7.17: Writing Up Results and Creating Illustrations
|
|
|
- Brendan Marshall
- 6 years ago
- Views:
Transcription
1 7.17: Writing Up Results and Creating Illustrations A Results Exercise: Kansas and Pancakes Write a 5-sentence paragraph describing the results illustrated in this figure: - Describe the figure: highlights? trends? conclusions? - Be sure to include a topic and a concluding sentence; watch for structure and coherence. Elevation (mm) Elevation (m) W-E Distance (m) Distance (mm) Fonstad et al. (2003) AIR 9 (3): 16. 1
2 from from 2
3 What is the Purpose of the Results Section? Objectivity: Make the data, just the data, easy to find. Some readers want to interpret your data themselves rather than accepting the interpretation presented in the discussion. Description: Describe the data presented in figures and tables. What Differentiates Results from the Methods? Methods = How the data were accumulated. Results = What data were accumulated. Readers expect to find the answers to your research questions in your Results section. 3
4 What Differentiates Results from Discussion? Results = Data Presentation ( Experiments showed that.... ) Discussion = Data Interpretation ( Experiments suggest that.... ) However, you still need to choose which data to present in your Results Section (an act of interpretation!). What are the Contents of a Results Section? A brief description of the experiment or rationale at the beginning of each subsection ( In order to.... As a result, we found that....). The data (in past tense). Descriptive text for FEW determinations. Tables or graphs for REPETITIVE determinations. The data that your methods indicated you would produce (and answering the questions you established in your introduction). 4
5 What are some qualities of a well-written Results section? Methods and Results Correspond. i.e., no experimental results for which there are no methods, and vice versa. Results are presented in a logical order. e.g., most important first, most fundamental first, etc. Results focus on the question(s) or hypothesis introduced earlier in the paper. What are some pitfalls of a Results section? Overstating the results (e.g., Figure 1 clearly shows ) Reporting irrelevant results Although it is sometimes useful to report experiments that didn t work. Omitting visual organizers Such as subheads. Including inappropriate illustrations (data best described in text). Including methods and/or discussion. Overlap is acceptable in some circumstances. 5
6 Results Example 1: Creating a context for the results Results I hypothesize that CG7593 acetylates certain lysine residues of the histone protein, therefore neutralizing them, disrupting histone-dna interaction, and allowing HeT-A to bind to telomeric DNA. CG7593 may or may not be involved in directing HeT-A to the telomeres. According to the hypothesis, I expect that CG7593 localizes in the nucleus and that in its absence, the entry of HeT-A into the nucleus would not be affected. The first steps in performing the experiments to test the hypothesis were verifications of HeT-A-GFP construct to be transfected into Schneider 2 cells, SD10812 EST from which CG7593 was amplified, and the created CG7593 dsrna. HeT-A-GFP construct verification SD10812 EST verification CG7593 dsrna verification HeT-A protein localization in CG7593 knock down Schneider 2 cell cultures Viability Analysis Results Example 2 RESULTS Pendulin and HeT-A were previously shown to interact in a yeast 2-hybrid screen. Pendulin encodes importin-_, which is involved in the translocation of proteins through the nuclear pore (Quimby and Corbett, 2001). The possible role of pendulin in the localization of HeT-A to the nucleus was studied via visualization of HeT-A with fluorescence microscopy and RNAi inhibition of pendulin translation in S2 cells. HeT-A Verification HeT-A Expression in S2 cells EST Verification Effect of RNAi on HeT-A expression in S2 cells Production and Transfection of GFP:Pendulin Construct Production and Transfection of Truncated GFP:Pendulin Deletion Derivatives Estimation of Cell Viability RT-PCR 6
7 Creating effective illustrations in Project Lab What s the Purpose of Illustrations? Condense large amounts of information Convince readers of your findings (by showing data quality). Focus attention on certain findings (e.g., relationship between values). Simplify complex findings. Promote thinking and discussion. Illustration Caveat: The most beautiful illustration cannot hide lousy content--content is key. 7
8 What are Some Pitfalls of Figures and Legends? Figures: Not mentioned in text. Textual data inconsistent with figures. Mislabeling. Symbols, data points, unreadable or cluttered. Ugliness (failure to get help from graphic designer). Legends: Reiterate results section Written in shorthand, abbreviated form rather than whole sentences. Choose the Most Effective Type of Illustration for a Given Goal To accomplish this: To present exact values, raw data, or data which do not fit into any simple pattern. To summarize trends, show interactions between two or more variables, relate data to constants, or emphasize an overall pattern rather than specific measurements. To dramatize differences or draw comparisons. To illustrate complex relationships, spatial configurations, pathways, processes, or interactions. To compare or contrast. Choose one of these: Table, list Line graph Bar graph Diagram Pictograph, pie chart, bar graph 8
9 Choose the Most Effective Type of Illustration (cont.) To accomplish this: Choose one of these: To show sequential processes. Flowchart To classify information. Table, list, pictograph To describe parts or circuits. Schematic To describe a process, organization, or Pictograph, flowchart, block model. diagram. To describe a change of state. Line graph, bar graph To describe proportions. Pie chart, bar graph To describe relationships. Table, line graph, block diagram To describe causation. Flowchart, pictograph To describe an entire object. Schematic, drawing, photograph To show the vertical or horizontal Flowchart, drawing tree, block hierarchy within an object, idea, or diagram. organization. Provide context for your illustrations in the body of your paper. Refer explicitly to the illustration (e.g., see Table 1, as shown in Figure 3. ) Tell the reader: How the graphic advances, supports, clarifies, or summarizes your discussion. Why it is important. What it means. How it supports your argument. 9
10 Gells and Cells: some figures from previous 7.17 projects Figure 1. Full length Tel2p digests showing the expected bands. Lanes 1 and 4 contain the 1kb ladder. Lanes 2 and 3 are Tel2p cut with BamHI. The gel shows the expected 7.5 kb and 1.8 kb bands. Gells and Cells: some figures from previous 7.17 projects Figure 4. Verification of Htt Constructs. The htt gene + psr11 expression construct was digested with EcoRI and BamHI. The expected DNA band sizes present in the gel electrophoresed digest were the size of the insert (mrfp-htt Q138 or mrfp-htt Q15) and the vector (psr11). Lane A has the expected band sizes for the mrfp-htt Q15 construct: 2.5Kb (insert DNA) and 5.1Kb (vector DNA). Lane B has the expected band sizes for the mrfp-htt Q138 construct: 2.8Kb (insert DNA) and 5.1Kb (vector DNA). 10
11 Gells and Cells: some figures from previous 7.17 projects kb 2kb 1.5kb 5kb Figure 2. NotI + XbaI digest verification of Mlp60A-YFP and Mlp60A-CFP constructs. Four clones each of Mlp60A-YFP and Mlp60A-CFP were analyzed by sequentially digesting with NotI and XbaI. Lanes 1 and 12 show 1kb DNA ladder. Lanes 2 and 7 show control digests of psr24 and psr25 respectively. Lanes 3 through 6 show experimental digests of Mlp60A-CFP clones 1-4 respectively, and lanes 8-11 show experimental digests of Mlp60A-YFP clones 1-4 respectively. The expected fragment sizes are 4.2kb + 1.7kb for the control digests, and 4.2kb + 2kb for the experimental digests. This analysis shows that all clones contain the correct construct. Gells and Cells: some figures from previous 7.17 projects Figure 8. Intracellular localization images of N-terminus Tel2 derivative and HetGag cotransfections. Left: HetGag-CFP dots form in the nucleus. Center: N-terminus Tel2- YFP mainly localizes in the nucleus. Aggregation at some of the Het dots is visible. Right: Overlay of N-terminus Tel2 and HetGag on a DIC and DAPI (red) image. Colocalization is seen as light-blue dots. 11
12 Gells and Cells: some figures from previous 7.17 projects a b c d e f g h Transfections with the R546A mutant. a,b: When transfected alone, the HetGagRA-YFP (green) was able form het-dots and -bodies. c,d: In many cells, however, the mutant protein formed nuclear clusters, like the HetGag MHR protein. e,f: In cells cotransfected with the R456A mutant (green) and TARTGag-CFP (red), the two proteins colocalized (yellow) efficiently. g,h: The mutant protein (green) and the HetGag deletion derivative, when cotransfected, co-localized to some extent to the nuclear het-dots and het-bodies (not shown). Some of the deletion derivative remained in the cytoplasm in loose clusters. Gells and Cells: some figures from previous 7.17 projects (b) (a) (c) Figure 11. Comparison of the localization of filamentous actin with that of Mlp60A- YFP. Cells were transfected with Mlp60A-YFP (green) and stained with both DAPI (blue) and rhodamine-phalloidin (red), which binds to filamentous actin. Transfected cell in (a) shows localization of Mlp60A-YFP throughout the cell but accumulating in the nucleus. Rhodamine-phalloidin staining in (b) shows filamentous actin localizing mostly at the periphery of the cell. Combination of image (a) and (b) in (c) shows that Mlp60A-YFP and filamentous actin do not colocalize. 12
7.17: Writing Materials & Methods Spring A Methods Section Exercise
 7.17: Writing Materials & Methods Spring 2006 Neal Lerner, nlerner@mit.edu, x2-2939 A Methods Section Exercise 1. Draw a relatively simple picture. 2. Write an account of how you drew that picture. 3.
