IPA : Maximizing the Biological Interpretation of Gene, Transcript & Protein Expression Data with IPA
|
|
|
- August Lambert
- 6 years ago
- Views:
Transcription
1 IPA : Maximizing the Biological Interpretation of Gene, Transcript & Protein Expression Data with IPA Marisa Chen Account Manager Qiagen Advanced Genomics Marisa.Chen@qiagen.com (203) Dev Mistry, Ph.D. Field Applications Scientist Qiagen Advanced Genomics Devendra.Mistry@qiagen.com
2 Overview Introduction to IPA Search and Explore Growing a network out of a molecule Bioprofiler (Advanced Analytics) Large Dataset Analysis Uploading your dataset(s) and starting a core analysis Core Analysis Canonical Pathways Upstream Regulators Causal Network (Advanced Analytics) Diseases and Functions Regulator Effect Networks Comparison Analysis Questions/Answer Title, Location, Date 4
3 Introduction Title, Location, Date 6
4 When do you use IPA? QIAGEN Sample Prep Assay Data Sequence- Level Statistics Biology of Interest (Genes, Variants, etc.) Annotation & Comparative (Statistical) Analysis Annotation & Biological Interpretation Upstream Analysis Primary Secondary Tertiary Insight Sample QIAGEN
5 Cell-Junction CT pathway ILK pathway Cancer Cellular Movement Invasion Cell Death TGF SNAI1 MYC P53 What can IPA do? Upstream Regulators IPA Biological Processes Canonical Pathways Large RNA seq dataset in form of a huge pile of papers Systematic analysis by IPA in form of organized binders on a bookshelf
6 Foundation behind all of Ingenuity s products Ingenuity Content The Ingenuity Knowledge Base The Ingenuity Ontology Ingenuity Findings Ingenuity Expert Findings Manually curated Findings that are reviewed, from the full-text, rich with contextual details, and are derived from top journals. Ingenuity ExpertAssist Findings Automated text Findings that are reviewed, from abstracts, timely, and cover a broad range of publications. Ingenuity Modeled Knowledge Ingenuity Expert Knowledge Content we model such as pathways, toxicity lists, etc. Ingenuity Supported Third Party Information Content areas include Protein-Protein, mirna, biomarker, clinical trial information, and others Species: human, mouse and rat Data from other species can be mapped to human, mouse and rat orthologues
7 Species Supported Species Supported Human, Mouse, Rat in full content IPA uses HomoloGene to map other identifiers to human/mouse/rat orthologs (though supporting content for the additional species will be specific to human, mouse, and rat) Arabidopsis thaliana Bos taurus (bovine) Caenorhabditis elegans Gallus gallus (chicken) Pan troglodytes (chimpanzee) Danio rerio (zebrafish) Canis lupus familiaris (canine) Drosophila melanogaster Macaca mulatta (Rhesus Monkey) Saccharomyces cerevisiae Schizosaccharomyces pombe Introduction to QIAGEN Ingenuity & IPA
8 # of publications Peer-reviewed publications citing QIAGEN s Ingenuity products 12,513 publications and growing! , thru Dec 2014
9 Two different types of analyses by IPA Deep pathway understanding of a single gene/protein Biological understanding of large data sets Title, Location, Date 12
10 How can IPA help you? Deep pathway understanding of a single gene/protein Drug/therapeutic target discovery 13
11 How can IPA help you? Biological understanding of large data sets Differential gene expression, array and RNAseq (transcriptomics) Differential protein expression (proteomics) Metabolomics mirna expression Gene List Chip-seq sirna screening Methylation Protein phosphorylation Title, Location, Date 16
12 Gene/Protein Expression Analysis IPA Core Analysis Canonical Pathway Analysis Predicts pathways that are changing based on gene expression New tools to predict directional effects on the pathway (MAP overlay tool) Upstream Regulator Analysis Predicts what regulators caused changes in gene expression Predicts directional state of regulator Creates de novo pathways based on upstream regulators (Mechanistic Networks) Diseases and Functions Analysis Predicts effected biology (cellular processes, biological functions) based on gene expression and predicts directional change on that effect Increase in apoptosis Decrease in proliferation Regulator Effects Networks Models pathway interactions from predicted upstream regulators, through differentially expressed genes, to biological processes Predicts non-directional gene interaction map Upstream Regulator Diseases & Functions
13 Change Preferences 1. Scroll down 2. Change memory to >1000mb 3. Save Title, Location, Date 21
14 Don t worry too much about notes or if you fall behind during the point and click training. We have manuals/videos for everything. Title, Location, Date 22
15 Uploading your dataset 23
16 Suggested Format for uploading RNA-seq data Required Recommended Max RPKM recommended for RNA seq
17 Case Study RNA Seq: Claudin Low vs Luminal Breast cancer cell lines Title, Location, Date 29
18 Adapted from Aroeira et al. J Am Soc Nephrol 18: , 2007 Epithelial to Mesenchymal Transition Luminal Breast cancer HER2-enriched Basal Mesenchymal / stem cell-like breast cancer Luminal cell lines Ratio Claudin-low to Luminal 5 vs 5 cell lines, RNA-Seq data Claudin-low cell lines Proprietary and Confidential 30
19 IPA Analysis Verify the biology Can IPA identify cancer and EMT related pathways and biological functions in this dataset? What are some of the relevant pathways? What are some of the relevant biological functions? Identification of transcriptional regulators What are the transcriptional regulators that are causing the gene expression changes in this dataset? Are they activated or inhibited? Hypothesis generation Are the predicted upstream regulators increasing or decreasing downstream biological functions? Title, Location, Date 31
20 P value and Z Score Title, Location, Date 32
21 Overlapping P-value Overlapping Molecules Genes in your filtered dataset Genes in the reference universe Genes from Knowledge Base Genes from previous literature that belong to A canonical pathway OR Downstream of an upstream regulator OR Upstream of a disease or function Different from the Expression P-value uploaded with your dataset Calculated using Fisher s exact test The statistical test looks for an unexpectedly large overlap given the number of molecules in each category p-values should be insignificant (<0.05) for random datasets Gene expression direction is not taken into account for this calculation
22 Z-score: Activation Prediction Gene expression from Knowledge Base (literature) Gene expression in your dataset score for the consistent and -1 for the inconsistent relationships = (7-1)/ 8 = 2.12 (= predicted activation) z-score is a statistical measure of the match between expected relationship direction and observed gene expression z-score > 2 or < -2 is considered significant Note that the actual z-score is weighted by the underlying findings, the relationship bias, and dataset bias
23 Upstream Regulators vs. Causal Networks Leveraging the network to create more upstream regulators Advanced Analytics: Causal Network Analysis Upstream Regulators Causal Network Master Regulator Regulator Dataset Regulators
24 Comparison Analysis
25 Canonical Pathways
26 Upstream Analysis
27 Diseases and Functions
28 Regulator Effects
29 Network Analysis
30 Cell-Junction CT pathway ILK pathway Cancer Cellular Movement Invasion Cell Death TGF SNAI1 MYC P53 What can IPA do? Cell Upstream Regulators Nature IPA Science JBC Biological Processes Canonical Pathways
31 Contact Us am-5pm Pacific Time (M-F) QIAGEN Redwood City 1700 Seaport Blvd., 3rd Floor Redwood City CA 94063, USA 45
32 Help > Legend Introduction to QIAGEN Ingenuity & IPA
IPA Advanced Training Course
 IPA Advanced Training Course Academia Sinica 2015 Oct Gene( 陳冠文 ) Supervisor and IPA certified analyst 1 Review for Introductory Training course Searching Building a Pathway Editing a Pathway for Publication
IPA Advanced Training Course Academia Sinica 2015 Oct Gene( 陳冠文 ) Supervisor and IPA certified analyst 1 Review for Introductory Training course Searching Building a Pathway Editing a Pathway for Publication
From Variants to Pathways: Agilent GeneSpring GX s Variant Analysis Workflow
 From Variants to Pathways: Agilent GeneSpring GX s Variant Analysis Workflow Technical Overview Import VCF Introduction Next-generation sequencing (NGS) studies have created unanticipated challenges with
From Variants to Pathways: Agilent GeneSpring GX s Variant Analysis Workflow Technical Overview Import VCF Introduction Next-generation sequencing (NGS) studies have created unanticipated challenges with
Gene Regulation Solutions. Microarrays and Next-Generation Sequencing
 Gene Regulation Solutions Microarrays and Next-Generation Sequencing Gene Regulation Solutions The Microarrays Advantage Microarrays Lead the Industry in: Comprehensive Content SurePrint G3 Human Gene
Gene Regulation Solutions Microarrays and Next-Generation Sequencing Gene Regulation Solutions The Microarrays Advantage Microarrays Lead the Industry in: Comprehensive Content SurePrint G3 Human Gene
AP BIOLOGY. Investigation #3 Comparing DNA Sequences to Understand Evolutionary Relationships with BLAST. Slide 1 / 32. Slide 2 / 32.
