Predicting Microarray Signals by Physical Modeling. Josh Deutsch. University of California. Santa Cruz
|
|
|
- Henry McCoy
- 6 years ago
- Views:
Transcription
1 Predicting Microarray Signals by Physical Modeling Josh Deutsch University of California Santa Cruz Predicting Microarray Signals by Physical Modeling p.1/39
2 Collaborators Shoudan Liang NASA Ames Onuttom Narayan UCSC Predicting Microarray Signals by Physical Modeling p.2/39
3 Outline Background Gene transcription and regulation What are microarrays Example application: Cancer diagnosis The problem: how to improve sensitivity? Microarrays in detail Physical constraints on how well they can work Different approaches to signal extraction Physical model of the system Different types of binding Experimental comparison Comparison with other approaches Conclusions Predicting Microarray Signals by Physical Modeling p.3/39
4 Gene transcription Transcription factors DNA RNA polymerase A T C A T G T A C G T A T A G T A C A T G C A T DNA messenger RNA (mrna) DNA is read by RNA polymerase pre-mrna junk A U C A U G U A C G U A spliceosome mrna Predicting Microarray Signals by Physical Modeling p.4/39
5 Gene transcription DNA A T C A T G T A C G T A T A G T A C A T G C A T DNA messenger RNA (mrna) Transcription factors RNA polymerase DNA is read by RNA polymerase pre-mrna junk A U C A U G U A C G U A Producing pre-mrna spliceosome mrna Predicting Microarray Signals by Physical Modeling p.4/39
6 Gene transcription DNA A T C A T G T A C G T A T A G T A C A T G C A T DNA messenger RNA (mrna) Transcription factors RNA polymerase DNA is read by RNA polymerase pre-mrna junk A U C A U G U A C G U A Producing pre-mrna spliceosome Which is spliced to get rid of junk mrna Predicting Microarray Signals by Physical Modeling p.4/39
7 Gene transcription protein mrna Ribosome The ribosome translates every three base pairs into one amino acid e.g. CAU Histidine. Predicting Microarray Signals by Physical Modeling p.5/39
8 Gene transcription protein mrna Ribosome The ribosome translates every three base pairs into one amino acid e.g. CAU Histidine. (Miller, Hamkalo, and Thomas) Predicting Microarray Signals by Physical Modeling p.5/39
9 Gene regulation promoter site TATAAAA DNA Sequence specific transcription factors Transcription factors RNA polymerase Many genes and cell types use the same proteins to regulate transcription. A specific combination of these is probably needed to turn on a gene. Predicting Microarray Signals by Physical Modeling p.6/39
10 Genetic networks The state of the cell can be understood from the way different proteins regulate each other and respond to external inputs. Predicting Microarray Signals by Physical Modeling p.7/39
11 What are microarrays? Gene 1 Gene 2 Gene 3 Gene 4 Fabricate a chip by synthesizing 25 base pair oligomers attached to a substrate. Predicting Microarray Signals by Physical Modeling p.8/39
12 What are microarrays? Gene 1 Gene 2 Gene 3 Gene 4 Fabricate a chip by synthesizing 25 base pair oligomers attached to a substrate. Predicting Microarray Signals by Physical Modeling p.8/39
13 Linkage to substrate E. Southern, K. Mir, M. Shchepinov Nature Genetics (1999). The oligomer probes are attached to the surface via polymeric linker molecules. Their density is about 1 probe every 40 square angstroms. Predicting Microarray Signals by Physical Modeling p.9/39
14 Sample preparation Extract mrna from cell samples mrna Predicting Microarray Signals by Physical Modeling p.10/39
15 Sample preparation Extract mrna from cell samples mrna mrna reverse transcriptase cdna Predicting Microarray Signals by Physical Modeling p.10/39
16 Hybridization to array Add fluorescent tags to targets: Predicting Microarray Signals by Physical Modeling p.11/39
17 Hybridization to array Add fluorescent tags to targets: Then hybridize to probes: Predicting Microarray Signals by Physical Modeling p.11/39
18 Linkage to substrate E. Southern, K. Mir, M. Shchepinov Nature Genetics (1999). In many experiments, target molecules can be longer or shorter than probe molecules. When they re longer, secondary structure formation can also occur. Predicting Microarray Signals by Physical Modeling p.12/39
19 A real microarray From Liming Shi /em gene-chips.com Predicting Microarray Signals by Physical Modeling p.13/39
20 Affymetrix gene-chip K.A. Baggerly (2002) This is a cell U95Av2 chip. Note the vertical bands which are evidence of misalignment with the scanner. Predicting Microarray Signals by Physical Modeling p.14/39
21 Application: Cancer diagnosis Microarrays have been used to differentially diagnose different kinds of cancer, for example Diffuse large B-cell Lymphoma Ovarian Cancer Leukemia Breast Cancer Small Round Blue Cell Tumors Predicting Microarray Signals by Physical Modeling p.15/39
22 Application: Cancer diagnosis Microarrays have been used to differentially diagnose different kinds of cancer, for example Diffuse large B-cell Lymphoma Ovarian Cancer Leukemia Breast Cancer Small Round Blue Cell Tumors Often it is hard to distinguish different sub-types of cancer by other means. These distinctions are very important because they often determine the course of treatment. Predicting Microarray Signals by Physical Modeling p.15/39
23 Small Round Blue Cell Tumors Recent work by Khan et al has used microarrays to diagnose four types of pediatric tumors, Small Round Blue Cell Tumors (SRBCT): neuroblastoma Ewing family tumors Rhabdomyosarcoma non-hodgkin lymphoma Using 63 samples for training and 20 for testing, Khan et al were able to correctly predict all data using only 96 genes from a pool of many thousands. Predicting Microarray Signals by Physical Modeling p.16/39
24 Prediction using minimal gene set EWS BL NB RMS Results for the prediction of samples for for different kinds of SRBCT. Using GESSES (J.M. Deutsch Bioinformatics (2003)). Predicting Microarray Signals by Physical Modeling p.17/39
25 High sensitivity needed Many genes critical to the function of a cell are only present in minute concentrations, for example in the initial stages of a regulatory cascade. With many orders of magnitude of concentrations present for different kinds of mrna, this pushes the limits of microarray technology. Predicting Microarray Signals by Physical Modeling p.18/39
26 High sensitivity needed Many genes critical to the function of a cell are only present in minute concentrations, for example in the initial stages of a regulatory cascade. With many orders of magnitude of concentrations present for different kinds of mrna, this pushes the limits of microarray technology input output level index Data from 1532 Series Affymetrix spike-in human gene experiments. Predicting Microarray Signals by Physical Modeling p.18/39
27 High sensitivity needed Many genes critical to the function of a cell are only present in minute concentrations, for example in the initial stages of a regulatory cascade. With many orders of magnitude of concentrations present for different kinds of mrna, this pushes the limits of microarray technology input output 1000 level index Data from 1532 Series Affymetrix spike-in human gene experiments. Predicting Microarray Signals by Physical Modeling p.18/39
28 Affymetrix Latin Square Experiments Groups of Transcripts pm Concentration A B C D E F G H I J K L M N GeneChip Experiment Predicting Microarray Signals by Physical Modeling p.19/39
29 Is this variation all noise? This variation is reproducible: Predicting Microarray Signals by Physical Modeling p.20/39
30 Is this variation all noise? This variation is reproducible: input output output 1000 level index Predicting Microarray Signals by Physical Modeling p.20/39
31 Is this variation all noise? This variation is reproducible: input output output 1000 level index It is likely due to the physicochemical differences between target and probe molecules. The output measures the amount of binding. In the case of these Affymetrix arrays, the hybridization is between crna targets in solution and DNA probes. Clearly not all target molecules are binding to the probes. Predicting Microarray Signals by Physical Modeling p.20/39
32 Why not design it to be less variable? It is hard finding optimal conditions for a microarray to work. An important parameter is the melting temperature for DNA/RNA hybridization of a probe and complementary target molecule. At temperatures << melting temperature, the time scale for relaxation becomes very long. The system becomes too sticky and irreversible. (Oligomers are 25 base pairs). Predicting Microarray Signals by Physical Modeling p.21/39
33 Why not design it to be less variable? It is hard finding optimal conditions for a microarray to work. An important parameter is the melting temperature for DNA/RNA hybridization of a probe and complementary target molecule. At temperatures << melting temperature, the time scale for relaxation becomes very long. The system becomes too sticky and irreversible. (Oligomers are 25 base pairs). At temperature >> melting temperature, nothing hybridizes. Predicting Microarray Signals by Physical Modeling p.21/39
34 Why not design it to be less variable? It is hard finding optimal conditions for a microarray to work. An important parameter is the melting temperature for DNA/RNA hybridization of a probe and complementary target molecule. At temperatures << melting temperature, the time scale for relaxation becomes very long. The system becomes too sticky and irreversible. (Oligomers are 25 base pairs). At temperature >> melting temperature, nothing hybridizes. Therefore it must operate just under a typical melting temperature Predicting Microarray Signals by Physical Modeling p.21/39
35 Why not design it to be less variable? It is hard finding optimal conditions for a microarray to work. An important parameter is the melting temperature for DNA/RNA hybridization of a probe and complementary target molecule. At temperatures << melting temperature, the time scale for relaxation becomes very long. The system becomes too sticky and irreversible. (Oligomers are 25 base pairs). At temperature >> melting temperature, nothing hybridizes. Therefore it must operate just under a typical melting temperature The melting temperature depends on the probe s chemical sequence. Predicting Microarray Signals by Physical Modeling p.21/39
36 Why not design it to be less variable? It is hard finding optimal conditions for a microarray to work. An important parameter is the melting temperature for DNA/RNA hybridization of a probe and complementary target molecule. At temperatures << melting temperature, the time scale for relaxation becomes very long. The system becomes too sticky and irreversible. (Oligomers are 25 base pairs). At temperature >> melting temperature, nothing hybridizes. Therefore it must operate just under a typical melting temperature The melting temperature depends on the probe s chemical sequence. Because of fluctuation in the melting temperature, probes with lower than average melting curves will show less hybridization. Predicting Microarray Signals by Physical Modeling p.21/39