7.17: Writing Materials & Methods Spring 2006 Neal Lerner, nlerner@mit.edu, x2-2939 A Methods Section Exercise 1. Draw a relatively simple picture. 2. Write an account of how you drew that picture. 3.
Photo credit: Theresa Walunas,
 Photo credit: Theresa Walunas, http://www.keyboardbiologist.net/knitblog/ 1 Photo: http://rufair.rutgers.edu/images/evaluation.jpg 2 3 Types of illustrations and their uses: -tables for specific numbers,
Photo credit: Theresa Walunas, http://www.keyboardbiologist.net/knitblog/ 1 Photo: http://rufair.rutgers.edu/images/evaluation.jpg 2 3 Types of illustrations and their uses: -tables for specific numbers,
Photo credit: Theresa Walunas,
 Photo credit: Theresa Walunas, http://www.keyboardbiologist.net/knitblog/ 1 The table on the left is useless because the only useful (i.e., nonzero) data are found in a much smaller range of temperatures.
Photo credit: Theresa Walunas, http://www.keyboardbiologist.net/knitblog/ 1 The table on the left is useless because the only useful (i.e., nonzero) data are found in a much smaller range of temperatures.
Schematic representation of the endogenous PALB2 locus and gene-disruption constructs
 Supplementary Figures Supplementary Figure 1. Generation of PALB2 -/- and BRCA2 -/- /PALB2 -/- DT40 cells. (A) Schematic representation of the endogenous PALB2 locus and gene-disruption constructs carrying
Supplementary Figures Supplementary Figure 1. Generation of PALB2 -/- and BRCA2 -/- /PALB2 -/- DT40 cells. (A) Schematic representation of the endogenous PALB2 locus and gene-disruption constructs carrying
SUPPLEMENTARY INFORMATION
 doi: 10.1038/nature06721 SUPPLEMENTARY INFORMATION. Supplemental Figure Legends Supplemental Figure 1 The distribution of hatx-1[82q] in Cos7 cells. Cos7 cells are co-transfected with hatx-1[82q]-gfp (green)
doi: 10.1038/nature06721 SUPPLEMENTARY INFORMATION. Supplemental Figure Legends Supplemental Figure 1 The distribution of hatx-1[82q] in Cos7 cells. Cos7 cells are co-transfected with hatx-1[82q]-gfp (green)
transcription and the promoter occupancy of Smad proteins. (A) HepG2 cells were co-transfected with the wwp-luc reporter, and FLAG-tagged FHL1,
 Supplementary Data Supplementary Figure Legends Supplementary Figure 1 FHL-mediated TGFβ-responsive reporter transcription and the promoter occupancy of Smad proteins. (A) HepG2 cells were co-transfected
Supplementary Data Supplementary Figure Legends Supplementary Figure 1 FHL-mediated TGFβ-responsive reporter transcription and the promoter occupancy of Smad proteins. (A) HepG2 cells were co-transfected
3.1 The Role of the Enzymatic Activity of Sir2 for Efficient. The Sir proteins are recruited to DNA by site-specific DNA-binding proteins and
 35 3 RESULTS 3.1 The Role of the Enzymatic Activity of Sir2 for Efficient Association of the SIR Complex with DNA The Sir proteins are recruited to DNA by site-specific DNA-binding proteins and subsequently
35 3 RESULTS 3.1 The Role of the Enzymatic Activity of Sir2 for Efficient Association of the SIR Complex with DNA The Sir proteins are recruited to DNA by site-specific DNA-binding proteins and subsequently
Figure S1. Verification of ihog Mutation by Protein Immunoblotting Figure S2. Verification of ihog and boi
 Figure S1. Verification of ihog Mutation by Protein Immunoblotting Extracts from S2R+ cells, embryos, and adults were analyzed by immunoprecipitation and immunoblotting with anti-ihog antibody. The Ihog
Figure S1. Verification of ihog Mutation by Protein Immunoblotting Extracts from S2R+ cells, embryos, and adults were analyzed by immunoprecipitation and immunoblotting with anti-ihog antibody. The Ihog
NAME TA SEC Problem Set 4 FRIDAY October 15, Answers to this problem set must be inserted into the box outside
 MIT Biology Department 7.012: Introductory Biology - Fall 2004 Instructors: Professor Eric Lander, Professor Robert A. Weinberg, Dr. Claudette Gardel NAME TA SEC 7.012 Problem Set 4 FRIDAY October 15,
MIT Biology Department 7.012: Introductory Biology - Fall 2004 Instructors: Professor Eric Lander, Professor Robert A. Weinberg, Dr. Claudette Gardel NAME TA SEC 7.012 Problem Set 4 FRIDAY October 15,
Figure S1: Insert sequences for GR1, GR2, Qc3c_AgeI and GR3 vectors
 Figure S1: Insert sequences for GR1, GR2, Qc3c_AgeI and GR3 vectors A Vector GR1 SacI BamHI CTCTGGCTAACTAGGC Insert 5/7nt - G TCGAGAGACCGATTGATCCG Insert 5/7nt - CCTAG 1 G CAACAACAACAACAACAACAACAACAACAACAACAACAACAACAACAACAACAACAACAACAAC
Figure S1: Insert sequences for GR1, GR2, Qc3c_AgeI and GR3 vectors A Vector GR1 SacI BamHI CTCTGGCTAACTAGGC Insert 5/7nt - G TCGAGAGACCGATTGATCCG Insert 5/7nt - CCTAG 1 G CAACAACAACAACAACAACAACAACAACAACAACAACAACAACAACAACAACAACAACAACAAC
Supplementary Figure S1. Immunodetection of full-length XA21 and the XA21 C-terminal cleavage product.
