Analysis of Microarray Data
|
|
|
- Janel Rich
- 6 years ago
- Views:
Transcription
1 Analysis of Microarray Data Lecture 3: Visualization and Functional Analysis George Bell, Ph.D. Senior Bioinformatics Scientist Bioinformatics and Research Computing Whitehead Institute
2 Outline Review Visualizing all the data What to do with a set of interesting genes? Basic annotation Comparing lists Genome mapping Obtaining and analyzing promoters Gene Ontology and pathway analysis Other expression data 2
3 Generic Microarray Pipeline Design experiment Prepare samples and perform hybridizations Quantify scanned slide image Calculate expression values Normalize Handle low-level expression values Merge data for replicates Determine differentially expressed genes Cluster interesting data not covered in course 3
4 Review Preliminary filtering? Measuring differential expression: Correcting for multiple hypothesis testing Fold change, t-test, ANOVA Bonferroni, False Discovery Rate, etc. Filtering; identifying interesting genes Distance measures for clustering Clustering/segmentation types and methods What is the best analysis pipeline? Why are you doing the experiment? Are you being reasonable with the statistics? 4
5 Why draw figures? Get a global perspective of the experiments Quality control: check for low-quality data and errors Compare raw and normalized data Compare controls: are they homogeneous? Help decide how to filter data Look at a subset of data in detail 5
6 Intensity histogram Median = 6.6 Median = 100 6
7 Intensity histogram Most genes have low expression levels Using log 2 scale to transform data more normal distribution more helpful interpretation One way to observe overall intensity of chip How to choose genes with no expression? 7
8 Intensity scatterplot 8
9 Intensity scatterplot Compares intensity on two colors or chips Genes with similar expression are on the diagonal Use log-transformed expression values Genes with lower expression noisier expression harder to call significant 9
10 R-I and M-A plots 10
11 R-I and M-A plots Compares intensity on two colors or chips Like an intensity scatterplot rotated 45º R (ratio) = log(chip1 / chip2) I (intensity) = log(chip1 * chip2) M = log 2 (chip1 / chip2) A = ½(log 2 (chip1*chip2)) Popularized with lowess normalization Easier to intrepret than an intensity scatterplot 11
12 Volcano plot 12
13 Volcano plot Scatterplot showing differential expression statistics and fold change Visualize effects of filtering genes by both measures Using fold change vs. statistical measures for differential expression produce very different results 13
14 Boxplots Raw and median-normalized log 2 (expression values) 14
15 Boxplots Display summary statistics about the distribution of each chip: Median Quartiles (25% and 75% percentiles) Extreme values (>3 quartiles from median) Note that mean-normalized chips wouldn t have the same median Easy in R; much harder to do in Excel 15
16 Chip images Affymetrix U95A chip hybridized with fetal brain Image generated from.cel file Helpful for quality control 16
17 Heatmaps experiments genes 17
18 Using distance measurements Genes with most similar profiles to GPR37 18
19 Functional Analysis: intro After data is normalized, compared, filtered, clustered, and differentially expressed genes are found, what happens next? Driven by experimental questions Specificity of hypothesis testing increases power of statistical tests One general question: what s special about the differentially expressed genes? 19
20 Annotation using sequence databases Gene data can be translated into IDs from a wide variety of sequence databases: LocusLink, Ensembl, UniGene, RefSeq, genome databases Each database in turn links to a lot of different types of data Use Excel or programming tools to do this quickly Web links, instead of actual data, can also be used. What s the difference between these databases? How can all this data be integrated? 20
21 Venn diagrams Show intersection(s) between at least 2 sets Typical figure More informative figure 21
22 Mapping genes to the genome Genomic locations of differentially expressed genes Human genome, May
23 Promoter extraction Prerequisite of any promoter analysis Requires a sequenced genome and a complete, mapped cdna sequence Promoter is defined in this context as upstream regulatory sequence Extract genomic DNA using a genome browser: UCSC, Ensembl, NCBI, GBrowse, etc. Functional promoter needs to be determined experimentally 23
24 Promoter analysis TRANSFAC contains curated binding data Transcription factor binding sites can be predicted matrix (probabilities of each nt at each site) pattern (fuzzy consensus of binding site) Functional sites tend to be evolutionarily conserved Functional promoter activity needs to be verified experimentally 24
25 Gene Ontology GO is a systematic way to describe protein (gene) function GO comprises ontologies and annotations The ontologies: Molecular function Biological process Cellular component Ontologies are like hierarchies except that a child can have more than one parent. Annotation sources: publications (TAS), bioinformatics (IEA), genetics (IGI), assays (IDA), phenotypes (IMP), etc. 25
26 Gene Ontology enrichment analysis Unbiased method to ask question, What s so special about my set of genes? Obtain GO annotation (most specific term(s)) for genes in your set Climb an ontology to get all parents (more general, induced terms) Look at occurrence of each term in your set compared to terms in population (all genes or all genes on your chip) Are some terms over-represented? Ex: sample:10/100 pop1: 600/6000 pop2: 15/
27 Pathway enrichment analysis Unbiased method to ask question, Is my set of genes especially involved in specific pathways? First step: Link genes to pathways Are some pathways over-represented? Caveats What is meant by pathway"? Multiple DBs with varied annotations Annotations are very incomplete 27
28 Enrichment analysis on sorted expression data Input 1: complete sorted gene list no threshold value or definition of significance Input 2: set of biologically meaningful gene sets pathway, genome location, function,... Is the rank of genes from any gene set in your sorted list non-random? Example: GSEA Broad Institute 28
29 Comparisons with other expression studies Array repositories: GEO (NCBI), ArrayExpress (EBI), WADE (WIBR) Search for genes, chips, types of experiments, species View or download data Normalize but still expect noise Check medians and distribution of data It s much easier to make comparisons within an experiment than between experiments 29
30 Summary Plots: histogram, scatter, R-I, volcano, box Other visualizations: whole chip, heatmaps, bar graphs, Venn diagrams Annotation to sequence DBs Genome mapping Promoter extraction and analysis GO and pathway enrichment analysis Comparison with published studies 30
31 More information Course page: Bioconductor short courses: BaRC analysis tools: Gene Ontology Consortium website: Causton HC et al. Microarray Gene Expression Data Analysis: A Beginner s Guide. Blackwell, Parmigiana G et al. The Analysis of Gene Expression Data: Methods and Software. Springer,
32 Exercises Graphing all data Scatterplot R-I (M-A) plot Volcano plot Functional analysis Annotation Comparisons Genome mapping Promoter extraction and analysis GO and pathway analysis Using other expression studies 32
Outline. Analysis of Microarray Data. Most important design question. General experimental issues
 Outline Analysis of Microarray Data Lecture 1: Experimental Design and Data Normalization Introduction to microarrays Experimental design Data normalization Other data transformation Exercises George Bell,
Outline Analysis of Microarray Data Lecture 1: Experimental Design and Data Normalization Introduction to microarrays Experimental design Data normalization Other data transformation Exercises George Bell,
Analysis of Microarray Data
