INTRODUCCIÓ A LES TECNOLOGIES DE 'NEXT GENERATION SEQUENCING'
|
|
|
- Aubrey Mosley
- 6 years ago
- Views:
Transcription
1 INTRODUCCIÓ A LES TECNOLOGIES DE 'NEXT GENERATION SEQUENCING' Bioinformàtica per a la Recerca Biomèdica Ricardo Gonzalo Sanz ricardo.gonzalo@vhir.org 14/12/2016
2 1. Introduction to NGS 2. First Generation Sequencing 3. Second Generation Sequencing 4. Third Generation Sequencing 5. Sequencing generation face to face 6. Applications of NGS techniques 7. A (very) brief introduction to DoE
3 1. Introduction to NGS 2. First Generation Sequencing 3. Second Generation Sequencing 4. Third Generation Sequencing 5. Sequencing generation face to face 6. Applications of NGS techniques 7. A (very) brief introduction to DoE
4 1. Introduction to NGS Diagnosis/Prognosis What happens? Why happens?
5 1. Introduction to NGS Diagnosis/Prognosis Treatment What happens? Why happens? Prevention
6 1. Introduction to NGS Diagnosis Signs Symptoms Medical examination Medical tests Many medicines are not as effective as expected in all patients Some patients may suffer from serious adverse reactions
7 1. Introduction to NGS Personalized medicine era -The right therapeutic strategy for the right person at the right time -Predisposition to disease -Early and targeted prevention Biomarker identification: Diagnostic Susceptibility/risk (prevention) Prognostic (indolent vs. aggressive) Predictive (response)
8 1. Introduction to NGS Genomics is a branch of genetics that enables the study of genomes of whole organisms. Gregor Mendel It differs from classical genetics in that it considers the hereditary material of an organism in global rather than one gene or one gene product at a time.
9 1. Introduction to NGS Deoxyribose nucleic acid (DNA)
10 1. Introduction to NGS Genome sequencing Genome sequencing is figuring out the order of DNA nucleotides, or bases, in a genome the order of As, Cs, Gs, and Ts that make up an organism's DNA.
11 1. Introduction to NGS Why is genome sequencing so important? Sequencing the genome is an important step towards understanding it. The genome sequence will help scientists find genes much more easily and quickly. Study the entire genome sequence will help them understand how the genome as a whole works how genes work together to direct the growth, development and maintenance of an entire organism. Genes account for less than 25 percent of the DNA in the genome, and so knowing the entire genome sequence will help scientists study the parts of the genome outside the genes.
12 1. Introduction to NGS Human Genome Project The Human Genome Project (HGP) was the international, collaborative research program whose goal was the complete mapping and understanding of all the genes of human beings. Begin in 1990 First draft in February 2001 full sequence April 2003 It catalyzes the developing and improving of sequencing techniques
13 1. Introduction to NGS
14 1. Introduction to NGS
15 1. Introduction to NGS Consortia-based projects
16 1. Introduction to NGS 3.234,83 Mb (haploid) $ 2,7 billion 1st generation 2nd generation 3rd generation
17 1. Introduction to NGS Cost of sequencing Size of the genome Whole genome/exome/ de novo Data quality Moore s Law: the number of transistors in an integrated circuit doubles approximately every two years (1965).
18 1. Introduction to NGS Cost of sequencing Cost of human genome sequence: HGP: 0.5-1x10 9 $ 2006: 14x10 6 $ 2016: $
19 1. Introduction to NGS 2. First Generation Sequencing 3. Second Generation Sequencing 4. Third Generation Sequencing 5. Sequencing generation face to face 6. Applications of NGS techniques 7. A (very) brief introduction to DoE
20 2. First Generation Sequencing Sanger sequencing It is a method of DNA sequencing based on the selective incorporation of chainterminating dideoxynucleotides by DNA polymerase during in vitro DNA replication. Developed by Frederick Sanger and colleagues in 1977, it was the most widely used sequencing method for approximately 25 years. Frederick Sanger received two Nobel prizes (in the same category), for his work on protein sequencing and DNA sequencing
21 2. First Generation Sequencing Sanger sequencing. How it works? A DNAp enzyme is used to replicate a ssdna. In the mix reaction, there exist normal and modified nucleotides. Random incorporation of modified nucleotides stops the synthesis reaction. Each generated fragment will have a different length that could be distinguish by gel electrophoresis. <1kb read
22 1. Introduction to NGS 2. First Generation Sequencing 3. Second Generation Sequencing 4. Third Generation Sequencing 5. Sequencing generation face to face 6. Applications of NGS techniques 7. A (very) brief introduction to DoE
23 3. Second Generation Sequencing Why second generation sequencing? One of the main problems of Sanger sequencing was dealing with larger sequence output using of gels or polymers as separation media limited number of samples which could be handled in parallel difficulties with complete automation of the sample preparation These limitations triggered the efforts to develop new techniques
24 3. Second Generation Sequencing Main characteristics of NGS: high speed and throughput shorter reads more accuracy much higher degree of sequence coverage easier assemblies of the data generated Huge data storage demands
25 3.Second Generation Sequencing High througput 2ond generation instruments Benchtop ROCHE x GS FLX+ 454 x GS Junior 454 Illumina NextSeq500 HiSeq HiSeq X-Ten MiniSeq Life Technologies x SOLID5500xl Ion S5 and S5 XL IonPGM
26 3. Second Generation Sequencing Basic NGS workflow. 1. Library Preparation It is prepared by random fragmentation of DNA or cdna sample, followed by 5 and 3 adapter ligation. Adapter-ligated fragments are then PCR amplified and gel purified 2. Clonal amplification Each DNA fragment is amplified millions of times. 3. Sequencing The nucleotides incorporated are read by the detector
27 3. Second Generation Sequencing Library preparation: i. Fragmenting/selecting the target sequences: i. Physical (acoustic shearing, sonications. I.e. Covaris) ii. Enzymatic (i.e Nextera) iii. Chemical (less frequent) ii. Converting target to dsdna (if necessary) iii. End-repair and adaptors ligation (barcode for multplexing) iv. Quantification and QC of the final library product
28 3. Second Generation Sequencing Library preparation: Useful for multiplexing samples Depending on the final application and method used for sequencing, different kits are available
29 3. Second Generation Sequencing Clonal amplification: For empcr based systems (Ion/PGM, 454) High-speed shaker -1 starting effective fragment per microreactor - ~10 6 microreactors per ml - All processed in parallel (Clonal amplification)
30 3. Second Generation Sequencing Clonal amplification: No empty beads Clonal amplification?? No beads containing more than one amplified fragment 1) Titration: constant number of beads vs. different DNA starting quantities 2) Optimal enrichment: one single fragment amplified millions of times in one single bead
31 3. Second Generation Sequencing Clonal amplification:
32 3. Second Generation Sequencing Clonal amplification:
33 3. Second Generation Sequencing Clonal amplification: Bridge PCR (Illumina):
34 3. Second Generation Sequencing Sequencing: Roche 454 (pyrosequencing, sequencing by synthesis) Roche-454 technology: pyrosequencing (sequencing by synthesis) Metal coated PTP reduces crosstalk 29 μm well diameter (20/bead) 3,400,000 wells per PTP
35 3. Second Generation Sequencing Sequencing: Roche 454 (pyrosequencing, sequencing by synthesis) CCD Camera flowgram (signal intensity is proportional to the number of nucleotides incorporated in the sequence) - throughput limited by the nº of wells in the PTP - errors in homopolymers (454) - long sequences (up to 1000bp) - low throughput, very expensive reagents - required for some specific applications, advisable for others ( de novo sequencing)
36 3. Second Generation Sequencing Sequencing: Illumina (dye terminator nt, sequencing by synthesis) - Limited by the fragment length than can efectively bridge - Labelled nucleotides are not incorporated as efficiently as native ones - Short sequences (improving, specially in MiSeq PE 2x300 bp) -Strand-specific errors, substitutions towards the end of the read, base substitution errors (sistematic error GGT >GGG) -Scalable set of machines suitable for nearly all the applications -High throughput, expensive machines, cost per Mb OK
37 3. Second Generation Sequencing Illumina workflow. B8E&v=fCd6B5HRaZ8
38 3. Second Generation Sequencing Sequencing: Ion S5/PGM Beads containing the clonally amplified library are loaded onto the chip The chip is run in the machine
39 3. Second Generation Sequencing Sequencing: Ion Torrent
40 3. Second Generation Sequencing Sequencing: Ion Torrent
41 3. Second Generation Sequencing Small NGS platforms GS Junior Plus (Roche) Read Length: 700 pb Output: 70 Mb Running time: 18 hours Ion Torrent PGM (LifeTechnol.) Read length: 35 to 400 bp Output: 30Mb 2Gb Running time: 2,3-4,4 hours Applications: Amplicon sequencing Targeted sequencing Metagenomics (16S) Small genome (i.e. Bacteria, fungi..) sequencing RNA profiling MiSeq (Illumina) Read Length: 2x150 bp Output: Gb Running time: 4-24 hours Source: Supplier webpage
42 3. Second Generation Sequencing High througput NGS Platforms NGS platforms GS FLX+ Read length: 700 bp Output: 1 Mb Running time: 23 hours Ion S5 XL Read length: Up to 200 bp Output: Gb Run time: 2.5 hours HiSeq 4000 Read length: 2x150 bp Output: Gb Run time: <1-3.5 days Applications: Amplicon sequencing Targeted sequencing Metagenomics (16S) Genomes Exomes, transcriptome Source: Supplier webpage
43 1. Introduction to NGS 2. First Generation Sequencing 3. Second Generation Sequencing 4. Third Generation Sequencing 5. Sequencing generation face to face 6. Applications of NGS techniques 7. A (very) brief introduction to DoE
