Tech Note. Using the ncounter Analysis System with FFPE Samples for Gene Expression Analysis. ncounter Gene Expression. Molecules That Count
|
|
|
- Patricia Powers
- 6 years ago
- Views:
Transcription
1 ncounter Gene Expression Tech Note Using the ncounter Analysis System with FFPE Samples for Gene Expression Analysis Introduction For the past several decades, pathologists have kept samples obtained from tissue resections and biopsies. Typically, these are archived as Formalin-Fixed Paraffin Embedded (FFPE) blocks, and in some cases are sectioned and preserved on slides. These archived FFPE clinical samples are stable for years, and are valuable resources for subsequent investigations. Diagnosis, treatment, disease progression, and other clinical information are associated with many of these samples, making them extremely valuable for clinical research. Moreover, pathologists currently use FFPE blocks and slices as the preferred method to preserve samples from new patients. The ability of new molecular diagnostic tests to use FFPE samples as input material would greatly increase the usability of these tests. However, the fixation and embedding process modifies and degrades RNA, presenting challenges for gene expression studies using these samples. Other currently available gene expression profiling methods such as PCR-based techniques and microarrays have shown a significant decrease in the quality of results when using FFPE samples as input material. In this technical note, we demonstrate that the ncounter System accurately measures gene expression in RNA extracted from FFPE samples - even though RNA from FFPE samples is usually more degraded than RNA from other sources. Our results show high correlations of transcript levels between freshly prepared RNA and RNA extracted from matched FFPE samples across various tissues. We also show that the ncounter System can accurately measure transcript levels in crude extracts derived from FFPE samples, without RNA purification. Finally, we present guidelines for assessing RNA quality prior to running an ncounter analysis. ncounter Analysis System Overview The NanoString ncounter Analysis System delivers direct, multiplexed measurement of gene expression, providing digital readouts of the relative abundance of hundreds of mrna transcripts simultaneously. The ncounter Analysis System is based on gene-specific probe pairs that are hybridized to the sample in solution. The protocol eliminates any enzymatic reactions that might introduce bias in the results (FIGURE, Step ). The Reporter Probe carries the fluorescent signal; the Capture Probe allows the complex to be immobilized for data collection. Up to 55 pairs of probes specific for a particular set of genes are combined with a series of internal controls to form a CodeSet. After hybridization of the CodeSet with target mrna, samples are transferred to the ncounter Prep Station (FIGURE, Step ) where excess probes are removed and probe / target complexes are aligned and immobilized in the ncounter Cartridge. Cartridges are then placed in the ncounter Digital Analyzer for data collection (FIGURE, Step 3). Each target molecule of interest is identified by the color code generated by six ordered fluorescent spots present on the reporter probe. The Reporter Probes on the surface of the cartridge are then counted and tabulated for each target molecule. Hybridize Purify and Immobilize Count Molecules That Count Translational Research Gene Expression mirna Expression Copy Number Variation
2 Frozen Tissue / Cryopreserved Sections / FFPE Sections: Matched human heart tissue samples preserved as flash-frozen tissue, cryopreserved sections, and FFPE sections were obtained from iochain Institute Inc. Total RNA was purified from flash-frozen tissues using the QIAGEN RNeasy Kit according to the manufacturer s instructions. RNA was extracted from cryopreserved sections using the QIAGEN Fibrous Tissue RNeasy Mini-Kit according to the manufacturer s instructions. RNA was extracted from FFPE slices using the QIAGEN RNeasy FFPE Kit, according to the manufacturer s instructions. One hundred nanograms of RNA from each sample was hybridized in triplicate to a 96-plex human gene CodeSet and processed on the ncounter platform. The counts were averaged for each set of triplicates and normalized according to the standard protocol (Geiss et al., 8). Tumor / Normal; Frozen Tissue / FFPE Sections: Purified total RNA samples from frozen and FFPE biopsies of normal lung, lung tumor, normal liver, and liver tumor were provided by a customer. One hundred nanograms of RNA from each sample was hybridized to a 8-plex human gene CodeSet, processed on the ncounter platform, and normalized to beta-actin according to the standard protocol. Crude FFPE Extracts: Crude FFPE extracts were prepared from heart and brain FFPE slices by deparaffinizing the slices and digesting with proteinase K. For comparison, total RNA was extracted from frozen heart and brain tissue and FFPE slices. The FFPE crude extracts and total RNA samples were hybrized to a 96- plex human gene CodeSet (different from that used for heart), processed on the ncounter platform, and normalized to positive controls according to the standard protocol. RNA Degradation Studies: Controlled RNA degradation studies were conducted by incubating Universal Human Reference RNA (Stratagene ) and Human rain Reference RNA (Ambion ) at 95 C for three minutes. One hundred nanograms of RNA from each sample was hybridized to a 96-plex human gene CodeSet, processed on the ncounter platform, and normalized according to the standard protocol. RNA Quality: To assess RNA quality, all samples were run on an Agilent ioanalyzer. The degree of RNA integrity was assessed using the smear analysis function in the Agilent Expert Software to measure the percentage of RNA molecules that were greater than 3bp. ncounter Analysis of FFPE Total Heart RNA We obtained tissue samples from a single human heart preserved as flash-frozen pieces, cryropreserved slices, and FFPE slices. We first assessed the amount of degradation in RNA from each sample by smear analysis using the Agilent ioanalyzer. In the RNA from fresh heart tissue, 9 percent of the transcripts were larger than 3bp and the 8s and 8s ribosomal RNA peaks were intact. The RNA from frozen heart sections also had intact ribosomal RNA peaks and 95 percent was greater than 3bp. In the RNA from the heart FFPE sections, only 6 percent was greater than 3bp. Also, the 8s and 8s ribosomal peaks were missing, indicating that RNA in the FFPE sample was degraded (FIGURE A). Gene expression in total RNA from the frozen pieces of human heart tissue was compared to expression in RNA from cryopreserved samples and FFPE slices. The ncounter assay results for RNA from cryopreserved slices and FFPE samples were highly correlated with results for total RNA from flash-frozen tissue (FIGURE ). 3 heart FFPE heart total RNA heart frozen 5 5 Counts from Preserved Slices Cryropreserved y =.7878x R =.9956 FFPE Preserved y =.58x R = FIGURE A. ioanalyzer analysis of RNA extracted from flash-frozen (grey line), cryropreserved (green line) and FFPE (orange line) archived human heart samples. FIGURE. Comparison of counts of 96 genes in ng of total heart RNA versus ng total RNA prepared from frozen heart slices (green points) and total RNA prepared from FFPE heart slices (orange points).
