To assess the localization of Citrine fusion proteins, we performed antibody staining to
|
|
|
- Erica Byrd
- 6 years ago
- Views:
Transcription
1 Trinh et al 1 SUPPLEMENTAL MATERIAL FlipTraps recapitulate endogenous protein localization To assess the localization of Citrine fusion proteins, we performed antibody staining to compare the expression of Citrine fusion proteins with the endogenous proteins. One challenge of immunohistochemistry experiments is the availability of antibodies that will recognize the endogenous zebrafish protein. To expand the number of comparisons possible, we have also compared localization of Citrine fusion proteins with antibody staining to endogenous zebrafish proteins that localize to the same subcellular compartments as the trap gene. Two examples are shown in Supplemental Figure 3. Gt(ctnna-citrine) ct3a is a trap of the -catenin gene, a component of the cadherin-catenin complex, co-localizing with E-cadherin and b-catenin in epithelial cells (citation). Immunohistochemistry with an antibody to -catenin show the same localization pattern as the Ctnna-Citrine fusion protein (Supplemental Figure 3A-C). Gt(lap2b-citrine) ct3b is a trap of the lamina-associated polypeptide 2b (LAP2b). LAP2 family members localize to the inner nuclear membrane. Immunofluorescence labeling with an antibody to the nuclear envelope protein LaminB co-localizes with the LAP2b-Citrine expression (Supplemental Figure 3D-F). The co-localization of the Citrine fusion protein with the endogenous protein indicates that FlipTrap fusions are properly localized. Functional Cerulean-Cre fusion to assess expression of Cre recombinase We developed a Cerulean-Cre fusion protein to enable the tracking of Cre recombinase expression in the developing embryo. To stably express Cerulean-Cre in the embryo, we have constructed a Tol2-based integration vector that used the ubiquitous beta- 1
2 Trinh et al 2 actin2 promoter to drive expression of Cerulean-Cre (Supplemental Figure 5A). Using this construct, we isolated a stable line, Tg(bactin2:cerulean-cre) ct5000, that exhibited ubiquitous Cerulean-Cre expression (Supplemental Figure 5B). Cerulean-Cre fusion protein localizes to the nucleus of all cells in the developing embryo (Supplemental Figure 5C, E). Importantly, breeding adult Tg(bactin2:cerulean-cre) ct5000 fish to FlipTrap lines results in the recombination of the lox sites, converting Citrine to mcherry fusion proteins (Figure 4F-I). SUPPLEMENTAL MATERIALS AND METHODS Construction of FlipTrap vector The following pairs of primers were used to perform fusion PCR of the citrine, mcherry, -tubulin polya, rassf8 splice acceptor and donor, lox and FRT sequences to create the FlipTrap vector: F3-F:5 agtctacgcccccaactgagagaactcaaaggttaccccagttggggcactacatcgattcaggaacctc acagactgc-3 and F3-R: 5 -tcccgggtgaatgtgtagcgaccagttcg-3, F4-F: 5 tcgctacacattcacccgggataacttcgtatagcatacattatacgaagttatccggagtgagcaagggc gaggagctg-3 and F4-R: 5 -acctgtagcccaatgtttgaaggaccggtcttgtacagctcgtccatgc-3, F5-F: 5 -ttcaaacattgggctacaggtgagtgtacctgaagaaagaacacaatttacttacacttacatttcaaacagat- 3 and F5-R:5 aaaagtagattagttacgtaataacttcgtataaagtatcctatacgaagttatggatccctttgggtag tttacaata-3, F6-F: 5 tacgtaactaatctacttttccttcatcccaagtggttggaagccaaacctgccaaagattagtaagcaga-3 and F6-R: 5 -taagatatcaatatccactaaatgtctaaaactg-3, 2
3 Trinh et al 3 F7-F: 5 tagtggatattgatatcttacttgtacagctcgtcca-3 and F7-R:5 atgataatatggccacaaccatgg tgagcaagggcgagga-3 F8-F:5 -ctcaccatggttgtggccatattatcatc-3 and F8-R:5 catacattatacgaagttatctagatagct gactgagcccctctccctcccccccccctaacgttac-3. SUPPLEMENTAL FIGURE LEGENDS: Supplemental Figure 1. Flow-chart for FlipTrap screen FlipTrap vector was injected with transposase mrna (tol2 or Ac) into the one-cell stage embryo to generate mosaic F 0 adults. The F 0 were crossed to wildtype. The F 1 embryos are screened for Citrine expression with a fluorescent dissecting scope and documented. Positive citrine embryos were subjected to a secondary screen in which the embryos were counter-stained with the vital stain, BodipyTR-methyl ester and imaged by confocal microscopy to obtain subcellular localization data. Because the screen is non-invasive, positive F 1 embryos can be raised after imaging to establish the line. They were also processed for RACE analysis to identify the trapped genes. Supplemental Figure 2: Differential spatio-temporal subcellular localization of Mapkapk2-Citrine. (A, B) Wide-field fluorescence image of Gt(mapkapk2-citrine) ct52b embryo at 26hpf (A) and 32hpf (B). Mapkapk2-Citrine is ubiquitously expressed but only heightened in the notochord at 26hpf (arrow). (C-H) Confocal image of Gt(mapkapk2-citrine) ct52b embryos 3
4 Trinh et al 4 counter-stained with bodipytr-methyl ester (red) at 26hpf (C-E) and 56hpf (F-H). (C, F) Mapkapk2-citrine expression in the trunk (D, G) BodipyTR-methyl ester staining the trunk. (E, I) Merge of images from C, D and F, G, respectively. At 26hpf, Mapkapk2- citrine is localized to the membrane of the vacuolating notochord cells. The membrane localization in the notochord cells is not observed in older, 56hpf embryos. Scale bar, 20 m. Supplemental Figure 3: Co-localization of Citrine fusion protein with endogenous proteins. Confocal images of Gt(ctnna-citrine) ct3a (A-C) and Gt(lap2b-citrine) ct3b (D-E) embryos immunostained for -catenin (B) and LaminB (E). (A) Ctnna-Citrine expression (green) is localized to the cortex of cells. (B) The pattern of antibody staining for -catenin appears similar to that of the Ctnna-Citrine fusion protein (A). (D) LAP2 -Citrine expression (green) is localized to the nuclear envelope. (E)The antibody staining of LaminB is similar to LAP2 -Citrine. (C) and (F) are merged images of (A,B) and (D,E) respectively. Scale bar, 20 m. Supplemental Figure 4: Citrine fusions are functional (A) 94.2% of FlipTrap lines are viable as homozygous of Citrine fusion (N=122). (B-E) Brightfield image of wildtype (B) and three FlipTrap lines (C-E) that exhibit developmental defects and are non-viable as homozygous Citrine fusion. (C) Gt(nt5c2- citrine) ct103b is a trap of 5'-nucleotidase, cytosolic IIa (nt5c2) in which the FlipTrap vector 4
5 Trinh et al 5 has inserted between exons 9 and 10 of 18 exons, which disrupts (HAD) domain. (D) Gt(dnm1l-citrine) ct111a is a trap of dynamin 1-like (dnm1l) that has inserted between exons 10 and 11 of 18 exons and disrupts the dynamin-central domain in Dnm1l. (E) Gt(rpl17-citrine) ct122b is a trap of a novel ribosomal protein in which the citrine exon creates a fusion protein that disrupts a ribosomal L22 domain that critical for interaction with 23S rrna. Scale bar, 50 m. Supplemental Figure 5: A stable transgene expressing Cerulean-Cre fusion. (A) Schematic of ubiquitously expressing Cerulean-cre construction that contains Tol2 transposable elements (outlined blue rectangle), minimal beta-actin2 promoter (light orange) that containing the first two exons of the beta-actin2 gene (orange), and coding sequence for cerulean-cre (teal). (B) Wide-field fluorescence image of Tg(bactin2:cerulean-cre) ct5000 embryo at 32hpf. (C-D) Confocal image of Tg(bactin2:cerulean-cre) ct5000 with expression of Cerulean-cre in the otic vesicle (C) and trunk (D) showing nuclear localization of Cre. Scale bar, 20 m (C,D) and 50 m (B). Supplemental Figure 6: Cre-lox mediated recombination of FlipTrap in of high mobility group AT-hook 2 (hmga2). (A,B) Schematic of hmga2 locus with FlipTrap insertion before (A) and after Cre recombination (B). Predicted protein domains of hmga2 gene as determined by Simple Modular Architecture Research Tool (SMART) with Citrine (A, green bar) and mcherry (B, red bar) protein fusion. AT-hook DNA binding motif are shown as blue ovals. Insertion of the FlipTrap vector has occurred between exon 3 and 4. The Cre induced 5
