BIOLOGY. Chapter 16 GenesExpression
|
|
|
- Derrick Boyd
- 6 years ago
- Views:
Transcription
1 BIOLOGY Chapter 16 GenesExpression
2 CAMPBELL BIOLOGY TENTH EDITION Reece Urry Cain Wasserman Minorsky Jackson 18 Gene Expression 2014 Pearson Education, Inc.
3 Figure 16.1 Differential Gene Expression results in you Genetic content is the same, Not all genes are expressed in every cell differential gene expression patterns
4 Fig / 16.1 Control of Gene Differential Gene Expression
5 Figure 18.2 Precursor Remember Negative Feedback Regulation? Feedback inhibition Enzyme 1 trpe Enzyme 2 trpd trpc Regulation of gene expression Enzyme 3 trpb trpa Tryptophan (a) Regulation of enzyme activity (b) Regulation of enzyme production
6 Figure 18.3 / 16.3 Prokaryotic control of gene expression Promoter mrna 5 trpr Regulatory gene 3 RNA polymerase Promoter Operator Start codon mrna 5 trp operon Genes of operon trpe trpd trpc trpb trpa Stop codon Protein Inactive repressor (a) Tryptophan absent, repressor inactive, operon on E D C B A Polypeptide subunits that make up enzymes for tryptophan synthesis mrna 5 Protein trpr 3 Tryptophan (corepressor) (b) Tryptophan present, repressor active, operon off trpe Active repressor No RNA made Bacterial Gene Expression Control: Operon: cluster of genes & includes: Promoter region (TATA/operator) Operator (on/off switch) Genes being expressed Two types of Operons Repressible always ON Repressor inactive Inducible always OFF Repressor active Corepressor Regulatory Gene upstream
7 Figure 18.3a Tryptophan (Trp) operon Repressible Operon Always ON Repressor inactive transcription of genes enzyme for tryptophan synthesis trp operon Promoter Regulatory gene Promoter Genes of operon trpr trpe trpd trpc trpb trpa mrna 5 3 RNA polymerase Operator Start codon mrna 5 Stop codon Protein Inactive repressor (a) Tryptophan absent, repressor inactive, operon on E D C B A Polypeptide subunits that make up enzymes for tryptophan synthesis
8 Figure 18.3b Trp operon Repressible Operon Corepressor + Active repressor turned OFF (repressed) Too much tryptophan trp acts as a corepressor, activating repressor mrna 5 trpr 3 trpe No RNA made Protein Tryptophan (corepressor) Active repressor (b) Tryptophan present, repressor active, operon off
9 Figure 18.4 / 16.4 & 16.5 Regulatory gene l a Ic Promoter Operator Lactose operon Inducible Operon Always OFF IacZ mrna 5 3 RNA polymerase No RNA made Protein Active repressor (a) Lactose absent, repressor active, operon off l a Ic lac operon lacz lacy laca mrna RNA polymerase Start codon Stop codon mrna 5 Protein β-galactosidase Permease Transacetylase Allolactose (inducer) Inactive repressor (b) Lactose present, repressor inactive, operon on
10 Figure 18.4a lac operon: Inducible Operon Active repressor genes turned OFF No Lactose no gene expression, no enzymes to breakdown lactose Regulatory gene Promoter Operator lac I IacZ mrna 5 3 RNA polymerase No RNA made Protein Active repressor (a) Lactose absent, repressor active, operon off
11 Figure 18.4b lac operon: Inducible Operon INactive repressor genes turned ON Lactose (Inducer) present repressor inactive gene expression (induced) lac operon lac I lacz lacy laca mrna 3 RNA polymerase Start codon Stop codon 5 mrna 5 Protein β-galactosidase Permease Transacetylase Inactive repressor Allolactose (inducer) (b) Lactose present, repressor inactive, operon on
12 Figure 18.5 / 16.4 Positive control of lac operon by Catabolite Activator Protein (CAP) lac I CAP-binding site camp Promoter Active CAP Operator RNA polymerase binds and transcribes lacz Inactive CAP Allolactose Inactive lac repressor (a) Lactose present, glucose scarce (camp level high): abundant lac mrna synthesized Promoter lac I lacz CAP-binding site Inactive CAP Operator RNA polymerase less likely to bind Inactive lac repressor (b) Lactose present, glucose present (camp level low): little lac mrna synthesized
13 Figure 18.5a / 16.4 lac I Promoter Positive control of lac operon by Catabolite Activator Protein (CAP) Operator lacz CAP-binding site camp Active CAP RNA polymerase binds and transcribes Inactive CAP Allolactose Inactive lac repressor (a) Lactose present, glucose scarce (camp level high): abundant lac mrna synthesized CAP helps regulate operons (increases when glucose is low)
14 Figure 18.5b Positive control of the lac operon by CAP Positive control of lac operon by Catabolite Activator Protein (CAP) lac I Promoter lacz CAP-binding site Inactive CAP Operator RNA polymerase less likely to bind Inactive lac repressor (b) Lactose present, glucose present (camp level low): little lac mrna synthesized CAP helps regulate operons (when glucose is high, CAP detaches) When glucose levels increase, CAP detaches, normal expression
15 Fig Promoter Lac operon inducible operon Operator laci CAP-binding site camp Active CAP lacz RNA polymerase binds and transcribes Inactive CAP Allolactose Inactive lac repressor (a) Lactose present, glucose scarce (camp level high): abundant lac mrna synthesized laci CAP-binding site Inactive CAP Promoter Operator lacz RNA polymerase less likely to bind Inactive lac repressor (b) Lactose present, glucose present (camp level low): little lac mrna synthesized
16 Figure 16.5 Transcription of the lac operon regulated Expression only occurs when: glucose is limited lactose is present (alternative fuel source)
17 Figure 18.6 Control of gene expression Signal Chromatin Chromatin modification: unpacking Gene available for transcription Transcription Cap RNA NUCLEUS CYTOPLASM Degradation of mrna Exon Intron Primary transcript RNA processing Tail mrna in nucleus Transport to cytoplasm mrna in cytoplasm Translation Polypeptide Protein processing Degradation of protein Active protein Transport to cellular destination Cellular function (such as enzymatic activity or structural support)
18 Figure 18.6 Control of gene expression Signal Chromatin Chromatin modification: unpacking Gene available for transcription 1) Chromatin modification (epigenetics) 2) Pre-transcription (transcriptional factor & activators) 3) Post-transcription (RNA Processing) 4) Pre-translation & mrna degradation 5) Post-translation (protein processing) 6) Protein degradation Cap RNA NUCLEUS CYTOPLASM Degradation of mrna Exon Intron Transcription Primary transcript RNA processing Tail mrna in nucleus Transport to cytoplasm mrna in cytoplasm Translation Polypeptide Protein processing Degradation of protein Active protein Transport to cellular destination Cellular function (such as enzymatic activity or structural support)
19 Figure 18.6a Control of gene expression Signal Chromatin 1) Chromatin modification Histone acetylation methylation Epigenetic inheritance Gene available for transcription Cap RNA Chromatin modification: unpacking Exon Intron Transcription Primary transcript RNA processing Tail mrna in nucleus NUCLEUS Transport to cytoplasm CYTOPLASM
20 Figure 18.7 / 16.7 Histone Acetylation favors transcription Histone tails Amino acids available for chemical modification double helix Nucleosome (end view) (a) Histone tails protrude outward from a nucleosome Acetyl groups Unacetylated histones (side view) Acetylated histones (b) Acetylation of histone tails promotes loose chromatin structure that permits transcription
21 Figure 16.8 Epigenetics
22 Epigenetic & Methylation Chromosome 15 Angelman Syndrome Maternal expressed Paternal silenced Prader-Willi Syndrome Paternal expressed Maternal silenced (deletion)
23 Figure 18.6a Control of gene expression Signal Chromatin 1) Chromatin modification 2) Pre-transcription Transcription initiation Chromatin modification: unpacking Gene available for transcription Transcription Cap RNA Exon Intron Primary transcript RNA processing Tail mrna in nucleus NUCLEUS Transport to cytoplasm CYTOPLASM
