Supplementary Materials
|
|
|
- Gavin Kristian Williamson
- 6 years ago
- Views:
Transcription
1 Supplementary Materials Construction of Synthetic Nucleoli in Human Cells Reveals How a Major Functional Nuclear Domain is Formed and Propagated Through Cell Divisision Authors: Alice Grob, Christine Colleran and Brian McStay* Centre for Chromosome Biology, School of Natural Sciences, National University of Ireland Galway, Ireland. Supplemental Materials contains: Supplemental Figures and Legends 1-12 Supplemental Fig. 1 relates to Materials and Methods Supplemental Fig. 2 and 3 relates to Fig. 1 Supplemental Fig. 4 relates to Fig. 2 Supplemental Fig. 5 and 6 relates to Fig. 3 Supplemental Fig. 7 relates to Fig. 4 Supplemental Fig. 8 and 9 relates to Fig. 4 and 5 Supplemental Fig. 10 relates to Fig. 5 Supplemental Fig. 11 relates to Fig. 6 Supplemental Fig. 12 and 13 relates to Fig. 7 Supplemental Table 1 relates to Materials and Methods Supplemental Video 1 relates to Fig. 6 1
2 A! Human 5ʼETS probe! 2.9kb NotI fragment! from 5ʼ external transcribed spacer! (+270/+3170)! IGS probe! 11.9kb Eco RI fragment from intergenic spacer! (+31497/+43464)!! q-arm BAC probes! 13q11 (AC018739)! 15q11.2 (AC068446)! 22q11.2 (AC013360)! centromere! DJ BAC probe! (CT476834)! B! The neo-nor cassette (20.4kb)! Mouse coding sequences! Human promoter! ITS1! ITS2! 3ʼETS! XEn elements! 5ʼETS! 18S! 5.8S! 28S! Transcriptional! terminator! Mouse 5ʼETS probe! 2.9kb AcuI-SalI fragment! from 5ʼ external transcribed spacer! (+120/+3100)! Mouse 28S probe! 35-mer oligonucleotide! (+11028/+11063)! XEn probe! Mouse ITS2 probe! 1.1kb PCR fragment! (+7036/+8123)! Mouse 3ʼETS probe! 0.5kb PCR fragment! (+12854/+13404)! Supplemental Figure S1. Schematic representation of hybridization probes. (A) Hybridization sites of probes used to visualize and identify individual acrocentric chromosomes (q-arm and DJ BAC), endogenous NORs (IGS) and endogenous pre-rrna (human 5 ETS). The 43972bp sequence used as a reference rdna repeat corresponds to nucleotides of BAC clone RP11-337M7 (Acc. No. AL592188). Next generation sequencing of nucleolar DNA has confirmed that this sequence is a representative rdna repeat (McStay unpublished data). (B) Location of hybridization probes used to identify neo-nors (XEn) by DNA-FISH and their derived transcripts (mouse 5 ETS, ITS2, 28S and 3 ETS) by RNA-FISH. The coordinates of mouse probes correspond to the sequence of a complete mouse rdna repeat (Acc. No. BK000964). 2!
3 * Supplemental Figure S2. UBF depletion induces the appearance of a sub-g1 cell population. (A) Western blot of proteins extracted from HT1080 cells and UBF-KD cells cultured with 1µg/ ml Dox for 48h, 96h and 144h. Molecular weight markers in kda are indicated on the left. Anti- UBF antibodies revealed a ~50-fold depletion of UBF1 (97kDa) and UBF2 (94kDa) while anti- RPA43 antibodies reveal the conserved level of pol I subunit RPA43 (43kDa). (B) FACS analysis of HT1080 cells and UBF-KD cells cultured with 1µg/ml Dox for 24h, 48h, 72h and 96h. Following 72h of culture with 1µg/ml Dox, G2/M cells decrease while sub-g1 cells accumulate (red *). 3!
4 α-ubf! IGS! Merge! Merge with DAPI! UBF-KD! +2ng/ml Dox! HT1080! Supplemental Figure S3. UBF down-regulation releases rdna repeats from the nucleolus. Combined 3D-immuno FISH performed on HT1080 and UBF-KD cells cultured with 2ng/ml Dox reveals that dissociated rdna repeats identified using an IGS probe (arrowheads) are devoid of UBF. 4!
5 B! Cont.! UBF sirna! Merge! Merge with DAPI! rdna! DAPI! rdna! DAPI! Silver! UBF sirna! UBF sirna! DAPI! Cont. sirna! D! C! Control sirna! α-ubf! UBF sirna! - UBF! α-fibrillarin! Cont. sirna! A! Supplemental Figure S4. Formation of 2 constriction is UBF-dependent in Ptk-2 cells. (A) A cdna clone encoding Ptk UBF was cloned and sequenced in order to design 3 sirnas. Western blot of proteins extracted from Ptk-2 cells transfected with control or UBF sirnas shows a 20fold UBF depletion. (B) Immuno-staining of sirna transfected Ptk-2 cells identifies the nucleolar remnants after UBF depletion. (C) Metaphase chromosome spreads prepared from control and UBF sirna transfected Ptk-2 cells were hybridized with a human 18S and 28S rdna probe. Arrowheads indicate the NOR. UBF depletion induces the loss of the 2 constriction associated with the single NOR in these cells. (D) Silver-staining (arrowheads) of metaphase spreads from Ptk-2 cells indicates that UBF depletion results in the loss of NOR silver-staining. 5!
6 *! Supplemental Figure S5. Chromosomal mapping and characterization of neo-nors. (A) FISH experiments performed on metaphase chromosome spreads prepared from each neo- NOR line classify them into acrocentric (a1-a3) and metacentric (m1-m3) clones. XEn probe labels neo-nors, while a human IGS probe identifies endogenous NORs. BAC clones RP11-42OH1 (AC18739), RP11-32B5 (AC068446) and RP11-278E23 (AC013360) identify respectively the q-arms of acrocentric chromosomes 13, 15 and 22. Chromosome painting with chromosome paints prepared from chromosome specific DOP-PCR products (kind gift from M. and T. Cremer) identify metacentric chromosome 4. (B) The size and integrity of each neo- NOR was determined by Southern blotting. Genomic DNA from HT1080 and neo-nor cell lines were digested to release neo-nor transcription units and probed with sequences derived from the mouse 5 ETS. Similarly digested pneo-nor plasmid served as a size marker. Note the high degree of sequence rearrangement. (C) Table indicating the chromosomal location and the estimated copy number of intact transcription units integrated in neo-nor lines. (D) Silver-staining of metaphase spreads from pseudo-nor clone 3D and neo-nor clone m1. Pseudo-NOR and neo-nor are indicated by arrowheads. Note the silver-staining of an endogenous NOR (*) in the lower right panel. 6!
7 Supplemental Figure S6. Transcription across the neo-nor cassette. (A) RNA FISH experiments performed on mouse 3T3 and human HT1080 cells demonstrate the specificity of mouse (green) and human (red) 5 ETS probes and identify neo-nor-derived transcripts in neo-nor clones a1-a3, m2 and m3. (B) RNA FISH using mouse ITS2 probe identifies neo-nor-derived transcripts in clone m1. (C) RNA FISH using mouse 3 ETS probe identifies neo-nor-derived transcripts in clones a1-a3 and m1-m3. 7!
8 Supplemental Figure S7. Neo-NORs recruit FC, DFC and GC components to build neonucleoli. Combined 3D-immuno RNA FISH reveals that FC/DFC factors coupling rdna transcription to pre-rrna processing, t-utp10 and Treacle, together with DFC factors, Nap57 and U3-55K, and GC factor Nucleolin, colocalize with neo-nor-derived transcripts in clone m1. 8!
9 Supplemental Figure S8. Efficient Maturation of neo-nor-derived pre-rrna. (A) Quantitation of the S1 nuclease protection assays presented in Fig. 5A. Note the similar ratio between cleaved and uncleaved transcripts in mouse and neo-nor lines. (B) Table indicating the estimated percentage of neo-nor-derived 28S rrnas among different RNA fractions extracted from the 6 neo-nor clones. 9!