44 4. Third Generation Sequencing TGS: Single molecule sequencing Advantages: Less sample preparation (no PCR) No amplification No PCR errors Fewer contamination issues No GC-bias Analyze every sample (unpcrable, unclonable) Analyze low quality DNA (forensics samples, archeological) Absolute quantification Sequence RNA directly
45 4. Third Generation Sequencing Helicos Genetic Analysis system Workflow similar to Illumina, but without bridge amplification: relative slow and expensive short reads Bankruptcy in 2002
46 4. Third Generation Sequencing SMRT (Pacific bioscience) Quick library construction (6 hours) Run time: from 30 min to 6 hours Sequencing by synthesis No library amplification Long reads: 5 Kb 20 Kb High error rate (but improving!) Source: Supplier webpage
47 4. Third Generation Sequencing SMRT (Pacific bioscience)
48 4. Third Generation Sequencing Oxford nanopore alpha-hemolysin Heptameric protein with a pore of inner diameter 1nm Pore diameter same scale of DNA Protein nanopores can be adapted. The company has optimised its large-scale production. DNA, RNA, mirna and protein analysis Changes in membrane voltage are measured
49 4. Third Generation Sequencing Oxford nanopore SmidgION MinION PromethION Simple 10 minute sample prep available Read length: hundreds of kb Very fast, but high error rates. Easy to work in the field (Ebola viruses sequenced in Guinea 2 days after sample collection, Quick J, 2016)
50 4. Third Generation Sequencing Oxford nanopore
51 4. Third Generation Sequencing High througput 3rd generation instruments Benchtop Pacific Bioscience Sequel system xford Nanopore Technologies PromethION GridION minion
52 1. Introduction to NGS 2. First Generation Sequencing 3. Second Generation Sequencing 4. Third Generation Sequencing 5. Sequencing generation face to face 6. Applications of NGS techniques 7. A (very) brief introduction to DoE
53 5. Sequencing Generation face to face Speed variation by technology
54 5. Sequencing Generation face to face
55 5.Sequencing Generation face to face
56 5.Sequencing Generation face to face 1. DNA fragmentation. 1. DNA fragmentation 1. DNA fragmentation 2. Vector cloning, bacterial transformation and growth, DNA isolation and purification 2. In vitro adaptor ligation + clonal amplification 2. y 3. in vitro adaptor ligation. NO AMPLIFICATION. Massive parallel sequencing. 3. Sequencing (chainterminating ddnucleotides) ssdna DNA primer Polimerase dntps Fluorescent ddntps 3. Massive paralel sequencing 4. Image processing Capillary electrophoresis (1 Sequence/Capilar. Up to 384 DNA samples in a single run) CTATGCTCG 4. Image processing and data analysis 4. Image processing and data analysis. FGS SGS TGS
57 5.Sequencing Generation face to face Release dates vs Machine outputs per run
58 1. Introduction to NGS 2. First Generation Sequencing 3. Second Generation Sequencing 4. Third Generation Sequencing 5. Sequencing generation face to face 6. Applications of NGS techniques 7. A (very) brief introduction to DoE
59 6. Applications of NGS techniques
60 6. Applications of NGS techniques
61 6. Applications of NGS techniques
62 6. Applications of NGS techniques Whole Genome sequencing De novo sequencing Resequencing
63 6. Applications of NGS techniques Whole Genome sequencing Complete characterization of the entire genome. Beijing Genomics Institute ( Does it look cute, we ll sequence it ) Since now, genome-wide association studies (microarray-based) for identifying disease associations across the whole genome (4 million markers). Now it is better to sequence the whole genome (3.2 billion bases) It can be applied not only for human genomes, also for plant, microbial The rapid drop in sequencing costs allow researchers to sequence a genome quickly. PROS Global genome picture, no systematic missing of information Useful for diseases that involve multiple genetic phenomena CONS: Can miss variants in the exonics regions due to lower coverage 2nd generation NGS: some regions cannot be sequenced/assembled (i.e. repetitive regions, GC rich regions ) More expensive and time consuming
64 6. Applications of NGS techniques Expression analysis Since now, performed with microarrays technology Sensitivity of sequence based studies is limited by the depth of sequencing Different approaches can be used: Targeted sequencing Exome sequencing RNA-seq
65 6. Applications of NGS techniques Targeted sequencing Only a subset of genes or regions of the genome are isolated and sequenced. It allows to focus times, expenses and data analysis on specific areas of interest. Enables sequencing at much higher coverage levels. Target sequencing panels can be purchased with preselected content or can be custom designed.
66 6. Applications of NGS techniques Exome sequencing PROS Cheaper and faster Simplified bioinformatics analysis (Often) useful for Mendelian disorders Increases coverage in coding regions: variants are more easily detected CONS: Misses non-coding variation and structural variation Biases due to library methology used Sometimes unsuccessful to interpret clinical data
67 6. Applications of NGS techniques RNA sequencing (RNA-Seq) RNA-seq works by sequencing every RNA molecule and profiling the expression of a particular gene by counting the number of time its transcripts have been sequenced. RNA-seq applications: Differential gene expression analysis (DGE) SNPs, splice variants (resolution at base-level) Detection of novel transcripts and isoforms Transcriptome of not previously studied organisms Map exon/intron boundaries, splice junctions Detection of allele specific expression patterns
68 6. Applications of NGS techniques RNAseq Very high dynamic range (10 5 to 10 7 )
69 6. Applications of NGS techniques Epigenomics Epigenome: different chromatin states at the whole genome level Epigenetic modifications: histone variants, post-translational modifications of aminoacids on the N-terminal tail of histones, covalent modifications of the DNA bases Some approaches: ChIPSeq: transcription factor binding and histone modifications Bisulfite conversion: unmet-c converted to U. Single CpG resolution. Gold standard for DNA methylation analysis. MeDIP: enrichment for the methylated/unmethylated fractions of the genome using 5-methyl-c antibodies
70 6. Applications of NGS techniques The promise of single cell sequencing... Ability to detect mutations in single cells in a population Now could be possible with TGS Tumour genomes are highly heterogeneous Haplotype determination (HLA) for bone marrow transplants Mosaics in disease (Alzheimer) Preimplantation genetic diagnosis Cell line heterogenicity Nature Methods Vol.11(1) 2014
71 6. Applications of NGS techniques Metagenomics Is a way to make an inventory of what (DNA) is present in a sample Two approaches: Sequence it all Focus on specific conserved sequences (ribosomal genes) Technology facilitates the study of the consequences of environmental changes and the causes of the changes.
72 1. Introduction to NGS 2. First Generation Sequencing 3. Second Generation Sequencing 4. Third Generation Sequencing 5. Sequencing generation face to face 6. Applications of NGS techniques 7. A (very) brief introduction to DoE
73 7. A (very) brief introduction to DoE The (statistical) design of experiments is an efficient procedure for planning experiments so that the data obtained can be analyzed to yield valid and objective conclusions Why are many life scientists so adverse to thinking about design? It is common to think that time spent designing experiments would be better spent actually doing experiments
74 7. A (very) brief introduction to DoE Variability types that play in an experiment: Planned systematic variability: This is the differences in response between treatments applied. Noise variability: random noise. Differences between two consecutives measures. We cannot avoid that. Systematic variability not planned: Produce a systematic variation in the results. A priori the reason is not known. It can be avoided with the randomization and the local control.
75 7. A (very) brief introduction to DoE Important steps to define before begin the experiment: Establish the main objectives of the experiment. Avoid collateral problems Identify all the noise sources: Treatment, experimental errors, Allocate each experimental unit which each treatment Clarify the type of response expected in each treatment Determinate the number of individuals in each group Run a pilot study
76 7. A (very) brief introduction to DoE Important steps to define before begin the experiment: Establish the main objectives of the experiment. Avoid collateral problems Identify all the noise sources: Treatment, experimental errors, Allocate each experimental unit which each treatment Clarify the type of response expected in each treatment Determinate the number of individuals in each group Run a pilot study
77 7. A (very) brief introduction to DoE Important steps to define before begin the experiment: Establish the main objectives of the experiment. Avoid collateral problems Identify all the noise sources: Treatment, experimental errors, Allocate each experimental unit which each treatment Clarify the type of response expected in each treatment Determinate the number of individuals in each group Run a pilot study
78 7. A (very) brief introduction to DoE Important steps to define before begin the experiment: Establish the main objectives of the experiment. Avoid collateral problems Identify all the noise sources: Treatment, experimental errors, Allocate each experimental unit which each treatment Clarify the type of response expected in each treatment Determinate the number of individuals in each group Run a pilot study How the data will be statistically analysed.