6 Trinh et al 6 mutant allele leads to the production of Hmga2-mCherry that retains the three DNA binding motif. (C, D) Wide-field fluorescent image of Gt(hmga-citrine) ct29a (C) and Gt(hmga-citrine) ct29ar (D) embryo taken with YFP and RFP filters, respectively. (E, F) Confocal image of the otic vesicle in Gt(hmga-citrine) ct29a (E) and Gt(hmga-citrine) ct29ar (F) in which the cell membranes are labeled with a membrane localized Cerulean protein (blue). Both the full-length Hmga2-citrine and truncated Hmga2-mCherry fusion proteins are localized to the nucleus of the epithelium of the otic vesicle and as expected, the Gt(hmga-citrine) ct29ar homozygous embryos are viable. Supplemental Figure 7: Clathrin heavy chain A is essential for neural and vasculature development. (A-D) Brightfield image of wildtype (A, C) and homozygous Gt(cltca-citrine) ct116ar (B, D) embryos at 32hpf. (B)Head region of homozygous Gt(cltca-mCherry) ct116ar embryos showing a smaller forebrain and midbrain in comparison to wildtype (A). (D) Tail region homozygous Gt(cltca-mCherry) ct116ar embryos show dilated caudal vein (arrow). 6
A CRISPR/Cas9 Vector System for Tissue-Specific Gene Disruption in Zebrafish
 Developmental Cell Supplemental Information A CRISPR/Cas9 Vector System for Tissue-Specific Gene Disruption in Zebrafish Julien Ablain, Ellen M. Durand, Song Yang, Yi Zhou, and Leonard I. Zon % larvae
Developmental Cell Supplemental Information A CRISPR/Cas9 Vector System for Tissue-Specific Gene Disruption in Zebrafish Julien Ablain, Ellen M. Durand, Song Yang, Yi Zhou, and Leonard I. Zon % larvae
Supplementary Figure 1. Isolation of GFPHigh cells.
 Supplementary Figure 1. Isolation of GFP High cells. (A) Schematic diagram of cell isolation based on Wnt signaling activity. Colorectal cancer (CRC) cell lines were stably transduced with lentivirus encoding
Supplementary Figure 1. Isolation of GFP High cells. (A) Schematic diagram of cell isolation based on Wnt signaling activity. Colorectal cancer (CRC) cell lines were stably transduced with lentivirus encoding
The Human Protein PRR14 Tethers Heterochromatin to the Nuclear Lamina During Interphase and Mitotic Exit
 Cell Reports, Volume 5 Supplemental Information The Human Protein PRR14 Tethers Heterochromatin to the Nuclear Lamina During Interphase and Mitotic Exit Andrey Poleshko, Katelyn M. Mansfield, Caroline
Cell Reports, Volume 5 Supplemental Information The Human Protein PRR14 Tethers Heterochromatin to the Nuclear Lamina During Interphase and Mitotic Exit Andrey Poleshko, Katelyn M. Mansfield, Caroline
Introducing new DNA into the genome requires cloning the donor sequence, delivery of the cloned DNA into the cell, and integration into the genome.
 Key Terms Chapter 32: Genetic Engineering Cloning describes propagation of a DNA sequence by incorporating it into a hybrid construct that can be replicated in a host cell. A cloning vector is a plasmid
Key Terms Chapter 32: Genetic Engineering Cloning describes propagation of a DNA sequence by incorporating it into a hybrid construct that can be replicated in a host cell. A cloning vector is a plasmid
Supplementary Figure S1. Immunodetection of full-length XA21 and the XA21 C-terminal cleavage product.
 Supplementary Information Supplementary Figure S1. Immunodetection of full-length XA21 and the XA21 C-terminal cleavage product. Total protein extracted from Kitaake wild type and rice plants carrying
Supplementary Information Supplementary Figure S1. Immunodetection of full-length XA21 and the XA21 C-terminal cleavage product. Total protein extracted from Kitaake wild type and rice plants carrying
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10163 Supplementary Table 1 Efficiency of vector construction. Process wells recovered efficiency (%) Recombineering* 480 461 96 Intermediate plasmids 461 381 83 Recombineering efficiency
doi:10.1038/nature10163 Supplementary Table 1 Efficiency of vector construction. Process wells recovered efficiency (%) Recombineering* 480 461 96 Intermediate plasmids 461 381 83 Recombineering efficiency
Supplementary Table 1. The Q-PCR primer sequence is summarized in the following table.
 Supplementary Table 1. The Q-PCR primer sequence is summarized in the following table. Name Sequence (5-3 ) Application Flag-u ggactacaaggacgacgatgac Shared upstream primer for all the amplifications of
Supplementary Table 1. The Q-PCR primer sequence is summarized in the following table. Name Sequence (5-3 ) Application Flag-u ggactacaaggacgacgatgac Shared upstream primer for all the amplifications of
Trasposable elements: Uses of P elements Problem set B at the end
 Trasposable elements: Uses of P elements Problem set B at the end P-elements have revolutionized the way Drosophila geneticists conduct their research. Here, we will discuss just a few of the approaches
Trasposable elements: Uses of P elements Problem set B at the end P-elements have revolutionized the way Drosophila geneticists conduct their research. Here, we will discuss just a few of the approaches
8/21/2014. From Gene to Protein
 From Gene to Protein Chapter 17 Objectives Describe the contributions made by Garrod, Beadle, and Tatum to our understanding of the relationship between genes and enzymes Briefly explain how information
From Gene to Protein Chapter 17 Objectives Describe the contributions made by Garrod, Beadle, and Tatum to our understanding of the relationship between genes and enzymes Briefly explain how information
Gene Expression: Transcription
 Gene Expression: Transcription The majority of genes are expressed as the proteins they encode. The process occurs in two steps: Transcription = DNA RNA Translation = RNA protein Taken together, they make
Gene Expression: Transcription The majority of genes are expressed as the proteins they encode. The process occurs in two steps: Transcription = DNA RNA Translation = RNA protein Taken together, they make
Bio 101 Sample questions: Chapter 10
 Bio 101 Sample questions: Chapter 10 1. Which of the following is NOT needed for DNA replication? A. nucleotides B. ribosomes C. Enzymes (like polymerases) D. DNA E. all of the above are needed 2 The information
Bio 101 Sample questions: Chapter 10 1. Which of the following is NOT needed for DNA replication? A. nucleotides B. ribosomes C. Enzymes (like polymerases) D. DNA E. all of the above are needed 2 The information
GM130 Is Required for Compartmental Organization of Dendritic Golgi Outposts
 Current Biology, Volume 24 Supplemental Information GM130 Is Required for Compartmental Organization of Dendritic Golgi Outposts Wei Zhou, Jin Chang, Xin Wang, Masha G. Savelieff, Yinyin Zhao, Shanshan
Current Biology, Volume 24 Supplemental Information GM130 Is Required for Compartmental Organization of Dendritic Golgi Outposts Wei Zhou, Jin Chang, Xin Wang, Masha G. Savelieff, Yinyin Zhao, Shanshan
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature09937 a Name Position Primersets 1a 1b 2 3 4 b2 Phenotype Genotype b Primerset 1a D T C R I E 10000 8000 6000 5000 4000 3000 2500 2000 1500 1000 800 Donor (D)
SUPPLEMENTARY INFORMATION doi:10.1038/nature09937 a Name Position Primersets 1a 1b 2 3 4 b2 Phenotype Genotype b Primerset 1a D T C R I E 10000 8000 6000 5000 4000 3000 2500 2000 1500 1000 800 Donor (D)
Figure S1. Figure S2. Figure S3 HB Anti-FSP27 (COOH-terminal peptide) Ab. Anti-GST-FSP27(45-127) Ab.