 Analysis of Microarray Data Lecture 1: Experimental Design and Data Normalization George Bell, Ph.D. Senior Bioinformatics Scientist Bioinformatics and Research Computing Whitehead Institute Outline Introduction
Analysis of Microarray Data Lecture 1: Experimental Design and Data Normalization George Bell, Ph.D. Senior Bioinformatics Scientist Bioinformatics and Research Computing Whitehead Institute Outline Introduction
Next-Generation Sequencing Gene Expression Analysis Using Agilent GeneSpring GX
 Next-Generation Sequencing Gene Expression Analysis Using Agilent GeneSpring GX Technical Overview Introduction RNA Sequencing (RNA-Seq) is one of the most commonly used next-generation sequencing (NGS)
Next-Generation Sequencing Gene Expression Analysis Using Agilent GeneSpring GX Technical Overview Introduction RNA Sequencing (RNA-Seq) is one of the most commonly used next-generation sequencing (NGS)
Microarray Analysis of Gene Expression in Huntington's Disease Peripheral Blood - a Platform Comparison. CodeLink compatible
 Microarray Analysis of Gene Expression in Huntington's Disease Peripheral Blood - a Platform Comparison CodeLink compatible Microarray Analysis of Gene Expression in Huntington's Disease Peripheral Blood
Microarray Analysis of Gene Expression in Huntington's Disease Peripheral Blood - a Platform Comparison CodeLink compatible Microarray Analysis of Gene Expression in Huntington's Disease Peripheral Blood
From Variants to Pathways: Agilent GeneSpring GX s Variant Analysis Workflow
 From Variants to Pathways: Agilent GeneSpring GX s Variant Analysis Workflow Technical Overview Import VCF Introduction Next-generation sequencing (NGS) studies have created unanticipated challenges with
From Variants to Pathways: Agilent GeneSpring GX s Variant Analysis Workflow Technical Overview Import VCF Introduction Next-generation sequencing (NGS) studies have created unanticipated challenges with
BCHM 6280 Tutorial: Gene specific information using NCBI, Ensembl and genome viewers
 BCHM 6280 Tutorial: Gene specific information using NCBI, Ensembl and genome viewers Web resources: NCBI database: http://www.ncbi.nlm.nih.gov/ Ensembl database: http://useast.ensembl.org/index.html UCSC
BCHM 6280 Tutorial: Gene specific information using NCBI, Ensembl and genome viewers Web resources: NCBI database: http://www.ncbi.nlm.nih.gov/ Ensembl database: http://useast.ensembl.org/index.html UCSC
Stefano Monti. Workshop Format
 Gad Getz Stefano Monti Michael Reich {gadgetz,smonti,mreich}@broad.mit.edu http://www.broad.mit.edu/~smonti/aws Broad Institute of MIT & Harvard October 18-20, 2006 Cambridge, MA Workshop Format Morning
Gad Getz Stefano Monti Michael Reich {gadgetz,smonti,mreich}@broad.mit.edu http://www.broad.mit.edu/~smonti/aws Broad Institute of MIT & Harvard October 18-20, 2006 Cambridge, MA Workshop Format Morning
EECS730: Introduction to Bioinformatics
 EECS730: Introduction to Bioinformatics Lecture 14: Microarray Some slides were adapted from Dr. Luke Huan (University of Kansas), Dr. Shaojie Zhang (University of Central Florida), and Dr. Dong Xu and
EECS730: Introduction to Bioinformatics Lecture 14: Microarray Some slides were adapted from Dr. Luke Huan (University of Kansas), Dr. Shaojie Zhang (University of Central Florida), and Dr. Dong Xu and
Analysis of a Tiling Regulation Study in Partek Genomics Suite 6.6
 Analysis of a Tiling Regulation Study in Partek Genomics Suite 6.6 The example data set used in this tutorial consists of 6 technical replicates from the same human cell line, 3 are SP1 treated, and 3
Analysis of a Tiling Regulation Study in Partek Genomics Suite 6.6 The example data set used in this tutorial consists of 6 technical replicates from the same human cell line, 3 are SP1 treated, and 3
Gene Expression Data Analysis (I)
 Gene Expression Data Analysis (I) Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Bioinformatics tasks Biological question Experiment design Microarray experiment
Gene Expression Data Analysis (I) Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Bioinformatics tasks Biological question Experiment design Microarray experiment
BABELOMICS: Microarray Data Analysis
 BABELOMICS: Microarray Data Analysis Madrid, 21 June 2010 Martina Marbà mmarba@cipf.es Bioinformatics and Genomics Department Centro de Investigación Príncipe Felipe (CIPF) (Valencia, Spain) DNA Microarrays
BABELOMICS: Microarray Data Analysis Madrid, 21 June 2010 Martina Marbà mmarba@cipf.es Bioinformatics and Genomics Department Centro de Investigación Príncipe Felipe (CIPF) (Valencia, Spain) DNA Microarrays
Exercise1 ArrayExpress Archive - High-throughput sequencing example
 ArrayExpress and Atlas practical: querying and exporting gene expression data at the EBI Gabriella Rustici gabry@ebi.ac.uk This practical will introduce you to the data content and query functionality
ArrayExpress and Atlas practical: querying and exporting gene expression data at the EBI Gabriella Rustici gabry@ebi.ac.uk This practical will introduce you to the data content and query functionality
NCBI web resources I: databases and Entrez
 NCBI web resources I: databases and Entrez Yanbin Yin Most materials are downloaded from ftp://ftp.ncbi.nih.gov/pub/education/ 1 Homework assignment 1 Two parts: Extract the gene IDs reported in table
NCBI web resources I: databases and Entrez Yanbin Yin Most materials are downloaded from ftp://ftp.ncbi.nih.gov/pub/education/ 1 Homework assignment 1 Two parts: Extract the gene IDs reported in table
IPA Advanced Training Course
 IPA Advanced Training Course Academia Sinica 2015 Oct Gene( 陳冠文 ) Supervisor and IPA certified analyst 1 Review for Introductory Training course Searching Building a Pathway Editing a Pathway for Publication
IPA Advanced Training Course Academia Sinica 2015 Oct Gene( 陳冠文 ) Supervisor and IPA certified analyst 1 Review for Introductory Training course Searching Building a Pathway Editing a Pathway for Publication
Gene Regulation Solutions. Microarrays and Next-Generation Sequencing
 Gene Regulation Solutions Microarrays and Next-Generation Sequencing Gene Regulation Solutions The Microarrays Advantage Microarrays Lead the Industry in: Comprehensive Content SurePrint G3 Human Gene
Gene Regulation Solutions Microarrays and Next-Generation Sequencing Gene Regulation Solutions The Microarrays Advantage Microarrays Lead the Industry in: Comprehensive Content SurePrint G3 Human Gene
Whole Transcriptome Analysis of Illumina RNA- Seq Data. Ryan Peters Field Application Specialist
 Whole Transcriptome Analysis of Illumina RNA- Seq Data Ryan Peters Field Application Specialist Partek GS in your NGS Pipeline Your Start-to-Finish Solution for Analysis of Next Generation Sequencing Data
Whole Transcriptome Analysis of Illumina RNA- Seq Data Ryan Peters Field Application Specialist Partek GS in your NGS Pipeline Your Start-to-Finish Solution for Analysis of Next Generation Sequencing Data
Microarray Technique. Some background. M. Nath
 Microarray Technique Some background M. Nath Outline Introduction Spotting Array Technique GeneChip Technique Data analysis Applications Conclusion Now Blind Guess? Functional Pathway Microarray Technique
Microarray Technique Some background M. Nath Outline Introduction Spotting Array Technique GeneChip Technique Data analysis Applications Conclusion Now Blind Guess? Functional Pathway Microarray Technique
CS 5984: Application of Basic Clustering Algorithms to Find Expression Modules in Cancer
 CS 5984: Application of Basic Clustering Algorithms to Find Expression Modules in Cancer T. M. Murali January 31, 2006 Innovative Application of Hierarchical Clustering A module map showing conditional
CS 5984: Application of Basic Clustering Algorithms to Find Expression Modules in Cancer T. M. Murali January 31, 2006 Innovative Application of Hierarchical Clustering A module map showing conditional
What we ll do today. Types of stem cells. Do engineered ips and ES cells have. What genes are special in stem cells?