37 Affymetrix s approach mrna sequence:...attctcaggatactgcccgttactgtctatggtcggcttcgaatgatactctcgtatatcgatcggcttatacgcgattatacgc... Probe intensities Perfect Match Mismatch TACTGTCTATGGTCGGCTTCGAATG TACTGTCTATGGACGGCTTCGAATG Predicting Microarray Signals by Physical Modeling p.22/39
38 Affymetrix uses statistics Include probes that have one base altered so that it does not hybridize as readily to the target sequence. Predicting Microarray Signals by Physical Modeling p.23/39
39 Affymetrix uses statistics Include probes that have one base altered so that it does not hybridize as readily to the target sequence. Make many probes (typically 16) per mrna, to average out over this variation as follow: Predicting Microarray Signals by Physical Modeling p.23/39
40 Affymetrix uses statistics Include probes that have one base altered so that it does not hybridize as readily to the target sequence. Make many probes (typically 16) per mrna, to average out over this variation as follow: Affymetrix MAS5.0 uses "One step Tukey s biweight estimate". Predicting Microarray Signals by Physical Modeling p.23/39
41 Affymetrix uses statistics Include probes that have one base altered so that it does not hybridize as readily to the target sequence. Make many probes (typically 16) per mrna, to average out over this variation as follow: Affymetrix MAS5.0 uses "One step Tukey s biweight estimate". It subtracts off the mismatch signal from the perfect match in cases where the perfect match is larger. Predicting Microarray Signals by Physical Modeling p.23/39
42 Affymetrix uses statistics Include probes that have one base altered so that it does not hybridize as readily to the target sequence. Make many probes (typically 16) per mrna, to average out over this variation as follow: Affymetrix MAS5.0 uses "One step Tukey s biweight estimate". It subtracts off the mismatch signal from the perfect match in cases where the perfect match is larger. Do statistics on these numbers to try to obtain a good estimate for the input target concentration. Predicting Microarray Signals by Physical Modeling p.23/39
43 Affymetrix uses statistics Include probes that have one base altered so that it does not hybridize as readily to the target sequence. Make many probes (typically 16) per mrna, to average out over this variation as follow: Affymetrix MAS5.0 uses "One step Tukey s biweight estimate". It subtracts off the mismatch signal from the perfect match in cases where the perfect match is larger. Do statistics on these numbers to try to obtain a good estimate for the input target concentration. This approach ignores the fact that these variations are largely reproducible and depend on the sequence of the probes. Predicting Microarray Signals by Physical Modeling p.23/39
44 Using sequence information Zhang, Miles, and Aldape (Nature Biotechnology 2003) invented a model to try to predict hybridization intensities using sequence information. The model they use postulates an "energy" of binding that depends on position along the sequence. One expects that the ends to be less tightly bound than the middle. There are two kinds of energy: Predicting Microarray Signals by Physical Modeling p.24/39
45 Using sequence information Zhang, Miles, and Aldape (Nature Biotechnology 2003) invented a model to try to predict hybridization intensities using sequence information. The model they use postulates an "energy" of binding that depends on position along the sequence. One expects that the ends to be less tightly bound than the middle. There are two kinds of energy: Non-specific term that gives the base line binding when the matching target is not present. Predicting Microarray Signals by Physical Modeling p.24/39
46 Using sequence information Zhang, Miles, and Aldape (Nature Biotechnology 2003) invented a model to try to predict hybridization intensities using sequence information. The model they use postulates an "energy" of binding that depends on position along the sequence. One expects that the ends to be less tightly bound than the middle. There are two kinds of energy: Non-specific term that gives the base line binding when the matching target is not present. Specific term that has a linear dependence of input target concentration on the measured signal. Predicting Microarray Signals by Physical Modeling p.24/39
47 Using sequence information Zhang, Miles, and Aldape (Nature Biotechnology 2003) invented a model to try to predict hybridization intensities using sequence information. The model they use postulates an "energy" of binding that depends on position along the sequence. One expects that the ends to be less tightly bound than the middle. There are two kinds of energy: Non-specific term that gives the base line binding when the matching target is not present. Specific term that has a linear dependence of input target concentration on the measured signal. Minimizing over (16*2 + 24*2 + 3) parameters, they obtain results far better than the MAS5.0 approach of Affymetrix. Predicting Microarray Signals by Physical Modeling p.24/39
48 Using sequence information Zhang, Miles, and Aldape (Nature Biotechnology 2003) invented a model to try to predict hybridization intensities using sequence information. The model they use postulates an "energy" of binding that depends on position along the sequence. One expects that the ends to be less tightly bound than the middle. There are two kinds of energy: Non-specific term that gives the base line binding when the matching target is not present. Specific term that has a linear dependence of input target concentration on the measured signal. Minimizing over (16*2 + 24*2 + 3) parameters, they obtain results far better than the MAS5.0 approach of Affymetrix. But does this model capture the physics? Predicting Microarray Signals by Physical Modeling p.24/39
49 Important physical effects Predicting Microarray Signals by Physical Modeling p.25/39
50 Important physical effects Nonspecific Binding Predicting Microarray Signals by Physical Modeling p.25/39
51 Important physical effects Target Target Binding Nonspecific Binding Predicting Microarray Signals by Physical Modeling p.25/39
52 Important physical effects Target Target Binding Nonspecific Binding Nonspecific binding and target-target binding are important effects to consider. Predicting Microarray Signals by Physical Modeling p.25/39
53 Evidence for target target binding output input Probes asymptote at different maximum values often correlating with their sequences being similar. Predicting Microarray Signals by Physical Modeling p.26/39
54 Evidence for nonspecific binding output input When the input target concentration is zero or small, there is still a substantial signal from the corresponding probes. Predicting Microarray Signals by Physical Modeling p.27/39
55 Partial zippering Because the binding energy is strongly dependent on base pair composition near the melting temperature, we expect that molecules are only partially bound at any one time. Predicting Microarray Signals by Physical Modeling p.28/39
56 Partial zippering Because the binding energy is strongly dependent on base pair composition near the melting temperature, we expect that molecules are only partially bound at any one time. For a segment bound from monomer n to monomer m, the free energy of the nearest neighbor model is: G mn = n 1 i=m ɛ(i, i + 1) + ɛ initiation Predicting Microarray Signals by Physical Modeling p.28/39
57 Partition function The partition function for a target molecule hybridized to a probe, Z = exp( β G) gives the total free energy for binding. It sums over all possible zippered states starting at m and ending at m. For a probe with a total of N monomers: Predicting Microarray Signals by Physical Modeling p.29/39
58 Partition function The partition function for a target molecule hybridized to a probe, Z = exp( β G) gives the total free energy for binding. It sums over all possible zippered states starting at m and ending at m. For a probe with a total of N monomers: Z = N e β G mn = N e β(p n 1 i=m ɛ(i,i+1)+ɛ initiation) m<n=2 m<n=2 sum over all begin and end points Predicting Microarray Signals by Physical Modeling p.29/39
59 Partition function The partition function for a target molecule hybridized to a probe, Z = exp( β G) gives the total free energy for binding. It sums over all possible zippered states starting at m and ending at m. For a probe with a total of N monomers: Z = N e β G mn = N e β(p n 1 i=m ɛ(i,i+1)+ɛ initiation) m<n=2 m<n=2 using the nearest neighbor model Predicting Microarray Signals by Physical Modeling p.29/39
60 Partition function The partition function for a target molecule hybridized to a probe, Z = exp( β G) gives the total free energy for binding. It sums over all possible zippered states starting at m and ending at m. For a probe with a total of N monomers: Z = N e β G mn = N e β(p n 1 i=m ɛ(i,i+1)+ɛ initiation) m<n=2 m<n=2 Predicting Microarray Signals by Physical Modeling p.29/39
61 Partition function The partition function for a target molecule hybridized to a probe, Z = exp( β G) gives the total free energy for binding. It sums over all possible zippered states starting at m and ending at m. For a probe with a total of N monomers: Z = N e β G mn = N e β(p n 1 i=m ɛ(i,i+1)+ɛ initiation) m<n=2 m<n=2 This depends strongly on the the temperature and the probe sequence. It can be efficiently computed in O(N) operations using a recursion relation for each probe considered. Predicting Microarray Signals by Physical Modeling p.29/39
62 Fraction bound Z (or G) and the chemical potential give the affinity for binding. That is the fraction f of bound probes is f = 1 G 1 + e β( G µ) Probe Solution µ = const + log(concentration)/β is determined by the requirement that µ solution = µ probes Predicting Microarray Signals by Physical Modeling p.30/39
63 Nonspecific free energy A probe can still bind, though more weakly to target sequences that have mismatches. The above method of calculation still applies. Predicting Microarray Signals by Physical Modeling p.31/39
64 Nonspecific free energy A probe can still bind, though more weakly to target sequences that have mismatches. The above method of calculation still applies. However we don t know the concentrations of the target solution because Affymetrix has added uncharacterized concentrations of human RNA as a background. Predicting Microarray Signals by Physical Modeling p.31/39
65 Nonspecific free energy A probe can still bind, though more weakly to target sequences that have mismatches. The above method of calculation still applies. However we don t know the concentrations of the target solution because Affymetrix has added uncharacterized concentrations of human RNA as a background. If the binding is weak, when can replace Z with an annealed average over different sequences. This leads to an effective nearest neighbor model for this non-specific binding G NS mn = n 1 i=m ɛ NS (i, i + 1) + ɛ NS initiation Here the ɛ NS s have to be empirically determined. Predicting Microarray Signals by Physical Modeling p.31/39
