction The vector Lenti-U6 sgrna-ef1 -Puro is designed for expression
Allele-specific locus binding and genome editing by CRISPR at the
 Supplementary Information Allele-specific locus binding and genome editing by CRISPR at the p6ink4a locus Toshitsugu Fujita, Miyuki Yuno, and Hodaka Fujii Supplementary Figure Legends Supplementary Figure
Supplementary Information Allele-specific locus binding and genome editing by CRISPR at the p6ink4a locus Toshitsugu Fujita, Miyuki Yuno, and Hodaka Fujii Supplementary Figure Legends Supplementary Figure
pslic-purop2a Lentivirus Plasmid Construction Kit (Cat# LT2002)
 pslic-purop2a Lentivirus Plasmid Construction Kit (Cat# LT2002) [ provides pslic-purop2a lentivirus plasmid construction kit to let user create the lentivirus plasmid expressing the gene of interest. This
pslic-purop2a Lentivirus Plasmid Construction Kit (Cat# LT2002) [ provides pslic-purop2a lentivirus plasmid construction kit to let user create the lentivirus plasmid expressing the gene of interest. This
pgm-t Cloning Kit Cat. # : GVT202 Size : 20 Reactions Store at -20
 pgm-t Cloning Kit Cat. # : GVT202 Size : 20 Reactions Store at -20 1 Kit Contents Contents pgm-t Cloning Kit pgm-t Vector (50 ng/μl) 20 μl T4 DNA Ligase (3 U/μl) 20 μl 10X T4 DNA Ligation Buffer 30 μl
pgm-t Cloning Kit Cat. # : GVT202 Size : 20 Reactions Store at -20 1 Kit Contents Contents pgm-t Cloning Kit pgm-t Vector (50 ng/μl) 20 μl T4 DNA Ligase (3 U/μl) 20 μl 10X T4 DNA Ligation Buffer 30 μl
Automated Electrophoresis Applications for Synthetic Biology
 Automated Electrophoresis Applications for Synthetic Biology Melissa Huang Liu, Ph.D. Product Manager, Electrophoresis Agilent Technologies 1 Outline 2100 Bioanalyzer & 4200 TapeStation Systems Automated
Automated Electrophoresis Applications for Synthetic Biology Melissa Huang Liu, Ph.D. Product Manager, Electrophoresis Agilent Technologies 1 Outline 2100 Bioanalyzer & 4200 TapeStation Systems Automated
Fast and efficient site-directed mutagenesis with Platinum SuperFi DNA Polymerase
 APPLICATION NOTE Platinum Superi Polymerase ast and efficient site-directed mutagenesis with Platinum Superi Polymerase Introduction Site-directed mutagenesis is one of the most essential techniques to
APPLICATION NOTE Platinum Superi Polymerase ast and efficient site-directed mutagenesis with Platinum Superi Polymerase Introduction Site-directed mutagenesis is one of the most essential techniques to
Biotechnology. Chapter 20. Biology Eighth Edition Neil Campbell and Jane Reece. PowerPoint Lecture Presentations for
 Chapter 20 Biotechnology PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from Joan Sharp Copyright
Chapter 20 Biotechnology PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from Joan Sharp Copyright
TrueORF TM cdna Clones and PrecisionShuttle TM Vector System
 TrueORF TM cdna Clones and PrecisionShuttle TM Vector System Application Guide Table of Contents Package Contents and Storage Conditions... 2 Related, Optional Reagents... 2 Related Products... 2 Available
TrueORF TM cdna Clones and PrecisionShuttle TM Vector System Application Guide Table of Contents Package Contents and Storage Conditions... 2 Related, Optional Reagents... 2 Related Products... 2 Available
Molecular Biology Techniques Supporting IBBE
 Molecular Biology Techniques Supporting IBBE Jared Cartwright Protein Production Lab Head Contact Details: email jared.cartwright@york.ac.uk Phone 01904 328797 Presentation Aims Gene synthesis Cloning
Molecular Biology Techniques Supporting IBBE Jared Cartwright Protein Production Lab Head Contact Details: email jared.cartwright@york.ac.uk Phone 01904 328797 Presentation Aims Gene synthesis Cloning
Lentiviral CRISPR guild RNA Cloning Kit for constructing CRISPR targeting grna lentivectors
 Lentiviral CRISPR guild RNA Cloning Kit for constructing CRISPR targeting grna lentivectors Cat# Product Name Amount Application grna-h1-gb grna-h1-gp grna-h1-rb grna-h1-rp grna-h1-puro grna-h1-bsd grna-u6-gb
Lentiviral CRISPR guild RNA Cloning Kit for constructing CRISPR targeting grna lentivectors Cat# Product Name Amount Application grna-h1-gb grna-h1-gp grna-h1-rb grna-h1-rp grna-h1-puro grna-h1-bsd grna-u6-gb
Chapter 6 - Molecular Genetic Techniques
 Chapter 6 - Molecular Genetic Techniques Two objects of molecular & genetic technologies For analysis For generation Molecular genetic technologies! For analysis DNA gel electrophoresis Southern blotting
Chapter 6 - Molecular Genetic Techniques Two objects of molecular & genetic technologies For analysis For generation Molecular genetic technologies! For analysis DNA gel electrophoresis Southern blotting
Cold Fusion Cloning Kit. Cat. #s MC100A-1, MC101A-1. User Manual
 Fusion Cloning technology Cold Fusion Cloning Kit Store the master mixture and positive controls at -20 C Store the competent cells at -80 C. (ver. 120909) A limited-use label license covers this product.