 New Jersey Center for Teaching and Learning Slide 1 / 32 Progressive Science Initiative This material is made freely available at www.njctl.org and is intended for the non-commercial use of students and
New Jersey Center for Teaching and Learning Slide 1 / 32 Progressive Science Initiative This material is made freely available at www.njctl.org and is intended for the non-commercial use of students and
Bioinformatics Analysis of Nano-based Omics Data
 Bioinformatics Analysis of Nano-based Omics Data Penny Nymark, Pekka Kohonen, Vesa Hongisto and Roland Grafström Hands-on Workshop on Nano Safety Assessment, 29 th September, 2016, National Technical University
Bioinformatics Analysis of Nano-based Omics Data Penny Nymark, Pekka Kohonen, Vesa Hongisto and Roland Grafström Hands-on Workshop on Nano Safety Assessment, 29 th September, 2016, National Technical University
Training Account. Account: ~ Password: ingenuity123. Sample & Assay Technologies
 Training Account Account: asininca21@ingenuity.com ~ asinica40@ingenuity.com Password: ingenuity123 1 IPA Introductory Training Course Academia Sinica 2014 September Chris (Yu-Lun Kuo) 2 About me Chris
Training Account Account: asininca21@ingenuity.com ~ asinica40@ingenuity.com Password: ingenuity123 1 IPA Introductory Training Course Academia Sinica 2014 September Chris (Yu-Lun Kuo) 2 About me Chris
QIAGEN s NGS Solutions for Biomarkers NGS & Bioinformatics team QIAGEN (Suzhou) Translational Medicine Co.,Ltd
 QIAGEN s NGS Solutions for Biomarkers NGS & Bioinformatics team QIAGEN (Suzhou) Translational Medicine Co.,Ltd 1 Our current NGS & Bioinformatics Platform 2 Our NGS workflow and applications 3 QIAGEN s
QIAGEN s NGS Solutions for Biomarkers NGS & Bioinformatics team QIAGEN (Suzhou) Translational Medicine Co.,Ltd 1 Our current NGS & Bioinformatics Platform 2 Our NGS workflow and applications 3 QIAGEN s
Whole Transcriptome Analysis of Illumina RNA- Seq Data. Ryan Peters Field Application Specialist
 Whole Transcriptome Analysis of Illumina RNA- Seq Data Ryan Peters Field Application Specialist Partek GS in your NGS Pipeline Your Start-to-Finish Solution for Analysis of Next Generation Sequencing Data
Whole Transcriptome Analysis of Illumina RNA- Seq Data Ryan Peters Field Application Specialist Partek GS in your NGS Pipeline Your Start-to-Finish Solution for Analysis of Next Generation Sequencing Data
SureSilencing sirna Array Technology Overview
 SureSilencing sirna Array Technology Overview Pathway-Focused sirna-based RNA Interference Topics to be Covered Who is SuperArray? Brief Introduction to RNA Interference Challenges Facing RNA Interference
SureSilencing sirna Array Technology Overview Pathway-Focused sirna-based RNA Interference Topics to be Covered Who is SuperArray? Brief Introduction to RNA Interference Challenges Facing RNA Interference
Next-Generation Sequencing Gene Expression Analysis Using Agilent GeneSpring GX
 Next-Generation Sequencing Gene Expression Analysis Using Agilent GeneSpring GX Technical Overview Introduction RNA Sequencing (RNA-Seq) is one of the most commonly used next-generation sequencing (NGS)
Next-Generation Sequencing Gene Expression Analysis Using Agilent GeneSpring GX Technical Overview Introduction RNA Sequencing (RNA-Seq) is one of the most commonly used next-generation sequencing (NGS)
Welcome to the NGS webinar series
 Welcome to the NGS webinar series Webinar 1 NGS: Introduction to technology, and applications NGS Technology Webinar 2 Targeted NGS for Cancer Research NGS in cancer Webinar 3 NGS: Data analysis for genetic
Welcome to the NGS webinar series Webinar 1 NGS: Introduction to technology, and applications NGS Technology Webinar 2 Targeted NGS for Cancer Research NGS in cancer Webinar 3 NGS: Data analysis for genetic
China National Grid --- BioNode. Jun Wang Beijing Genomics Institute
 China National Grid --- BioNode Jun Wang Beijing Genomics Institute Core of life science and bio-tech: Getting, Mining, Applying the basic life information Old China meets New China? Sequencing, sequencing,
China National Grid --- BioNode Jun Wang Beijing Genomics Institute Core of life science and bio-tech: Getting, Mining, Applying the basic life information Old China meets New China? Sequencing, sequencing,
High Profile Publishing in Molecular Biology. Hélène Hodak, Marina Ostankovitch,
 High Profile Publishing in Molecular Biology Hélène Hodak, h.hodak@elsevier.com Marina Ostankovitch, m.ostankovitch@elsevier.com The field of Molecular Biology, inception to current trends Preparing a
High Profile Publishing in Molecular Biology Hélène Hodak, h.hodak@elsevier.com Marina Ostankovitch, m.ostankovitch@elsevier.com The field of Molecular Biology, inception to current trends Preparing a
Complete automation for NGS interpretation and reporting with evidence-based clinical decision support
 Brochure Bioinformatics for Clinical Oncology Testing Complete automation for NGS interpretation and reporting with evidence-based clinical decision support Sample to Insight Powering clinical insights
Brochure Bioinformatics for Clinical Oncology Testing Complete automation for NGS interpretation and reporting with evidence-based clinical decision support Sample to Insight Powering clinical insights
TIGR THE INSTITUTE FOR GENOMIC RESEARCH
 Introduction to Genome Annotation: Overview of What You Will Learn This Week C. Robin Buell May 21, 2007 Types of Annotation Structural Annotation: Defining genes, boundaries, sequence motifs e.g. ORF,
Introduction to Genome Annotation: Overview of What You Will Learn This Week C. Robin Buell May 21, 2007 Types of Annotation Structural Annotation: Defining genes, boundaries, sequence motifs e.g. ORF,
High-throughput scale. Desktop simplicity.