3 NanoString Technologies TECH NOTE A Counts from FFPE Tissue Normal Lung y =.8x R =.936 Lung Tumor y =.3x +.65 R =.9398 Counts from FFPE Tissue Normal Liver y =.8x R =.96 LiverTumor y =.95x R =.9859 FIGURE 3. Correlations between counts for RNA samples extracted from fresh-frozen or FFPE tissues and hybridized to a 8-gene CodeSet. A) Counts for RNA from FFPE and fresh-frozen tissue for normal lung (orange points) and lung tumor (green points). ) Counts for RNA from FFPE and fresh-frozen tissue for normal liver (orange points) and liver tumor (green points). When compared to total RNA, the expression levels of genes in the RNA from cryopreserved slices showed excellent correlation (R =.995). RNA from the FFPE samples was also highly correlated to total RNA (R =.976). However, there was an overall reduction in raw counts in the FFPE sample as compared with the frozen slices. This is illustrated by the reduced slope of the regression analysis for the FFPE sample and is likely due to its greater degree of RNA degradation. Since there is good correlation of expression among genes in the entire set, this reduction does not affect analyses of fold changes in expression levels. Also, the reduction in counts can be corrected by adding more input RNA (data not shown). The high correlations in counts and fold changes across these samples show that the ncounter System can accurately measure gene expression even when the target RNA is degraded. ncounter Analysis of FFPE Total Lung and Liver RNA We further analyzed the performance of FFPE-derived RNA on the ncounter System in collaboration with a customer. Purified total RNA samples from frozen and FFPE biopsies of normal lung, lung tumor, normal liver, and liver tumor were provided to NanoString. These samples were analyzed using a 8-gene CodeSet, normalized to beta-actin for RNA content, and the counts in the frozen- and FFPE-paired tissues were compared (FIGURE 3). In each case, the beta-actin-normalized counts were very comparable; the correlation coefficients between frozen and FFPE samples were all greater than.9. ecause these samples were from different patients, some of the differences may also be due to biological variation. These results show that RNA from FFPE samples performs well on the ncounter system, despite the greater RNA degradation inherent in FFPE samples. ncounter Analysis of Crude FFPE Extracts without RNA Purification ecause of the enzyme-free nature of the ncounter System, good results can be obtained with cell and tissue extracts and unpurified RNA (see Sample Flexibility technical note). This raised the possibility that the ncounter System could also accurately analyze FFPE samples without RNA purification. To verify this, we hybridized crude extracts from FFPE slices of human heart and brain in an ncounter assay with a different 96-gene CodeSet. For comparison, we also hybridized purified total heart and total brain RNA extracted from the frozen tissue and the FFPE slices. A Counts in FFPE Sample rain FFPE Purified RNA y =.869x +.3 R =.933 rain FFPE Extract y =.7758x +.6 R =.98 Counts in FFPE Sample Heart FFPE Purified RNA y =.8389x -.79 R =.959 Heart FFPE Extract y =.795x R =.96 R =.995 R =.995 FIGURE. Correlation of counts from FFPE extracts to counts from purified RNA from FFPE slices and flash-frozen tissue. A) Correlation between brain FFPE extract (orange) and brain FFPE purified RNA (green) to purified RNA from frozen brain tissue. (A inset) Correlation between brain FFPE extract and purified RNA from brain FFPE tissue. ) Correlation between heart FFPE extract (orange) and heart FFPE purified RNA (green) to purifed RNA from frozen heart tissue. ( inset) Correlation between heart FFPE extract and purified RNA from heart FFPE tissue. Molecules That Count 3
4 The ncounter System performed well when using FFPE extracts as starting material. The counts from FFPE extract were highly correlated to counts from RNA purified from FFPE slices and from flash-frozen tissue (FIGURE ). In brain and heart, the correlation between the FFPE extract and RNA purified from FFPE was very high (R =.99) (FIGURE A inset and inset). The correlation between the brain FFPE extract and total RNA from frozen brain tissue was also high (R =.93) (FIGURE A). In heart, the correlation between FFPE extract and total RNA from frozen heart tissue was again high (R =.96) (FIGURE ). Since the counts from FFPE extracts, RNA from FFPE, and total RNA from frozen tissue were all highly correlated, we expected the fold-changes derived from these counts to be highly correlated as well. We compared the ratios of transcript abundance in brain versus heart in the three sample preparation methods (FIGURE C). The correlation between fold changes for RNA from frozen tissue and RNA from FFPE tissue was very high (R =.9); the correlation of fold changes for RNA from frozen tissue and RNA from FFPE extracts was slightly lower, but still good (R =.86). C Log Fold Change (Purified Total RNA) Purified FFPE RNA y =.98x +.3 R =.96 FFPE Extract y =.96x -.98 R =.8688 FIGURE. C) Fold changes of counts in heart versus brain. Fold changes for RNA purified from frozen heart and brain tissue are on the x-axis. Fold changes for FFPE extracts (orange points) or purified RNA from FFPE samples (green points) are on the y-axis. These results suggest that crude FFPE extracts containing degraded RNA can be used to accurately quantify gene expression levels and fold changes on the ncounter platform. The results for FFPE extracts show strong correlation with intact and purified RNA samples. Controlled RNA Degradation Study on the ncounter ecause RNA extracted from FFPE samples is usually more degraded than RNA from other sources, we conducted a controlled experiment to further characterize the effects of degraded RNA on the ncounter assay. We degraded the Universal Human Reference (UHR) and Human rain Reference (RN) by incubating at 95 C. After three minutes, RNA in the treated samples was substantially degraded. In the UHR sample, only 56 percent was greater than 3bp and in the RN sample, only 57 percent was greater than 3bp. For comparison, the untreated UHR and RN samples were intact, with 9 and 97 percent greater than 3bp, respectively. We hybridized each of the samples with a 98-gene CodeSet, and analyzed the correlation between intact and fragmented samples for UHR and RN. From the degraded UHR sample, we recovered approximately 7 percent of the counts of the intact UHR sample, and the counts were highly correlated (R =.98). In the degraded RN sample, we recovered approximately 6 percent of the counts of the intact RN sample and the correlation was again very high (R =.98)(FIGURE 5A). Finally, we compared the fold changes for degraded UHR versus degraded RN and intact UHR versus intact RN (FIGURE 5). ecause the correlation of counts in the degraded and intact samples was high, we also measured excellent correlation between the fold changes of counts in the UHR and RN samples (R =.958). These experiments demonstrate that the ncounter system can accurately measure gene expression even in samples where the RNA is substantially fragmented Log Fold Change (FFPE Sample) A Counts in Fragmented RNA UHR y =.77x +.79 R =.988 RN y =.66x R =.98 Log Fold Change (Fragmented RNA) y =.9383x +.79 R = Log Fold Change (Unfragmented RNA) FIGURE 5. ncounter analysis of heat-fragmented Universal Reference RNA (UHR) and Human rain (RN) RNA using a 98-gene CodeSet. A) Correlation of counts between unfragmented and fragmented UHR sample (green points) and RN sample (orange points). ) Fold change comparison between UHR and RN for counts from fragmented and unfragmented samples.