 Do engineered ips and ES cells have similar molecular signatures? What we ll do today Research questions in stem cell biology Comparing expression and epigenetics in stem cells asuring gene expression
Do engineered ips and ES cells have similar molecular signatures? What we ll do today Research questions in stem cell biology Comparing expression and epigenetics in stem cells asuring gene expression
Exploration and Analysis of DNA Microarray Data
 Exploration and Analysis of DNA Microarray Data Dhammika Amaratunga Senior Research Fellow in Nonclinical Biostatistics Johnson & Johnson Pharmaceutical Research & Development Javier Cabrera Associate
Exploration and Analysis of DNA Microarray Data Dhammika Amaratunga Senior Research Fellow in Nonclinical Biostatistics Johnson & Johnson Pharmaceutical Research & Development Javier Cabrera Associate
Microarray Analysis of Gene Expression in Huntington's Disease Peripheral Blood - a Platform Comparison
 Microarray Analysis of Gene Expression in Huntington's Disease Peripheral Blood - a Platform Comparison Thank you for waiting. The presentation will be starting in a few minutes at 9AM Pacific Daylight
Microarray Analysis of Gene Expression in Huntington's Disease Peripheral Blood - a Platform Comparison Thank you for waiting. The presentation will be starting in a few minutes at 9AM Pacific Daylight
Do engineered ips and ES cells have similar molecular signatures?
 Do engineered ips and ES cells have similar molecular signatures? Comparing expression and epigenetics in stem cells George Bell, Ph.D. Bioinformatics and Research Computing 2012 Spring Lecture Series
Do engineered ips and ES cells have similar molecular signatures? Comparing expression and epigenetics in stem cells George Bell, Ph.D. Bioinformatics and Research Computing 2012 Spring Lecture Series
Functional analysis using EBI Metagenomics
 Functional analysis using EBI Metagenomics Contents Tutorial information... 2 Tutorial learning objectives... 2 An introduction to functional analysis using EMG... 3 What are protein signatures?... 3 Assigning
Functional analysis using EBI Metagenomics Contents Tutorial information... 2 Tutorial learning objectives... 2 An introduction to functional analysis using EMG... 3 What are protein signatures?... 3 Assigning
Designing Complex Omics Experiments
 Designing Complex Omics Experiments Xiangqin Cui Section on Statistical Genetics Department of Biostatistics University of Alabama at Birmingham 6/15/2015 Some slides are from previous lectures given by
Designing Complex Omics Experiments Xiangqin Cui Section on Statistical Genetics Department of Biostatistics University of Alabama at Birmingham 6/15/2015 Some slides are from previous lectures given by
B I O I N F O R M A T I C S
 B I O I N F O R M A T I C S Kristel Van Steen, PhD 2 Montefiore Institute - Systems and Modeling GIGA - Bioinformatics ULg kristel.vansteen@ulg.ac.be SUPPLEMENTARY CHAPTER: DATA BASES AND MINING 1 What
B I O I N F O R M A T I C S Kristel Van Steen, PhD 2 Montefiore Institute - Systems and Modeling GIGA - Bioinformatics ULg kristel.vansteen@ulg.ac.be SUPPLEMENTARY CHAPTER: DATA BASES AND MINING 1 What
Functional Enrichment Analysis & Candidate Gene Ranking
 Functional Enrichment Analysis & Candidate Gene Ranking Anil Jegga Biomedical Informatics Contact Information: Anil Jegga Biomedical Informatics Room # 232, S Building 10th Floor CCHMC Homepage: http://anil.cchmc.org
Functional Enrichment Analysis & Candidate Gene Ranking Anil Jegga Biomedical Informatics Contact Information: Anil Jegga Biomedical Informatics Room # 232, S Building 10th Floor CCHMC Homepage: http://anil.cchmc.org
Microarray Gene Expression Analysis at CNIO
 Microarray Gene Expression Analysis at CNIO Orlando Domínguez Genomics Unit Biotechnology Program, CNIO 8 May 2013 Workflow, from samples to Gene Expression data Experimental design user/gu/ubio Samples
Microarray Gene Expression Analysis at CNIO Orlando Domínguez Genomics Unit Biotechnology Program, CNIO 8 May 2013 Workflow, from samples to Gene Expression data Experimental design user/gu/ubio Samples
ProtPlot A Tissue Molecular Anatomy Program Java-based Data Mining Tool: Screen Shots
 ProtPlot A Tissue Molecular Anatomy Program Java-based Data Mining Tool: Screen Shots **** DRAFT - undergoing revision **** Peter F. Lemkin, Ph.D. (1), Djamel Medjahed Ph.D. (2) 1 NCI-Frederick; 2 SAIC-Frederick,
ProtPlot A Tissue Molecular Anatomy Program Java-based Data Mining Tool: Screen Shots **** DRAFT - undergoing revision **** Peter F. Lemkin, Ph.D. (1), Djamel Medjahed Ph.D. (2) 1 NCI-Frederick; 2 SAIC-Frederick,
Using 2-way ANOVA to dissect gene expression following myocardial infarction in mice
 Using 2-way ANOVA to dissect gene expression following myocardial infarction in mice Thank you for waiting. The presentation will be starting in a few minutes at 9AM Pacific Daylight Time. During this
Using 2-way ANOVA to dissect gene expression following myocardial infarction in mice Thank you for waiting. The presentation will be starting in a few minutes at 9AM Pacific Daylight Time. During this
Gene-centered resources at NCBI
 COURSE OF BIOINFORMATICS a.a. 2014-2015 Gene-centered resources at NCBI We searched Accession Number: M60495 AT NCBI Nucleotide Gene has been implemented at NCBI to organize information about genes, serving
COURSE OF BIOINFORMATICS a.a. 2014-2015 Gene-centered resources at NCBI We searched Accession Number: M60495 AT NCBI Nucleotide Gene has been implemented at NCBI to organize information about genes, serving
less sensitive than RNA-seq but more robust analysis pipelines expensive but quantitiatve standard but typically not high throughput
 Chapter 11: Gene Expression The availability of an annotated genome sequence enables massively parallel analysis of gene expression. The expression of all genes in an organism can be measured in one experiment.
