The Human Protein PRR14 Tethers Heterochromatin to the Nuclear Lamina During Interphase and Mitotic Exit
|
|
|
- Erika Davis
- 6 years ago
- Views:
Transcription
1 Cell Reports, Volume 5 Supplemental Information The Human Protein PRR14 Tethers Heterochromatin to the Nuclear Lamina During Interphase and Mitotic Exit Andrey Poleshko, Katelyn M. Mansfield, Caroline C. Burlingame, Mark D. Andrake, Neil R. Shah, and Richard A. Katz EXTENDED EXPERIMENTAL PROCEDURES Antibodies The following primary antibodies were used in immunostaining: anti-gfp (Abcam ab290), anti-lamina/c (SantaCruz sc-7292), anti-laminb (SantaCruz sc-6216), anti-laminb1 (Abcam ab16048), anti-laminb2 (Abcam ab8983), anti- -Tubulin (Sigma-Aldrich T5168), anti-myc tag (Abcam ab9132), anti-prr (Sigma-Aldrich SAB ), anti-prr (Sigma-Aldrich SAB ), anti-lbr (Abcam ab122919). sirna Sequences PRR14 sirna sequences (Dharmacon, ThermoScientific): #1 5 -CAACAAGAATTACCAATCA, #2 5 - GCTAGAAGATGTCATGGCT, #3 5 -GAGGATAGTTCGCCAGCCA, #4 5 -GGACTGCCTCGACCAATCA. Lamin A/C sirna sequences: 5 -CAGGCAGTCTGCTGAGAGGAA, 5 -CCCACCAAAGTTCACCCTGAA, 5 - AACTGGACTTCCAGAAGAACA, 5 -CCAGGAGCTTCTGGACATCAA. Lamin B1 sirna sequences: 5 - AACGCGCTTGGTAGAGGTGGA, 5 -CAGGGAGAGGAGGTTGCTCAA, 5 -CCCGAGCATCCTCAAGTCGTA, 5 -CTGGCGAAGATGTGAAGGTTA. Lamin B2 sirna sequences: 5 - CAGCGCCAGGTTAACGCTGAA, 5 - TCGGCAATAGCTCACCGTTTA, 5 -CACGCGGCAGTTCTTTGTTAA, 5 -CGCCTACAAGTTCACGCCCAA. LBR sirna sequences: 5 -TAAGGTAGCTTTAGCAATTAA, 5 -TAGCAATTAAATTCTAGTGAA, 5 - AGCGTTGTTGACGACCATGGA, 5 -CAGCGTGTGCCCTACCGTATA, 5 -CAGGCCGACATTAAGGAAGCA. HP1 sirna SMARTpools (Dharmacon, ThermoScientific): CBX1 (HP1beta) M , CBX3 (HP1gamma) M , and CBX5 (HP1alpha) M SiControl, control sirna (QIAGEN): AllStars Negative Control sirna SI Image Analysis and Quantitation Image analysis was performed using Nikon EZ-C1 v3.8 (Nikon Instruments, Inc., Melville, NY) and MetaXpress v.3.1 software (MDS Analytical Technologies, Sunnyvale, CA). For Figures 1C and S4 the line profile tool was used. For Figures 2B and 2C, nuclear pixel-to-pixel channel overlap was quantitated for GFP-PRR14 (green) and mrfp-hp1a (red) (panel 2B), and PRR14 N-GFP (green) and anti-h3k9me3 (red) (panel 2C). For these two panels multiple cells were analyzed (n = 15), and the results are presented graphically in the respective panels. For Figure 2D, channel overlap for nuclear PRR14 N-GFP and two substituted forms (green) were each compared to Hoechst (blue). Multiple cells were analyzed (n = 15) for each comparison, and highly significant differences (p < ) between wt and substituted proteins were observed as presented graphically in panel 2D. For Figure 2E, the effect of PRR14 sirna knockdown on peripheral H3K9me3 chromatin was measured by comparing H3K9me3 staining (green) with Lamin A/C staining (red). To quantitate loss of H3K9me3 at the nuclear periphery, a mask of the peripheral nuclear region was first created using the red channel. The fraction of the green signal within the peripheral region could then be compared to the total green signal intensity. Individual cells that had been treated with either control sirna or PRR14 sirna were compared (also see Figure S4). Multiple images were analyzed for each treatment (n = 25), and differences in H3K9me3 distribution between control and PRR14 sirna-treated cells were highly significant (p-value < ) (Figure 2E). For Figure 2F, the effects on PRR14 localization after control, Lamin A/C, Lamin B1 and Lamin B2 sirna knockdown were determined by comparing the PRR14 N-GFP signal (green) and Lamin B signal (red) channels (n = 16 for each treatment). Highly significant differences (p-value < ) between the control and Lamin A/C sirna knockdowns were observed, as presented graphically in Figure 2F. For Figure S2D, channel overlap for nuclear PRR14 N-GFP (green) to Hoechst (blue) was compared in the control and HP1 knockdown samples. Multiple cells were analyzed (n = 20) for each sample, and highly significant differences (p < ) between control and HP1 knockdown sample were
2 detected. For Figure S2E, the effect of LBR, Lamin A/C and PRR14 sirna knockdowns on peripheral H3K9me3 chromatin was measured similarly as described in Figure 2E, using Lamin B signal (red) to create a peripheral region mask. Multiple images were analyzed for each treatment (n = 20), and differences in H3K9me3 distribution between sirna control and sirna target samples were *p = and **p < , respectively as shown on figure S2E. SUPPLEMENTAL MOVIE LEGEND Movie S1. Binding of PRR14 to Mitotic Chromatin at the Onset of Anaphase Left panel, DNA (Hoechst staining). Middle panel, N-GFP-tagged PRR14 protein. Right panel, merge. HeLa cells stably expressing N-GFP-tagged PRR14 protein were synchronized using a double thymidine block. The metaphaseanaphase-telophase transition is shown.
3 Figure S1. Detection of Native PRR14 in HeLa Cells and Role in Nuclear Lamina Structure, Related to Figure 1 (A) HeLa cells were transfected with myc-tagged PRR14, and cells were permeabilized using differential conditions that allow antibodies to penetrate either the plasma membrane only (digitonin), or both the plasma and nuclear membranes (Triton-X 100). PRR14 protein and Lamin A/C were detected only after Triton X-100 treatment, indicating that the tagged PRR14 was concentrated at the inner nuclear periphery. Control experiments confirmed conditional access to cytoplasmic components or the nuclear lamina (data not shown). (B) Native PRR14 was detected using a peptide antibody corresponding to positions Two different secondary antibodies were used to confirm that staining at the nuclear periphery was specific. (C) The efficiencies of two PRR14 antibodies targeting amino acids and were compared using transfected N-terminal GFP-tagged PRR14. Peptide antibody to positions detected transfected PRR14 at the periphery, while the antibody favored detection of nucleoplasmic PRR14. Transfected GFP-tagged PRR14 reacted weakly with both antibodies. For example, a cell with moderate levels of GFPtagged PRR14 expression shows no significant staining with the anti-prr14 antibody (arrow). We conclude that the inability to detect native PRR14 with the antibody is due to weak reactivity. We also conclude that the epitope for the antibody can be blocked in some cells by nuclear lamina association. (D) GFP reactivation assay is shown (Poleshko et al., 2010). Treatment with PRR14 sirnas #1 and #4 resulted in reactivation of an epigenetically silent GFP reporter gene. (E) Treatment with independent PRR14 sirnas (#1 and #4), as well as a PRR14 sirna mix caused detachment of the nucleus from the actin cytoskeleton in HeLa cells, as well as binucleation. SiRNA #3 produced a weaker, but reproducible effect. No significant effect on microtubules was detected. sicontrol, control non-targeting sirna. (F) Cells were treated with control or the PRR14 sirna mix, and stained for actin and α-tubulin simultaneously.
