X-ray structures of fructosyl peptide oxidases revealing residues responsible for gating oxygen access in the oxidative half reaction
|
|
|
- Colin Ryan
- 6 years ago
- Views:
Transcription
1 X-ray structures of fructosyl peptide oxidases revealing residues responsible for gating oxygen access in the oxidative half reaction Tomohisa Shimasaki 1, Hiromi Yoshida 2, Shigehiro Kamitori 2 & Koji Sode 1 * 1 Department of Biotechnology and Life Science, Graduate School of Engineering, Tokyo University of Agriculture and Technology, , Nakamachi, Koganei, Tokyo , Japan 2 Life Science Research Center and Faculty of Medicine, Kagawa University, Ikenobe, Miki-cho, Kita-gun, Kagawa, Japan *Corresponding author Correspondence to: sode@cc.tuat.ac.jp 1
2 Lys376/ Lys376/ Lys380/ Met375/ Lys378 Thr18/ Thr18/ Thr18/ Thr19/ Thr16 BDF-Lys Lys376 (partially conserved) Ile58/ Ile58/ Ile58/ Ile57/ Val54 Thr18 Ile58 (main chain conserved) Ala51 (main chain conserved) FSA / / / Lys56/ Lys53 PnFPOX WT with bound ACY PnFOPX N56A EtFPOX Amadoriase I with bound BDF-Lys Amadoriase II with bound FSA ACY Ala51/ Ala51/ Ala51/ Ala50/ Ala47 Asn56/ Asn56Ala/ Asn56/ Asn55/ Asn52 Ser50 (partially conserved) Asn56 Ser50/ Ser50/ Ser50/ Ala49/ Ser46 Asp41 / / Lys278/ Lys281/ Lys276 Asp41/ Asp41/ Asp41/ Asp40/ Asp37 Cys343 Ser47 / / / Asp53/ Asp50 His239 Cys343/ Cys343/ Cys347/ Cys342/ Cys335 Ser47/ Serr47/ Ser47/ Ser46/ Ser43 His239/ His239/ His243/ His245/ His240 Supplementary Figure S1 Structural comparison around at the si-face among PnFPOX WT with bound ACY (yellow), N56A (cyan), EtFPOX (light purple), amadoriase I with bound BDF-Lys (magenta) and amadoriase II with bound FSA (light green). Six residues (, Asn56,, His239,, Cys343) in the vicinity of at the si-face are structurally conserved in these enzymes. Especially, four of them (, Asn56, and ) are conserved in sequence alignment of eukaryotic FAOD/FPOX, according to Kim et al 12. ACY: acetate BDF: beta-d-fructopyranose Lys: Lysine FSA: 1-s-(carboxymethyl)-1-thio-beta-D-fructopyranose 2
3 (a) (b) (c) (d) O1 CH3 O2 O5 C C4 3 O5 C6 5 4 C Acetate 1 1 Lys BDF FSA Supplementary Figure S2 The comparison of bound ligands in active site. (a) The bound acetate in wild-type PnFPOX (b) The bound BDF-Lys in amadoliase I (PDB: 4XWZ) (c) The bound FSA in amadoriase II (PDB: 3DJE). (d) The bound ligands of (b) and (c) were superimposed onto (a). BDF: beta-d-fructopyranose Lys: Lysine FSA: 1-s-(carboxymethyl)-1-thio-beta-D-fructopyranose 3
4 (a) (b) (c) W1 W2 OD2 OD2 OD1 W1 W2 OD2 OD1 OD1 Asn σ 3.5 σ 4.0 σ Supplementary Figure S3 SA-omit maps of the,, W1 and W2 in wild-type PnFPOX contoured at 3.0, 3.5, 4.0 σ (a, b, c, respectively). The possibility of the alternative conformation of was shown, however, the distance between W2 and OD2 of in alternative conformation is 1.1 Å ((a), (b)). In the SA-omit maps at 4.0 σ, the positions of W2 and were confirmed (c). At the condition of 0.5 occupancy, the b-factors of OD1 and OD1 (OD2 and OD2 ) of were and (27.99 and 37.33), respectively, and the b-factor of water molecule (W2) with 0.5 occupancy was Although we chose the current position of as main position in our current model from the SA-omit maps, the water molecule (W2) and the side chain of could occupy the position alternatively. 4
5 (a) (b) Ala51 Ala51 N5 O2 Cys343 Cys343 W W3 W W4 W1 W4 Asn56 His239 Ala56 His239 (c) W6 W5 W6 W5 Ser43 Cys315 His45 W2 Arg49 W1 Cl- Gly46 Thr48 W5 W6 Lys265 Supplementary Figure S4 Oxygen binding-site comparison between wild-type PnFPOX, N56A, and monomeric sarcosine oxidase (MSOX) with bound chloride. (a) The structure of wild-type PnFPOX. Red spheres indicate water molecules. (b) The structure of PnFPOX N56A. Light pink spheres indicate water molecules. (c) The structure of MSOX with bound chloride (PDB: 3QSM). Light green sphere indicates chloride. Magenta spheres indicate water molecules. Selected hydrogen bonds and salt bridge interactions (< 3.53 Å ) are indicated by dotted lines with distances. Blue solid lines indicate the distance between selected atoms.
6 Supplementary Figure S5 SDS-PAGE analysis of water soluble fraction of cell lysate derived from recombinant E.coli expressing either wild-type or mutant enzyme. Lane M contains molecular-mass markers. Water soluble fraction derived from cell lysate of E.coli expressing either wild-type or mutant enzyme, containing 7.5 mg was applied in each lane. The red arrow indicates the expected molecular weight (48kDa) of PnFPOX WT and mutants. Lane 1, WT; Lane 2, D54E; Lane 3, D54N; Lane 4, D54A; Lane 5 D54H; Lane 6, D54V; Lane 7, D54S; Lane 8, D54Y; Lane 9, D54F.
7 (a) (b) (c) W6/W6 W5/W5 His239 Asn56 W1/W1 W2 W4/W4 Ala56 W3 Supplementary Figure S6 Analysis of protein channels of wild-type PnFPOX and N56A (a) Surface model of wild-type PnFPOX with oxygen accessible channels (light green and blue dotted spheres). The entrance of short and plausible main channel is indicated with blue arrow. (b) Surface model of PnFPOX N56A with oxygen accessible channel (blue, cyan, light green, and red dotted spheres). The entrance of short and plausible main channel in wild-type PnFPOX is indicated with blue arrow. The main channel is closed in PnFPOX N56A. (c) Superimposition of the PnFPOX N56A (cyan stick) with water molecules (pink spheres) onto the surface model of wild-type PnFPOX (yellow stick) with water molecules (red spheres) and main oxygen accessible channel (blue dotted spheres with arrow). In PnFPOX N56A, the movement of and blocks a short and plausible main channel observed in wild-type PnFPOX (blue dotted path with arrow). Another long path would enable oxygen to reach the active site instead of the closed path in PnFPOX N56A, but it would not be effective for competition with dye mediator for dehydrogenase activity.
8 (a) (b) (c) W6 W5 Leu55 His239 Met59 Asn56 W1 W2 W3 W4 Asn53 Ala51 Supplementary Figure S7 Analysis of protein channels of wild-type PnFPOX in the presence/absence of NaCl The surface models with oxygen accessible channels between the structures of wild-type PnFPOX in the presence (green stick and surface) / absence (yellow stick and surface) of NaCl were compared. For comparison, the structure of Mol A in wild-type PnFPOX with NaCl was used, since the electron densities of the side chains of and were invisible in Mol B of wild-type PnFPOX with NaCl. The orientation of the side of was not clear due to the weak electron density in Mol A. The entrance of short and plausible main channel is indicated with blue arrow. (a) The surface model of wild-type PnFPOX with NaCl (green stick and surface). Oxygen accessible channels were shown in blue, light green, cyan, magenta, red, yellow, orange, purple, and other dotted spheres. (b) The surface model of wild-type PnFPOX (yellow stick and surface) with water molecules (red spheres). Oxygen accessible channels were shown in light green and blue dotted spheres. (c) Superimposition of the region (Asn53 Met59) of the wild-type PnFPOX in the presence of NaCl (green stick) onto the corresponding region of the structure of wild-type PnFPOX in the absence of NaCl (yellow stick) and with water molecules (red spheres). Both molecules were shown in line models. In the vicinity of isoalloxazine ring of in wild-type PnFPOX in the presence of NaCl (green stick), the region (Asn53 Met59) including and was slightly moved, and the orientations of side chains of Asn53,, Leu55 were changed. Although the orientation of was not clear due to the weak electron density, the orientation of was also moved. The addition of NaCl caused a structural change of the region and gave more possible oxygen accessible channels leading to the putative main gate consisted a pair of and.