Chapter 11: Gene Expression The availability of an annotated genome sequence enables massively parallel analysis of gene expression. The expression of all genes in an organism can be measured in one experiment.
Engineering Genetic Circuits
 Engineering Genetic Circuits I use the book and slides of Chris J. Myers Lecture 0: Preface Chris J. Myers (Lecture 0: Preface) Engineering Genetic Circuits 1 / 19 Samuel Florman Engineering is the art
Engineering Genetic Circuits I use the book and slides of Chris J. Myers Lecture 0: Preface Chris J. Myers (Lecture 0: Preface) Engineering Genetic Circuits 1 / 19 Samuel Florman Engineering is the art
DNA Microarray Data Oligonucleotide Arrays
 DNA Microarray Data Oligonucleotide Arrays Sandrine Dudoit, Robert Gentleman, Rafael Irizarry, and Yee Hwa Yang Bioconductor Short Course 2003 Copyright 2002, all rights reserved Biological question Experimental
DNA Microarray Data Oligonucleotide Arrays Sandrine Dudoit, Robert Gentleman, Rafael Irizarry, and Yee Hwa Yang Bioconductor Short Course 2003 Copyright 2002, all rights reserved Biological question Experimental
Introduction to Microarray Data Analysis and Gene Networks. Alvis Brazma European Bioinformatics Institute
 Introduction to Microarray Data Analysis and Gene Networks Alvis Brazma European Bioinformatics Institute A brief outline of this course What is gene expression, why it s important Microarrays and how
Introduction to Microarray Data Analysis and Gene Networks Alvis Brazma European Bioinformatics Institute A brief outline of this course What is gene expression, why it s important Microarrays and how
Gene Signal Estimates from Exon Arrays
 Gene Signal Estimates from Exon Arrays I. Introduction: With exon arrays like the GeneChip Human Exon 1.0 ST Array, researchers can examine the transcriptional profile of an entire gene (Figure 1). Being
Gene Signal Estimates from Exon Arrays I. Introduction: With exon arrays like the GeneChip Human Exon 1.0 ST Array, researchers can examine the transcriptional profile of an entire gene (Figure 1). Being
Protein Bioinformatics Part I: Access to information
 Protein Bioinformatics Part I: Access to information 260.655 April 6, 2006 Jonathan Pevsner, Ph.D. pevsner@kennedykrieger.org Outline [1] Proteins at NCBI RefSeq accession numbers Cn3D to visualize structures
Protein Bioinformatics Part I: Access to information 260.655 April 6, 2006 Jonathan Pevsner, Ph.D. pevsner@kennedykrieger.org Outline [1] Proteins at NCBI RefSeq accession numbers Cn3D to visualize structures
Types of Databases - By Scope
 Biological Databases Bioinformatics Workshop 2009 Chi-Cheng Lin, Ph.D. Department of Computer Science Winona State University clin@winona.edu Biological Databases Data Domains - By Scope - By Level of
Biological Databases Bioinformatics Workshop 2009 Chi-Cheng Lin, Ph.D. Department of Computer Science Winona State University clin@winona.edu Biological Databases Data Domains - By Scope - By Level of
RNA Sequencing Analyses & Mapping Uncertainty
 RNA Sequencing Analyses & Mapping Uncertainty Adam McDermaid 1/26 RNA-seq Pipelines Collection of tools for analyzing raw RNA-seq data Tier 1 Quality Check Data Trimming Tier 2 Read Alignment Assembly
RNA Sequencing Analyses & Mapping Uncertainty Adam McDermaid 1/26 RNA-seq Pipelines Collection of tools for analyzing raw RNA-seq data Tier 1 Quality Check Data Trimming Tier 2 Read Alignment Assembly
Introduction to Bioinformatics. Fabian Hoti 6.10.
 Introduction to Bioinformatics Fabian Hoti 6.10. Analysis of Microarray Data Introduction Different types of microarrays Experiment Design Data Normalization Feature selection/extraction Clustering Introduction
Introduction to Bioinformatics Fabian Hoti 6.10. Analysis of Microarray Data Introduction Different types of microarrays Experiment Design Data Normalization Feature selection/extraction Clustering Introduction
Gene Annotation and Gene Set Analysis
 Gene Annotation and Gene Set Analysis After you obtain a short list of genes/clusters/classifiers what next? For each gene, you may ask What it is What is does What processes is it involved in Which chromosome
Gene Annotation and Gene Set Analysis After you obtain a short list of genes/clusters/classifiers what next? For each gene, you may ask What it is What is does What processes is it involved in Which chromosome
Identifying Candidate Informative Genes for Biomarker Prediction of Liver Cancer
 Identifying Candidate Informative Genes for Biomarker Prediction of Liver Cancer Nagwan M. Abdel Samee 1, Nahed H. Solouma 2, Mahmoud Elhefnawy 3, Abdalla S. Ahmed 4, Yasser M. Kadah 5 1 Computer Engineering
Identifying Candidate Informative Genes for Biomarker Prediction of Liver Cancer Nagwan M. Abdel Samee 1, Nahed H. Solouma 2, Mahmoud Elhefnawy 3, Abdalla S. Ahmed 4, Yasser M. Kadah 5 1 Computer Engineering
Bioinformatics for Proteomics. Ann Loraine
 Bioinformatics for Proteomics Ann Loraine aloraine@uab.edu What is bioinformatics? The science of collecting, processing, organizing, storing, analyzing, and mining biological information, especially data
Bioinformatics for Proteomics Ann Loraine aloraine@uab.edu What is bioinformatics? The science of collecting, processing, organizing, storing, analyzing, and mining biological information, especially data
Humboldt Universität zu Berlin. Grundlagen der Bioinformatik SS Microarrays. Lecture
 Humboldt Universität zu Berlin Microarrays Grundlagen der Bioinformatik SS 2017 Lecture 6 09.06.2017 Agenda 1.mRNA: Genomic background 2.Overview: Microarray 3.Data-analysis: Quality control & normalization
Humboldt Universität zu Berlin Microarrays Grundlagen der Bioinformatik SS 2017 Lecture 6 09.06.2017 Agenda 1.mRNA: Genomic background 2.Overview: Microarray 3.Data-analysis: Quality control & normalization
Package GeneExpressionSignature
 Package GeneExpressionSignature December 30, 2017 Title Gene Expression Signature based Similarity Metric Version 1.25.0 Date 2012-10-24 Author Yang Cao Maintainer Yang Cao , Fei
Package GeneExpressionSignature December 30, 2017 Title Gene Expression Signature based Similarity Metric Version 1.25.0 Date 2012-10-24 Author Yang Cao Maintainer Yang Cao , Fei
SIMS2003. Instructors:Rus Yukhananov, Alex Loguinov BWH, Harvard Medical School. Introduction to Microarray Technology.