79 7. A (very) brief introduction to DoE Sample Treatment Sex Batch Sample Treatment Sex Batch 1 A Male 1 1 A Male 1 2 A Male 1 2 A Female 2 3 A Male 1 3 A Male 2 4 A Male 1 4 A Female 1 5 B Female 2 5 B Male 1 6 B Female 2 6 B Female 2 7 B Female 2 7 B Male 2 8 B Female 2 8 B Female 1 Treatment are confounded between sex and batch Treatment is well balanced
80 7. A (very) brief introduction to DoE What do you need to ask before starting a NGS experiment? -What do I want to sequence? Whole genome, exome, metagenome, epigenome, RNAseq... -How many samples? -Length of read required? -Quality and quantity of starting material? -Size of nucleic acids to sequence -Amount of sequence needed: coverage (Depth of) Coverage: how many times a particular base is sequenced. 30x = each base has been read by 30 sequences (in average) (Breadth of) Coverage: amount of the target sequence that has been covered (with a given coverage)
Deep Sequencing technologies
 Deep Sequencing technologies Gabriela Salinas 30 October 2017 Transcriptome and Genome Analysis Laboratory http://www.uni-bc.gwdg.de/index.php?id=709 Microarray and Deep-Sequencing Core Facility University
Deep Sequencing technologies Gabriela Salinas 30 October 2017 Transcriptome and Genome Analysis Laboratory http://www.uni-bc.gwdg.de/index.php?id=709 Microarray and Deep-Sequencing Core Facility University
Third Generation Sequencing
 Third Generation Sequencing By Mohammad Hasan Samiee Aref Medical Genetics Laboratory of Dr. Zeinali History of DNA sequencing 1953 : Discovery of DNA structure by Watson and Crick 1973 : First sequence
Third Generation Sequencing By Mohammad Hasan Samiee Aref Medical Genetics Laboratory of Dr. Zeinali History of DNA sequencing 1953 : Discovery of DNA structure by Watson and Crick 1973 : First sequence
Overview of Next Generation Sequencing technologies. Céline Keime
 Overview of Next Generation Sequencing technologies Céline Keime keime@igbmc.fr Next Generation Sequencing < Second generation sequencing < General principle < Sequencing by synthesis - Illumina < Sequencing
Overview of Next Generation Sequencing technologies Céline Keime keime@igbmc.fr Next Generation Sequencing < Second generation sequencing < General principle < Sequencing by synthesis - Illumina < Sequencing
Research school methods seminar Genomics and Transcriptomics
 Research school methods seminar Genomics and Transcriptomics Stephan Klee 19.11.2014 2 3 4 5 Genetics, Genomics what are we talking about? Genetics and Genomics Study of genes Role of genes in inheritence
Research school methods seminar Genomics and Transcriptomics Stephan Klee 19.11.2014 2 3 4 5 Genetics, Genomics what are we talking about? Genetics and Genomics Study of genes Role of genes in inheritence
Next Generation Sequencing. Jeroen Van Houdt - Leuven 13/10/2017
 Next Generation Sequencing Jeroen Van Houdt - Leuven 13/10/2017 Landmarks in DNA sequencing 1953 Discovery of DNA double helix structure 1977 A Maxam and W Gilbert "DNA seq by chemical degradation" F Sanger"DNA
Next Generation Sequencing Jeroen Van Houdt - Leuven 13/10/2017 Landmarks in DNA sequencing 1953 Discovery of DNA double helix structure 1977 A Maxam and W Gilbert "DNA seq by chemical degradation" F Sanger"DNA
Next-generation sequencing Technology Overview
 Next-generation sequencing Technology Overview UQ Winter School 2018 Christopher Noune, PhD AGRF Melbourne christopher.noune@agrf.org.au What is NGS? Ion Torrent PGM (Thermo-Fisher) MiSeq (Illumina) High-Throughput
Next-generation sequencing Technology Overview UQ Winter School 2018 Christopher Noune, PhD AGRF Melbourne christopher.noune@agrf.org.au What is NGS? Ion Torrent PGM (Thermo-Fisher) MiSeq (Illumina) High-Throughput
Matthew Tinning Australian Genome Research Facility. July 2012
 Next-Generation Sequencing: an overview of technologies and applications Matthew Tinning Australian Genome Research Facility July 2012 History of Sequencing Where have we been? 1869 Discovery of DNA 1909
Next-Generation Sequencing: an overview of technologies and applications Matthew Tinning Australian Genome Research Facility July 2012 History of Sequencing Where have we been? 1869 Discovery of DNA 1909
DNA-Sequencing. Technologies & Devices. Matthias Platzer. Genome Analysis Leibniz Institute on Aging - Fritz Lipmann Institute (FLI)
 DNA-Sequencing Technologies & Devices Matthias Platzer Genome Analysis Leibniz Institute on Aging - Fritz Lipmann Institute (FLI) Genome analysis DNA sequencing platforms ABI 3730xl 4/2004 & 6/2006 1 Mb/day,
DNA-Sequencing Technologies & Devices Matthias Platzer Genome Analysis Leibniz Institute on Aging - Fritz Lipmann Institute (FLI) Genome analysis DNA sequencing platforms ABI 3730xl 4/2004 & 6/2006 1 Mb/day,
The Journey of DNA Sequencing. Chromosomes. What is a genome? Genome size. H. Sunny Sun
 The Journey of DNA Sequencing H. Sunny Sun What is a genome? Genome is the total genetic complement of a living organism. The nuclear genome comprises approximately 3.2 * 10 9 nucleotides of DNA, divided
The Journey of DNA Sequencing H. Sunny Sun What is a genome? Genome is the total genetic complement of a living organism. The nuclear genome comprises approximately 3.2 * 10 9 nucleotides of DNA, divided
DNA-Sequencing. Technologies & Devices. Matthias Platzer. Genome Analysis Leibniz Institute on Aging - Fritz Lipmann Institute (FLI)
 DNA-Sequencing Technologies & Devices Matthias Platzer Genome Analysis Leibniz Institute on Aging - Fritz Lipmann Institute (FLI) Genome analysis DNA sequencing platforms ABI 3730xl 4/2004 & 6/2006 1 Mb/day,
DNA-Sequencing Technologies & Devices Matthias Platzer Genome Analysis Leibniz Institute on Aging - Fritz Lipmann Institute (FLI) Genome analysis DNA sequencing platforms ABI 3730xl 4/2004 & 6/2006 1 Mb/day,
Welcome to the NGS webinar series
 Welcome to the NGS webinar series Webinar 1 NGS: Introduction to technology, and applications NGS Technology Webinar 2 Targeted NGS for Cancer Research NGS in cancer Webinar 3 NGS: Data analysis for genetic
Welcome to the NGS webinar series Webinar 1 NGS: Introduction to technology, and applications NGS Technology Webinar 2 Targeted NGS for Cancer Research NGS in cancer Webinar 3 NGS: Data analysis for genetic
High Throughput Sequencing Technologies. J Fass UCD Genome Center Bioinformatics Core Monday September 15, 2014
 High Throughput Sequencing Technologies J Fass UCD Genome Center Bioinformatics Core Monday September 15, 2014 Sequencing Explosion www.genome.gov/sequencingcosts http://t.co/ka5cvghdqo Sequencing Explosion
High Throughput Sequencing Technologies J Fass UCD Genome Center Bioinformatics Core Monday September 15, 2014 Sequencing Explosion www.genome.gov/sequencingcosts http://t.co/ka5cvghdqo Sequencing Explosion
High Throughput Sequencing Technologies. J Fass UCD Genome Center Bioinformatics Core Monday June 16, 2014
 High Throughput Sequencing Technologies J Fass UCD Genome Center Bioinformatics Core Monday June 16, 2014 Sequencing Explosion www.genome.gov/sequencingcosts http://t.co/ka5cvghdqo Sequencing Explosion
High Throughput Sequencing Technologies J Fass UCD Genome Center Bioinformatics Core Monday June 16, 2014 Sequencing Explosion www.genome.gov/sequencingcosts http://t.co/ka5cvghdqo Sequencing Explosion
Next Generation Sequencing (NGS)
 Next Generation Sequencing (NGS) Fernando Alvarez Sección Biomatemática, Facultad de Ciencias, UdelaR 1 Uruguay Montevide o 3 TANGO World Champ 1930 1950 (Maraca 4 Next Generation Sequencing module Next
Next Generation Sequencing (NGS) Fernando Alvarez Sección Biomatemática, Facultad de Ciencias, UdelaR 1 Uruguay Montevide o 3 TANGO World Champ 1930 1950 (Maraca 4 Next Generation Sequencing module Next
DNA-Sequencing. Technologies & Devices
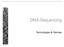 DNA-Sequencing Technologies & Devices Genome analysis DNA sequencing platforms ABI 3730xl 4/2004 & 6/2006 1 Mb/day, 850 nt reads 2 Mb/day, 550 nt reads Roche/454 GS FLX 12/2006 800 Mb/23h, 800 nt reads