9 (a) (b) (c) W6 W5 Leu55 (d) Ala56 His239 Met59 CL Ile58 W4 W1 NA Pro66 Gly60 Met59 Ile58 W1 W2 Asn53 Ala51 Arg94 CL Gly156 Supplementary Figure S8 Analysis of protein channels of PnFPOX N56A in the presence/absence of NaCl The surface models with oxygen accessible channel between the structures of PnFPOX N56A in the presence (blue stick and surface) / absence (cyan stick and white surface) of NaCl were compared. The region in the vicinity of occurred conformational change in Mol A of PnFPOX N56A with NaCl, and the structure of Mol A was used for comparison to see any difference. The entrance of short and plausible main channel belonging to wild-type PnFPOX is indicated with blue arrow. (a) The surface model of PnFPOX N56A in the presence of NaCl (blue stick and surface). Water molecules (magenta spheres) and oxygen accessible channels (blue, light green, cyan and red dotted spheres) were shown. The bound chlorides were shown in orange spheres. (b) The surface model of PnFPOX N56A (cyan stick and white surface) with water molecules (pink spheres). Oxygen accessible channels were shown in blue, cyan, light green, and red dotted spheres. (c) The region (Asn53 Met59) of PnFPOX N56A in the presence of NaCl (blue stick) with water molecules (magenta spheres), the bound chloride (orange sphere) and the bound sodium (yellow sphere) was superimposed onto the corresponding region of the structure of PnFPOX N56A in the absence of NaCl (cyan stick) and with water molecules (pink spheres with the number). Both molecules were shown in line models. (d) Comparison of the structure in the vicinity of the bound chloride between PnFPOX N56A in the presence /absence of NaCl. In the vicinity of the isoalloxazine ring of, the region (Asn53 Pro66) of PnFPOX N56A (cyan line) caused conformational change by addition of NaCl (PnFPOX N56A with NaCl, blue stick), giving the disorder of the side chains and shift of main chains. The bound chloride (orange sphere) was found in slightly opened area surrounded by Ile58, Met59, Gly60, Arg94, Gly156. The bound sodium interacted with carbonyl oxygen of Asn53. The addition of NaCl would create other plausible oxygen channels, but it caused the conformational change of the flexible loop region which might affect oxygen paths for oxygen activity.
10 1/V Oxidase activity (U/mg) Substrate (mm) /[S] Supplementary Figure S9. Michaelis-Menten and Lineweaver-Burk plots of oxidase activity of PnFPOX WT. 10
11 Supplementary Table S1. Relative activities and Dh/Ox values of PnFPOX mutants Activity relative to wild-type PnFPOX (%) 1) Dh/Ox 2) Oxidase activity Dehydrogenase activity WT D54E D54N D54A D54H D54V D54S D54Y n.d. 3) 0.64 n.a. 4) D54F n.d. 3) 0.97 n.a. 4) 1) These values were calculated based on the oxidase or dehydrogenase activity of wild-type enzyme sample, which were prepared from cell lysate of recombinant E.coli. The values derived from wild-type enzyme were used as 100%. 2) Dh/Ox values were calculated based on the comparison of oxidase activities and dye-mediated dehydrogenase activities observed in the cell lysate of recombinant E.coli expressing either wild-type or mutant enzyme with 1.0 mm substrate concentration. 3) n.d. : not detected 4) n.a. ; not available 11
12 Supplementary Table S2. Data collection and refinement statistics. Data collection Wild-type PnFPOX with NaCl N56A with NaCl Beamline MicroMax-007 HF MicroMax-007 HF Temperature (K) Wavelength (Å) Resolution range (Å) ( ) ( ) No. of measured refs. 58, ,813 No. of unique refs. 16,927 80,576 Redundancy 3.5 (3.5) 3.6 (3.5) Completeness (%) 96.7 (97.8) 96.0 (93.3) Mean I o /s(i o ) 7.1 (4.0) 6.6 (1.8) R merge (%) 16.9 (47.6) 8.0 (51.4) Space group P2 1 P2 1 a = a = Unit cell parameters b = b = a, b, c (Å) c = c = b ( ) b = b = Refinement Resolution range (Å) ( ) ( ) No. of refs. 16,073 (1,194) 76,559 (5,448) Completeness (%) 96.8 (97.7) 96.0 (93.5) R factor (%) 25.1 (32.5) (35.5) R free (%) 30.1 (38.6) (40.6) RMSD bond lengths (Å) RMSD bond angles (º) Ramachandran plot Most favoured region (%) Additional allowed region (%) B-factor (Å 2 ) Protein Ligand () 18.2 (2 molecules) 24.2 (2 molecules) Ligand (ACY) (2 molecules) Ligand (CL) (5 molecules) Ligand (NA) (4 molecules) Water PDB code - 5XAO Values in parentheses are of the high-resolution bin. R merge = Σ h Σ i [ I i (h) - <I(h)> / Σ h Σ i I i (h)], in which I i is the ith measurement and <I(h)> is the weighted mean of all measurements of I(h).
13 Supplementary Table S3. Primer sequences for mutant construction D54A sense D54A antisense D54E sense D54E antisense D54F sense D54F antisense D54H sense D54H antisense D54N sense D54N antisense D54S sense D54S antisense D54V sense D54V antisense D54Y sense D54Y antisense N56A sense N56A antisense 5 -CAATCTGCAGGCAATGCGCTGAATAAGATTATG-3 5 -CATAATCTTATTCAGCGCATTGCCTGCAGATTG-3 5 -CAATCTGCAGGCAATGAACTGAATAAGATTATG-3 5 -CATAATCTTATTCAGTTCATTGCCTGCAGATTG-3 5 -CAATCTGCAGGCAATTTCCTGAATAAGATTATG-3 5 -CATAATCTTATTCAGGAAATTGCCTGCAGATTG-3 5 -AGGCAATCACCTGAATAAGATTATGGGCGTT-3 5 -AACGCCCATAATCTTATTCAGGTGATTGCCT-3 5 -CAATCTGCAGGCAATAACCTGAATAAGATTATG-3 5 -CATAATCTTATTCAGGTTATTGCCTGCAGATTG-3 5 -GCAGGCAATTCTCTGAATAAGATTATGGGCGTT-3 5 -AACGCCCATAATCTTATTCAGAGAATTGCCTGC-3 5 -CAATCTGCAGGCAATGTTCTGAATAAGATTATG-3 5 -CATAATCTTATTCAGAACATTGCCTGCAGATTG-3 5 -CAATCTGCAGGCAATTACCTGAATAAGATTATG-3 5 -CATAATCTTATTCAGGTAATTGCCTGCAGATTG-3 5 -AGGCAATGACCTGGCGAAGATTATGGGCG-3 5 -CGCCCATAATCTTCGCCAGGTCATTGCCT-3 13
Suppl. Figure 1: RCC1 sequence and sequence alignments. (a) Amino acid
 Supplementary Figures Suppl. Figure 1: RCC1 sequence and sequence alignments. (a) Amino acid sequence of Drosophila RCC1. Same colors are for Figure 1 with sequence of β-wedge that interacts with Ran in
Supplementary Figures Suppl. Figure 1: RCC1 sequence and sequence alignments. (a) Amino acid sequence of Drosophila RCC1. Same colors are for Figure 1 with sequence of β-wedge that interacts with Ran in
Supplementary Figure 1
 Supplementary Figure 1 2 Supplementary Figure 1: Sequence alignment of HsHSD17B8 and HsCBR4 of with KAR orthologs. The secondary structure elements as calculated by DSSP and residue numbers are displayed
Supplementary Figure 1 2 Supplementary Figure 1: Sequence alignment of HsHSD17B8 and HsCBR4 of with KAR orthologs. The secondary structure elements as calculated by DSSP and residue numbers are displayed
has only one nucleotide, U20, between G19 and A21, while trna Glu CUC has two
 SPPLEMENTRY INFORMTION doi:1.138/nature9411 Supplementary Discussion The structural characteristics of trn Gln G in comparison to trn Glu 16 in trn Gln G is directed towards the G19 56 pair, or the outer
SPPLEMENTRY INFORMTION doi:1.138/nature9411 Supplementary Discussion The structural characteristics of trn Gln G in comparison to trn Glu 16 in trn Gln G is directed towards the G19 56 pair, or the outer
Supplementary Figure 1. Electron microscopy of gb-698glyco/1g2 Fab complex. a)
 Supplementary Figure 1. Electron microscopy of gb-698glyco/1g2 Fab complex. a) Representative images of 2D class averages of gb-698glyc bound to 1G2 Fab. Top views of the complex were underrepresented
Supplementary Figure 1. Electron microscopy of gb-698glyco/1g2 Fab complex. a) Representative images of 2D class averages of gb-698glyc bound to 1G2 Fab. Top views of the complex were underrepresented
Structure and Function of the First Full-Length Murein Peptide Ligase (Mpl) Cell Wall Recycling Protein