 SIMS2003 Instructors:Rus Yukhananov, Alex Loguinov BWH, Harvard Medical School Introduction to Microarray Technology. Lecture 1 I. EXPERIMENTAL DETAILS II. ARRAY CONSTRUCTION III. IMAGE ANALYSIS Lecture
SIMS2003 Instructors:Rus Yukhananov, Alex Loguinov BWH, Harvard Medical School Introduction to Microarray Technology. Lecture 1 I. EXPERIMENTAL DETAILS II. ARRAY CONSTRUCTION III. IMAGE ANALYSIS Lecture
Leonardo Mariño-Ramírez, PhD NCBI / NLM / NIH. BIOL 7210 A Computational Genomics 2/18/2015
 Leonardo Mariño-Ramírez, PhD NCBI / NLM / NIH BIOL 7210 A Computational Genomics 2/18/2015 The $1,000 genome is here! http://www.illumina.com/systems/hiseq-x-sequencing-system.ilmn Bioinformatics bottleneck
Leonardo Mariño-Ramírez, PhD NCBI / NLM / NIH BIOL 7210 A Computational Genomics 2/18/2015 The $1,000 genome is here! http://www.illumina.com/systems/hiseq-x-sequencing-system.ilmn Bioinformatics bottleneck
Bioinformatics Analysis of Nano-based Omics Data
 Bioinformatics Analysis of Nano-based Omics Data Penny Nymark, Pekka Kohonen, Vesa Hongisto and Roland Grafström Hands-on Workshop on Nano Safety Assessment, 29 th September, 2016, National Technical University
Bioinformatics Analysis of Nano-based Omics Data Penny Nymark, Pekka Kohonen, Vesa Hongisto and Roland Grafström Hands-on Workshop on Nano Safety Assessment, 29 th September, 2016, National Technical University
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Origin use and efficiency are similar among WT, rrm3, pif1-m2, and pif1-m2; rrm3 strains. A. Analysis of fork progression around confirmed and likely origins (from cerevisiae.oridb.org).
Supplementary Figure 1 Origin use and efficiency are similar among WT, rrm3, pif1-m2, and pif1-m2; rrm3 strains. A. Analysis of fork progression around confirmed and likely origins (from cerevisiae.oridb.org).
Why learn sequence database searching? Searching Molecular Databases with BLAST
 Why learn sequence database searching? Searching Molecular Databases with BLAST What have I cloned? Is this really!my gene"? Basic Local Alignment Search Tool How BLAST works Interpreting search results
Why learn sequence database searching? Searching Molecular Databases with BLAST What have I cloned? Is this really!my gene"? Basic Local Alignment Search Tool How BLAST works Interpreting search results
Compendium of Immune Signatures Identifies Conserved and Species-Specific Biology in Response to Inflammation
 Immunity Supplemental Information Compendium of Immune Signatures Identifies Conserved and Species-Specific Biology in Response to Inflammation Jernej Godec, Yan Tan, Arthur Liberzon, Pablo Tamayo, Sanchita
Immunity Supplemental Information Compendium of Immune Signatures Identifies Conserved and Species-Specific Biology in Response to Inflammation Jernej Godec, Yan Tan, Arthur Liberzon, Pablo Tamayo, Sanchita
Le proteine regolative variano nei vari tipi cellulari e in funzione degli stimoli ambientali
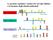 Le proteine regolative variano nei vari tipi cellulari e in funzione degli stimoli ambientali Tipo cellulare 1 Tipo cellulare 2 Tipo cellulare 3 DNA-protein Crosslink Lisi Frammentazione Immunopurificazione
Le proteine regolative variano nei vari tipi cellulari e in funzione degli stimoli ambientali Tipo cellulare 1 Tipo cellulare 2 Tipo cellulare 3 DNA-protein Crosslink Lisi Frammentazione Immunopurificazione
Overview of Health Informatics. ITI BMI-Dept
 Overview of Health Informatics ITI BMI-Dept Fellowship Week 5 Overview of Health Informatics ITI, BMI-Dept Day 10 7/5/2010 2 Agenda 1-Bioinformatics Definitions 2-System Biology 3-Bioinformatics vs Computational
Overview of Health Informatics ITI BMI-Dept Fellowship Week 5 Overview of Health Informatics ITI, BMI-Dept Day 10 7/5/2010 2 Agenda 1-Bioinformatics Definitions 2-System Biology 3-Bioinformatics vs Computational
1. Introduction. 1 Page 1
 1. INTRODUCTION Osteoblast differentiation is the key component of the bone formation, growth and remodeling. During this incompletely understood process a subpopulation of multipotential mesenchymal stem
1. INTRODUCTION Osteoblast differentiation is the key component of the bone formation, growth and remodeling. During this incompletely understood process a subpopulation of multipotential mesenchymal stem
Regulation of eukaryotic transcription:
 Promoter definition by mass genome annotation data: in silico primer extension EMBNET course Bioinformatics of transcriptional regulation Jan 28 2008 Christoph Schmid Regulation of eukaryotic transcription:
Promoter definition by mass genome annotation data: in silico primer extension EMBNET course Bioinformatics of transcriptional regulation Jan 28 2008 Christoph Schmid Regulation of eukaryotic transcription:
CS273B: Deep Learning in Genomics and Biomedicine. Recitation 1 30/9/2016
 CS273B: Deep Learning in Genomics and Biomedicine. Recitation 1 30/9/2016 Topics Genetic variation Population structure Linkage disequilibrium Natural disease variants Genome Wide Association Studies Gene
CS273B: Deep Learning in Genomics and Biomedicine. Recitation 1 30/9/2016 Topics Genetic variation Population structure Linkage disequilibrium Natural disease variants Genome Wide Association Studies Gene
Introduction to microarrays. Overview The analysis process Limitations Extensions (NGS)
 Introduction to microarrays Overview The analysis process Limitations Extensions (NGS) Outline An overview (a review) of microarrays Experiments with microarrays The data analysis process Microarray limitations
Introduction to microarrays Overview The analysis process Limitations Extensions (NGS) Outline An overview (a review) of microarrays Experiments with microarrays The data analysis process Microarray limitations
DNA Microarrays Introduction Part 2. Todd Lowe BME/BIO 210 April 11, 2007
 DNA Microarrays Introduction Part 2 Todd Lowe BME/BIO 210 April 11, 2007 Reading Assigned For Friday, please read two papers and be prepared to discuss in detail: Comprehensive Identification of Cell Cycle-related
DNA Microarrays Introduction Part 2 Todd Lowe BME/BIO 210 April 11, 2007 Reading Assigned For Friday, please read two papers and be prepared to discuss in detail: Comprehensive Identification of Cell Cycle-related
Expression Array System
 Integrated Science for Gene Expression Applied Biosystems Expression Array System Expression Array System SEE MORE GENES The most complete, most sensitive system for whole genome expression analysis. The
Integrated Science for Gene Expression Applied Biosystems Expression Array System Expression Array System SEE MORE GENES The most complete, most sensitive system for whole genome expression analysis. The
Introduction to Next Generation Sequencing (NGS) Data Analysis and Pathway Analysis. Jenny Wu
 Introduction to Next Generation Sequencing (NGS) Data Analysis and Pathway Analysis Jenny Wu Outline Introduction to NGS data analysis in Cancer Genomics NGS applications in cancer research Typical NGS
Introduction to Next Generation Sequencing (NGS) Data Analysis and Pathway Analysis Jenny Wu Outline Introduction to NGS data analysis in Cancer Genomics NGS applications in cancer research Typical NGS
Risk Management User Guide
 Risk Management User Guide Version 17 December 2017 Contents About This Guide... 5 Risk Overview... 5 Creating Projects for Risk Management... 5 Project Templates Overview... 5 Add a Project Template...