5 A. Percent Counts Relative to Control 8 6 Correlation with Control (R ) Fragmentation (Percent Larger than 3bp) Fragmentation (Percent Larger than 3bp) FIGURE 6. Meta-analysis of degree of RNA fragmentation and data quality in ncounter assays. A) Percent counts relative to intact RNA as a function of RNA fragmentation. ) Correlation between counts from degraded and intact RNA as a function of RNA fragmentation. Correlating ncounter Assay Performance with RNA Degradation Although the ncounter System generates data from fragmented RNA that correlates well with intact RNA, at some point the RNA becomes too degraded to produce good correlations. Therefore, we characterized the maximum amount of RNA degradation that still produces highly correlated data with intact RNA. We performed a meta-analysis of all our results using fragmented RNA, including some data described above and other data from degraded and FFPE-extracted samples not shown in this paper. We first compared the percentage of counts recovered from degraded RNA relative to intact RNA to the percentage of transcripts greater than 3bp (FIGURE 6A). Although there is much variation, it is clear that as RNA becomes more degraded, fewer counts are recovered. We found that if more than 5 percent of the RNA is greater than 3bp, the correlation with intact RNA can be.9 or greater (FIGURE 6). If less than 5 percent is greater than 3bp, good results can still be obtained, but the correlation with intact RNA may be lower. To obtain better results with such samples, it may be useful to increase the amount of RNA used in the hybridization. This would increase the number of fragments with intact target binding sites. Although these results are typical of what the ncounter System generates, results with different CodeSets and gene expression levels may vary from those presented here. Conclusions Gene expression analysis of FFPE samples is especially challenging because the RNA is usually more degraded than RNA from other sources. In this technical note, we demonstrated that the enzyme-free nature of the ncounter System enables accurate gene expression analysis of FFPE samples. On the ncounter System, expression data from RNA purified from FFPE samples is highly correlated with data from RNA from matched fresh samples. Furthermore, the ncounter System delivers accurate gene expression data directly from crude FFPE extracts, without requiring RNA purification. These capabilities make the ncounter System ideal for various types of clinical research studies and signature development, especially in cancer research. Acknowledgements We acknowledge the Sloan-Kettering Institute for providing purified RNA from frozen and FFPE biopsies of normal and tumor samples from lung and liver. References Geiss GK et al., Direct multiplexed measurement of gene expression with color-coded probe pairs. Nat iotech 6(3):37-35 (8). NanoString Technologies, Sample Flexibility for Gene Expression Analysis with the ncounter Analysis System. (9). NanoString Technologies, Inc. Contact Us Sales Contacts 53 Fairview Ave N Suite Seattle, Washington 989 info@nanostring.com United States: us.sales@nanostring.com Tel: (888) Europe: europe.sales@nanostring.com Fax: (6) Japan: japan.sales@nanostring.com Other Regions: info@nanostring.com NanoString Technologies, Inc. All rights reserved. NanoString, NanoString Technologies, ncounter, Molecules That Count, nsolver, Plex, ChIP-String and mirge are registered trademarks or trademarks of NanoString Technologies, Inc., ( NanoString ) in the United States and/or other countries. All other trademarks and or service marks not owned by NanoString that appear in this document are the property of their respective owners. The manufacture, use and or sale of NanoString product(s) may be subject to one or more patents or pending patent applications owned by NanoString or licensed to NanoString from Life Technologies Corporation and other third parties. For research use only. Not for use in diagnostic procedures. v.7
Sample Flexibility for Gene Expression Analysis with the ncounter Analysis System
 Technical Note Sample Flexibility for Gene Expression Analysis with the ncounter Analysis System Sensitive Gene Expression Analysis From Crude Preparations and Fragmented RNA Cell Lysates Tissue Homogenates
Technical Note Sample Flexibility for Gene Expression Analysis with the ncounter Analysis System Sensitive Gene Expression Analysis From Crude Preparations and Fragmented RNA Cell Lysates Tissue Homogenates
ncounter Elements TagSets General Purpose Reagents PACKAGE INSERT Molecules That Count NanoString Technologies, Inc.
 ncounter Elements TagSets General Purpose Reagents PACKAGE INSERT NanoString Technologies, Inc. 530 Fairview Ave N Suite 2000 Seattle, Washington 98109 www.nanostring.com Tel: 206.378.6266 888.358.6266
ncounter Elements TagSets General Purpose Reagents PACKAGE INSERT NanoString Technologies, Inc. 530 Fairview Ave N Suite 2000 Seattle, Washington 98109 www.nanostring.com Tel: 206.378.6266 888.358.6266
Developing an Accurate and Precise Companion Diagnostic Assay for Targeted Therapies in DLBCL
 Developing an Accurate and Precise Companion Diagnostic Assay for Targeted Therapies in DLBCL James Storhoff, Ph.D. Senior Manager, Diagnostic Test Development World Cdx, Boston, Sep. 10th Molecules That
Developing an Accurate and Precise Companion Diagnostic Assay for Targeted Therapies in DLBCL James Storhoff, Ph.D. Senior Manager, Diagnostic Test Development World Cdx, Boston, Sep. 10th Molecules That
Fusion Gene Analysis. ncounter Elements. Molecules That Count WHITE PAPER. v1.0 OCTOBER 2014 NanoString Technologies, Inc.