 High-throughput scale. Desktop simplicity. NextSeq 500 System. Flexible power. Speed and simplicity for whole-genome, exome, and transcriptome sequencing. Harness the power of next-generation sequencing.
High-throughput scale. Desktop simplicity. NextSeq 500 System. Flexible power. Speed and simplicity for whole-genome, exome, and transcriptome sequencing. Harness the power of next-generation sequencing.
Microarray Gene Expression Analysis at CNIO
 Microarray Gene Expression Analysis at CNIO Orlando Domínguez Genomics Unit Biotechnology Program, CNIO 8 May 2013 Workflow, from samples to Gene Expression data Experimental design user/gu/ubio Samples
Microarray Gene Expression Analysis at CNIO Orlando Domínguez Genomics Unit Biotechnology Program, CNIO 8 May 2013 Workflow, from samples to Gene Expression data Experimental design user/gu/ubio Samples
Guided tour to Ensembl
 Guided tour to Ensembl Introduction Introduction to the Ensembl project Walk-through of the browser Variations and Functional Genomics Comparative Genomics BioMart Ensembl Genome browser http://www.ensembl.org
Guided tour to Ensembl Introduction Introduction to the Ensembl project Walk-through of the browser Variations and Functional Genomics Comparative Genomics BioMart Ensembl Genome browser http://www.ensembl.org
3. human genomics clone genes associated with genetic disorders. 4. many projects generate ordered clones that cover genome
 Lectures 30 and 31 Genome analysis I. Genome analysis A. two general areas 1. structural 2. functional B. genome projects a status report 1. 1 st sequenced: several viral genomes 2. mitochondria and chloroplasts
Lectures 30 and 31 Genome analysis I. Genome analysis A. two general areas 1. structural 2. functional B. genome projects a status report 1. 1 st sequenced: several viral genomes 2. mitochondria and chloroplasts
Ion S5 and Ion S5 XL Systems
 Ion S5 and Ion S5 XL Systems Targeted sequencing has never been simpler Explore the Ion S5 and Ion S5 XL Systems Adopting next-generation sequencing (NGS) in your lab is now simpler than ever The Ion S5
Ion S5 and Ion S5 XL Systems Targeted sequencing has never been simpler Explore the Ion S5 and Ion S5 XL Systems Adopting next-generation sequencing (NGS) in your lab is now simpler than ever The Ion S5
Introduction to Bioinformatics
 Introduction to Bioinformatics 260.602.01 September 1, 2006 Jonathan Pevsner, Ph.D. pevsner@kennedykrieger.org Teaching assistants Hugh Cahill (hugh@jhu.edu) Jennifer Turney (jturney@jhsph.edu) Meg Zupancic
Introduction to Bioinformatics 260.602.01 September 1, 2006 Jonathan Pevsner, Ph.D. pevsner@kennedykrieger.org Teaching assistants Hugh Cahill (hugh@jhu.edu) Jennifer Turney (jturney@jhsph.edu) Meg Zupancic
Sanger vs Next-Gen Sequencing
 Tools and Algorithms in Bioinformatics GCBA815/MCGB815/BMI815, Fall 2017 Week-8: Next-Gen Sequencing RNA-seq Data Analysis Babu Guda, Ph.D. Professor, Genetics, Cell Biology & Anatomy Director, Bioinformatics
Tools and Algorithms in Bioinformatics GCBA815/MCGB815/BMI815, Fall 2017 Week-8: Next-Gen Sequencing RNA-seq Data Analysis Babu Guda, Ph.D. Professor, Genetics, Cell Biology & Anatomy Director, Bioinformatics
Interpreting RNA-seq data (Browser Exercise II)
 Interpreting RNA-seq data (Browser Exercise II) In previous exercises, you spent some time learning about gene pages and examining genes in the context of the GBrowse genome browser. It is important to
Interpreting RNA-seq data (Browser Exercise II) In previous exercises, you spent some time learning about gene pages and examining genes in the context of the GBrowse genome browser. It is important to
Ion S5 and Ion S5 XL Systems
 Ion S5 and Ion S5 XL Systems Targeted sequencing has never been simpler Introducing the Ion S5 and Ion S5 XL systems Now, adopting next-generation sequencing in your lab is simpler than ever. The Ion S5
Ion S5 and Ion S5 XL Systems Targeted sequencing has never been simpler Introducing the Ion S5 and Ion S5 XL systems Now, adopting next-generation sequencing in your lab is simpler than ever. The Ion S5
Cancer Genetics Solutions
 Cancer Genetics Solutions Cancer Genetics Solutions Pushing the Boundaries in Cancer Genetics Cancer is a formidable foe that presents significant challenges. The complexity of this disease can be daunting
Cancer Genetics Solutions Cancer Genetics Solutions Pushing the Boundaries in Cancer Genetics Cancer is a formidable foe that presents significant challenges. The complexity of this disease can be daunting
Microarray Analysis of Gene Expression in Huntington's Disease Peripheral Blood - a Platform Comparison. CodeLink compatible
 Microarray Analysis of Gene Expression in Huntington's Disease Peripheral Blood - a Platform Comparison CodeLink compatible Microarray Analysis of Gene Expression in Huntington's Disease Peripheral Blood
Microarray Analysis of Gene Expression in Huntington's Disease Peripheral Blood - a Platform Comparison CodeLink compatible Microarray Analysis of Gene Expression in Huntington's Disease Peripheral Blood
Introduction to Next Generation Sequencing (NGS) Data Analysis and Pathway Analysis. Jenny Wu
 Introduction to Next Generation Sequencing (NGS) Data Analysis and Pathway Analysis Jenny Wu Outline Introduction to NGS data analysis in Cancer Genomics NGS applications in cancer research Typical NGS
Introduction to Next Generation Sequencing (NGS) Data Analysis and Pathway Analysis Jenny Wu Outline Introduction to NGS data analysis in Cancer Genomics NGS applications in cancer research Typical NGS
MICROARRAYS+SEQUENCING
 MICROARRAYS+SEQUENCING The most efficient way to advance genomics research Down to a Science. www.affymetrix.com/downtoascience Affymetrix GeneChip Expression Technology Complementing your Next-Generation
MICROARRAYS+SEQUENCING The most efficient way to advance genomics research Down to a Science. www.affymetrix.com/downtoascience Affymetrix GeneChip Expression Technology Complementing your Next-Generation
Accessible answers. Targeted sequencing: accelerating and amplifying answers for oncology research
 Accessible answers Targeted sequencing: accelerating and amplifying answers for oncology research Help advance precision medicine Accelerate results with Ion Torrent NGS Life without cancer. This is our
Accessible answers Targeted sequencing: accelerating and amplifying answers for oncology research Help advance precision medicine Accelerate results with Ion Torrent NGS Life without cancer. This is our
PAREXEL GENOMIC MEDICINE SERVICES. Applying genomics to enhance your drug development journey