Risk Management User Guide Version 17 December 2017 Contents About This Guide... 5 Risk Overview... 5 Creating Projects for Risk Management... 5 Project Templates Overview... 5 Add a Project Template...
LIR 832: MINITAB WORKSHOP
 LIR 832: MINITAB WORKSHOP Opening Minitab Minitab will be in the Start Menu under Net Apps. Opening the Data Go to the following web site: http://www.msu.edu/course/lir/832/datasets.htm Right-click and
LIR 832: MINITAB WORKSHOP Opening Minitab Minitab will be in the Start Menu under Net Apps. Opening the Data Go to the following web site: http://www.msu.edu/course/lir/832/datasets.htm Right-click and
Microarray Data Mining and Gene Regulatory Network Analysis
 The University of Southern Mississippi The Aquila Digital Community Dissertations Spring 5-2011 Microarray Data Mining and Gene Regulatory Network Analysis Ying Li University of Southern Mississippi Follow
The University of Southern Mississippi The Aquila Digital Community Dissertations Spring 5-2011 Microarray Data Mining and Gene Regulatory Network Analysis Ying Li University of Southern Mississippi Follow
MATH 5610, Computational Biology
 MATH 5610, Computational Biology Lecture 2 Intro to Molecular Biology (cont) Stephen Billups University of Colorado at Denver MATH 5610, Computational Biology p.1/24 Announcements Error on syllabus Class
MATH 5610, Computational Biology Lecture 2 Intro to Molecular Biology (cont) Stephen Billups University of Colorado at Denver MATH 5610, Computational Biology p.1/24 Announcements Error on syllabus Class
Lecture #1. Introduction to microarray technology
 Lecture #1 Introduction to microarray technology Outline General purpose Microarray assay concept Basic microarray experimental process cdna/two channel arrays Oligonucleotide arrays Exon arrays Comparing
Lecture #1 Introduction to microarray technology Outline General purpose Microarray assay concept Basic microarray experimental process cdna/two channel arrays Oligonucleotide arrays Exon arrays Comparing
Long and short/small RNA-seq data analysis
 Long and short/small RNA-seq data analysis GEF5, 4.9.2015 Sami Heikkinen, PhD, Dos. Topics 1. RNA-seq in a nutshell 2. Long vs short/small RNA-seq 3. Bioinformatic analysis work flows GEF5 / Heikkinen
Long and short/small RNA-seq data analysis GEF5, 4.9.2015 Sami Heikkinen, PhD, Dos. Topics 1. RNA-seq in a nutshell 2. Long vs short/small RNA-seq 3. Bioinformatic analysis work flows GEF5 / Heikkinen
Introduction to genome biology
 Introduction to genome biology Lisa Stubbs We ve found most genes; but what about the rest of the genome? Genome size* 12 Mb 95 Mb 170 Mb 1500 Mb 2700 Mb 3200 Mb #coding genes ~7000 ~20000 ~14000 ~26000
Introduction to genome biology Lisa Stubbs We ve found most genes; but what about the rest of the genome? Genome size* 12 Mb 95 Mb 170 Mb 1500 Mb 2700 Mb 3200 Mb #coding genes ~7000 ~20000 ~14000 ~26000
Exploring Similarities of Conserved Domains/Motifs
 Exploring Similarities of Conserved Domains/Motifs Sotiria Palioura Abstract Traditionally, proteins are represented as amino acid sequences. There are, though, other (potentially more exciting) representations;
Exploring Similarities of Conserved Domains/Motifs Sotiria Palioura Abstract Traditionally, proteins are represented as amino acid sequences. There are, though, other (potentially more exciting) representations;
Gene Expression Technology
 Gene Expression Technology Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Gene expression Gene expression is the process by which information from a gene
Gene Expression Technology Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Gene expression Gene expression is the process by which information from a gene
Application Note. NGS Analysis of B-Cell Receptors & Antibodies by AptaAnalyzer -BCR
 Reduce to the Best Application Note NGS Analysis of B-Cell Receptors & Antibodies by AptaAnalyzer -BCR The software AptaAnalyzer harnesses next generation sequencing (NGS) data to monitor the immune response
Reduce to the Best Application Note NGS Analysis of B-Cell Receptors & Antibodies by AptaAnalyzer -BCR The software AptaAnalyzer harnesses next generation sequencing (NGS) data to monitor the immune response
Post-assembly Data Analysis
 Assembled transcriptome Post-assembly Data Analysis Quantification: get expression for each gene in each sample Genes differentially expressed between samples Clustering/network analysis Identifying over-represented
Assembled transcriptome Post-assembly Data Analysis Quantification: get expression for each gene in each sample Genes differentially expressed between samples Clustering/network analysis Identifying over-represented
Data Sheet. GeneChip Human Genome U133 Arrays
 GeneChip Human Genome Arrays AFFYMETRIX PRODUCT FAMILY > ARRAYS > Data Sheet GeneChip Human Genome U133 Arrays The Most Comprehensive Coverage of the Human Genome in Two Flexible Formats: Single-array
GeneChip Human Genome Arrays AFFYMETRIX PRODUCT FAMILY > ARRAYS > Data Sheet GeneChip Human Genome U133 Arrays The Most Comprehensive Coverage of the Human Genome in Two Flexible Formats: Single-array
massir: MicroArray Sample Sex Identifier
 massir: MicroArray Sample Sex Identifier Sam Buckberry October 30, 2017 Contents 1 The Problem 2 2 Importing data and beginning the analysis 2 3 Extracting the Y chromosome probe data 3 4 Predicting the
massir: MicroArray Sample Sex Identifier Sam Buckberry October 30, 2017 Contents 1 The Problem 2 2 Importing data and beginning the analysis 2 3 Extracting the Y chromosome probe data 3 4 Predicting the
ELE4120 Bioinformatics. Tutorial 5
 ELE4120 Bioinformatics Tutorial 5 1 1. Database Content GenBank RefSeq TPA UniProt 2. Database Searches 2 Databases A common situation for alignment is to search through a database to retrieve the similar
ELE4120 Bioinformatics Tutorial 5 1 1. Database Content GenBank RefSeq TPA UniProt 2. Database Searches 2 Databases A common situation for alignment is to search through a database to retrieve the similar
Training materials.