Fusion Cloning technology Cold Fusion Cloning Kit Store the master mixture and positive controls at -20 C Store the competent cells at -80 C. (ver. 120909) A limited-use label license covers this product.
KOD -Plus- Mutagenesis Kit
 Instruction manual KOD -Plus- Mutagenesis Kit 0811 F0936K KOD -Plus- Mutagenesis Kit SMK-101 20 reactions Store at -20 C Contents [1] Introduction [2] Flow chart [3] Components [4] Notes [5] Protocol 1.
Instruction manual KOD -Plus- Mutagenesis Kit 0811 F0936K KOD -Plus- Mutagenesis Kit SMK-101 20 reactions Store at -20 C Contents [1] Introduction [2] Flow chart [3] Components [4] Notes [5] Protocol 1.
For designing necessary primers and oligos for grna, online CRISPR designing tools can be used (e.g.
 DESIGN OF grnas For designing necessary primers and oligos for grna, online CRISPR designing tools can be used (e.g. http://crispr.mit.edu/, https://benchling.com). Assembly of grna transcriptional cassettes
DESIGN OF grnas For designing necessary primers and oligos for grna, online CRISPR designing tools can be used (e.g. http://crispr.mit.edu/, https://benchling.com). Assembly of grna transcriptional cassettes
Incorporating SeqStudio Genetic Analyzer and Sanger sequencing into genome editing workflows
 Incorporating SeqStudio Genetic Analyzer and Sanger sequencing into genome editing workflows Stephen Jackson, Ph.D 27 May 2017 The world leader in serving science Key Applications for Genome Editing Research
Incorporating SeqStudio Genetic Analyzer and Sanger sequencing into genome editing workflows Stephen Jackson, Ph.D 27 May 2017 The world leader in serving science Key Applications for Genome Editing Research
Alt-Ring the genome with CRISPR- Cas9
 Alt-Ring the genome with CRISPR- Cas9 Genome editing done easily and fast CHIA Jin Ngee Regional Application Specialist 1 Integrated DNA Technologies The Custom Biology Company 2 IDT was founded by a scientist
Alt-Ring the genome with CRISPR- Cas9 Genome editing done easily and fast CHIA Jin Ngee Regional Application Specialist 1 Integrated DNA Technologies The Custom Biology Company 2 IDT was founded by a scientist
CRISPR/Cas9 Gene Editing Tools
 CRISPR/Cas9 Gene Editing Tools - Guide-it Products for Successful CRISPR/Cas9 Gene Editing - Why choose Guide-it products? Optimized methods designed for speed and ease of use Complete kits that don t
CRISPR/Cas9 Gene Editing Tools - Guide-it Products for Successful CRISPR/Cas9 Gene Editing - Why choose Guide-it products? Optimized methods designed for speed and ease of use Complete kits that don t
Construct Design and Cloning Guide for Cas9-triggered homologous recombination
 Construct Design and Cloning Guide for Cas9-triggered homologous recombination Written by Dan Dickinson (ddickins@live.unc.edu) and last updated December 2013. Reference: Dickinson DJ, Ward JD, Reiner
Construct Design and Cloning Guide for Cas9-triggered homologous recombination Written by Dan Dickinson (ddickins@live.unc.edu) and last updated December 2013. Reference: Dickinson DJ, Ward JD, Reiner
Transfection of CRISPR/Cas9 Nuclease NLS ribonucleoprotein (RNP) into adherent mammalian cells using Lipofectamine RNAiMAX
 Transfection of CRISPR/Cas9 Nuclease NLS ribonucleoprotein (RNP) into adherent mammalian cells using Lipofectamine RNAiMAX INTRODUCTION The CRISPR/Cas genome editing system consists of a single guide RNA
Transfection of CRISPR/Cas9 Nuclease NLS ribonucleoprotein (RNP) into adherent mammalian cells using Lipofectamine RNAiMAX INTRODUCTION The CRISPR/Cas genome editing system consists of a single guide RNA
pgm-t Cloning Kit For direct cloning of PCR products generated by Taq DNA polymerases For research use only Cat. # : GVT202 Size : 20 Reactions