10 Supplemental Figure S9. Probing strategy for detection of neo-nor-derived 18S and 28S rrnas. (A) The upper panel correspond to a schematic representation of the species specific primers used for RT-PCR based identification of mouse 18S rrna. The gel illustrates the specificity of these primers for mouse 18S rrna in RT-PCR. (B) Schematic representation of the oligonucleotide used to probe neo-nor-derived 28S rrnas. 10!
11 HT1080! A! WCE! N! PMT!R! 62! 49! 38! 28! 14! * * Rib. proteins! Purification steps! B! Polysomes! 60S! 40S! 28S! 18S! direction of sedimentation! Supplemental Figure S10. Ribosome and polysome purification. (A) Coomassiestained SDS protein gel of whole cell extract (WCE), nuclear (N), post mitochondrial (PMT) and ribosome (R) fractions prepared from HT1080 cells. The low-molecularweight histone bands (*) and the size range of ribosomal proteins are indicated. (B) Polysomal fractions were identified by electrophoresis of RNA aliquots extracted from polysome sucrose gradient fractions. The example shown is from clone m1. Polysome, small 40S and large 60S ribosome subunit fractions are indicated. 11!
12 Supplemental Figure S11. Nucleolar segregation upon Actinomycin D (AMD) treatment. AMD treatment reveals the compartmentalized structure of nucleoli as illustrated in cartoon form (upper panel). FC/DFC regions are visualized with UBF antibodies (green), while GC regions are visualized with Nop52 antibodies (red). 12!
13 Control sirna! Control sirna! α-ubf! XEn! α-fib! XEn! XEn! XEn! UBF sirna! UBF sirna! α-ubf! XEn! α-fib! XEn! XEn! XEn! Supplemental Figure S12. UBF depletion induces neo-nor condensation and loss of neonucleoli. Combined 3D-immuno FISH experiments performed on clone m1 cells transfected with control or UBF sirnas reveal the condensation of neo-nors (XEn probe) and their failure to form neo-nucleoli recruiting UBF and Fibrillarin (Fib) upon UBF sirna knockdown. Images shown are restricted to the nuclear region containing neo-nors. 13!
14 A! M-5ʼETS! H-5ʼETS! UBF sirna! Control sirna! B! Percentage of cells! with active neo-nor! #!!" +!" *!" )!" (!" '!" &!" %!" $!" #!"!",-./0-1"23456" 789"23456" Supplemental Figure S13. UBF depletion silences neo-nor transcription. (A) RNA FISH performed on clone m1 cells transfected with control or UBF sirnas. Mouse and human 5 ETS probes (M- and H-5 ETS respectively) reveal the lost of neo-nor transcription upon UBF sirna knockdown. (B) A graph representing the percentage of cells with transcriptionally active neo-nors in clone m1 cells transfected with control or UBF sirna. 14!
15 Supplemental Table S1. Oligonucleotide probes. List and sequences of the oligonucleotide probes used for 3D-immuno RNA FISH, S1 nuclease protection assays and Northern blotting. Note that the probes testing endogenous human and ectopic neo-nor rdna transcription start (+1) correspond respectively to nucleotides -18/+42 and -19/ !
16 Supplemental Video S1: neo-nor territory. A video showing the 3D arrangement of neo- NOR organization within large mature nucleoli. 3D-RNA FISH was performed on neo-nor clone m1. Neo-NOR derived transcripts were detected using a mouse 5 ETS probe (green) transcripts from endogenous NORs were detected using a human 5 ETS probe (red). This video was prepared from 3D images of deconvolved Z-stacks rotated to create a series of bookmarks. The movie is an animation of the transition between these bookmarks. 16!
Molecular Cell Biology - Problem Drill 11: Recombinant DNA
 Molecular Cell Biology - Problem Drill 11: Recombinant DNA Question No. 1 of 10 1. Which of the following statements about the sources of DNA used for molecular cloning is correct? Question #1 (A) cdna
Molecular Cell Biology - Problem Drill 11: Recombinant DNA Question No. 1 of 10 1. Which of the following statements about the sources of DNA used for molecular cloning is correct? Question #1 (A) cdna
Contents... vii. List of Figures... xii. List of Tables... xiv. Abbreviatons... xv. Summary... xvii. 1. Introduction In vitro evolution...
 vii Contents Contents... vii List of Figures... xii List of Tables... xiv Abbreviatons... xv Summary... xvii 1. Introduction...1 1.1 In vitro evolution... 1 1.2 Phage Display Technology... 3 1.3 Cell surface
vii Contents Contents... vii List of Figures... xii List of Tables... xiv Abbreviatons... xv Summary... xvii 1. Introduction...1 1.1 In vitro evolution... 1 1.2 Phage Display Technology... 3 1.3 Cell surface
Figure S1. Figure S2. Figure S3 HB Anti-FSP27 (COOH-terminal peptide) Ab. Anti-GST-FSP27(45-127) Ab.
 / 36B4 mrna ratio Figure S1 * 2. 1.6 1.2.8 *.4 control TNFα BRL49653 Figure S2 Su bw AT p iw Anti- (COOH-terminal peptide) Ab Blot : Anti-GST-(45-127) Ab β-actin Figure S3 HB2 HW AT BA T Figure S4 A TAG
/ 36B4 mrna ratio Figure S1 * 2. 1.6 1.2.8 *.4 control TNFα BRL49653 Figure S2 Su bw AT p iw Anti- (COOH-terminal peptide) Ab Blot : Anti-GST-(45-127) Ab β-actin Figure S3 HB2 HW AT BA T Figure S4 A TAG
Computational Biology I LSM5191
 Computational Biology I LSM5191 Lecture 5 Notes: Genetic manipulation & Molecular Biology techniques Broad Overview of: Enzymatic tools in Molecular Biology Gel electrophoresis Restriction mapping DNA
Computational Biology I LSM5191 Lecture 5 Notes: Genetic manipulation & Molecular Biology techniques Broad Overview of: Enzymatic tools in Molecular Biology Gel electrophoresis Restriction mapping DNA
pdsipher and pdsipher -GFP shrna Vector User s Guide
 pdsipher and pdsipher -GFP shrna Vector User s Guide NOTE: PLEASE READ THE ENTIRE PROTOCOL CAREFULLY BEFORE USE Page 1. Introduction... 1 2. Vector Overview... 1 3. Vector Maps 2 4. Materials Provided...
pdsipher and pdsipher -GFP shrna Vector User s Guide NOTE: PLEASE READ THE ENTIRE PROTOCOL CAREFULLY BEFORE USE Page 1. Introduction... 1 2. Vector Overview... 1 3. Vector Maps 2 4. Materials Provided...