 / 36B4 mrna ratio Figure S1 * 2. 1.6 1.2.8 *.4 control TNFα BRL49653 Figure S2 Su bw AT p iw Anti- (COOH-terminal peptide) Ab Blot : Anti-GST-(45-127) Ab β-actin Figure S3 HB2 HW AT BA T Figure S4 A TAG
/ 36B4 mrna ratio Figure S1 * 2. 1.6 1.2.8 *.4 control TNFα BRL49653 Figure S2 Su bw AT p iw Anti- (COOH-terminal peptide) Ab Blot : Anti-GST-(45-127) Ab β-actin Figure S3 HB2 HW AT BA T Figure S4 A TAG
Effective Gene Trapping Mediated by Sleeping Beauty Transposon
 Effective Gene Trapping Mediated by Sleeping Beauty Transposon Guili Song 1,2, Qing Li 1, Yong Long 1, Qilin Gu 1,2, Perry B. Hackett 3, Zongbin Cui 1 * 1 The Key Laboratory of Aquatic Biodiversity and
Effective Gene Trapping Mediated by Sleeping Beauty Transposon Guili Song 1,2, Qing Li 1, Yong Long 1, Qilin Gu 1,2, Perry B. Hackett 3, Zongbin Cui 1 * 1 The Key Laboratory of Aquatic Biodiversity and
Protein Synthesis & Gene Expression
 DNA provides the instructions for how to build proteins Each gene dictates how to build a single protein in prokaryotes The sequence of nucleotides (AGCT) in DNA dictates the order of amino acids that
DNA provides the instructions for how to build proteins Each gene dictates how to build a single protein in prokaryotes The sequence of nucleotides (AGCT) in DNA dictates the order of amino acids that
Supplementary Fig. S1. Schematic representation of mouse lines Pax6 fl/fl and mrx-cre used in this study. (A) To generate Pax6 fl/ fl
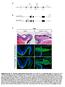 Supplementary Fig. S1. Schematic representation of mouse lines Pax6 fl/fl and mrx-cre used in this study. (A) To generate Pax6 fl/ fl, loxp sites flanking exons 3-6 (red arrowheads) were introduced into
Supplementary Fig. S1. Schematic representation of mouse lines Pax6 fl/fl and mrx-cre used in this study. (A) To generate Pax6 fl/ fl, loxp sites flanking exons 3-6 (red arrowheads) were introduced into
Theoretical cloning project
 Theoretical cloning project Needed to get credits Make it up yourself, don't copy Possible to do in groups of 2-4 students If you need help or an idea, ask! If you have no idea what to clone, I can give
Theoretical cloning project Needed to get credits Make it up yourself, don't copy Possible to do in groups of 2-4 students If you need help or an idea, ask! If you have no idea what to clone, I can give
Cambridge University Press
 Figure 1.1. Model of RNAi pathway in C. elegans. Transmembrane protein SID-1 allows dsrna to enter the cell. In the cytoplasm,dsrna gets processed by DCR-1,existing in a complex with RDE-4,RDE-1 and DRH-1.
Figure 1.1. Model of RNAi pathway in C. elegans. Transmembrane protein SID-1 allows dsrna to enter the cell. In the cytoplasm,dsrna gets processed by DCR-1,existing in a complex with RDE-4,RDE-1 and DRH-1.
Fig Ch 17: From Gene to Protein
 Fig. 17-1 Ch 17: From Gene to Protein Basic Principles of Transcription and Translation RNA is the intermediate between genes and the proteins for which they code Transcription is the synthesis of RNA
Fig. 17-1 Ch 17: From Gene to Protein Basic Principles of Transcription and Translation RNA is the intermediate between genes and the proteins for which they code Transcription is the synthesis of RNA
Transcription in Eukaryotes
 Transcription in Eukaryotes Biology I Hayder A Giha Transcription Transcription is a DNA-directed synthesis of RNA, which is the first step in gene expression. Gene expression, is transformation of the
Transcription in Eukaryotes Biology I Hayder A Giha Transcription Transcription is a DNA-directed synthesis of RNA, which is the first step in gene expression. Gene expression, is transformation of the
The Use of Genetically-Modified Mouse Models to Study the Actin Cytoskeleton
 The Use of Genetically-Modified Mouse Models to Study the Actin Cytoskeleton Anthony Kee (PhD) Cellular and Genetic Medicine Unit School of Medical Sciences (a.kee@unsw.edu.au) 2017 Structure of the Prac
The Use of Genetically-Modified Mouse Models to Study the Actin Cytoskeleton Anthony Kee (PhD) Cellular and Genetic Medicine Unit School of Medical Sciences (a.kee@unsw.edu.au) 2017 Structure of the Prac
NAME TA SEC Problem Set 4 FRIDAY October 15, Answers to this problem set must be inserted into the box outside
 MIT Biology Department 7.012: Introductory Biology - Fall 2004 Instructors: Professor Eric Lander, Professor Robert A. Weinberg, Dr. Claudette Gardel NAME TA SEC 7.012 Problem Set 4 FRIDAY October 15,
MIT Biology Department 7.012: Introductory Biology - Fall 2004 Instructors: Professor Eric Lander, Professor Robert A. Weinberg, Dr. Claudette Gardel NAME TA SEC 7.012 Problem Set 4 FRIDAY October 15,
Short hairpin RNA (shrna) against MMP14. Lentiviral plasmids containing shrna
 Supplemental Materials and Methods Short hairpin RNA (shrna) against MMP14. Lentiviral plasmids containing shrna (Mission shrna, Sigma) against mouse MMP14 were transfected into HEK293 cells using FuGene6
Supplemental Materials and Methods Short hairpin RNA (shrna) against MMP14. Lentiviral plasmids containing shrna (Mission shrna, Sigma) against mouse MMP14 were transfected into HEK293 cells using FuGene6
Supplemental Information. Antagonistic Activities of Sox2. and Brachyury Control the Fate Choice. of Neuro-Mesodermal Progenitors
 Developmental Cell, Volume 42 Supplemental Information Antagonistic Activities of Sox2 and Brachyury Control the Fate Choice of Neuro-Mesodermal Progenitors Frederic Koch, Manuela Scholze, Lars Wittler,
Developmental Cell, Volume 42 Supplemental Information Antagonistic Activities of Sox2 and Brachyury Control the Fate Choice of Neuro-Mesodermal Progenitors Frederic Koch, Manuela Scholze, Lars Wittler,
Protocol for tissue-specific gene disruption in zebrafish
 Protocol for tissue-specific gene disruption in zebrafish Overview This protocol describes a method to inactivate genes in zebrafish in a tissue-specific manner. It can be used to analyze mosaic loss-of-function
Protocol for tissue-specific gene disruption in zebrafish Overview This protocol describes a method to inactivate genes in zebrafish in a tissue-specific manner. It can be used to analyze mosaic loss-of-function
7.17: Writing Up Results and Creating Illustrations
 7.17: Writing Up Results and Creating Illustrations A Results Exercise: Kansas and Pancakes Write a 5-sentence paragraph describing the results illustrated in this figure: - Describe the figure: highlights?
7.17: Writing Up Results and Creating Illustrations A Results Exercise: Kansas and Pancakes Write a 5-sentence paragraph describing the results illustrated in this figure: - Describe the figure: highlights?