66 Target-target binding For similar reasons we don t know the concentration of targets. Therefore we replace those interactions with an effective annealed model. G T T mn = n 1 i=m ɛ T T (i, i + 1) + ɛ T T initiation Predicting Microarray Signals by Physical Modeling p.32/39
67 The full model Given an initial set of concentrations for different target molecules, our model will predict the observed intensities of hybridization of targets to probe molecules. The parameters in the model are: Energies in the nearest neighbor model ɛ(b 1, b 2 ) e.g. ɛ(g, T ). There are 16 possibilities and 3 sets of terms. specific, non-specific, and target-target. Predicting Microarray Signals by Physical Modeling p.33/39
68 The full model Given an initial set of concentrations for different target molecules, our model will predict the observed intensities of hybridization of targets to probe molecules. The parameters in the model are: Energies in the nearest neighbor model ɛ(b 1, b 2 ) e.g. ɛ(g, T ). There are 16 possibilities and 3 sets of terms. specific, non-specific, and target-target. 3 Initiation factors. Predicting Microarray Signals by Physical Modeling p.33/39
69 The full model Given an initial set of concentrations for different target molecules, our model will predict the observed intensities of hybridization of targets to probe molecules. The parameters in the model are: Energies in the nearest neighbor model ɛ(b 1, b 2 ) e.g. ɛ(g, T ). There are 16 possibilities and 3 sets of terms. specific, non-specific, and target-target. 3 Initiation factors. Predicting Microarray Signals by Physical Modeling p.33/39
70 The full model Given an initial set of concentrations for different target molecules, our model will predict the observed intensities of hybridization of targets to probe molecules. The parameters in the model are: Energies in the nearest neighbor model ɛ(b 1, b 2 ) e.g. ɛ(g, T ). There are 16 possibilities and 3 sets of terms. specific, non-specific, and target-target. 3 Initiation factors. Small additive background constant, Predicting Microarray Signals by Physical Modeling p.33/39
71 The full model Given an initial set of concentrations for different target molecules, our model will predict the observed intensities of hybridization of targets to probe molecules. The parameters in the model are: Energies in the nearest neighbor model ɛ(b 1, b 2 ) e.g. ɛ(g, T ). There are 16 possibilities and 3 sets of terms. specific, non-specific, and target-target. 3 Initiation factors. Small additive background constant, Number of probe molecules, Predicting Microarray Signals by Physical Modeling p.33/39
72 The full model Given an initial set of concentrations for different target molecules, our model will predict the observed intensities of hybridization of targets to probe molecules. The parameters in the model are: Energies in the nearest neighbor model ɛ(b 1, b 2 ) e.g. ɛ(g, T ). There are 16 possibilities and 3 sets of terms. specific, non-specific, and target-target. 3 Initiation factors. Small additive background constant, Number of probe molecules, Proportionality factor between binding and light intensity. Predicting Microarray Signals by Physical Modeling p.33/39
73 The full model Given an initial set of concentrations for different target molecules, our model will predict the observed intensities of hybridization of targets to probe molecules. The parameters in the model are: Energies in the nearest neighbor model ɛ(b 1, b 2 ) e.g. ɛ(g, T ). There are 16 possibilities and 3 sets of terms. specific, non-specific, and target-target. 3 Initiation factors. Small additive background constant, Number of probe molecules, Proportionality factor between binding and light intensity. A total of 54 parameters, to fit 2464 data points from the Latin Square experiments. Predicting Microarray Signals by Physical Modeling p.33/39
74 Parameter fitting We minimize the fitness function: I = (log(predicted conc.) log(observed conc.)) 2 all data with respect to all parameters. This is hard because there are many minima in parameter space. This can be solved using simulating annealing monte carlo, and parallel tempering. Predicting Microarray Signals by Physical Modeling p.34/39
75 Parallel tempering N 3 N 2 N 1 N T Simulate N copies of a system at temperatures T 1, T 2,..., T N. Predicting Microarray Signals by Physical Modeling p.35/39
76 Parallel tempering N 3 N 2 N 1 N T Simulate N copies of a system at temperatures T 1, T 2,..., T N. Exchange neighboring system configurations depending on the relative energies of the systems E and the temperature difference β with probability p = { exp( β E) for β E < 0, 1 otherwise. Predicting Microarray Signals by Physical Modeling p.35/39
77 Parallel tempering N 3 N 2 N 1 N T Simulate N copies of a system at temperatures T 1, T 2,..., T N. Exchange neighboring system configurations depending on the relative energies of the systems E and the temperature difference β with probability p = { exp( β E) for β E < 0, 1 otherwise. It is very efficient at finding low energy states of systems with many degrees of freedom Predicting Microarray Signals by Physical Modeling p.35/39
78 Results I min = level level index real pred input index real pred input Predicting Microarray Signals by Physical Modeling p.36/39
79 Leave out one Predict first transcript (16 probes) training on all the other data. level real pred tex index Predicting Microarray Signals by Physical Modeling p.37/39
80 Partial binding The ends of the hybridized targets tend to be frayed. Predicting Microarray Signals by Physical Modeling p.38/39
81 Partial binding The ends of the hybridized targets tend to be frayed. The fraction of the time a nucleotide is unbound depends on the sequence. Predicting Microarray Signals by Physical Modeling p.38/39
82 Partial binding The ends of the hybridized targets tend to be frayed. The fraction of the time a nucleotide is unbound depends on the sequence fraction unbound position Predicting Microarray Signals by Physical Modeling p.38/39
83 Conclusions The output of Affymetrix gene chips can be understood in terms of the physicochemical properties of equilibrium statistical mechanics. The processes involve Predicting Microarray Signals by Physical Modeling p.39/39
84 Conclusions The output of Affymetrix gene chips can be understood in terms of the physicochemical properties of equilibrium statistical mechanics. The processes involve Partially zippered RNA targets and DNA probes. Predicting Microarray Signals by Physical Modeling p.39/39
85 Conclusions The output of Affymetrix gene chips can be understood in terms of the physicochemical properties of equilibrium statistical mechanics. The processes involve Partially zippered RNA targets and DNA probes. Non-specific binding of other target molecules. Predicting Microarray Signals by Physical Modeling p.39/39
86 Conclusions The output of Affymetrix gene chips can be understood in terms of the physicochemical properties of equilibrium statistical mechanics. The processes involve Partially zippered RNA targets and DNA probes. Non-specific binding of other target molecules. Binding of target molecules to each other. Predicting Microarray Signals by Physical Modeling p.39/39
87 Conclusions The output of Affymetrix gene chips can be understood in terms of the physicochemical properties of equilibrium statistical mechanics. The processes involve Partially zippered RNA targets and DNA probes. Non-specific binding of other target molecules. Binding of target molecules to each other. Taking into account the nonlinearity (saturation) of binding. Predicting Microarray Signals by Physical Modeling p.39/39
88 Conclusions The output of Affymetrix gene chips can be understood in terms of the physicochemical properties of equilibrium statistical mechanics. The processes involve Partially zippered RNA targets and DNA probes. Non-specific binding of other target molecules. Binding of target molecules to each other. Taking into account the nonlinearity (saturation) of binding. This understanding leads a model that predicts mrna levels well. Predicting Microarray Signals by Physical Modeling p.39/39
89 Conclusions The output of Affymetrix gene chips can be understood in terms of the physicochemical properties of equilibrium statistical mechanics. The processes involve Partially zippered RNA targets and DNA probes. Non-specific binding of other target molecules. Binding of target molecules to each other. Taking into account the nonlinearity (saturation) of binding. This understanding leads a model that predicts mrna levels well. It is hoped that this model will be turned into software that will benefit researchers wanting a more accurate determination of mrna levels in their experiments. Predicting Microarray Signals by Physical Modeling p.39/39
3.1.4 DNA Microarray Technology
 3.1.4 DNA Microarray Technology Scientists have discovered that one of the differences between healthy and cancer is which genes are turned on in each. Scientists can compare the gene expression patterns
3.1.4 DNA Microarray Technology Scientists have discovered that one of the differences between healthy and cancer is which genes are turned on in each. Scientists can compare the gene expression patterns
Gene Expression Technology
 Gene Expression Technology Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Gene expression Gene expression is the process by which information from a gene
Gene Expression Technology Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Gene expression Gene expression is the process by which information from a gene
Methods of Biomaterials Testing Lesson 3-5. Biochemical Methods - Molecular Biology -
 Methods of Biomaterials Testing Lesson 3-5 Biochemical Methods - Molecular Biology - Chromosomes in the Cell Nucleus DNA in the Chromosome Deoxyribonucleic Acid (DNA) DNA has double-helix structure The
Methods of Biomaterials Testing Lesson 3-5 Biochemical Methods - Molecular Biology - Chromosomes in the Cell Nucleus DNA in the Chromosome Deoxyribonucleic Acid (DNA) DNA has double-helix structure The
Introduction to BioMEMS & Medical Microdevices DNA Microarrays and Lab-on-a-Chip Methods
 Introduction to BioMEMS & Medical Microdevices DNA Microarrays and Lab-on-a-Chip Methods Companion lecture to the textbook: Fundamentals of BioMEMS and Medical Microdevices, by Prof., http://saliterman.umn.edu/
Introduction to BioMEMS & Medical Microdevices DNA Microarrays and Lab-on-a-Chip Methods Companion lecture to the textbook: Fundamentals of BioMEMS and Medical Microdevices, by Prof., http://saliterman.umn.edu/
DNA Arrays Affymetrix GeneChip System
 DNA Arrays Affymetrix GeneChip System chip scanner Affymetrix Inc. hybridization Affymetrix Inc. data analysis Affymetrix Inc. mrna 5' 3' TGTGATGGTGGGAATTGGGTCAGAAGGACTGTGGGCGCTGCC... GGAATTGGGTCAGAAGGACTGTGGC
DNA Arrays Affymetrix GeneChip System chip scanner Affymetrix Inc. hybridization Affymetrix Inc. data analysis Affymetrix Inc. mrna 5' 3' TGTGATGGTGGGAATTGGGTCAGAAGGACTGTGGGCGCTGCC... GGAATTGGGTCAGAAGGACTGTGGC
Introduction to Bioinformatics. Fabian Hoti 6.10.