 Fusion Gene Analysis NanoString Technologies, Inc. Fusion Gene Analysis ncounter Elements v1.0 OCTOBER 2014 NanoString Technologies, Inc., Seattle WA 98109 FOR RESEARCH USE ONLY. Not for use in diagnostic
Fusion Gene Analysis NanoString Technologies, Inc. Fusion Gene Analysis ncounter Elements v1.0 OCTOBER 2014 NanoString Technologies, Inc., Seattle WA 98109 FOR RESEARCH USE ONLY. Not for use in diagnostic
ncounter Vantage 3D RNA:Protein Immune Cell Signaling Panel for Cell Suspensions
 ncounter Vantage 3D RNA:Protein Immune Cell Signaling Panel for Cell Suspensions with Universal Cell Capture Kit Intracellular Compatible User Manual NanoString Technologies, Inc. 530 Fairview Ave North
ncounter Vantage 3D RNA:Protein Immune Cell Signaling Panel for Cell Suspensions with Universal Cell Capture Kit Intracellular Compatible User Manual NanoString Technologies, Inc. 530 Fairview Ave North
FFPE DNA Extraction kit
 FFPE DNA Extraction kit Instruction Manual FFPE DNA Extraction kit Cat. No. C20000030, Format 50 rxns Version 1 / 14.08.13 Content Introduction 4 Overview of FFPE DNA extraction workflow 4 Kit contents
FFPE DNA Extraction kit Instruction Manual FFPE DNA Extraction kit Cat. No. C20000030, Format 50 rxns Version 1 / 14.08.13 Content Introduction 4 Overview of FFPE DNA extraction workflow 4 Kit contents
Comprehensive Report on FFPE Extraction Methods
 Technical Note Comprehensive Report on FFPE Extraction Methods Mutation detection based on DNA or RNA sequencing is becoming increasingly important in both human disease research and clinical settings.
Technical Note Comprehensive Report on FFPE Extraction Methods Mutation detection based on DNA or RNA sequencing is becoming increasingly important in both human disease research and clinical settings.
NanoString Sample Submission Guidelines
 High quality ncounter data is achieved with high quality starting material. The guidelines set forth in this document will assist you in preparing your samples for your NanoString run. Please adhere to
High quality ncounter data is achieved with high quality starting material. The guidelines set forth in this document will assist you in preparing your samples for your NanoString run. Please adhere to
NanoString Assay Sample Type Assay Input Amount Required Sample Amount/Concentration Total RNA 100ng no less than 150ng, normalized to 20ng/µl
 High quality ncounter data is achieved with high quality starting material. The guidelines set forth in this document will assist you in preparing your samples for your NanoString run. Please adhere to
High quality ncounter data is achieved with high quality starting material. The guidelines set forth in this document will assist you in preparing your samples for your NanoString run. Please adhere to
FirePlex mirna Assay. Multiplex microrna profiling from low sample inputs
 FirePlex mirna Assay Multiplex microrna profiling from low sample inputs Abstract We introduce a new assay for multiplex microrna (mirna) discovery and verification that enables simultaneous profiling
FirePlex mirna Assay Multiplex microrna profiling from low sample inputs Abstract We introduce a new assay for multiplex microrna (mirna) discovery and verification that enables simultaneous profiling
3.1.4 DNA Microarray Technology
 3.1.4 DNA Microarray Technology Scientists have discovered that one of the differences between healthy and cancer is which genes are turned on in each. Scientists can compare the gene expression patterns
3.1.4 DNA Microarray Technology Scientists have discovered that one of the differences between healthy and cancer is which genes are turned on in each. Scientists can compare the gene expression patterns
Unlocking your FFPE archive
 Unlocking your FFPE archive Critical factors for molecular analysis of FFPE samples Sample & Assay Technologies Table of contents 1. 2. Introduction Important considerations when preparing and archiving
Unlocking your FFPE archive Critical factors for molecular analysis of FFPE samples Sample & Assay Technologies Table of contents 1. 2. Introduction Important considerations when preparing and archiving
SOLiD Total RNA-Seq Kit SOLiD RNA Barcoding Kit
 SOLiD Total RNA-Seq Kit SOLiD RNA Barcoding Kit Agenda SOLiD Total RNAseq Kit Overview Kit Configurations Barcoding Kit Introduction New Small RNA and WT Workflow Small RNA Workflow Step-by-step Workflow
SOLiD Total RNA-Seq Kit SOLiD RNA Barcoding Kit Agenda SOLiD Total RNAseq Kit Overview Kit Configurations Barcoding Kit Introduction New Small RNA and WT Workflow Small RNA Workflow Step-by-step Workflow
MMLV Reverse Transcriptase 1st-Strand cdna Synthesis Kit
 MMLV Reverse Transcriptase 1st-Strand cdna Synthesis Kit Cat. No. MM070150 Available exclusively thru Lucigen. lucigen.com/epibio www.lucigen.com MA265E MMLV Reverse Transcriptase 1st-Strand cdna Synthesis
MMLV Reverse Transcriptase 1st-Strand cdna Synthesis Kit Cat. No. MM070150 Available exclusively thru Lucigen. lucigen.com/epibio www.lucigen.com MA265E MMLV Reverse Transcriptase 1st-Strand cdna Synthesis
MicroRNA profiling directly from low amounts of plasma or serum using the Multiplex Circulating mirna Assay with Firefly particle technology
 Technical note MicroRNA profiling directly from low amounts of plasma or serum using the Multiplex Circulating mirna Assay with Firefly particle technology Abstract We introduce a new assay that enables
Technical note MicroRNA profiling directly from low amounts of plasma or serum using the Multiplex Circulating mirna Assay with Firefly particle technology Abstract We introduce a new assay that enables
SMARTer Ultra Low RNA Kit for Illumina Sequencing Two powerful technologies combine to enable sequencing with ultra-low levels of RNA
 SMARTer Ultra Low RNA Kit for Illumina Sequencing Two powerful technologies combine to enable sequencing with ultra-low levels of RNA The most sensitive cdna synthesis technology, combined with next-generation
SMARTer Ultra Low RNA Kit for Illumina Sequencing Two powerful technologies combine to enable sequencing with ultra-low levels of RNA The most sensitive cdna synthesis technology, combined with next-generation
Assay I nput Amount. varied (amplification 1 cell or 10pg RNA 10pg- 10ng. 100ng. 1-3ul. varied (5-10µL)
 High quality ncounter data is achieved with samples that meet NanoString assay requirements. The guidelines set forth in this document will assist you in preparing your samples prior to submission to NanoString.