 pgm-t Cloning Kit For direct cloning of PCR products generated by Taq DNA polymerases Cat. # : GVT202 Size : 20 Reactions Store at -20 For research use only 1 pgm-t Cloning Kit Cat. No.: GVT202 Kit Contents
pgm-t Cloning Kit For direct cloning of PCR products generated by Taq DNA polymerases Cat. # : GVT202 Size : 20 Reactions Store at -20 For research use only 1 pgm-t Cloning Kit Cat. No.: GVT202 Kit Contents
Genome Editing with Programmable Nucleases. Jin-Soo Kim Department of Chemistry Seoul National University
 Genome Editing with Programmable Nucleases Jin-Soo Kim Department of Chemistry Seoul National University 1 Method of the Year 2011: Engineered Nucleases RNA-guided Cas9 Endonuclease 3 FokI and the First
Genome Editing with Programmable Nucleases Jin-Soo Kim Department of Chemistry Seoul National University 1 Method of the Year 2011: Engineered Nucleases RNA-guided Cas9 Endonuclease 3 FokI and the First
Journal of Experimental Microbiology and Immunology (JEMI) Vol. 6:20-25 Copyright December 2004, M&I UBC
 Preparing Plasmid Constructs to Investigate the Characteristics of Thiol Reductase and Flavin Reductase With Regard to Solubilizing Insoluble Proteinase Inhibitor 2 in Bacterial Protein Overexpression
Preparing Plasmid Constructs to Investigate the Characteristics of Thiol Reductase and Flavin Reductase With Regard to Solubilizing Insoluble Proteinase Inhibitor 2 in Bacterial Protein Overexpression
Chapter 20 Recombinant DNA Technology. Copyright 2009 Pearson Education, Inc.
 Chapter 20 Recombinant DNA Technology Copyright 2009 Pearson Education, Inc. 20.1 Recombinant DNA Technology Began with Two Key Tools: Restriction Enzymes and DNA Cloning Vectors Recombinant DNA refers
Chapter 20 Recombinant DNA Technology Copyright 2009 Pearson Education, Inc. 20.1 Recombinant DNA Technology Began with Two Key Tools: Restriction Enzymes and DNA Cloning Vectors Recombinant DNA refers
Genome editing. Knock-ins
 Genome editing Knock-ins Experiment design? Should we even do it? In mouse or rat, the HR-mediated knock-in of homologous fragments derived from a donor vector functions well. However, HR-dependent knock-in
Genome editing Knock-ins Experiment design? Should we even do it? In mouse or rat, the HR-mediated knock-in of homologous fragments derived from a donor vector functions well. However, HR-dependent knock-in
(Supplementary Methods online)
 (Supplementary Methods online) Production and purification of either LC-antisense or control molecules Recombinant phagemids and the phagemid vector were transformed into XL-1 Blue competent bacterial
(Supplementary Methods online) Production and purification of either LC-antisense or control molecules Recombinant phagemids and the phagemid vector were transformed into XL-1 Blue competent bacterial
The GeneEditor TM in vitro Mutagenesis System: Site- Directed Mutagenesis Using Altered Beta-Lactamase Specificity
 Promega Notes Magazine Number 62, 1997, p. 02 The GeneEditor TM in vitro Mutagenesis System: Site- Directed Mutagenesis Using Altered Beta-Lactamase Specificity By Christine Andrews and Scott Lesley Promega
Promega Notes Magazine Number 62, 1997, p. 02 The GeneEditor TM in vitro Mutagenesis System: Site- Directed Mutagenesis Using Altered Beta-Lactamase Specificity By Christine Andrews and Scott Lesley Promega
Lab Book igem Stockholm Lysostaphin. Week 8
 Lysostaphin Week 8 Summarized below are the experiments conducted this week in chronological order. Click on the experiment name to view it. To go back to this summary, click Summary in the footer. Summary