 Introduction to Bioinformatics Fabian Hoti 6.10. Analysis of Microarray Data Introduction Different types of microarrays Experiment Design Data Normalization Feature selection/extraction Clustering Introduction
Introduction to Bioinformatics Fabian Hoti 6.10. Analysis of Microarray Data Introduction Different types of microarrays Experiment Design Data Normalization Feature selection/extraction Clustering Introduction
Bioinformatics III Structural Bioinformatics and Genome Analysis. PART II: Genome Analysis. Chapter 7. DNA Microarrays
 Bioinformatics III Structural Bioinformatics and Genome Analysis PART II: Genome Analysis Chapter 7. DNA Microarrays 7.1 Motivation 7.2 DNA Microarray History and current states 7.3 DNA Microarray Techniques
Bioinformatics III Structural Bioinformatics and Genome Analysis PART II: Genome Analysis Chapter 7. DNA Microarrays 7.1 Motivation 7.2 DNA Microarray History and current states 7.3 DNA Microarray Techniques
DNA Microarray Data Oligonucleotide Arrays
 DNA Microarray Data Oligonucleotide Arrays Sandrine Dudoit, Robert Gentleman, Rafael Irizarry, and Yee Hwa Yang Bioconductor Short Course 2003 Copyright 2002, all rights reserved Biological question Experimental
DNA Microarray Data Oligonucleotide Arrays Sandrine Dudoit, Robert Gentleman, Rafael Irizarry, and Yee Hwa Yang Bioconductor Short Course 2003 Copyright 2002, all rights reserved Biological question Experimental
Lecture #1. Introduction to microarray technology
 Lecture #1 Introduction to microarray technology Outline General purpose Microarray assay concept Basic microarray experimental process cdna/two channel arrays Oligonucleotide arrays Exon arrays Comparing
Lecture #1 Introduction to microarray technology Outline General purpose Microarray assay concept Basic microarray experimental process cdna/two channel arrays Oligonucleotide arrays Exon arrays Comparing
CHAPTER 9 DNA Technologies
 CHAPTER 9 DNA Technologies Recombinant DNA Artificially created DNA that combines sequences that do not occur together in the nature Basis of much of the modern molecular biology Molecular cloning of genes
CHAPTER 9 DNA Technologies Recombinant DNA Artificially created DNA that combines sequences that do not occur together in the nature Basis of much of the modern molecular biology Molecular cloning of genes
Technical Review. Real time PCR
 Technical Review Real time PCR Normal PCR: Analyze with agarose gel Normal PCR vs Real time PCR Real-time PCR, also known as quantitative PCR (qpcr) or kinetic PCR Key feature: Used to amplify and simultaneously
Technical Review Real time PCR Normal PCR: Analyze with agarose gel Normal PCR vs Real time PCR Real-time PCR, also known as quantitative PCR (qpcr) or kinetic PCR Key feature: Used to amplify and simultaneously
Gene Expression Transcription/Translation Protein Synthesis
 Gene Expression Transcription/Translation Protein Synthesis 1. Describe how genetic information is transcribed into sequences of bases in RNA molecules and is finally translated into sequences of amino
Gene Expression Transcription/Translation Protein Synthesis 1. Describe how genetic information is transcribed into sequences of bases in RNA molecules and is finally translated into sequences of amino
Molecular Cell Biology - Problem Drill 11: Recombinant DNA
 Molecular Cell Biology - Problem Drill 11: Recombinant DNA Question No. 1 of 10 1. Which of the following statements about the sources of DNA used for molecular cloning is correct? Question #1 (A) cdna
Molecular Cell Biology - Problem Drill 11: Recombinant DNA Question No. 1 of 10 1. Which of the following statements about the sources of DNA used for molecular cloning is correct? Question #1 (A) cdna
Themes: RNA and RNA Processing. Messenger RNA (mrna) What is a gene? RNA is very versatile! RNA-RNA interactions are very important!
 Themes: RNA is very versatile! RNA and RNA Processing Chapter 14 RNA-RNA interactions are very important! Prokaryotes and Eukaryotes have many important differences. Messenger RNA (mrna) Carries genetic
Themes: RNA is very versatile! RNA and RNA Processing Chapter 14 RNA-RNA interactions are very important! Prokaryotes and Eukaryotes have many important differences. Messenger RNA (mrna) Carries genetic
Computational Biology I LSM5191
 Computational Biology I LSM5191 Lecture 5 Notes: Genetic manipulation & Molecular Biology techniques Broad Overview of: Enzymatic tools in Molecular Biology Gel electrophoresis Restriction mapping DNA
Computational Biology I LSM5191 Lecture 5 Notes: Genetic manipulation & Molecular Biology techniques Broad Overview of: Enzymatic tools in Molecular Biology Gel electrophoresis Restriction mapping DNA
Lecture Four. Molecular Approaches I: Nucleic Acids
 Lecture Four. Molecular Approaches I: Nucleic Acids I. Recombinant DNA and Gene Cloning Recombinant DNA is DNA that has been created artificially. DNA from two or more sources is incorporated into a single
Lecture Four. Molecular Approaches I: Nucleic Acids I. Recombinant DNA and Gene Cloning Recombinant DNA is DNA that has been created artificially. DNA from two or more sources is incorporated into a single
Feature Selection of Gene Expression Data for Cancer Classification: A Review
 Available online at www.sciencedirect.com ScienceDirect Procedia Computer Science 50 (2015 ) 52 57 2nd International Symposium on Big Data and Cloud Computing (ISBCC 15) Feature Selection of Gene Expression
Available online at www.sciencedirect.com ScienceDirect Procedia Computer Science 50 (2015 ) 52 57 2nd International Symposium on Big Data and Cloud Computing (ISBCC 15) Feature Selection of Gene Expression
Transcription in Eukaryotes
 Transcription in Eukaryotes Biology I Hayder A Giha Transcription Transcription is a DNA-directed synthesis of RNA, which is the first step in gene expression. Gene expression, is transformation of the
Transcription in Eukaryotes Biology I Hayder A Giha Transcription Transcription is a DNA-directed synthesis of RNA, which is the first step in gene expression. Gene expression, is transformation of the
Protein Synthesis: Transcription and Translation
 Protein Synthesis: Transcription and Translation Proteins In living things, proteins are in charge of the expression of our traits (hair/eye color, ability to make insulin, predisposition for cancer, etc.)
Protein Synthesis: Transcription and Translation Proteins In living things, proteins are in charge of the expression of our traits (hair/eye color, ability to make insulin, predisposition for cancer, etc.)
Bio 101 Sample questions: Chapter 10
 Bio 101 Sample questions: Chapter 10 1. Which of the following is NOT needed for DNA replication? A. nucleotides B. ribosomes C. Enzymes (like polymerases) D. DNA E. all of the above are needed 2 The information
Bio 101 Sample questions: Chapter 10 1. Which of the following is NOT needed for DNA replication? A. nucleotides B. ribosomes C. Enzymes (like polymerases) D. DNA E. all of the above are needed 2 The information
EECS730: Introduction to Bioinformatics
 EECS730: Introduction to Bioinformatics Lecture 14: Microarray Some slides were adapted from Dr. Luke Huan (University of Kansas), Dr. Shaojie Zhang (University of Central Florida), and Dr. Dong Xu and
EECS730: Introduction to Bioinformatics Lecture 14: Microarray Some slides were adapted from Dr. Luke Huan (University of Kansas), Dr. Shaojie Zhang (University of Central Florida), and Dr. Dong Xu and
Multiple choice questions (numbers in brackets indicate the number of correct answers)
 1 Multiple choice questions (numbers in brackets indicate the number of correct answers) February 1, 2013 1. Ribose is found in Nucleic acids Proteins Lipids RNA DNA (2) 2. Most RNA in cells is transfer
1 Multiple choice questions (numbers in brackets indicate the number of correct answers) February 1, 2013 1. Ribose is found in Nucleic acids Proteins Lipids RNA DNA (2) 2. Most RNA in cells is transfer
Replication Review. 1. What is DNA Replication? 2. Where does DNA Replication take place in eukaryotic cells?