2054, Chap. 14, page 1
 2054, Chap. 14, page 1 I. Recombinant DNA technology (Chapter 14) A. recombinant DNA technology = collection of methods used to perform genetic engineering 1. genetic engineering = deliberate modification
2054, Chap. 14, page 1 I. Recombinant DNA technology (Chapter 14) A. recombinant DNA technology = collection of methods used to perform genetic engineering 1. genetic engineering = deliberate modification
XactEdit Cas9 Nuclease with NLS User Manual
 XactEdit Cas9 Nuclease with NLS User Manual An RNA-guided recombinant endonuclease for efficient targeted DNA cleavage Catalog Numbers CE1000-50K, CE1000-50, CE1000-250, CE1001-250, CE1001-1000 Table of
XactEdit Cas9 Nuclease with NLS User Manual An RNA-guided recombinant endonuclease for efficient targeted DNA cleavage Catalog Numbers CE1000-50K, CE1000-50, CE1000-250, CE1001-250, CE1001-1000 Table of
SUPPLEMENTARY INFORMATION. Transcriptional output transiently spikes upon mitotic exit
 SUPPLEMENTARY INFORMATION Transcriptional output transiently spikes upon mitotic exit Viola Vaňková Hausnerová 1, 2, Christian Lanctôt 1* 1 BIOCEV and Department of Cell Biology, Faculty of Science, Charles
SUPPLEMENTARY INFORMATION Transcriptional output transiently spikes upon mitotic exit Viola Vaňková Hausnerová 1, 2, Christian Lanctôt 1* 1 BIOCEV and Department of Cell Biology, Faculty of Science, Charles
CHAPTER 9 DNA Technologies
 CHAPTER 9 DNA Technologies Recombinant DNA Artificially created DNA that combines sequences that do not occur together in the nature Basis of much of the modern molecular biology Molecular cloning of genes
CHAPTER 9 DNA Technologies Recombinant DNA Artificially created DNA that combines sequences that do not occur together in the nature Basis of much of the modern molecular biology Molecular cloning of genes
Lecture Four. Molecular Approaches I: Nucleic Acids
 Lecture Four. Molecular Approaches I: Nucleic Acids I. Recombinant DNA and Gene Cloning Recombinant DNA is DNA that has been created artificially. DNA from two or more sources is incorporated into a single
Lecture Four. Molecular Approaches I: Nucleic Acids I. Recombinant DNA and Gene Cloning Recombinant DNA is DNA that has been created artificially. DNA from two or more sources is incorporated into a single
Multiple choice questions (numbers in brackets indicate the number of correct answers)
 1 Multiple choice questions (numbers in brackets indicate the number of correct answers) February 1, 2013 1. Ribose is found in Nucleic acids Proteins Lipids RNA DNA (2) 2. Most RNA in cells is transfer
1 Multiple choice questions (numbers in brackets indicate the number of correct answers) February 1, 2013 1. Ribose is found in Nucleic acids Proteins Lipids RNA DNA (2) 2. Most RNA in cells is transfer
Learning Objectives. Define RNA interference. Define basic terminology. Describe molecular mechanism. Define VSP and relevance
 Learning Objectives Define RNA interference Define basic terminology Describe molecular mechanism Define VSP and relevance Describe role of RNAi in antigenic variation A Nobel Way to Regulate Gene Expression
Learning Objectives Define RNA interference Define basic terminology Describe molecular mechanism Define VSP and relevance Describe role of RNAi in antigenic variation A Nobel Way to Regulate Gene Expression
NAME TA SEC Problem Set 4 FRIDAY October 15, Answers to this problem set must be inserted into the box outside
 MIT Biology Department 7.012: Introductory Biology - Fall 2004 Instructors: Professor Eric Lander, Professor Robert A. Weinberg, Dr. Claudette Gardel NAME TA SEC 7.012 Problem Set 4 FRIDAY October 15,
MIT Biology Department 7.012: Introductory Biology - Fall 2004 Instructors: Professor Eric Lander, Professor Robert A. Weinberg, Dr. Claudette Gardel NAME TA SEC 7.012 Problem Set 4 FRIDAY October 15,
Construction of plant complementation vector and generation of transgenic plants
 MATERIAL S AND METHODS Plant materials and growth conditions Arabidopsis ecotype Columbia (Col0) was used for this study. SALK_072009, SALK_076309, and SALK_027645 were obtained from the Arabidopsis Biological
MATERIAL S AND METHODS Plant materials and growth conditions Arabidopsis ecotype Columbia (Col0) was used for this study. SALK_072009, SALK_076309, and SALK_027645 were obtained from the Arabidopsis Biological
Supplementary Figure 1. RAD51 and RAD51 paralogs are enriched spontaneously onto
 Supplementary Figure legends Supplementary Figure 1. and paralogs are enriched spontaneously onto the S-phase chromatin during DN replication. () Chromatin fractionation was carried out as described in
Supplementary Figure legends Supplementary Figure 1. and paralogs are enriched spontaneously onto the S-phase chromatin during DN replication. () Chromatin fractionation was carried out as described in
GFP CCD2 GFP IP:GFP
 D1 D2 1 75 95 148 178 492 GFP CCD1 CCD2 CCD2 GFP D1 D2 GFP D1 D2 Beclin 1 IB:GFP IP:GFP Supplementary Figure 1: Mapping domains required for binding to HEK293T cells are transfected with EGFP-tagged mutant
D1 D2 1 75 95 148 178 492 GFP CCD1 CCD2 CCD2 GFP D1 D2 GFP D1 D2 Beclin 1 IB:GFP IP:GFP Supplementary Figure 1: Mapping domains required for binding to HEK293T cells are transfected with EGFP-tagged mutant
Supplementary Figure S1. Immunodetection of full-length XA21 and the XA21 C-terminal cleavage product.
 Supplementary Information Supplementary Figure S1. Immunodetection of full-length XA21 and the XA21 C-terminal cleavage product. Total protein extracted from Kitaake wild type and rice plants carrying
Supplementary Information Supplementary Figure S1. Immunodetection of full-length XA21 and the XA21 C-terminal cleavage product. Total protein extracted from Kitaake wild type and rice plants carrying
A 5 P degradation hot spot influences molecular farming of anticancerogenic nuclease TBN1 in tobacco cells
 Plant Cell, Tissue and Organ Culture (PCTOC) A 5 P degradation hot spot influences molecular farming of anticancerogenic nuclease TBN1 in tobacco cells Anna Týcová a,b, Rajen J. J. Piernikarczyk c, Michael
Plant Cell, Tissue and Organ Culture (PCTOC) A 5 P degradation hot spot influences molecular farming of anticancerogenic nuclease TBN1 in tobacco cells Anna Týcová a,b, Rajen J. J. Piernikarczyk c, Michael
Confocal immunofluorescence microscopy
 Confocal immunofluorescence microscopy HL-6 and cells were cultured and cytospun onto glass slides. The cells were double immunofluorescence stained for Mt NPM1 and fibrillarin (nucleolar marker). Briefly,
Confocal immunofluorescence microscopy HL-6 and cells were cultured and cytospun onto glass slides. The cells were double immunofluorescence stained for Mt NPM1 and fibrillarin (nucleolar marker). Briefly,
Chromosomes. Ms. Gunjan M. Chaudhari
 Chromosomes Ms. Gunjan M. Chaudhari Chromsomes Chromosome structure Chromatin structure Chromosome variations The new cytogenetics Prokaryotic chromosomes Circular double helix Complexed with protein in
Chromosomes Ms. Gunjan M. Chaudhari Chromsomes Chromosome structure Chromatin structure Chromosome variations The new cytogenetics Prokaryotic chromosomes Circular double helix Complexed with protein in
Bacterial DNA replication
 Bacterial DNA replication Summary: What problems do these proteins solve? Tyr OH attacks PO4 and forms a covalent intermediate Structural changes in the protein open the gap by 20 Å! 1 Summary: What problems
Bacterial DNA replication Summary: What problems do these proteins solve? Tyr OH attacks PO4 and forms a covalent intermediate Structural changes in the protein open the gap by 20 Å! 1 Summary: What problems
NUCLEUS. Fig. 2. Various stages in the condensation of chromatin
 NUCLEUS Animal cells contain DNA in nucleus (contains ~ 98% of cell DNA) and mitochondrion. Both compartments are surrounded by an envelope (double membrane). Nuclear DNA represents some linear molecules
NUCLEUS Animal cells contain DNA in nucleus (contains ~ 98% of cell DNA) and mitochondrion. Both compartments are surrounded by an envelope (double membrane). Nuclear DNA represents some linear molecules
Thyroid peroxidase gene expression is induced by lipopolysaccharide involving Nuclear Factor (NF)-κB p65 subunit phosphorylation
 1 2 3 4 5 SUPPLEMENTAL DATA Thyroid peroxidase gene expression is induced by lipopolysaccharide involving Nuclear Factor (NF)-κB p65 subunit phosphorylation Magalí Nazar, Juan Pablo Nicola, María Laura
1 2 3 4 5 SUPPLEMENTAL DATA Thyroid peroxidase gene expression is induced by lipopolysaccharide involving Nuclear Factor (NF)-κB p65 subunit phosphorylation Magalí Nazar, Juan Pablo Nicola, María Laura
Roche Molecular Biochemicals Technical Note No. LC 10/2000
 Roche Molecular Biochemicals Technical Note No. LC 10/2000 LightCycler Overview of LightCycler Quantification Methods 1. General Introduction Introduction Content Definitions This Technical Note will introduce
Roche Molecular Biochemicals Technical Note No. LC 10/2000 LightCycler Overview of LightCycler Quantification Methods 1. General Introduction Introduction Content Definitions This Technical Note will introduce
ASPP1 Fw GGTTGGGAATCCACGTGTTG ASPP1 Rv GCCATATCTTGGAGCTCTGAGAG
 Supplemental Materials and Methods Plasmids: the following plasmids were used in the supplementary data: pwzl-myc- Lats2 (Aylon et al, 2006), pretrosuper-vector and pretrosuper-shp53 (generous gift of
Supplemental Materials and Methods Plasmids: the following plasmids were used in the supplementary data: pwzl-myc- Lats2 (Aylon et al, 2006), pretrosuper-vector and pretrosuper-shp53 (generous gift of
Supporting Online Material for
 www.sciencemag.org/cgi/content/full/1137999/dc1 Supporting Online Material for Disrupting the Pairing Between let-7 and Enhances Oncogenic Transformation Christine Mayr, Michael T. Hemann, David P. Bartel*
www.sciencemag.org/cgi/content/full/1137999/dc1 Supporting Online Material for Disrupting the Pairing Between let-7 and Enhances Oncogenic Transformation Christine Mayr, Michael T. Hemann, David P. Bartel*
SANTA CRUZ BIOTECHNOLOGY, INC.