 PAREXEL GENOMIC MEDICINE SERVICES Applying genomics to enhance your drug development journey YOUR JOURNEY. OUR MISSION. Genomic expertise to simplify the route to product approval and maximize patient
PAREXEL GENOMIC MEDICINE SERVICES Applying genomics to enhance your drug development journey YOUR JOURNEY. OUR MISSION. Genomic expertise to simplify the route to product approval and maximize patient
Towards unbiased biomarker discovery
 Towards unbiased biomarker discovery High-throughput molecular profiling technologies are routinely applied for biomarker discovery to make the drug discovery process more efficient and enable personalised
Towards unbiased biomarker discovery High-throughput molecular profiling technologies are routinely applied for biomarker discovery to make the drug discovery process more efficient and enable personalised
Non-Organic-Based Isolation of Mammalian microrna using Norgen s microrna Purification Kit
 Application Note 13 RNA Sample Preparation Non-Organic-Based Isolation of Mammalian microrna using Norgen s microrna Purification Kit B. Lam, PhD 1, P. Roberts, MSc 1 Y. Haj-Ahmad, M.Sc., Ph.D 1,2 1 Norgen
Application Note 13 RNA Sample Preparation Non-Organic-Based Isolation of Mammalian microrna using Norgen s microrna Purification Kit B. Lam, PhD 1, P. Roberts, MSc 1 Y. Haj-Ahmad, M.Sc., Ph.D 1,2 1 Norgen
Cytomics in Action: Cytokine Network Cytometry
 Cytomics in Action: Cytokine Network Cytometry Jonni S. Moore, Ph.D. Director, Clinical and Research Flow Cytometry and PathBioResource Associate Professor of Pathology & Laboratory Medicine University
Cytomics in Action: Cytokine Network Cytometry Jonni S. Moore, Ph.D. Director, Clinical and Research Flow Cytometry and PathBioResource Associate Professor of Pathology & Laboratory Medicine University
Gene Expression Analysis Superior Solutions for any Project
 Gene Expression Analysis Superior Solutions for any Project Find Your Perfect Match ArrayXS Global Array-to-Go Focussed Comprehensive: detect the whole transcriptome reliably Certified: discover exceptional
Gene Expression Analysis Superior Solutions for any Project Find Your Perfect Match ArrayXS Global Array-to-Go Focussed Comprehensive: detect the whole transcriptome reliably Certified: discover exceptional
Product selection guide Ion S5 and Ion S5 XL Systems
 Product selection guide Ion S5 and s Cancer genomics research Molecular profiling Ion AmpliSeq Ready-to-Use Panels Ion AmpliSeq Comprehensive Cancer Panel Cat. No. 4477685 Ion AmpliSeq Made-to-Order Panels
Product selection guide Ion S5 and s Cancer genomics research Molecular profiling Ion AmpliSeq Ready-to-Use Panels Ion AmpliSeq Comprehensive Cancer Panel Cat. No. 4477685 Ion AmpliSeq Made-to-Order Panels
Machine learning applications in genomics: practical issues & challenges. Yuzhen Ye School of Informatics and Computing, Indiana University
 Machine learning applications in genomics: practical issues & challenges Yuzhen Ye School of Informatics and Computing, Indiana University Reference Machine learning applications in genetics and genomics
Machine learning applications in genomics: practical issues & challenges Yuzhen Ye School of Informatics and Computing, Indiana University Reference Machine learning applications in genetics and genomics
Leonardo Mariño-Ramírez, PhD NCBI / NLM / NIH. BIOL 7210 A Computational Genomics 2/18/2015
 Leonardo Mariño-Ramírez, PhD NCBI / NLM / NIH BIOL 7210 A Computational Genomics 2/18/2015 The $1,000 genome is here! http://www.illumina.com/systems/hiseq-x-sequencing-system.ilmn Bioinformatics bottleneck
Leonardo Mariño-Ramírez, PhD NCBI / NLM / NIH BIOL 7210 A Computational Genomics 2/18/2015 The $1,000 genome is here! http://www.illumina.com/systems/hiseq-x-sequencing-system.ilmn Bioinformatics bottleneck
E2ES to Accelerate Next-Generation Genome Analysis in Clinical Research
 www.hcltech.com E2ES to Accelerate Next-Generation Genome Analysis in Clinical Research whitepaper April 2015 TABLE OF CONTENTS Introduction 3 Challenges associated with NGS data analysis 3 HCL s NGS Solution
www.hcltech.com E2ES to Accelerate Next-Generation Genome Analysis in Clinical Research whitepaper April 2015 TABLE OF CONTENTS Introduction 3 Challenges associated with NGS data analysis 3 HCL s NGS Solution
The 150+ Tomato Genome (re-)sequence Project; Lessons Learned and Potential
 The 150+ Tomato Genome (re-)sequence Project; Lessons Learned and Potential Applications Richard Finkers Researcher Plant Breeding, Wageningen UR Plant Breeding, P.O. Box 16, 6700 AA, Wageningen, The Netherlands,
The 150+ Tomato Genome (re-)sequence Project; Lessons Learned and Potential Applications Richard Finkers Researcher Plant Breeding, Wageningen UR Plant Breeding, P.O. Box 16, 6700 AA, Wageningen, The Netherlands,
Next Generation Sequencing. Target Enrichment
 Next Generation Sequencing Target Enrichment Next Generation Sequencing Your Partner in Every Step from Sample to Data NGS: Revolutionizing Genetic Analysis with Single-Molecule Resolution Next generation
Next Generation Sequencing Target Enrichment Next Generation Sequencing Your Partner in Every Step from Sample to Data NGS: Revolutionizing Genetic Analysis with Single-Molecule Resolution Next generation
Developing an Accurate and Precise Companion Diagnostic Assay for Targeted Therapies in DLBCL
 Developing an Accurate and Precise Companion Diagnostic Assay for Targeted Therapies in DLBCL James Storhoff, Ph.D. Senior Manager, Diagnostic Test Development World Cdx, Boston, Sep. 10th Molecules That
Developing an Accurate and Precise Companion Diagnostic Assay for Targeted Therapies in DLBCL James Storhoff, Ph.D. Senior Manager, Diagnostic Test Development World Cdx, Boston, Sep. 10th Molecules That
Bioinformatics Advice on Experimental Design
 Bioinformatics Advice on Experimental Design Where do I start? Please refer to the following guide to better plan your experiments for good statistical analysis, best suited for your research needs. Statistics
Bioinformatics Advice on Experimental Design Where do I start? Please refer to the following guide to better plan your experiments for good statistical analysis, best suited for your research needs. Statistics
Supplementary Fig. 1 related to Fig. 1 Clinical relevance of lncrna candidate
 Supplementary Figure Legends Supplementary Fig. 1 related to Fig. 1 Clinical relevance of lncrna candidate BC041951 in gastric cancer. (A) The flow chart for selected candidate lncrnas in 660 up-regulated
Supplementary Figure Legends Supplementary Fig. 1 related to Fig. 1 Clinical relevance of lncrna candidate BC041951 in gastric cancer. (A) The flow chart for selected candidate lncrnas in 660 up-regulated