 Replication Review 1. What is DNA Replication? 2. Where does DNA Replication take place in eukaryotic cells? 3. Where does DNA Replication take place in the cell cycle? 4. 4. What guides DNA Replication?
Replication Review 1. What is DNA Replication? 2. Where does DNA Replication take place in eukaryotic cells? 3. Where does DNA Replication take place in the cell cycle? 4. 4. What guides DNA Replication?
The Polymerase Chain Reaction. Chapter 6: Background
 The Polymerase Chain Reaction Chapter 6: Background Invention of PCR Kary Mullis Mile marker 46.58 in April of 1983 Pulled off the road and outlined a way to conduct DNA replication in a tube Worked for
The Polymerase Chain Reaction Chapter 6: Background Invention of PCR Kary Mullis Mile marker 46.58 in April of 1983 Pulled off the road and outlined a way to conduct DNA replication in a tube Worked for
DNA replication: Enzymes link the aligned nucleotides by phosphodiester bonds to form a continuous strand.
 DNA replication: Copying genetic information for transmission to the next generation Occurs in S phase of cell cycle Process of DNA duplicating itself Begins with the unwinding of the double helix to expose
DNA replication: Copying genetic information for transmission to the next generation Occurs in S phase of cell cycle Process of DNA duplicating itself Begins with the unwinding of the double helix to expose
Chapter 17. PCR the polymerase chain reaction and its many uses. Prepared by Woojoo Choi
 Chapter 17. PCR the polymerase chain reaction and its many uses Prepared by Woojoo Choi Polymerase chain reaction 1) Polymerase chain reaction (PCR): artificial amplification of a DNA sequence by repeated
Chapter 17. PCR the polymerase chain reaction and its many uses Prepared by Woojoo Choi Polymerase chain reaction 1) Polymerase chain reaction (PCR): artificial amplification of a DNA sequence by repeated
Non-Organic-Based Isolation of Mammalian microrna using Norgen s microrna Purification Kit
 Application Note 13 RNA Sample Preparation Non-Organic-Based Isolation of Mammalian microrna using Norgen s microrna Purification Kit B. Lam, PhD 1, P. Roberts, MSc 1 Y. Haj-Ahmad, M.Sc., Ph.D 1,2 1 Norgen
Application Note 13 RNA Sample Preparation Non-Organic-Based Isolation of Mammalian microrna using Norgen s microrna Purification Kit B. Lam, PhD 1, P. Roberts, MSc 1 Y. Haj-Ahmad, M.Sc., Ph.D 1,2 1 Norgen
AP Biology. Chapter 20. Biotechnology: DNA Technology & Genomics. Biotechnology. The BIG Questions. Evolution & breeding of food plants
 What do you notice about these phrases? radar racecar Madam I m Adam Able was I ere I saw Elba a man, a plan, a canal, Panama Was it a bar or a bat I saw? Chapter 20. Biotechnology: DNA Technology & enomics
What do you notice about these phrases? radar racecar Madam I m Adam Able was I ere I saw Elba a man, a plan, a canal, Panama Was it a bar or a bat I saw? Chapter 20. Biotechnology: DNA Technology & enomics
Preprocessing Affymetrix GeneChip Data. Affymetrix GeneChip Design. Terminology TGTGATGGTGGGGAATGGGTCAGAAGGCCTCCGATGCGCCGATTGAGAAT
 Preprocessing Affymetrix GeneChip Data Credit for some of today s materials: Ben Bolstad, Leslie Cope, Laurent Gautier, Terry Speed and Zhijin Wu Affymetrix GeneChip Design 5 3 Reference sequence TGTGATGGTGGGGAATGGGTCAGAAGGCCTCCGATGCGCCGATTGAGAAT
Preprocessing Affymetrix GeneChip Data Credit for some of today s materials: Ben Bolstad, Leslie Cope, Laurent Gautier, Terry Speed and Zhijin Wu Affymetrix GeneChip Design 5 3 Reference sequence TGTGATGGTGGGGAATGGGTCAGAAGGCCTCCGATGCGCCGATTGAGAAT
Intelligent DNA Chips: Logical Operation of Gene Expression Profiles on DNA Computers
 Genome Informatics 11: 33 42 (2000) 33 Intelligent DNA Chips: Logical Operation of Gene Expression Profiles on DNA Computers Yasubumi Sakakibara 1 Akira Suyama 2 yasu@j.dendai.ac.jp suyama@dna.c.u-tokyo.ac.jp
Genome Informatics 11: 33 42 (2000) 33 Intelligent DNA Chips: Logical Operation of Gene Expression Profiles on DNA Computers Yasubumi Sakakibara 1 Akira Suyama 2 yasu@j.dendai.ac.jp suyama@dna.c.u-tokyo.ac.jp
Chapter 15 Gene Technologies and Human Applications
 Chapter Outline Chapter 15 Gene Technologies and Human Applications Section 1: The Human Genome KEY IDEAS > Why is the Human Genome Project so important? > How do genomics and gene technologies affect
Chapter Outline Chapter 15 Gene Technologies and Human Applications Section 1: The Human Genome KEY IDEAS > Why is the Human Genome Project so important? > How do genomics and gene technologies affect
DNA. translation. base pairing rules for DNA Replication. thymine. cytosine. amino acids. The building blocks of proteins are?
 2 strands, has the 5-carbon sugar deoxyribose, and has the nitrogen base Thymine. The actual process of assembling the proteins on the ribosome is called? DNA translation Adenine pairs with Thymine, Thymine
2 strands, has the 5-carbon sugar deoxyribose, and has the nitrogen base Thymine. The actual process of assembling the proteins on the ribosome is called? DNA translation Adenine pairs with Thymine, Thymine
Recent technology allow production of microarrays composed of 70-mers (essentially a hybrid of the two techniques)
 Microarrays and Transcript Profiling Gene expression patterns are traditionally studied using Northern blots (DNA-RNA hybridization assays). This approach involves separation of total or polya + RNA on
Microarrays and Transcript Profiling Gene expression patterns are traditionally studied using Northern blots (DNA-RNA hybridization assays). This approach involves separation of total or polya + RNA on
qpcr Quantitative PCR or Real-time PCR Gives a measurement of PCR product at end of each cycle real time
 qpcr qpcr Quantitative PCR or Real-time PCR Gives a measurement of PCR product at end of each cycle real time Differs from endpoint PCR gel on last cycle Used to determines relative amount of template
qpcr qpcr Quantitative PCR or Real-time PCR Gives a measurement of PCR product at end of each cycle real time Differs from endpoint PCR gel on last cycle Used to determines relative amount of template
M I C R O B I O L O G Y WITH DISEASES BY TAXONOMY, THIRD EDITION
 M I C R O B I O L O G Y WITH DISEASES BY TAXONOMY, THIRD EDITION Chapter 7 Microbial Genetics Lecture prepared by Mindy Miller-Kittrell, University of Tennessee, Knoxville The Structure and Replication
M I C R O B I O L O G Y WITH DISEASES BY TAXONOMY, THIRD EDITION Chapter 7 Microbial Genetics Lecture prepared by Mindy Miller-Kittrell, University of Tennessee, Knoxville The Structure and Replication
Review of Protein (one or more polypeptide) A polypeptide is a long chain of..
 Gene expression Review of Protein (one or more polypeptide) A polypeptide is a long chain of.. In a protein, the sequence of amino acid determines its which determines the protein s A protein with an enzymatic
Gene expression Review of Protein (one or more polypeptide) A polypeptide is a long chain of.. In a protein, the sequence of amino acid determines its which determines the protein s A protein with an enzymatic
CHAPTER 20 DNA TECHNOLOGY AND GENOMICS. Section A: DNA Cloning
 Section A: DNA Cloning 1. DNA technology makes it possible to clone genes for basic research and commercial applications: an overview 2. Restriction enzymes are used to make recombinant DNA 3. Genes can
Section A: DNA Cloning 1. DNA technology makes it possible to clone genes for basic research and commercial applications: an overview 2. Restriction enzymes are used to make recombinant DNA 3. Genes can
Humboldt Universität zu Berlin. Grundlagen der Bioinformatik SS Microarrays. Lecture
 Humboldt Universität zu Berlin Microarrays Grundlagen der Bioinformatik SS 2017 Lecture 6 09.06.2017 Agenda 1.mRNA: Genomic background 2.Overview: Microarray 3.Data-analysis: Quality control & normalization
Humboldt Universität zu Berlin Microarrays Grundlagen der Bioinformatik SS 2017 Lecture 6 09.06.2017 Agenda 1.mRNA: Genomic background 2.Overview: Microarray 3.Data-analysis: Quality control & normalization
From Gene to Protein Transcription and Translation
 Name: Hour: From Gene to Protein Transcription and Translation Introduction: In this activity you will learn how the genes in our DNA influence our characteristics. For example, how can a gene cause albinism
Name: Hour: From Gene to Protein Transcription and Translation Introduction: In this activity you will learn how the genes in our DNA influence our characteristics. For example, how can a gene cause albinism
Make the protein through the genetic dogma process.