 TECHNICAL SERVICE GUIDE: Western Blotting 2. What size bands were expected and what size bands were detected? 3. Was the blot blank or was a dark background or non-specific bands seen? 4. Did this same
TECHNICAL SERVICE GUIDE: Western Blotting 2. What size bands were expected and what size bands were detected? 3. Was the blot blank or was a dark background or non-specific bands seen? 4. Did this same
Quantitative Real Time PCR USING SYBR GREEN
 Quantitative Real Time PCR USING SYBR GREEN SYBR Green SYBR Green is a cyanine dye that binds to double stranded DNA. When it is bound to D.S. DNA it has a much greater fluorescence than when bound to
Quantitative Real Time PCR USING SYBR GREEN SYBR Green SYBR Green is a cyanine dye that binds to double stranded DNA. When it is bound to D.S. DNA it has a much greater fluorescence than when bound to
Supplementary Information
 Journal : Nature Biotechnology Supplementary Information Targeted genome engineering in human cells with RNA-guided endonucleases Seung Woo Cho, Sojung Kim, Jong Min Kim, and Jin-Soo Kim* National Creative
Journal : Nature Biotechnology Supplementary Information Targeted genome engineering in human cells with RNA-guided endonucleases Seung Woo Cho, Sojung Kim, Jong Min Kim, and Jin-Soo Kim* National Creative
Supplementary Fig. 1 related to Fig. 1 Clinical relevance of lncrna candidate
 Supplementary Figure Legends Supplementary Fig. 1 related to Fig. 1 Clinical relevance of lncrna candidate BC041951 in gastric cancer. (A) The flow chart for selected candidate lncrnas in 660 up-regulated
Supplementary Figure Legends Supplementary Fig. 1 related to Fig. 1 Clinical relevance of lncrna candidate BC041951 in gastric cancer. (A) The flow chart for selected candidate lncrnas in 660 up-regulated
Supplementary Fig. 1. Characteristics of transcription elongation by YonO. a. YonO forms a saltstable EC. Immobilized ECs were washed with
 Supplementary Fig. 1. Characteristics of transcription elongation by YonO. a. YonO forms a saltstable EC. Immobilized ECs were washed with transcription buffer with or without a high salt concentration
Supplementary Fig. 1. Characteristics of transcription elongation by YonO. a. YonO forms a saltstable EC. Immobilized ECs were washed with transcription buffer with or without a high salt concentration
Coleman et al., Supplementary Figure 1
 Coleman et al., Supplementary Figure 1 BrdU Merge G1 Early S Mid S Supplementary Figure 1. Sequential destruction of CRL4 Cdt2 targets during the G1/S transition. HCT116 cells were synchronized by sequential
Coleman et al., Supplementary Figure 1 BrdU Merge G1 Early S Mid S Supplementary Figure 1. Sequential destruction of CRL4 Cdt2 targets during the G1/S transition. HCT116 cells were synchronized by sequential
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION Dynamic Phosphorylation of HP1 Regulates Mitotic Progression in Human Cells Supplementary Figures Supplementary Figure 1. NDR1 interacts with HP1. (a) Immunoprecipitation using
SUPPLEMENTARY INFORMATION Dynamic Phosphorylation of HP1 Regulates Mitotic Progression in Human Cells Supplementary Figures Supplementary Figure 1. NDR1 interacts with HP1. (a) Immunoprecipitation using
Supplemental Data. Regulating Gene Expression. through RNA Nuclear Retention
 Supplemental Data Regulating Gene Expression through RNA Nuclear Retention Kannanganattu V. Prasanth, Supriya G. Prasanth, Zhenyu Xuan, Stephen Hearn, Susan M. Freier, C. Frank Bennett, Michael Q. Zhang,
Supplemental Data Regulating Gene Expression through RNA Nuclear Retention Kannanganattu V. Prasanth, Supriya G. Prasanth, Zhenyu Xuan, Stephen Hearn, Susan M. Freier, C. Frank Bennett, Michael Q. Zhang,
A novel two-step genome editing strategy with CRISPR-Cas9 provides new insights into telomerase action and TERT gene expression
 Xi et al. Genome Biology (2015) 16:231 DOI 10.1186/s13059-015-0791-1 RESEARCH A novel two-step genome editing strategy with CRISPR-Cas9 provides new insights into telomerase action and TERT gene expression
Xi et al. Genome Biology (2015) 16:231 DOI 10.1186/s13059-015-0791-1 RESEARCH A novel two-step genome editing strategy with CRISPR-Cas9 provides new insights into telomerase action and TERT gene expression
The Human Protein PRR14 Tethers Heterochromatin to the Nuclear Lamina During Interphase and Mitotic Exit
 Cell Reports, Volume 5 Supplemental Information The Human Protein PRR14 Tethers Heterochromatin to the Nuclear Lamina During Interphase and Mitotic Exit Andrey Poleshko, Katelyn M. Mansfield, Caroline
Cell Reports, Volume 5 Supplemental Information The Human Protein PRR14 Tethers Heterochromatin to the Nuclear Lamina During Interphase and Mitotic Exit Andrey Poleshko, Katelyn M. Mansfield, Caroline
supplementary information
 DOI: 1.138/ncb1839 a b Control 1 2 3 Control 1 2 3 Fbw7 Smad3 1 2 3 4 1 2 3 4 c d IGF-1 IGF-1Rβ IGF-1Rβ-P Control / 1 2 3 4 Real-time RT-PCR Relative quantity (IGF-1/ mrna) 2 1 IGF-1 1 2 3 4 Control /
DOI: 1.138/ncb1839 a b Control 1 2 3 Control 1 2 3 Fbw7 Smad3 1 2 3 4 1 2 3 4 c d IGF-1 IGF-1Rβ IGF-1Rβ-P Control / 1 2 3 4 Real-time RT-PCR Relative quantity (IGF-1/ mrna) 2 1 IGF-1 1 2 3 4 Control /
3I03 - Eukaryotic Genetics Repetitive DNA
 Repetitive DNA Satellite DNA Minisatellite DNA Microsatellite DNA Transposable elements LINES, SINES and other retrosequences High copy number genes (e.g. ribosomal genes, histone genes) Multifamily member
Repetitive DNA Satellite DNA Minisatellite DNA Microsatellite DNA Transposable elements LINES, SINES and other retrosequences High copy number genes (e.g. ribosomal genes, histone genes) Multifamily member
Supplementary Figure 1: MYCER protein expressed from the transgene can enhance
 Relative luciferase activity Relative luciferase activity MYC is a critical target FBXW7 MYC Supplementary is a critical Figures target 1-7. FBXW7 Supplementary Material A E-box sequences 1 2 3 4 5 6 HSV-TK
Relative luciferase activity Relative luciferase activity MYC is a critical target FBXW7 MYC Supplementary is a critical Figures target 1-7. FBXW7 Supplementary Material A E-box sequences 1 2 3 4 5 6 HSV-TK
Figure S1: NUN preparation yields nascent, unadenylated RNA with a different profile from Total RNA.