4 Figure S2. Mapping of PRR14 Domains, Subcellular Localization, and Effects of PRR14 Knockdown on H3K9me3 Heterochromatin, Related to Figure 2 (A) PRR14 map is shown (yellow, proline rich region; light blue, conserved region; red, HP1-binding motif; dark blue, NLSs). Substitutions and small deletions are indicated with red arrow heads. (B) Confocal imaging of substituted and truncated GFP-tagged PRR14 proteins. Representative images are shown based on > 15 examples. PSORT analysis detected three potential NLSs at positions in the N-terminal fragment, in the central region (not shown), and in the C-terminal fragment. Consistent with the NLS predictions, N-terminal (positions 1-135) and C-terminal fragments (positions ) both localize to the nucleus. Site-specific mutagenesis of the candidate signals at positions and resulted in nuclear import defects of the respective fragments. In the context of the full length N-terminal GFP fusion, disruption of the candidate N-terminal NLS caused a partial nuclear import defect, while deletion of the C-terminal NLS had no significant effect. The N-terminal NLS substitution, a large fraction of cells showed cytoplasmic accumulation, while some cells showed nuclear accumulation consistent with nuclear entry of the NLS substituted protein
5 during mitosis. The and fragments both showed nuclear lamina localization as well as internal tube-like structures similar to the wild type protein. (C) Quantitation of total nuclear signal intensity of GFP-tagged PRR14 after knockdown of Lamin A/C using MetaXpress software. Statistical analysis indicates that Lamin A/C knockdown did not cause significant changes in total signal intensity, N.S., not significant (n = 15 for each sample). (D) HeLa cells stably expressing the GFP-tagged PRR fragment were transfected with sirnas targeting all three HP1 isoforms. Confocal imaging of live cells is shown. Dense chromatin regions were detected with Hoechst (blue). The results are expressed as the fraction of colocalizing green and blue signals in individual cells (n = 20). ** indicates a p-value < for differences between control and HP1 sirna knockdown cells. (E) Measurement of disassociation of H3K9me3-marked chromatin from nuclear perphiphery after sirna-based knockdown of LBR, Lamin A/C, LBR plus Lamin A/C or PRR14. Experimental design and statistical analyses are as described in Figure 2E. Results of statistical analyses are indicated: N.S., not significant; *p = 0.018, **p < (n = 20 for each sample).
6 Figure S3. Confocal Imaging of GFP-Tagged PRR14 Proteins during Mitosis, Related to Figure 3 (A) Time lapse confocal imaging (thirty second intervals) of HeLa cells stably expressing N-terminal GFP-tagged PRR14. The metaphase to anaphase to telophase transitions are shown. GFP (top, green); DNA visualized with Hoechst (middle, blue). Merged images of the first 4 minutes of anaphase are shown (bottom). (B) Mitotic behavior of C-terminal GFP-tagged PRR14, stably expressed in HeLa cells. The following abbreviations are used to indicate stages of mitosis: interphase, I; prophase, P; prometaphase, PM; metaphase, M; anaphase, A; early telophase, telophase, and late telophase, ET, T, LT; daughter cells, D. (C) Mitotic behavior of N-terminal GFP-tagged PRR14, stably expressed in RPE1 cells. Methods and stage designations are as in panel B. The behavior of PRR14 is similar to that observed with N-terminal GFP-tagged PRR14 in HeLa cells (Figure 3A). In these experiments (panel B and C), the GFP signal was enhanced with anti-gfp antibody.
7 Figure S4. Image Analysis of Detachment of H3K9me3-Marked Chromatin from the Nuclear Lamina after PRR14 sirna Knockdown, Related to Figure 2E (A) Shown are representative images of the experiment described in Figure 2E. HeLa cells were transfected with PRR14 sirna and control sirnas, and after 72 hrs, co-staining with anti-h3k9me3 (green) and Lamin A/C (red) antibodies was carried out. Confocal images are shown along with a merged image of the red and green channels. (B) Line profile scans of signal intensity in the regions indicated in panel A were analyzed using Nikon EZ-C1 FreeViewer v3.3 software. Superscripts a and b correspond to scans generated from the indicate lines in panel A. Scans are consistent with detachment of H3K9me3 chromatin from the nuclear lamina after PRR14 knockdown. (C) Three dimensional image profiles were generated using MetaXpress software. Superscript c corresponds to images in panel A. The profiles are consistent with detachment of H3K9me3 marked chromatin from the nuclear periphery after PRR14 knockdown. (D) Quantitation of total nuclear signal intensity of H3K9me3 after knockdown of PRR14 using MetaXpress software. Statistical analysis indicates that PRR14 knockdown did not cause significant changes in total signal intensity (N.S., not significant). (E) Western blot analysis of PRR14 sirna knockdown is shown. No major changes in the total level of H3K9me3 histone mark were observed.
8 Figure S5. Proposed Model for PRR14 Function in Tethering, and Mitotic Specification of HP1-H3K9me2/3-Marked Heterochromatin for Attachment at the Nuclear Lamina, Related to Figures 2 and 3 Color coding of PRR14 regions is as described in Figure 2A. The model first depicts PRR14 tethering HP1-H3K9me3-marked heterochromatin to the nuclear lamina in interphase (left). The association of PRR14 with the nuclear lamina may be direct or indirect. In prophase and metaphase, the tether components disassemble to allow chromosome condensation. In anaphase, HP1 and PRR14 begin to rebind to chromosomes through the H3K9me2/3 mark, which is retained throughout mitosis. In mid- to late-telophase the pre-assembled PRR14-HP1-H3K9me3 heterochromatin complex reassociates with the formed nuclear lamina (arrow), thus reestablishing the tethered state.
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION Dynamic Phosphorylation of HP1 Regulates Mitotic Progression in Human Cells Supplementary Figures Supplementary Figure 1. NDR1 interacts with HP1. (a) Immunoprecipitation using
SUPPLEMENTARY INFORMATION Dynamic Phosphorylation of HP1 Regulates Mitotic Progression in Human Cells Supplementary Figures Supplementary Figure 1. NDR1 interacts with HP1. (a) Immunoprecipitation using
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION Legends for Supplementary Tables. Supplementary Table 1. An excel file containing primary screen data. Worksheet 1, Normalized quantification data from a duplicated screen: valid
SUPPLEMENTARY INFORMATION Legends for Supplementary Tables. Supplementary Table 1. An excel file containing primary screen data. Worksheet 1, Normalized quantification data from a duplicated screen: valid
Description: Nuclear morphology and dynamics in nontargeting sirna transfected cells. HeLa Kyoto
 Title of file for HTML: Supplementary Information Description: Supplementary Figures and Supplementary Tables Title of file for HTML: Supplementary Movie 1 Description: Nuclear morphology and dynamics
Title of file for HTML: Supplementary Information Description: Supplementary Figures and Supplementary Tables Title of file for HTML: Supplementary Movie 1 Description: Nuclear morphology and dynamics
Chapter 3. DNA Replication & The Cell Cycle
 Chapter 3 DNA Replication & The Cell Cycle DNA Replication and the Cell Cycle Before cells divide, they must duplicate their DNA // the genetic material DNA is organized into strands called chromosomes
Chapter 3 DNA Replication & The Cell Cycle DNA Replication and the Cell Cycle Before cells divide, they must duplicate their DNA // the genetic material DNA is organized into strands called chromosomes
Supplementary Table 1. The Q-PCR primer sequence is summarized in the following table.
 Supplementary Table 1. The Q-PCR primer sequence is summarized in the following table. Name Sequence (5-3 ) Application Flag-u ggactacaaggacgacgatgac Shared upstream primer for all the amplifications of
Supplementary Table 1. The Q-PCR primer sequence is summarized in the following table. Name Sequence (5-3 ) Application Flag-u ggactacaaggacgacgatgac Shared upstream primer for all the amplifications of
Cell Division. Use Target Reading Skills. This section explains how cells grow and divide.
 Name Date Class Cell Processes Guided Reading and Study Cell Division This section explains how cells grow and divide. Use Target Reading Skills As you read, make a cycle diagram that shows the events
Name Date Class Cell Processes Guided Reading and Study Cell Division This section explains how cells grow and divide. Use Target Reading Skills As you read, make a cycle diagram that shows the events
Section 10. Junaid Malek, M.D.