 Make the protein through the genetic dogma process. Coding Strand 5 AGCAATCATGGATTGGGTACATTTGTAACTGT 3 Template Strand mrna Protein Complete the table. DNA strand DNA s strand G mrna A C U G T A T Amino
Make the protein through the genetic dogma process. Coding Strand 5 AGCAATCATGGATTGGGTACATTTGTAACTGT 3 Template Strand mrna Protein Complete the table. DNA strand DNA s strand G mrna A C U G T A T Amino
Chapter 8 Lecture Outline. Transcription, Translation, and Bioinformatics
 Chapter 8 Lecture Outline Transcription, Translation, and Bioinformatics Replication, Transcription, Translation n Repetitive processes Build polymers of nucleotides or amino acids n All have 3 major steps
Chapter 8 Lecture Outline Transcription, Translation, and Bioinformatics Replication, Transcription, Translation n Repetitive processes Build polymers of nucleotides or amino acids n All have 3 major steps
Introduction to Microarray Data Analysis and Gene Networks. Alvis Brazma European Bioinformatics Institute
 Introduction to Microarray Data Analysis and Gene Networks Alvis Brazma European Bioinformatics Institute A brief outline of this course What is gene expression, why it s important Microarrays and how
Introduction to Microarray Data Analysis and Gene Networks Alvis Brazma European Bioinformatics Institute A brief outline of this course What is gene expression, why it s important Microarrays and how
Recombinant DNA Technology. The Role of Recombinant DNA Technology in Biotechnology. yeast. Biotechnology. Recombinant DNA technology.
 PowerPoint Lecture Presentations prepared by Mindy Miller-Kittrell, North Carolina State University C H A P T E R 8 Recombinant DNA Technology The Role of Recombinant DNA Technology in Biotechnology Biotechnology?
PowerPoint Lecture Presentations prepared by Mindy Miller-Kittrell, North Carolina State University C H A P T E R 8 Recombinant DNA Technology The Role of Recombinant DNA Technology in Biotechnology Biotechnology?
Microarray Technique. Some background. M. Nath
 Microarray Technique Some background M. Nath Outline Introduction Spotting Array Technique GeneChip Technique Data analysis Applications Conclusion Now Blind Guess? Functional Pathway Microarray Technique
Microarray Technique Some background M. Nath Outline Introduction Spotting Array Technique GeneChip Technique Data analysis Applications Conclusion Now Blind Guess? Functional Pathway Microarray Technique
Student Exploration: RNA and Protein Synthesis Due Wednesday 11/27/13
 http://www.explorelearning.com Name: Period : Student Exploration: RNA and Protein Synthesis Due Wednesday 11/27/13 Vocabulary: Define these terms in complete sentences on a separate piece of paper: amino
http://www.explorelearning.com Name: Period : Student Exploration: RNA and Protein Synthesis Due Wednesday 11/27/13 Vocabulary: Define these terms in complete sentences on a separate piece of paper: amino
Neural Networks and Applications in Bioinformatics. Yuzhen Ye School of Informatics and Computing, Indiana University
 Neural Networks and Applications in Bioinformatics Yuzhen Ye School of Informatics and Computing, Indiana University Contents Biological problem: promoter modeling Basics of neural networks Perceptrons
Neural Networks and Applications in Bioinformatics Yuzhen Ye School of Informatics and Computing, Indiana University Contents Biological problem: promoter modeling Basics of neural networks Perceptrons
PROTEIN SYNTHESIS. copyright cmassengale
 PROTEIN SYNTHESIS 1 DNA and Genes 2 Roles of RNA and DNA DNA is the MASTER PLAN RNA is the BLUEPRINT of the Master Plan 3 RNA Differs from DNA RNA has a sugar ribose DNA has a sugar deoxyribose 4 Other
PROTEIN SYNTHESIS 1 DNA and Genes 2 Roles of RNA and DNA DNA is the MASTER PLAN RNA is the BLUEPRINT of the Master Plan 3 RNA Differs from DNA RNA has a sugar ribose DNA has a sugar deoxyribose 4 Other
Chapter 14 Active Reading Guide From Gene to Protein
 Name: AP Biology Mr. Croft Chapter 14 Active Reading Guide From Gene to Protein This is going to be a very long journey, but it is crucial to your understanding of biology. Work on this chapter a single
Name: AP Biology Mr. Croft Chapter 14 Active Reading Guide From Gene to Protein This is going to be a very long journey, but it is crucial to your understanding of biology. Work on this chapter a single
LATE-PCR. Linear-After-The-Exponential
 LATE-PCR Linear-After-The-Exponential A Patented Invention of the Laboratory of Human Genetics and Reproductive Biology Lab. Director: Lawrence J. Wangh, Ph.D. Department of Biology, Brandeis University,
LATE-PCR Linear-After-The-Exponential A Patented Invention of the Laboratory of Human Genetics and Reproductive Biology Lab. Director: Lawrence J. Wangh, Ph.D. Department of Biology, Brandeis University,
Transcription Eukaryotic Cells
 Transcription Eukaryotic Cells Packet #20 1 Introduction Transcription is the process in which genetic information, stored in a strand of DNA (gene), is copied into a strand of RNA. Protein-encoding genes
Transcription Eukaryotic Cells Packet #20 1 Introduction Transcription is the process in which genetic information, stored in a strand of DNA (gene), is copied into a strand of RNA. Protein-encoding genes
Chapter 14: Gene Expression: From Gene to Protein
 Chapter 14: Gene Expression: From Gene to Protein This is going to be a very long journey, but it is crucial to your understanding of biology. Work on this chapter a single concept at a time, and expect
Chapter 14: Gene Expression: From Gene to Protein This is going to be a very long journey, but it is crucial to your understanding of biology. Work on this chapter a single concept at a time, and expect
Genetic Engineering & Recombinant DNA
 Genetic Engineering & Recombinant DNA Chapter 10 Copyright The McGraw-Hill Companies, Inc) Permission required for reproduction or display. Applications of Genetic Engineering Basic science vs. Applied
Genetic Engineering & Recombinant DNA Chapter 10 Copyright The McGraw-Hill Companies, Inc) Permission required for reproduction or display. Applications of Genetic Engineering Basic science vs. Applied
Protein Synthesis. Lab Exercise 12. Introduction. Contents. Objectives
 Lab Exercise Protein Synthesis Contents Objectives 1 Introduction 1 Activity.1 Overview of Process 2 Activity.2 Transcription 2 Activity.3 Translation 3 Resutls Section 4 Introduction Having information
Lab Exercise Protein Synthesis Contents Objectives 1 Introduction 1 Activity.1 Overview of Process 2 Activity.2 Transcription 2 Activity.3 Translation 3 Resutls Section 4 Introduction Having information
Protein Synthesis Notes
 Protein Synthesis Notes Protein Synthesis: Overview Transcription: synthesis of mrna under the direction of DNA. Translation: actual synthesis of a polypeptide under the direction of mrna. Transcription
Protein Synthesis Notes Protein Synthesis: Overview Transcription: synthesis of mrna under the direction of DNA. Translation: actual synthesis of a polypeptide under the direction of mrna. Transcription
What does PLIER really do?
 What does PLIER really do? Terry M. Therneau Karla V. Ballman Technical Report #75 November 2005 Copyright 2005 Mayo Foundation 1 Abstract Motivation: Our goal was to understand why the PLIER algorithm
What does PLIER really do? Terry M. Therneau Karla V. Ballman Technical Report #75 November 2005 Copyright 2005 Mayo Foundation 1 Abstract Motivation: Our goal was to understand why the PLIER algorithm
The Double Helix. DNA and RNA, part 2. Part A. Hint 1. The difference between purines and pyrimidines. Hint 2. Distinguish purines from pyrimidines
 DNA and RNA, part 2 Due: 3:00pm on Wednesday, September 24, 2014 You will receive no credit for items you complete after the assignment is due. Grading Policy The Double Helix DNA, or deoxyribonucleic
DNA and RNA, part 2 Due: 3:00pm on Wednesday, September 24, 2014 You will receive no credit for items you complete after the assignment is due. Grading Policy The Double Helix DNA, or deoxyribonucleic
RNA and Protein Synthesis
 RNA and Protein Synthesis CTE: Agriculture and Natural Resources: C5.3 Understand various cell actions, such as osmosis and cell division. C5.4 Compare and contrast plant and animal cells, bacteria, and
RNA and Protein Synthesis CTE: Agriculture and Natural Resources: C5.3 Understand various cell actions, such as osmosis and cell division. C5.4 Compare and contrast plant and animal cells, bacteria, and
TRANSCRIPTION AND TRANSLATION
 TRANSCRIPTION AND TRANSLATION Bell Ringer (5 MINUTES) 1. Have your homework (any missing work) out on your desk and ready to turn in 2. Draw and label a nucleotide. 3. Summarize the steps of DNA replication.
TRANSCRIPTION AND TRANSLATION Bell Ringer (5 MINUTES) 1. Have your homework (any missing work) out on your desk and ready to turn in 2. Draw and label a nucleotide. 3. Summarize the steps of DNA replication.