24 Figure 18.8 Control of gene expression 2) Regulation of Transcription Initiation (pre-transcription) Transcriptional factors bind to enhancers Transcriptional factors bind Enhancer (group of distal control elements) Proximal control elements Transcription start site Exon Intron Exon Intron Poly-A signal sequence Exon Transcription termination region Upstream Primary RNA transcript (pre-mrna) Promoter 5 Exon Intron Exon Transcription Intron Downstream Poly-A signal Exon Cleaved 3 end of primary transcript Intron RNA RNA processing Coding segment mrna G P P P 5 Cap 5 UTR Start codon Stop codon AAA AAA 3 UTR Poly-A tail 3
25 Figure / 16.9 Activators Promoter Enhancer Distal control element TATA box Gene Control of Gene Expression 2) Regulation of Transcription Initiation (pre-transcription) Transcriptional factors bind to enhancers Activators stimulates transcription bending protein General transcription factors Group of mediator proteins RNA polymerase II RNA polymerase II Transcription initiation complex RNA synthesis
26 Figure Cell specific transcription in both cells contains the albumin gene and the crystallin gene: Control elements Enhancer for albumin gene Promoter Enhancer for crystallin gene Promoter Albumin gene Crystallin gene LIVER CELL NUCLEUS Available activators LENS CELL NUCLEUS Available activators Albumin gene expressed Albumin gene not expressed (a) Liver cell Crystallin gene not expressed (b) Lens cell Crystallin gene expressed
27 Figure 18.6a Control of gene expression Signal Chromatin 1) Chromatin modification 2) Pre-transcription Transcription initiation 3) Post-transcription RNA Processing Chromatin modification: unpacking Gene available for transcription Transcription Cap RNA Exon Intron Primary transcript RNA processing Tail mrna in nucleus NUCLEUS Transport to cytoplasm CYTOPLASM
28 Fig Control of Gene Expression 3) Post-transcription RNA processing Enhancer (distal control elements) Upstream Proximal control elements Promoter Exon Intron Exon Poly-A signal sequence Termination region Intron Exon Transcription Downstream Primary RNA transcript 5 Exon Intron Exon Intron Exon RNA processing Cleaved 3 end of primary transcript Intron RNA Poly-A signal Coding segment mrna 5 Cap 5 UTR Start codon Stop codon 3 UTR Poly-A tail 3
29 Post-transcription alternative splicing & mrna degradation Fig / Exons Troponin T gene Primary RNA transcript RNA splicing mrna or
30 Figure There are five basic modes of alternative splicing.
31 Figure 18.6b Control of gene expression CYTOPLASM mrna in cytoplasm Degradation of mrna Translation 1) Chromatin modification 2) Pre-transcription Transcription initiation 3) Post-transcription RNA Processing 4) Pre-translation Degradation of protein Translation initiation & mrna degradation Polypeptide Protein processing Active protein Transport to cellular destination Cellular function (such as enzymatic activity or structural support)
32 Fig Hairpin mirna Hydrogen bond Dicer 5 3 (a) Primary mirna transcript mirna mirnaprotein complex Pre-translation & mrna degradation RNAi RNA interference caused by: mirna blocks translation sirna blocks transcription Both: degrade mrna & chromatin modification mrna degraded Translation blocked (b) Generation and function of mirnas
33 Figure mirna mirnaprotein complex 1 The mirna binds to a target mrna. OR mrna degraded Translation blocked 2 If bases are completely complementary, mrna is degraded. If match is less than complete, translation is blocked.
34 Figure 18.6b Control of gene expression CYTOPLASM mrna in cytoplasm 1) Chromatin modification 2) Pre-transcription 3) Post-transcription 4) Pre-translation 5) Protein Processing Degradation of mrna Degradation of protein Translation Polypeptide Protein processing Active protein Transport to cellular destination Cellular function (such as enzymatic activity or structural support)
35 Figure 18.6b Control of gene expression CYTOPLASM mrna in cytoplasm 1) Chromatin modification 2) Pre-transcription 3) Post-transcription 4) Pre-translation 5) Protein Processing 6) Protein Degradation Degradation of mrna Degradation of protein Translation Polypeptide Protein processing Active protein Transport to cellular destination Cellular function (such as enzymatic activity or structural support)
36 Fig / Post-translation protein degradation Ubiquitin Proteasome Proteasome and ubiquitin to be recycled Protein to be degraded Ubiquitinated protein Protein entering a proteasome Protein fragments (peptides)
37 Fig Nucleus Master regulatory gene myod Other muscle-specific genes Embryonic precursor cell OFF OFF mrna OFF Myoblast (determined) MyoD protein (transcription factor) mrna mrna mrna mrna Part of a muscle fiber (fully differentiated cell) MyoD Another transcription factor Myosin, other muscle proteins, and cell cycle blocking proteins
38 Fig Nucleus Master regulatory gene myod Determination & Differentiation of cells Other muscle-specific genes Embryonic precursor cell OFF OFF mrna OFF Myoblast (determined) MyoD protein (transcription factor) mrna mrna mrna mrna Part of a muscle fiber (fully differentiated cell) MyoD Another transcription factor Myosin, other muscle proteins, and cell cycle blocking proteins
39 Fig Growth factor 3 G protein GTP Ras GTP Ras MUTATION Hyperactive Ras protein (product of oncogene) issues signals on its own 2 Receptor 4 Protein kinases (phosphorylation cascade) 5 Transcription factor (activator) NUCLEUS Gene expression Protein that stimulates the cell cycle (a) Cell cycle stimulating pathway 2 Protein kinases MUTATION UV light 3 Active form of p53 Defective or missing transcription factor, such as p53, cannot activate transcription 1 damage in genome Protein that inhibits the cell cycle (b) Cell cycle inhibiting pathway EFFECTS OF MUTATIONS Protein overexpressed Protein absent Cell cycle overstimulated Increased cell division Cell cycle not inhibited (c) Effects of mutations
40 Fig c EFFECTS OF MUTATIONS Protein overexpressed Protein absent Cell cycle overstimulated Increased cell division Cell cycle not inhibited (c) Effects of mutations
41 Fig Colon EFFECTS OF MUTATIONS Colon wall 1 Loss of tumorsuppressor gene APC (or other) 2 Activation of ras oncogene 4 Loss of tumor-suppressor gene p53 Normal colon epithelial cells Small benign growth (polyp) 3 Loss of tumor-suppressor gene DCC Larger benign growth (adenoma) 5 Additional mutations Malignant tumor (carcinoma)
42 Fig
43 Fig. 18-UN1 Operon Promoter Operator RNA polymerase A Genes B C A B C Polypeptides
44 Fig. 18-UN2 Promoter Genes expressed Operator Genes Inactive repressor: no corepressor present Genes not expressed Corepressor Active repressor: corepressor bound
45 Fig. 18-UN3 Genes not expressed Promoter Genes expressed Operator Genes Active repressor: no inducer present Inactive repressor: inducer bound
46 Figure 18.UN09 Chromatin modification Genes in highly compacted chromatin are generally not transcribed. Histone acetylation seems to loosen chromatin structure, enhancing transcription. methylation generally reduces transcripton. Transcription Regulation of transcription initiation: control elements in enhancers bind specific transcription factors. Bending of the enables activators to contact proteins at the promoter, initiating transcription. Coordinate regulation: CHROMATIN MODIFICATION Enhancer for liver-specific genes Enhancer for lens-specific genes mrna DEGRADATION TRANSCRIPTION RNA PROCESSING TRANSLATION PROTEIN PROCESSING AND DEGRADATION mrna degradation Each mrna has a characteristic life span, determined in part by sequences in the 5 and 3 UTRs. RNA processing Alternative RNA splicing: Primary RNA transcript mrna OR Translation Initiation of translation can be controlled via regulation of initiation factors. Protein processing and degradation Protein processing and degradation are subject to regulation.