 Paper Presentation PLoS ONE 2011 Structure and Function of the First Full-Length Murein Peptide Ligase (Mpl) Cell Wall Recycling Protein Debanu Das, Mireille Herve, Julie Feuerhelm, etc. and Dominique
Paper Presentation PLoS ONE 2011 Structure and Function of the First Full-Length Murein Peptide Ligase (Mpl) Cell Wall Recycling Protein Debanu Das, Mireille Herve, Julie Feuerhelm, etc. and Dominique
SUPPLEMENTARY INFORMATION Figures. Supplementary Figure 1 a. Page 1 of 30. Nature Chemical Biology: doi: /nchembio.2528
 SUPPLEMENTARY INFORMATION Figures Supplementary Figure 1 a b c Page 1 of 0 11 Supplementary Figure 1: Biochemical characterisation and binding validation of the reversible USP inhibitor 1. a, Biochemical
SUPPLEMENTARY INFORMATION Figures Supplementary Figure 1 a b c Page 1 of 0 11 Supplementary Figure 1: Biochemical characterisation and binding validation of the reversible USP inhibitor 1. a, Biochemical
Ensemble refinement shows conformational flexibility in crystal structures of human complement factor D
 Supplementary Information for Ensemble refinement shows conformational flexibility in crystal structures of human complement factor D Federico Forneris a,b, B. Tom Burnley a,b,c and Piet Gros a * a Crystal
Supplementary Information for Ensemble refinement shows conformational flexibility in crystal structures of human complement factor D Federico Forneris a,b, B. Tom Burnley a,b,c and Piet Gros a * a Crystal
6-Foot Mini Toober Activity
 Big Idea The interaction between the substrate and enzyme is highly specific. Even a slight change in shape of either the substrate or the enzyme may alter the efficient and selective ability of the enzyme
Big Idea The interaction between the substrate and enzyme is highly specific. Even a slight change in shape of either the substrate or the enzyme may alter the efficient and selective ability of the enzyme
SUPPLEMENTARY INFORMATION
 doi: 10.1038/nature06147 SUPPLEMENTARY INFORMATION Figure S1 The genomic and domain structure of Dscam. The Dscam gene comprises 24 exons, encoding a signal peptide (SP), 10 IgSF domains, 6 fibronectin
doi: 10.1038/nature06147 SUPPLEMENTARY INFORMATION Figure S1 The genomic and domain structure of Dscam. The Dscam gene comprises 24 exons, encoding a signal peptide (SP), 10 IgSF domains, 6 fibronectin
In silico measurements of twist and bend. moduli for beta solenoid protein self-
 In silico measurements of twist and bend moduli for beta solenoid protein self- assembly units Leonard P. Heinz, Krishnakumar M. Ravikumar, and Daniel L. Cox Department of Physics and Institute for Complex
In silico measurements of twist and bend moduli for beta solenoid protein self- assembly units Leonard P. Heinz, Krishnakumar M. Ravikumar, and Daniel L. Cox Department of Physics and Institute for Complex
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10963 Supplementary Table 1 Data collection, phasing and refinement statistics. Crystal Native Derivative-1 (OsO 4 ) Derivative-2 (Orange-Pt) Data collection Space group C2 C2 C2 Cell
doi:10.1038/nature10963 Supplementary Table 1 Data collection, phasing and refinement statistics. Crystal Native Derivative-1 (OsO 4 ) Derivative-2 (Orange-Pt) Data collection Space group C2 C2 C2 Cell
466 Asn (N) to Ala (A) Generate beta dimer Interface
 Table S1: Amino acid changes to the HexA α-subunit to convert the dimer interface from α to β and to introduce the putative GM2A binding surface from β- onto the α- subunit Residue position (α-numbering)
Table S1: Amino acid changes to the HexA α-subunit to convert the dimer interface from α to β and to introduce the putative GM2A binding surface from β- onto the α- subunit Residue position (α-numbering)
Supplementary Fig. S1. SAMHD1c has a more potent dntpase activity than. SAMHD1c. Purified recombinant SAMHD1c and SAMHD1c proteins (with
 Supplementary Fig. S1. SAMHD1c has a more potent dntpase activity than SAMHD1c. Purified recombinant SAMHD1c and SAMHD1c proteins (with concentration of 800nM) were incubated with 1mM dgtp for the indicated
Supplementary Fig. S1. SAMHD1c has a more potent dntpase activity than SAMHD1c. Purified recombinant SAMHD1c and SAMHD1c proteins (with concentration of 800nM) were incubated with 1mM dgtp for the indicated
Supplementary Information Titles
 Supplementary Information Titles Please list each supplementary item and its title or caption, in the order shown below. Note that we do NOT copy edit or otherwise change supplementary information, and
Supplementary Information Titles Please list each supplementary item and its title or caption, in the order shown below. Note that we do NOT copy edit or otherwise change supplementary information, and
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature10258 Supplementary Figure 1 Reconstitution of the CENP-A nucleosome with recombinant human histones H2A, H2B, H4, and CENP-A. a, Purified recombinant human
SUPPLEMENTARY INFORMATION doi:10.1038/nature10258 Supplementary Figure 1 Reconstitution of the CENP-A nucleosome with recombinant human histones H2A, H2B, H4, and CENP-A. a, Purified recombinant human
Q22M, T44W, R81G, H83G, T84M, N130G, N172M, A234S, T236L, C9G, V48G, L50W, T78S, S101E, E237M, T265S, W267F 7, 11, , 239,
 111 Table 4-1. Summary of design calculations for Kemp elimination enzymes. The residues allowed for each required catalytic contact are indicated along with the actual catalytic residue chosen from the
111 Table 4-1. Summary of design calculations for Kemp elimination enzymes. The residues allowed for each required catalytic contact are indicated along with the actual catalytic residue chosen from the
Solutions to 7.02 Quiz II 10/27/05
 Solutions to 7.02 Quiz II 10/27/05 Class Average = 83 Standard Deviation = 9 Range Grade % 87-100 A 43 74-86 B 39 55-73 C 17 > 54 D 1 Question 1 (56 points) While studying deep sea bacteria, you discover
Solutions to 7.02 Quiz II 10/27/05 Class Average = 83 Standard Deviation = 9 Range Grade % 87-100 A 43 74-86 B 39 55-73 C 17 > 54 D 1 Question 1 (56 points) While studying deep sea bacteria, you discover
Dr. R. Sankar, BSE 631 (2018)
 Pauling, Corey and Branson Diffraction of DNA http://www.nature.com/scitable/topicpage/dna-is-a-structure-that-encodes-biological-6493050 In short, stereochemistry is important in determining which helices
Pauling, Corey and Branson Diffraction of DNA http://www.nature.com/scitable/topicpage/dna-is-a-structure-that-encodes-biological-6493050 In short, stereochemistry is important in determining which helices
Supporting Information Contents
 Supporting Information Choy Theng Loh, Kiyoshi Ozawa, Kellie L. Tuck, Nicholas Barlow, Thomas Huber, Gottfried Otting, and Bim Graham Lanthanide tags for site-specific ligation to an unnatural amino acid
Supporting Information Choy Theng Loh, Kiyoshi Ozawa, Kellie L. Tuck, Nicholas Barlow, Thomas Huber, Gottfried Otting, and Bim Graham Lanthanide tags for site-specific ligation to an unnatural amino acid
Supporting Information
 Supporting Information A Facile and Sensitive Near-Infrared Fluorescence Probe for the Detection of Endogenous Alkaline Phosphatase Activity in Vivo Song-Jiao Li, Chun-Yan Li,*,, Yong-Fei Li, Junjie Fei,
Supporting Information A Facile and Sensitive Near-Infrared Fluorescence Probe for the Detection of Endogenous Alkaline Phosphatase Activity in Vivo Song-Jiao Li, Chun-Yan Li,*,, Yong-Fei Li, Junjie Fei,
Supplementary Data for Monti, et al.