Functional Genomics Overview RORY STARK PRINCIPAL BIOINFORMATICS ANALYST CRUK CAMBRIDGE INSTITUTE 18 SEPTEMBER 2017
 Functional Genomics Overview RORY STARK PRINCIPAL BIOINFORMATICS ANALYST CRUK CAMBRIDGE INSTITUTE 18 SEPTEMBER 2017 Agenda What is Functional Genomics? RNA Transcription/Gene Expression Measuring Gene
Functional Genomics Overview RORY STARK PRINCIPAL BIOINFORMATICS ANALYST CRUK CAMBRIDGE INSTITUTE 18 SEPTEMBER 2017 Agenda What is Functional Genomics? RNA Transcription/Gene Expression Measuring Gene
Chapter 17: From Gene to Protein
 Name Period This is going to be a very long journey, but it is crucial to your understanding of biology. Work on this chapter a single concept at a time, and expect to spend at least 6 hours to truly master
Name Period This is going to be a very long journey, but it is crucial to your understanding of biology. Work on this chapter a single concept at a time, and expect to spend at least 6 hours to truly master
Bundle 6 Test Review
 Bundle 6 Test Review DNA vs. RNA DNA Replication Gene Mutations- Protein Synthesis 1. Label the different components and complete the complimentary base pairing. What is this molecule called? Deoxyribonucleic
Bundle 6 Test Review DNA vs. RNA DNA Replication Gene Mutations- Protein Synthesis 1. Label the different components and complete the complimentary base pairing. What is this molecule called? Deoxyribonucleic
DNA Transcription. Visualizing Transcription. The Transcription Process
 DNA Transcription By: Suzanne Clancy, Ph.D. 2008 Nature Education Citation: Clancy, S. (2008) DNA transcription. Nature Education 1(1) If DNA is a book, then how is it read? Learn more about the DNA transcription
DNA Transcription By: Suzanne Clancy, Ph.D. 2008 Nature Education Citation: Clancy, S. (2008) DNA transcription. Nature Education 1(1) If DNA is a book, then how is it read? Learn more about the DNA transcription
7.2 Protein Synthesis. From DNA to Protein Animation
 7.2 Protein Synthesis From DNA to Protein Animation Proteins Why are proteins so important? They break down your food They build up muscles They send signals through your brain that control your body They
7.2 Protein Synthesis From DNA to Protein Animation Proteins Why are proteins so important? They break down your food They build up muscles They send signals through your brain that control your body They
Transcription. Unit: DNA. Central Dogma. 2. Transcription converts DNA into RNA. What is a gene? What is transcription? 1/7/2016
 Warm Up Questions 1. Where is DNA located? 2. Name the 3 parts of a nucleotide. 3. Enzymes can catalyze many different reactions (T or F) 4. How many variables should you have in an experiment? 5. A red
Warm Up Questions 1. Where is DNA located? 2. Name the 3 parts of a nucleotide. 3. Enzymes can catalyze many different reactions (T or F) 4. How many variables should you have in an experiment? 5. A red
RNA & PROTEIN SYNTHESIS
 RNA & PROTEIN SYNTHESIS DNA & RNA Genes are coded DNA instructions that control the production of proteins within the cell. The first step in decoding these genetic messages is to copy part of the nucleotide
RNA & PROTEIN SYNTHESIS DNA & RNA Genes are coded DNA instructions that control the production of proteins within the cell. The first step in decoding these genetic messages is to copy part of the nucleotide
DNA & Protein Synthesis #21
 Name: Period: Date: Living Environment Lab DNA & Protein Synthesis #21 Introduction Of all the molecules that is in the body, DNA is perhaps the most important. DNA or dioxiribosenucleic acid is important
Name: Period: Date: Living Environment Lab DNA & Protein Synthesis #21 Introduction Of all the molecules that is in the body, DNA is perhaps the most important. DNA or dioxiribosenucleic acid is important
Do you remember. What is a gene? What is RNA? How does it differ from DNA? What is protein?
 Lesson 1 - RNA Do you remember What is a gene? What is RNA? How does it differ from DNA? What is protein? Gene Segment of DNA that codes for building a protein DNA code is copied into RNA form, and RNA
Lesson 1 - RNA Do you remember What is a gene? What is RNA? How does it differ from DNA? What is protein? Gene Segment of DNA that codes for building a protein DNA code is copied into RNA form, and RNA
Nucleic Acids: DNA and RNA
 Nucleic Acids: DNA and RNA Living organisms are complex systems. Hundreds of thousands of proteins exist inside each one of us to help carry out our daily functions. These proteins are produced locally,
Nucleic Acids: DNA and RNA Living organisms are complex systems. Hundreds of thousands of proteins exist inside each one of us to help carry out our daily functions. These proteins are produced locally,
Ch. 10 Notes DNA: Transcription and Translation
 Ch. 10 Notes DNA: Transcription and Translation GOALS Compare the structure of RNA with that of DNA Summarize the process of transcription Relate the role of codons to the sequence of amino acids that
Ch. 10 Notes DNA: Transcription and Translation GOALS Compare the structure of RNA with that of DNA Summarize the process of transcription Relate the role of codons to the sequence of amino acids that
DNA makes RNA makes Proteins. The Central Dogma
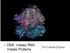 DNA makes RNA makes Proteins The Central Dogma TRANSCRIPTION DNA RNA transcript RNA polymerase RNA PROCESSING Exon RNA transcript (pre-mrna) Intron Aminoacyl-tRNA synthetase NUCLEUS CYTOPLASM FORMATION
DNA makes RNA makes Proteins The Central Dogma TRANSCRIPTION DNA RNA transcript RNA polymerase RNA PROCESSING Exon RNA transcript (pre-mrna) Intron Aminoacyl-tRNA synthetase NUCLEUS CYTOPLASM FORMATION
TRANSCRIPTION COMPARISON OF DNA & RNA TRANSCRIPTION. Umm AL Qura University. Sugar Ribose Deoxyribose. Bases AUCG ATCG. Strand length Short Long
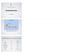 Umm AL Qura University TRANSCRIPTION Dr Neda Bogari TRANSCRIPTION COMPARISON OF DNA & RNA RNA DNA Sugar Ribose Deoxyribose Bases AUCG ATCG Strand length Short Long No. strands One Two Helix Single Double
Umm AL Qura University TRANSCRIPTION Dr Neda Bogari TRANSCRIPTION COMPARISON OF DNA & RNA RNA DNA Sugar Ribose Deoxyribose Bases AUCG ATCG Strand length Short Long No. strands One Two Helix Single Double
Protein Synthesis. OpenStax College
 OpenStax-CNX module: m46032 1 Protein Synthesis OpenStax College This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution License 3.0 By the end of this section, you will
OpenStax-CNX module: m46032 1 Protein Synthesis OpenStax College This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution License 3.0 By the end of this section, you will
Transcription is the first step of gene expression, in which a particular segment of DNA is copied into RNA by the enzyme, RNA polymerase.
 Transcription in Bacteria Transcription in Bacteria Transcription in Bacteria Transcription is the first step of gene expression, in which a particular segment of DNA is copied into RNA by the enzyme,
Transcription in Bacteria Transcription in Bacteria Transcription in Bacteria Transcription is the first step of gene expression, in which a particular segment of DNA is copied into RNA by the enzyme,
Bio11 Announcements. Ch 21: DNA Biology and Technology. DNA Functions. DNA and RNA Structure. How do DNA and RNA differ? What are genes?
 Bio11 Announcements TODAY Genetics (review) and quiz (CP #4) Structure and function of DNA Extra credit due today Next week in lab: Case study presentations Following week: Lab Quiz 2 Ch 21: DNA Biology
Bio11 Announcements TODAY Genetics (review) and quiz (CP #4) Structure and function of DNA Extra credit due today Next week in lab: Case study presentations Following week: Lab Quiz 2 Ch 21: DNA Biology
STUDY GUIDE SECTION 10-1 Discovery of DNA
 STUDY GUIDE SECTION 10-1 Discovery of DNA Name Period Date Multiple Choice-Write the correct letter in the blank. 1. The virulent strain of the bacterium S. pneumoniae causes disease because it a. has
STUDY GUIDE SECTION 10-1 Discovery of DNA Name Period Date Multiple Choice-Write the correct letter in the blank. 1. The virulent strain of the bacterium S. pneumoniae causes disease because it a. has
If Dna Has The Instructions For Building Proteins Why Is Mrna Needed
 If Dna Has The Instructions For Building Proteins Why Is Mrna Needed if a strand of DNA has the sequence CGGTATATC, then the complementary each strand of DNA contains the info needed to produce the complementary
If Dna Has The Instructions For Building Proteins Why Is Mrna Needed if a strand of DNA has the sequence CGGTATATC, then the complementary each strand of DNA contains the info needed to produce the complementary
DNA RNA PROTEIN SYNTHESIS -NOTES-
 DNA RNA PROTEIN SYNTHESIS -NOTES- THE COMPONENTS AND STRUCTURE OF DNA DNA is made up of units called nucleotides. Nucleotides are made up of three basic components:, called deoxyribose in DNA In DNA, there
DNA RNA PROTEIN SYNTHESIS -NOTES- THE COMPONENTS AND STRUCTURE OF DNA DNA is made up of units called nucleotides. Nucleotides are made up of three basic components:, called deoxyribose in DNA In DNA, there
Department. Zoology & Biotechnology QUESTION BANK BIOTECHNOLOGY SEMESTER-V
 Department of Zoology & Biotechnology QUESTION BANK BIOTECHNOLOGY SEMESTER-V Unit-1 Genetic Material Different forms of DNA(DNA topology):- B-form, Z-form, D-form; Gene structure-introns,exaons and pseudogenes:
Department of Zoology & Biotechnology QUESTION BANK BIOTECHNOLOGY SEMESTER-V Unit-1 Genetic Material Different forms of DNA(DNA topology):- B-form, Z-form, D-form; Gene structure-introns,exaons and pseudogenes:
Optimizing Synthetic DNA for Metabolic Engineering Applications. Howard Salis Penn State University
 Optimizing Synthetic DNA for Metabolic Engineering Applications Howard Salis Penn State University Synthetic Biology Specify a function Build a genetic system (a DNA molecule) Genetic Pseudocode call producequorumsignal(luxi
Optimizing Synthetic DNA for Metabolic Engineering Applications Howard Salis Penn State University Synthetic Biology Specify a function Build a genetic system (a DNA molecule) Genetic Pseudocode call producequorumsignal(luxi