Exercise1 ArrayExpress Archive - High-throughput sequencing example
 ArrayExpress and Atlas practical: querying and exporting gene expression data at the EBI Gabriella Rustici gabry@ebi.ac.uk This practical will introduce you to the data content and query functionality
ArrayExpress and Atlas practical: querying and exporting gene expression data at the EBI Gabriella Rustici gabry@ebi.ac.uk This practical will introduce you to the data content and query functionality
Introduction to Bioinformatics
 Introduction to Bioinformatics Alla L Lapidus, Ph.D. SPbSU St. Petersburg Term Bioinformatics Term Bioinformatics was invented by Paulien Hogeweg (Полина Хогевег) and Ben Hesper in 1970 as "the study of
Introduction to Bioinformatics Alla L Lapidus, Ph.D. SPbSU St. Petersburg Term Bioinformatics Term Bioinformatics was invented by Paulien Hogeweg (Полина Хогевег) and Ben Hesper in 1970 as "the study of
Updates to the NIH Guidelines for Research Involving Recombinant DNA Molecules (NIH Guidelines)
 Updates to the NIH Guidelines for Research Involving Recombinant DNA Molecules (NIH Guidelines) Jacqueline Corrigan-Curay, J.D. M.D. Acting Director Office of Biotechnology Activities National Institute
Updates to the NIH Guidelines for Research Involving Recombinant DNA Molecules (NIH Guidelines) Jacqueline Corrigan-Curay, J.D. M.D. Acting Director Office of Biotechnology Activities National Institute
Genetic Exercises. SNPs and Population Genetics
 Genetic Exercises SNPs and Population Genetics Single Nucleotide Polymorphisms (SNPs) in EuPathDB can be used to characterize similarities and differences within a group of isolates or that distinguish
Genetic Exercises SNPs and Population Genetics Single Nucleotide Polymorphisms (SNPs) in EuPathDB can be used to characterize similarities and differences within a group of isolates or that distinguish
Functional Genomics Overview RORY STARK PRINCIPAL BIOINFORMATICS ANALYST CRUK CAMBRIDGE INSTITUTE 18 SEPTEMBER 2017
 Functional Genomics Overview RORY STARK PRINCIPAL BIOINFORMATICS ANALYST CRUK CAMBRIDGE INSTITUTE 18 SEPTEMBER 2017 Agenda What is Functional Genomics? RNA Transcription/Gene Expression Measuring Gene
Functional Genomics Overview RORY STARK PRINCIPAL BIOINFORMATICS ANALYST CRUK CAMBRIDGE INSTITUTE 18 SEPTEMBER 2017 Agenda What is Functional Genomics? RNA Transcription/Gene Expression Measuring Gene
Course Presentation. Ignacio Medina Presentation
 Course Index Introduction Agenda Analysis pipeline Some considerations Introduction Who we are Teachers: Marta Bleda: Computational Biologist and Data Analyst at Department of Medicine, Addenbrooke's Hospital
Course Index Introduction Agenda Analysis pipeline Some considerations Introduction Who we are Teachers: Marta Bleda: Computational Biologist and Data Analyst at Department of Medicine, Addenbrooke's Hospital
Personalized Medicine
 Personalized Medicine Dr. Pablo Mentzinis, Director Government Relations, SAP SE Courtesy by Dominik Bertram, Marc von der Linden, Péter Adorján, SAP June 2016 1. Traditional medicine vs. personalized
Personalized Medicine Dr. Pablo Mentzinis, Director Government Relations, SAP SE Courtesy by Dominik Bertram, Marc von der Linden, Péter Adorján, SAP June 2016 1. Traditional medicine vs. personalized
NCBI web resources I: databases and Entrez
 NCBI web resources I: databases and Entrez Yanbin Yin Most materials are downloaded from ftp://ftp.ncbi.nih.gov/pub/education/ 1 Homework assignment 1 Two parts: Extract the gene IDs reported in table
NCBI web resources I: databases and Entrez Yanbin Yin Most materials are downloaded from ftp://ftp.ncbi.nih.gov/pub/education/ 1 Homework assignment 1 Two parts: Extract the gene IDs reported in table
EXECUTIVE SUMMARY GLOBAL GENOMICS AND BIOINFORMATICS RESEARCH INSTITUTE. May 24, 2017
 EXECUTIVE SUMMARY GLOBAL GENOMICS AND BIOINFORMATICS RESEARCH INSTITUTE May 24, 2017 Introduction to the Institute Inova Health System Foundation ( Inova ), The Rector and Visitors of the University of
EXECUTIVE SUMMARY GLOBAL GENOMICS AND BIOINFORMATICS RESEARCH INSTITUTE May 24, 2017 Introduction to the Institute Inova Health System Foundation ( Inova ), The Rector and Visitors of the University of
Maximizing opportunities towards achieving clinical success D R U G D I S C O V E R Y. Report Price Publication date
 F o r a c l e a r e r m a r k e t p e r s p e c t i v e Early Stage Drug Safety Strategies & Risk Management Maximizing opportunities towards achieving clinical success D R U G D I S C O V E R Y Report
F o r a c l e a r e r m a r k e t p e r s p e c t i v e Early Stage Drug Safety Strategies & Risk Management Maximizing opportunities towards achieving clinical success D R U G D I S C O V E R Y Report
ATIP Avenir Program 2018 Young group leader
 ATIP Avenir Program 2018 Young group leader Important dates - October 17th (4 pm) 2017 : opening of the registrations online - November 23 th 2017: deadline for the online submission, the mailing of the
ATIP Avenir Program 2018 Young group leader Important dates - October 17th (4 pm) 2017 : opening of the registrations online - November 23 th 2017: deadline for the online submission, the mailing of the
MOLECULAR BIOLOGY OF EUKARYOTES 2016 SYLLABUS
 03-442 Lectures: MWF 9:30-10:20 a.m. Doherty Hall 2105 03-742 Advanced Discussion Section: Time and place to be announced Probably Mon 4-6 p.m. or 6-8p.m.? Once we establish who is taking the advanced
03-442 Lectures: MWF 9:30-10:20 a.m. Doherty Hall 2105 03-742 Advanced Discussion Section: Time and place to be announced Probably Mon 4-6 p.m. or 6-8p.m.? Once we establish who is taking the advanced
Gene expression connectivity mapping and its application to Cat-App
 Gene expression connectivity mapping and its application to Cat-App Shu-Dong Zhang Northern Ireland Centre for Stratified Medicine University of Ulster Outline TITLE OF THE PRESENTATION Gene expression
Gene expression connectivity mapping and its application to Cat-App Shu-Dong Zhang Northern Ireland Centre for Stratified Medicine University of Ulster Outline TITLE OF THE PRESENTATION Gene expression
Applications of the Ion AmpliSeq Immune Repertoire Assay Plus TCRβ
 Applications of the Ion AmpliSeq Immune Repertoire Assay Plus TCRβ Timothy Looney, PhD Staff Scientist, Clinical Next-Generation Sequencing Division Thermo Fisher Scientific The world leader in serving
Applications of the Ion AmpliSeq Immune Repertoire Assay Plus TCRβ Timothy Looney, PhD Staff Scientist, Clinical Next-Generation Sequencing Division Thermo Fisher Scientific The world leader in serving