 Supplementary Information Supplementary Figure S1. Immunodetection of full-length XA21 and the XA21 C-terminal cleavage product. Total protein extracted from Kitaake wild type and rice plants carrying
Supplementary Information Supplementary Figure S1. Immunodetection of full-length XA21 and the XA21 C-terminal cleavage product. Total protein extracted from Kitaake wild type and rice plants carrying
Lab 9 Restriction Enzyme Analysis
 Name Assignment # Lab 9 Restriction Enzyme Analysis http://www.phschool.com/science/biology_place/labbench/lab6/concepts2.html 1) Define restriction enzyme 2) Define recognition sequence 3) Label the images
Name Assignment # Lab 9 Restriction Enzyme Analysis http://www.phschool.com/science/biology_place/labbench/lab6/concepts2.html 1) Define restriction enzyme 2) Define recognition sequence 3) Label the images
Sarker et al. Supplementary Material. Subcellular Fractionation
 Supplementary Material Subcellular Fractionation Transfected 293T cells were harvested with phosphate buffered saline (PBS) and centrifuged at 2000 rpm (500g) for 3 min. The pellet was washed, re-centrifuged
Supplementary Material Subcellular Fractionation Transfected 293T cells were harvested with phosphate buffered saline (PBS) and centrifuged at 2000 rpm (500g) for 3 min. The pellet was washed, re-centrifuged
hnrnp D/AUF1 Rabbit IgG hnrnp M
 Mouse IgG Goat IgG Rabbit IgG Mouse IgG hnrnp F Goat IgG Mouse IgG Kb 6 4 3 2 15 5 Supplementary Figure S1. In vivo binding of TERRA-bound RBPs to target RNAs. Immunoprecipitation (IP) assay using 3 mg
Mouse IgG Goat IgG Rabbit IgG Mouse IgG hnrnp F Goat IgG Mouse IgG Kb 6 4 3 2 15 5 Supplementary Figure S1. In vivo binding of TERRA-bound RBPs to target RNAs. Immunoprecipitation (IP) assay using 3 mg
Chapter 10 (Part II) Gene Isolation and Manipulation
 Biology 234 J. G. Doheny Chapter 10 (Part II) Gene Isolation and Manipulation Practice Questions: Answer the following questions with one or two sentences. 1. What does PCR stand for? 2. What does the
Biology 234 J. G. Doheny Chapter 10 (Part II) Gene Isolation and Manipulation Practice Questions: Answer the following questions with one or two sentences. 1. What does PCR stand for? 2. What does the
Revision Checklist for Science Signaling Research Manuscripts: Data Requirements and Style Guidelines
 Revision Checklist for Science Signaling Research Manuscripts: Data Requirements and Style Guidelines Further information can be found at: http://stke.sciencemag.org/sites/default/files/researcharticlerevmsinstructions_0.pdf.
Revision Checklist for Science Signaling Research Manuscripts: Data Requirements and Style Guidelines Further information can be found at: http://stke.sciencemag.org/sites/default/files/researcharticlerevmsinstructions_0.pdf.
Genome Sequence Assembly
 Genome Sequence Assembly Learning Goals: Introduce the field of bioinformatics Familiarize the student with performing sequence alignments Understand the assembly process in genome sequencing Introduction:
Genome Sequence Assembly Learning Goals: Introduce the field of bioinformatics Familiarize the student with performing sequence alignments Understand the assembly process in genome sequencing Introduction:
The Human Protein PRR14 Tethers Heterochromatin to the Nuclear Lamina During Interphase and Mitotic Exit
 Cell Reports, Volume 5 Supplemental Information The Human Protein PRR14 Tethers Heterochromatin to the Nuclear Lamina During Interphase and Mitotic Exit Andrey Poleshko, Katelyn M. Mansfield, Caroline
Cell Reports, Volume 5 Supplemental Information The Human Protein PRR14 Tethers Heterochromatin to the Nuclear Lamina During Interphase and Mitotic Exit Andrey Poleshko, Katelyn M. Mansfield, Caroline
PHT1;2-CFP YFP-PHF + PHT1;2-CFP YFP-PHF
 YFP-PHF1 CFP-PHT1;2 PHT1;2-CFP YFP-PHF + PHT1;2-CFP YFP-PHF + CFP-PHT1;2 Negative control!-gfp Supplemental Figure 1: PHT1;2 accumulation is PHF1 dependent. Immunoblot analysis on total protein extract
YFP-PHF1 CFP-PHT1;2 PHT1;2-CFP YFP-PHF + PHT1;2-CFP YFP-PHF + CFP-PHT1;2 Negative control!-gfp Supplemental Figure 1: PHT1;2 accumulation is PHF1 dependent. Immunoblot analysis on total protein extract
To investigate the heredity of the WFP gene, we selected plants that were homozygous
 Supplementary information Supplementary Note ST-12 WFP allele is semi-dominant To investigate the heredity of the WFP gene, we selected plants that were homozygous for chromosome 1 of Nipponbare and heterozygous
Supplementary information Supplementary Note ST-12 WFP allele is semi-dominant To investigate the heredity of the WFP gene, we selected plants that were homozygous for chromosome 1 of Nipponbare and heterozygous
Supplemental Figure 1 A
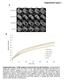 Supplemental Figure A prebleach postbleach 2 min 6 min 3 min mh2a.-gfp mh2a.2-gfp mh2a2-gfp GFP-H2A..9 Relative Intensity.8.7.6.5 mh2a. GFP n=8.4 mh2a.2 GFP n=4.3 mh2a2 GFP n=2.2 GFP H2A n=24. GFP n=7.
Supplemental Figure A prebleach postbleach 2 min 6 min 3 min mh2a.-gfp mh2a.2-gfp mh2a2-gfp GFP-H2A..9 Relative Intensity.8.7.6.5 mh2a. GFP n=8.4 mh2a.2 GFP n=4.3 mh2a2 GFP n=2.2 GFP H2A n=24. GFP n=7.
7.02/ RECOMBINANT DNA METHODS EXAM KEY
 MIT Department of Biology 7.02 Experimental Biology & Communication, Spring 2005 7.02/10.702 RECOMBINANT DNA METHODS EXAM KEY Regrade requests are due to the instructor in the 7.02 teaching lab by the
MIT Department of Biology 7.02 Experimental Biology & Communication, Spring 2005 7.02/10.702 RECOMBINANT DNA METHODS EXAM KEY Regrade requests are due to the instructor in the 7.02 teaching lab by the
Supplemental Data. Farmer et al. (2010) Plant Cell /tpc
 Supplemental Figure 1. Amino acid sequence comparison of RAD23 proteins. Identical and similar residues are shown in the black and gray boxes, respectively. Dots denote gaps. The sequence of plant Ub is
Supplemental Figure 1. Amino acid sequence comparison of RAD23 proteins. Identical and similar residues are shown in the black and gray boxes, respectively. Dots denote gaps. The sequence of plant Ub is
MIT Department of Biology 7.013: Introductory Biology - Spring 2005 Instructors: Professor Hazel Sive, Professor Tyler Jacks, Dr.
 MIT Department of Biology 7.01: Introductory Biology - Spring 2005 Instructors: Professor Hazel Sive, Professor Tyler Jacks, Dr. Claudette Gardel iv) Would Xba I be useful for cloning? Why or why not?
MIT Department of Biology 7.01: Introductory Biology - Spring 2005 Instructors: Professor Hazel Sive, Professor Tyler Jacks, Dr. Claudette Gardel iv) Would Xba I be useful for cloning? Why or why not?
Supplementary Methods
 Supplementary Methods Reverse transcribed Quantitative PCR. Total RNA was isolated from bone marrow derived macrophages using RNeasy Mini Kit (Qiagen), DNase-treated (Promega RQ1), and reverse transcribed
Supplementary Methods Reverse transcribed Quantitative PCR. Total RNA was isolated from bone marrow derived macrophages using RNeasy Mini Kit (Qiagen), DNase-treated (Promega RQ1), and reverse transcribed
SUPPLEMENTARY EXPEMENTAL PROCEDURES
 SUPPLEMENTARY EXPEMENTAL PROCEDURES Plasmids- Total RNAs were extracted from HeLaS3 cells and reverse-transcribed using Superscript III Reverse Transcriptase (Invitrogen) to obtain DNA template for the
SUPPLEMENTARY EXPEMENTAL PROCEDURES Plasmids- Total RNAs were extracted from HeLaS3 cells and reverse-transcribed using Superscript III Reverse Transcriptase (Invitrogen) to obtain DNA template for the
Writing Your Honors Thesis
 Writing Your Honors Thesis Your thesis should be in the common scientific paper format using FIVE separate sections Abstract Introduction Materials and Methods Results Discussion Keep in mind----your thesis
Writing Your Honors Thesis Your thesis should be in the common scientific paper format using FIVE separate sections Abstract Introduction Materials and Methods Results Discussion Keep in mind----your thesis
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION Legends for Supplementary Tables. Supplementary Table 1. An excel file containing primary screen data. Worksheet 1, Normalized quantification data from a duplicated screen: valid
SUPPLEMENTARY INFORMATION Legends for Supplementary Tables. Supplementary Table 1. An excel file containing primary screen data. Worksheet 1, Normalized quantification data from a duplicated screen: valid
Figure S1. Figure S2. Figure S3 HB Anti-FSP27 (COOH-terminal peptide) Ab. Anti-GST-FSP27(45-127) Ab.