High quality ncounter data is achieved with samples that meet NanoString assay requirements. The guidelines set forth in this document will assist you in preparing your samples prior to submission to NanoString.
Pinpoint Slide DNA Isolation System Catalog No. D3001
 INSTRUCTIONS Pinpoint Slide DNA Isolation System Catalog No. D3001 Highlights Easily isolates genomic DNA in any targeted microscopic tissue area on a slide. The simple procedure combines Pinpoint tissue
INSTRUCTIONS Pinpoint Slide DNA Isolation System Catalog No. D3001 Highlights Easily isolates genomic DNA in any targeted microscopic tissue area on a slide. The simple procedure combines Pinpoint tissue
NEBNext rrna Depletion Kit (Human/Mouse/Rat)
 LIBRARY PREPARATION NEBNext rrna Depletion Kit (Human/Mouse/Rat) Instruction Manual NEB #E6310S/L/X, #E6350S/L/X 6/24/96 reactions Version 4.0 4/17 be INSPIRED drive DISCOVERY stay GENUINE This product
LIBRARY PREPARATION NEBNext rrna Depletion Kit (Human/Mouse/Rat) Instruction Manual NEB #E6310S/L/X, #E6350S/L/X 6/24/96 reactions Version 4.0 4/17 be INSPIRED drive DISCOVERY stay GENUINE This product
ReliaPrep FFPE gdna Miniprep System
 TECHNICAL MANUAL ReliaPrep FFPE gdna Miniprep System Instructions for Use of Products A2351 and A2352 Revised 12/15 TM352 ReliaPrep FFPE gdna Miniprep System All technical literature is available at: www.promega.com/protocols/
TECHNICAL MANUAL ReliaPrep FFPE gdna Miniprep System Instructions for Use of Products A2351 and A2352 Revised 12/15 TM352 ReliaPrep FFPE gdna Miniprep System All technical literature is available at: www.promega.com/protocols/
Gene Regulation Solutions. Microarrays and Next-Generation Sequencing
 Gene Regulation Solutions Microarrays and Next-Generation Sequencing Gene Regulation Solutions The Microarrays Advantage Microarrays Lead the Industry in: Comprehensive Content SurePrint G3 Human Gene
Gene Regulation Solutions Microarrays and Next-Generation Sequencing Gene Regulation Solutions The Microarrays Advantage Microarrays Lead the Industry in: Comprehensive Content SurePrint G3 Human Gene
DNA Microarray Technology
 CHAPTER 1 DNA Microarray Technology All living organisms are composed of cells. As a functional unit, each cell can make copies of itself, and this process depends on a proper replication of the genetic
CHAPTER 1 DNA Microarray Technology All living organisms are composed of cells. As a functional unit, each cell can make copies of itself, and this process depends on a proper replication of the genetic
TECHNICAL BULLETIN. SeqPlex DNA Amplification Kit for use with high throughput sequencing technologies. Catalog Number SEQX Storage Temperature 20 C
 SeqPlex DNA Amplification Kit for use with high throughput sequencing technologies Catalog Number SEQX Storage Temperature 20 C TECHNICAL BULLETIN Product Description The SeqPlex DNA Amplification Kit
SeqPlex DNA Amplification Kit for use with high throughput sequencing technologies Catalog Number SEQX Storage Temperature 20 C TECHNICAL BULLETIN Product Description The SeqPlex DNA Amplification Kit
Implementation of Automated Sample Quality Control in Whole Exome Sequencing
 Journal of Life Sciences 11 (2017) 261-268 doi: 10.17265/1934-7391/2017.06.001 D DAVID PUBLISHING Implementation of Automated Sample Quality Control in Whole Exome Sequencing Elisa Viering 1, Jana Molitor
Journal of Life Sciences 11 (2017) 261-268 doi: 10.17265/1934-7391/2017.06.001 D DAVID PUBLISHING Implementation of Automated Sample Quality Control in Whole Exome Sequencing Elisa Viering 1, Jana Molitor
Gene Expression Technology
 Gene Expression Technology Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Gene expression Gene expression is the process by which information from a gene
Gene Expression Technology Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Gene expression Gene expression is the process by which information from a gene
Optimizing FFPE DNA preparation for use in SureSelect XT2
 Optimizing PE DA preparation for use in SureSelect X2 Focus on the extraction of tumor-derived DA and preparation of target-enriched libraries for next generation sequencing Application ote For Research
Optimizing PE DA preparation for use in SureSelect X2 Focus on the extraction of tumor-derived DA and preparation of target-enriched libraries for next generation sequencing Application ote For Research
Roche Molecular Biochemicals Technical Note No. LC 10/2000
 Roche Molecular Biochemicals Technical Note No. LC 10/2000 LightCycler Overview of LightCycler Quantification Methods 1. General Introduction Introduction Content Definitions This Technical Note will introduce