 Training materials Ensembl training materials are protected by a CC BY license http://creativecommons.org/licenses/by/4.0/ If you wish to re-use these materials, please credit Ensembl for their creation
Training materials Ensembl training materials are protected by a CC BY license http://creativecommons.org/licenses/by/4.0/ If you wish to re-use these materials, please credit Ensembl for their creation
Technical Note. Performance Review of the GeneChip AutoLoader for the Affymetrix GeneChip Scanner Introduction
 GeneChip AutoLoader AFFYMETRIX PRODUCT FAMILY > > Technical Note Performance Review of the GeneChip AutoLoader for the Affymetrix GeneChip ner 3000 Designed for use with the GeneChip ner 3000, the GeneChip
GeneChip AutoLoader AFFYMETRIX PRODUCT FAMILY > > Technical Note Performance Review of the GeneChip AutoLoader for the Affymetrix GeneChip ner 3000 Designed for use with the GeneChip ner 3000, the GeneChip
USDA NRSP-8 BIOINFORMATICS COORDINATION PROGRAM ACTIVITIES
 USDA NRSP-8 BIOINFORMATICS COORDINATION PROGRAM ACTIVITIES Supported by Regional Research Funds, Hatch Act for the Period 10/1/15-9/30/16 James Reecy, Sue Lamont, Chris Tuggle, Max Rothschild and Fiona
USDA NRSP-8 BIOINFORMATICS COORDINATION PROGRAM ACTIVITIES Supported by Regional Research Funds, Hatch Act for the Period 10/1/15-9/30/16 James Reecy, Sue Lamont, Chris Tuggle, Max Rothschild and Fiona
MOLECULAR BIOLOGY OF EUKARYOTES 2016 SYLLABUS
 03-442 Lectures: MWF 9:30-10:20 a.m. Doherty Hall 2105 03-742 Advanced Discussion Section: Time and place to be announced Probably Mon 4-6 p.m. or 6-8p.m.? Once we establish who is taking the advanced
03-442 Lectures: MWF 9:30-10:20 a.m. Doherty Hall 2105 03-742 Advanced Discussion Section: Time and place to be announced Probably Mon 4-6 p.m. or 6-8p.m.? Once we establish who is taking the advanced
 Genome Annotation Genome annotation What is the function of each part of the genome? Where are the genes? What is the mrna sequence (transcription, splicing) What is the protein sequence? What does
Genome Annotation Genome annotation What is the function of each part of the genome? Where are the genes? What is the mrna sequence (transcription, splicing) What is the protein sequence? What does
Hands-On Four Investigating Inherited Diseases
 Hands-On Four Investigating Inherited Diseases The purpose of these exercises is to introduce bioinformatics databases and tools. We investigate an important human gene and see how mutations give rise
Hands-On Four Investigating Inherited Diseases The purpose of these exercises is to introduce bioinformatics databases and tools. We investigate an important human gene and see how mutations give rise
Technical note: Molecular Index counting adjustment methods
 Technical note: Molecular Index counting adjustment methods By Jue Fan, Jennifer Tsai, Eleen Shum Introduction. Overview of BD Precise assays BD Precise assays are fast, high-throughput, next-generation
Technical note: Molecular Index counting adjustment methods By Jue Fan, Jennifer Tsai, Eleen Shum Introduction. Overview of BD Precise assays BD Precise assays are fast, high-throughput, next-generation
Additional file 2. Figure 1: Receiver operating characteristic (ROC) curve using the top
 Additional file 2 Figure Legends: Figure 1: Receiver operating characteristic (ROC) curve using the top discriminatory features between HIV-infected (n=32) and HIV-uninfected (n=15) individuals. The top
Additional file 2 Figure Legends: Figure 1: Receiver operating characteristic (ROC) curve using the top discriminatory features between HIV-infected (n=32) and HIV-uninfected (n=15) individuals. The top
MulCom: a Multiple Comparison statistical test for microarray data in Bioconductor.
 MulCom: a Multiple Comparison statistical test for microarray data in Bioconductor. Claudio Isella, Tommaso Renzulli, Davide Corà and Enzo Medico May 3, 2016 Abstract Many microarray experiments compare
MulCom: a Multiple Comparison statistical test for microarray data in Bioconductor. Claudio Isella, Tommaso Renzulli, Davide Corà and Enzo Medico May 3, 2016 Abstract Many microarray experiments compare
TSSpredator User Guide v 1.00
 TSSpredator User Guide v 1.00 Alexander Herbig alexander.herbig@uni-tuebingen.de Kay Nieselt kay.nieselt@uni-tuebingen.de June 3, 2013 1 Getting Started TSSpredator is a tool for the comparative detection
TSSpredator User Guide v 1.00 Alexander Herbig alexander.herbig@uni-tuebingen.de Kay Nieselt kay.nieselt@uni-tuebingen.de June 3, 2013 1 Getting Started TSSpredator is a tool for the comparative detection
Association Mapping in Plants PLSC 731 Plant Molecular Genetics Phil McClean April, 2010
 Association Mapping in Plants PLSC 731 Plant Molecular Genetics Phil McClean April, 2010 Traditional QTL approach Uses standard bi-parental mapping populations o F2 or RI These have a limited number of
Association Mapping in Plants PLSC 731 Plant Molecular Genetics Phil McClean April, 2010 Traditional QTL approach Uses standard bi-parental mapping populations o F2 or RI These have a limited number of
Entrez Gene: gene-centered information at NCBI
 D54 D58 Nucleic Acids Research, 2005, Vol. 33, Database issue doi:10.1093/nar/gki031 Entrez Gene: gene-centered information at NCBI Donna Maglott*, Jim Ostell, Kim D. Pruitt and Tatiana Tatusova National
D54 D58 Nucleic Acids Research, 2005, Vol. 33, Database issue doi:10.1093/nar/gki031 Entrez Gene: gene-centered information at NCBI Donna Maglott*, Jim Ostell, Kim D. Pruitt and Tatiana Tatusova National
The Five Key Elements of a Successful Metabolomics Study
 The Five Key Elements of a Successful Metabolomics Study Metabolomics: Completing the Biological Picture Metabolomics is offering new insights into systems biology, empowering biomarker discovery, and
The Five Key Elements of a Successful Metabolomics Study Metabolomics: Completing the Biological Picture Metabolomics is offering new insights into systems biology, empowering biomarker discovery, and
DNA Arrays Affymetrix GeneChip System
 DNA Arrays Affymetrix GeneChip System chip scanner Affymetrix Inc. hybridization Affymetrix Inc. data analysis Affymetrix Inc. mrna 5' 3' TGTGATGGTGGGAATTGGGTCAGAAGGACTGTGGGCGCTGCC... GGAATTGGGTCAGAAGGACTGTGGC