47 Figure 18.UN09b Chromatin modification Genes in highly compacted chromatin are generally not transcribed. Histone acetylation seems to loosen chromatin structure, enhancing transcription. methylation generally reduces transcription. mrna degradation Each mrna has a characteristic life span, determined in part by sequences in the 5 and 3 UTRs. RNA processing Alternative RNA splicing: Primary RNA transcript mrna OR Translation Initiation of translation can be controlled via regulation of initiation factors. Protein processing and degradation Protein processing and degradation are subject to regulation.
48 Fig. 18-UN4 Chromatin modification Genes in highly compacted chromatin are generally not transcribed. Histone acetylation seems to loosen chromatin structure, enhancing transcription. Transcription Regulation of transcription initiation: control elements bind specific transcription factors. methylation generally reduces transcription. Bending of the enables activators to contact proteins at the promoter, initiating transcription. Coordinate regulation: Enhancer for liver-specific genes Enhancer for lens-specific genes Chromatin modification mrna degradation Transcription RNA processing Translation Primary RNA transcript mrna RNA processing Alternative RNA splicing: or Protein processing and degradation mrna degradation Each mrna has a characteristic life span, determined in part by sequences in the 5 and 3 UTRs. Translation Initiation of translation can be controlled via regulation of initiation factors. Protein processing and degradation Protein processing and degradation by proteasomes are subject to regulation.
49 Fig. 18-UN5 Chromatin modification Transcription Chromatin modification Small RNAs can promote the formation of heterochromatin in certain regions, blocking transcription. RNA processing Translation mirna or sirna can block the translation of specific mrnas. mrna degradation Translation Protein processing and degradation mrna degradation mirna or sirna can target specific mrnas for destruction.
Chapter 18: Regulation of Gene Expression. 1. Gene Regulation in Bacteria 2. Gene Regulation in Eukaryotes 3. Gene Regulation & Cancer
 Chapter 18: Regulation of Gene Expression 1. Gene Regulation in Bacteria 2. Gene Regulation in Eukaryotes 3. Gene Regulation & Cancer Gene Regulation Gene regulation refers to all aspects of controlling
Chapter 18: Regulation of Gene Expression 1. Gene Regulation in Bacteria 2. Gene Regulation in Eukaryotes 3. Gene Regulation & Cancer Gene Regulation Gene regulation refers to all aspects of controlling
Regulation of Gene Expression
 Chapter 18 Regulation of Gene Expression Edited by Shawn Lester PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley
Chapter 18 Regulation of Gene Expression Edited by Shawn Lester PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley
Chapter 18: Regulation of Gene Expression. Gene Regulation. Transcription Factors 3/21/2017
 Chapter 18: Regulation of Gene Expression 1. Gene Regulation in Bacteria 2. Gene Regulation in Eukaryotes 3. Gene Regulation & Cancer Gene Regulation Gene regulation refers to all aspects of controlling
Chapter 18: Regulation of Gene Expression 1. Gene Regulation in Bacteria 2. Gene Regulation in Eukaryotes 3. Gene Regulation & Cancer Gene Regulation Gene regulation refers to all aspects of controlling
Regulation of Gene Expression
 Slide 1 Chapter 18 Regulation of Gene Expression PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions
Slide 1 Chapter 18 Regulation of Gene Expression PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions
Regulation of Gene Expression
 CAMPBELL BIOLOGY IN FOCUS URRY CAIN WASSERMAN MINORSKY REECE 15 Regulation of Gene Expression Lecture Presentations by Kathleen Fitzpatrick and Nicole Tunbridge, Simon Fraser University SECOND EDITION
CAMPBELL BIOLOGY IN FOCUS URRY CAIN WASSERMAN MINORSKY REECE 15 Regulation of Gene Expression Lecture Presentations by Kathleen Fitzpatrick and Nicole Tunbridge, Simon Fraser University SECOND EDITION
Control of Eukaryotic Gene Expression (Learning Objectives)
 Control of Eukaryotic Gene Expression (Learning Objectives) 1. Compare and contrast chromatin and chromosome: composition, proteins involved and level of packing. Explain the structure and function of
Control of Eukaryotic Gene Expression (Learning Objectives) 1. Compare and contrast chromatin and chromosome: composition, proteins involved and level of packing. Explain the structure and function of
32 Gene regulation in Eukaryotes Lecture Outline 11/28/05. Gene Regulation in Prokaryotes and Eukarykotes
 3 Gene regulation in Eukaryotes Lecture Outline /8/05 Gene regulation in eukaryotes Chromatin remodeling More kinds of control elements Promoters, Enhancers, and Silencers Combinatorial control Cell-specific
3 Gene regulation in Eukaryotes Lecture Outline /8/05 Gene regulation in eukaryotes Chromatin remodeling More kinds of control elements Promoters, Enhancers, and Silencers Combinatorial control Cell-specific
Control of Gene Expression
 Control of Gene Expression 1 How Gene Regulation Works 2 Control of Gene Expression Controlling gene expression is often accomplished by controlling transcription initiation Regulatory proteins bind to
Control of Gene Expression 1 How Gene Regulation Works 2 Control of Gene Expression Controlling gene expression is often accomplished by controlling transcription initiation Regulatory proteins bind to
CHAPTER 13 LECTURE SLIDES
 CHAPTER 13 LECTURE SLIDES Prepared by Brenda Leady University of Toledo To run the animations you must be in Slideshow View. Use the buttons on the animation to play, pause, and turn audio/text on or off.
CHAPTER 13 LECTURE SLIDES Prepared by Brenda Leady University of Toledo To run the animations you must be in Slideshow View. Use the buttons on the animation to play, pause, and turn audio/text on or off.
CELL BIOLOGY - CLUTCH CH. 7 - GENE EXPRESSION.