1. The diagram below shows an error in the transcription of a DNA template to messenger RNA (mrna).
 1. The diagram below shows an error in the transcription of a DNA template to messenger RNA (mrna). Which statement best describes the error shown in the diagram? (A) The mrna strand contains the uracil
1. The diagram below shows an error in the transcription of a DNA template to messenger RNA (mrna). Which statement best describes the error shown in the diagram? (A) The mrna strand contains the uracil
Gene Signal Estimates from Exon Arrays
 Gene Signal Estimates from Exon Arrays I. Introduction: With exon arrays like the GeneChip Human Exon 1.0 ST Array, researchers can examine the transcriptional profile of an entire gene (Figure 1). Being
Gene Signal Estimates from Exon Arrays I. Introduction: With exon arrays like the GeneChip Human Exon 1.0 ST Array, researchers can examine the transcriptional profile of an entire gene (Figure 1). Being
REGULATION OF GENE EXPRESSION
 REGULATION OF GENE EXPRESSION Each cell of a living organism contains thousands of genes. But all genes do not function at a time. Genes function according to requirements of the cell. Genes control the
REGULATION OF GENE EXPRESSION Each cell of a living organism contains thousands of genes. But all genes do not function at a time. Genes function according to requirements of the cell. Genes control the
DNA is the genetic material. DNA structure. Chapter 7: DNA Replication, Transcription & Translation; Mutations & Ames test
 DNA is the genetic material Chapter 7: DNA Replication, Transcription & Translation; Mutations & Ames test Dr. Amy Rogers Bio 139 General Microbiology Hereditary information is carried by DNA Griffith/Avery
DNA is the genetic material Chapter 7: DNA Replication, Transcription & Translation; Mutations & Ames test Dr. Amy Rogers Bio 139 General Microbiology Hereditary information is carried by DNA Griffith/Avery
Click here to read the case study about protein synthesis.
 Click here to read the case study about protein synthesis. Big Question: How do cells use the genetic information stored in DNA to make millions of different proteins the body needs? Key Concept: Genetics
Click here to read the case study about protein synthesis. Big Question: How do cells use the genetic information stored in DNA to make millions of different proteins the body needs? Key Concept: Genetics
Unit VII DNA to RNA to protein The Central Dogma
 Unit VII DNA to RNA to protein The Central Dogma DNA Deoxyribonucleic acid, the material that contains information that determines inherited characteristics. A DNA molecule is shaped like a spiral staircase
Unit VII DNA to RNA to protein The Central Dogma DNA Deoxyribonucleic acid, the material that contains information that determines inherited characteristics. A DNA molecule is shaped like a spiral staircase
Transcription and Translation
 Biology Name: Morales Date: Period: Transcription and Translation Directions: Read the following and answer the questions in complete sentences. DNA is the molecule of heredity it determines an organism
Biology Name: Morales Date: Period: Transcription and Translation Directions: Read the following and answer the questions in complete sentences. DNA is the molecule of heredity it determines an organism
Roche Molecular Biochemicals Technical Note No. LC 10/2000
 Roche Molecular Biochemicals Technical Note No. LC 10/2000 LightCycler Overview of LightCycler Quantification Methods 1. General Introduction Introduction Content Definitions This Technical Note will introduce
Roche Molecular Biochemicals Technical Note No. LC 10/2000 LightCycler Overview of LightCycler Quantification Methods 1. General Introduction Introduction Content Definitions This Technical Note will introduce
Chapter 13 - Concept Mapping
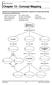 Chapter 13 - Concept Mapping Using the terms and phrases provided below, complete the concept map showing the discovery of DNA structure. amount of base pairs five-carbon sugar purine DNA polymerases Franklin
Chapter 13 - Concept Mapping Using the terms and phrases provided below, complete the concept map showing the discovery of DNA structure. amount of base pairs five-carbon sugar purine DNA polymerases Franklin
Goals of pharmacogenomics
 Goals of pharmacogenomics Use drugs better and use better drugs! People inherit/exhibit differences in drug: Absorption Metabolism and degradation of the drug Transport of drug to the target molecule Excretion
Goals of pharmacogenomics Use drugs better and use better drugs! People inherit/exhibit differences in drug: Absorption Metabolism and degradation of the drug Transport of drug to the target molecule Excretion
Activity A: Build a DNA molecule
 Name: Date: Student Exploration: Building DNA Vocabulary: double helix, DNA, enzyme, lagging strand, leading strand, mutation, nitrogenous base, nucleoside, nucleotide, replication Prior Knowledge Questions
Name: Date: Student Exploration: Building DNA Vocabulary: double helix, DNA, enzyme, lagging strand, leading strand, mutation, nitrogenous base, nucleoside, nucleotide, replication Prior Knowledge Questions
SPH 247 Statistical Analysis of Laboratory Data
 SPH 247 Statistical Analysis of Laboratory Data April 14, 2015 SPH 247 Statistical Analysis of Laboratory Data 1 Basic Design of Expression Arrays For each gene that is a target for the array, we have
SPH 247 Statistical Analysis of Laboratory Data April 14, 2015 SPH 247 Statistical Analysis of Laboratory Data 1 Basic Design of Expression Arrays For each gene that is a target for the array, we have
Reliable classification of two-class cancer data using evolutionary algorithms
 BioSystems 72 (23) 111 129 Reliable classification of two-class cancer data using evolutionary algorithms Kalyanmoy Deb, A. Raji Reddy Kanpur Genetic Algorithms Laboratory (KanGAL), Indian Institute of
BioSystems 72 (23) 111 129 Reliable classification of two-class cancer data using evolutionary algorithms Kalyanmoy Deb, A. Raji Reddy Kanpur Genetic Algorithms Laboratory (KanGAL), Indian Institute of
PROTEIN SYNTHESIS Flow of Genetic Information The flow of genetic information can be symbolized as: DNA RNA Protein
 PROTEIN SYNTHESIS Flow of Genetic Information The flow of genetic information can be symbolized as: DNA RNA Protein This is also known as: The central dogma of molecular biology Protein Proteins are made
PROTEIN SYNTHESIS Flow of Genetic Information The flow of genetic information can be symbolized as: DNA RNA Protein This is also known as: The central dogma of molecular biology Protein Proteins are made
AP Biology Gene Expression/Biotechnology REVIEW
 AP Biology Gene Expression/Biotechnology REVIEW Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Gene expression can be a. regulated before transcription.
AP Biology Gene Expression/Biotechnology REVIEW Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Gene expression can be a. regulated before transcription.
Gene Expression: Transcription
 Gene Expression: Transcription The majority of genes are expressed as the proteins they encode. The process occurs in two steps: Transcription = DNA RNA Translation = RNA protein Taken together, they make
Gene Expression: Transcription The majority of genes are expressed as the proteins they encode. The process occurs in two steps: Transcription = DNA RNA Translation = RNA protein Taken together, they make
Year III Pharm.D Dr. V. Chitra
 Year III Pharm.D Dr. V. Chitra 1 Genome entire genetic material of an individual Transcriptome set of transcribed sequences Proteome set of proteins encoded by the genome 2 Only one strand of DNA serves
Year III Pharm.D Dr. V. Chitra 1 Genome entire genetic material of an individual Transcriptome set of transcribed sequences Proteome set of proteins encoded by the genome 2 Only one strand of DNA serves
MODULE 1: INTRODUCTION TO THE GENOME BROWSER: WHAT IS A GENE?
 MODULE 1: INTRODUCTION TO THE GENOME BROWSER: WHAT IS A GENE? Lesson Plan: Title Introduction to the Genome Browser: what is a gene? JOYCE STAMM Objectives Demonstrate basic skills in using the UCSC Genome
MODULE 1: INTRODUCTION TO THE GENOME BROWSER: WHAT IS A GENE? Lesson Plan: Title Introduction to the Genome Browser: what is a gene? JOYCE STAMM Objectives Demonstrate basic skills in using the UCSC Genome
From Gene to Protein Transcription and Translation i
 How do genes influence our characteristics? From Gene to Protein Transcription and Translation i A gene is a segment of DNA that provides the instructions for making a protein. Proteins have many different
How do genes influence our characteristics? From Gene to Protein Transcription and Translation i A gene is a segment of DNA that provides the instructions for making a protein. Proteins have many different
Lecture 25 (11/15/17)
 Lecture 25 (11/15/17) Reading: Ch9; 328-332 Ch25; 990-995, 1005-1012 Problems: Ch9 (study-guide: applying); 1,2 Ch9 (study-guide: facts); 7,8 Ch25 (text); 1-3,5-7,9,10,13-15 Ch25 (study-guide: applying);
Lecture 25 (11/15/17) Reading: Ch9; 328-332 Ch25; 990-995, 1005-1012 Problems: Ch9 (study-guide: applying); 1,2 Ch9 (study-guide: facts); 7,8 Ch25 (text); 1-3,5-7,9,10,13-15 Ch25 (study-guide: applying);