 Section 10 Junaid Malek, M.D. Cell Division Make sure you understand: How do cells know when to divide? (What drives the cell cycle? Why is it important to regulate this?) How is DNA replication regulated?
Section 10 Junaid Malek, M.D. Cell Division Make sure you understand: How do cells know when to divide? (What drives the cell cycle? Why is it important to regulate this?) How is DNA replication regulated?
Figure S1. Figure S2. Figure S3 HB Anti-FSP27 (COOH-terminal peptide) Ab. Anti-GST-FSP27(45-127) Ab.
 / 36B4 mrna ratio Figure S1 * 2. 1.6 1.2.8 *.4 control TNFα BRL49653 Figure S2 Su bw AT p iw Anti- (COOH-terminal peptide) Ab Blot : Anti-GST-(45-127) Ab β-actin Figure S3 HB2 HW AT BA T Figure S4 A TAG
/ 36B4 mrna ratio Figure S1 * 2. 1.6 1.2.8 *.4 control TNFα BRL49653 Figure S2 Su bw AT p iw Anti- (COOH-terminal peptide) Ab Blot : Anti-GST-(45-127) Ab β-actin Figure S3 HB2 HW AT BA T Figure S4 A TAG
Supporting Information
 Supporting Information Shao et al. 10.1073/pnas.1504837112 SI Materials and Methods Immunofluorescence and Immunoblotting. For immunofluorescence, cells were fixed with 4% paraformaldehyde and permeabilized
Supporting Information Shao et al. 10.1073/pnas.1504837112 SI Materials and Methods Immunofluorescence and Immunoblotting. For immunofluorescence, cells were fixed with 4% paraformaldehyde and permeabilized
7.17: Writing Up Results and Creating Illustrations
 7.17: Writing Up Results and Creating Illustrations A Results Exercise: Kansas and Pancakes Write a 5-sentence paragraph describing the results illustrated in this figure: - Describe the figure: highlights?
7.17: Writing Up Results and Creating Illustrations A Results Exercise: Kansas and Pancakes Write a 5-sentence paragraph describing the results illustrated in this figure: - Describe the figure: highlights?
Cells and Tissues. Overview CELLS
 Cells and Tissues WIll The basic unit of structure and function in the human body is the cell. Each of a cell's parts, or organelles, as well as the entire cell, is organized to perform a specific function.
Cells and Tissues WIll The basic unit of structure and function in the human body is the cell. Each of a cell's parts, or organelles, as well as the entire cell, is organized to perform a specific function.
Immunofluorescence images of different core histones and different histone exchange assay.
 Molecular Cell, Volume 51 Supplemental Information Enhanced Chromatin Dynamics by FACT Promotes Transcriptional Restart after UV-Induced DNA Damage Christoffel Dinant, Giannis Ampatziadis-Michailidis,
Molecular Cell, Volume 51 Supplemental Information Enhanced Chromatin Dynamics by FACT Promotes Transcriptional Restart after UV-Induced DNA Damage Christoffel Dinant, Giannis Ampatziadis-Michailidis,
DNA STRUCTURE. Nucleotides: Nitrogenous Bases (Carry the Genetic Code) Expectation Sheet: DNA & Cell Cycle. I can statements: Basic Information:
 Expectation Sheet: DNA & Cell Cycle NAME: Test is 11/8/17 I can statements: I can discuss how DNA is found in all organisms and that the structure is common to all living things. I can diagram and label
Expectation Sheet: DNA & Cell Cycle NAME: Test is 11/8/17 I can statements: I can discuss how DNA is found in all organisms and that the structure is common to all living things. I can diagram and label
OCTOBER 22, 2010 VOLUME 285 NUMBER 43 JOURNAL OF BIOLOGICAL CHEMISTRY 33281
 THE JOURNAL OF BIOLOGICAL CHEMISTRY VOL. 285, NO. 43, pp. 33281 33293, October 22, 2010 2010 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in the U.S.A. Requirement for Lamin
THE JOURNAL OF BIOLOGICAL CHEMISTRY VOL. 285, NO. 43, pp. 33281 33293, October 22, 2010 2010 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in the U.S.A. Requirement for Lamin
7.06 Cell Biology EXAM #2 March 20, 2003
 7.06 Cell Biology EXAM #2 March 20, 2003 This is an open book exam, and you are allowed access to books, a calculator, and notes but not computers or any other types of electronic devices. Please write
7.06 Cell Biology EXAM #2 March 20, 2003 This is an open book exam, and you are allowed access to books, a calculator, and notes but not computers or any other types of electronic devices. Please write
NUCLEUS. Fig. 2. Various stages in the condensation of chromatin
 NUCLEUS Animal cells contain DNA in nucleus (contains ~ 98% of cell DNA) and mitochondrion. Both compartments are surrounded by an envelope (double membrane). Nuclear DNA represents some linear molecules
NUCLEUS Animal cells contain DNA in nucleus (contains ~ 98% of cell DNA) and mitochondrion. Both compartments are surrounded by an envelope (double membrane). Nuclear DNA represents some linear molecules
Supplementary Figures and Legends
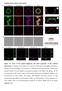 Supplementary Figures and Legends Figure S1. Tests of the optical alignment and focal properties of the confocal microscope. (A) Images of the optical cross-section of fluorescent microspheres differing
Supplementary Figures and Legends Figure S1. Tests of the optical alignment and focal properties of the confocal microscope. (A) Images of the optical cross-section of fluorescent microspheres differing
Genetics and Heredity. Mr. Gagnon
 Genetics and Heredity Mr. Gagnon Key Terms: Traits Heredity Genetics Purebred Genes Alleles Recessive Allele Dominant Allele Hybrids Key Concepts: What factors control the inheritance of traits in organisms?
Genetics and Heredity Mr. Gagnon Key Terms: Traits Heredity Genetics Purebred Genes Alleles Recessive Allele Dominant Allele Hybrids Key Concepts: What factors control the inheritance of traits in organisms?
Biology Lecture 2 Genes
 Genes Definitions o Gene: DNA that codes for a single polypeptide/mrna/rrna/trna o Euchromatin: region of DNA containing genes being actively transcribed o Heterochromatin: region of DNA containing genes
Genes Definitions o Gene: DNA that codes for a single polypeptide/mrna/rrna/trna o Euchromatin: region of DNA containing genes being actively transcribed o Heterochromatin: region of DNA containing genes
Chapter 13. The Nucleus. The nucleus is the hallmark of eukaryotic cells; the very term eukaryotic means having a "true nucleus".