Review of Protein (one or more polypeptide) A polypeptide is a long chain of..
 Gene expression Review of Protein (one or more polypeptide) A polypeptide is a long chain of.. In a protein, the sequence of amino acid determines its which determines the protein s A protein with an enzymatic
Gene expression Review of Protein (one or more polypeptide) A polypeptide is a long chain of.. In a protein, the sequence of amino acid determines its which determines the protein s A protein with an enzymatic
Confocal immunofluorescence microscopy
 Confocal immunofluorescence microscopy HL-6 and cells were cultured and cytospun onto glass slides. The cells were double immunofluorescence stained for Mt NPM1 and fibrillarin (nucleolar marker). Briefly,
Confocal immunofluorescence microscopy HL-6 and cells were cultured and cytospun onto glass slides. The cells were double immunofluorescence stained for Mt NPM1 and fibrillarin (nucleolar marker). Briefly,
Post-expansion antibody delivery, after epitope-preserving homogenization.
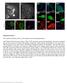 Supplementary Figure 1 Post-expansion antibody delivery, after epitope-preserving homogenization. (a, b) Wide-field fluorescence images of Thy1-YFP-expressing mouse brain hemisphere slice before expansion
Supplementary Figure 1 Post-expansion antibody delivery, after epitope-preserving homogenization. (a, b) Wide-field fluorescence images of Thy1-YFP-expressing mouse brain hemisphere slice before expansion
Correction: The Leukemia-Associated Mllt10/ Af10-Dot1l Are Tcf4/β-Catenin Coactivators Essential for Intestinal Homeostasis
 CORRECTION Correction: The Leukemia-Associated Mllt10/ Af10-Dot1l Are Tcf4/β-Catenin Coactivators Essential for Intestinal Homeostasis Tokameh Mahmoudi, Sylvia F. Boj, Pantelis Hatzis, Vivian S. W. Li,
CORRECTION Correction: The Leukemia-Associated Mllt10/ Af10-Dot1l Are Tcf4/β-Catenin Coactivators Essential for Intestinal Homeostasis Tokameh Mahmoudi, Sylvia F. Boj, Pantelis Hatzis, Vivian S. W. Li,
Supplemental Materials. Matrix Proteases Contribute to Progression of Pelvic Organ Prolapse in Mice and Humans
 Supplemental Materials Matrix Proteases Contribute to Progression of Pelvic Organ Prolapse in Mice and Humans Madhusudhan Budatha, Shayzreen Roshanravan, Qian Zheng, Cecilia Weislander, Shelby L. Chapman,
Supplemental Materials Matrix Proteases Contribute to Progression of Pelvic Organ Prolapse in Mice and Humans Madhusudhan Budatha, Shayzreen Roshanravan, Qian Zheng, Cecilia Weislander, Shelby L. Chapman,
GFP CCD2 GFP IP:GFP
 D1 D2 1 75 95 148 178 492 GFP CCD1 CCD2 CCD2 GFP D1 D2 GFP D1 D2 Beclin 1 IB:GFP IP:GFP Supplementary Figure 1: Mapping domains required for binding to HEK293T cells are transfected with EGFP-tagged mutant
D1 D2 1 75 95 148 178 492 GFP CCD1 CCD2 CCD2 GFP D1 D2 GFP D1 D2 Beclin 1 IB:GFP IP:GFP Supplementary Figure 1: Mapping domains required for binding to HEK293T cells are transfected with EGFP-tagged mutant
TRANSGENIC TECHNOLOGIES: Gene-targeting
 TRANSGENIC TECHNOLOGIES: Gene-targeting Reverse Genetics Wild-type Bmp7 -/- Forward Genetics Phenotype Gene or Mutations First Molecular Analysis Second Reverse Genetics Gene Phenotype or Molecular Analysis
TRANSGENIC TECHNOLOGIES: Gene-targeting Reverse Genetics Wild-type Bmp7 -/- Forward Genetics Phenotype Gene or Mutations First Molecular Analysis Second Reverse Genetics Gene Phenotype or Molecular Analysis
Supplemental Movie Legend.
 Supplemental Movie Legend. Transfected T cells were dropped onto SEE superantigen-pulsed Raji B cells (approximate location indicated by circle). Maximum-intensity projections from Z-stacks (17 slices,
Supplemental Movie Legend. Transfected T cells were dropped onto SEE superantigen-pulsed Raji B cells (approximate location indicated by circle). Maximum-intensity projections from Z-stacks (17 slices,
Supplemental Fig. 1. Transient expression of dynamic reporters of DNA methylation (DYNAMETs) in plants. (A) Schematic representation of potential
 Supplemental Fig. 1. Transient expression of dynamic reporters of DNA methylation (DYNAMETs) in plants. (A) Schematic representation of potential dynamic reporter of DNA methylation (DYNAMET). A DYNAMET
Supplemental Fig. 1. Transient expression of dynamic reporters of DNA methylation (DYNAMETs) in plants. (A) Schematic representation of potential dynamic reporter of DNA methylation (DYNAMET). A DYNAMET
Supplement Figure 1. Plin5 Plin2 Plin1. KDEL-DSRed. Plin-YFP. Merge
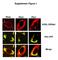 Supplement Figure 1 Plin5 Plin2 Plin1 KDEL-DSRed Plin-YFP Merge Supplement Figure 2 A. Plin5-Ab MitoTracker Merge AML12 B. Plin5-YFP Cytochrome c-cfp merge Supplement Figure 3 Ad.GFP Ad.Plin5 Supplement
Supplement Figure 1 Plin5 Plin2 Plin1 KDEL-DSRed Plin-YFP Merge Supplement Figure 2 A. Plin5-Ab MitoTracker Merge AML12 B. Plin5-YFP Cytochrome c-cfp merge Supplement Figure 3 Ad.GFP Ad.Plin5 Supplement
M I C R O B I O L O G Y WITH DISEASES BY TAXONOMY, THIRD EDITION
 M I C R O B I O L O G Y WITH DISEASES BY TAXONOMY, THIRD EDITION Chapter 7 Microbial Genetics Lecture prepared by Mindy Miller-Kittrell, University of Tennessee, Knoxville The Structure and Replication
M I C R O B I O L O G Y WITH DISEASES BY TAXONOMY, THIRD EDITION Chapter 7 Microbial Genetics Lecture prepared by Mindy Miller-Kittrell, University of Tennessee, Knoxville The Structure and Replication
RDH10 Oxidation of Vitamin A Is a Critical Control Step in Synthesis of Retinoic Acid during Mouse Embryogenesis
 RDH10 Oxidation of Vitamin A Is a Critical Control Step in Synthesis of Retinoic Acid during Mouse Embryogenesis Lisa L. Sandell 1,2 *, Megan L. Lynn 1, Kimberly E. Inman 1, William McDowell 1, Paul A.
RDH10 Oxidation of Vitamin A Is a Critical Control Step in Synthesis of Retinoic Acid during Mouse Embryogenesis Lisa L. Sandell 1,2 *, Megan L. Lynn 1, Kimberly E. Inman 1, William McDowell 1, Paul A.
MIT Department of Biology 7.013: Introductory Biology - Spring 2005 Instructors: Professor Hazel Sive, Professor Tyler Jacks, Dr.
 MIT Department of Biology 7.01: Introductory Biology - Spring 2005 Instructors: Professor Hazel Sive, Professor Tyler Jacks, Dr. Claudette Gardel iv) Would Xba I be useful for cloning? Why or why not?
MIT Department of Biology 7.01: Introductory Biology - Spring 2005 Instructors: Professor Hazel Sive, Professor Tyler Jacks, Dr. Claudette Gardel iv) Would Xba I be useful for cloning? Why or why not?