 !! www.clutchprep.com CONCEPT: CONTROL OF GENE EXPRESSION BASICS Gene expression is the process through which cells selectively to express some genes and not others Every cell in an organism is a clone
!! www.clutchprep.com CONCEPT: CONTROL OF GENE EXPRESSION BASICS Gene expression is the process through which cells selectively to express some genes and not others Every cell in an organism is a clone
Gene Expression. Lesson 6
 Gene Expression Lesson 6 Regulation of gene expression Gene regulation turning on or off specific genes depending on the requirements of an organism Housekeeping genes are always switched on (vital life
Gene Expression Lesson 6 Regulation of gene expression Gene regulation turning on or off specific genes depending on the requirements of an organism Housekeeping genes are always switched on (vital life
Control of Eukaryotic Genes. AP Biology
 Control of Eukaryotic Genes The BIG Questions How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different, specialized functions? Evolution
Control of Eukaryotic Genes The BIG Questions How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different, specialized functions? Evolution
GENE REGULATION slide shows by Kim Foglia modified Slides with blue edges are Kim s
 GENE REGULATION slide shows by Kim Foglia modified Slides with blue edges are Kim s 2007-2008 Bacterial metabolism Bacteria need to respond quickly to changes in their environment STOP GO if they have
GENE REGULATION slide shows by Kim Foglia modified Slides with blue edges are Kim s 2007-2008 Bacterial metabolism Bacteria need to respond quickly to changes in their environment STOP GO if they have
GENE REGULATION IN PROKARYOTES
 GENE REGULATION IN PROKARYOTES Prepared by Brenda Leady, University of Toledo Copyright (c) The McGraw-Hill Companies, Inc. Permission required for reproduction or display. 1 Gene regulation refers to
GENE REGULATION IN PROKARYOTES Prepared by Brenda Leady, University of Toledo Copyright (c) The McGraw-Hill Companies, Inc. Permission required for reproduction or display. 1 Gene regulation refers to
Section C: The Control of Gene Expression
 Section C: The Control of Gene Expression 1. Each cell of a multicellular eukaryote expresses only a small fraction of its genes 2. The control of gene expression can occur at any step in the pathway from
Section C: The Control of Gene Expression 1. Each cell of a multicellular eukaryote expresses only a small fraction of its genes 2. The control of gene expression can occur at any step in the pathway from
AP Biology Day Wednesday, November 2, 2016 Friday, November 3, 2016
 AP Biology Day 30-31 Wednesday, November 2, 2016 Friday, November 3, 2016 Do-Now 1. With your neighbors, plan out your response and jot down your ideas for the 2016 short FRQ 2 Part (a) The primary transcript
AP Biology Day 30-31 Wednesday, November 2, 2016 Friday, November 3, 2016 Do-Now 1. With your neighbors, plan out your response and jot down your ideas for the 2016 short FRQ 2 Part (a) The primary transcript
Chapter 11: Regulation of Gene Expression
 Chapter Review 1. It has long been known that there is probably a genetic link for alcoholism. Researchers studying rats have begun to elucidate this link. Briefly describe the genetic mechanism found
Chapter Review 1. It has long been known that there is probably a genetic link for alcoholism. Researchers studying rats have begun to elucidate this link. Briefly describe the genetic mechanism found
EUKARYOTIC GENE CONTROL
 EUKARYOTIC GENE CONTROL THE BIG QUESTIONS How are genes turned on and off? How do cells with the same DNA/ genes differentiate to perform completely different and specialized functions? GENE EXPRESSION
EUKARYOTIC GENE CONTROL THE BIG QUESTIONS How are genes turned on and off? How do cells with the same DNA/ genes differentiate to perform completely different and specialized functions? GENE EXPRESSION
Control of Eukaryotic Genes
 Control of Eukaryotic Genes 2007-2008 The BIG Questions How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different, specialized functions?
Control of Eukaryotic Genes 2007-2008 The BIG Questions How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different, specialized functions?
Control of Eukaryotic Genes. AP Biology
 Control of Eukaryotic Genes The BIG Questions How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different, specialized functions? Evolution
Control of Eukaryotic Genes The BIG Questions How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different, specialized functions? Evolution
Control of Eukaryotic Genes. AP Biology
 Control of Eukaryotic Genes The BIG Questions How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different, specialized functions? Evolution
Control of Eukaryotic Genes The BIG Questions How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different, specialized functions? Evolution
Chapter 18. Regulation of Gene Expression
 Chapter 18 Regulation of Gene Expression 2007-2008 Control of Prokaryotic (Bacterial) Genes 2007- Bacterial metabolism Bacteria need to respond quickly to changes in their environment STOP GO if they have
Chapter 18 Regulation of Gene Expression 2007-2008 Control of Prokaryotic (Bacterial) Genes 2007- Bacterial metabolism Bacteria need to respond quickly to changes in their environment STOP GO if they have
Transcription and Post Transcript Modification
 Transcription and Post Transcript Modification You Should Be Able To 1. Describe transcription. 2. Compare and contrast eukaryotic + prokaryotic transcription. 3. Explain mrna processing in eukaryotes.
Transcription and Post Transcript Modification You Should Be Able To 1. Describe transcription. 2. Compare and contrast eukaryotic + prokaryotic transcription. 3. Explain mrna processing in eukaryotes.
Control of Eukaryotic Genes. AP Biology
 Control of Eukaryotic Genes The BIG Questions How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different, specialized functions? Evolution
Control of Eukaryotic Genes The BIG Questions How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different, specialized functions? Evolution
Unit 7. Genetic Regulation, Development, and Biotechnology. AP Biology
 Unit 7 Genetic Regulation, Development, and Biotechnology The BIG Questions How are genes turned on & off in eukaryotes and prokaryotes? How do cells with the same genes differentiate to perform completely
Unit 7 Genetic Regulation, Development, and Biotechnology The BIG Questions How are genes turned on & off in eukaryotes and prokaryotes? How do cells with the same genes differentiate to perform completely
Gene Expression and Regulation - 1
 Gene Expression and Regulation - 1 We have been discussing the molecular structure of DNA and its function in DNA replication and in transcription. Earlier we discussed how genes interact in transmission
Gene Expression and Regulation - 1 We have been discussing the molecular structure of DNA and its function in DNA replication and in transcription. Earlier we discussed how genes interact in transmission
Biotechnology Unit 3: DNA to Proteins. From DNA to RNA
 From DNA to RNA Biotechnology Unit 3: DNA to Proteins I. After the discovery of the structure of DNA, the major question remaining was how does the stored in the 4 letter code of DNA direct the and of
From DNA to RNA Biotechnology Unit 3: DNA to Proteins I. After the discovery of the structure of DNA, the major question remaining was how does the stored in the 4 letter code of DNA direct the and of
7.1 The lac Operon 7-1
 7.1 The lac Operon The lac operon was the first operon discovered It contains 3 genes coding for E. coli proteins that permit the bacteria to use the sugar lactose Galactoside permease (lacy) which transports
7.1 The lac Operon The lac operon was the first operon discovered It contains 3 genes coding for E. coli proteins that permit the bacteria to use the sugar lactose Galactoside permease (lacy) which transports
Genetics Biology 331 Exam 3B Spring 2015
 Genetics Biology 331 Exam 3B Spring 2015 MULTIPLE CHOICE. Choose the one alternative that best completes the statement or answers the question. 1) DNA methylation may be a significant mode of genetic regulation
Genetics Biology 331 Exam 3B Spring 2015 MULTIPLE CHOICE. Choose the one alternative that best completes the statement or answers the question. 1) DNA methylation may be a significant mode of genetic regulation
Chapter 18: Regulation of Gene Expression
 Chapter 18: Regulation of Gene Expression Regulation of Metabolism Shuts off transcription Types of Feedback Negative feedback = body s response is to reduce the stimulus Ex: regulation of body temp, blood
Chapter 18: Regulation of Gene Expression Regulation of Metabolism Shuts off transcription Types of Feedback Negative feedback = body s response is to reduce the stimulus Ex: regulation of body temp, blood
Chapter 2. An Introduction to Genes and Genomes
 PowerPoint Lectures for Introduction to Biotechnology, Second Edition William J.Thieman and Michael A.Palladino Chapter 2 An Introduction to Genes and Genomes Lectures by Lara Dowland Chapter Contents
PowerPoint Lectures for Introduction to Biotechnology, Second Edition William J.Thieman and Michael A.Palladino Chapter 2 An Introduction to Genes and Genomes Lectures by Lara Dowland Chapter Contents
AP Biology. The BIG Questions. Chapter 19. Prokaryote vs. eukaryote genome. Prokaryote vs. eukaryote genome. Why turn genes on & off?
 The BIG Questions Chapter 19. Control of Eukaryotic Genome How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different, specialized functions?
The BIG Questions Chapter 19. Control of Eukaryotic Genome How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different, specialized functions?