 Chapter 13 The Nucleus The nucleus is the hallmark of eukaryotic cells; the very term eukaryotic means having a "true nucleus". Fig.13.1. The EM of the Nucleus of a Eukaryotic Cell 13.1. The Nuclear Envelope
Chapter 13 The Nucleus The nucleus is the hallmark of eukaryotic cells; the very term eukaryotic means having a "true nucleus". Fig.13.1. The EM of the Nucleus of a Eukaryotic Cell 13.1. The Nuclear Envelope
Chapter 5 DNA and Chromosomes
 Chapter 5 DNA and Chromosomes DNA as the genetic material Heat-killed bacteria can transform living cells S Smooth R Rough Fred Griffith, 1920 DNA is the genetic material Oswald Avery Colin MacLeod Maclyn
Chapter 5 DNA and Chromosomes DNA as the genetic material Heat-killed bacteria can transform living cells S Smooth R Rough Fred Griffith, 1920 DNA is the genetic material Oswald Avery Colin MacLeod Maclyn
Robust Spindle Alignment in Drosophila Neuroblasts by Ultrasensitive Activation of Pins
 Molecular Cell, Volume 43 Supplemental Information Robust Spindle Alignment in Drosophila Neuroblasts by Ultrasensitive Activation of Pins Nicholas R. Smith and Kenneth E. Prehoda Inventory of Supplemental
Molecular Cell, Volume 43 Supplemental Information Robust Spindle Alignment in Drosophila Neuroblasts by Ultrasensitive Activation of Pins Nicholas R. Smith and Kenneth E. Prehoda Inventory of Supplemental
CELLULAR PROCESSES; REPRODUCTION. Unit 5
 CELLULAR PROCESSES; REPRODUCTION Unit 5 Cell Cycle Chromosomes and their make up Crossover Cytokines Diploid (haploid diploid and karyotypes) Mitosis Meiosis What is Cancer? Somatic Cells THE CELL CYCLE
CELLULAR PROCESSES; REPRODUCTION Unit 5 Cell Cycle Chromosomes and their make up Crossover Cytokines Diploid (haploid diploid and karyotypes) Mitosis Meiosis What is Cancer? Somatic Cells THE CELL CYCLE
Aurora Kinase-A Inactivates DNA Damage-Induced
 Cancer Cell, Volume 21 Supplemental Information Aurora Kinase-A Inactivates DNA Damage-Induced Apoptosis and Spindle Assembly Checkpoint Response Functions of p73 Hiroshi Katayama, Jin Wang, Warapen Treekitkarnmongkol,
Cancer Cell, Volume 21 Supplemental Information Aurora Kinase-A Inactivates DNA Damage-Induced Apoptosis and Spindle Assembly Checkpoint Response Functions of p73 Hiroshi Katayama, Jin Wang, Warapen Treekitkarnmongkol,
The Conserved Isoleucine Valine Phenylalanine Motif Couples Activation State and Endocytic Functions of b-arrestins
 Traffic 2007 Blackwell Munksgaard # 2007 The Authors doi: 10.1111/j.1600-0854.2007.00578.x The Conserved Isoleucine Valine Phenylalanine Motif Couples Activation State and Endocytic Functions of b-arrestins
Traffic 2007 Blackwell Munksgaard # 2007 The Authors doi: 10.1111/j.1600-0854.2007.00578.x The Conserved Isoleucine Valine Phenylalanine Motif Couples Activation State and Endocytic Functions of b-arrestins
Kostandin V. Pajcini, Stephane Y. Corbel, Julien Sage, Jason H. Pomerantz, and Helen M. Blau
 Cell Stem Cell, Volume 7 Supplemental Information Transient Inactivation of Rb and ARF Yields Regenerative Cells from Postmitotic Mammalian Muscle Kostandin V. Pajcini, Stephane Y. Corbel, Julien Sage,
Cell Stem Cell, Volume 7 Supplemental Information Transient Inactivation of Rb and ARF Yields Regenerative Cells from Postmitotic Mammalian Muscle Kostandin V. Pajcini, Stephane Y. Corbel, Julien Sage,
Histone Analysis. Histone Analysis
 Histone Analysis Histone Analysis innovative tools designed to help unravel the histone code Histone Antibodies Histone Purification Recombinant Histones Histone Modifying Enzymes Histone Modification
Histone Analysis Histone Analysis innovative tools designed to help unravel the histone code Histone Antibodies Histone Purification Recombinant Histones Histone Modifying Enzymes Histone Modification
Supplemental Movie Legend.
 Supplemental Movie Legend. Transfected T cells were dropped onto SEE superantigen-pulsed Raji B cells (approximate location indicated by circle). Maximum-intensity projections from Z-stacks (17 slices,
Supplemental Movie Legend. Transfected T cells were dropped onto SEE superantigen-pulsed Raji B cells (approximate location indicated by circle). Maximum-intensity projections from Z-stacks (17 slices,
Chromosome territory relocation during DNA repair requires nuclear myosin 1 recruitment to chromatin mediated by ϒ-H2AX signaling
 7 9 Nucleic Acids Research,, Vol., No. 7 ublished online June doi:.9/nar/gkw7 hromosome territory relocation during DNA repair requires nuclear myosin recruitment to chromatin mediated by ϒ-HAX signaling
7 9 Nucleic Acids Research,, Vol., No. 7 ublished online June doi:.9/nar/gkw7 hromosome territory relocation during DNA repair requires nuclear myosin recruitment to chromatin mediated by ϒ-HAX signaling
6.2 Chromatin is divided into euchromatin and heterochromatin
 6.2 Chromatin is divided into euchromatin and heterochromatin Individual chromosomes can be seen only during mitosis. During interphase, the general mass of chromatin is in the form of euchromatin. Euchromatin
6.2 Chromatin is divided into euchromatin and heterochromatin Individual chromosomes can be seen only during mitosis. During interphase, the general mass of chromatin is in the form of euchromatin. Euchromatin
Xfect Protein Transfection Reagent
 Xfect Protein Transfection Reagent Mammalian Expression Systems Rapid, high-efficiency, low-toxicity protein transfection Transfect a large amount of active protein Virtually no cytotoxicity, unlike lipofection
Xfect Protein Transfection Reagent Mammalian Expression Systems Rapid, high-efficiency, low-toxicity protein transfection Transfect a large amount of active protein Virtually no cytotoxicity, unlike lipofection
sirna Transfection Into Primary Neurons Using Fuse-It-siRNA
 sirna Transfection Into Primary Neurons Using Fuse-It-siRNA This Application Note describes a protocol for sirna transfection into sensitive, primary cortical neurons using Fuse-It-siRNA. This innovative
sirna Transfection Into Primary Neurons Using Fuse-It-siRNA This Application Note describes a protocol for sirna transfection into sensitive, primary cortical neurons using Fuse-It-siRNA. This innovative
EUKARYOTIC REGULATION C H A P T E R 1 3
 EUKARYOTIC REGULATION C H A P T E R 1 3 EUKARYOTIC REGULATION Every cell in an organism contains a complete set of DNA. But it doesn t use all of the DNA it receives Each cell chooses different DNA sequences
EUKARYOTIC REGULATION C H A P T E R 1 3 EUKARYOTIC REGULATION Every cell in an organism contains a complete set of DNA. But it doesn t use all of the DNA it receives Each cell chooses different DNA sequences
PGAM5 tethers a ternary complex containing Keap1 and Nrf2 to mitochondria
 available at www.sciencedirect.com www.elsevier.com/locate/yexcr Research Article PGAM5 tethers a ternary complex containing Keap1 and Nrf2 to mitochondria Shih-Ching Lo, Mark Hannink Department of Biochemistry,
available at www.sciencedirect.com www.elsevier.com/locate/yexcr Research Article PGAM5 tethers a ternary complex containing Keap1 and Nrf2 to mitochondria Shih-Ching Lo, Mark Hannink Department of Biochemistry,
Chromatin. Structure and modification of chromatin. Chromatin domains
 Chromatin Structure and modification of chromatin Chromatin domains 2 DNA consensus 5 3 3 DNA DNA 4 RNA 5 ss RNA forms secondary structures with ds hairpins ds forms 6 of nucleic acids Form coiling bp/turn
Chromatin Structure and modification of chromatin Chromatin domains 2 DNA consensus 5 3 3 DNA DNA 4 RNA 5 ss RNA forms secondary structures with ds hairpins ds forms 6 of nucleic acids Form coiling bp/turn
Thyroid peroxidase gene expression is induced by lipopolysaccharide involving Nuclear Factor (NF)-κB p65 subunit phosphorylation
 1 2 3 4 5 SUPPLEMENTAL DATA Thyroid peroxidase gene expression is induced by lipopolysaccharide involving Nuclear Factor (NF)-κB p65 subunit phosphorylation Magalí Nazar, Juan Pablo Nicola, María Laura
1 2 3 4 5 SUPPLEMENTAL DATA Thyroid peroxidase gene expression is induced by lipopolysaccharide involving Nuclear Factor (NF)-κB p65 subunit phosphorylation Magalí Nazar, Juan Pablo Nicola, María Laura
Molecular Biology of the Gene
 Molecular Biology of the Gene : where the genetic information is stored, blueprint for making proteins. RNA: Always involved in protein synthesis Macromolecules (polymers!) Monomers (units): nucleotides
Molecular Biology of the Gene : where the genetic information is stored, blueprint for making proteins. RNA: Always involved in protein synthesis Macromolecules (polymers!) Monomers (units): nucleotides
Learning Objectives. Define RNA interference. Define basic terminology. Describe molecular mechanism. Define VSP and relevance
 Learning Objectives Define RNA interference Define basic terminology Describe molecular mechanism Define VSP and relevance Describe role of RNAi in antigenic variation A Nobel Way to Regulate Gene Expression
Learning Objectives Define RNA interference Define basic terminology Describe molecular mechanism Define VSP and relevance Describe role of RNAi in antigenic variation A Nobel Way to Regulate Gene Expression
immuno concepts Immuno Concepts is proud to announce that when present the characteristic SS-A/Ro pattern detected by the HEp-2000 substrate
 immuno concepts Immuno Concepts is proud to announce that when present the characteristic SS-A/Ro pattern detected by the HEp-2000 substrate is now considered confirmatory for the presence of antibodies
immuno concepts Immuno Concepts is proud to announce that when present the characteristic SS-A/Ro pattern detected by the HEp-2000 substrate is now considered confirmatory for the presence of antibodies
Confocal immunofluorescence microscopy
 Confocal immunofluorescence microscopy HL-6 and cells were cultured and cytospun onto glass slides. The cells were double immunofluorescence stained for Mt NPM1 and fibrillarin (nucleolar marker). Briefly,
Confocal immunofluorescence microscopy HL-6 and cells were cultured and cytospun onto glass slides. The cells were double immunofluorescence stained for Mt NPM1 and fibrillarin (nucleolar marker). Briefly,
Lecture 21: Epigenetics Nurture or Nature? Chromatin DNA methylation Histone Code Twin study X-chromosome inactivation Environemnt and epigenetics
 Lecture 21: Epigenetics Nurture or Nature? Chromatin DNA methylation Histone Code Twin study X-chromosome inactivation Environemnt and epigenetics Epigenetics represents the science for the studying heritable
Lecture 21: Epigenetics Nurture or Nature? Chromatin DNA methylation Histone Code Twin study X-chromosome inactivation Environemnt and epigenetics Epigenetics represents the science for the studying heritable
NIH Public Access Author Manuscript Cell Cycle. Author manuscript; available in PMC 2010 March 1.