DNA-Sequencing Technologies & Devices Genome analysis DNA sequencing platforms ABI 3730xl 4/2004 & 6/2006 1 Mb/day, 850 nt reads 2 Mb/day, 550 nt reads Roche/454 GS FLX 12/2006 800 Mb/23h, 800 nt reads
Aaron Liston, Oregon State University Botany 2012 Intro to Next Generation Sequencing Workshop
 Output (bp) Aaron Liston, Oregon State University Growth in Next-Gen Sequencing Capacity 3.5E+11 2002 2004 2006 2008 2010 3.0E+11 2.5E+11 2.0E+11 1.5E+11 1.0E+11 Adapted from Mardis, 2011, Nature 5.0E+10
Output (bp) Aaron Liston, Oregon State University Growth in Next-Gen Sequencing Capacity 3.5E+11 2002 2004 2006 2008 2010 3.0E+11 2.5E+11 2.0E+11 1.5E+11 1.0E+11 Adapted from Mardis, 2011, Nature 5.0E+10
DNA-Sequencing. Technologies & Devices
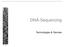 DNA-Sequencing Technologies & Devices Genome analysis DNA sequencing platforms ABI 3730xl 4/2004 & 6/2006 1 Mb/day, 850 nt reads 2 Mb/day, 550 nt reads Roche/454 GS FLX 12/2006 800 Mb/23h, 800 nt reads
DNA-Sequencing Technologies & Devices Genome analysis DNA sequencing platforms ABI 3730xl 4/2004 & 6/2006 1 Mb/day, 850 nt reads 2 Mb/day, 550 nt reads Roche/454 GS FLX 12/2006 800 Mb/23h, 800 nt reads
Sequencing technologies. Jose Blanca COMAV institute bioinf.comav.upv.es
 Sequencing technologies Jose Blanca COMAV institute bioinf.comav.upv.es Outline Sequencing technologies: Sanger 2nd generation sequencing: 3er generation sequencing: 454 Illumina SOLiD Ion Torrent PacBio
Sequencing technologies Jose Blanca COMAV institute bioinf.comav.upv.es Outline Sequencing technologies: Sanger 2nd generation sequencing: 3er generation sequencing: 454 Illumina SOLiD Ion Torrent PacBio
Next generation sequencing techniques" Toma Tebaldi Centre for Integrative Biology University of Trento
 Next generation sequencing techniques" Toma Tebaldi Centre for Integrative Biology University of Trento Mattarello September 28, 2009 Sequencing Fundamental task in modern biology read the information
Next generation sequencing techniques" Toma Tebaldi Centre for Integrative Biology University of Trento Mattarello September 28, 2009 Sequencing Fundamental task in modern biology read the information
Sequencing technologies. Jose Blanca COMAV institute bioinf.comav.upv.es
 Sequencing technologies Jose Blanca COMAV institute bioinf.comav.upv.es Outline Sequencing technologies: Sanger 2nd generation sequencing: 3er generation sequencing: 454 Illumina SOLiD Ion Torrent PacBio
Sequencing technologies Jose Blanca COMAV institute bioinf.comav.upv.es Outline Sequencing technologies: Sanger 2nd generation sequencing: 3er generation sequencing: 454 Illumina SOLiD Ion Torrent PacBio
Next Generation Sequencing. Tobias Österlund
 Next Generation Sequencing Tobias Österlund tobiaso@chalmers.se NGS part of the course Week 4 Friday 13/2 15.15-17.00 NGS lecture 1: Introduction to NGS, alignment, assembly Week 6 Thursday 26/2 08.00-09.45
Next Generation Sequencing Tobias Österlund tobiaso@chalmers.se NGS part of the course Week 4 Friday 13/2 15.15-17.00 NGS lecture 1: Introduction to NGS, alignment, assembly Week 6 Thursday 26/2 08.00-09.45
High Throughput Sequencing the Multi-Tool of Life Sciences. Lutz Froenicke DNA Technologies and Expression Analysis Cores UCD Genome Center
 High Throughput Sequencing the Multi-Tool of Life Sciences Lutz Froenicke DNA Technologies and Expression Analysis Cores UCD Genome Center Complementary Approaches Illumina Still-imaging of clusters (~1000
High Throughput Sequencing the Multi-Tool of Life Sciences Lutz Froenicke DNA Technologies and Expression Analysis Cores UCD Genome Center Complementary Approaches Illumina Still-imaging of clusters (~1000
Contact us for more information and a quotation
 GenePool Information Sheet #1 Installed Sequencing Technologies in the GenePool The GenePool offers sequencing service on three platforms: Sanger (dideoxy) sequencing on ABI 3730 instruments Illumina SOLEXA
GenePool Information Sheet #1 Installed Sequencing Technologies in the GenePool The GenePool offers sequencing service on three platforms: Sanger (dideoxy) sequencing on ABI 3730 instruments Illumina SOLEXA
Sequencing technologies. Jose Blanca COMAV institute bioinf.comav.upv.es
 Sequencing technologies Jose Blanca COMAV institute bioinf.comav.upv.es Outline Sequencing technologies: Sanger 2nd generation sequencing: 3er generation sequencing: 454 Illumina SOLiD Ion Torrent PacBio
Sequencing technologies Jose Blanca COMAV institute bioinf.comav.upv.es Outline Sequencing technologies: Sanger 2nd generation sequencing: 3er generation sequencing: 454 Illumina SOLiD Ion Torrent PacBio
Introduction to NGS. Josef K Vogt Slides by: Simon Rasmussen Next Generation Sequencing Analysis
 Introduction to NGS Josef K Vogt Slides by: Simon Rasmussen 2017 Life science data deluge Massive unstructured data from several areas DNA, patient journals, proteomics, imaging,... Impacts Industry, Environment,
Introduction to NGS Josef K Vogt Slides by: Simon Rasmussen 2017 Life science data deluge Massive unstructured data from several areas DNA, patient journals, proteomics, imaging,... Impacts Industry, Environment,
Introduction to Next Generation Sequencing (NGS)
 Introduction to Next eneration Sequencing (NS) Simon Rasmussen Assistant Professor enter for Biological Sequence analysis Technical University of Denmark 2012 Today 9.00-9.45: Introduction to NS, How it
Introduction to Next eneration Sequencing (NS) Simon Rasmussen Assistant Professor enter for Biological Sequence analysis Technical University of Denmark 2012 Today 9.00-9.45: Introduction to NS, How it
A Crash Course in NGS for GI Pathologists. Sandra O Toole
 A Crash Course in NGS for GI Pathologists Sandra O Toole The Sanger Technique First generation sequencing Uses dideoxynucleotides (dideoxyadenine, dideoxyguanine, etc) These are molecules that resemble
A Crash Course in NGS for GI Pathologists Sandra O Toole The Sanger Technique First generation sequencing Uses dideoxynucleotides (dideoxyadenine, dideoxyguanine, etc) These are molecules that resemble
High Throughput Sequencing Technologies. UCD Genome Center Bioinformatics Core Monday 15 June 2015
 High Throughput Sequencing Technologies UCD Genome Center Bioinformatics Core Monday 15 June 2015 Sequencing Explosion www.genome.gov/sequencingcosts http://t.co/ka5cvghdqo Sequencing Explosion 2011 PacBio
High Throughput Sequencing Technologies UCD Genome Center Bioinformatics Core Monday 15 June 2015 Sequencing Explosion www.genome.gov/sequencingcosts http://t.co/ka5cvghdqo Sequencing Explosion 2011 PacBio
High Throughput Sequencing Technologies. J Fass UCD Genome Center Bioinformatics Core Tuesday December 16, 2014
 High Throughput Sequencing Technologies J Fass UCD Genome Center Bioinformatics Core Tuesday December 16, 2014 Sequencing Explosion www.genome.gov/sequencingcosts http://t.co/ka5cvghdqo Sequencing Explosion
High Throughput Sequencing Technologies J Fass UCD Genome Center Bioinformatics Core Tuesday December 16, 2014 Sequencing Explosion www.genome.gov/sequencingcosts http://t.co/ka5cvghdqo Sequencing Explosion
Outline General NGS background and terms 11/14/2016 CONFLICT OF INTEREST. HLA region targeted enrichment. NGS library preparation methodologies
 Eric T. Weimer, PhD, D(ABMLI) Assistant Professor, Pathology & Laboratory Medicine, UNC School of Medicine Director, Molecular Immunology Associate Director, Clinical Flow Cytometry, HLA, and Immunology
Eric T. Weimer, PhD, D(ABMLI) Assistant Professor, Pathology & Laboratory Medicine, UNC School of Medicine Director, Molecular Immunology Associate Director, Clinical Flow Cytometry, HLA, and Immunology
Next-generation sequencing technologies
 Next-generation sequencing technologies NGS applications Illumina sequencing workflow Overview Sequencing by ligation Short-read NGS Sequencing by synthesis Illumina NGS Single-molecule approach Long-read