Control of Metabolic Processes
 Control of Metabolic Processes Harriet Wilson, Lecture Notes Bio. Sci. 4 - Microbiology Sierra College As described earlier, the metabolic processes occurring within living organisms (glycolysis, respiration,
Control of Metabolic Processes Harriet Wilson, Lecture Notes Bio. Sci. 4 - Microbiology Sierra College As described earlier, the metabolic processes occurring within living organisms (glycolysis, respiration,
Matakuliah Genetika (BIO612206) Jurusan Biologi FMIPA Universitas Lampung. Priyambodo, M.Sc. staff.unila.ac.id/priyambodo
 Matakuliah Genetika (BIO612206) Jurusan Biologi FMIPA Universitas Lampung Priyambodo, M.Sc. staff.unila.ac.id/priyambodo Prokariotik Eukariotik staff.unila.ac.id/priyambodo Regulasi ekspresi gen pada
Matakuliah Genetika (BIO612206) Jurusan Biologi FMIPA Universitas Lampung Priyambodo, M.Sc. staff.unila.ac.id/priyambodo Prokariotik Eukariotik staff.unila.ac.id/priyambodo Regulasi ekspresi gen pada
Chapter 13 - Regulation of Gene Expression
 Chapter 13 - Regulation of Gene Expression 1. Describe the typical components of an operon in an E. coli (prokaryotic) cell. (p. 238-239) a. regulator gene - b. promoter - c. operator - d. structural gene
Chapter 13 - Regulation of Gene Expression 1. Describe the typical components of an operon in an E. coli (prokaryotic) cell. (p. 238-239) a. regulator gene - b. promoter - c. operator - d. structural gene
GENE EXPRESSSION. Promoter sequence where RNA polymerase binds. Operator sequence that acts as a switch (yellow) OPERON
 GENE EXPRESSSION 1 GENE REGULATION IN PROKARYOTES Bacteria can turn genes on or off depending on their environment Prokaryotes have operons clusters of related genes and regulatory sequences Promoter sequence
GENE EXPRESSSION 1 GENE REGULATION IN PROKARYOTES Bacteria can turn genes on or off depending on their environment Prokaryotes have operons clusters of related genes and regulatory sequences Promoter sequence
Division Ave. High School AP Biology
 Control of Eukaryotic Genes 2007-2008 The BIG Questions n How are genes turned on & off in eukaryotes? n How do cells with the same genes differentiate to perform completely different, specialized functions?
Control of Eukaryotic Genes 2007-2008 The BIG Questions n How are genes turned on & off in eukaryotes? n How do cells with the same genes differentiate to perform completely different, specialized functions?
Unit IX Problem 3 Genetics: Basic Concepts in Molecular Biology
 Unit IX Problem 3 Genetics: Basic Concepts in Molecular Biology - The central dogma (principle) of molecular biology: Information from DNA are transcribed to mrna which will be further translated to synthesize
Unit IX Problem 3 Genetics: Basic Concepts in Molecular Biology - The central dogma (principle) of molecular biology: Information from DNA are transcribed to mrna which will be further translated to synthesize
Synthetic cells: do bacteria need all its genes? No.
 NO NEED TO REFER TO THE SLIDES. بسم هللا الرحمن الرحيم Do we need all the non coding regions of the DNA? Two weeks ago, they discovered that the genome of a plant is very small (recall that plant genome
NO NEED TO REFER TO THE SLIDES. بسم هللا الرحمن الرحيم Do we need all the non coding regions of the DNA? Two weeks ago, they discovered that the genome of a plant is very small (recall that plant genome
Prokaryotic Transcription
 Prokaryotic Transcription Transcription Basics DNA is the genetic material Nucleic acid Capable of self-replication and synthesis of RNA RNA is the middle man Nucleic acid Structure and base sequence are
Prokaryotic Transcription Transcription Basics DNA is the genetic material Nucleic acid Capable of self-replication and synthesis of RNA RNA is the middle man Nucleic acid Structure and base sequence are
Make the protein through the genetic dogma process.
 Make the protein through the genetic dogma process. Coding Strand 5 AGCAATCATGGATTGGGTACATTTGTAACTGT 3 Template Strand mrna Protein Complete the table. DNA strand DNA s strand G mrna A C U G T A T Amino
Make the protein through the genetic dogma process. Coding Strand 5 AGCAATCATGGATTGGGTACATTTGTAACTGT 3 Template Strand mrna Protein Complete the table. DNA strand DNA s strand G mrna A C U G T A T Amino
Gene Regulation in Eukaryotes
 Gene Regulation in Eukaryotes The latest estimates are that a human cell, a eukaryotic cell, contains 20,000 25,000 genes. Some of these are expressed in all cells all the time. These so-called housekeeping
Gene Regulation in Eukaryotes The latest estimates are that a human cell, a eukaryotic cell, contains 20,000 25,000 genes. Some of these are expressed in all cells all the time. These so-called housekeeping
Chapter 16: Gene Expression from Biology by OpenStax College is licensed under a Creative Commons Attribution 3.0 Unported license.
 Chapter 16: Gene Expression from Biology by OpenStax College is licensed under a Creative Commons Attribution 3.0 Unported license. 2013, Rice University. CHAPTER 16 GENE EXPRESSION 429 16 GENE EXPRESSION
Chapter 16: Gene Expression from Biology by OpenStax College is licensed under a Creative Commons Attribution 3.0 Unported license. 2013, Rice University. CHAPTER 16 GENE EXPRESSION 429 16 GENE EXPRESSION
Chapter 19. Control of Eukaryotic Genome. AP Biology
 Chapter 19. Control of Eukaryotic Genome The BIG Questions How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different, specialized functions?
Chapter 19. Control of Eukaryotic Genome The BIG Questions How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different, specialized functions?
Gene Circuits -2. Shu-Ping Lin, Ph.D. Institute of Biomedical Engineering
 Gene Circuits -2 Shu-Ping Lin, Ph.D. Institute of Biomedical Engineering E-mail: splin@dragon.nchu.edu.tw Website: http://web.nchu.edu.tw/pweb/users/splin/ Genes Flow of information in gene expression
Gene Circuits -2 Shu-Ping Lin, Ph.D. Institute of Biomedical Engineering E-mail: splin@dragon.nchu.edu.tw Website: http://web.nchu.edu.tw/pweb/users/splin/ Genes Flow of information in gene expression
3.5b. Regulation of Gene Expression CAMPBELL BIOLOGY IN FOCUS. Urry Cain Wasserman Minorsky Jackson Reece
 CAMPBELL BIOLOGY IN FOCUS Urry Cain Wasserman Minorsky Jackson Reece 3.5b Regulation of Gene Expression Lecture Presentations by Kathleen Fitzpatrick and Nicole Tunbridge Overview: Differential Expression
CAMPBELL BIOLOGY IN FOCUS Urry Cain Wasserman Minorsky Jackson Reece 3.5b Regulation of Gene Expression Lecture Presentations by Kathleen Fitzpatrick and Nicole Tunbridge Overview: Differential Expression
Themes: RNA and RNA Processing. Messenger RNA (mrna) What is a gene? RNA is very versatile! RNA-RNA interactions are very important!