 / 36B4 mrna ratio Figure S1 * 2. 1.6 1.2.8 *.4 control TNFα BRL49653 Figure S2 Su bw AT p iw Anti- (COOH-terminal peptide) Ab Blot : Anti-GST-(45-127) Ab β-actin Figure S3 HB2 HW AT BA T Figure S4 A TAG
/ 36B4 mrna ratio Figure S1 * 2. 1.6 1.2.8 *.4 control TNFα BRL49653 Figure S2 Su bw AT p iw Anti- (COOH-terminal peptide) Ab Blot : Anti-GST-(45-127) Ab β-actin Figure S3 HB2 HW AT BA T Figure S4 A TAG
Supplemental Figure 1 HDA18 has an HDAC domain and therefore has concentration dependent and TSA inhibited histone deacetylase activity.
 Supplemental Figure 1 HDA18 has an HDAC domain and therefore has concentration dependent and TSA inhibited histone deacetylase activity. (A) Amino acid alignment of HDA5, HDA15 and HDA18. The blue line
Supplemental Figure 1 HDA18 has an HDAC domain and therefore has concentration dependent and TSA inhibited histone deacetylase activity. (A) Amino acid alignment of HDA5, HDA15 and HDA18. The blue line
 DOI: 10.1038/ncb3259 A Ismail et al. Supplementary Figure 1 B 60000 45000 SSC 30000 15000 Live cells 0 0 15000 30000 45000 60000 FSC- PARR 60000 45000 PARR Width 30000 FSC- 15000 Single cells 0 0 15000
DOI: 10.1038/ncb3259 A Ismail et al. Supplementary Figure 1 B 60000 45000 SSC 30000 15000 Live cells 0 0 15000 30000 45000 60000 FSC- PARR 60000 45000 PARR Width 30000 FSC- 15000 Single cells 0 0 15000
b alternative classical none
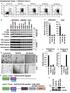 Supplementary Figure. 1: Related to Figure.1 a d e b alternative classical none NIK P-IkBa Total IkBa Tubulin P52 (Lighter) P52 (Darker) RelB (Lighter) RelB (Darker) HDAC1 Control-Sh RelB-Sh NF-kB2-Sh
Supplementary Figure. 1: Related to Figure.1 a d e b alternative classical none NIK P-IkBa Total IkBa Tubulin P52 (Lighter) P52 (Darker) RelB (Lighter) RelB (Darker) HDAC1 Control-Sh RelB-Sh NF-kB2-Sh
Engineering splicing factors with designed specificities
 nature methods Engineering splicing factors with designed specificities Yang Wang, Cheom-Gil Cheong, Traci M Tanaka Hall & Zefeng Wang Supplementary figures and text: Supplementary Figure 1 Supplementary
nature methods Engineering splicing factors with designed specificities Yang Wang, Cheom-Gil Cheong, Traci M Tanaka Hall & Zefeng Wang Supplementary figures and text: Supplementary Figure 1 Supplementary
Supplemental Data. Na Xu et al. (2016). Plant Cell /tpc
 Supplemental Figure 1. The weak fluorescence phenotype is not caused by the mutation in At3g60240. (A) A mutation mapped to the gene At3g60240. Map-based cloning strategy was used to map the mutated site
Supplemental Figure 1. The weak fluorescence phenotype is not caused by the mutation in At3g60240. (A) A mutation mapped to the gene At3g60240. Map-based cloning strategy was used to map the mutated site
Fig. S1. Effect of p120-catenin overexpression on the interaction of SCUBE2 with E-cadherin. The expression plasmid encoding FLAG.
 Fig. S1. Effect of p120-catenin overexpression on the interaction of SCUBE2 with E-cadherin. The expression plasmid encoding FLAG.SCUBE2, E-cadherin.Myc, or HA.p120-catenin was transfected in a combination
Fig. S1. Effect of p120-catenin overexpression on the interaction of SCUBE2 with E-cadherin. The expression plasmid encoding FLAG.SCUBE2, E-cadherin.Myc, or HA.p120-catenin was transfected in a combination
Somatic Primary pirna Biogenesis Driven by cis-acting RNA Elements and Trans-Acting Yb
 Cell Reports Supplemental Information Somatic Primary pirna Biogenesis Driven by cis-acting RNA Elements and Trans-Acting Yb Hirotsugu Ishizu, Yuka W. Iwasaki, Shigeki Hirakata, Haruka Ozaki, Wataru Iwasaki,
Cell Reports Supplemental Information Somatic Primary pirna Biogenesis Driven by cis-acting RNA Elements and Trans-Acting Yb Hirotsugu Ishizu, Yuka W. Iwasaki, Shigeki Hirakata, Haruka Ozaki, Wataru Iwasaki,
Grb2-Mediated Alteration in the Trafficking of AβPP: Insights from Grb2-AICD Interaction
 Journal of Alzheimer s Disease 20 (2010) 1 9 1 IOS Press Supplementary Material Grb2-Mediated Alteration in the Trafficking of AβPP: Insights from Grb2-AICD Interaction Mithu Raychaudhuri and Debashis
Journal of Alzheimer s Disease 20 (2010) 1 9 1 IOS Press Supplementary Material Grb2-Mediated Alteration in the Trafficking of AβPP: Insights from Grb2-AICD Interaction Mithu Raychaudhuri and Debashis
Rotation Report Sample Version 3. Due Date: August 9, Analysis of the Guanine Nucleotide Exchange Activity of the S. cerevisiae Ats1 Protein
 Rotation Report Sample Version 3 Due Date: August 9, 1998 Analysis of the Guanine Nucleotide Exchange Activity of the S. cerevisiae Ats1 Protein Anita H. Corbett Advisor: Amy Jones Rotation 1 Abstract:
Rotation Report Sample Version 3 Due Date: August 9, 1998 Analysis of the Guanine Nucleotide Exchange Activity of the S. cerevisiae Ats1 Protein Anita H. Corbett Advisor: Amy Jones Rotation 1 Abstract:
10X ligation buffer ligase 1 vector DNA insert DNA H 2 O. 10 µl Total Volume. 10X ligation buffer ligase 1 vector DNA insert DNA
 Biol/Chem 475 S07 Study problems for quiz 1 See also questions posed in lab handouts including ligase handout Answers to questions 1&2 included at the end of this document. 1. You plan to clone a 1.0 kb
Biol/Chem 475 S07 Study problems for quiz 1 See also questions posed in lab handouts including ligase handout Answers to questions 1&2 included at the end of this document. 1. You plan to clone a 1.0 kb
Supplementary Figure 1. (a) The qrt-pcr for lnc-2, lnc-6 and lnc-7 RNA level in DU145, 22Rv1, wild type HCT116 and HCT116 Dicer ex5 cells transfected
 Supplementary Figure 1. (a) The qrt-pcr for lnc-2, lnc-6 and lnc-7 RNA level in DU145, 22Rv1, wild type HCT116 and HCT116 Dicer ex5 cells transfected with the sirna against lnc-2, lnc-6, lnc-7, and the
Supplementary Figure 1. (a) The qrt-pcr for lnc-2, lnc-6 and lnc-7 RNA level in DU145, 22Rv1, wild type HCT116 and HCT116 Dicer ex5 cells transfected with the sirna against lnc-2, lnc-6, lnc-7, and the
Supplemental Figure legends Figure S1. (A) (B) (C) (D) Figure S2. Figure S3. (A-E) Figure S4. Figure S5. (A, C, E, G, I) (B, D, F, H, Figure S6.