Next-generation sequencing technologies NGS applications Illumina sequencing workflow Overview Sequencing by ligation Short-read NGS Sequencing by synthesis Illumina NGS Single-molecule approach Long-read
TREE CODE PRODUCT BROCHURE
 TREE CODE PRODUCT BROCHURE Single Molecule, Real-Time (SMRT) Sequencing technology offers: Long read sequencing ~10 Gb with 20 kb average read lengths for WGS ~20 Gb with 40 kb average read length for
TREE CODE PRODUCT BROCHURE Single Molecule, Real-Time (SMRT) Sequencing technology offers: Long read sequencing ~10 Gb with 20 kb average read lengths for WGS ~20 Gb with 40 kb average read length for
Phenotype analysis: biological-biochemical analysis. Genotype analysis: molecular and physical analysis
 1 Genetic Analysis Phenotype analysis: biological-biochemical analysis Behaviour under specific environmental conditions Behaviour of specific genetic configurations Behaviour of progeny in crosses - Genotype
1 Genetic Analysis Phenotype analysis: biological-biochemical analysis Behaviour under specific environmental conditions Behaviour of specific genetic configurations Behaviour of progeny in crosses - Genotype
Understanding the science and technology of whole genome sequencing
 Understanding the science and technology of whole genome sequencing Dag Undlien Department of Medical Genetics Oslo University Hospital University of Oslo and The Norwegian Sequencing Centre d.e.undlien@medisin.uio.no
Understanding the science and technology of whole genome sequencing Dag Undlien Department of Medical Genetics Oslo University Hospital University of Oslo and The Norwegian Sequencing Centre d.e.undlien@medisin.uio.no
High Throughput Sequencing the Multi-Tool of Life Sciences. Lutz Froenicke DNA Technologies and Expression Analysis Cores UCD Genome Center
 High Throughput Sequencing the Multi-Tool of Life Sciences Lutz Froenicke DNA Technologies and Expression Analysis Cores UCD Genome Center DNA Technologies & Expression Analysis Cores HT Sequencing (Illumina
High Throughput Sequencing the Multi-Tool of Life Sciences Lutz Froenicke DNA Technologies and Expression Analysis Cores UCD Genome Center DNA Technologies & Expression Analysis Cores HT Sequencing (Illumina
Phenotype analysis: biological-biochemical analysis. Genotype analysis: molecular and physical analysis
 1 Genetic Analysis Phenotype analysis: biological-biochemical analysis Behaviour under specific environmental conditions Behaviour of specific genetic configurations Behaviour of progeny in crosses - Genotype
1 Genetic Analysis Phenotype analysis: biological-biochemical analysis Behaviour under specific environmental conditions Behaviour of specific genetic configurations Behaviour of progeny in crosses - Genotype
Next-Generation Sequencing. Technologies
 Next-Generation Next-Generation Sequencing Technologies Sequencing Technologies Nicholas E. Navin, Ph.D. MD Anderson Cancer Center Dept. Genetics Dept. Bioinformatics Introduction to Bioinformatics GS011062
Next-Generation Next-Generation Sequencing Technologies Sequencing Technologies Nicholas E. Navin, Ph.D. MD Anderson Cancer Center Dept. Genetics Dept. Bioinformatics Introduction to Bioinformatics GS011062
Genome Resequencing. Rearrangements. SNPs, Indels CNVs. De novo genome Sequencing. Metagenomics. Exome Sequencing. RNA-seq Gene Expression
 Genome Resequencing De novo genome Sequencing SNPs, Indels CNVs Rearrangements Metagenomics RNA-seq Gene Expression Splice Isoform Abundance High Throughput Short Read Sequencing: Illumina Exome Sequencing
Genome Resequencing De novo genome Sequencing SNPs, Indels CNVs Rearrangements Metagenomics RNA-seq Gene Expression Splice Isoform Abundance High Throughput Short Read Sequencing: Illumina Exome Sequencing
Using New ThiNGS on Small Things. Shane Byrne
 Using New ThiNGS on Small Things Shane Byrne Next Generation Sequencing New Things Small Things NGS Next Generation Sequencing = 2 nd generation of sequencing 454 GS FLX, SOLiD, GAIIx, HiSeq, MiSeq, Ion
Using New ThiNGS on Small Things Shane Byrne Next Generation Sequencing New Things Small Things NGS Next Generation Sequencing = 2 nd generation of sequencing 454 GS FLX, SOLiD, GAIIx, HiSeq, MiSeq, Ion
Wet-lab Considerations for Illumina data analysis
 Wet-lab Considerations for Illumina data analysis Based on a presentation by Henriette O Geen Lutz Froenicke DNA Technologies and Expression Analysis Cores UCD Genome Center Complementary Approaches Illumina
Wet-lab Considerations for Illumina data analysis Based on a presentation by Henriette O Geen Lutz Froenicke DNA Technologies and Expression Analysis Cores UCD Genome Center Complementary Approaches Illumina
CM581A2: NEXT GENERATION SEQUENCING PLATFORMS AND LIBRARY GENERATION
 CM581A2: NEXT GENERATION SEQUENCING PLATFORMS AND LIBRARY GENERATION Fall 2015 Instructors: Coordinator: Carol Wilusz, Associate Professor MIP, CMB Instructor: Dan Sloan, Assistant Professor, Biology,
CM581A2: NEXT GENERATION SEQUENCING PLATFORMS AND LIBRARY GENERATION Fall 2015 Instructors: Coordinator: Carol Wilusz, Associate Professor MIP, CMB Instructor: Dan Sloan, Assistant Professor, Biology,
02 Agenda Item 03 Agenda Item
 01 Agenda Item 02 Agenda Item 03 Agenda Item SOLiD 3 System: Applications Overview April 12th, 2010 Jennifer Stover Field Application Specialist - SOLiD Applications Workflow for SOLiD Application Application
01 Agenda Item 02 Agenda Item 03 Agenda Item SOLiD 3 System: Applications Overview April 12th, 2010 Jennifer Stover Field Application Specialist - SOLiD Applications Workflow for SOLiD Application Application
Chapter 7. DNA Microarrays
 Bioinformatics III Structural Bioinformatics and Genome Analysis Chapter 7. DNA Microarrays 7.9 Next Generation Sequencing 454 Sequencing Solexa Illumina Solid TM System Sequencing Process of determining
Bioinformatics III Structural Bioinformatics and Genome Analysis Chapter 7. DNA Microarrays 7.9 Next Generation Sequencing 454 Sequencing Solexa Illumina Solid TM System Sequencing Process of determining
FUTURE PROSPECTS IN MOLECULAR INFECTIOUS DISEASES DIAGNOSIS
 FUTURE PROSPECTS IN MOLECULAR INFECTIOUS DISEASES DIAGNOSIS Richard L. Hodinka, Ph.D. University of South Carolina School of Medicine Greenville Greenville Health System, Greenville, SC hodinka@greenvillemed.sc.edu
FUTURE PROSPECTS IN MOLECULAR INFECTIOUS DISEASES DIAGNOSIS Richard L. Hodinka, Ph.D. University of South Carolina School of Medicine Greenville Greenville Health System, Greenville, SC hodinka@greenvillemed.sc.edu
Next Gen Sequencing. Expansion of sequencing technology. Contents
 Next Gen Sequencing Contents 1 Expansion of sequencing technology 2 The Next Generation of Sequencing: High-Throughput Technologies 3 High Throughput Sequencing Applied to Genome Sequencing (TEDed CC BY-NC-ND
Next Gen Sequencing Contents 1 Expansion of sequencing technology 2 The Next Generation of Sequencing: High-Throughput Technologies 3 High Throughput Sequencing Applied to Genome Sequencing (TEDed CC BY-NC-ND
HLA-Typing Strategies
 HLA-Typing Strategies Cologne, 13.5.2017 Joannis Mytilineos MD, PhD Department of Transplantation Immunology Institute for Clinical Transfusion Medicine and Immunogenetics German Red Cross Blood Transfusion
HLA-Typing Strategies Cologne, 13.5.2017 Joannis Mytilineos MD, PhD Department of Transplantation Immunology Institute for Clinical Transfusion Medicine and Immunogenetics German Red Cross Blood Transfusion
MHC Region. MHC expression: Class I: All nucleated cells and platelets Class II: Antigen presenting cells
 DNA based HLA typing methods By: Yadollah Shakiba, MD, PhD MHC Region MHC expression: Class I: All nucleated cells and platelets Class II: Antigen presenting cells Nomenclature of HLA Alleles Assigned
DNA based HLA typing methods By: Yadollah Shakiba, MD, PhD MHC Region MHC expression: Class I: All nucleated cells and platelets Class II: Antigen presenting cells Nomenclature of HLA Alleles Assigned
Applications of Next Generation Sequencing in Metagenomics Studies