Protein Synthesis
 HEBISD Student Expectations: Identify that RNA Is a nucleic acid with a single strand of nucleotides Contains the 5-carbon sugar ribose Contains the nitrogen bases A, G, C and U instead of T. The U is
HEBISD Student Expectations: Identify that RNA Is a nucleic acid with a single strand of nucleotides Contains the 5-carbon sugar ribose Contains the nitrogen bases A, G, C and U instead of T. The U is
Supplementary information to accompany: A novel role for the DNA repair gene Rad51 in Netrin-1 signalling
 Supplementary information to accompany: A novel role for the DNA repair gene Rad51 in Netrin-1 signalling Glendining KA 1, Markie D 2, Gardner RJM 4, Franz EA 3, Robertson SP 4, Jasoni CL 1 Supplementary
Supplementary information to accompany: A novel role for the DNA repair gene Rad51 in Netrin-1 signalling Glendining KA 1, Markie D 2, Gardner RJM 4, Franz EA 3, Robertson SP 4, Jasoni CL 1 Supplementary
Supplemental Information. Boundary Formation through a Direct. Threshold-Based Readout. of Mobile Small RNA Gradients
 Developmental Cell, Volume 43 Supplemental Information Boundary Formation through a Direct Threshold-Based Readout of Mobile Small RNA Gradients Damianos S. Skopelitis, Anna H. Benkovics, Aman Y. Husbands,
Developmental Cell, Volume 43 Supplemental Information Boundary Formation through a Direct Threshold-Based Readout of Mobile Small RNA Gradients Damianos S. Skopelitis, Anna H. Benkovics, Aman Y. Husbands,
Make the protein through the genetic dogma process.
 Make the protein through the genetic dogma process. Coding Strand 5 AGCAATCATGGATTGGGTACATTTGTAACTGT 3 Template Strand mrna Protein Complete the table. DNA strand DNA s strand G mrna A C U G T A T Amino
Make the protein through the genetic dogma process. Coding Strand 5 AGCAATCATGGATTGGGTACATTTGTAACTGT 3 Template Strand mrna Protein Complete the table. DNA strand DNA s strand G mrna A C U G T A T Amino
Nature Methods: doi: /nmeth Supplementary Figure 1. Retention of RNA with LabelX.
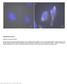 Supplementary Figure 1 Retention of RNA with LabelX. (a) Epi-fluorescence image of single molecule FISH (smfish) against GAPDH on HeLa cells expanded without LabelX treatment. (b) Epi-fluorescence image
Supplementary Figure 1 Retention of RNA with LabelX. (a) Epi-fluorescence image of single molecule FISH (smfish) against GAPDH on HeLa cells expanded without LabelX treatment. (b) Epi-fluorescence image
Supplemental Figure 1. Mutation in NLA Causes Increased Pi Uptake Activity and
 Supplemental Figure 1. Mutation in NLA Causes Increased Pi Uptake Activity and PHT1 Protein Amounts. (A) Shoot morphology of 19-day-old nla mutants under Pi-sufficient conditions. (B) [ 33 P]Pi uptake
Supplemental Figure 1. Mutation in NLA Causes Increased Pi Uptake Activity and PHT1 Protein Amounts. (A) Shoot morphology of 19-day-old nla mutants under Pi-sufficient conditions. (B) [ 33 P]Pi uptake
Bio11 Announcements. Ch 21: DNA Biology and Technology. DNA Functions. DNA and RNA Structure. How do DNA and RNA differ? What are genes?
 Bio11 Announcements TODAY Genetics (review) and quiz (CP #4) Structure and function of DNA Extra credit due today Next week in lab: Case study presentations Following week: Lab Quiz 2 Ch 21: DNA Biology
Bio11 Announcements TODAY Genetics (review) and quiz (CP #4) Structure and function of DNA Extra credit due today Next week in lab: Case study presentations Following week: Lab Quiz 2 Ch 21: DNA Biology
Themes: RNA and RNA Processing. Messenger RNA (mrna) What is a gene? RNA is very versatile! RNA-RNA interactions are very important!
 Themes: RNA is very versatile! RNA and RNA Processing Chapter 14 RNA-RNA interactions are very important! Prokaryotes and Eukaryotes have many important differences. Messenger RNA (mrna) Carries genetic
Themes: RNA is very versatile! RNA and RNA Processing Chapter 14 RNA-RNA interactions are very important! Prokaryotes and Eukaryotes have many important differences. Messenger RNA (mrna) Carries genetic
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION DOI: 10.1038/ncb3575 In the format provided by the authors and unedited. Supplementary Figure 1 Validation of key reagents and assays a, top, IHC with antibody recognizing specifically
SUPPLEMENTARY INFORMATION DOI: 10.1038/ncb3575 In the format provided by the authors and unedited. Supplementary Figure 1 Validation of key reagents and assays a, top, IHC with antibody recognizing specifically
Alternative Cleavage and Polyadenylation of RNA
 Developmental Cell 18 Supplemental Information The Spen Family Protein FPA Controls Alternative Cleavage and Polyadenylation of RNA Csaba Hornyik, Lionel C. Terzi, and Gordon G. Simpson Figure S1, related
Developmental Cell 18 Supplemental Information The Spen Family Protein FPA Controls Alternative Cleavage and Polyadenylation of RNA Csaba Hornyik, Lionel C. Terzi, and Gordon G. Simpson Figure S1, related
OPTICAL CONTROL OF TUMOR INDUCTION IN THE ZEBRAFISH
 OPTICAL CONTROL OF TUMOR INDUCTION IN THE ZEBRAFISH Zhiping Feng, 1,12 Suzy Nam, 2 Fatima Hamouri, 3,4 Isabelle Aujard, 5,6 Bertrand Ducos, 3,4 Sophie Vriz, 7,8 Michel Volovitch, 7,9 Ludovic Jullien, 5,6
OPTICAL CONTROL OF TUMOR INDUCTION IN THE ZEBRAFISH Zhiping Feng, 1,12 Suzy Nam, 2 Fatima Hamouri, 3,4 Isabelle Aujard, 5,6 Bertrand Ducos, 3,4 Sophie Vriz, 7,8 Michel Volovitch, 7,9 Ludovic Jullien, 5,6
TRANSCRIPTION COMPARISON OF DNA & RNA TRANSCRIPTION. Umm AL Qura University. Sugar Ribose Deoxyribose. Bases AUCG ATCG. Strand length Short Long
 Umm AL Qura University TRANSCRIPTION Dr Neda Bogari TRANSCRIPTION COMPARISON OF DNA & RNA RNA DNA Sugar Ribose Deoxyribose Bases AUCG ATCG Strand length Short Long No. strands One Two Helix Single Double
Umm AL Qura University TRANSCRIPTION Dr Neda Bogari TRANSCRIPTION COMPARISON OF DNA & RNA RNA DNA Sugar Ribose Deoxyribose Bases AUCG ATCG Strand length Short Long No. strands One Two Helix Single Double
ASPP1 Fw GGTTGGGAATCCACGTGTTG ASPP1 Rv GCCATATCTTGGAGCTCTGAGAG
 Supplemental Materials and Methods Plasmids: the following plasmids were used in the supplementary data: pwzl-myc- Lats2 (Aylon et al, 2006), pretrosuper-vector and pretrosuper-shp53 (generous gift of
Supplemental Materials and Methods Plasmids: the following plasmids were used in the supplementary data: pwzl-myc- Lats2 (Aylon et al, 2006), pretrosuper-vector and pretrosuper-shp53 (generous gift of
Chapter 13. From DNA to Protein
 Chapter 13 From DNA to Protein Proteins All proteins consist of polypeptide chains A linear sequence of amino acids Each chain corresponds to the nucleotide base sequenceof a gene The Path From Genes to
Chapter 13 From DNA to Protein Proteins All proteins consist of polypeptide chains A linear sequence of amino acids Each chain corresponds to the nucleotide base sequenceof a gene The Path From Genes to
Nature Neuroscience: doi: /nn Supplementary Figure 1
 Supplementary Figure 1 PCR-genotyping of the three mouse models used in this study and controls for behavioral experiments after semi-chronic Pten inhibition. a-c. DNA from App/Psen1 (a), Pten tg (b) and
Supplementary Figure 1 PCR-genotyping of the three mouse models used in this study and controls for behavioral experiments after semi-chronic Pten inhibition. a-c. DNA from App/Psen1 (a), Pten tg (b) and
A Level. A Level Biology. DNA Technology Questions. AQA, OCR, Edexcel. Name: Total Marks: Page 1
 AQA, OCR, Edexcel A Level A Level Biology DNA Technology Questions Name: Total Marks: Page 1 Q1.(a) (i) A mutation of a tumour suppressor gene can result in the formation of a tumour. Explain how.........(2)