Lysostaphin Week 8 Summarized below are the experiments conducted this week in chronological order. Click on the experiment name to view it. To go back to this summary, click Summary in the footer. Summary
Learning Objectives :
 Learning Objectives : Understand the basic differences between genomic and cdna libraries Understand how genomic libraries are constructed Understand the purpose for having overlapping DNA fragments in
Learning Objectives : Understand the basic differences between genomic and cdna libraries Understand how genomic libraries are constructed Understand the purpose for having overlapping DNA fragments in
Nature Biotechnology: doi: /nbt.4166
 Supplementary Figure 1 Validation of correct targeting at targeted locus. (a) by immunofluorescence staining of 2C-HR-CRISPR microinjected embryos cultured to the blastocyst stage. Embryos were stained
Supplementary Figure 1 Validation of correct targeting at targeted locus. (a) by immunofluorescence staining of 2C-HR-CRISPR microinjected embryos cultured to the blastocyst stage. Embryos were stained
7.1 Techniques for Producing and Analyzing DNA. SBI4U Ms. Ho-Lau
 7.1 Techniques for Producing and Analyzing DNA SBI4U Ms. Ho-Lau What is Biotechnology? From Merriam-Webster: the manipulation of living organisms or their components to produce useful usually commercial
7.1 Techniques for Producing and Analyzing DNA SBI4U Ms. Ho-Lau What is Biotechnology? From Merriam-Webster: the manipulation of living organisms or their components to produce useful usually commercial
Protocols. We used a buffer mix with polymerase (either Mango Mix (forhandler) or 2x High Fidelity) and the appropriate primers and template DNA.
 Protocols PCRs Colony PCRs We used a buffer mix with polymerase (either Mango Mix (forhandler) or 2x High Fidelity) and the appropriate primers and template DNA. For a 10 µl reaction 2x Mango Mix/2x High
Protocols PCRs Colony PCRs We used a buffer mix with polymerase (either Mango Mix (forhandler) or 2x High Fidelity) and the appropriate primers and template DNA. For a 10 µl reaction 2x Mango Mix/2x High
Improving CRISPR-Cas9 Gene Knockout with a Validated Guide RNA Algorithm
 Improving CRISPR-Cas9 Gene Knockout with a Validated Guide RNA Algorithm Anja Smith Director R&D Dharmacon, part of GE Healthcare Imagination at work crrna:tracrrna program Cas9 nuclease Active crrna is
Improving CRISPR-Cas9 Gene Knockout with a Validated Guide RNA Algorithm Anja Smith Director R&D Dharmacon, part of GE Healthcare Imagination at work crrna:tracrrna program Cas9 nuclease Active crrna is
Version(1(*(July(14 th,(2014(
 Morrisey(Lab(Protocol:(( Generating(Large((>1kb)(Genomic(Deletions(Using(CRISPRs( bydanswarr&davefrank GeneralComments ThisprotocolisintendedtousetheCRISPR@Cas9systemforgenerationoflarge genomicdeletions(>1kb),inordertoremoveorinactivateagenomicelementof
Morrisey(Lab(Protocol:(( Generating(Large((>1kb)(Genomic(Deletions(Using(CRISPRs( bydanswarr&davefrank GeneralComments ThisprotocolisintendedtousetheCRISPR@Cas9systemforgenerationoflarge genomicdeletions(>1kb),inordertoremoveorinactivateagenomicelementof
Application Note: Generating GFP-Tagged Human CD81 Tetraspanin Protein Using SBI s PrecisionX SmartNuclease System And HR Tagging Vectors
 Application Note: Generating GFP-Tagged Human CD81 Tetraspanin Protein Using SBI s PrecisionX SmartNuclease System And HR Tagging Vectors Table of Contents: I. Background Pg. 1 II. Analysis of Gene Target
Application Note: Generating GFP-Tagged Human CD81 Tetraspanin Protein Using SBI s PrecisionX SmartNuclease System And HR Tagging Vectors Table of Contents: I. Background Pg. 1 II. Analysis of Gene Target