 Applications of Next Generation Sequencing in Metagenomics Studies Francesca Rizzo, PhD Genomix4life Laboratory of Molecular Medicine and Genomics Department of Medicine and Surgery University of Salerno
Applications of Next Generation Sequencing in Metagenomics Studies Francesca Rizzo, PhD Genomix4life Laboratory of Molecular Medicine and Genomics Department of Medicine and Surgery University of Salerno
NGS technologies: a user s guide. Karim Gharbi & Mark Blaxter
 NGS technologies: a user s guide Karim Gharbi & Mark Blaxter genepool-manager@ed.ac.uk Natural history of sequencing 2 Brief history of sequencing 100s bp throughput 100 Gb 1977 1986 1995 1999 2005 2007
NGS technologies: a user s guide Karim Gharbi & Mark Blaxter genepool-manager@ed.ac.uk Natural history of sequencing 2 Brief history of sequencing 100s bp throughput 100 Gb 1977 1986 1995 1999 2005 2007
DNA-Sequenzierung. Technologien & Geräte
 DNA-Sequenzierung Technologien & Geräte Genome analysis DNA sequencing platforms ABI 3730xl 4/2004 & 6/2006 1 Mb/day, 850 nt reads 2 Mb/day, 550 nt reads Roche/454 GS FLX 12/2006 400 Mb/7h, 350 nt reads
DNA-Sequenzierung Technologien & Geräte Genome analysis DNA sequencing platforms ABI 3730xl 4/2004 & 6/2006 1 Mb/day, 850 nt reads 2 Mb/day, 550 nt reads Roche/454 GS FLX 12/2006 400 Mb/7h, 350 nt reads
The New Genome Analyzer IIx Delivering more data, faster, and easier than ever before. Jeremy Preston, PhD Marketing Manager, Sequencing
 The New Genome Analyzer IIx Delivering more data, faster, and easier than ever before Jeremy Preston, PhD Marketing Manager, Sequencing Illumina Genome Analyzer: a Paradigm Shift 2000x gain in efficiency
The New Genome Analyzer IIx Delivering more data, faster, and easier than ever before Jeremy Preston, PhD Marketing Manager, Sequencing Illumina Genome Analyzer: a Paradigm Shift 2000x gain in efficiency
Introduction to NGS. Simon Rasmussen Associate Professor DTU Bioinformatics Technical University of Denmark 2018
 Introduction to NGS Simon Rasmussen Associate Professor DTU Bioinformatics Technical University of Denmark 2018 Life science data deluge Massive unstructured data from several areas DNA, patient journals,
Introduction to NGS Simon Rasmussen Associate Professor DTU Bioinformatics Technical University of Denmark 2018 Life science data deluge Massive unstructured data from several areas DNA, patient journals,
DNA Sequencing by Ion Torrent. Marc Lavergne CHEM 4590
 DNA Sequencing by Ion Torrent Marc Lavergne CHEM 4590 OVERVIEW History DNA Synthesis and First-Gen Sequencing Technology Sequencing Signal Detection Advantages/Disadvantages Applications Current Research
DNA Sequencing by Ion Torrent Marc Lavergne CHEM 4590 OVERVIEW History DNA Synthesis and First-Gen Sequencing Technology Sequencing Signal Detection Advantages/Disadvantages Applications Current Research
RNA Sequencing. Next gen insight into transcriptomes , Elio Schijlen
 RNA Sequencing Next gen insight into transcriptomes 05-06-2013, Elio Schijlen Transcriptome complete set of transcripts in a cell, and their quantity, for a specific developmental stage or physiological
RNA Sequencing Next gen insight into transcriptomes 05-06-2013, Elio Schijlen Transcriptome complete set of transcripts in a cell, and their quantity, for a specific developmental stage or physiological
Targeted Sequencing in the NBS Laboratory
 Targeted Sequencing in the NBS Laboratory Christopher Greene, PhD Newborn Screening and Molecular Biology Branch Division of Laboratory Sciences Gene Sequencing in Public Health Newborn Screening February
Targeted Sequencing in the NBS Laboratory Christopher Greene, PhD Newborn Screening and Molecular Biology Branch Division of Laboratory Sciences Gene Sequencing in Public Health Newborn Screening February
Introducing QIAseq. Accelerate your NGS performance through Sample to Insight solutions. Sample to Insight
 Introducing QIAseq Accelerate your NGS performance through Sample to Insight solutions Sample to Insight From Sample to Insight let QIAGEN enhance your NGS-based research High-throughput next-generation
Introducing QIAseq Accelerate your NGS performance through Sample to Insight solutions Sample to Insight From Sample to Insight let QIAGEN enhance your NGS-based research High-throughput next-generation
Transcriptomics analysis with RNA seq: an overview Frederik Coppens
 Transcriptomics analysis with RNA seq: an overview Frederik Coppens Platforms Applications Analysis Quantification RNA content Platforms Platforms Short (few hundred bases) Long reads (multiple kilobases)
Transcriptomics analysis with RNA seq: an overview Frederik Coppens Platforms Applications Analysis Quantification RNA content Platforms Platforms Short (few hundred bases) Long reads (multiple kilobases)
Human genome sequence
 NGS: the basics Human genome sequence June 26th 2000: official announcement of the completion of the draft of the human genome sequence (truly finished in 2004) Francis Collins Craig Venter HGP: 3 billion
NGS: the basics Human genome sequence June 26th 2000: official announcement of the completion of the draft of the human genome sequence (truly finished in 2004) Francis Collins Craig Venter HGP: 3 billion
solid S Y S T E M s e q u e n c i n g See the Difference Discover the Quality Genome
 solid S Y S T E M s e q u e n c i n g See the Difference Discover the Quality Genome See the Difference With a commitment to your peace of mind, Life Technologies provides a portfolio of robust and scalable
solid S Y S T E M s e q u e n c i n g See the Difference Discover the Quality Genome See the Difference With a commitment to your peace of mind, Life Technologies provides a portfolio of robust and scalable
Molecular Biology and Functional Genomic Core Facility
 Molecular Biology and Functional Genomic Core Facility General Presentation Dr Odile Neyret Core Manager Myriam Rondeau Research Assistant Agnès Dumont Research Assistant Institut de recherche clinique
Molecular Biology and Functional Genomic Core Facility General Presentation Dr Odile Neyret Core Manager Myriam Rondeau Research Assistant Agnès Dumont Research Assistant Institut de recherche clinique
Ultrasequencing: methods and applications of the new generation sequencing platforms
 Ultrasequencing: methods and applications of the new generation sequencing platforms Nuria Tubío Santamaría Course: Genomics Universitat Autònoma de Barcelona 1 Introduction Clasical methods of sequencing:
Ultrasequencing: methods and applications of the new generation sequencing platforms Nuria Tubío Santamaría Course: Genomics Universitat Autònoma de Barcelona 1 Introduction Clasical methods of sequencing:
Functional Genomics Research Stream. Research Meetings: November 2 & 3, 2009 Next Generation Sequencing
 Functional Genomics Research Stream Research Meetings: November 2 & 3, 2009 Next Generation Sequencing Current Issues Research Meetings: Meet with me this Thursday or Friday. (bring laboratory notebook
Functional Genomics Research Stream Research Meetings: November 2 & 3, 2009 Next Generation Sequencing Current Issues Research Meetings: Meet with me this Thursday or Friday. (bring laboratory notebook
Introductie en Toepassingen van Next-Generation Sequencing in de Klinische Virologie. Sander van Boheemen Medical Microbiology
 Introductie en Toepassingen van Next-Generation Sequencing in de Klinische Virologie Sander van Boheemen Medical Microbiology Next-generation sequencing Next-generation sequencing (NGS), also known as
Introductie en Toepassingen van Next-Generation Sequencing in de Klinische Virologie Sander van Boheemen Medical Microbiology Next-generation sequencing Next-generation sequencing (NGS), also known as
Next-generation sequencing and quality control: An introduction 2016
 Next-generation sequencing and quality control: An introduction 2016 s.schmeier@massey.ac.nz http://sschmeier.com/bioinf-workshop/ Overview Typical workflow of a genomics experiment Genome versus transcriptome