Roche Molecular Biochemicals Technical Note No. LC 10/2000 LightCycler Overview of LightCycler Quantification Methods 1. General Introduction Introduction Content Definitions This Technical Note will introduce
QIAGEN s NGS Solutions for Biomarkers NGS & Bioinformatics team QIAGEN (Suzhou) Translational Medicine Co.,Ltd
 QIAGEN s NGS Solutions for Biomarkers NGS & Bioinformatics team QIAGEN (Suzhou) Translational Medicine Co.,Ltd 1 Our current NGS & Bioinformatics Platform 2 Our NGS workflow and applications 3 QIAGEN s
QIAGEN s NGS Solutions for Biomarkers NGS & Bioinformatics team QIAGEN (Suzhou) Translational Medicine Co.,Ltd 1 Our current NGS & Bioinformatics Platform 2 Our NGS workflow and applications 3 QIAGEN s
Chromatrap 96 DNA Micro Elution Purification Kit
 V 1.0 Chromatrap 96 DNA Micro Elution Purification Kit High throughput plates for ultra-pure DNA Protocol v1.0 Catalogue no. 500240 ADVANCEMENTS IN EPIGENETICS Contents Kit Components and Storage 3 Additional
V 1.0 Chromatrap 96 DNA Micro Elution Purification Kit High throughput plates for ultra-pure DNA Protocol v1.0 Catalogue no. 500240 ADVANCEMENTS IN EPIGENETICS Contents Kit Components and Storage 3 Additional
NRAS Codon 61 Mutation Analysis Reagents
 NRAS Codon 61 Mutation Analysis Reagents User Manual V1.0 Cat No. GP19 32 reactions 1 CONTENTS Introduction 4 Overview of Mutector TM Assay 5 Materials Provided 6 Materials Required 7 Equipment Required
NRAS Codon 61 Mutation Analysis Reagents User Manual V1.0 Cat No. GP19 32 reactions 1 CONTENTS Introduction 4 Overview of Mutector TM Assay 5 Materials Provided 6 Materials Required 7 Equipment Required
Use of BHQplus Probes to Optimize Multiplex Competitive qrt-pcr in Diagnostics. Dr. James C. Willey Chief Medical Advisor Accugenomics, Inc.
 Use of BHQplus Probes to Optimize Multiplex Competitive qrt-pcr in Diagnostics Dr. James C. Willey Chief Medical Advisor Accugenomics, Inc. Professor, Medicine and Pathology University of Toledo College
Use of BHQplus Probes to Optimize Multiplex Competitive qrt-pcr in Diagnostics Dr. James C. Willey Chief Medical Advisor Accugenomics, Inc. Professor, Medicine and Pathology University of Toledo College
LightCycler 480 qpcr Tools. Meeting the Challenge of Your Research
 LightCycler 480 qpcr Tools Meeting the Challenge of Your Research Find the Optimal LightCycler 480 Reagents for Your Research Application: Are you analyzing DNA DNA Nucleic acid isolation Manual processing
LightCycler 480 qpcr Tools Meeting the Challenge of Your Research Find the Optimal LightCycler 480 Reagents for Your Research Application: Are you analyzing DNA DNA Nucleic acid isolation Manual processing
NRAS Mutation Analysis Reagents (Codons 12 and 13)
 NRAS Mutation Analysis Reagents (Codons 12 and 13) User Manual V1.1 Cat No. GP18 32 reactions 1 CONTENTS Introduction 4 Overview of Mutector TM Assay 5 Materials Provided 6 Materials Required 7 Equipment
NRAS Mutation Analysis Reagents (Codons 12 and 13) User Manual V1.1 Cat No. GP18 32 reactions 1 CONTENTS Introduction 4 Overview of Mutector TM Assay 5 Materials Provided 6 Materials Required 7 Equipment
Real-Time PCR Workshop Gene Expression. Applications Absolute and Relative Quantitation
 Real-Time PCR Workshop Gene Expression Applications Absolute and Relative Quantitation Absolute Quantitation Easy to understand the data, difficult to develop/qualify the standards Relative Quantitation
Real-Time PCR Workshop Gene Expression Applications Absolute and Relative Quantitation Absolute Quantitation Easy to understand the data, difficult to develop/qualify the standards Relative Quantitation
Technical note: Molecular Index counting adjustment methods
 Technical note: Molecular Index counting adjustment methods By Jue Fan, Jennifer Tsai, Eleen Shum Introduction. Overview of BD Precise assays BD Precise assays are fast, high-throughput, next-generation
Technical note: Molecular Index counting adjustment methods By Jue Fan, Jennifer Tsai, Eleen Shum Introduction. Overview of BD Precise assays BD Precise assays are fast, high-throughput, next-generation
Improved Sensitivity for FFPE Microarray Analysis: ArrayGrade FFPE RNA Isolation
 Improved Sensitivity for FFPE Microarray Analysis: ArrayGrade FFPE RNA Isolation Improved RNA Isolation from FFPE Blocks and Slides for Greater Microarray Sensitivity By Savita Prabhakar, George Quellhorst,
Improved Sensitivity for FFPE Microarray Analysis: ArrayGrade FFPE RNA Isolation Improved RNA Isolation from FFPE Blocks and Slides for Greater Microarray Sensitivity By Savita Prabhakar, George Quellhorst,
Product # Specifications. Kit Specifications Maximum Column Binding Capacity (gdna) Maximum Column Loading Volume.