 NIH Public Access Author Manuscript Published in final edited form as: Cell Cycle. 2009 October 1; 8(19): 3133 3148. Temporal differences in DNA replication during the S phase using single fiber analysis
NIH Public Access Author Manuscript Published in final edited form as: Cell Cycle. 2009 October 1; 8(19): 3133 3148. Temporal differences in DNA replication during the S phase using single fiber analysis
Three major types of cytoskeleton
 The Cytoskeleton Organizes and stabilizes cells Pulls chromosomes apart Drives intracellular traffic Supports plasma membrane and nuclear envelope Enables cellular movement Guides growth of the plant cell
The Cytoskeleton Organizes and stabilizes cells Pulls chromosomes apart Drives intracellular traffic Supports plasma membrane and nuclear envelope Enables cellular movement Guides growth of the plant cell
Tumor Growth Suppression Through the Activation of p21, a Cyclin-Dependent Kinase Inhibitor
 Tumor Growth Suppression Through the Activation of p21, a Cyclin-Dependent Kinase Inhibitor Nicholas Love 11/28/01 A. What is p21? Introduction - p21 is a gene found on chromosome 6 at 6p21.2 - this gene
Tumor Growth Suppression Through the Activation of p21, a Cyclin-Dependent Kinase Inhibitor Nicholas Love 11/28/01 A. What is p21? Introduction - p21 is a gene found on chromosome 6 at 6p21.2 - this gene
mcherry Polyclonal Antibody Catalog Number PA Product data sheet
 Website: thermofisher.com Customer Service (US): 1 800 955 6288 ext. 1 Technical Support (US): 1 800 955 6288 ext. 441 mcherry Polyclonal Antibody Catalog Number PA5-34974 Product data sheet Details Size
Website: thermofisher.com Customer Service (US): 1 800 955 6288 ext. 1 Technical Support (US): 1 800 955 6288 ext. 441 mcherry Polyclonal Antibody Catalog Number PA5-34974 Product data sheet Details Size
RNA oligonucleotides and 2 -O-methylated oligonucleotides were synthesized by. 5 AGACACAAACACCAUUGUCACACUCCACAGC; Rand-2 OMe,
 Materials and methods Oligonucleotides and DNA constructs RNA oligonucleotides and 2 -O-methylated oligonucleotides were synthesized by Dharmacon Inc. (Lafayette, CO). The sequences were: 122-2 OMe, 5
Materials and methods Oligonucleotides and DNA constructs RNA oligonucleotides and 2 -O-methylated oligonucleotides were synthesized by Dharmacon Inc. (Lafayette, CO). The sequences were: 122-2 OMe, 5
Assays for studying mitochondrial health and function
 APPLICATION NOTE Fluorescence labeling and detection Assays for studying mitochondrial health and function Introduction Mitochondria play a critical role in maintaining normal cellular activities. Mitochondria
APPLICATION NOTE Fluorescence labeling and detection Assays for studying mitochondrial health and function Introduction Mitochondria play a critical role in maintaining normal cellular activities. Mitochondria
A Level. A Level Biology. Cells, Microscopes, Cell Cycle and Immunity Questions. AQA, OCR, Edexcel. Name: Total Marks: Page 1
 AQA, OCR, Edexcel A Level A Level Biology Cells, Microscopes, Cell Cycle and Immunity Questions Name: Total Marks: Page 1 Q1.The diagram shows a eukaryotic cell. (a) Complete the table by giving the letter
AQA, OCR, Edexcel A Level A Level Biology Cells, Microscopes, Cell Cycle and Immunity Questions Name: Total Marks: Page 1 Q1.The diagram shows a eukaryotic cell. (a) Complete the table by giving the letter
DNA Sequence-Dependent Compartmentalization and Silencing of Chromatin at the Nuclear Lamina
 DNA Sequence-Dependent Compartmentalization and Silencing of Chromatin at the Nuclear Lamina Joseph M. Zullo, 1,8 Ignacio A. Demarco, 3,8 Roger Piqué-Regi, 2,4 Daniel J. Gaffney, 2,4 Charles B. Epstein,
DNA Sequence-Dependent Compartmentalization and Silencing of Chromatin at the Nuclear Lamina Joseph M. Zullo, 1,8 Ignacio A. Demarco, 3,8 Roger Piqué-Regi, 2,4 Daniel J. Gaffney, 2,4 Charles B. Epstein,
Thursday 26 May 2016 Afternoon Time allowed: 1 hour 30 minutes
 Please write clearly in block capitals. Centre number Candidate number Surname Forename(s) Candidate signature AS BIOLOGY Paper 1 Thursday 26 May 2016 Afternoon Time allowed: 1 hour 30 minutes Materials
Please write clearly in block capitals. Centre number Candidate number Surname Forename(s) Candidate signature AS BIOLOGY Paper 1 Thursday 26 May 2016 Afternoon Time allowed: 1 hour 30 minutes Materials
Chapter 15. Cytoskeletal Systems. Lectures by Kathleen Fitzpatrick Simon Fraser University Pearson Education, Inc.