 Supplementary Data for Monti, et al. Supplementary Figure S1 Legend to Supplementary Figure S1 Tumor spectrum associated with germline p53 alleles (restricted to the 7 most frequent tissue targets). Structural
Supplementary Data for Monti, et al. Supplementary Figure S1 Legend to Supplementary Figure S1 Tumor spectrum associated with germline p53 alleles (restricted to the 7 most frequent tissue targets). Structural
Supplementary Information For. A genetically encoded tool for manipulation of NADP + /NADPH in living cells
 Supplementary Information For A genetically encoded tool for manipulation of NADP + /NADPH in living cells Valentin Cracan 1,2,3, Denis V. Titov 1,2,3, Hongying Shen 1,2,3, Zenon Grabarek 1* and Vamsi
Supplementary Information For A genetically encoded tool for manipulation of NADP + /NADPH in living cells Valentin Cracan 1,2,3, Denis V. Titov 1,2,3, Hongying Shen 1,2,3, Zenon Grabarek 1* and Vamsi
First&year&tutorial&in&Chemical&Biology&(amino&acids,&peptide&and&proteins)&! 1.&!
 First&year&tutorial&in&Chemical&Biology&(amino&acids,&peptide&and&proteins& 1.& a. b. c. d. e. 2.& a. b. c. d. e. f. & UsingtheCahn Ingold Prelogsystem,assignstereochemicaldescriptorstothe threeaminoacidsshownbelow.
First&year&tutorial&in&Chemical&Biology&(amino&acids,&peptide&and&proteins& 1.& a. b. c. d. e. 2.& a. b. c. d. e. f. & UsingtheCahn Ingold Prelogsystem,assignstereochemicaldescriptorstothe threeaminoacidsshownbelow.
Supplementary Table 1: List of CH3 domain interface residues in the first chain (A) and
 Supplementary Tables Supplementary Table 1: List of CH3 domain interface residues in the first chain (A) and their side chain contacting residues in the second chain (B) a Interface Res. in Contacting
Supplementary Tables Supplementary Table 1: List of CH3 domain interface residues in the first chain (A) and their side chain contacting residues in the second chain (B) a Interface Res. in Contacting
Case 7 A Storage Protein From Seeds of Brassica nigra is a Serine Protease Inhibitor
 Case 7 A Storage Protein From Seeds of Brassica nigra is a Serine Protease Inhibitor Focus concept Purification of a novel seed storage protein allows sequence analysis and determination of the protein
Case 7 A Storage Protein From Seeds of Brassica nigra is a Serine Protease Inhibitor Focus concept Purification of a novel seed storage protein allows sequence analysis and determination of the protein
Homology Modelling. Thomas Holberg Blicher NNF Center for Protein Research University of Copenhagen
 Homology Modelling Thomas Holberg Blicher NNF Center for Protein Research University of Copenhagen Why are Protein Structures so Interesting? They provide a detailed picture of interesting biological features,
Homology Modelling Thomas Holberg Blicher NNF Center for Protein Research University of Copenhagen Why are Protein Structures so Interesting? They provide a detailed picture of interesting biological features,
Nature Structural & Molecular Biology: doi: /nsmb.2307
 A novel locally-closed conformation of a bacterial pentameric proton-gated ion channel Marie S. Prevost*, Ludovic Sauguet*, Hugues Nury, Catherine Van Renterghem, Christèle Huon, Frederic Poitevin, Marc
A novel locally-closed conformation of a bacterial pentameric proton-gated ion channel Marie S. Prevost*, Ludovic Sauguet*, Hugues Nury, Catherine Van Renterghem, Christèle Huon, Frederic Poitevin, Marc
1/4/18 NUCLEIC ACIDS. Nucleic Acids. Nucleic Acids. ECS129 Instructor: Patrice Koehl
 NUCLEIC ACIDS ECS129 Instructor: Patrice Koehl Nucleic Acids Nucleotides DNA Structure RNA Synthesis Function Secondary structure Tertiary interactions Wobble hypothesis DNA RNA Replication Transcription
NUCLEIC ACIDS ECS129 Instructor: Patrice Koehl Nucleic Acids Nucleotides DNA Structure RNA Synthesis Function Secondary structure Tertiary interactions Wobble hypothesis DNA RNA Replication Transcription
NUCLEIC ACIDS. ECS129 Instructor: Patrice Koehl
 NUCLEIC ACIDS ECS129 Instructor: Patrice Koehl Nucleic Acids Nucleotides DNA Structure RNA Synthesis Function Secondary structure Tertiary interactions Wobble hypothesis DNA RNA Replication Transcription
NUCLEIC ACIDS ECS129 Instructor: Patrice Koehl Nucleic Acids Nucleotides DNA Structure RNA Synthesis Function Secondary structure Tertiary interactions Wobble hypothesis DNA RNA Replication Transcription
Supplementary Table 1. DNA sequence synthesized to express the Zika virus NS5 protein.
 Supplementary Table 1. DNA sequence synthesized to express the Zika virus NS5 protein. MR766 NS5 sequence 1 ACAGAGAACAGATTGGTGGTGGAGGTGGGACGGGAGAGACTCTGGGAGAGAAGTGGAAAG 61 CTCGTCTGAATCAGATGTCGGCCCTGGAGTTCTACTCTTATAAAAAGTCAGGTATCACTG
Supplementary Table 1. DNA sequence synthesized to express the Zika virus NS5 protein. MR766 NS5 sequence 1 ACAGAGAACAGATTGGTGGTGGAGGTGGGACGGGAGAGACTCTGGGAGAGAAGTGGAAAG 61 CTCGTCTGAATCAGATGTCGGCCCTGGAGTTCTACTCTTATAAAAAGTCAGGTATCACTG
Amino Acids and Proteins
 Various Functions of Proteins SB203 Amino Acids and Proteins Jirundon Yuvaniyama, Ph.D. Department of Biochemistry Faculty of Science Mahidol University Enzymes Transport proteins utrient and storage proteins
Various Functions of Proteins SB203 Amino Acids and Proteins Jirundon Yuvaniyama, Ph.D. Department of Biochemistry Faculty of Science Mahidol University Enzymes Transport proteins utrient and storage proteins
Immune system IgGs. Carla Cortinas, Eva Espigulé, Guillem Lopez-Grado, Margalida Roig, Valentina Salas. Group 2
 Immune system IgGs Carla Cortinas, Eva Espigulé, Guillem Lopez-Grado, Margalida Roig, Valentina Salas Group 2 Index 1. Introduction 1.1. 1.2. 1.3. 1.4. 2. Immunoglobulins IgG formation IgG subclasses Structural
Immune system IgGs Carla Cortinas, Eva Espigulé, Guillem Lopez-Grado, Margalida Roig, Valentina Salas Group 2 Index 1. Introduction 1.1. 1.2. 1.3. 1.4. 2. Immunoglobulins IgG formation IgG subclasses Structural
Supplementary Information for. Structure of human tyrosylprotein sulfotransferase-2 reveals the mechanism of protein tyrosine sulfation reaction
 Supplementary Information for Structure of human tyrosylprotein sulfotransferase-2 reveals the mechanism of protein tyrosine sulfation reaction Takamasa Teramoto, Yukari Fujikawa, Yoshirou Kawaguchi, Katsuhisa
Supplementary Information for Structure of human tyrosylprotein sulfotransferase-2 reveals the mechanism of protein tyrosine sulfation reaction Takamasa Teramoto, Yukari Fujikawa, Yoshirou Kawaguchi, Katsuhisa
Homology Modelling. Thomas Holberg Blicher NNF Center for Protein Research University of Copenhagen
 Homology Modelling Thomas Holberg Blicher NNF Center for Protein Research University of Copenhagen Why are Protein Structures so Interesting? They provide a detailed picture of interesting biological features,
Homology Modelling Thomas Holberg Blicher NNF Center for Protein Research University of Copenhagen Why are Protein Structures so Interesting? They provide a detailed picture of interesting biological features,
Supplementary Information. Structural basis for duplex RNA recognition and cleavage by A.