 3430 Schmon Parkway Thorold, ON, Canada L2V 4Y6 Phone: 866-667-4362 (905) 227-8848 Fax: (905) 227-1061 Email: techsupport@norgenbiotek.com FFPE DNA Purification Kit Product # 47400 Product Insert Norgen
3430 Schmon Parkway Thorold, ON, Canada L2V 4Y6 Phone: 866-667-4362 (905) 227-8848 Fax: (905) 227-1061 Email: techsupport@norgenbiotek.com FFPE DNA Purification Kit Product # 47400 Product Insert Norgen
For Research Use Only Ver
 INSTRUCTION MANUAL ZR-Duet DNA/RNA MiniPrep Plus Catalog No. D7003 Highlights Efficient isolation and separation of DNA and RNA from any cells, tissue, blood, and biological fluids. High quality DNA and
INSTRUCTION MANUAL ZR-Duet DNA/RNA MiniPrep Plus Catalog No. D7003 Highlights Efficient isolation and separation of DNA and RNA from any cells, tissue, blood, and biological fluids. High quality DNA and
DNA Genotyping from Human FFPE Samples Reliable and Reproducible
 DNA Genotyping from Human FFPE Samples Reliable and Reproducible Use this optimized protocol to obtain reliable and reproducible SNP Genotyping data from those difficult human FFPE samples. 1 FFPE DNA Isolation
DNA Genotyping from Human FFPE Samples Reliable and Reproducible Use this optimized protocol to obtain reliable and reproducible SNP Genotyping data from those difficult human FFPE samples. 1 FFPE DNA Isolation
Optimized Covaris truxtrac FFPE RNA Protocol for Isolating RNA from Laser-Capture Microdissected (LCM) FFPE Tissue for Next Generation Sequencing
 Optimized truxtrac FFPE RNA Protocol for Isolating RNA from Laser-Capture Microdissected (LCM) FFPE Tissue for Next Generation Sequencing Enni Markkanen Institute of Veterinary Pharmacology and Toxicology,
Optimized truxtrac FFPE RNA Protocol for Isolating RNA from Laser-Capture Microdissected (LCM) FFPE Tissue for Next Generation Sequencing Enni Markkanen Institute of Veterinary Pharmacology and Toxicology,
electrophoresis tech Effect of RNA Degradation on Data Quality in Quantitative PCR and Microarray Experiments
 electrophoresis tech note 5452 Effect of RNA Degradation on Data Quality in Quantitative PCR and Microarray Experiments Jeff Gingrich, Teresa Rubio, and Cathleen Karlak, Bio-Rad Laboratories, Inc., Hercules,
electrophoresis tech note 5452 Effect of RNA Degradation on Data Quality in Quantitative PCR and Microarray Experiments Jeff Gingrich, Teresa Rubio, and Cathleen Karlak, Bio-Rad Laboratories, Inc., Hercules,
Illumina TruSeq RNA Access Library Prep Kit Automated on the Biomek FX P Dual-Hybrid Liquid Handler
 Illumina TruSeq RNA Access Library Prep Kit Automated on the Biomek FX P Dual-Hybrid Liquid Handler Introduction David Horvath, M.S., Senior Applications Scientist, Beckman Coulter, Inc.; Tim Hill, Scientist,
Illumina TruSeq RNA Access Library Prep Kit Automated on the Biomek FX P Dual-Hybrid Liquid Handler Introduction David Horvath, M.S., Senior Applications Scientist, Beckman Coulter, Inc.; Tim Hill, Scientist,
NEBNext Magnesium RNA Fragmentation Module
 SAMPLE PREPARATION NEBNext Magnesium RNA Fragmentation Module Instruction Manual NEB #E6150S 200 reactions NEBNext Magnesium RNA Fragmentation Module Table of Contents: Description....2 Applications....2
SAMPLE PREPARATION NEBNext Magnesium RNA Fragmentation Module Instruction Manual NEB #E6150S 200 reactions NEBNext Magnesium RNA Fragmentation Module Table of Contents: Description....2 Applications....2
Accessible answers. Targeted sequencing: accelerating and amplifying answers for oncology research
 Accessible answers Targeted sequencing: accelerating and amplifying answers for oncology research Help advance precision medicine Accelerate results with Ion Torrent NGS Life without cancer. This is our
Accessible answers Targeted sequencing: accelerating and amplifying answers for oncology research Help advance precision medicine Accelerate results with Ion Torrent NGS Life without cancer. This is our
Using Low Input of Poly (A) + RNA and Total RNA for Oligonucleotide Microarrays Application
 Using Low Input of Poly (A) + RNA and Total RNA for Oligonucleotide Microarrays Application Gene Expression Author Michelle M. Chen Agilent Technologies, Inc. 3500 Deer Creek Road, MS 25U-7 Palo Alto,
Using Low Input of Poly (A) + RNA and Total RNA for Oligonucleotide Microarrays Application Gene Expression Author Michelle M. Chen Agilent Technologies, Inc. 3500 Deer Creek Road, MS 25U-7 Palo Alto,
Technical Review. Real time PCR
 Technical Review Real time PCR Normal PCR: Analyze with agarose gel Normal PCR vs Real time PCR Real-time PCR, also known as quantitative PCR (qpcr) or kinetic PCR Key feature: Used to amplify and simultaneously
Technical Review Real time PCR Normal PCR: Analyze with agarose gel Normal PCR vs Real time PCR Real-time PCR, also known as quantitative PCR (qpcr) or kinetic PCR Key feature: Used to amplify and simultaneously
Quality of Cancer RNA Samples Is Essential for Molecular Classification Based on Microarray Results Application
 Quality of Cancer RNA Samples Is Essential for Molecular Classification Based on Microarray Results Application Pathology Author Jenny Xiao Agilent Technologies, Inc. Deer Creek Road MC U-7 Palo Alto,
Quality of Cancer RNA Samples Is Essential for Molecular Classification Based on Microarray Results Application Pathology Author Jenny Xiao Agilent Technologies, Inc. Deer Creek Road MC U-7 Palo Alto,
MANUAL SOLUTIONS BIOMARKER ANALYSIS BY RNA IN SITU HYBRIDIZATION
 RNASCOPE MANUAL SOLUTIONS BIOMARKER ANALYSIS BY RNA IN SITU HYBRIDIZATION Get Quantitative Molecular Detection with Morphological Context in a Single Assay Advanced Cell Diagnostics RNAscope is a proprietary