 Supplemental Figure legends Figure S1. Map-based cloning and complementation testing for ZOP1. (A) ZOP1 was mapped to a ~273-kb interval on Chromosome 1. In the interval, a single-nucleotide G to A substitution
Supplemental Figure legends Figure S1. Map-based cloning and complementation testing for ZOP1. (A) ZOP1 was mapped to a ~273-kb interval on Chromosome 1. In the interval, a single-nucleotide G to A substitution
1. A brief overview of sequencing biochemistry
 Supplementary reading materials on Genome sequencing (optional) The materials are from Mark Blaxter s lecture notes on Sequencing strategies and Primary Analysis 1. A brief overview of sequencing biochemistry
Supplementary reading materials on Genome sequencing (optional) The materials are from Mark Blaxter s lecture notes on Sequencing strategies and Primary Analysis 1. A brief overview of sequencing biochemistry
Revised: RG-RV2 by Fukuhara et al.
 Supplemental Figure 1 The generation of Spns2 conditional knockout mice. (A) Schematic representation of the wild type Spns2 locus (Spns2 + ), the targeted allele, the floxed allele (Spns2 f ) and the
Supplemental Figure 1 The generation of Spns2 conditional knockout mice. (A) Schematic representation of the wild type Spns2 locus (Spns2 + ), the targeted allele, the floxed allele (Spns2 f ) and the
MOLECULAR BIOLOGY OF EUKARYOTES 2016 SYLLABUS
 03-442 Lectures: MWF 9:30-10:20 a.m. Doherty Hall 2105 03-742 Advanced Discussion Section: Time and place to be announced Probably Mon 4-6 p.m. or 6-8p.m.? Once we establish who is taking the advanced
03-442 Lectures: MWF 9:30-10:20 a.m. Doherty Hall 2105 03-742 Advanced Discussion Section: Time and place to be announced Probably Mon 4-6 p.m. or 6-8p.m.? Once we establish who is taking the advanced
Supplementary Figure 1 Collision-induced dissociation (CID) mass spectra of peptides from PPK1, PPK2, PPK3 and PPK4 respectively.
 Supplementary Figure 1 lision-induced dissociation (CID) mass spectra of peptides from PPK1, PPK, PPK3 and PPK respectively. % of nuclei with signal / field a 5 c ppif3:gus pppk1:gus 0 35 30 5 0 15 10
Supplementary Figure 1 lision-induced dissociation (CID) mass spectra of peptides from PPK1, PPK, PPK3 and PPK respectively. % of nuclei with signal / field a 5 c ppif3:gus pppk1:gus 0 35 30 5 0 15 10
7.012 Problem Set 5. Question 1
 Name Section 7.012 Problem Set 5 Question 1 While studying the problem of infertility, you attempt to isolate a hypothetical rabbit gene that accounts for the prolific reproduction of rabbits. After much
Name Section 7.012 Problem Set 5 Question 1 While studying the problem of infertility, you attempt to isolate a hypothetical rabbit gene that accounts for the prolific reproduction of rabbits. After much
MID-TERM EXAMINATION
 PLNT3140 INTRODUCTORY CYTOGENETICS MID-TERM EXAMINATION 1 p.m. to 2:15 p.m. Thursday, October 18, 2012 Answer any combination of questions totalling to exactly 100 points. If you answer questions totalling
PLNT3140 INTRODUCTORY CYTOGENETICS MID-TERM EXAMINATION 1 p.m. to 2:15 p.m. Thursday, October 18, 2012 Answer any combination of questions totalling to exactly 100 points. If you answer questions totalling
7.06 Cell Biology EXAM #2 March 20, 2003
 7.06 Cell Biology EXAM #2 March 20, 2003 This is an open book exam, and you are allowed access to books, a calculator, and notes but not computers or any other types of electronic devices. Please write
7.06 Cell Biology EXAM #2 March 20, 2003 This is an open book exam, and you are allowed access to books, a calculator, and notes but not computers or any other types of electronic devices. Please write
CD93 and dystroglycan cooperation in human endothelial cell adhesion and migration
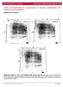 /, Supplementary Advance Publications Materials 2016 CD93 and dystroglycan cooperation in human endothelial cell adhesion and migration Supplementary Materials Supplementary Figure S1: In ECs CD93 silencing
/, Supplementary Advance Publications Materials 2016 CD93 and dystroglycan cooperation in human endothelial cell adhesion and migration Supplementary Materials Supplementary Figure S1: In ECs CD93 silencing
Fig. S1. CrgA intracellular levels in M. tuberculosis. Ten and twenty micrograms of
 Supplementary data Fig. S1. CrgA intracellular levels in M. tuberculosis. Ten and twenty micrograms of cell free protein lysates from WT M. tuberculosis (Rv) together with various known concentrations
Supplementary data Fig. S1. CrgA intracellular levels in M. tuberculosis. Ten and twenty micrograms of cell free protein lysates from WT M. tuberculosis (Rv) together with various known concentrations
Supplemental Figure 1 Human REEP family of proteins can be divided into two distinct subfamilies. Residues (single letter amino acid code) identical
 Supplemental Figure Human REEP family of proteins can be divided into two distinct subfamilies. Residues (single letter amino acid code) identical in all six REEPs are highlighted in green. Additional
Supplemental Figure Human REEP family of proteins can be divided into two distinct subfamilies. Residues (single letter amino acid code) identical in all six REEPs are highlighted in green. Additional
T H E J O U R N A L O F C E L L B I O L O G Y
 Supplemental material Thompson et al., http://www.jcb.org/cgi/content/full/jcb.200909067/dc1 T H E J O U R N A L O F C E L L B I O L O G Y Figure S1. Modification-specific antibodies do not detect unmodified
Supplemental material Thompson et al., http://www.jcb.org/cgi/content/full/jcb.200909067/dc1 T H E J O U R N A L O F C E L L B I O L O G Y Figure S1. Modification-specific antibodies do not detect unmodified
Understanding the Cellular Mechanism of the Excess Microsporocytes I (EMSI) Gene. Andrew ElBardissi, The Pennsylvania State University
 Understanding the Cellular Mechanism of the Excess Microsporocytes I (EMSI) Gene Andrew ElBardissi, The Pennsylvania State University Abstract: Hong Ma, The Pennsylvania State University The Excess Microsporocytes
Understanding the Cellular Mechanism of the Excess Microsporocytes I (EMSI) Gene Andrew ElBardissi, The Pennsylvania State University Abstract: Hong Ma, The Pennsylvania State University The Excess Microsporocytes
Nature Methods: doi: /nmeth Supplementary Figure 1. Construction of a sensitive TetR mediated auxotrophic off-switch.
 Supplementary Figure 1 Construction of a sensitive TetR mediated auxotrophic off-switch. A Production of the Tet repressor in yeast when conjugated to either the LexA4 or LexA8 promoter DNA binding sequences.
Supplementary Figure 1 Construction of a sensitive TetR mediated auxotrophic off-switch. A Production of the Tet repressor in yeast when conjugated to either the LexA4 or LexA8 promoter DNA binding sequences.
The drawing of RNA and cdna is worth 3 points, if there is no second strand of cdna or no oligo dc or dg linker added, 1 point will be deducted.