 Chapter 15 Cytoskeletal Systems Lectures by Kathleen Fitzpatrick Simon Fraser University Table 15-1 - Microtubules Table 15-1 - Microfilaments Table 15-1 Intermediate Filaments Table 15-3 Microtubules
Chapter 15 Cytoskeletal Systems Lectures by Kathleen Fitzpatrick Simon Fraser University Table 15-1 - Microtubules Table 15-1 - Microfilaments Table 15-1 Intermediate Filaments Table 15-3 Microtubules
Alternative Cleavage and Polyadenylation of RNA
 Developmental Cell 18 Supplemental Information The Spen Family Protein FPA Controls Alternative Cleavage and Polyadenylation of RNA Csaba Hornyik, Lionel C. Terzi, and Gordon G. Simpson Figure S1, related
Developmental Cell 18 Supplemental Information The Spen Family Protein FPA Controls Alternative Cleavage and Polyadenylation of RNA Csaba Hornyik, Lionel C. Terzi, and Gordon G. Simpson Figure S1, related
RNA polymerase II precursor mrna (hnrna) introns introns U1RNP. splicing
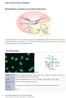 DNA RNA polymerase II precursor mrna (hnrna) introns introns U2RNP splicing mrna U1RNP U5RNP U4/U6RNP splicing The genetic information of DNA is transcribed to precursor mrna (heterogeneous nuclear RNA,
DNA RNA polymerase II precursor mrna (hnrna) introns introns U2RNP splicing mrna U1RNP U5RNP U4/U6RNP splicing The genetic information of DNA is transcribed to precursor mrna (heterogeneous nuclear RNA,
Supporting Information
 Supporting Information Signal mingle: Micropatterns of BMP-2 and fibronectin on soft biopolymeric films regulate myoblast shape and SMAD signaling. Vincent Fitzpatrick, Laure Fourel, Olivier Destaing,
Supporting Information Signal mingle: Micropatterns of BMP-2 and fibronectin on soft biopolymeric films regulate myoblast shape and SMAD signaling. Vincent Fitzpatrick, Laure Fourel, Olivier Destaing,
DNA is a functional genetic material as it:
 DNA DNA is a functional genetic material as it: varies between species and individuals can store information remains constant within a species Replicates undergoes mutations 1 `It has not escaped our notice
DNA DNA is a functional genetic material as it: varies between species and individuals can store information remains constant within a species Replicates undergoes mutations 1 `It has not escaped our notice
Spectral Separation of Multifluorescence Labels with the LSM 510 META
 Microscopy from Carl Zeiss Spectral Separation of Multifluorescence Labels with the LSM 510 META Indians living in the South American rain forest can distinguish between almost 200 hues of green in their
Microscopy from Carl Zeiss Spectral Separation of Multifluorescence Labels with the LSM 510 META Indians living in the South American rain forest can distinguish between almost 200 hues of green in their
Electronic Supplementary Information
 Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2017 Electronic Supplementary Information Dissecting binding of a β-barrel outer membrane
Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2017 Electronic Supplementary Information Dissecting binding of a β-barrel outer membrane
SUPPLEMENTAL MATERIALS
 SUPPLEMENL MERILS Eh-seq: RISPR epitope tagging hip-seq of DN-binding proteins Daniel Savic, E. hristopher Partridge, Kimberly M. Newberry, Sophia. Smith, Sarah K. Meadows, rian S. Roberts, Mark Mackiewicz,
SUPPLEMENL MERILS Eh-seq: RISPR epitope tagging hip-seq of DN-binding proteins Daniel Savic, E. hristopher Partridge, Kimberly M. Newberry, Sophia. Smith, Sarah K. Meadows, rian S. Roberts, Mark Mackiewicz,
Supplementary Information
 Journal : Nature Biotechnology Supplementary Information Targeted genome engineering in human cells with RNA-guided endonucleases Seung Woo Cho, Sojung Kim, Jong Min Kim, and Jin-Soo Kim* National Creative
Journal : Nature Biotechnology Supplementary Information Targeted genome engineering in human cells with RNA-guided endonucleases Seung Woo Cho, Sojung Kim, Jong Min Kim, and Jin-Soo Kim* National Creative
Supplemental Data Length Control of the Metaphase Spindle
 Supplemental Data Length Control of the Metaphase Spindle S1 Gohta Goshima, Roy Wollman, Nico Stuurman, Jonathan M. Scholey, and Ronald D. Vale Supplemental Experimental Procedures Cell Culture, RNA Interference,
Supplemental Data Length Control of the Metaphase Spindle S1 Gohta Goshima, Roy Wollman, Nico Stuurman, Jonathan M. Scholey, and Ronald D. Vale Supplemental Experimental Procedures Cell Culture, RNA Interference,
Small-Molecule Drug Target Identification/Deconvolution Technologies
 Small-Molecule Drug Target Identification/Deconvolution Technologies Case-Studies Shantani Target ID Technology Tool Box Target Deconvolution is not Trivial = A single Tool / Technology May Not necessarily
Small-Molecule Drug Target Identification/Deconvolution Technologies Case-Studies Shantani Target ID Technology Tool Box Target Deconvolution is not Trivial = A single Tool / Technology May Not necessarily
Supplementary material to Alterations in the properties of the cell membrane due to glycosphingolipid accumulation in a model of Gaucher disease
 Supplementary material to Alterations in the properties of the cell membrane due to glycosphingolipid accumulation in a model of Gaucher disease Gyula Batta, Lilla Soltész, Tamás Kovács, Tamás Bozó, Zoltán
Supplementary material to Alterations in the properties of the cell membrane due to glycosphingolipid accumulation in a model of Gaucher disease Gyula Batta, Lilla Soltész, Tamás Kovács, Tamás Bozó, Zoltán
10-2 Cell Division (Pages )
 10-2 Cell Division (Pages 244-245) What do you think would happen if a cell were simply to split into two, without any advance preparation? Would each daughter cell have everything it needed to survive?
10-2 Cell Division (Pages 244-245) What do you think would happen if a cell were simply to split into two, without any advance preparation? Would each daughter cell have everything it needed to survive?
GM130 Is Required for Compartmental Organization of Dendritic Golgi Outposts
 Current Biology, Volume 24 Supplemental Information GM130 Is Required for Compartmental Organization of Dendritic Golgi Outposts Wei Zhou, Jin Chang, Xin Wang, Masha G. Savelieff, Yinyin Zhao, Shanshan
Current Biology, Volume 24 Supplemental Information GM130 Is Required for Compartmental Organization of Dendritic Golgi Outposts Wei Zhou, Jin Chang, Xin Wang, Masha G. Savelieff, Yinyin Zhao, Shanshan
Purification of Lactate Dehydrogenase
 Dominican University of California Dominican Scholar Scholarly & Creative Works Conference 2018 Scholarly and Creative Works Conference 2016 Apr 15th, 1:30 PM - 2:00 PM Purification of Lactate Dehydrogenase
Dominican University of California Dominican Scholar Scholarly & Creative Works Conference 2018 Scholarly and Creative Works Conference 2016 Apr 15th, 1:30 PM - 2:00 PM Purification of Lactate Dehydrogenase
TECHNICAL BULLETIN. U2OS GFP-NUP98 Osteosarcoma Cell Line with 3XFLAG -GFPtagged. Catalog Number CLL1136 Storage Temperature 196 C (liquid nitrogen)
 U2OS GFP-NUP98 Osteosarcoma Cell Line with 3XFLAG -GFPtagged NUP98 Catalog Number CLL1136 Storage Temperature 196 C (liquid nitrogen) TECHNICAL BULLETIN Product Description This product is a human U2OS
U2OS GFP-NUP98 Osteosarcoma Cell Line with 3XFLAG -GFPtagged NUP98 Catalog Number CLL1136 Storage Temperature 196 C (liquid nitrogen) TECHNICAL BULLETIN Product Description This product is a human U2OS
PARP-1 (cleaved) Human In-Cell ELISA Kit (IR)
 ab110215 PARP-1 (cleaved) Human In-Cell ELISA Kit (IR) Instructions for Use For the quantitative measurement of Human PARP-1 (cleaved) concentrations in cultured adherent and suspension cells. This product
ab110215 PARP-1 (cleaved) Human In-Cell ELISA Kit (IR) Instructions for Use For the quantitative measurement of Human PARP-1 (cleaved) concentrations in cultured adherent and suspension cells. This product
Mammalian Orc1 Protein Is Selectively Released from Chromatin and Ubiquitinated during the S-to-M Transition in the Cell Division Cycle
 MOLECULAR AND CELLULAR BIOLOGY, Jan. 2002, p. 105 116 Vol. 22, No. 1 0270-7306/02/$04.00 0 DOI: 10.1128/MCB.22.1.105 116.2002 Mammalian Orc1 Protein Is Selectively Released from Chromatin and Ubiquitinated
MOLECULAR AND CELLULAR BIOLOGY, Jan. 2002, p. 105 116 Vol. 22, No. 1 0270-7306/02/$04.00 0 DOI: 10.1128/MCB.22.1.105 116.2002 Mammalian Orc1 Protein Is Selectively Released from Chromatin and Ubiquitinated
Short hairpin RNA (shrna) against MMP14. Lentiviral plasmids containing shrna
 Supplemental Materials and Methods Short hairpin RNA (shrna) against MMP14. Lentiviral plasmids containing shrna (Mission shrna, Sigma) against mouse MMP14 were transfected into HEK293 cells using FuGene6