AQA, OCR, Edexcel A Level A Level Biology DNA Technology Questions Name: Total Marks: Page 1 Q1.(a) (i) A mutation of a tumour suppressor gene can result in the formation of a tumour. Explain how.........(2)
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION Dynamic Phosphorylation of HP1 Regulates Mitotic Progression in Human Cells Supplementary Figures Supplementary Figure 1. NDR1 interacts with HP1. (a) Immunoprecipitation using
SUPPLEMENTARY INFORMATION Dynamic Phosphorylation of HP1 Regulates Mitotic Progression in Human Cells Supplementary Figures Supplementary Figure 1. NDR1 interacts with HP1. (a) Immunoprecipitation using
Construction of plant complementation vector and generation of transgenic plants
 MATERIAL S AND METHODS Plant materials and growth conditions Arabidopsis ecotype Columbia (Col0) was used for this study. SALK_072009, SALK_076309, and SALK_027645 were obtained from the Arabidopsis Biological
MATERIAL S AND METHODS Plant materials and growth conditions Arabidopsis ecotype Columbia (Col0) was used for this study. SALK_072009, SALK_076309, and SALK_027645 were obtained from the Arabidopsis Biological
A subclass of HSP70s regulate development and abiotic stress responses in Arabidopsis thaliana
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 Journal of Plant Research A subclass of HSP70s regulate development and abiotic stress responses in Arabidopsis thaliana Linna Leng 1 Qianqian Liang
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 Journal of Plant Research A subclass of HSP70s regulate development and abiotic stress responses in Arabidopsis thaliana Linna Leng 1 Qianqian Liang
Gene Expression Technology
 Gene Expression Technology Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Gene expression Gene expression is the process by which information from a gene
Gene Expression Technology Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Gene expression Gene expression is the process by which information from a gene
2054, Chap. 14, page 1
 2054, Chap. 14, page 1 I. Recombinant DNA technology (Chapter 14) A. recombinant DNA technology = collection of methods used to perform genetic engineering 1. genetic engineering = deliberate modification
2054, Chap. 14, page 1 I. Recombinant DNA technology (Chapter 14) A. recombinant DNA technology = collection of methods used to perform genetic engineering 1. genetic engineering = deliberate modification
Ramp1 EPD0843_4_B11. EUCOMM/KOMP-CSD Knockout-First Genotyping
 EUCOMM/KOMP-CSD Knockout-First Genotyping Introduction The majority of animals produced from the EUCOMM/KOMP-CSD ES cell resource contain the Knockout-First-Reporter Tagged Insertion allele. As well as
EUCOMM/KOMP-CSD Knockout-First Genotyping Introduction The majority of animals produced from the EUCOMM/KOMP-CSD ES cell resource contain the Knockout-First-Reporter Tagged Insertion allele. As well as
Usp14 EPD0582_2_G09. EUCOMM/KOMP-CSD Knockout-First Genotyping
 EUCOMM/KOMP-CSD Knockout-First Genotyping Introduction The majority of animals produced from the EUCOMM/KOMP-CSD ES cell resource contain the Knockout-First-Reporter Tagged Insertion allele. As well as
EUCOMM/KOMP-CSD Knockout-First Genotyping Introduction The majority of animals produced from the EUCOMM/KOMP-CSD ES cell resource contain the Knockout-First-Reporter Tagged Insertion allele. As well as
! Allele Interactions
 Chapter 4!Extensions to Mendelian Genetics! Allele Interactions 1 INTRODUCTION Mendelian inheritance describes inheritance patterns that obey two laws Law of segregation Law of independent assortment Simple
Chapter 4!Extensions to Mendelian Genetics! Allele Interactions 1 INTRODUCTION Mendelian inheritance describes inheritance patterns that obey two laws Law of segregation Law of independent assortment Simple
Figure S1. sporulation frequency (%) 80. sme2-5. sme2-3. sme2
 Figure S1 N WT -5-3 - DSRless mei4 100 ssm4 MS2loop () (kb) 1.0 0.5 sporulation frequency (%) 80 60 40 20 rrna 1 2 3 4 5 6 7 8 0 WT -5-3 -DSRless Figure S1. The DSR motifs in are crucial for its function.
Figure S1 N WT -5-3 - DSRless mei4 100 ssm4 MS2loop () (kb) 1.0 0.5 sporulation frequency (%) 80 60 40 20 rrna 1 2 3 4 5 6 7 8 0 WT -5-3 -DSRless Figure S1. The DSR motifs in are crucial for its function.
Cell autonomous vs. cell non-autonomous gene function
 Cell autonomous vs. cell non-autonomous gene function In multicellular organisms, it is important to know in what cell(s) the activity of a gene is required. - while RNA expression can be highly informative,
Cell autonomous vs. cell non-autonomous gene function In multicellular organisms, it is important to know in what cell(s) the activity of a gene is required. - while RNA expression can be highly informative,
Transcription Eukaryotic Cells
 Transcription Eukaryotic Cells Packet #20 1 Introduction Transcription is the process in which genetic information, stored in a strand of DNA (gene), is copied into a strand of RNA. Protein-encoding genes
Transcription Eukaryotic Cells Packet #20 1 Introduction Transcription is the process in which genetic information, stored in a strand of DNA (gene), is copied into a strand of RNA. Protein-encoding genes
WORKING WITH THE FIGURES. 1. In Figure 8-3, why are the arrows for genes 1 and 2 pointing in opposite directions?
 8 RNA: Transcription and Processing WORKING WITH THE FIGURES 1. In Figure 8-3, why are the arrows for genes 1 and 2 pointing in opposite directions? The arrows for genes 1 and 2 indicate the direction
8 RNA: Transcription and Processing WORKING WITH THE FIGURES 1. In Figure 8-3, why are the arrows for genes 1 and 2 pointing in opposite directions? The arrows for genes 1 and 2 indicate the direction
Differential Gene Expression
 Biology 4361 Developmental Biology Differential Gene Expression June 19, 2008 Differential Gene Expression Overview Chromatin structure Gene anatomy RNA processing and protein production Initiating transcription:
Biology 4361 Developmental Biology Differential Gene Expression June 19, 2008 Differential Gene Expression Overview Chromatin structure Gene anatomy RNA processing and protein production Initiating transcription:
Isolation, culture, and transfection of primary mammary epithelial organoids
 Supplementary Experimental Procedures Isolation, culture, and transfection of primary mammary epithelial organoids Primary mammary epithelial organoids were prepared from 8-week-old CD1 mice (Charles River)
Supplementary Experimental Procedures Isolation, culture, and transfection of primary mammary epithelial organoids Primary mammary epithelial organoids were prepared from 8-week-old CD1 mice (Charles River)
CHAPTER 9 DNA Technologies
 CHAPTER 9 DNA Technologies Recombinant DNA Artificially created DNA that combines sequences that do not occur together in the nature Basis of much of the modern molecular biology Molecular cloning of genes
CHAPTER 9 DNA Technologies Recombinant DNA Artificially created DNA that combines sequences that do not occur together in the nature Basis of much of the modern molecular biology Molecular cloning of genes
Gene mutation and DNA polymorphism
 Gene mutation and DNA polymorphism Outline of this chapter Gene Mutation DNA Polymorphism Gene Mutation Definition Major Types Definition A gene mutation is a change in the nucleotide sequence that composes
Gene mutation and DNA polymorphism Outline of this chapter Gene Mutation DNA Polymorphism Gene Mutation Definition Major Types Definition A gene mutation is a change in the nucleotide sequence that composes
Supplemental figures Supplemental Figure 1: Fluorescence recovery for FRAP experiments depicted in Figure 1.