 1. a) The 3 end of mrna usually ends up copied in a cdna because the first strand of cdna is synthesized by reverse transcriptase using oligo dt as the primer. The enzyme will fall off after a while resulting
1. a) The 3 end of mrna usually ends up copied in a cdna because the first strand of cdna is synthesized by reverse transcriptase using oligo dt as the primer. The enzyme will fall off after a while resulting
MCB 102 University of California, Berkeley August 11 13, Problem Set 8
 MCB 102 University of California, Berkeley August 11 13, 2009 Isabelle Philipp Handout Problem Set 8 The answer key will be posted by Tuesday August 11. Try to solve the problem sets always first without
MCB 102 University of California, Berkeley August 11 13, 2009 Isabelle Philipp Handout Problem Set 8 The answer key will be posted by Tuesday August 11. Try to solve the problem sets always first without
Supplementary Materials
 Supplementary Materials Construction of Synthetic Nucleoli in Human Cells Reveals How a Major Functional Nuclear Domain is Formed and Propagated Through Cell Divisision Authors: Alice Grob, Christine Colleran
Supplementary Materials Construction of Synthetic Nucleoli in Human Cells Reveals How a Major Functional Nuclear Domain is Formed and Propagated Through Cell Divisision Authors: Alice Grob, Christine Colleran
Value Correct Answer Feedback. Student Response. A. Dicer enzyme. complex. C. the Dicer-RISC complex D. none of the above
 1 RNA mediated interference is a post-transcriptional gene silencing mechanism Which component of the RNAi pathway have been implicated in cleavage of the target mrna? A Dicer enzyme B the RISC-siRNA complex
1 RNA mediated interference is a post-transcriptional gene silencing mechanism Which component of the RNAi pathway have been implicated in cleavage of the target mrna? A Dicer enzyme B the RISC-siRNA complex
Supplementary Figure 1. Nature Structural & Molecular Biology: doi: /nsmb.3494
 Supplementary Figure 1 Pol structure-function analysis (a) Inactivating polymerase and helicase mutations do not alter the stability of Pol. Flag epitopes were introduced using CRISPR/Cas9 gene targeting
Supplementary Figure 1 Pol structure-function analysis (a) Inactivating polymerase and helicase mutations do not alter the stability of Pol. Flag epitopes were introduced using CRISPR/Cas9 gene targeting
Supplemental materials
 Supplemental materials Materials and methods for supplemental figures Yeast two-hybrid assays TAP46-PP2Ac interactions I. The TAP46 was used as the bait and the full-length cdnas of the five C subunits
Supplemental materials Materials and methods for supplemental figures Yeast two-hybrid assays TAP46-PP2Ac interactions I. The TAP46 was used as the bait and the full-length cdnas of the five C subunits
Antisense RNA Targeting the First Periplasmic Domain of YidC in Escherichia coli Appears to Induce Filamentation but Does Not Affect Cell Viability
 Antisense RNA Targeting the First Periplasmic Domain of YidC in Escherichia coli Appears to Induce Filamentation but Does Not Affect Cell Viability Riaaz Lalani, Nathaniel Susilo, Elisa Xiao, Andrea Xu
Antisense RNA Targeting the First Periplasmic Domain of YidC in Escherichia coli Appears to Induce Filamentation but Does Not Affect Cell Viability Riaaz Lalani, Nathaniel Susilo, Elisa Xiao, Andrea Xu
AD BD TOC1. Supplementary Figure 1: Yeast two-hybrid assays showing the interaction between
 AD X BD TOC1 AD BD X PIFΔAD PIF TOC1 TOC1 PIFΔAD PIF N TOC1 TOC1 C1 PIFΔAD PIF C1 TOC1 TOC1 C PIFΔAD PIF C TOC1 Supplementary Figure 1: Yeast two-hybrid assays showing the interaction between PIF and TOC1
AD X BD TOC1 AD BD X PIFΔAD PIF TOC1 TOC1 PIFΔAD PIF N TOC1 TOC1 C1 PIFΔAD PIF C1 TOC1 TOC1 C PIFΔAD PIF C TOC1 Supplementary Figure 1: Yeast two-hybrid assays showing the interaction between PIF and TOC1
 Name: Date: Virtual Student Guide http://www.phschool.com/science/biology_place/labbench/index.html AP Biology Laboratory 6 Part II DNA Electrophoresis Introduction In this laboratory you will use some
Name: Date: Virtual Student Guide http://www.phschool.com/science/biology_place/labbench/index.html AP Biology Laboratory 6 Part II DNA Electrophoresis Introduction In this laboratory you will use some
Supplementary Information
 Supplementary Information Supplementary Figure 1: Over-expression of CD300f in NIH3T3 cells enhances their capacity to phagocytize AC. (a) NIH3T3 cells were stably transduced by EV, CD300f WT or CD300f
Supplementary Information Supplementary Figure 1: Over-expression of CD300f in NIH3T3 cells enhances their capacity to phagocytize AC. (a) NIH3T3 cells were stably transduced by EV, CD300f WT or CD300f
KEY CONCEPTS AND PROCESS SKILLS. 1. Blood types can be used as evidence about identity and about family relationships.
 Evidence from DNA 40- to 1 2 50-minute sessions 69 M O D E L I N G ACTIVITY OVERVIEW SUMMARY Students learn how DNA fingerprinting is done by performing a simulation of the process used to generate different
Evidence from DNA 40- to 1 2 50-minute sessions 69 M O D E L I N G ACTIVITY OVERVIEW SUMMARY Students learn how DNA fingerprinting is done by performing a simulation of the process used to generate different
SUPPLEMENTARY INFORMATION
 doi:.38/nature899 Supplementary Figure Suzuki et al. a c p7 -/- / WT ratio (+)/(-) p7 -/- / WT ratio Log X 3. Fold change by treatment ( (+)/(-)) Log X.5 3-3. -. b Fold change by treatment ( (+)/(-)) 8
doi:.38/nature899 Supplementary Figure Suzuki et al. a c p7 -/- / WT ratio (+)/(-) p7 -/- / WT ratio Log X 3. Fold change by treatment ( (+)/(-)) Log X.5 3-3. -. b Fold change by treatment ( (+)/(-)) 8
Construction of plant complementation vector and generation of transgenic plants
 MATERIAL S AND METHODS Plant materials and growth conditions Arabidopsis ecotype Columbia (Col0) was used for this study. SALK_072009, SALK_076309, and SALK_027645 were obtained from the Arabidopsis Biological
MATERIAL S AND METHODS Plant materials and growth conditions Arabidopsis ecotype Columbia (Col0) was used for this study. SALK_072009, SALK_076309, and SALK_027645 were obtained from the Arabidopsis Biological
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/4/9/eaat5401/dc1 Supplementary Materials for GLK-IKKβ signaling induces dimerization and translocation of the AhR-RORγt complex in IL-17A induction and autoimmune
advances.sciencemag.org/cgi/content/full/4/9/eaat5401/dc1 Supplementary Materials for GLK-IKKβ signaling induces dimerization and translocation of the AhR-RORγt complex in IL-17A induction and autoimmune
Loss of Cul3 in Primary Fibroblasts
 PSU McNair Scholars Online Journal Volume 5 Issue 1 Humans Being: People, Places, Perspectives and Processes Article 15 2011 Loss of Cul3 in Primary Fibroblasts Paula Hanna Portland State University Let
PSU McNair Scholars Online Journal Volume 5 Issue 1 Humans Being: People, Places, Perspectives and Processes Article 15 2011 Loss of Cul3 in Primary Fibroblasts Paula Hanna Portland State University Let
Supplementary data. sienigma. F-Enigma F-EnigmaSM. a-p53
 Supplementary data Supplemental Figure 1 A sienigma #2 sienigma sicontrol a-enigma - + ++ - - - - - - + ++ - - - - - - ++ B sienigma F-Enigma F-EnigmaSM a-flag HLK3 cells - - - + ++ + ++ - + - + + - -
Supplementary data Supplemental Figure 1 A sienigma #2 sienigma sicontrol a-enigma - + ++ - - - - - - + ++ - - - - - - ++ B sienigma F-Enigma F-EnigmaSM a-flag HLK3 cells - - - + ++ + ++ - + - + + - -
2. (So) get (fragments with gene) R / required gene. Accept: allele for gene / same gene 2
 M.(a). Cut (DNA) at same (base) sequence / (recognition) sequence; Accept: cut DNA at same place. (So) get (fragments with gene) R / required gene. Accept: allele for gene / same gene (b). Each has / they
M.(a). Cut (DNA) at same (base) sequence / (recognition) sequence; Accept: cut DNA at same place. (So) get (fragments with gene) R / required gene. Accept: allele for gene / same gene (b). Each has / they
Biology 201 (Genetics) Exam #3 120 points 20 November Read the question carefully before answering. Think before you write.