Genome Biology and Biotechnology
 Genome Biology and Biotechnology 10. The proteome Prof. M. Zabeau Department of Plant Systems Biology Flanders Interuniversity Institute for Biotechnology (VIB) University of Gent International course
Genome Biology and Biotechnology 10. The proteome Prof. M. Zabeau Department of Plant Systems Biology Flanders Interuniversity Institute for Biotechnology (VIB) University of Gent International course
Protein Bioinformatics Part I: Access to information
 Protein Bioinformatics Part I: Access to information 260.655 April 6, 2006 Jonathan Pevsner, Ph.D. pevsner@kennedykrieger.org Outline [1] Proteins at NCBI RefSeq accession numbers Cn3D to visualize structures
Protein Bioinformatics Part I: Access to information 260.655 April 6, 2006 Jonathan Pevsner, Ph.D. pevsner@kennedykrieger.org Outline [1] Proteins at NCBI RefSeq accession numbers Cn3D to visualize structures
Next Generation Sequencing. Jeroen Van Houdt - Leuven 13/10/2017
 Next Generation Sequencing Jeroen Van Houdt - Leuven 13/10/2017 Landmarks in DNA sequencing 1953 Discovery of DNA double helix structure 1977 A Maxam and W Gilbert "DNA seq by chemical degradation" F Sanger"DNA
Next Generation Sequencing Jeroen Van Houdt - Leuven 13/10/2017 Landmarks in DNA sequencing 1953 Discovery of DNA double helix structure 1977 A Maxam and W Gilbert "DNA seq by chemical degradation" F Sanger"DNA
Peptide libraries: applications, design options and considerations. Laura Geuss, PhD May 5, 2015, 2:00-3:00 pm EST
 Peptide libraries: applications, design options and considerations Laura Geuss, PhD May 5, 2015, 2:00-3:00 pm EST Overview 1 2 3 4 5 Introduction Peptide library basics Peptide library design considerations
Peptide libraries: applications, design options and considerations Laura Geuss, PhD May 5, 2015, 2:00-3:00 pm EST Overview 1 2 3 4 5 Introduction Peptide library basics Peptide library design considerations
Gene Expression on the Fluidigm BioMark HD
 Gene Expression on the Fluidigm BioMark HD Overview Introduction to Fluidigm James Miller Advantages of the technology Running a Fluidigm gene expression project Paul Lacaze Assay design, chemistry, experimental
Gene Expression on the Fluidigm BioMark HD Overview Introduction to Fluidigm James Miller Advantages of the technology Running a Fluidigm gene expression project Paul Lacaze Assay design, chemistry, experimental
Training materials.
 Training materials Ensembl training materials are protected by a CC BY license http://creativecommons.org/licenses/by/4.0/ If you wish to re-use these materials, please credit Ensembl for their creation
Training materials Ensembl training materials are protected by a CC BY license http://creativecommons.org/licenses/by/4.0/ If you wish to re-use these materials, please credit Ensembl for their creation
Introductory Next Gen Workshop
 Introductory Next Gen Workshop http://www.illumina.ucr.edu/ http://www.genomics.ucr.edu/ Workshop Objectives Workshop aimed at those who are new to Illumina sequencing and will provide: - a basic overview
Introductory Next Gen Workshop http://www.illumina.ucr.edu/ http://www.genomics.ucr.edu/ Workshop Objectives Workshop aimed at those who are new to Illumina sequencing and will provide: - a basic overview
Differential Gene Expression
 Biology 4361 Developmental Biology Differential Gene Expression September 28, 2006 Chromatin Structure ~140 bp ~60 bp Transcriptional Regulation: 1. Packing prevents access CH 3 2. Acetylation ( C O )
Biology 4361 Developmental Biology Differential Gene Expression September 28, 2006 Chromatin Structure ~140 bp ~60 bp Transcriptional Regulation: 1. Packing prevents access CH 3 2. Acetylation ( C O )
ALSO: look at figure 5-11 showing exonintron structure of the beta globin gene
 S08 Biology 205 6/4/08 Reading Assignment Chapter 7: From DNA to Protein: How cells read the genome pg 237-243 on exons and introns (you are not responsible for the biochemistry of splicing: figures 7-15,16
S08 Biology 205 6/4/08 Reading Assignment Chapter 7: From DNA to Protein: How cells read the genome pg 237-243 on exons and introns (you are not responsible for the biochemistry of splicing: figures 7-15,16
Lecture #1. Introduction to microarray technology
 Lecture #1 Introduction to microarray technology Outline General purpose Microarray assay concept Basic microarray experimental process cdna/two channel arrays Oligonucleotide arrays Exon arrays Comparing
Lecture #1 Introduction to microarray technology Outline General purpose Microarray assay concept Basic microarray experimental process cdna/two channel arrays Oligonucleotide arrays Exon arrays Comparing
Gene-centered resources at NCBI
 COURSE OF BIOINFORMATICS a.a. 2014-2015 Gene-centered resources at NCBI We searched Accession Number: M60495 AT NCBI Nucleotide Gene has been implemented at NCBI to organize information about genes, serving
COURSE OF BIOINFORMATICS a.a. 2014-2015 Gene-centered resources at NCBI We searched Accession Number: M60495 AT NCBI Nucleotide Gene has been implemented at NCBI to organize information about genes, serving
Build Your Own Gene Expression Analysis Panels
 Build Your Own Gene Expression Analysis Panels George J. Quellhorst, Jr. Ph.D. Associate Director, R&D Agenda What Will We Discuss? Introduction Building Your Own Gene List Getting Started Increasing Coverage
Build Your Own Gene Expression Analysis Panels George J. Quellhorst, Jr. Ph.D. Associate Director, R&D Agenda What Will We Discuss? Introduction Building Your Own Gene List Getting Started Increasing Coverage
Genetic Variability of MicroRNA Genes in 15 Animal Species
 51 Ivyspring International Publisher Research Paper Journal of Genomics 2015; 3: 51-56. doi: 10.7150/jgen.11246 Genetic Variability of MicroRNA Genes in 15 Animal Species Minja Zorc, Jana Obsteter, Peter
51 Ivyspring International Publisher Research Paper Journal of Genomics 2015; 3: 51-56. doi: 10.7150/jgen.11246 Genetic Variability of MicroRNA Genes in 15 Animal Species Minja Zorc, Jana Obsteter, Peter
BCHM 6280 Tutorial: Gene specific information using NCBI, Ensembl and genome viewers
 BCHM 6280 Tutorial: Gene specific information using NCBI, Ensembl and genome viewers Web resources: NCBI database: http://www.ncbi.nlm.nih.gov/ Ensembl database: http://useast.ensembl.org/index.html UCSC
BCHM 6280 Tutorial: Gene specific information using NCBI, Ensembl and genome viewers Web resources: NCBI database: http://www.ncbi.nlm.nih.gov/ Ensembl database: http://useast.ensembl.org/index.html UCSC
Using semantic web technology to accelerate plant breeding.