 Supplemental figures Supplemental Figure 1: Fluorescence recovery for FRAP experiments depicted in Figure 1. Percent of original fluorescence was plotted as a function of time following photobleaching
Supplemental figures Supplemental Figure 1: Fluorescence recovery for FRAP experiments depicted in Figure 1. Percent of original fluorescence was plotted as a function of time following photobleaching
X2-C/X1-Y X2-C/VCAM-Y. FRET efficiency. Ratio YFP/CFP
 FRET efficiency.7.6..4.3.2 X2-C/X1-Y X2-C/VCAM-Y.1 1 2 3 Ratio YFP/CFP Supplemental Data 1. Analysis of / heterodimers in live cells using FRET. FRET saturation curves were obtained using cells transiently
FRET efficiency.7.6..4.3.2 X2-C/X1-Y X2-C/VCAM-Y.1 1 2 3 Ratio YFP/CFP Supplemental Data 1. Analysis of / heterodimers in live cells using FRET. FRET saturation curves were obtained using cells transiently
Transport of Potato Lipoxygenase into the Vacuole Larsen, Mia Kruse Guldstrand; Welinder, Karen Gjesing; Jørgensen, Malene
 Aalborg Universitet Transport of Potato Lipoxygenase into the Vacuole Larsen, Mia Kruse Guldstrand; Welinder, Karen Gjesing; Jørgensen, Malene Publication date: 2009 Document Version Publisher's PDF, also
Aalborg Universitet Transport of Potato Lipoxygenase into the Vacuole Larsen, Mia Kruse Guldstrand; Welinder, Karen Gjesing; Jørgensen, Malene Publication date: 2009 Document Version Publisher's PDF, also
Table S1. List of primers used in this study
 Table S1. List of primers used in this study Name KanMx-F2 KanMx-A2 FEN1-DG-S FEN1-DG-A SUR4-DG-S SUR4-DG-A CaARG4-R1130 CaARG4-F61 CaHIS1-DR CaHIS1-ter CaFEN1-US1 CaFEN1-UA1 CaFEN1-DS2 CaFEN1-DA2 CaFEN1-DG-S
Table S1. List of primers used in this study Name KanMx-F2 KanMx-A2 FEN1-DG-S FEN1-DG-A SUR4-DG-S SUR4-DG-A CaARG4-R1130 CaARG4-F61 CaHIS1-DR CaHIS1-ter CaFEN1-US1 CaFEN1-UA1 CaFEN1-DS2 CaFEN1-DA2 CaFEN1-DG-S
Year III Pharm.D Dr. V. Chitra
 Year III Pharm.D Dr. V. Chitra 1 Genome entire genetic material of an individual Transcriptome set of transcribed sequences Proteome set of proteins encoded by the genome 2 Only one strand of DNA serves
Year III Pharm.D Dr. V. Chitra 1 Genome entire genetic material of an individual Transcriptome set of transcribed sequences Proteome set of proteins encoded by the genome 2 Only one strand of DNA serves
The Genetic Code and Transcription. Chapter 12 Honors Genetics Ms. Susan Chabot
 The Genetic Code and Transcription Chapter 12 Honors Genetics Ms. Susan Chabot TRANSCRIPTION Copy SAME language DNA to RNA Nucleic Acid to Nucleic Acid TRANSLATION Copy DIFFERENT language RNA to Amino
The Genetic Code and Transcription Chapter 12 Honors Genetics Ms. Susan Chabot TRANSCRIPTION Copy SAME language DNA to RNA Nucleic Acid to Nucleic Acid TRANSLATION Copy DIFFERENT language RNA to Amino
Figure S1. Immunoblotting showing proper expression of tagged R and AVR proteins in N. benthamiana.
 SUPPLEMENTARY FIGURES Figure S1. Immunoblotting showing proper expression of tagged R and AVR proteins in N. benthamiana. YFP:, :CFP, HA: and (A) and HA, CFP and YFPtagged and AVR1-CO39 (B) were expressed
SUPPLEMENTARY FIGURES Figure S1. Immunoblotting showing proper expression of tagged R and AVR proteins in N. benthamiana. YFP:, :CFP, HA: and (A) and HA, CFP and YFPtagged and AVR1-CO39 (B) were expressed
Supplemental Data. Aung et al. (2011). Plant Cell /tpc
 35S pro:pmd1-yfp 10 µm Supplemental Figure 1. -terminal YFP fusion of PMD1 (PMD1-YFP, in green) is localized to the cytosol. grobacterium cells harboring 35S pro :PMD1-YFP were infiltrated into tobacco
35S pro:pmd1-yfp 10 µm Supplemental Figure 1. -terminal YFP fusion of PMD1 (PMD1-YFP, in green) is localized to the cytosol. grobacterium cells harboring 35S pro :PMD1-YFP were infiltrated into tobacco
pgbkt7 Anti- Myc AH109 strain (KDa) 50
 pgbkt7 (KDa) 50 37 Anti- Myc AH109 strain Supplementary Figure 1. Protein expression of CRN and TDR in yeast. To analyse the protein expression of CRNKD and TDRKD, total proteins extracted from yeast culture
pgbkt7 (KDa) 50 37 Anti- Myc AH109 strain Supplementary Figure 1. Protein expression of CRN and TDR in yeast. To analyse the protein expression of CRNKD and TDRKD, total proteins extracted from yeast culture
Protein Synthesis Notes
 Protein Synthesis Notes Protein Synthesis: Overview Transcription: synthesis of mrna under the direction of DNA. Translation: actual synthesis of a polypeptide under the direction of mrna. Transcription
Protein Synthesis Notes Protein Synthesis: Overview Transcription: synthesis of mrna under the direction of DNA. Translation: actual synthesis of a polypeptide under the direction of mrna. Transcription
PHT1;2-CFP YFP-PHF + PHT1;2-CFP YFP-PHF
 YFP-PHF1 CFP-PHT1;2 PHT1;2-CFP YFP-PHF + PHT1;2-CFP YFP-PHF + CFP-PHT1;2 Negative control!-gfp Supplemental Figure 1: PHT1;2 accumulation is PHF1 dependent. Immunoblot analysis on total protein extract
YFP-PHF1 CFP-PHT1;2 PHT1;2-CFP YFP-PHF + PHT1;2-CFP YFP-PHF + CFP-PHT1;2 Negative control!-gfp Supplemental Figure 1: PHT1;2 accumulation is PHF1 dependent. Immunoblot analysis on total protein extract
Chapter 13. The Nucleus. The nucleus is the hallmark of eukaryotic cells; the very term eukaryotic means having a "true nucleus".