 Summary of Supplemental Information Figure S1: NUN preparation yields nascent, unadenylated RNA with a different profile from Total RNA. Figure S2: rrna removal procedure is effective for clearing out
Summary of Supplemental Information Figure S1: NUN preparation yields nascent, unadenylated RNA with a different profile from Total RNA. Figure S2: rrna removal procedure is effective for clearing out
Supplementary Figures Montero et al._supplementary Figure 1
 Montero et al_suppl. Info 1 Supplementary Figures Montero et al._supplementary Figure 1 Montero et al_suppl. Info 2 Supplementary Figure 1. Transcripts arising from the structurally conserved subtelomeres
Montero et al_suppl. Info 1 Supplementary Figures Montero et al._supplementary Figure 1 Montero et al_suppl. Info 2 Supplementary Figure 1. Transcripts arising from the structurally conserved subtelomeres
Sensitivity vs Specificity
 Viral Detection Animal Inoculation Culturing the Virus Definitive Length of time Serology Detecting antibodies to the infectious agent Detecting Viral Proteins Western Blot ELISA Detecting the Viral Genome
Viral Detection Animal Inoculation Culturing the Virus Definitive Length of time Serology Detecting antibodies to the infectious agent Detecting Viral Proteins Western Blot ELISA Detecting the Viral Genome
Microbial Diversity and Assessment (III) Spring, 2007 Guangyi Wang, Ph.D. POST103B
 Microbial Diversity and Assessment (III) Spring, 2007 Guangyi Wang, Ph.D. POST103B guangyi@hawaii.edu http://www.soest.hawaii.edu/marinefungi/ocn403webpage.htm Overview of Last Lecture Taxonomy (three
Microbial Diversity and Assessment (III) Spring, 2007 Guangyi Wang, Ph.D. POST103B guangyi@hawaii.edu http://www.soest.hawaii.edu/marinefungi/ocn403webpage.htm Overview of Last Lecture Taxonomy (three
Genome Sequence Assembly
 Genome Sequence Assembly Learning Goals: Introduce the field of bioinformatics Familiarize the student with performing sequence alignments Understand the assembly process in genome sequencing Introduction:
Genome Sequence Assembly Learning Goals: Introduce the field of bioinformatics Familiarize the student with performing sequence alignments Understand the assembly process in genome sequencing Introduction:
Enzymatic assembly of DNA molecules up to several hundred kilobases
 nature methods Enzymatic assembly of DNA molecules up to several hundred kilobases Daniel G Gibson, Lei Young, Ray-Yuan Chuang, J Craig Venter, Clyde A Hutchison III & Hamilton O Smith Supplementary figures
nature methods Enzymatic assembly of DNA molecules up to several hundred kilobases Daniel G Gibson, Lei Young, Ray-Yuan Chuang, J Craig Venter, Clyde A Hutchison III & Hamilton O Smith Supplementary figures
1. Cross-linking and cell harvesting
 ChIP is a powerful tool that allows the specific matching of proteins or histone modifications to regions of the genome. Chromatin is isolated and antibodies to the antigen of interest are used to determine
ChIP is a powerful tool that allows the specific matching of proteins or histone modifications to regions of the genome. Chromatin is isolated and antibodies to the antigen of interest are used to determine
Supplementary information to accompany: A novel role for the DNA repair gene Rad51 in Netrin-1 signalling
 Supplementary information to accompany: A novel role for the DNA repair gene Rad51 in Netrin-1 signalling Glendining KA 1, Markie D 2, Gardner RJM 4, Franz EA 3, Robertson SP 4, Jasoni CL 1 Supplementary
Supplementary information to accompany: A novel role for the DNA repair gene Rad51 in Netrin-1 signalling Glendining KA 1, Markie D 2, Gardner RJM 4, Franz EA 3, Robertson SP 4, Jasoni CL 1 Supplementary
Supplemental Materials. Matrix Proteases Contribute to Progression of Pelvic Organ Prolapse in Mice and Humans
 Supplemental Materials Matrix Proteases Contribute to Progression of Pelvic Organ Prolapse in Mice and Humans Madhusudhan Budatha, Shayzreen Roshanravan, Qian Zheng, Cecilia Weislander, Shelby L. Chapman,
Supplemental Materials Matrix Proteases Contribute to Progression of Pelvic Organ Prolapse in Mice and Humans Madhusudhan Budatha, Shayzreen Roshanravan, Qian Zheng, Cecilia Weislander, Shelby L. Chapman,
ENCODE RBP Antibody Characterization Guidelines
 ENCODE RBP Antibody Characterization Guidelines Approved on November 18, 2016 Background An integral part of the ENCODE Project is to characterize the antibodies used in the experiments. This document
ENCODE RBP Antibody Characterization Guidelines Approved on November 18, 2016 Background An integral part of the ENCODE Project is to characterize the antibodies used in the experiments. This document
SUPPLEMENTAL MATERIALS
 SUPPLEMENL MERILS Eh-seq: RISPR epitope tagging hip-seq of DN-binding proteins Daniel Savic, E. hristopher Partridge, Kimberly M. Newberry, Sophia. Smith, Sarah K. Meadows, rian S. Roberts, Mark Mackiewicz,
SUPPLEMENL MERILS Eh-seq: RISPR epitope tagging hip-seq of DN-binding proteins Daniel Savic, E. hristopher Partridge, Kimberly M. Newberry, Sophia. Smith, Sarah K. Meadows, rian S. Roberts, Mark Mackiewicz,
RNA Structure and the Versatility of RNA. Mitesh Shrestha
 RNA Structure and the Versatility of RNA Mitesh Shrestha Ribonucleic Acid (RNA) Nitrogenous Bases (Adenine, Uracil, Guanine, Cytosine) Ribose Sugar Ribonucleic Acid (RNA) Phosphate Group RNA world Hypothesis
RNA Structure and the Versatility of RNA Mitesh Shrestha Ribonucleic Acid (RNA) Nitrogenous Bases (Adenine, Uracil, Guanine, Cytosine) Ribose Sugar Ribonucleic Acid (RNA) Phosphate Group RNA world Hypothesis
Viral RNAi suppressor reversibly binds sirna to. outcompete Dicer and RISC via multiple-turnover
 Supplementary Data Viral RNAi suppressor reversibly binds sirna to outcompete Dicer and RISC via multiple-turnover Renata A. Rawlings 1,2, Vishalakshi Krishnan 2 and Nils G. Walter 2 * 1 Biophysics and
Supplementary Data Viral RNAi suppressor reversibly binds sirna to outcompete Dicer and RISC via multiple-turnover Renata A. Rawlings 1,2, Vishalakshi Krishnan 2 and Nils G. Walter 2 * 1 Biophysics and
Rapid Learning Center Presents. Teach Yourself AP Biology in 24 Hours
 Rapid Learning Center Chemistry :: Biology :: Physics :: Math Rapid Learning Center Presents Teach Yourself AP Biology in 24 Hours 1/35 *AP is a registered trademark of the College Board, which does not
Rapid Learning Center Chemistry :: Biology :: Physics :: Math Rapid Learning Center Presents Teach Yourself AP Biology in 24 Hours 1/35 *AP is a registered trademark of the College Board, which does not