Next-generation sequencing and quality control: An introduction 2016 s.schmeier@massey.ac.nz http://sschmeier.com/bioinf-workshop/ Overview Typical workflow of a genomics experiment Genome versus transcriptome
Bioinformatics Advice on Experimental Design
 Bioinformatics Advice on Experimental Design Where do I start? Please refer to the following guide to better plan your experiments for good statistical analysis, best suited for your research needs. Statistics
Bioinformatics Advice on Experimental Design Where do I start? Please refer to the following guide to better plan your experiments for good statistical analysis, best suited for your research needs. Statistics
GENOMICS WORKFLOW SOLUTIONS THAT GO WHERE THE SCIENCE LEADS. Genomics Solutions Portfolio
 GENOMICS WORKFLOW SOLUTIONS THAT GO WHERE THE SCIENCE LEADS Genomics Solutions Portfolio WORKFLOW SOLUTIONS FROM EXTRACTION TO ANALYSIS Application-based answers for every step of your workflow Scientists
GENOMICS WORKFLOW SOLUTIONS THAT GO WHERE THE SCIENCE LEADS Genomics Solutions Portfolio WORKFLOW SOLUTIONS FROM EXTRACTION TO ANALYSIS Application-based answers for every step of your workflow Scientists
Ultrasequencing: Methods and Applications of the New Generation Sequencing Platforms
 Ultrasequencing: Methods and Applications of the New Generation Sequencing Platforms Laura Moya Andérico Master in Advanced Genetics Genomics Class December 16 th, 2015 Brief Overview First-generation
Ultrasequencing: Methods and Applications of the New Generation Sequencing Platforms Laura Moya Andérico Master in Advanced Genetics Genomics Class December 16 th, 2015 Brief Overview First-generation
Outline. General principles of clonal sequencing Analysis principles Applications CNV analysis Genome architecture
 The use of new sequencing technologies for genome analysis Chris Mattocks National Genetics Reference Laboratory (Wessex) NGRL (Wessex) 2008 Outline General principles of clonal sequencing Analysis principles
The use of new sequencing technologies for genome analysis Chris Mattocks National Genetics Reference Laboratory (Wessex) NGRL (Wessex) 2008 Outline General principles of clonal sequencing Analysis principles
Sequencing techniques and applications
 I519 Introduction to Bioinformatics Sequencing techniques and applications Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Contents Sequencing techniques Sanger sequencing Next generation
I519 Introduction to Bioinformatics Sequencing techniques and applications Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Contents Sequencing techniques Sanger sequencing Next generation
Sample to Insight. Dr. Bhagyashree S. Birla NGS Field Application Scientist
 Dr. Bhagyashree S. Birla NGS Field Application Scientist bhagyashree.birla@qiagen.com NGS spans a broad range of applications DNA Applications Human ID Liquid biopsy Biomarker discovery Inherited and somatic
Dr. Bhagyashree S. Birla NGS Field Application Scientist bhagyashree.birla@qiagen.com NGS spans a broad range of applications DNA Applications Human ID Liquid biopsy Biomarker discovery Inherited and somatic
The Expanded Illumina Sequencing Portfolio New Sample Prep Solutions and Workflow
 The Expanded Illumina Sequencing Portfolio New Sample Prep Solutions and Workflow Marcus Hausch, Ph.D. 2010 Illumina, Inc. All rights reserved. Illumina, illuminadx, Solexa, Making Sense Out of Life, Oligator,
The Expanded Illumina Sequencing Portfolio New Sample Prep Solutions and Workflow Marcus Hausch, Ph.D. 2010 Illumina, Inc. All rights reserved. Illumina, illuminadx, Solexa, Making Sense Out of Life, Oligator,
Next Generation Sequencing. Simon Rasmussen Assistant Professor Center for Biological Sequence analysis Technical University of Denmark
 Next eneration Sequencing Simon Rasmussen Assistant Professor enter for Biological Sequence analysis Technical University of Denmark DNA Sequencing DNA sequencing Reading the order of bases in DNA fragments
Next eneration Sequencing Simon Rasmussen Assistant Professor enter for Biological Sequence analysis Technical University of Denmark DNA Sequencing DNA sequencing Reading the order of bases in DNA fragments
SEQUENCING FROM SAMPLE TO SEQUENCE READY
 SEQUENCING FROM SAMPLE TO SEQUENCE READY ACCESS ARRAY SYSTEM HIGH-QUALITY LIBRARIES NOT ONCE, BUT EVERY TIME n The highest quality amplicons more sensitive, accurate, and specific n Full support for all
SEQUENCING FROM SAMPLE TO SEQUENCE READY ACCESS ARRAY SYSTEM HIGH-QUALITY LIBRARIES NOT ONCE, BUT EVERY TIME n The highest quality amplicons more sensitive, accurate, and specific n Full support for all
RIPTIDE HIGH THROUGHPUT RAPID LIBRARY PREP (HT-RLP)
 Application Note: RIPTIDE HIGH THROUGHPUT RAPID LIBRARY PREP (HT-RLP) Introduction: Innovations in DNA sequencing during the 21st century have revolutionized our ability to obtain nucleotide information
Application Note: RIPTIDE HIGH THROUGHPUT RAPID LIBRARY PREP (HT-RLP) Introduction: Innovations in DNA sequencing during the 21st century have revolutionized our ability to obtain nucleotide information
Axygen AxyPrep Magnetic Bead Purification Kits. A Corning Brand
 Axygen AxyPrep Magnetic Bead Purification Kits A Corning Brand D Sample Prep Solutions for Genomics Obtaining Pure Nucleic Acids from Your Sample is Precious The purification of high quality DNA is the
Axygen AxyPrep Magnetic Bead Purification Kits A Corning Brand D Sample Prep Solutions for Genomics Obtaining Pure Nucleic Acids from Your Sample is Precious The purification of high quality DNA is the
Integrated NGS Sample Preparation Solutions for Limiting Amounts of RNA and DNA. March 2, Steven R. Kain, Ph.D. ABRF 2013
 Integrated NGS Sample Preparation Solutions for Limiting Amounts of RNA and DNA March 2, 2013 Steven R. Kain, Ph.D. ABRF 2013 NuGEN s Core Technologies Selective Sequence Priming Nucleic Acid Amplification
Integrated NGS Sample Preparation Solutions for Limiting Amounts of RNA and DNA March 2, 2013 Steven R. Kain, Ph.D. ABRF 2013 NuGEN s Core Technologies Selective Sequence Priming Nucleic Acid Amplification
Genetics and Genomics in Medicine Chapter 3. Questions & Answers
 Genetics and Genomics in Medicine Chapter 3 Multiple Choice Questions Questions & Answers Question 3.1 Which of the following statements, if any, is false? a) Amplifying DNA means making many identical
Genetics and Genomics in Medicine Chapter 3 Multiple Choice Questions Questions & Answers Question 3.1 Which of the following statements, if any, is false? a) Amplifying DNA means making many identical
DNA METHYLATION RESEARCH TOOLS
 SeqCap Epi Enrichment System Revolutionize your epigenomic research DNA METHYLATION RESEARCH TOOLS Methylated DNA The SeqCap Epi System is a set of target enrichment tools for DNA methylation assessment
SeqCap Epi Enrichment System Revolutionize your epigenomic research DNA METHYLATION RESEARCH TOOLS Methylated DNA The SeqCap Epi System is a set of target enrichment tools for DNA methylation assessment
Next Generation Sequencing Lecture Saarbrücken, 19. March Sequencing Platforms
 Next Generation Sequencing Lecture Saarbrücken, 19. March 2012 Sequencing Platforms Contents Introduction Sequencing Workflow Platforms Roche 454 ABI SOLiD Illumina Genome Anlayzer / HiSeq Problems Quality
Next Generation Sequencing Lecture Saarbrücken, 19. March 2012 Sequencing Platforms Contents Introduction Sequencing Workflow Platforms Roche 454 ABI SOLiD Illumina Genome Anlayzer / HiSeq Problems Quality
Galaxy Workshop
 Galaxy Workshop 1-8-13 Intros: Tom Bair thomas-bair@uiowa.edu Ann Black-Ziegelbein annblack@eng.uiowa.edu Srinivas Maddhi srinivas-maddhi@uiowa.edu What is galaxy good for Access to resources Documentation
Galaxy Workshop 1-8-13 Intros: Tom Bair thomas-bair@uiowa.edu Ann Black-Ziegelbein annblack@eng.uiowa.edu Srinivas Maddhi srinivas-maddhi@uiowa.edu What is galaxy good for Access to resources Documentation
GENETICS - CLUTCH CH.15 GENOMES AND GENOMICS.