Supplemental Materials and Methods Short hairpin RNA (shrna) against MMP14. Lentiviral plasmids containing shrna (Mission shrna, Sigma) against mouse MMP14 were transfected into HEK293 cells using FuGene6
Transforming Growth Factor -Independent Shuttling of Smad4 between the Cytoplasm and Nucleus
 MOLECULAR AND CELLULAR BIOLOGY, Dec. 2000, p. 9041 9054 Vol. 20, No. 23 0270-7306/00/$04.00 0 Copyright 2000, American Society for Microbiology. All Rights Reserved. Transforming Growth Factor -Independent
MOLECULAR AND CELLULAR BIOLOGY, Dec. 2000, p. 9041 9054 Vol. 20, No. 23 0270-7306/00/$04.00 0 Copyright 2000, American Society for Microbiology. All Rights Reserved. Transforming Growth Factor -Independent
mir-24-mediated down-regulation of H2AX suppresses DNA repair
 Supplemental Online Material mir-24-mediated down-regulation of H2AX suppresses DNA repair in terminally differentiated blood cells Ashish Lal 1,4, Yunfeng Pan 2,4, Francisco Navarro 1,4, Derek M. Dykxhoorn
Supplemental Online Material mir-24-mediated down-regulation of H2AX suppresses DNA repair in terminally differentiated blood cells Ashish Lal 1,4, Yunfeng Pan 2,4, Francisco Navarro 1,4, Derek M. Dykxhoorn
Nuclear Organization of Mammalian Genomes: Polar Chromosome Territories Build Up Functionally Distinct Higher Order Compartments
 Nuclear Organization of Mammalian Genomes: Polar Chromosome Territories Build Up Functionally Distinct Higher Order Compartments Nicolas Sadoni,* Sabine Langer,* Christine Fauth,* Giorgio Bernardi, Thomas
Nuclear Organization of Mammalian Genomes: Polar Chromosome Territories Build Up Functionally Distinct Higher Order Compartments Nicolas Sadoni,* Sabine Langer,* Christine Fauth,* Giorgio Bernardi, Thomas
Resolution of Microscopes Visible light is nm Dry lens(0.5na), green(530nm light)=0.65µm=650nm for oil lens (1.4NA) UV light (300nm) = 0.13µm f
 Microscopes and Microscopy MCB 380 Good information sources: Alberts-Molecular Biology of the Cell http://micro.magnet.fsu.edu/primer/ http://www.microscopyu.com/ Approaches to Problems in Cell Biology
Microscopes and Microscopy MCB 380 Good information sources: Alberts-Molecular Biology of the Cell http://micro.magnet.fsu.edu/primer/ http://www.microscopyu.com/ Approaches to Problems in Cell Biology
Intermediate Filaments in Motion: Observations of Intermediate Filaments in Cells Using Green Fluorescent Protein-Vimentin V
 Molecular Biology of the Cell Vol. 10, 1289 1295, May 1999 Intermediate Filaments in Motion: Observations of Intermediate Filaments in Cells Using Green Fluorescent Protein-Vimentin V Jayme L. Martys,
Molecular Biology of the Cell Vol. 10, 1289 1295, May 1999 Intermediate Filaments in Motion: Observations of Intermediate Filaments in Cells Using Green Fluorescent Protein-Vimentin V Jayme L. Martys,
7.06 Problem Set #3, Spring 2005
 7.06 Problem Set #3, Spring 2005 1. The Drosophila compound eye is composed of about 800 units called ommatidia. Each ommatidium contains eight photoreceptor neurons (R1 through R8), which develop in a
7.06 Problem Set #3, Spring 2005 1. The Drosophila compound eye is composed of about 800 units called ommatidia. Each ommatidium contains eight photoreceptor neurons (R1 through R8), which develop in a
ENCODE RBP Antibody Characterization Guidelines
 ENCODE RBP Antibody Characterization Guidelines Approved on November 18, 2016 Background An integral part of the ENCODE Project is to characterize the antibodies used in the experiments. This document
ENCODE RBP Antibody Characterization Guidelines Approved on November 18, 2016 Background An integral part of the ENCODE Project is to characterize the antibodies used in the experiments. This document
Division Ave. High School AP Biology
 Control of Eukaryotic Genes 2007-2008 The BIG Questions n How are genes turned on & off in eukaryotes? n How do cells with the same genes differentiate to perform completely different, specialized functions?
Control of Eukaryotic Genes 2007-2008 The BIG Questions n How are genes turned on & off in eukaryotes? n How do cells with the same genes differentiate to perform completely different, specialized functions?
SANTA CRUZ BIOTECHNOLOGY, INC.
 TECHNICAL SERVICE GUIDE: Western Blotting 2. What size bands were expected and what size bands were detected? 3. Was the blot blank or was a dark background or non-specific bands seen? 4. Did this same
TECHNICAL SERVICE GUIDE: Western Blotting 2. What size bands were expected and what size bands were detected? 3. Was the blot blank or was a dark background or non-specific bands seen? 4. Did this same
1. Describe the structure of DNA. Be sure to include what forms the skeleton and how are the strands held together? 2. Compare and contrast
 1. Describe the structure of DNA. Be sure to include what forms the skeleton and how are the strands held together? 2. Compare and contrast chromosomes, chromatids, genes, and alleles. 3. Compare and contrast
1. Describe the structure of DNA. Be sure to include what forms the skeleton and how are the strands held together? 2. Compare and contrast chromosomes, chromatids, genes, and alleles. 3. Compare and contrast
Figure S1. MUT-16 localization in L4 hermaphrodite, adult hermaphrodite, and adult male germlines. MUT-16 DAPI. Phillips et al. S-1. male.
 Supplementary Material for Phillips et al. Figure S1. MUT-16 localization in L4 hermaphrodite, adult hermaphrodite, and adult male germlines. Figure S2. Mutator foci and P granules localize independently
Supplementary Material for Phillips et al. Figure S1. MUT-16 localization in L4 hermaphrodite, adult hermaphrodite, and adult male germlines. Figure S2. Mutator foci and P granules localize independently
Supporting Online Material for
 www.sciencemag.org/cgi/content/full/1137999/dc1 Supporting Online Material for Disrupting the Pairing Between let-7 and Enhances Oncogenic Transformation Christine Mayr, Michael T. Hemann, David P. Bartel*
www.sciencemag.org/cgi/content/full/1137999/dc1 Supporting Online Material for Disrupting the Pairing Between let-7 and Enhances Oncogenic Transformation Christine Mayr, Michael T. Hemann, David P. Bartel*
DNA Replication. The Organization of DNA. Recall:
 Recall: The Organization of DNA DNA Replication Chromosomal form appears only during mitosis, and is used in karyotypes. folded back upon itself (chromosomes) coiled around itself (chromatin) wrapped around
Recall: The Organization of DNA DNA Replication Chromosomal form appears only during mitosis, and is used in karyotypes. folded back upon itself (chromosomes) coiled around itself (chromatin) wrapped around
Use of Phase Contrast Imaging to Track Morphological Cellular Changes due to Apoptotic Activity
 A p p l i c a t i o n N o t e Use of Phase Contrast Imaging to Track Morphological Cellular Changes due to Apoptotic Activity Brad Larson and Peter Banks, Applications Department, BioTek Instruments, Inc.,
A p p l i c a t i o n N o t e Use of Phase Contrast Imaging to Track Morphological Cellular Changes due to Apoptotic Activity Brad Larson and Peter Banks, Applications Department, BioTek Instruments, Inc.,
ENCODE DCC Antibody Validation Document
 ENCODE DCC Antibody Validation Document Date of Submission 09/12/12 Name: Trupti Kawli Email: trupti@stanford.edu Lab Snyder Antibody Name: SREBP1 (sc-8984) Target: SREBP1 Company/ Source: Santa Cruz Biotechnology
ENCODE DCC Antibody Validation Document Date of Submission 09/12/12 Name: Trupti Kawli Email: trupti@stanford.edu Lab Snyder Antibody Name: SREBP1 (sc-8984) Target: SREBP1 Company/ Source: Santa Cruz Biotechnology
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/3/6/ra27/dc Supplementary Materials for AAA+ Proteins and Coordinate PIKK Activity and Function in Nonsense-Mediated mrna Decay Natsuko Izumi, Akio Yamashita,*
www.sciencesignaling.org/cgi/content/full/3/6/ra27/dc Supplementary Materials for AAA+ Proteins and Coordinate PIKK Activity and Function in Nonsense-Mediated mrna Decay Natsuko Izumi, Akio Yamashita,*
Synchronisation of BY-2 cell culture
 Synchronisation of BY-2 cell culture Introduction Nicotiana tobacum cell suspension culture Bright Yellow 2 (BY-2) Synchronisation with aphidicolin to achieve up to 70% synchrony Aphidicolin arrests cells
Synchronisation of BY-2 cell culture Introduction Nicotiana tobacum cell suspension culture Bright Yellow 2 (BY-2) Synchronisation with aphidicolin to achieve up to 70% synchrony Aphidicolin arrests cells
MODULE 1: INTRODUCTION TO THE GENOME BROWSER: WHAT IS A GENE?