 Supplementary Information Structural asis for duplex RNA recognition and cleavage y A. fulgidus C3PO Eneida arizotto 1, Edward D Lowe 1 & James S Parker 1 1 Department of Biochemistry University of Oxford
Supplementary Information Structural asis for duplex RNA recognition and cleavage y A. fulgidus C3PO Eneida arizotto 1, Edward D Lowe 1 & James S Parker 1 1 Department of Biochemistry University of Oxford
Supplementary Fig. 1. Initial electron density maps for the NOX-D20:mC5a complex obtained after SAD-phasing. (a) Initial experimental electron
 Supplementary Fig. 1. Initial electron density maps for the NOX-D20:mC5a complex obtained after SAD-phasing. (a) Initial experimental electron density map obtained after SAD-phasing and density modification
Supplementary Fig. 1. Initial electron density maps for the NOX-D20:mC5a complex obtained after SAD-phasing. (a) Initial experimental electron density map obtained after SAD-phasing and density modification
Structural basis of a novel PD-L1 nanobody for immune checkpoint blockade
 Structural basis of a novel PD-L1 nanobody for immune checkpoint blockade Supplementary Materials Supplementary methods Table S1-S Figure S1-S 1 1 1 1 1 1 1 1 1 0 1 0 Supplementary Methods Competitive
Structural basis of a novel PD-L1 nanobody for immune checkpoint blockade Supplementary Materials Supplementary methods Table S1-S Figure S1-S 1 1 1 1 1 1 1 1 1 0 1 0 Supplementary Methods Competitive
SUPPLEMENTARY INFORMATION
 Supplementary Table 1. Crystallographic statistics CRM1-SNUPN complex Space group P6 4 22 a=b=250.4, c=190.4 Data collection statistics: CRM1-selenomethionine SNUPN MAD data Peak Inflection Remote Native
Supplementary Table 1. Crystallographic statistics CRM1-SNUPN complex Space group P6 4 22 a=b=250.4, c=190.4 Data collection statistics: CRM1-selenomethionine SNUPN MAD data Peak Inflection Remote Native
Basic concepts of molecular biology
 Basic concepts of molecular biology Gabriella Trucco Email: gabriella.trucco@unimi.it Life The main actors in the chemistry of life are molecules called proteins nucleic acids Proteins: many different
Basic concepts of molecular biology Gabriella Trucco Email: gabriella.trucco@unimi.it Life The main actors in the chemistry of life are molecules called proteins nucleic acids Proteins: many different
Chapter 8. One-Dimensional Structural Properties of Proteins in the Coarse-Grained CABS Model. Sebastian Kmiecik and Andrzej Kolinski.
 Chapter 8 One-Dimensional Structural Properties of Proteins in the Coarse-Grained CABS Model Abstract Despite the significant increase in computational power, molecular modeling of protein structure using
Chapter 8 One-Dimensional Structural Properties of Proteins in the Coarse-Grained CABS Model Abstract Despite the significant increase in computational power, molecular modeling of protein structure using
Case 7 A Storage Protein From Seeds of Brassica nigra is a Serine Protease Inhibitor Last modified 29 September 2005
 Case 7 A Storage Protein From Seeds of Brassica nigra is a Serine Protease Inhibitor Last modified 9 September 005 Focus concept Purification of a novel seed storage protein allows sequence analysis and
Case 7 A Storage Protein From Seeds of Brassica nigra is a Serine Protease Inhibitor Last modified 9 September 005 Focus concept Purification of a novel seed storage protein allows sequence analysis and
SUPPLEMENTARY INFORMATION
 This supplementary information is an extension of the letter with the same title and includes further discussion on the comparison of our designed Fe B Mb (computer model and crystal structure) with the
This supplementary information is an extension of the letter with the same title and includes further discussion on the comparison of our designed Fe B Mb (computer model and crystal structure) with the
Supplementary Materials. Figure S1 Chemical structures of cisplatin and carboplatin.
 Supplementary Materials Figure S1 Chemical structures of cisplatin and carboplatin. Figure S2 (a) RMS difference plot between the RT and 100K structures for the carboplatin_dmso case after 13 months of
Supplementary Materials Figure S1 Chemical structures of cisplatin and carboplatin. Figure S2 (a) RMS difference plot between the RT and 100K structures for the carboplatin_dmso case after 13 months of
Acceleration of protein folding by four orders of magnitude through a single amino acid substitution
 Acceleration of protein folding by four orders of magnitude through a single amino acid substitution Daniel J. A. Roderer 1, Martin A. Schärer 1, Marina Rubini 2 * and Rudi Glockshuber 1 AUTHOR ADDRESS
Acceleration of protein folding by four orders of magnitude through a single amino acid substitution Daniel J. A. Roderer 1, Martin A. Schärer 1, Marina Rubini 2 * and Rudi Glockshuber 1 AUTHOR ADDRESS
DNA/Protein Binding, Molecular Docking and in Vitro Anti-cancer Activity of some Thioether-Dipyrrinato Complexes
 DNA/Protein Binding, Molecular Docking and in Vitro Anti-cancer Activity of some Thioether-Dipyrrinato Complexes Rakesh Kumar Gupta, Gunjan Sharma, ξ Rampal Pandey, Amit Kumar, Biplob Koch, ξ Pei- Zhou
DNA/Protein Binding, Molecular Docking and in Vitro Anti-cancer Activity of some Thioether-Dipyrrinato Complexes Rakesh Kumar Gupta, Gunjan Sharma, ξ Rampal Pandey, Amit Kumar, Biplob Koch, ξ Pei- Zhou
7.014 Solution Set 4
 7.014 Solution Set 4 Question 1 Shown below is a fragment of the sequence of a hypothetical bacterial gene. This gene encodes production of HWDWN, protein essential for metabolizing sugar yummose. The
7.014 Solution Set 4 Question 1 Shown below is a fragment of the sequence of a hypothetical bacterial gene. This gene encodes production of HWDWN, protein essential for metabolizing sugar yummose. The
HEK293T. Fig. 1 in the
 Supplementary Information Supplementary Figure 1 Zinc uptake assay of hzip4 and hzip4-δecd transiently expressed in HEK293T cells. The results of one representative e experiment are shown in Fig. 1 in
Supplementary Information Supplementary Figure 1 Zinc uptake assay of hzip4 and hzip4-δecd transiently expressed in HEK293T cells. The results of one representative e experiment are shown in Fig. 1 in
Supporting Online Material. Av1 and Av2 were isolated and purified under anaerobic conditions according to
 Supporting Online Material Materials and Methods Av1 and Av2 were isolated and purified under anaerobic conditions according to published protocols (S1). Crystals of nf-, pcp- and adp-av2:av1 complexes
Supporting Online Material Materials and Methods Av1 and Av2 were isolated and purified under anaerobic conditions according to published protocols (S1). Crystals of nf-, pcp- and adp-av2:av1 complexes
SUPPLEMENTARY INFORMATION. Reengineering Protein Interfaces Yields Copper-Inducible Ferritin Cage Assembly
 SUPPLEMENTARY INFORMATION Reengineering Protein Interfaces Yields Copper-Inducible Ferritin Cage Assembly Dustin J. E. Huard, Kathleen M. Kane and F. Akif Tezcan* Department of Chemistry and Biochemistry,
SUPPLEMENTARY INFORMATION Reengineering Protein Interfaces Yields Copper-Inducible Ferritin Cage Assembly Dustin J. E. Huard, Kathleen M. Kane and F. Akif Tezcan* Department of Chemistry and Biochemistry,
Figure S1
 Supplementary Figure 1 The distribution of chlorophyll containing complexes eluted from DEAE-cellulose column in a sucrose gradient tube Six pigment-containing bands were resolved and identified as: B1,
Supplementary Figure 1 The distribution of chlorophyll containing complexes eluted from DEAE-cellulose column in a sucrose gradient tube Six pigment-containing bands were resolved and identified as: B1,
Amino Acid Sequences and Evolutionary Relationships
 Amino Acid Sequences and Evolutionary Relationships One technique used to determine evolutionary relationships is to study the biochemical similarity of organisms. Though molds, aardvarks, and humans appear