 Name KEY Section Biology 201 (Genetics) Exam #3 120 points 20 November 2006 Read the question carefully before answering. Think before you write. You will have up to 50 minutes to take this exam. After
Name KEY Section Biology 201 (Genetics) Exam #3 120 points 20 November 2006 Read the question carefully before answering. Think before you write. You will have up to 50 minutes to take this exam. After
oligonucleotide primers listed in Supplementary Table 2 and cloned into ppcr-script (Stratagene) before digestion
 Supplementary Methods. Construction of BiFC vectors The full-length ARC3 cdna, ARC3 1-1794 and ARC3 1-1083 were amplified from ppcr-script/arc3 using the oligonucleotide primers listed in Supplementary
Supplementary Methods. Construction of BiFC vectors The full-length ARC3 cdna, ARC3 1-1794 and ARC3 1-1083 were amplified from ppcr-script/arc3 using the oligonucleotide primers listed in Supplementary
SBI4U Culminating Activity Part 1: Genetic Engineering of a Recombinant Plasmid Name:
 SBI4U Culminating Activity Part 1: Genetic Engineering of a Recombinant Plasmid Name: Background Read through The Major Steps of Cloning of DNA on page 290 and examine the figure on page 291. This is the
SBI4U Culminating Activity Part 1: Genetic Engineering of a Recombinant Plasmid Name: Background Read through The Major Steps of Cloning of DNA on page 290 and examine the figure on page 291. This is the
SUPPLEMENTARY INFORMATION
 Secondary mutations as a mechanism of cisplatin resistance in BRCA2-mutated cancers Wataru Sakai, Elizabeth M. Swisher, Beth Y. Karlan, Mukesh K. Agarwal, Jake Higgins, Cynthia Friedman, Emily Villegas,
Secondary mutations as a mechanism of cisplatin resistance in BRCA2-mutated cancers Wataru Sakai, Elizabeth M. Swisher, Beth Y. Karlan, Mukesh K. Agarwal, Jake Higgins, Cynthia Friedman, Emily Villegas,
Molecular Techniques. 3 Goals in Molecular Biology. Nucleic Acids: DNA and RNA. Disclaimer Nucleic Acids Proteins
 Molecular Techniques Disclaimer Nucleic Acids Proteins Houpt, CMN, 9-30-11 3 Goals in Molecular Biology Identify All nucleic acids (and proteins) are chemically identical in aggregate - need to identify
Molecular Techniques Disclaimer Nucleic Acids Proteins Houpt, CMN, 9-30-11 3 Goals in Molecular Biology Identify All nucleic acids (and proteins) are chemically identical in aggregate - need to identify
Supporting Information. Sequence Independent Cloning and Posttranslational. Enzymatic Ligation
 Supporting Information Sequence Independent Cloning and Posttranslational Modification of Repetitive Protein Polymers through Sortase and Sfp-mediated Enzymatic Ligation Wolfgang Ott,,, Thomas Nicolaus,,
Supporting Information Sequence Independent Cloning and Posttranslational Modification of Repetitive Protein Polymers through Sortase and Sfp-mediated Enzymatic Ligation Wolfgang Ott,,, Thomas Nicolaus,,
- 1 - Supplemental Data
 - 1-1 Supplemental Data 2 3 4 5 6 7 8 9 Supplemental Figure S1. Differential expression of AtPIP Genes in DC3000-inoculated plants. Gene expression in leaves was analyzed by real-time RT-PCR and expression
- 1-1 Supplemental Data 2 3 4 5 6 7 8 9 Supplemental Figure S1. Differential expression of AtPIP Genes in DC3000-inoculated plants. Gene expression in leaves was analyzed by real-time RT-PCR and expression
Supplementary Table 1. The Q-PCR primer sequence is summarized in the following table.
 Supplementary Table 1. The Q-PCR primer sequence is summarized in the following table. Name Sequence (5-3 ) Application Flag-u ggactacaaggacgacgatgac Shared upstream primer for all the amplifications of
Supplementary Table 1. The Q-PCR primer sequence is summarized in the following table. Name Sequence (5-3 ) Application Flag-u ggactacaaggacgacgatgac Shared upstream primer for all the amplifications of
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature12119 SUPPLEMENTARY FIGURES AND LEGENDS pre-let-7a- 1 +14U pre-let-7a- 1 Ddx3x Dhx30 Dis3l2 Elavl1 Ggt5 Hnrnph 2 Osbpl5 Puf60 Rnpc3 Rpl7 Sf3b3 Sf3b4 Tia1 Triobp U2af1 U2af2 1 6 2 4 3
doi:10.1038/nature12119 SUPPLEMENTARY FIGURES AND LEGENDS pre-let-7a- 1 +14U pre-let-7a- 1 Ddx3x Dhx30 Dis3l2 Elavl1 Ggt5 Hnrnph 2 Osbpl5 Puf60 Rnpc3 Rpl7 Sf3b3 Sf3b4 Tia1 Triobp U2af1 U2af2 1 6 2 4 3
Specific Aims Assignment Developing a Specific Aims Section
 Specific Aims Assignment Developing a Specific Aims Section A Specific Aims section is a 1-page document that functions as an abbreviated version of a full grant application, and is generally considered
Specific Aims Assignment Developing a Specific Aims Section A Specific Aims section is a 1-page document that functions as an abbreviated version of a full grant application, and is generally considered
1. Why do DNA restriction fragments and plasmids separate when analyzed by gel electrophoresis?
 INTRODUCTION When biologists clone a gene in order to produce human insulin, they create a recombinant plasmid that has the insulin gene. To do so, they use restriction enzymes to create DNA fragments
INTRODUCTION When biologists clone a gene in order to produce human insulin, they create a recombinant plasmid that has the insulin gene. To do so, they use restriction enzymes to create DNA fragments
A Beginners Guide to Writing Scientific Papers
 A Beginners Guide to Writing Scientific Papers Forword The writing of a scientific manuscript is a mental exercise of structural thinking. It requires thoughtfulness, mental discipline and carefulness.
A Beginners Guide to Writing Scientific Papers Forword The writing of a scientific manuscript is a mental exercise of structural thinking. It requires thoughtfulness, mental discipline and carefulness.
7.013 Practice Quiz
 MIT Department of Biology 7.013: Introductory Biology - Spring 2005 Instructors: Professor Hazel Sive, Professor Tyler Jacks, Dr. Claudette Gardel 7.013 Practice Quiz 2 2004 1 Question 1 A. The primer
MIT Department of Biology 7.013: Introductory Biology - Spring 2005 Instructors: Professor Hazel Sive, Professor Tyler Jacks, Dr. Claudette Gardel 7.013 Practice Quiz 2 2004 1 Question 1 A. The primer
Lesson 3 Gel Electrophoresis of Amplified PCR Samples and Staining of Agarose Gels
 Lesson 3 Gel Electrophoresis of Amplified PCR Samples and Staining of Agarose Gels What Are You Looking At? Before you analyze your PCR products, let s take a look at the target sequence being explored.
Lesson 3 Gel Electrophoresis of Amplified PCR Samples and Staining of Agarose Gels What Are You Looking At? Before you analyze your PCR products, let s take a look at the target sequence being explored.
Supplementary Figure S1. N-terminal fragments of LRRK1 bind to Grb2.