 Using semantic web technology to accelerate plant breeding. Pierre-Yves Chibon 1,2,3, Benoît Carrères 1, Heleena de Weerd 1, Richard G. F. Visser 1,2,3, and Richard Finkers 1,3 1 Wageningen UR Plant Breeding,
Using semantic web technology to accelerate plant breeding. Pierre-Yves Chibon 1,2,3, Benoît Carrères 1, Heleena de Weerd 1, Richard G. F. Visser 1,2,3, and Richard Finkers 1,3 1 Wageningen UR Plant Breeding,
The Pathways to Understanding Diseases
 FOR PHARMA & LIFE SCIENCES White paper The Pathways to Understanding Diseases Deciphering complex biological processes Executive Summary Understanding complex biological processes is critical whether doing
FOR PHARMA & LIFE SCIENCES White paper The Pathways to Understanding Diseases Deciphering complex biological processes Executive Summary Understanding complex biological processes is critical whether doing
University of Pennsylvania Perelman School of Medicine High Throughput Screening Core
 University of Pennsylvania Perelman School of Medicine High Throughput Screening Core Sara Cherry, Ph. D. David C. Schultz, Ph. D. dschultz@mail.med.upenn.edu (215) 573 9641 67 John Morgan Building Mission
University of Pennsylvania Perelman School of Medicine High Throughput Screening Core Sara Cherry, Ph. D. David C. Schultz, Ph. D. dschultz@mail.med.upenn.edu (215) 573 9641 67 John Morgan Building Mission
Microarray Analysis of Gene Expression in Huntington's Disease Peripheral Blood - a Platform Comparison
 Microarray Analysis of Gene Expression in Huntington's Disease Peripheral Blood - a Platform Comparison Thank you for waiting. The presentation will be starting in a few minutes at 9AM Pacific Daylight
Microarray Analysis of Gene Expression in Huntington's Disease Peripheral Blood - a Platform Comparison Thank you for waiting. The presentation will be starting in a few minutes at 9AM Pacific Daylight
Integration of heterogeneous omics data
 Integration of heterogeneous omics data Andrea Rau March 11, 2016 Formation doctorale: Biologie expe rimentale animale et mode lisation pre dictive andrea.rau@jouy.inra.fr Integration of heterogeneous
Integration of heterogeneous omics data Andrea Rau March 11, 2016 Formation doctorale: Biologie expe rimentale animale et mode lisation pre dictive andrea.rau@jouy.inra.fr Integration of heterogeneous
Bioinformatics of Transcriptional Regulation
 Bioinformatics of Transcriptional Regulation Carl Herrmann IPMB & DKFZ c.herrmann@dkfz.de Wechselwirkung von Maßnahmen und Auswirkungen Einflussmöglichkeiten in einem Dialog From genes to active compounds
Bioinformatics of Transcriptional Regulation Carl Herrmann IPMB & DKFZ c.herrmann@dkfz.de Wechselwirkung von Maßnahmen und Auswirkungen Einflussmöglichkeiten in einem Dialog From genes to active compounds
Comments on Use of Databases for Establishing the Clinical Relevance of Human Genetic Variants
 Division of Dockets Management (HFA-305) Food and Drug Administration 5630 Fishers Lane, Rm. 1061 Rockville, MD 20852 The American Society of Human Genetics 9650 Rockville Pike Bethesda, MD 20814 24 December
Division of Dockets Management (HFA-305) Food and Drug Administration 5630 Fishers Lane, Rm. 1061 Rockville, MD 20852 The American Society of Human Genetics 9650 Rockville Pike Bethesda, MD 20814 24 December
Themes: RNA and RNA Processing. Messenger RNA (mrna) What is a gene? RNA is very versatile! RNA-RNA interactions are very important!
 Themes: RNA is very versatile! RNA and RNA Processing Chapter 14 RNA-RNA interactions are very important! Prokaryotes and Eukaryotes have many important differences. Messenger RNA (mrna) Carries genetic
Themes: RNA is very versatile! RNA and RNA Processing Chapter 14 RNA-RNA interactions are very important! Prokaryotes and Eukaryotes have many important differences. Messenger RNA (mrna) Carries genetic
Functional analysis using EBI Metagenomics
 Functional analysis using EBI Metagenomics Contents Tutorial information... 2 Tutorial learning objectives... 2 An introduction to functional analysis using EMG... 3 What are protein signatures?... 3 Assigning
Functional analysis using EBI Metagenomics Contents Tutorial information... 2 Tutorial learning objectives... 2 An introduction to functional analysis using EMG... 3 What are protein signatures?... 3 Assigning
Thermo Scientific Dharmacon SMARTvector 2.0 Lentiviral shrna Particles
 Thermo Scientific Dharmacon SMARTvector 2.0 Lentiviral shrna Particles Long-term gene silencing shrna-specific design algorithm High titer, purified particles Thermo Scientific Dharmacon SMARTvector shrna
Thermo Scientific Dharmacon SMARTvector 2.0 Lentiviral shrna Particles Long-term gene silencing shrna-specific design algorithm High titer, purified particles Thermo Scientific Dharmacon SMARTvector shrna
New Frontiers in Personalized Medicine
 New Frontiers in Personalized Medicine Oracle Open World Shanghai 2013 Neil de Crescenzo SVP and GM 1 Safe Harbor Statement The following is intended to outline our general product direction. It is intended
New Frontiers in Personalized Medicine Oracle Open World Shanghai 2013 Neil de Crescenzo SVP and GM 1 Safe Harbor Statement The following is intended to outline our general product direction. It is intended
Assessing De-Novo Transcriptome Assemblies
 Assessing De-Novo Transcriptome Assemblies Shawn T. O Neil Center for Genome Research and Biocomputing Oregon State University Scott J. Emrich University of Notre Dame 100K Contigs, Perfect 1M Contigs,
Assessing De-Novo Transcriptome Assemblies Shawn T. O Neil Center for Genome Research and Biocomputing Oregon State University Scott J. Emrich University of Notre Dame 100K Contigs, Perfect 1M Contigs,
The Institute of Microbiology of the Czech Academy of Sciences is looking for. four POSTDOCTORAL FELLOW positions in the fields of:
 4 Postdoctoral Fellow positions in microbiology, molecular biology and ecology The Institute of Microbiology of the Czech Academy of Sciences, Prague, Czech Republic The Institute of Microbiology of the
4 Postdoctoral Fellow positions in microbiology, molecular biology and ecology The Institute of Microbiology of the Czech Academy of Sciences, Prague, Czech Republic The Institute of Microbiology of the
RNA-Seq with the Tuxedo Suite
 RNA-Seq with the Tuxedo Suite Monica Britton, Ph.D. Sr. Bioinformatics Analyst September 2015 Workshop The Basic Tuxedo Suite References Trapnell C, et al. 2009 TopHat: discovering splice junctions with