2. In Figure 10-4, why is edna made only from mrna and not also from trnas and ribosomal RNAs?
 2. In Figure 10-4, why is edna made only from mrna and not also from trnas and ribosomal RNAs? Answer: edna is made from mrna and not from trnas or rrnas because polyt primers are used to prime the first
2. In Figure 10-4, why is edna made only from mrna and not also from trnas and ribosomal RNAs? Answer: edna is made from mrna and not from trnas or rrnas because polyt primers are used to prime the first
Learning Objectives :
 Learning Objectives : Understand the basic differences between genomic and cdna libraries Understand how genomic libraries are constructed Understand the purpose for having overlapping DNA fragments in
Learning Objectives : Understand the basic differences between genomic and cdna libraries Understand how genomic libraries are constructed Understand the purpose for having overlapping DNA fragments in
Transcriptional Regulation in Eukaryotes
 Transcriptional Regulation in Eukaryotes Concepts, Strategies, and Techniques Michael Carey Stephen T. Smale COLD SPRING HARBOR LABORATORY PRESS NEW YORK 2000 Cold Spring Harbor Laboratory Press, 0-87969-537-4
Transcriptional Regulation in Eukaryotes Concepts, Strategies, and Techniques Michael Carey Stephen T. Smale COLD SPRING HARBOR LABORATORY PRESS NEW YORK 2000 Cold Spring Harbor Laboratory Press, 0-87969-537-4
Genetic Engineering & Recombinant DNA
 Genetic Engineering & Recombinant DNA Chapter 10 Copyright The McGraw-Hill Companies, Inc) Permission required for reproduction or display. Applications of Genetic Engineering Basic science vs. Applied
Genetic Engineering & Recombinant DNA Chapter 10 Copyright The McGraw-Hill Companies, Inc) Permission required for reproduction or display. Applications of Genetic Engineering Basic science vs. Applied
3. human genomics clone genes associated with genetic disorders. 4. many projects generate ordered clones that cover genome
 Lectures 30 and 31 Genome analysis I. Genome analysis A. two general areas 1. structural 2. functional B. genome projects a status report 1. 1 st sequenced: several viral genomes 2. mitochondria and chloroplasts
Lectures 30 and 31 Genome analysis I. Genome analysis A. two general areas 1. structural 2. functional B. genome projects a status report 1. 1 st sequenced: several viral genomes 2. mitochondria and chloroplasts
Supplementary Figures and Legends
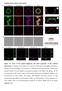 Supplementary Figures and Legends Figure S1. Tests of the optical alignment and focal properties of the confocal microscope. (A) Images of the optical cross-section of fluorescent microspheres differing
Supplementary Figures and Legends Figure S1. Tests of the optical alignment and focal properties of the confocal microscope. (A) Images of the optical cross-section of fluorescent microspheres differing
DNA Transcription. Visualizing Transcription. The Transcription Process
 DNA Transcription By: Suzanne Clancy, Ph.D. 2008 Nature Education Citation: Clancy, S. (2008) DNA transcription. Nature Education 1(1) If DNA is a book, then how is it read? Learn more about the DNA transcription
DNA Transcription By: Suzanne Clancy, Ph.D. 2008 Nature Education Citation: Clancy, S. (2008) DNA transcription. Nature Education 1(1) If DNA is a book, then how is it read? Learn more about the DNA transcription
Chapter 15 Gene Technologies and Human Applications
 Chapter Outline Chapter 15 Gene Technologies and Human Applications Section 1: The Human Genome KEY IDEAS > Why is the Human Genome Project so important? > How do genomics and gene technologies affect
Chapter Outline Chapter 15 Gene Technologies and Human Applications Section 1: The Human Genome KEY IDEAS > Why is the Human Genome Project so important? > How do genomics and gene technologies affect
Sept 2. Structure and Organization of Genomes. Today: Genetic and Physical Mapping. Sept 9. Forward and Reverse Genetics. Genetic and Physical Mapping
 Sept 2. Structure and Organization of Genomes Today: Genetic and Physical Mapping Assignments: Gibson & Muse, pp.4-10 Brown, pp. 126-160 Olson et al., Science 245: 1434 New homework:due, before class,
Sept 2. Structure and Organization of Genomes Today: Genetic and Physical Mapping Assignments: Gibson & Muse, pp.4-10 Brown, pp. 126-160 Olson et al., Science 245: 1434 New homework:due, before class,
Contents. 1 Basic Molecular Microbiology of Bacteria... 1 Exp. 1.1 Isolation of Genomic DNA Introduction Principle...
 Contents 1 Basic Molecular Microbiology of Bacteria... 1 Exp. 1.1 Isolation of Genomic DNA... 1 Introduction... 1 Principle... 1 Reagents Required and Their Role... 2 Procedure... 3 Observation... 4 Result
Contents 1 Basic Molecular Microbiology of Bacteria... 1 Exp. 1.1 Isolation of Genomic DNA... 1 Introduction... 1 Principle... 1 Reagents Required and Their Role... 2 Procedure... 3 Observation... 4 Result
SuperScript IV Reverse Transcriptase as a better alternative to AMV-based enzymes
 WHITE PAPER SuperScript IV Reverse Transcriptase SuperScript IV Reverse Transcriptase as a better alternative to AMV-based enzymes Abstract Reverse transcriptases (RTs) from avian myeloblastosis virus
WHITE PAPER SuperScript IV Reverse Transcriptase SuperScript IV Reverse Transcriptase as a better alternative to AMV-based enzymes Abstract Reverse transcriptases (RTs) from avian myeloblastosis virus
Supplementary Table 1. The Q-PCR primer sequence is summarized in the following table.
 Supplementary Table 1. The Q-PCR primer sequence is summarized in the following table. Name Sequence (5-3 ) Application Flag-u ggactacaaggacgacgatgac Shared upstream primer for all the amplifications of
Supplementary Table 1. The Q-PCR primer sequence is summarized in the following table. Name Sequence (5-3 ) Application Flag-u ggactacaaggacgacgatgac Shared upstream primer for all the amplifications of
Lecture 25 (11/15/17)
 Lecture 25 (11/15/17) Reading: Ch9; 328-332 Ch25; 990-995, 1005-1012 Problems: Ch9 (study-guide: applying); 1,2 Ch9 (study-guide: facts); 7,8 Ch25 (text); 1-3,5-7,9,10,13-15 Ch25 (study-guide: applying);
Lecture 25 (11/15/17) Reading: Ch9; 328-332 Ch25; 990-995, 1005-1012 Problems: Ch9 (study-guide: applying); 1,2 Ch9 (study-guide: facts); 7,8 Ch25 (text); 1-3,5-7,9,10,13-15 Ch25 (study-guide: applying);
Synthetic Biology. Sustainable Energy. Therapeutics Industrial Enzymes. Agriculture. Accelerating Discoveries, Expanding Possibilities. Design.