 Chapter 13 The Nucleus The nucleus is the hallmark of eukaryotic cells; the very term eukaryotic means having a "true nucleus". Fig.13.1. The EM of the Nucleus of a Eukaryotic Cell 13.1. The Nuclear Envelope
Chapter 13 The Nucleus The nucleus is the hallmark of eukaryotic cells; the very term eukaryotic means having a "true nucleus". Fig.13.1. The EM of the Nucleus of a Eukaryotic Cell 13.1. The Nuclear Envelope
Heme utilization in the Caenorhabditis elegans hypodermal cells is facilitated by hemeresponsive
 Supplemental Data Heme utilization in the Caenorhabditis elegans hypodermal cells is facilitated by hemeresponsive gene-2 Caiyong Chen 1, Tamika K. Samuel 1, Michael Krause 2, Harry A. Dailey 3, and Iqbal
Supplemental Data Heme utilization in the Caenorhabditis elegans hypodermal cells is facilitated by hemeresponsive gene-2 Caiyong Chen 1, Tamika K. Samuel 1, Michael Krause 2, Harry A. Dailey 3, and Iqbal
CHAPTER 20 DNA TECHNOLOGY AND GENOMICS. Section A: DNA Cloning
 Section A: DNA Cloning 1. DNA technology makes it possible to clone genes for basic research and commercial applications: an overview 2. Restriction enzymes are used to make recombinant DNA 3. Genes can
Section A: DNA Cloning 1. DNA technology makes it possible to clone genes for basic research and commercial applications: an overview 2. Restriction enzymes are used to make recombinant DNA 3. Genes can
Eukaryotic Gene Structure
 Eukaryotic Gene Structure Terminology Genome entire genetic material of an individual Transcriptome set of transcribed sequences Proteome set of proteins encoded by the genome 2 Gene Basic physical and
Eukaryotic Gene Structure Terminology Genome entire genetic material of an individual Transcriptome set of transcribed sequences Proteome set of proteins encoded by the genome 2 Gene Basic physical and
Figure S1. gfp tola and pal mcherry can complement deletion mutants of tola and pal respectively. (A)When strain LS4522 was grown in the presence of
 Figure S1. gfp tola and pal mcherry can complement deletion mutants of tola and pal respectively. (A)When strain LS4522 was grown in the presence of xylose, inducing expression of gfp-tola, localization
Figure S1. gfp tola and pal mcherry can complement deletion mutants of tola and pal respectively. (A)When strain LS4522 was grown in the presence of xylose, inducing expression of gfp-tola, localization
Supplemental Table 1 Gene Symbol FDR corrected p-value PLOD1 CSRP2 PFKP ADFP ADM C10orf10 GPI LOX PLEKHA2 WIPF1
 Supplemental Table 1 Gene Symbol FDR corrected p-value PLOD1 4.52E-18 PDK1 6.77E-18 CSRP2 4.42E-17 PFKP 1.23E-14 MSH2 3.79E-13 NARF_A 5.56E-13 ADFP 5.56E-13 FAM13A1 1.56E-12 FAM29A_A 1.22E-11 CA9 1.54E-11
Supplemental Table 1 Gene Symbol FDR corrected p-value PLOD1 4.52E-18 PDK1 6.77E-18 CSRP2 4.42E-17 PFKP 1.23E-14 MSH2 3.79E-13 NARF_A 5.56E-13 ADFP 5.56E-13 FAM13A1 1.56E-12 FAM29A_A 1.22E-11 CA9 1.54E-11
Developmental Biology 3230 Exam 1 (Feb. 6) NAME
 DevelopmentalBiology3230Exam1(Feb.6)NAME 1. (10pts) What is a Fate Map? How would you experimentally acquire the data to draw a Fate Map? Explain what a Fate Map does and does not tell you about the mechanisms
DevelopmentalBiology3230Exam1(Feb.6)NAME 1. (10pts) What is a Fate Map? How would you experimentally acquire the data to draw a Fate Map? Explain what a Fate Map does and does not tell you about the mechanisms
TRANSCRIPTION AND PROCESSING OF RNA
 TRANSCRIPTION AND PROCESSING OF RNA 1. The steps of gene expression. 2. General characterization of transcription: steps, components of transcription apparatus. 3. Transcription of eukaryotic structural
TRANSCRIPTION AND PROCESSING OF RNA 1. The steps of gene expression. 2. General characterization of transcription: steps, components of transcription apparatus. 3. Transcription of eukaryotic structural
I. Prokaryotic Gene Regulation. Figure 1: Operon. Operon:
 I. Prokaryotic Gene Regulation Figure 1: Operon Operon: a) Regulatory Elements consist of an Operator that serves as the on-off switch for the genes of the operon. Also contains a promoter for the Structural
I. Prokaryotic Gene Regulation Figure 1: Operon Operon: a) Regulatory Elements consist of an Operator that serves as the on-off switch for the genes of the operon. Also contains a promoter for the Structural
Supplementary Figure 1. (a) The qrt-pcr for lnc-2, lnc-6 and lnc-7 RNA level in DU145, 22Rv1, wild type HCT116 and HCT116 Dicer ex5 cells transfected
 Supplementary Figure 1. (a) The qrt-pcr for lnc-2, lnc-6 and lnc-7 RNA level in DU145, 22Rv1, wild type HCT116 and HCT116 Dicer ex5 cells transfected with the sirna against lnc-2, lnc-6, lnc-7, and the
Supplementary Figure 1. (a) The qrt-pcr for lnc-2, lnc-6 and lnc-7 RNA level in DU145, 22Rv1, wild type HCT116 and HCT116 Dicer ex5 cells transfected with the sirna against lnc-2, lnc-6, lnc-7, and the
SUPPLEMENTAL MATERIALS
 SUPPLEMENL MERILS Eh-seq: RISPR epitope tagging hip-seq of DN-binding proteins Daniel Savic, E. hristopher Partridge, Kimberly M. Newberry, Sophia. Smith, Sarah K. Meadows, rian S. Roberts, Mark Mackiewicz,
SUPPLEMENL MERILS Eh-seq: RISPR epitope tagging hip-seq of DN-binding proteins Daniel Savic, E. hristopher Partridge, Kimberly M. Newberry, Sophia. Smith, Sarah K. Meadows, rian S. Roberts, Mark Mackiewicz,
7.2 Protein Synthesis. From DNA to Protein Animation
 7.2 Protein Synthesis From DNA to Protein Animation Proteins Why are proteins so important? They break down your food They build up muscles They send signals through your brain that control your body They
7.2 Protein Synthesis From DNA to Protein Animation Proteins Why are proteins so important? They break down your food They build up muscles They send signals through your brain that control your body They
Transcription. Unit: DNA. Central Dogma. 2. Transcription converts DNA into RNA. What is a gene? What is transcription? 1/7/2016
 Warm Up Questions 1. Where is DNA located? 2. Name the 3 parts of a nucleotide. 3. Enzymes can catalyze many different reactions (T or F) 4. How many variables should you have in an experiment? 5. A red
Warm Up Questions 1. Where is DNA located? 2. Name the 3 parts of a nucleotide. 3. Enzymes can catalyze many different reactions (T or F) 4. How many variables should you have in an experiment? 5. A red
IP-Free Electra DAUGHTER Vectors Mammalian with CMV promoter
 IP-Free Electra DAUGHTER Vectors Mammalian with CMV promoter Electra cloning DNA2.0 has developed a simple one-tube universal cloning process that can be performed in a 5 minute bench-top reaction with
IP-Free Electra DAUGHTER Vectors Mammalian with CMV promoter Electra cloning DNA2.0 has developed a simple one-tube universal cloning process that can be performed in a 5 minute bench-top reaction with
 Watson BM Gene Capitolo 11 Watson et al., BIOLOGIA MOLECOLARE DEL GENE, Zanichelli editore S.p.A. ? Le proteine della trasposizione Watson et al., BIOLOGIA MOLECOLARE DEL GENE, Zanichelli editore S.p.A.
Watson BM Gene Capitolo 11 Watson et al., BIOLOGIA MOLECOLARE DEL GENE, Zanichelli editore S.p.A. ? Le proteine della trasposizione Watson et al., BIOLOGIA MOLECOLARE DEL GENE, Zanichelli editore S.p.A.
Multiple choice questions (numbers in brackets indicate the number of correct answers)
 1 Multiple choice questions (numbers in brackets indicate the number of correct answers) February 1, 2013 1. Ribose is found in Nucleic acids Proteins Lipids RNA DNA (2) 2. Most RNA in cells is transfer
1 Multiple choice questions (numbers in brackets indicate the number of correct answers) February 1, 2013 1. Ribose is found in Nucleic acids Proteins Lipids RNA DNA (2) 2. Most RNA in cells is transfer