 Themes: RNA is very versatile! RNA and RNA Processing Chapter 14 RNA-RNA interactions are very important! Prokaryotes and Eukaryotes have many important differences. Messenger RNA (mrna) Carries genetic
Themes: RNA is very versatile! RNA and RNA Processing Chapter 14 RNA-RNA interactions are very important! Prokaryotes and Eukaryotes have many important differences. Messenger RNA (mrna) Carries genetic
Gene Expression. Chapters 11 & 12: Gene Conrtrol and DNA Technology. Cloning. Honors Biology Fig
 Chapters & : Conrtrol and Technology Honors Biology 0 Cloning Produced by asexual reproduction and so it is genetically identical to the parent st large cloned mammal: Dolly the sheep Animals that are
Chapters & : Conrtrol and Technology Honors Biology 0 Cloning Produced by asexual reproduction and so it is genetically identical to the parent st large cloned mammal: Dolly the sheep Animals that are
Chapter 11. How Genes Are Controlled. Lectures by Edward J. Zalisko
 Chapter 11 How Genes Are Controlled PowerPoint Lectures for Campbell Essential Biology, Fifth Edition, and Campbell Essential Biology with Physiology, Fourth Edition Eric J. Simon, Jean L. Dickey, and
Chapter 11 How Genes Are Controlled PowerPoint Lectures for Campbell Essential Biology, Fifth Edition, and Campbell Essential Biology with Physiology, Fourth Edition Eric J. Simon, Jean L. Dickey, and
Developmental Biology BY1101 P. Murphy
 Developmental Biology BY1101 P. Murphy Lecture 7 Cellular differentiation and the regulation of gene expression. In this lecture we looked at two main questions: How is gene expression regulated? (revision
Developmental Biology BY1101 P. Murphy Lecture 7 Cellular differentiation and the regulation of gene expression. In this lecture we looked at two main questions: How is gene expression regulated? (revision
Gene Expression: Transcription, Translation, RNAs and the Genetic Code
 Lecture 28-29 Gene Expression: Transcription, Translation, RNAs and the Genetic Code Central dogma of molecular biology During transcription, the information in a DNA sequence (a gene) is copied into a
Lecture 28-29 Gene Expression: Transcription, Translation, RNAs and the Genetic Code Central dogma of molecular biology During transcription, the information in a DNA sequence (a gene) is copied into a
Fig Ch 17: From Gene to Protein
 Fig. 17-1 Ch 17: From Gene to Protein Basic Principles of Transcription and Translation RNA is the intermediate between genes and the proteins for which they code Transcription is the synthesis of RNA
Fig. 17-1 Ch 17: From Gene to Protein Basic Principles of Transcription and Translation RNA is the intermediate between genes and the proteins for which they code Transcription is the synthesis of RNA
Differences between prokaryotes & eukaryotes. Gene function
 GENE REGULATION Differences between prokaryotes & eukaryotes Gene function Description of Prokaryotic Chromosome and E.coli Review Differences between Prokaryotic & Eukaryotic Chromosomes Four differences
GENE REGULATION Differences between prokaryotes & eukaryotes Gene function Description of Prokaryotic Chromosome and E.coli Review Differences between Prokaryotic & Eukaryotic Chromosomes Four differences
Biol 3301 Genetics Exam #2A October 26, 2004
 Biol 3301 Genetics Exam #2A October 26, 2004 This exam consists of 40 multiple choice questions worth 2.5 points each, for a total of 100 points. Good luck. Name SS# 1. Which of the following statements
Biol 3301 Genetics Exam #2A October 26, 2004 This exam consists of 40 multiple choice questions worth 2.5 points each, for a total of 100 points. Good luck. Name SS# 1. Which of the following statements
Molecular Cell Biology - Problem Drill 09: Gene Expression in Prokaryotes
 Molecular Cell Biology - Problem Drill 09: Gene Expression in Prokaryotes Question No. 1 of 10 1. Which of the following statements about gene expression in prokaryotes is correct? Question #1 (A) In prokaryotes,
Molecular Cell Biology - Problem Drill 09: Gene Expression in Prokaryotes Question No. 1 of 10 1. Which of the following statements about gene expression in prokaryotes is correct? Question #1 (A) In prokaryotes,
I. Prokaryotic Gene Regulation. Figure 1: Operon. Operon:
 I. Prokaryotic Gene Regulation Figure 1: Operon Operon: a) Regulatory Elements consist of an Operator that serves as the on-off switch for the genes of the operon. Also contains a promoter for the Structural
I. Prokaryotic Gene Regulation Figure 1: Operon Operon: a) Regulatory Elements consist of an Operator that serves as the on-off switch for the genes of the operon. Also contains a promoter for the Structural
30 Gene expression: Transcription
 30 Gene expression: Transcription Gene structure. o Exons coding region of DNA. o Introns non-coding region of DNA. o Introns are interspersed between exons of a single gene. o Promoter region helps enzymes
30 Gene expression: Transcription Gene structure. o Exons coding region of DNA. o Introns non-coding region of DNA. o Introns are interspersed between exons of a single gene. o Promoter region helps enzymes
Molecular Genetics Quiz #1 SBI4U K T/I A C TOTAL
 Name: Molecular Genetics Quiz #1 SBI4U K T/I A C TOTAL Part A: Multiple Choice (15 marks) Circle the letter of choice that best completes the statement or answers the question. One mark for each correct
Name: Molecular Genetics Quiz #1 SBI4U K T/I A C TOTAL Part A: Multiple Choice (15 marks) Circle the letter of choice that best completes the statement or answers the question. One mark for each correct
B. Incorrect! Centromeric DNA is largely heterochromatin, which is inactive DNA.
 MCAT Biology - Problem Drill 06: Molecular Biology of Eukaryotes Question No. 1 of 10 1. Which type of DNA would have the highest level of expression? Question #01 (A) Heterochromatin. (B) Centromeric
MCAT Biology - Problem Drill 06: Molecular Biology of Eukaryotes Question No. 1 of 10 1. Which type of DNA would have the highest level of expression? Question #01 (A) Heterochromatin. (B) Centromeric
Quick Review of Protein Synthesis
 Collin College BIOL. 2401 Quick Review of Protein Synthesis. Proteins and Protein Synthesis Proteins are the molecular units that do most of the work in a cell. They function as molecular catalysts, help
Collin College BIOL. 2401 Quick Review of Protein Synthesis. Proteins and Protein Synthesis Proteins are the molecular units that do most of the work in a cell. They function as molecular catalysts, help
From DNA to Protein: Genotype to Phenotype
 12 From DNA to Protein: Genotype to Phenotype 12.1 What Is the Evidence that Genes Code for Proteins? The gene-enzyme relationship is one-gene, one-polypeptide relationship. Example: In hemoglobin, each
12 From DNA to Protein: Genotype to Phenotype 12.1 What Is the Evidence that Genes Code for Proteins? The gene-enzyme relationship is one-gene, one-polypeptide relationship. Example: In hemoglobin, each
Lecture 9 Controlling gene expression
 Lecture 9 Controlling gene expression BIOLOGY Campbell, Reece and Mitchell Chapter 18 334- (352-356) Every cell in your body contains the same number of genes approximately 35, 000 DNA is wound around
Lecture 9 Controlling gene expression BIOLOGY Campbell, Reece and Mitchell Chapter 18 334- (352-356) Every cell in your body contains the same number of genes approximately 35, 000 DNA is wound around
Transcription in Eukaryotes
 Transcription in Eukaryotes Biology I Hayder A Giha Transcription Transcription is a DNA-directed synthesis of RNA, which is the first step in gene expression. Gene expression, is transformation of the
Transcription in Eukaryotes Biology I Hayder A Giha Transcription Transcription is a DNA-directed synthesis of RNA, which is the first step in gene expression. Gene expression, is transformation of the
Chapter 19 Genetic Regulation of the Eukaryotic Genome. A. Bergeron AP Biology PCHS
 Chapter 19 Genetic Regulation of the Eukaryotic Genome A. Bergeron AP Biology PCHS 2 Do Now - Eukaryotic Transcription Regulation The diagram below shows five genes (with their enhancers) from the genome
Chapter 19 Genetic Regulation of the Eukaryotic Genome A. Bergeron AP Biology PCHS 2 Do Now - Eukaryotic Transcription Regulation The diagram below shows five genes (with their enhancers) from the genome
17.5 Eukaryotic Transcription Initiation Is Regulated by Transcription Factors That Bind to Cis-Acting Sites
 17.5 Eukaryotic Transcription Initiation Is Regulated by Transcription Factors That Bind to Cis-Acting Sites 1 Section 17.5 Transcription regulatory proteins, transcription factors, target cis-acting sites
17.5 Eukaryotic Transcription Initiation Is Regulated by Transcription Factors That Bind to Cis-Acting Sites 1 Section 17.5 Transcription regulatory proteins, transcription factors, target cis-acting sites
REGULATION OF GENE EXPRESSION
 REGULATION OF GENE EXPRESSION Each cell of a living organism contains thousands of genes. But all genes do not function at a time. Genes function according to requirements of the cell. Genes control the
REGULATION OF GENE EXPRESSION Each cell of a living organism contains thousands of genes. But all genes do not function at a time. Genes function according to requirements of the cell. Genes control the
AP Biology Gene Expression/Biotechnology REVIEW
 AP Biology Gene Expression/Biotechnology REVIEW Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Gene expression can be a. regulated before transcription.