RNA-Seq with the Tuxedo Suite Monica Britton, Ph.D. Sr. Bioinformatics Analyst September 2015 Workshop The Basic Tuxedo Suite References Trapnell C, et al. 2009 TopHat: discovering splice junctions with
RNA-Seq analysis using R: Differential expression and transcriptome assembly
 RNA-Seq analysis using R: Differential expression and transcriptome assembly Beibei Chen Ph.D BICF 12/7/2016 Agenda Brief about RNA-seq and experiment design Gene oriented analysis Gene quantification
RNA-Seq analysis using R: Differential expression and transcriptome assembly Beibei Chen Ph.D BICF 12/7/2016 Agenda Brief about RNA-seq and experiment design Gene oriented analysis Gene quantification
RNA spike-in controls & analysis methods for trustworthy genome-scale measurements
 RNA spike-in controls & analysis methods for trustworthy genome-scale measurements Sarah A. Munro, Ph.D. Genome-Scale Measurements Group ABRF Meeting March 29, 2015 Overview External RNA Controls Consortium
RNA spike-in controls & analysis methods for trustworthy genome-scale measurements Sarah A. Munro, Ph.D. Genome-Scale Measurements Group ABRF Meeting March 29, 2015 Overview External RNA Controls Consortium
HCT116 SW48 Nutlin: p53
 Figure S HCT6 SW8 Nutlin: - + - + p GAPDH Figure S. Nutlin- treatment induces p protein. HCT6 and SW8 cells were left untreated or treated for 8 hr with Nutlin- ( µm) to up-regulate p. Whole cell lysates
Figure S HCT6 SW8 Nutlin: - + - + p GAPDH Figure S. Nutlin- treatment induces p protein. HCT6 and SW8 cells were left untreated or treated for 8 hr with Nutlin- ( µm) to up-regulate p. Whole cell lysates
Mapping strategies for sequence reads
 Mapping strategies for sequence reads Ernest Turro University of Cambridge 21 Oct 2013 Quantification A basic aim in genomics is working out the contents of a biological sample. 1. What distinct elements
Mapping strategies for sequence reads Ernest Turro University of Cambridge 21 Oct 2013 Quantification A basic aim in genomics is working out the contents of a biological sample. 1. What distinct elements
less sensitive than RNA-seq but more robust analysis pipelines expensive but quantitiatve standard but typically not high throughput
 Chapter 11: Gene Expression The availability of an annotated genome sequence enables massively parallel analysis of gene expression. The expression of all genes in an organism can be measured in one experiment.
Chapter 11: Gene Expression The availability of an annotated genome sequence enables massively parallel analysis of gene expression. The expression of all genes in an organism can be measured in one experiment.
Product Catalog # Description List Price (JPY) Primer Assays (desalted)
 Assays and Controls Primer Assay Pricing Primer Assays (desalted) 10025636 Primer assay desalted, 200 reactions 16,000 (Wet-lab validated human, mouse, and rat) 10025637 Primer assay desalted, 1,000 reactions
Assays and Controls Primer Assay Pricing Primer Assays (desalted) 10025636 Primer assay desalted, 200 reactions 16,000 (Wet-lab validated human, mouse, and rat) 10025637 Primer assay desalted, 1,000 reactions
ACCELERATING GENOMIC ANALYSIS ON THE CLOUD. Enabling the PanCancer Analysis of Whole Genomes (PCAWG) consortia to analyze thousands of genomes
 ACCELERATING GENOMIC ANALYSIS ON THE CLOUD Enabling the PanCancer Analysis of Whole Genomes (PCAWG) consortia to analyze thousands of genomes Enabling the PanCancer Analysis of Whole Genomes (PCAWG) consortia
ACCELERATING GENOMIC ANALYSIS ON THE CLOUD Enabling the PanCancer Analysis of Whole Genomes (PCAWG) consortia to analyze thousands of genomes Enabling the PanCancer Analysis of Whole Genomes (PCAWG) consortia
Supplementary Figure 1. Characterization of EVs (a) Phase-contrast electron microscopy was used to visualize resuspended EV pellets.
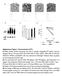 Supplementary Figure 1. Characterization of EVs (a) Phase-contrast electron microscopy was used to visualize resuspended EV pellets. Scale bar represent 100 nm. The sizes of EVs from MDA-MB-231-D3H1 (D3H1),
Supplementary Figure 1. Characterization of EVs (a) Phase-contrast electron microscopy was used to visualize resuspended EV pellets. Scale bar represent 100 nm. The sizes of EVs from MDA-MB-231-D3H1 (D3H1),
Introduction to C. elegans and RNA interference
 Introduction to C. elegans and RNA interference Why study model organisms? The problem: In order to understand biology, we need to learn about the function of the underlying genes How can we find out what
Introduction to C. elegans and RNA interference Why study model organisms? The problem: In order to understand biology, we need to learn about the function of the underlying genes How can we find out what
Package goseq. R topics documented: December 23, 2017
 Package goseq December 23, 2017 Version 1.30.0 Date 2017/09/04 Title Gene Ontology analyser for RNA-seq and other length biased data Author Matthew Young Maintainer Nadia Davidson ,
Package goseq December 23, 2017 Version 1.30.0 Date 2017/09/04 Title Gene Ontology analyser for RNA-seq and other length biased data Author Matthew Young Maintainer Nadia Davidson ,
Long and short/small RNA-seq data analysis
 Long and short/small RNA-seq data analysis GEF5, 4.9.2015 Sami Heikkinen, PhD, Dos. Topics 1. RNA-seq in a nutshell 2. Long vs short/small RNA-seq 3. Bioinformatic analysis work flows GEF5 / Heikkinen
Long and short/small RNA-seq data analysis GEF5, 4.9.2015 Sami Heikkinen, PhD, Dos. Topics 1. RNA-seq in a nutshell 2. Long vs short/small RNA-seq 3. Bioinformatic analysis work flows GEF5 / Heikkinen
1. Introduction. 1 Page 1
 1. INTRODUCTION Osteoblast differentiation is the key component of the bone formation, growth and remodeling. During this incompletely understood process a subpopulation of multipotential mesenchymal stem
1. INTRODUCTION Osteoblast differentiation is the key component of the bone formation, growth and remodeling. During this incompletely understood process a subpopulation of multipotential mesenchymal stem
Single Cell Genomics (SCG): Market Size, Segmentation, Growth, Competition and Trends ( )
 Single Cell Genomics (SCG): Market Size, Segmentation, Growth, Competition and Trends (2013-2021) May, 2017 3 rd Edition Information contained in this market report is believed to be reliable at the time
Single Cell Genomics (SCG): Market Size, Segmentation, Growth, Competition and Trends (2013-2021) May, 2017 3 rd Edition Information contained in this market report is believed to be reliable at the time