Amino Acid Sequences and Evolutionary Relationships One technique used to determine evolutionary relationships is to study the biochemical similarity of organisms. Though molds, aardvarks, and humans appear
11 questions for a total of 120 points
 Your Name: BYS 201, Final Exam, May 3, 2010 11 questions for a total of 120 points 1. 25 points Take a close look at these tables of amino acids. Some of them are hydrophilic, some hydrophobic, some positive
Your Name: BYS 201, Final Exam, May 3, 2010 11 questions for a total of 120 points 1. 25 points Take a close look at these tables of amino acids. Some of them are hydrophilic, some hydrophobic, some positive
Algorithms in Bioinformatics ONE Transcription Translation
 Algorithms in Bioinformatics ONE Transcription Translation Sami Khuri Department of Computer Science San José State University sami.khuri@sjsu.edu Biology Review DNA RNA Proteins Central Dogma Transcription
Algorithms in Bioinformatics ONE Transcription Translation Sami Khuri Department of Computer Science San José State University sami.khuri@sjsu.edu Biology Review DNA RNA Proteins Central Dogma Transcription
Thr Gly Tyr. Gly Lys Asn
 Your unique body characteristics (traits), such as hair color or blood type, are determined by the proteins your body produces. Proteins are the building blocks of life - in fact, about 45% of the human
Your unique body characteristics (traits), such as hair color or blood type, are determined by the proteins your body produces. Proteins are the building blocks of life - in fact, about 45% of the human
Laboratory Evolution of Robust and Enantioselective Baeyer-Villiger Monooxygenases for Asymmetric Catalysis
 Laboratory Evolution of Robust and Enantioselective Baeyer-Villiger Monooxygenases for Asymmetric Catalysis Induced fit docking model Manfred T. Reetz* and Sheng Wu Max-Planck-Institut für Kohlenforschung
Laboratory Evolution of Robust and Enantioselective Baeyer-Villiger Monooxygenases for Asymmetric Catalysis Induced fit docking model Manfred T. Reetz* and Sheng Wu Max-Planck-Institut für Kohlenforschung
Colchicine. Colchicine. a b c d e
 α1-tub Colchicine Laulimalide RB3 β1-tub Vinblastine α2-tub Taxol Colchicine β2-tub Maytansine a b c d e Supplementary Figure 1 Structures of microtubule and tubulin (a)the cartoon of head to tail arrangement
α1-tub Colchicine Laulimalide RB3 β1-tub Vinblastine α2-tub Taxol Colchicine β2-tub Maytansine a b c d e Supplementary Figure 1 Structures of microtubule and tubulin (a)the cartoon of head to tail arrangement
Nature Structural & Molecular Biology: doi: /nsmb.2548
 Supplementary Figure 1. Structure of GltPhout. (a) Stereo view of a slice through a single GltPhout protomer shown in stick representation along with 2Fo-Fc and anomalous difference electron maps. The
Supplementary Figure 1. Structure of GltPhout. (a) Stereo view of a slice through a single GltPhout protomer shown in stick representation along with 2Fo-Fc and anomalous difference electron maps. The
BC 367, Exam 2 November 13, Part I. Multiple Choice (3 pts each)- Please circle the single best answer.
 Name BC 367, Exam 2 November 13, 2008 Part I. Multiple Choice (3 pts each)- Please circle the single best answer. 1. The enzyme pyruvate dehydrogenase catalyzes the following reaction. What kind of enzyme
Name BC 367, Exam 2 November 13, 2008 Part I. Multiple Choice (3 pts each)- Please circle the single best answer. 1. The enzyme pyruvate dehydrogenase catalyzes the following reaction. What kind of enzyme
Supplementary materials. Computational study of β-n-acetylhexosaminidase from. Talaromyces flavus, a glycosidase with high substrate flexibility
 Supplementary materials Computational study of β-n-acetylhexosaminidase from Talaromyces flavus, a glycosidase with high substrate flexibility Natallia Kulik 1,*, Kristýna Slámová 2, Rüdiger Ettrich 1,3,
Supplementary materials Computational study of β-n-acetylhexosaminidase from Talaromyces flavus, a glycosidase with high substrate flexibility Natallia Kulik 1,*, Kristýna Slámová 2, Rüdiger Ettrich 1,3,
Structural bioinformatics
 Structural bioinformatics Why structures? The representation of the molecules in 3D is more informative New properties of the molecules are revealed, which can not be detected by sequences Eran Eyal Plant
Structural bioinformatics Why structures? The representation of the molecules in 3D is more informative New properties of the molecules are revealed, which can not be detected by sequences Eran Eyal Plant
Virtual bond representation
 Today s subjects: Virtual bond representation Coordination number Contact maps Sidechain packing: is it an instrumental way of selecting and consolidating a fold? ASA of proteins Interatomic distances
Today s subjects: Virtual bond representation Coordination number Contact maps Sidechain packing: is it an instrumental way of selecting and consolidating a fold? ASA of proteins Interatomic distances
Programme Good morning and summary of last week Levels of Protein Structure - I Levels of Protein Structure - II
 Programme 8.00-8.10 Good morning and summary of last week 8.10-8.30 Levels of Protein Structure - I 8.30-9.00 Levels of Protein Structure - II 9.00-9.15 Break 9.15-11.15 Exercise: Building a protein model
Programme 8.00-8.10 Good morning and summary of last week 8.10-8.30 Levels of Protein Structure - I 8.30-9.00 Levels of Protein Structure - II 9.00-9.15 Break 9.15-11.15 Exercise: Building a protein model
Description of Changes and Corrections for PDB File Format Version 4.0. Provisional Document April 12, 2011
 Description of Changes and Corrections for PDB File Format Version 4.0 Provisional Document April 12, 2011 The wwpdb has reviewed the PDB archive and created a new set of corrected files that will be released
Description of Changes and Corrections for PDB File Format Version 4.0 Provisional Document April 12, 2011 The wwpdb has reviewed the PDB archive and created a new set of corrected files that will be released
Amino Acid Sequences and Evolutionary Relationships
 Amino Acid Sequences and Evolutionary Relationships Pre-Lab Discussion Homologous structures -- those structures believed to have a common origin but not necessarily a common function -- provide some of
Amino Acid Sequences and Evolutionary Relationships Pre-Lab Discussion Homologous structures -- those structures believed to have a common origin but not necessarily a common function -- provide some of
Protein analysis. Dr. Mamoun Ahram Summer semester, Resources This lecture Campbell and Farrell s Biochemistry, Chapters 5
 Protein analysis Dr. Mamoun Ahram Summer semester, 2015-2016 Resources This lecture Campbell and Farrell s Biochemistry, Chapters 5 Bases of protein separation Proteins can be purified on the basis Solubility
Protein analysis Dr. Mamoun Ahram Summer semester, 2015-2016 Resources This lecture Campbell and Farrell s Biochemistry, Chapters 5 Bases of protein separation Proteins can be purified on the basis Solubility
Computational Methods for Protein Structure Prediction
 Computational Methods for Protein Structure Prediction Ying Xu 2017/12/6 1 Outline introduction to protein structures the problem of protein structure prediction why it is possible to predict protein structures
Computational Methods for Protein Structure Prediction Ying Xu 2017/12/6 1 Outline introduction to protein structures the problem of protein structure prediction why it is possible to predict protein structures
Molecular design principles underlying β-strand swapping. in the adhesive dimerization of cadherins
 Supplementary information for: Molecular design principles underlying β-strand swapping in the adhesive dimerization of cadherins Jeremie Vendome 1,2,3,5, Shoshana Posy 1,2,3,5,6, Xiangshu Jin, 1,3 Fabiana