AP Biology Gene Expression/Biotechnology REVIEW Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Gene expression can be a. regulated before transcription.
Transcription is the first stage of gene expression
 Transcription is the first stage of gene expression RNA synthesis is catalyzed by RNA polymerase, which pries the DNA strands apart and hooks together the RNA nucleotides The RNA is complementary to the
Transcription is the first stage of gene expression RNA synthesis is catalyzed by RNA polymerase, which pries the DNA strands apart and hooks together the RNA nucleotides The RNA is complementary to the
Big Idea 3C Basic Review
 Big Idea 3C Basic Review 1. A gene is a. A sequence of DNA that codes for a protein. b. A sequence of amino acids that codes for a protein. c. A sequence of codons that code for nucleic acids. d. The end
Big Idea 3C Basic Review 1. A gene is a. A sequence of DNA that codes for a protein. b. A sequence of amino acids that codes for a protein. c. A sequence of codons that code for nucleic acids. d. The end
Lecture for Wednesday. Dr. Prince BIOL 1408
 Lecture for Wednesday Dr. Prince BIOL 1408 THE FLOW OF GENETIC INFORMATION FROM DNA TO RNA TO PROTEIN Copyright 2009 Pearson Education, Inc. Genes are expressed as proteins A gene is a segment of DNA that
Lecture for Wednesday Dr. Prince BIOL 1408 THE FLOW OF GENETIC INFORMATION FROM DNA TO RNA TO PROTEIN Copyright 2009 Pearson Education, Inc. Genes are expressed as proteins A gene is a segment of DNA that
Chapter 17. From Gene to Protein. Slide 1. Slide 2. Slide 3. Gene Expression. Which of the following is the best example of gene expression? Why?
 Slide 1 Chapter 17 From Gene to Protein PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from
Slide 1 Chapter 17 From Gene to Protein PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from
GENETICS - CLUTCH CH.10 TRANSCRIPTION.
 !! www.clutchprep.com CONCEPT: OVERVIEW OF TRANSCRIPTION Transcription is the process of using DNA as a template to RNA RNA polymerase is the enzyme that transcribes DNA - There are many different types
!! www.clutchprep.com CONCEPT: OVERVIEW OF TRANSCRIPTION Transcription is the process of using DNA as a template to RNA RNA polymerase is the enzyme that transcribes DNA - There are many different types
Gene Regulation Biology
 Gene Regulation Biology Potential and Limitations of Cell Re-programming in Cancer Research Eric Blanc KCL April 13, 2010 Eric Blanc (KCL) Gene Regulation Biology April 13, 2010 1 / 21 Outline 1 The Central
Gene Regulation Biology Potential and Limitations of Cell Re-programming in Cancer Research Eric Blanc KCL April 13, 2010 Eric Blanc (KCL) Gene Regulation Biology April 13, 2010 1 / 21 Outline 1 The Central
Chapter 17 Lecture. Concepts of Genetics. Tenth Edition. Regulation of Gene Expression in Eukaryotes
 Chapter 17 Lecture Concepts of Genetics Tenth Edition Regulation of Gene Expression in Eukaryotes Chapter Contents 17.1 Eukaryotic Gene Regulation Can Occur at Any of the Steps Leading from DNA to Protein
Chapter 17 Lecture Concepts of Genetics Tenth Edition Regulation of Gene Expression in Eukaryotes Chapter Contents 17.1 Eukaryotic Gene Regulation Can Occur at Any of the Steps Leading from DNA to Protein
Review of Protein (one or more polypeptide) A polypeptide is a long chain of..
 Gene expression Review of Protein (one or more polypeptide) A polypeptide is a long chain of.. In a protein, the sequence of amino acid determines its which determines the protein s A protein with an enzymatic
Gene expression Review of Protein (one or more polypeptide) A polypeptide is a long chain of.. In a protein, the sequence of amino acid determines its which determines the protein s A protein with an enzymatic
DNA makes RNA makes Proteins. The Central Dogma
 DNA makes RNA makes Proteins The Central Dogma TRANSCRIPTION DNA RNA transcript RNA polymerase RNA PROCESSING Exon RNA transcript (pre-mrna) Intron Aminoacyl-tRNA synthetase NUCLEUS CYTOPLASM FORMATION
DNA makes RNA makes Proteins The Central Dogma TRANSCRIPTION DNA RNA transcript RNA polymerase RNA PROCESSING Exon RNA transcript (pre-mrna) Intron Aminoacyl-tRNA synthetase NUCLEUS CYTOPLASM FORMATION
CH 17 :From Gene to Protein
 CH 17 :From Gene to Protein Defining a gene gene gene Defining a gene is problematic because one gene can code for several protein products, some genes code only for RNA, two genes can overlap, and there
CH 17 :From Gene to Protein Defining a gene gene gene Defining a gene is problematic because one gene can code for several protein products, some genes code only for RNA, two genes can overlap, and there
I. Gene Expression Figure 1: Central Dogma of Molecular Biology
 I. Gene Expression Figure 1: Central Dogma of Molecular Biology Central Dogma: Gene Expression: RNA Structure RNA nucleotides contain the pentose sugar Ribose instead of deoxyribose. Contain the bases
I. Gene Expression Figure 1: Central Dogma of Molecular Biology Central Dogma: Gene Expression: RNA Structure RNA nucleotides contain the pentose sugar Ribose instead of deoxyribose. Contain the bases
TRANSCRIPTION AND PROCESSING OF RNA
 TRANSCRIPTION AND PROCESSING OF RNA 1. The steps of gene expression. 2. General characterization of transcription: steps, components of transcription apparatus. 3. Transcription of eukaryotic structural
TRANSCRIPTION AND PROCESSING OF RNA 1. The steps of gene expression. 2. General characterization of transcription: steps, components of transcription apparatus. 3. Transcription of eukaryotic structural
Resources. This lecture Campbell and Farrell's Biochemistry, Chapter 11
 Transcription Resources This lecture Campbell and Farrell's Biochemistry, Chapter 11 2 Definition of a gene The entire nucleic acid sequence that is necessary for the synthesis of a functional polypeptide
Transcription Resources This lecture Campbell and Farrell's Biochemistry, Chapter 11 2 Definition of a gene The entire nucleic acid sequence that is necessary for the synthesis of a functional polypeptide
GENE REGULATION. Gene regulation occurs at the level of transcription or production of mrna
 GENE REGULATION Virtually every cell in your body contains a complete set of genes But they are not all turned on in every tissue Each cell in your body expresses only a small subset of genes at any time
GENE REGULATION Virtually every cell in your body contains a complete set of genes But they are not all turned on in every tissue Each cell in your body expresses only a small subset of genes at any time
REGULATION OF PROTEIN SYNTHESIS. II. Eukaryotes
 REGULATION OF PROTEIN SYNTHESIS II. Eukaryotes Complexities of eukaryotic gene expression! Several steps needed for synthesis of mrna! Separation in space of transcription and translation! Compartmentation
REGULATION OF PROTEIN SYNTHESIS II. Eukaryotes Complexities of eukaryotic gene expression! Several steps needed for synthesis of mrna! Separation in space of transcription and translation! Compartmentation
REGULATION OF GENE EXPRESSION
 REGULATION OF GENE EXPRESSION Each cell of a living organism contains thousands of genes. But all genes do not function at a time. Genes function according to requirements of the cell. Genes control the
REGULATION OF GENE EXPRESSION Each cell of a living organism contains thousands of genes. But all genes do not function at a time. Genes function according to requirements of the cell. Genes control the
Unit II Problem 3 Genetics: Summary of Basic Concepts in Molecular Biology
 Unit II Problem 3 Genetics: Summary of Basic Concepts in Molecular Biology - The central dogma (principle) of molecular biology: Information from DNA are transcribed to mrna which will be further translated
Unit II Problem 3 Genetics: Summary of Basic Concepts in Molecular Biology - The central dogma (principle) of molecular biology: Information from DNA are transcribed to mrna which will be further translated
Chapter 17. From Gene to Protein
 Chapter 17 From Gene to Protein Overview: The Flow of Genetic Information The information content of DNA is in the form of specific sequences of nucleotides The DNA inherited by an organism leads to specific
Chapter 17 From Gene to Protein Overview: The Flow of Genetic Information The information content of DNA is in the form of specific sequences of nucleotides The DNA inherited by an organism leads to specific
Chapter 12 Packet DNA 1. What did Griffith conclude from his experiment? 2. Describe the process of transformation.