 Synthetic Biology Accelerating Discoveries, Expanding Possibilities Sustainable Energy Therapeutics Industrial Enzymes Agriculture Design Build Generate Solutions to Advance Synthetic Biology Research
Synthetic Biology Accelerating Discoveries, Expanding Possibilities Sustainable Energy Therapeutics Industrial Enzymes Agriculture Design Build Generate Solutions to Advance Synthetic Biology Research
RNA oligonucleotides and 2 -O-methylated oligonucleotides were synthesized by. 5 AGACACAAACACCAUUGUCACACUCCACAGC; Rand-2 OMe,
 Materials and methods Oligonucleotides and DNA constructs RNA oligonucleotides and 2 -O-methylated oligonucleotides were synthesized by Dharmacon Inc. (Lafayette, CO). The sequences were: 122-2 OMe, 5
Materials and methods Oligonucleotides and DNA constructs RNA oligonucleotides and 2 -O-methylated oligonucleotides were synthesized by Dharmacon Inc. (Lafayette, CO). The sequences were: 122-2 OMe, 5
Non-Organic-Based Isolation of Mammalian microrna using Norgen s microrna Purification Kit
 Application Note 13 RNA Sample Preparation Non-Organic-Based Isolation of Mammalian microrna using Norgen s microrna Purification Kit B. Lam, PhD 1, P. Roberts, MSc 1 Y. Haj-Ahmad, M.Sc., Ph.D 1,2 1 Norgen
Application Note 13 RNA Sample Preparation Non-Organic-Based Isolation of Mammalian microrna using Norgen s microrna Purification Kit B. Lam, PhD 1, P. Roberts, MSc 1 Y. Haj-Ahmad, M.Sc., Ph.D 1,2 1 Norgen
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION Legends for Supplementary Tables. Supplementary Table 1. An excel file containing primary screen data. Worksheet 1, Normalized quantification data from a duplicated screen: valid
SUPPLEMENTARY INFORMATION Legends for Supplementary Tables. Supplementary Table 1. An excel file containing primary screen data. Worksheet 1, Normalized quantification data from a duplicated screen: valid
Nuclear Organization and Gene Expression Dr. David L. Spector
 NUCLEAR ORGANIZATION AND GENE EXPRESSION David L. Spector, Ph.D. Cold Spring Harbor Laboratory One Bungtown Road Cold Spring Harbor, New York 11724 Visit our website at www.cshl.edu/spectorlab 1 Orphanides
NUCLEAR ORGANIZATION AND GENE EXPRESSION David L. Spector, Ph.D. Cold Spring Harbor Laboratory One Bungtown Road Cold Spring Harbor, New York 11724 Visit our website at www.cshl.edu/spectorlab 1 Orphanides
Biology Lecture 2 Genes
 Genes Definitions o Gene: DNA that codes for a single polypeptide/mrna/rrna/trna o Euchromatin: region of DNA containing genes being actively transcribed o Heterochromatin: region of DNA containing genes
Genes Definitions o Gene: DNA that codes for a single polypeptide/mrna/rrna/trna o Euchromatin: region of DNA containing genes being actively transcribed o Heterochromatin: region of DNA containing genes
Chapter 20: Biotechnology
 Name Period The AP Biology exam has reached into this chapter for essay questions on a regular basis over the past 15 years. Student responses show that biotechnology is a difficult topic. This chapter
Name Period The AP Biology exam has reached into this chapter for essay questions on a regular basis over the past 15 years. Student responses show that biotechnology is a difficult topic. This chapter
Concepts: What are RFLPs and how do they act like genetic marker loci?
 Restriction Fragment Length Polymorphisms (RFLPs) -1 Readings: Griffiths et al: 7th Edition: Ch. 12 pp. 384-386; Ch.13 pp404-407 8th Edition: pp. 364-366 Assigned Problems: 8th Ch. 11: 32, 34, 38-39 7th
Restriction Fragment Length Polymorphisms (RFLPs) -1 Readings: Griffiths et al: 7th Edition: Ch. 12 pp. 384-386; Ch.13 pp404-407 8th Edition: pp. 364-366 Assigned Problems: 8th Ch. 11: 32, 34, 38-39 7th
Correction: The Leukemia-Associated Mllt10/ Af10-Dot1l Are Tcf4/β-Catenin Coactivators Essential for Intestinal Homeostasis
 CORRECTION Correction: The Leukemia-Associated Mllt10/ Af10-Dot1l Are Tcf4/β-Catenin Coactivators Essential for Intestinal Homeostasis Tokameh Mahmoudi, Sylvia F. Boj, Pantelis Hatzis, Vivian S. W. Li,
CORRECTION Correction: The Leukemia-Associated Mllt10/ Af10-Dot1l Are Tcf4/β-Catenin Coactivators Essential for Intestinal Homeostasis Tokameh Mahmoudi, Sylvia F. Boj, Pantelis Hatzis, Vivian S. W. Li,
7.17: Writing Up Results and Creating Illustrations
 7.17: Writing Up Results and Creating Illustrations A Results Exercise: Kansas and Pancakes Write a 5-sentence paragraph describing the results illustrated in this figure: - Describe the figure: highlights?
7.17: Writing Up Results and Creating Illustrations A Results Exercise: Kansas and Pancakes Write a 5-sentence paragraph describing the results illustrated in this figure: - Describe the figure: highlights?
AP Biology Gene Expression/Biotechnology REVIEW
 AP Biology Gene Expression/Biotechnology REVIEW Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Gene expression can be a. regulated before transcription.
AP Biology Gene Expression/Biotechnology REVIEW Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Gene expression can be a. regulated before transcription.
TRANSGENIC TECHNOLOGIES: Gene-targeting
 TRANSGENIC TECHNOLOGIES: Gene-targeting Reverse Genetics Wild-type Bmp7 -/- Forward Genetics Phenotype Gene or Mutations First Molecular Analysis Second Reverse Genetics Gene Phenotype or Molecular Analysis
TRANSGENIC TECHNOLOGIES: Gene-targeting Reverse Genetics Wild-type Bmp7 -/- Forward Genetics Phenotype Gene or Mutations First Molecular Analysis Second Reverse Genetics Gene Phenotype or Molecular Analysis
Bio-Reagent Services. Custom Gene Services. Gateway to Smooth Molecular Biology! Your Innovation Partner in Drug Discovery!
 Bio-Reagent Services Custom Gene Services Gateway to Smooth Molecular Biology! Gene Synthesis Mutagenesis Mutant Libraries Plasmid Preparation sirna and mirna Services Large-scale DNA Sequencing GenPool
Bio-Reagent Services Custom Gene Services Gateway to Smooth Molecular Biology! Gene Synthesis Mutagenesis Mutant Libraries Plasmid Preparation sirna and mirna Services Large-scale DNA Sequencing GenPool
American Society of Cytopathology Core Curriculum in Molecular Biology
 American Society of Cytopathology Core Curriculum in Molecular Biology American Society of Cytopathology Core Curriculum in Molecular Biology Chapter 3 Molecular Techniques Separation and Detection, Part
American Society of Cytopathology Core Curriculum in Molecular Biology American Society of Cytopathology Core Curriculum in Molecular Biology Chapter 3 Molecular Techniques Separation and Detection, Part
P HENIX. PHENIX PCR Enzyme Guide Tools For Life Science Discovery RESEARCH PRODUCTS
 PHENIX PCR Enzyme Guide PHENIX offers a broad line of premium quality PCR Enzymes. This PCR Enzyme Guide will help simplify your polymerase selection process. Each DNA Polymerase has different characteristics
PHENIX PCR Enzyme Guide PHENIX offers a broad line of premium quality PCR Enzymes. This PCR Enzyme Guide will help simplify your polymerase selection process. Each DNA Polymerase has different characteristics
Concepts and Methods in Developmental Biology
 Biology 4361 Developmental Biology Concepts and Methods in Developmental Biology June 16, 2009 Conceptual and Methodological Tools Concepts Genomic equivalence Differential gene expression Differentiation/de-differentiation
Biology 4361 Developmental Biology Concepts and Methods in Developmental Biology June 16, 2009 Conceptual and Methodological Tools Concepts Genomic equivalence Differential gene expression Differentiation/de-differentiation
Cloning and Sequencing of the Gene Encoding Curvaticin FS47, an Anti- Listerial Bacteriocin Produced by Lactobacillus curvatus FS47
 Cloning and Sequencing of the Gene Encoding Curvaticin FS47, an Anti- Listerial Bacteriocin Produced by Lactobacillus curvatus FS47 S. Macwana, L. Ma, M.A. Cousin, and P.M. Muriana Story in Brief Curvaticin
Cloning and Sequencing of the Gene Encoding Curvaticin FS47, an Anti- Listerial Bacteriocin Produced by Lactobacillus curvatus FS47 S. Macwana, L. Ma, M.A. Cousin, and P.M. Muriana Story in Brief Curvaticin