Supplementary information for: Molecular design principles underlying β-strand swapping in the adhesive dimerization of cadherins Jeremie Vendome 1,2,3,5, Shoshana Posy 1,2,3,5,6, Xiangshu Jin, 1,3 Fabiana
Solutions to Problem Set 1
 MIT Department of Biology 7.014 Introductory Biology, Spring 004 Question 1 Solutions to 7.014 Problem Set 1 a) Describe the conditions of the atmosphere on prebiotic earth and how these conditions differ
MIT Department of Biology 7.014 Introductory Biology, Spring 004 Question 1 Solutions to 7.014 Problem Set 1 a) Describe the conditions of the atmosphere on prebiotic earth and how these conditions differ
7.014 Quiz II Handout
 7.014 Quiz II Handout Quiz II: Wednesday, March 17 12:05-12:55 54-100 **This will be a closed book exam** Quiz Review Session: Friday, March 12 7:00-9:00 pm room 54-100 Open Tutoring Session: Tuesday,
7.014 Quiz II Handout Quiz II: Wednesday, March 17 12:05-12:55 54-100 **This will be a closed book exam** Quiz Review Session: Friday, March 12 7:00-9:00 pm room 54-100 Open Tutoring Session: Tuesday,
Speaker: Yu-Chen Ku Professor: Ching-Tsan Huang, Ph. D. Source: Biochemical Journal (2007) 402, Date: December 4th, 2007
 Directed evolution and structural analysis of N-carbamoyl-D-amino acid amidohydrolase provide insights into recombinant protein solubility in Escherichia coli Speaker: Yu-Chen Ku Professor: Ching-Tsan
Directed evolution and structural analysis of N-carbamoyl-D-amino acid amidohydrolase provide insights into recombinant protein solubility in Escherichia coli Speaker: Yu-Chen Ku Professor: Ching-Tsan
doi: /nature09408 Figure S1
 doi:10.1038/nature09408 A Figure S1 www.nature.com/nature 1 RESEARCH SUPPLEMENTARY INFORMATION Figure S1 Primary sequence, and structure of NorM VC. A. Amino acid sequence alignment of NorM VC with selected
doi:10.1038/nature09408 A Figure S1 www.nature.com/nature 1 RESEARCH SUPPLEMENTARY INFORMATION Figure S1 Primary sequence, and structure of NorM VC. A. Amino acid sequence alignment of NorM VC with selected
2018 Protein Modeling Exam Key
 2018 Protein Modeling Exam Key Multiple Choice: 1. Which of the following amino acids has a negative charge at ph 7? a. Gln b. Glu c. Ser d. Cys 2. Which of the following is an example of secondary structure?
2018 Protein Modeling Exam Key Multiple Choice: 1. Which of the following amino acids has a negative charge at ph 7? a. Gln b. Glu c. Ser d. Cys 2. Which of the following is an example of secondary structure?
LIST OF ACRONYMS & ABBREVIATIONS
 LIST OF ACRONYMS & ABBREVIATIONS ALA ARG ASN ATD CRD CYS GLN GLU GLY GPCR HIS hstr ILE LEU LYS MET mglur1 NHDC PDB PHE PRO SER T1R2 T1R3 TMD TRP TYR THR 7-TM VFTM ZnSO 4 Alanine, A Arginine, R Asparagines,
LIST OF ACRONYMS & ABBREVIATIONS ALA ARG ASN ATD CRD CYS GLN GLU GLY GPCR HIS hstr ILE LEU LYS MET mglur1 NHDC PDB PHE PRO SER T1R2 T1R3 TMD TRP TYR THR 7-TM VFTM ZnSO 4 Alanine, A Arginine, R Asparagines,
Fundamentals of Protein Structure
 Outline Fundamentals of Protein Structure Yu (Julie) Chen and Thomas Funkhouser Princeton University CS597A, Fall 2005 Protein structure Primary Secondary Tertiary Quaternary Forces and factors Levels
Outline Fundamentals of Protein Structure Yu (Julie) Chen and Thomas Funkhouser Princeton University CS597A, Fall 2005 Protein structure Primary Secondary Tertiary Quaternary Forces and factors Levels
The Stringent Response
 The Stringent Response When amino acids are limiting a response is triggered to shut down a wide range of biosynthetic processes This process is called the Stringent Response It results in the synthesis
The Stringent Response When amino acids are limiting a response is triggered to shut down a wide range of biosynthetic processes This process is called the Stringent Response It results in the synthesis
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Multiple sequence alignments of four Swi2/Snf2 subfamily proteins, ScChd1, SsoRad54 and the RNA helicase Vasa. The sequence alignments of the Swi2/Snf2 subfamily proteins, ScChd1
Supplementary Figure 1 Multiple sequence alignments of four Swi2/Snf2 subfamily proteins, ScChd1, SsoRad54 and the RNA helicase Vasa. The sequence alignments of the Swi2/Snf2 subfamily proteins, ScChd1
SUPPLEMENTARY INFORMATION
 Structure of a tyrosyl-trna synthetase splicing factor bound to a group I intron RNA Paul J. Paukstelis 1, Jui-Hui Chen 2, Elaine Chase 2, Alan M. Lambowitz 1,*, and Barbara L. Golden 2,*,. 1 Institute
Structure of a tyrosyl-trna synthetase splicing factor bound to a group I intron RNA Paul J. Paukstelis 1, Jui-Hui Chen 2, Elaine Chase 2, Alan M. Lambowitz 1,*, and Barbara L. Golden 2,*,. 1 Institute
Conformational changes in IgE contribute to its. uniquely slow dissociation rate from receptor FcεRI
 Conformational changes in IgE contribute to its uniquely slow dissociation rate from receptor FcεRI M.D. Holdom, A.M. Davies, J.E. Nettleship, S.C. Bagby, B. Dhaliwal, E. Girardi, J. Hunt, H.J. Gould,
Conformational changes in IgE contribute to its uniquely slow dissociation rate from receptor FcεRI M.D. Holdom, A.M. Davies, J.E. Nettleship, S.C. Bagby, B. Dhaliwal, E. Girardi, J. Hunt, H.J. Gould,
Diversity in DNA recognition by p53 revealed by crystal structures with Hoogsteen base pairs
 SUPPLEMENTARY INFORMATION Diversity in DNA recognition by p53 revealed by crystal structures with Hoogsteen base pairs Malka Kitayner 1, 3, Haim Rozenberg 1, 3, Remo Rohs 2, 3, Oded Suad 1, Dov Rabinovich
SUPPLEMENTARY INFORMATION Diversity in DNA recognition by p53 revealed by crystal structures with Hoogsteen base pairs Malka Kitayner 1, 3, Haim Rozenberg 1, 3, Remo Rohs 2, 3, Oded Suad 1, Dov Rabinovich
Homework. A bit about the nature of the atoms of interest. Project. The role of electronega<vity
 Homework Why cited articles are especially useful. citeulike science citation index When cutting and pasting less is more. Project Your protein: I will mail these out this weekend If you haven t gotten
Homework Why cited articles are especially useful. citeulike science citation index When cutting and pasting less is more. Project Your protein: I will mail these out this weekend If you haven t gotten
Supplementary materials for Structure of an open clamp type II topoisomerase-dna complex provides a mechanism for DNA capture and transport
 Supplementary materials for Structure of an open clamp type II topoisomerase-dna complex provides a mechanism for DNA capture and transport Ivan Laponogov 1,2, Dennis A. Veselkov 1, Isabelle M-T. Crevel
Supplementary materials for Structure of an open clamp type II topoisomerase-dna complex provides a mechanism for DNA capture and transport Ivan Laponogov 1,2, Dennis A. Veselkov 1, Isabelle M-T. Crevel
Abbreviations. Inhibitor synthesis
 Supplementary Material I for: Bailey et al. manuscript An analysis of subdomain orientation, conformational change and disorder in relation to crystal packing of aspartic proteinases. Abbreviations Boc
Supplementary Material I for: Bailey et al. manuscript An analysis of subdomain orientation, conformational change and disorder in relation to crystal packing of aspartic proteinases. Abbreviations Boc
The molecular basis of lysine 48 ubiquitin chain synthesis by Ube2K
 Supplementary Information The molecular basis of lysine 48 ubiquitin chain synthesis by Adam J. Middleton, Catherine L. Day* Department of Biochemistry, Otago School of Medical Sciences, University of
Supplementary Information The molecular basis of lysine 48 ubiquitin chain synthesis by Adam J. Middleton, Catherine L. Day* Department of Biochemistry, Otago School of Medical Sciences, University of
human Cdc45 Figure 1c. (c)
 1 Details of the refined crystallographic model of human Cdc45 and comparison of its active-site region with that of bacterial RecJ. (a) Stereo view of a representative example of the final 2F o -F c electron