 Chapter 12 Packet DNA and RNA Name Period California State Standards covered by this chapter: Cell Biology 1. The fundamental life processes of plants and animals depend on a variety of chemical reactions
Chapter 12 Packet DNA and RNA Name Period California State Standards covered by this chapter: Cell Biology 1. The fundamental life processes of plants and animals depend on a variety of chemical reactions
Transcription. The sugar molecule found in RNA is ribose, rather than the deoxyribose found in DNA.
 Transcription RNA (ribonucleic acid) is a key intermediary between a DNA sequence and a polypeptide. RNA is an informational polynucleotide similar to DNA, but it differs from DNA in three ways: RNA generally
Transcription RNA (ribonucleic acid) is a key intermediary between a DNA sequence and a polypeptide. RNA is an informational polynucleotide similar to DNA, but it differs from DNA in three ways: RNA generally
(c) 2014 Dr. Alice Heicklen & Dr. Deborah Mowshowitz, Columbia University, New York, NY. Last update 02/26/ :57 PM
 C2006/F2402 '14 OUTLINE OF LECTURE #11 (c) 2014 Dr. Alice Heicklen & Dr. Deborah Mowshowitz, Columbia University, New York, NY. Last update 02/26/2014 12:57 PM Handouts: 10C -- Typical Eukaryotic Gene,
C2006/F2402 '14 OUTLINE OF LECTURE #11 (c) 2014 Dr. Alice Heicklen & Dr. Deborah Mowshowitz, Columbia University, New York, NY. Last update 02/26/2014 12:57 PM Handouts: 10C -- Typical Eukaryotic Gene,
There are four major types of introns. Group I introns, found in some rrna genes, are self-splicing: they can catalyze their own removal.
 1 2 Continuous genes - Intron: Many eukaryotic genes contain coding regions called exons and noncoding regions called intervening sequences or introns. The average human gene contains from eight to nine
1 2 Continuous genes - Intron: Many eukaryotic genes contain coding regions called exons and noncoding regions called intervening sequences or introns. The average human gene contains from eight to nine
M I C R O B I O L O G Y WITH DISEASES BY TAXONOMY, THIRD EDITION
 M I C R O B I O L O G Y WITH DISEASES BY TAXONOMY, THIRD EDITION Chapter 7 Microbial Genetics Lecture prepared by Mindy Miller-Kittrell, University of Tennessee, Knoxville The Structure and Replication
M I C R O B I O L O G Y WITH DISEASES BY TAXONOMY, THIRD EDITION Chapter 7 Microbial Genetics Lecture prepared by Mindy Miller-Kittrell, University of Tennessee, Knoxville The Structure and Replication
DNA Function: Information Transmission
 DNA Function: Information Transmission DNA is called the code of life. What does it code for? *the information ( code ) to make proteins! Why are proteins so important? Nearly every function of a living
DNA Function: Information Transmission DNA is called the code of life. What does it code for? *the information ( code ) to make proteins! Why are proteins so important? Nearly every function of a living
How to Use This Presentation
 How to Use This Presentation To View the presentation as a slideshow with effects select View on the menu bar and click on Slide Show. To advance through the presentation, click the right-arrow key or
How to Use This Presentation To View the presentation as a slideshow with effects select View on the menu bar and click on Slide Show. To advance through the presentation, click the right-arrow key or
BCH 4054 Fall 2000 Chapter 31 Lecture Notes
 BCH 4054 Fall 2000 Chapter 31 Lecture Notes 1 Chapter 31 Transcription and Regulation of Gene Expression 2 Messenger RNA Central Dogma (Francis Crick, 1958) DNA RNA Protein (Fig 31.1) Jacob-Monod Hypothesis:
BCH 4054 Fall 2000 Chapter 31 Lecture Notes 1 Chapter 31 Transcription and Regulation of Gene Expression 2 Messenger RNA Central Dogma (Francis Crick, 1958) DNA RNA Protein (Fig 31.1) Jacob-Monod Hypothesis:
Protein Synthesis Notes
 Protein Synthesis Notes Protein Synthesis: Overview Transcription: synthesis of mrna under the direction of DNA. Translation: actual synthesis of a polypeptide under the direction of mrna. Transcription
Protein Synthesis Notes Protein Synthesis: Overview Transcription: synthesis of mrna under the direction of DNA. Translation: actual synthesis of a polypeptide under the direction of mrna. Transcription
Chapter 15 Gene Regulation in Prokaryotes
 Chapter 15 Gene Regulation in Prokaryotes 17-1 Sections to study 15.1 The elements of prokaryotic gene expression 15.2 Regulation of transcription initiation via DNA-binding proteins 15.3 RNA-mediated
Chapter 15 Gene Regulation in Prokaryotes 17-1 Sections to study 15.1 The elements of prokaryotic gene expression 15.2 Regulation of transcription initiation via DNA-binding proteins 15.3 RNA-mediated
Wednesday, November 22, 17. Exons and Introns
 Exons and Introns Introns and Exons Exons: coded regions of DNA that get transcribed and translated into proteins make up 5% of the genome Introns and Exons Introns: non-coded regions of DNA Must be removed
Exons and Introns Introns and Exons Exons: coded regions of DNA that get transcribed and translated into proteins make up 5% of the genome Introns and Exons Introns: non-coded regions of DNA Must be removed
Enzyme that uses RNA as a template to synthesize a complementary DNA
 Biology 105: Introduction to Genetics PRACTICE FINAL EXAM 2006 Part I: Definitions Homology: Comparison of two or more protein or DNA sequence to ascertain similarities in sequences. If two genes have
Biology 105: Introduction to Genetics PRACTICE FINAL EXAM 2006 Part I: Definitions Homology: Comparison of two or more protein or DNA sequence to ascertain similarities in sequences. If two genes have
Gene expression DNA RNA. Protein DNA. Replication. Initiation Elongation Processing Export. DNA RNA Protein. Transcription. Degradation.
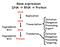 Gene expression DNA RNA Protein DNA DNA Degradation RNA Degradation Protein Replication Transcription Translation Initiation Elongation Processing Export Initiation Elongation Processing Targeting Chapter
Gene expression DNA RNA Protein DNA DNA Degradation RNA Degradation Protein Replication Transcription Translation Initiation Elongation Processing Export Initiation Elongation Processing Targeting Chapter