 Myc- HA-Grb2 Mr(K) 105 IP HA 75 25 105 1-1163 1-595 - + - + - + 1164-1989 Blot Myc HA total lysate 75 25 Myc HA Supplementary Figure S1. N-terminal fragments of bind to Grb2. COS7 cells were cotransfected
Myc- HA-Grb2 Mr(K) 105 IP HA 75 25 105 1-1163 1-595 - + - + - + 1164-1989 Blot Myc HA total lysate 75 25 Myc HA Supplementary Figure S1. N-terminal fragments of bind to Grb2. COS7 cells were cotransfected
SUPPLEMENTARY INFORMATION
 Materials and Methods Transgenic Plant Materials and DNA Constructs. VEX1::H2B-GFP, ACA3::H2B-GFP, KRP6::H2B-GFP, KRP6::mock21ts-GFP, KRP6::TE21ts- GFP and KRP6::miR161ts-GFP constructs were generated
Materials and Methods Transgenic Plant Materials and DNA Constructs. VEX1::H2B-GFP, ACA3::H2B-GFP, KRP6::H2B-GFP, KRP6::mock21ts-GFP, KRP6::TE21ts- GFP and KRP6::miR161ts-GFP constructs were generated
(phosphatase tensin) domain is shown in dark gray, the FH1 domain in black, and the
 Supplemental Figure 1. Predicted Domain Organization of the AFH14 Protein. (A) Schematic representation of the predicted domain organization of AFH14. The PTEN (phosphatase tensin) domain is shown in dark
Supplemental Figure 1. Predicted Domain Organization of the AFH14 Protein. (A) Schematic representation of the predicted domain organization of AFH14. The PTEN (phosphatase tensin) domain is shown in dark
Mission (Im)possible: Plasmid Mapping Student Materials
 Mission (Im)possible: Plasmid Mapping Student Materials Introduction... 2 Pre-Lab Questions... 6 Lab Protocol... 7 Data Collection Worksheet... 11 Post-Lab Questions and Analysis... 12 Last updated: August
Mission (Im)possible: Plasmid Mapping Student Materials Introduction... 2 Pre-Lab Questions... 6 Lab Protocol... 7 Data Collection Worksheet... 11 Post-Lab Questions and Analysis... 12 Last updated: August
Supplementary Fig. 1. Schematic structure of TRAIP and RAP80. The prey line below TRAIP indicates bait and the two lines above RAP80 highlight the
 Supplementary Fig. 1. Schematic structure of TRAIP and RAP80. The prey line below TRAIP indicates bait and the two lines above RAP80 highlight the prey clones identified in the yeast two hybrid screen.
Supplementary Fig. 1. Schematic structure of TRAIP and RAP80. The prey line below TRAIP indicates bait and the two lines above RAP80 highlight the prey clones identified in the yeast two hybrid screen.
Transport of Potato Lipoxygenase into the Vacuole Larsen, Mia Kruse Guldstrand; Welinder, Karen Gjesing; Jørgensen, Malene
 Aalborg Universitet Transport of Potato Lipoxygenase into the Vacuole Larsen, Mia Kruse Guldstrand; Welinder, Karen Gjesing; Jørgensen, Malene Publication date: 2009 Document Version Publisher's PDF, also
Aalborg Universitet Transport of Potato Lipoxygenase into the Vacuole Larsen, Mia Kruse Guldstrand; Welinder, Karen Gjesing; Jørgensen, Malene Publication date: 2009 Document Version Publisher's PDF, also
Supplemental Data. Hu et al. Plant Cell (2017) /tpc
 1 2 3 4 Supplemental Figure 1. DNA gel blot analysis of homozygous transgenic plants. (Supports Figure 1.) 5 6 7 8 Rice genomic DNA was digested with the restriction enzymes EcoRⅠ and BamHⅠ. Lanes in the
1 2 3 4 Supplemental Figure 1. DNA gel blot analysis of homozygous transgenic plants. (Supports Figure 1.) 5 6 7 8 Rice genomic DNA was digested with the restriction enzymes EcoRⅠ and BamHⅠ. Lanes in the
Supplementary Figure 1. GST pull-down analysis of the interaction of GST-cIAP1 (A, B), GSTcIAP1
 Legends Supplementary Figure 1. GST pull-down analysis of the interaction of GST- (A, B), GST mutants (B) or GST- (C) with indicated proteins. A, B, Cell lysate from untransfected HeLa cells were loaded
Legends Supplementary Figure 1. GST pull-down analysis of the interaction of GST- (A, B), GST mutants (B) or GST- (C) with indicated proteins. A, B, Cell lysate from untransfected HeLa cells were loaded
A RRM1 H2AX DAPI. RRM1 H2AX DAPI Merge. Cont. sirna RRM1
 A H2AX DAPI H2AX DAPI Merge Cont sirna Figure S1: Accumulation of RRM1 at DNA damage sites (A) HeLa cells were subjected to in situ detergent extraction without IR irradiation, and immunostained with the
A H2AX DAPI H2AX DAPI Merge Cont sirna Figure S1: Accumulation of RRM1 at DNA damage sites (A) HeLa cells were subjected to in situ detergent extraction without IR irradiation, and immunostained with the
BeANs Lab Brief Guidelines for Report/Paper Writing
 BeANs Lab Brief Guidelines for Report/Paper Writing Prepared by Sierin Lim Last updated: 14-Aug-13 Please make sure that you did not copy from any source. Rule of Thumbs Paper size: A4 or letter Font:
BeANs Lab Brief Guidelines for Report/Paper Writing Prepared by Sierin Lim Last updated: 14-Aug-13 Please make sure that you did not copy from any source. Rule of Thumbs Paper size: A4 or letter Font:
Enzyme that uses RNA as a template to synthesize a complementary DNA
 Biology 105: Introduction to Genetics PRACTICE FINAL EXAM 2006 Part I: Definitions Homology: Comparison of two or more protein or DNA sequence to ascertain similarities in sequences. If two genes have
Biology 105: Introduction to Genetics PRACTICE FINAL EXAM 2006 Part I: Definitions Homology: Comparison of two or more protein or DNA sequence to ascertain similarities in sequences. If two genes have
Supplemental Information. Autoregulatory Feedback Controls. Sequential Action of cis-regulatory Modules. at the brinker Locus
 Developmental Cell, Volume 26 Supplemental Information Autoregulatory Feedback Controls Sequential Action of cis-regulatory Modules at the brinker Locus Leslie Dunipace, Abbie Saunders, Hilary L. Ashe,
Developmental Cell, Volume 26 Supplemental Information Autoregulatory Feedback Controls Sequential Action of cis-regulatory Modules at the brinker Locus Leslie Dunipace, Abbie Saunders, Hilary L. Ashe,
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION 1 Supplementary Figure S1. Proportion of HEK293T transfectants showing fluorescent foci after heat sho Blinded slides were counted for the appearance of fluorescent foci of tagrfp-t
SUPPLEMENTARY INFORMATION 1 Supplementary Figure S1. Proportion of HEK293T transfectants showing fluorescent foci after heat sho Blinded slides were counted for the appearance of fluorescent foci of tagrfp-t
Appendix A. Exogenous matrix binding assays with. various putative MARs
 Appendix A Exogenous matrix binding assays with various putative MARs Introduction Matrix attachment regions of DNA are defined operationally by their affinity for the nuclear matrix. Identification of
Appendix A Exogenous matrix binding assays with various putative MARs Introduction Matrix attachment regions of DNA are defined operationally by their affinity for the nuclear matrix. Identification of
Supplementary Information. A superfolding Spinach2 reveals the dynamic nature of. trinucleotide repeat RNA
 Supplementary Information A superfolding Spinach2 reveals the dynamic nature of trinucleotide repeat RNA Rita L. Strack 1, Matthew D. Disney 2 & Samie R. Jaffrey 1 1 Department of Pharmacology, Weill Medical
Supplementary Information A superfolding Spinach2 reveals the dynamic nature of trinucleotide repeat RNA Rita L. Strack 1, Matthew D. Disney 2 & Samie R. Jaffrey 1 1 Department of Pharmacology, Weill Medical