1 Details of the refined crystallographic model of human Cdc45 and comparison of its active-site region with that of bacterial RecJ. (a) Stereo view of a representative example of the final 2F o -F c electron
SUPPLEMENTARY MATERIAL
 SUPPLEMENTARY MATERIAL Purification and biochemical characterization of acid phosphatase-i from seeds of Nelumbo nucifera Sanaullah Khan a*, Shahnaz Asmat c, Sajida Batool a, Mushtaq Ahmed b a Department
SUPPLEMENTARY MATERIAL Purification and biochemical characterization of acid phosphatase-i from seeds of Nelumbo nucifera Sanaullah Khan a*, Shahnaz Asmat c, Sajida Batool a, Mushtaq Ahmed b a Department
Name Section Problem Set 3
 Name Section 7.013 Problem Set 3 The completed problem must be turned into the wood box outside 68120 by 4:40 pm, Thursday, March 13. Problem sets will not be accepted late. Question 1 Based upon your
Name Section 7.013 Problem Set 3 The completed problem must be turned into the wood box outside 68120 by 4:40 pm, Thursday, March 13. Problem sets will not be accepted late. Question 1 Based upon your
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
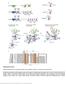 Supplementary Figure 1 Domain architecture and conformational states of the decapping complex, as revealed by structural studies. (a) Domain organization of Schizosaccharomyces pombe (Sp) and Saccharomyces
Supplementary Figure 1 Domain architecture and conformational states of the decapping complex, as revealed by structural studies. (a) Domain organization of Schizosaccharomyces pombe (Sp) and Saccharomyces
Packing of Secondary Structures
 7.88 Lecture Notes - 5 7.24/7.88J/5.48J The Protein Folding and Human Disease Packing of Secondary Structures Packing of Helices against sheets Packing of sheets against sheets Parallel Orthogonal Table:
7.88 Lecture Notes - 5 7.24/7.88J/5.48J The Protein Folding and Human Disease Packing of Secondary Structures Packing of Helices against sheets Packing of sheets against sheets Parallel Orthogonal Table:
Lecture 7: Affinity Chromatography-II
 Lecture 7: Affinity Chromatography-II We have studied basics of affinity purification during last lecture. The current lecture is continuation of last lecture and we will cover following: 1. Few specific
Lecture 7: Affinity Chromatography-II We have studied basics of affinity purification during last lecture. The current lecture is continuation of last lecture and we will cover following: 1. Few specific
Crystal Structures of the sugar epimerase for rare sugar production in complexes with deoxy sugars
 The nd International Symposium Space Science of High Quality Protein Crystallization Technology Crystal Structures of the sugar epimerase for rare sugar production in complexes with deoxy sugars ~ Crystal
The nd International Symposium Space Science of High Quality Protein Crystallization Technology Crystal Structures of the sugar epimerase for rare sugar production in complexes with deoxy sugars ~ Crystal
Problem Set Unit The base ratios in the DNA and RNA for an onion (Allium cepa) are given below.
 Problem Set Unit 3 Name 1. Which molecule is found in both DNA and RNA? A. Ribose B. Uracil C. Phosphate D. Amino acid 2. Which molecules form the nucleotide marked in the diagram? A. phosphate, deoxyribose
Problem Set Unit 3 Name 1. Which molecule is found in both DNA and RNA? A. Ribose B. Uracil C. Phosphate D. Amino acid 2. Which molecules form the nucleotide marked in the diagram? A. phosphate, deoxyribose
CFSSP: Chou and Fasman Secondary Structure Prediction server
 Wide Spectrum, Vol. 1, No. 9, (2013) pp 15-19 CFSSP: Chou and Fasman Secondary Structure Prediction server T. Ashok Kumar Department of Bioinformatics, Noorul Islam College of Arts and Science, Kumaracoil
Wide Spectrum, Vol. 1, No. 9, (2013) pp 15-19 CFSSP: Chou and Fasman Secondary Structure Prediction server T. Ashok Kumar Department of Bioinformatics, Noorul Islam College of Arts and Science, Kumaracoil
High-resolution structure of a papaya plant-defence barwin-like protein solved by in-house sulphur-sad phasing
 High-resolution structure of a papaya plant-defence barwin-like protein solved by in-house sulphur-sad phasing Joëlle Huet, Emmanuel Jean Teinkela Mbosso, Sameh Soror, Franck Meyer, Yvan Looze, René Wintjens,
High-resolution structure of a papaya plant-defence barwin-like protein solved by in-house sulphur-sad phasing Joëlle Huet, Emmanuel Jean Teinkela Mbosso, Sameh Soror, Franck Meyer, Yvan Looze, René Wintjens,
If you wish to have extra practice with swiss pdb viewer or to familiarize yourself with how to use the program here is a tutorial:
 Name (s): Swiss PDB viewer assignment chapter 4. If you wish to have extra practice with swiss pdb viewer or to familiarize yourself with how to use the program here is a tutorial: http://spdbv.vital-it.ch/themolecularlevel/spvtut/index.html
Name (s): Swiss PDB viewer assignment chapter 4. If you wish to have extra practice with swiss pdb viewer or to familiarize yourself with how to use the program here is a tutorial: http://spdbv.vital-it.ch/themolecularlevel/spvtut/index.html
Basic concepts of molecular biology
 Basic concepts of molecular biology Gabriella Trucco Email: gabriella.trucco@unimi.it What is life made of? 1665: Robert Hooke discovered that organisms are composed of individual compartments called cells
Basic concepts of molecular biology Gabriella Trucco Email: gabriella.trucco@unimi.it What is life made of? 1665: Robert Hooke discovered that organisms are composed of individual compartments called cells
Reduced structural flexibility for exonuclease deficient DNA polymerase III mutant
 Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 2018 Reduced structural flexibility for exonuclease deficient DNA polymerase III mutant
Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 2018 Reduced structural flexibility for exonuclease deficient DNA polymerase III mutant
1) The penicillin family of antibiotics, discovered by Alexander Fleming in 1928, has the following general structure: O O
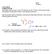 ame: TF ame: LS1a Fall 06 Problem Set #3 Due Friday 10/13 at noon in your TF s drop box on the 2 nd floor of the Science Center All questions including the (*extra*) ones should be turned in 1) The penicillin
ame: TF ame: LS1a Fall 06 Problem Set #3 Due Friday 10/13 at noon in your TF s drop box on the 2 nd floor of the Science Center All questions including the (*extra*) ones should be turned in 1) The penicillin
 Problem: The GC base pairs are more stable than AT base pairs. Why? 5. Triple-stranded DNA was first observed in 1957. Scientists later discovered that the formation of triplestranded DNA involves a type
Problem: The GC base pairs are more stable than AT base pairs. Why? 5. Triple-stranded DNA was first observed in 1957. Scientists later discovered that the formation of triplestranded DNA involves a type
Alpha-helices, beta-sheets and U-turns within a protein are stabilized by (hint: two words).
 1 Quiz1 Q1 2011 Alpha-helices, beta-sheets and U-turns within a protein are stabilized by (hint: two words) Value Correct Answer 1 noncovalent interactions 100% Equals hydrogen bonds (100%) Equals H-bonds
1 Quiz1 Q1 2011 Alpha-helices, beta-sheets and U-turns within a protein are stabilized by (hint: two words) Value Correct Answer 1 noncovalent interactions 100% Equals hydrogen bonds (100%) Equals H-bonds
IB HL Biology Test: Topics 1 and 3
 October 26, 2011 IB HL Biology Test: Topics 1 and 3 Multiple Choice Identify the choice that best completes the statement or answers the question. 1. What conditions must be met for the t-test to be applied?
October 26, 2011 IB HL Biology Test: Topics 1 and 3 Multiple Choice Identify the choice that best completes the statement or answers the question. 1. What conditions must be met for the t-test to be applied?