 MODULE 1: INTRODUCTION TO THE GENOME BROWSER: WHAT IS A GENE? Lesson Plan: Title Introduction to the Genome Browser: what is a gene? JOYCE STAMM Objectives Demonstrate basic skills in using the UCSC Genome
MODULE 1: INTRODUCTION TO THE GENOME BROWSER: WHAT IS A GENE? Lesson Plan: Title Introduction to the Genome Browser: what is a gene? JOYCE STAMM Objectives Demonstrate basic skills in using the UCSC Genome
Fig. 16-7a. 5 end Hydrogen bond 3 end. 1 nm. 3.4 nm nm
 Fig. 16-7a end Hydrogen bond end 1 nm 3.4 nm 0.34 nm (a) Key features of DNA structure end (b) Partial chemical structure end Fig. 16-8 Adenine (A) Thymine (T) Guanine (G) Cytosine (C) Concept 16.2: Many
Fig. 16-7a end Hydrogen bond end 1 nm 3.4 nm 0.34 nm (a) Key features of DNA structure end (b) Partial chemical structure end Fig. 16-8 Adenine (A) Thymine (T) Guanine (G) Cytosine (C) Concept 16.2: Many
Microtubule Minus-End Stabilization by Polymerization-Driven CAMSAP Deposition
 Article Microtubule Minus-End Stabilization by Polymerization-Driven CAMSAP Deposition Kai Jiang, 1 Shasha Hua, 1 Renu Mohan, 1 Ilya Grigoriev, 1 Kah Wai Yau, 1 Qingyang Liu, 1,2 Eugene A. Katrukha, 1
Article Microtubule Minus-End Stabilization by Polymerization-Driven CAMSAP Deposition Kai Jiang, 1 Shasha Hua, 1 Renu Mohan, 1 Ilya Grigoriev, 1 Kah Wai Yau, 1 Qingyang Liu, 1,2 Eugene A. Katrukha, 1
Supplemental Material for Xue et al. List. Supplemental Figure legends. Figure S1. Related to Figure 1. Figure S2. Related to Figure 3
 Supplemental Material for Xue et al List Supplemental Figure legends Figure S1. Related to Figure 1 Figure S. Related to Figure 3 Figure S3. Related to Figure 4 Figure S4. Related to Figure 4 Figure S5.
Supplemental Material for Xue et al List Supplemental Figure legends Figure S1. Related to Figure 1 Figure S. Related to Figure 3 Figure S3. Related to Figure 4 Figure S4. Related to Figure 4 Figure S5.
Acetylated Microtubules Are Preferentially Bundled Leading to Enhanced
 Biophysical Journal, Volume 113 Supplemental Information Acetylated Microtubules Are Preferentially Bundled Leading to Enhanced Kinesin-1 Motility Linda Balabanian, Christopher L. Berger, and Adam G. Hendricks
Biophysical Journal, Volume 113 Supplemental Information Acetylated Microtubules Are Preferentially Bundled Leading to Enhanced Kinesin-1 Motility Linda Balabanian, Christopher L. Berger, and Adam G. Hendricks
A CRISPR/Cas9 Vector System for Tissue-Specific Gene Disruption in Zebrafish
 Developmental Cell Supplemental Information A CRISPR/Cas9 Vector System for Tissue-Specific Gene Disruption in Zebrafish Julien Ablain, Ellen M. Durand, Song Yang, Yi Zhou, and Leonard I. Zon % larvae
Developmental Cell Supplemental Information A CRISPR/Cas9 Vector System for Tissue-Specific Gene Disruption in Zebrafish Julien Ablain, Ellen M. Durand, Song Yang, Yi Zhou, and Leonard I. Zon % larvae
Page 1. Name: 1) Which letter indicates a cell structure that directly controls the movement of molecules into and out of the cell?
 Name: 1) Which letter indicates a cell structure that directly controls the movement of molecules into and out of the cell? A) A B) B C) C D) D 2) A single-celled organism is represented in the diagram
Name: 1) Which letter indicates a cell structure that directly controls the movement of molecules into and out of the cell? A) A B) B C) C D) D 2) A single-celled organism is represented in the diagram
Nature Neuroscience: doi: /nn Supplementary Figure 1
 Supplementary Figure 1 PCR-genotyping of the three mouse models used in this study and controls for behavioral experiments after semi-chronic Pten inhibition. a-c. DNA from App/Psen1 (a), Pten tg (b) and
Supplementary Figure 1 PCR-genotyping of the three mouse models used in this study and controls for behavioral experiments after semi-chronic Pten inhibition. a-c. DNA from App/Psen1 (a), Pten tg (b) and
Introduction to Basic Human Genetics. Professor Hanan Hamamy Department of Genetic Medicine and Development Geneva University Switzerland
 Introduction to Basic Human Genetics Professor Hanan Hamamy Department of Genetic Medicine and Development Geneva University Switzerland Training Course in Sexual and Reproductive Health Research Geneva
Introduction to Basic Human Genetics Professor Hanan Hamamy Department of Genetic Medicine and Development Geneva University Switzerland Training Course in Sexual and Reproductive Health Research Geneva
Viral RNAi suppressor reversibly binds sirna to. outcompete Dicer and RISC via multiple-turnover
 Supplementary Data Viral RNAi suppressor reversibly binds sirna to outcompete Dicer and RISC via multiple-turnover Renata A. Rawlings 1,2, Vishalakshi Krishnan 2 and Nils G. Walter 2 * 1 Biophysics and
Supplementary Data Viral RNAi suppressor reversibly binds sirna to outcompete Dicer and RISC via multiple-turnover Renata A. Rawlings 1,2, Vishalakshi Krishnan 2 and Nils G. Walter 2 * 1 Biophysics and
Lab 1A: Microscopy I. Name: Section:
 Lab 1A: Microscopy I A response is required for each item marked: (# ). Your grade for the lab 1 report (1A and 1B combined) will be the fraction of correct responses on a 50 point scale[(# correct/# total)
Lab 1A: Microscopy I A response is required for each item marked: (# ). Your grade for the lab 1 report (1A and 1B combined) will be the fraction of correct responses on a 50 point scale[(# correct/# total)
DNA: The Genetic Material. Chapter 10
 DNA: The Genetic Material Chapter 10 DNA as the Genetic Material DNA was first extracted from nuclei in 1870 named nuclein after their source. Chemical analysis determined that DNA was a weak acid rich
DNA: The Genetic Material Chapter 10 DNA as the Genetic Material DNA was first extracted from nuclei in 1870 named nuclein after their source. Chemical analysis determined that DNA was a weak acid rich
The common structure of a DNA nucleotide. Hewitt
 GENETICS Unless otherwise noted* the artwork and photographs in this slide show are original and by Burt Carter. Permission is granted to use them for non-commercial, non-profit educational purposes provided
GENETICS Unless otherwise noted* the artwork and photographs in this slide show are original and by Burt Carter. Permission is granted to use them for non-commercial, non-profit educational purposes provided