GenBuilder TM Cloning Kit User Manual
 GenBuilder TM Cloning Kit User Manual Cat.no L00701 Version 11242017 Ⅰ. Introduction... 2 I.1 Product Information... 2 I.2 Kit Contents and Storage... 2 I.3 GenBuilder Cloning Kit Workflow... 2 Ⅱ. DNA
GenBuilder TM Cloning Kit User Manual Cat.no L00701 Version 11242017 Ⅰ. Introduction... 2 I.1 Product Information... 2 I.2 Kit Contents and Storage... 2 I.3 GenBuilder Cloning Kit Workflow... 2 Ⅱ. DNA
CRISPR/Cas9 Gene Editing Tools
 CRISPR/Cas9 Gene Editing Tools - Separations Simply Spectacular INDELS Identify indels Determine if one or both copies of your gene have indels The Guide-it Genotype Confirmation Kit: Simple detection
CRISPR/Cas9 Gene Editing Tools - Separations Simply Spectacular INDELS Identify indels Determine if one or both copies of your gene have indels The Guide-it Genotype Confirmation Kit: Simple detection
Development of Positive Control for Hepatitis B Virus
 Human Journals Research Article December 2015 Vol.:2, Issue:2 All rights are reserved by Saurabh Bandhavkar et al. Development of Positive Control for Hepatitis B Virus Keywords: Hepatitis B virus, pbluescript,
Human Journals Research Article December 2015 Vol.:2, Issue:2 All rights are reserved by Saurabh Bandhavkar et al. Development of Positive Control for Hepatitis B Virus Keywords: Hepatitis B virus, pbluescript,
Supporting Information
 Supporting Information Development of a 2,4-Dinitrotoluene-Responsive Synthetic Riboswitch in E. coli cells Molly E. Davidson, Svetlana V. Harbaugh, Yaroslav G. Chushak, Morley O. Stone, Nancy Kelley-
Supporting Information Development of a 2,4-Dinitrotoluene-Responsive Synthetic Riboswitch in E. coli cells Molly E. Davidson, Svetlana V. Harbaugh, Yaroslav G. Chushak, Morley O. Stone, Nancy Kelley-
VDL101.3 CLONING TRANSGENE INTO pad5f35
 Purpose 1.1. The purpose of this protocol is to transfer a transgene from the pshuttlex plasmid to pad5/f35. 1.2. The starting material is 10 μg plasmid DNA. 1.3. This procedure is routinely performed
Purpose 1.1. The purpose of this protocol is to transfer a transgene from the pshuttlex plasmid to pad5/f35. 1.2. The starting material is 10 μg plasmid DNA. 1.3. This procedure is routinely performed
Supplementary Protocol: CIRCLE-seq Library Preparation
 Supplementary Protocol: CIRCLE-seq Library Preparation Reagent Gentra Puregene Tissue Kit Qubit dsdna BR Assay Kit Agencourt AMPure XP magnetic beads High throughput, with bead, PCR-free Library Preparation
Supplementary Protocol: CIRCLE-seq Library Preparation Reagent Gentra Puregene Tissue Kit Qubit dsdna BR Assay Kit Agencourt AMPure XP magnetic beads High throughput, with bead, PCR-free Library Preparation
A universal chassis plasmid pwh34 was first constructed to facilitate the
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 File name: Supplementary method Plasmids construction A universal chassis plasmid pwh34 was first constructed to facilitate the construction
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 File name: Supplementary method Plasmids construction A universal chassis plasmid pwh34 was first constructed to facilitate the construction
YG1 Control. YG1 PstI XbaI. YG5 PstI XbaI Water Buffer DNA Enzyme1 Enzyme2
 9/9/03 Aim: Digestion and gel extraction of YG, YG3, YG5 and 8/C. Strain: E. coli DH5α Plasmid: Bba_J600, psbc3 4,, 3 6 7, 8 9 0, 3, 4 8/C 8/C SpeI PstI 3 3 x3 YG YG PstI XbaI.5 3.5 x YG3 YG3 PstI XbaI
9/9/03 Aim: Digestion and gel extraction of YG, YG3, YG5 and 8/C. Strain: E. coli DH5α Plasmid: Bba_J600, psbc3 4,, 3 6 7, 8 9 0, 3, 4 8/C 8/C SpeI PstI 3 3 x3 YG YG PstI XbaI.5 3.5 x YG3 YG3 PstI XbaI