 !! www.clutchprep.com CONCEPT: OVERVIEW OF GENOMICS Genomics is the study of genomes in their entirety Bioinformatics is the analysis of the information content of genomes - Genes, regulatory sequences,
!! www.clutchprep.com CONCEPT: OVERVIEW OF GENOMICS Genomics is the study of genomes in their entirety Bioinformatics is the analysis of the information content of genomes - Genes, regulatory sequences,
DNA Sequencing and Assembly
 DNA Sequencing and Assembly CS 262 Lecture Notes, Winter 2016 February 2nd, 2016 Scribe: Mark Berger Abstract In this lecture, we survey a variety of different sequencing technologies, including their
DNA Sequencing and Assembly CS 262 Lecture Notes, Winter 2016 February 2nd, 2016 Scribe: Mark Berger Abstract In this lecture, we survey a variety of different sequencing technologies, including their
Genome Sequencing. I: Methods. MMG 835, SPRING 2016 Eukaryotic Molecular Genetics. George I. Mias
 Genome Sequencing I: Methods MMG 835, SPRING 2016 Eukaryotic Molecular Genetics George I. Mias Department of Biochemistry and Molecular Biology gmias@msu.edu Sequencing Methods Cost of Sequencing Wetterstrand
Genome Sequencing I: Methods MMG 835, SPRING 2016 Eukaryotic Molecular Genetics George I. Mias Department of Biochemistry and Molecular Biology gmias@msu.edu Sequencing Methods Cost of Sequencing Wetterstrand
Sequencing technologies
 Sequencing technologies part of High-Throughput Analyzes of Genome Sequenzes Computational EvoDevo University of Leipzig Leipzig, WS 2014/15 Sanger Sequencing (Chain Termination Method) Sequencing of one
Sequencing technologies part of High-Throughput Analyzes of Genome Sequenzes Computational EvoDevo University of Leipzig Leipzig, WS 2014/15 Sanger Sequencing (Chain Termination Method) Sequencing of one
GENOMICS WORKFLOW SOLUTIONS THAT GO WHERE THE SCIENCE LEADS. Genomics Solutions Portfolio
 GENOMICS WORKFLOW SOLUTIONS THAT GO WHERE THE SCIENCE LEADS Genomics Solutions Portfolio WORKFLOW SOLUTIONS FROM EXTRACTION TO ANALYSIS Application-based answers for every step of your workflow Scientists
GENOMICS WORKFLOW SOLUTIONS THAT GO WHERE THE SCIENCE LEADS Genomics Solutions Portfolio WORKFLOW SOLUTIONS FROM EXTRACTION TO ANALYSIS Application-based answers for every step of your workflow Scientists
Wheat CAP Gene Expression with RNA-Seq
 Wheat CAP Gene Expression with RNA-Seq July 9 th -13 th, 2018 Overview of the workshop, Alina Akhunova http://www.ksre.k-state.edu/igenomics/workshops/ RNA-Seq Workshop Activities Lectures Laboratory Molecular
Wheat CAP Gene Expression with RNA-Seq July 9 th -13 th, 2018 Overview of the workshop, Alina Akhunova http://www.ksre.k-state.edu/igenomics/workshops/ RNA-Seq Workshop Activities Lectures Laboratory Molecular
Lecture 7. Next-generation sequencing technologies
 Lecture 7 Next-generation sequencing technologies Next-generation sequencing technologies General principles of short-read NGS Construct a library of fragments Generate clonal template populations Massively
Lecture 7 Next-generation sequencing technologies Next-generation sequencing technologies General principles of short-read NGS Construct a library of fragments Generate clonal template populations Massively
Joint RuminOmics/Rumen Microbial Genomics Network Workshop
 Joint RuminOmics/Rumen Microbial Genomics Network Workshop Microbiome analysis - Amplicon sequencing Dr. Sinéad Waters Animal and Bioscience Research Department, Teagasc Grange, Ireland Prof. Leluo Guan
Joint RuminOmics/Rumen Microbial Genomics Network Workshop Microbiome analysis - Amplicon sequencing Dr. Sinéad Waters Animal and Bioscience Research Department, Teagasc Grange, Ireland Prof. Leluo Guan
Surely Better Target Enrichment from Sample to Sequencer
 sureselect TARGET ENRICHMENT solutions Surely Better Target Enrichment from Sample to Sequencer Agilent s market leading SureSelect platform provides a complete portfolio of catalog to custom products,
sureselect TARGET ENRICHMENT solutions Surely Better Target Enrichment from Sample to Sequencer Agilent s market leading SureSelect platform provides a complete portfolio of catalog to custom products,
1. Introduction Gene regulation Genomics and genome analyses
 1. Introduction Gene regulation Genomics and genome analyses 2. Gene regulation tools and methods Regulatory sequences and motif discovery TF binding sites Databases 3. Technologies Microarrays Deep sequencing
1. Introduction Gene regulation Genomics and genome analyses 2. Gene regulation tools and methods Regulatory sequences and motif discovery TF binding sites Databases 3. Technologies Microarrays Deep sequencing
Bio(tech) Interlude. 3 Nobel Prizes: PCR: Kary Mullis, 1993 Electrophoresis: A.W.K. Tiselius, 1948 DNA Sequencing: Frederick Sanger, 1980
 Bio(tech) Interlude 3 Nobel Prizes: PCR: Kary Mullis, 1993 Electrophoresis: A.W.K. Tiselius, 1948 DNA Sequencing: Frederick Sanger, 1980 PCR 1: 25ºC G 2: 95ºC A 3: 60ºC T 5 3 A A G 3 G T C 5 T T T 6: 72ºC
Bio(tech) Interlude 3 Nobel Prizes: PCR: Kary Mullis, 1993 Electrophoresis: A.W.K. Tiselius, 1948 DNA Sequencing: Frederick Sanger, 1980 PCR 1: 25ºC G 2: 95ºC A 3: 60ºC T 5 3 A A G 3 G T C 5 T T T 6: 72ºC
RNA-Seq data analysis course September 7-9, 2015
 RNA-Seq data analysis course September 7-9, 2015 Peter-Bram t Hoen (LUMC) Jan Oosting (LUMC) Celia van Gelder, Jacintha Valk (BioSB) Anita Remmelzwaal (LUMC) Expression profiling DNA mrna protein Comprehensive
RNA-Seq data analysis course September 7-9, 2015 Peter-Bram t Hoen (LUMC) Jan Oosting (LUMC) Celia van Gelder, Jacintha Valk (BioSB) Anita Remmelzwaal (LUMC) Expression profiling DNA mrna protein Comprehensive
Considerations for Illumina library preparation. Henriette O Geen June 20, 2014 UCD Genome Center
 Considerations for Illumina library preparation Henriette O Geen June 20, 2014 UCD Genome Center Diversity of applications De novo genome Sequencing ranscriptome Expression Splice Isoform bundance Genotyping
Considerations for Illumina library preparation Henriette O Geen June 20, 2014 UCD Genome Center Diversity of applications De novo genome Sequencing ranscriptome Expression Splice Isoform bundance Genotyping
resequencing storage SNP ncrna metagenomics private trio de novo exome ncrna RNA DNA bioinformatics RNA-seq comparative genomics
 RNA Sequencing T TM variation genetics validation SNP ncrna metagenomics private trio de novo exome mendelian ChIP-seq RNA DNA bioinformatics custom target high-throughput resequencing storage ncrna comparative
RNA Sequencing T TM variation genetics validation SNP ncrna metagenomics private trio de novo exome mendelian ChIP-seq RNA DNA bioinformatics custom target high-throughput resequencing storage ncrna comparative
The Genome Analysis Centre. Building Excellence in Genomics and Computa5onal Bioscience
 Building Excellence in Genomics and Computa5onal Bioscience Resequencing approaches Sarah Ayling Crop Genomics and Diversity sarah.ayling@tgac.ac.uk Why re- sequence plants? To iden
Building Excellence in Genomics and Computa5onal Bioscience Resequencing approaches Sarah Ayling Crop Genomics and Diversity sarah.ayling@tgac.ac.uk Why re- sequence plants? To iden
 www.illumina.com/hiseq www.illumina.com FOR RESEARCH USE ONLY 2012 2014 Illumina, Inc. All rights reserved. Illumina, BaseSpace, cbot, CSPro, Genetic Energy, HiSeq, Nextera, TruSeq, the pumpkin orange
www.illumina.com/hiseq www.illumina.com FOR RESEARCH USE ONLY 2012 2014 Illumina, Inc. All rights reserved. Illumina, BaseSpace, cbot, CSPro, Genetic Energy, HiSeq, Nextera, TruSeq, the pumpkin orange
CMPS 6630: Introduction to Computational Biology and Bioinformatics. High-Throughput Sequencing and Applications
 CMPS 6630: Introduction to Computational Biology and Bioinformatics High-Throughput Sequencing and Applications Sanger (1982) introduced chaintermination sequencing. Main idea: Obtain fragments of all
CMPS 6630: Introduction to Computational Biology and Bioinformatics High-Throughput Sequencing and Applications Sanger (1982) introduced chaintermination sequencing. Main idea: Obtain fragments of all
Gene Expression Technology
 Gene Expression Technology Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Gene expression Gene expression is the process by which information from a gene
Gene Expression Technology Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Gene expression Gene expression is the process by which information from a gene
Get to Know Your DNA. Every Single Fragment.
 HaloPlex HS NGS Target Enrichment System Get to Know Your DNA. Every Single Fragment. High sensitivity detection of rare variants using molecular barcodes How Does Molecular Barcoding Work? HaloPlex HS
HaloPlex HS NGS Target Enrichment System Get to Know Your DNA. Every Single Fragment. High sensitivity detection of rare variants using molecular barcodes How Does Molecular Barcoding Work? HaloPlex HS