Purification of alpha-1 antitrypsin using an antibody based affinity chromatography medium
 Purification of alpha-1 antitrypsin using an antibody based affinity chromatography medium Ulrika Meyer a, Hanna Wlad a, Sven Blokland b, Frank J.M. Detmers b and Henrik Ihre a a GE Healthcare Bio-Sciences
Purification of alpha-1 antitrypsin using an antibody based affinity chromatography medium Ulrika Meyer a, Hanna Wlad a, Sven Blokland b, Frank J.M. Detmers b and Henrik Ihre a a GE Healthcare Bio-Sciences
Recent technology allow production of microarrays composed of 70-mers (essentially a hybrid of the two techniques)
 Microarrays and Transcript Profiling Gene expression patterns are traditionally studied using Northern blots (DNA-RNA hybridization assays). This approach involves separation of total or polya + RNA on
Microarrays and Transcript Profiling Gene expression patterns are traditionally studied using Northern blots (DNA-RNA hybridization assays). This approach involves separation of total or polya + RNA on
Chapter 5 DNA and Chromosomes
 Chapter 5 DNA and Chromosomes DNA as the genetic material Heat-killed bacteria can transform living cells S Smooth R Rough Fred Griffith, 1920 DNA is the genetic material Oswald Avery Colin MacLeod Maclyn
Chapter 5 DNA and Chromosomes DNA as the genetic material Heat-killed bacteria can transform living cells S Smooth R Rough Fred Griffith, 1920 DNA is the genetic material Oswald Avery Colin MacLeod Maclyn
Ribosomal DNA Integrating raav-rdna Vectors Allow for Stable Transgene Expression
 original article The American Society of Gene & Cell Therapy Ribosomal DNA Integrating raav-rdna Vectors Allow for Stable Transgene Expression Leszek Lisowski, Ashley Lau,2, Zhongya Wang 3, Yue Zhang,
original article The American Society of Gene & Cell Therapy Ribosomal DNA Integrating raav-rdna Vectors Allow for Stable Transgene Expression Leszek Lisowski, Ashley Lau,2, Zhongya Wang 3, Yue Zhang,
Supplementary Information
 Supplementary Information Deletion of the B-B and C-C regions of inverted terminal repeats reduces raav productivity but increases transgene expression Qingzhang Zhou 1, Wenhong Tian 2, Chunguo Liu 3,
Supplementary Information Deletion of the B-B and C-C regions of inverted terminal repeats reduces raav productivity but increases transgene expression Qingzhang Zhou 1, Wenhong Tian 2, Chunguo Liu 3,
Supplementary Figure 1. (a) The qrt-pcr for lnc-2, lnc-6 and lnc-7 RNA level in DU145, 22Rv1, wild type HCT116 and HCT116 Dicer ex5 cells transfected
 Supplementary Figure 1. (a) The qrt-pcr for lnc-2, lnc-6 and lnc-7 RNA level in DU145, 22Rv1, wild type HCT116 and HCT116 Dicer ex5 cells transfected with the sirna against lnc-2, lnc-6, lnc-7, and the
Supplementary Figure 1. (a) The qrt-pcr for lnc-2, lnc-6 and lnc-7 RNA level in DU145, 22Rv1, wild type HCT116 and HCT116 Dicer ex5 cells transfected with the sirna against lnc-2, lnc-6, lnc-7, and the
Bio 101 Sample questions: Chapter 10
 Bio 101 Sample questions: Chapter 10 1. Which of the following is NOT needed for DNA replication? A. nucleotides B. ribosomes C. Enzymes (like polymerases) D. DNA E. all of the above are needed 2 The information
Bio 101 Sample questions: Chapter 10 1. Which of the following is NOT needed for DNA replication? A. nucleotides B. ribosomes C. Enzymes (like polymerases) D. DNA E. all of the above are needed 2 The information
Chapter 3. Enzyme manipulation of DNA and RNA
 Chapter 3 Enzyme manipulation of DNA and RNA To measure incorporation of radioactivity (to see if the probe is good or not for hybridization) Acid precipitation method: - Add sonicated salmon sperm DNA
Chapter 3 Enzyme manipulation of DNA and RNA To measure incorporation of radioactivity (to see if the probe is good or not for hybridization) Acid precipitation method: - Add sonicated salmon sperm DNA
HiPer RT-PCR Teaching Kit
 HiPer RT-PCR Teaching Kit Product Code: HTBM024 Number of experiments that can be performed: 5 Duration of Experiment: Protocol: 4 hours Agarose Gel Electrophoresis: 45 minutes Storage Instructions: The
HiPer RT-PCR Teaching Kit Product Code: HTBM024 Number of experiments that can be performed: 5 Duration of Experiment: Protocol: 4 hours Agarose Gel Electrophoresis: 45 minutes Storage Instructions: The
Biotechnology and Genomics in Public Health. Sharon S. Krag, PhD Johns Hopkins University
 This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike License. Your use of this material constitutes acceptance of that license and the conditions of use of materials on this
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike License. Your use of this material constitutes acceptance of that license and the conditions of use of materials on this
RNA Isolation and Technology Applications. Nadine Nassif Senior Research Scientist Promega Corporation
 RNA Isolation and Technology Applications Nadine Nassif Senior Research Scientist Promega Corporation verview Brief overview of basic RNA/DNA chemistry. verview of total and poly(a+) RNA isolation. Discuss
RNA Isolation and Technology Applications Nadine Nassif Senior Research Scientist Promega Corporation verview Brief overview of basic RNA/DNA chemistry. verview of total and poly(a+) RNA isolation. Discuss
High Pure PCR Template Preparation Kit for preparation of 100 nucleic acid samples Cat. No
 for preparation of 100 nucleic acid samples Cat. No. 1 796 88 Principle Cells are lysed during a short incubation with Proteinase K in the presence of a chaotropic salt (guanidine HCl), which immediately
for preparation of 100 nucleic acid samples Cat. No. 1 796 88 Principle Cells are lysed during a short incubation with Proteinase K in the presence of a chaotropic salt (guanidine HCl), which immediately
Gene Expression Technology
 Gene Expression Technology Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Gene expression Gene expression is the process by which information from a gene
Gene Expression Technology Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Gene expression Gene expression is the process by which information from a gene
Epigenetic Regulation of Retrotransposons within the Nucleolus of Drosophila
 MOLECULAR AND CELLULAR BIOLOGY, Oct. 2008, p. 6452 6461 Vol. 28, No. 20 0270-7306/08/$08.00 0 doi:10.1128/mcb.01015-08 Copyright 2008, American Society for Microbiology. All Rights Reserved. Epigenetic
MOLECULAR AND CELLULAR BIOLOGY, Oct. 2008, p. 6452 6461 Vol. 28, No. 20 0270-7306/08/$08.00 0 doi:10.1128/mcb.01015-08 Copyright 2008, American Society for Microbiology. All Rights Reserved. Epigenetic
The preparation of native chromatin from cultured human cells.
 Native chromatin immunoprecipitation protocol The preparation of native chromatin from cultured human cells. All solutions need to be ice cold. Sucrose containing solutions must be made up fresh on the
Native chromatin immunoprecipitation protocol The preparation of native chromatin from cultured human cells. All solutions need to be ice cold. Sucrose containing solutions must be made up fresh on the
BS 50 Genetics and Genomics Week of Nov 29
 BS 50 Genetics and Genomics Week of Nov 29 Additional Practice Problems for Section Problem 1. A linear piece of DNA is digested with restriction enzymes EcoRI and HinDIII, and the products are separated
BS 50 Genetics and Genomics Week of Nov 29 Additional Practice Problems for Section Problem 1. A linear piece of DNA is digested with restriction enzymes EcoRI and HinDIII, and the products are separated
CLASS 3.5: 03/29/07 EUKARYOTIC TRANSCRIPTION I: PROMOTERS AND ENHANCERS
 CLASS 3.5: 03/29/07 EUKARYOTIC TRANSCRIPTION I: PROMOTERS AND ENHANCERS A. Promoters and Polymerases (RNA pols): 1. General characteristics - Initiation of transcription requires a. Transcription factors
CLASS 3.5: 03/29/07 EUKARYOTIC TRANSCRIPTION I: PROMOTERS AND ENHANCERS A. Promoters and Polymerases (RNA pols): 1. General characteristics - Initiation of transcription requires a. Transcription factors