DNA Arrays Affymetrix GeneChip System chip scanner Affymetrix Inc. hybridization Affymetrix Inc. data analysis Affymetrix Inc. mrna 5' 3' TGTGATGGTGGGAATTGGGTCAGAAGGACTGTGGGCGCTGCC... GGAATTGGGTCAGAAGGACTGTGGC
Supporting Information
 Supporting Information Ho et al. 1.173/pnas.81288816 SI Methods Sequences of shrna hairpins: Brg shrna #1: ccggcggctcaagaaggaagttgaactcgagttcaacttccttcttgacgnttttg (TRCN71383; Open Biosystems). Brg shrna
Supporting Information Ho et al. 1.173/pnas.81288816 SI Methods Sequences of shrna hairpins: Brg shrna #1: ccggcggctcaagaaggaagttgaactcgagttcaacttccttcttgacgnttttg (TRCN71383; Open Biosystems). Brg shrna
Copy Number Analysis in Partek Genomics Suite 6.6
 Copy Number Analysis in Partek Genomics Suite 6.6 Introduction At a simple level, copy number analysis can be thought of as a gene dosage question: where are the regions of the genome that have altered
Copy Number Analysis in Partek Genomics Suite 6.6 Introduction At a simple level, copy number analysis can be thought of as a gene dosage question: where are the regions of the genome that have altered
3.1.4 DNA Microarray Technology
 3.1.4 DNA Microarray Technology Scientists have discovered that one of the differences between healthy and cancer is which genes are turned on in each. Scientists can compare the gene expression patterns
3.1.4 DNA Microarray Technology Scientists have discovered that one of the differences between healthy and cancer is which genes are turned on in each. Scientists can compare the gene expression patterns
ALGORITHMS IN BIO INFORMATICS. Chapman & Hall/CRC Mathematical and Computational Biology Series A PRACTICAL INTRODUCTION. CRC Press WING-KIN SUNG
 Chapman & Hall/CRC Mathematical and Computational Biology Series ALGORITHMS IN BIO INFORMATICS A PRACTICAL INTRODUCTION WING-KIN SUNG CRC Press Taylor & Francis Group Boca Raton London New York CRC Press
Chapman & Hall/CRC Mathematical and Computational Biology Series ALGORITHMS IN BIO INFORMATICS A PRACTICAL INTRODUCTION WING-KIN SUNG CRC Press Taylor & Francis Group Boca Raton London New York CRC Press
Machine Learning in Computational Biology CSC 2431
 Machine Learning in Computational Biology CSC 2431 Lecture 9: Combining biological datasets Instructor: Anna Goldenberg What kind of data integration is there? What kind of data integration is there? SNPs
Machine Learning in Computational Biology CSC 2431 Lecture 9: Combining biological datasets Instructor: Anna Goldenberg What kind of data integration is there? What kind of data integration is there? SNPs
Lees J.A., Vehkala M. et al., 2016 In Review
 Sequence element enrichment analysis to determine the genetic basis of bacterial phenotypes Lees J.A., Vehkala M. et al., 2016 In Review Journal Club Triinu Kõressaar 16.03.2016 Introduction Bacterial
Sequence element enrichment analysis to determine the genetic basis of bacterial phenotypes Lees J.A., Vehkala M. et al., 2016 In Review Journal Club Triinu Kõressaar 16.03.2016 Introduction Bacterial
Protein-Protein-Interaction Networks. Ulf Leser, Samira Jaeger
 Protein-Protein-Interaction Networks Ulf Leser, Samira Jaeger SHK Stelle frei Ab 1.9.2015, 2 Jahre, 41h/Monat Verbundprojekt MaptTorNet: Pankreatische endokrine Tumore Insb. statistische Aufbereitung und
Protein-Protein-Interaction Networks Ulf Leser, Samira Jaeger SHK Stelle frei Ab 1.9.2015, 2 Jahre, 41h/Monat Verbundprojekt MaptTorNet: Pankreatische endokrine Tumore Insb. statistische Aufbereitung und
Jobs and Skills in the Bay Area
 Jobs and Skills in the Bay Area Project Goals For my final project, I decided to work with a dataset of jobs and skills data that was collected as part of my Master s Final Project. In our Master s Final
Jobs and Skills in the Bay Area Project Goals For my final project, I decided to work with a dataset of jobs and skills data that was collected as part of my Master s Final Project. In our Master s Final
SPH 247 Statistical Analysis of Laboratory Data
 SPH 247 Statistical Analysis of Laboratory Data April 14, 2015 SPH 247 Statistical Analysis of Laboratory Data 1 Basic Design of Expression Arrays For each gene that is a target for the array, we have
SPH 247 Statistical Analysis of Laboratory Data April 14, 2015 SPH 247 Statistical Analysis of Laboratory Data 1 Basic Design of Expression Arrays For each gene that is a target for the array, we have
RNA spike-in controls & analysis methods for trustworthy genome-scale measurements
 RNA spike-in controls & analysis methods for trustworthy genome-scale measurements Sarah A. Munro, Ph.D. Genome-Scale Measurements Group ABRF Meeting March 29, 2015 Overview External RNA Controls Consortium
RNA spike-in controls & analysis methods for trustworthy genome-scale measurements Sarah A. Munro, Ph.D. Genome-Scale Measurements Group ABRF Meeting March 29, 2015 Overview External RNA Controls Consortium
Functional Genomics Overview RORY STARK PRINCIPAL BIOINFORMATICS ANALYST CRUK CAMBRIDGE INSTITUTE 18 SEPTEMBER 2017
 Functional Genomics Overview RORY STARK PRINCIPAL BIOINFORMATICS ANALYST CRUK CAMBRIDGE INSTITUTE 18 SEPTEMBER 2017 Agenda What is Functional Genomics? RNA Transcription/Gene Expression Measuring Gene
Functional Genomics Overview RORY STARK PRINCIPAL BIOINFORMATICS ANALYST CRUK CAMBRIDGE INSTITUTE 18 SEPTEMBER 2017 Agenda What is Functional Genomics? RNA Transcription/Gene Expression Measuring Gene
COMPUTATIONAL ANALYSIS OF MICROARRAY DATA
 COMPUTATIONAL ANALYSIS OF MICROARRAY DATA John Quackenbush Microarray experiments are providing unprecedented quantities of genome-wide data on gene-expression patterns. Although this technique has been
COMPUTATIONAL ANALYSIS OF MICROARRAY DATA John Quackenbush Microarray experiments are providing unprecedented quantities of genome-wide data on gene-expression patterns. Although this technique has been
Recent technology allow production of microarrays composed of 70-mers (essentially a hybrid of the two techniques)
 Microarrays and Transcript Profiling Gene expression patterns are traditionally studied using Northern blots (DNA-RNA hybridization assays). This approach involves separation of total or polya + RNA on
Microarrays and Transcript Profiling Gene expression patterns are traditionally studied using Northern blots (DNA-RNA hybridization assays). This approach involves separation of total or polya + RNA on