RNASCOPE MANUAL SOLUTIONS BIOMARKER ANALYSIS BY RNA IN SITU HYBRIDIZATION Get Quantitative Molecular Detection with Morphological Context in a Single Assay Advanced Cell Diagnostics RNAscope is a proprietary
AmpliScribe T 7 Aminoallyl-RNA Transcription Kit
 Cat. No. AA50125 The AmpliScribe T7 Aminoallyl-RNA Transcription Kit enables high-yield production of aminoallyl-labeled RNA. The kit utilizes Epicentre s high yielding AmpliScribe T7-Flash in vitro transcription
Cat. No. AA50125 The AmpliScribe T7 Aminoallyl-RNA Transcription Kit enables high-yield production of aminoallyl-labeled RNA. The kit utilizes Epicentre s high yielding AmpliScribe T7-Flash in vitro transcription
Introduction To Real-Time Quantitative PCR (qpcr)
 Introduction To Real-Time Quantitative PCR (qpcr) Samuel Rulli, Ph.D. Samuel.Rulli@QIAGEN.com Technical Support: BRCsupport@qiagen.com The products described in this webinar are intended for molecular
Introduction To Real-Time Quantitative PCR (qpcr) Samuel Rulli, Ph.D. Samuel.Rulli@QIAGEN.com Technical Support: BRCsupport@qiagen.com The products described in this webinar are intended for molecular
QuickExtract FFPE RNA Extraction Kit
 QuickExtract FFPE RNA Extraction Kit Cat. Nos. QFR82805 and QFR82050 Connect with Epicentre on our blog (epicentral.blogspot.com), Facebook (facebook.com/epicentrebio), and Twitter (@EpicentreBio). www.epicentre.com
QuickExtract FFPE RNA Extraction Kit Cat. Nos. QFR82805 and QFR82050 Connect with Epicentre on our blog (epicentral.blogspot.com), Facebook (facebook.com/epicentrebio), and Twitter (@EpicentreBio). www.epicentre.com
Oncomine cfdna Assays Part III: Variant Analysis
 Oncomine cfdna Assays Part III: Variant Analysis USER GUIDE for use with: Oncomine Lung cfdna Assay Oncomine Colon cfdna Assay Oncomine Breast cfdna Assay Catalog Numbers A31149, A31182, A31183 Publication
Oncomine cfdna Assays Part III: Variant Analysis USER GUIDE for use with: Oncomine Lung cfdna Assay Oncomine Colon cfdna Assay Oncomine Breast cfdna Assay Catalog Numbers A31149, A31182, A31183 Publication
Quantitative Real Time PCR USING SYBR GREEN
 Quantitative Real Time PCR USING SYBR GREEN SYBR Green SYBR Green is a cyanine dye that binds to double stranded DNA. When it is bound to D.S. DNA it has a much greater fluorescence than when bound to
Quantitative Real Time PCR USING SYBR GREEN SYBR Green SYBR Green is a cyanine dye that binds to double stranded DNA. When it is bound to D.S. DNA it has a much greater fluorescence than when bound to
MICROARRAYS+SEQUENCING
 MICROARRAYS+SEQUENCING The most efficient way to advance genomics research Down to a Science. www.affymetrix.com/downtoascience Affymetrix GeneChip Expression Technology Complementing your Next-Generation
MICROARRAYS+SEQUENCING The most efficient way to advance genomics research Down to a Science. www.affymetrix.com/downtoascience Affymetrix GeneChip Expression Technology Complementing your Next-Generation
REAL TIME PCR USING SYBR GREEN
 REAL TIME PCR USING SYBR GREEN 1 THE PROBLEM NEED TO QUANTITATE DIFFERENCES IN mrna EXPRESSION SMALL AMOUNTS OF mrna LASER CAPTURE SMALL AMOUNTS OF TISSUE PRIMARY CELLS PRECIOUS REAGENTS 2 THE PROBLEM
REAL TIME PCR USING SYBR GREEN 1 THE PROBLEM NEED TO QUANTITATE DIFFERENCES IN mrna EXPRESSION SMALL AMOUNTS OF mrna LASER CAPTURE SMALL AMOUNTS OF TISSUE PRIMARY CELLS PRECIOUS REAGENTS 2 THE PROBLEM
NAG-1 (GDF-15, MIC-1) Prostate Cancer ELISA kit
 NAG-1 (GDF-15, MIC-1) Prostate Cancer ELISA kit Catalog Number: NG1 Store at -0 C. FOR RESEARCH USE ONLY v. 1081 Introduction This sandwich ELISA kit is for determination of NAG-1 (GDF-15, MIC-1) levels
NAG-1 (GDF-15, MIC-1) Prostate Cancer ELISA kit Catalog Number: NG1 Store at -0 C. FOR RESEARCH USE ONLY v. 1081 Introduction This sandwich ELISA kit is for determination of NAG-1 (GDF-15, MIC-1) levels
PlexSet Reagents User Manual
 PlexSet Reagents User Manual For Research Use Only. Not for use in diagnostic procedures. MAN-10040-03 October 2017 ncounter PlexSet Reagents for Gene Expression User Manual MAN-10040-03 PlexSet Reagents
PlexSet Reagents User Manual For Research Use Only. Not for use in diagnostic procedures. MAN-10040-03 October 2017 ncounter PlexSet Reagents for Gene Expression User Manual MAN-10040-03 PlexSet Reagents
TransIT -mrna Transfection Kit
 INTRODUCTION TransIT -mrna Transfection Kit is designed to transfect RNA into a broad range of cell types with minimal cellular toxicity. RNA delivery avoids transcriptional regulation effects by directly
INTRODUCTION TransIT -mrna Transfection Kit is designed to transfect RNA into a broad range of cell types with minimal cellular toxicity. RNA delivery avoids transcriptional regulation effects by directly
Cancer Genetics Solutions
 Cancer Genetics Solutions Cancer Genetics Solutions Pushing the Boundaries in Cancer Genetics Cancer is a formidable foe that presents significant challenges. The complexity of this disease can be daunting
Cancer Genetics Solutions Cancer Genetics Solutions Pushing the Boundaries in Cancer Genetics Cancer is a formidable foe that presents significant challenges. The complexity of this disease can be daunting
Nucleopore FFPE DNA Miniprep Kit
 H A N D B O O K GX-4076 50 Prep Genetix Biotech Asia Pvt. Ltd. 71/1, First Floor, Shivaji Marg, Najafgarh Road, New Delhi - 110015 Phone : +91-11-45027000 Fax : +91-11-25419631 E-mail : info@genetixbiotech.com
H A N D B O O K GX-4076 50 Prep Genetix Biotech Asia Pvt. Ltd. 71/1, First Floor, Shivaji Marg, Najafgarh Road, New Delhi - 110015 Phone : +91-11-45027000 Fax : +91-11-25419631 E-mail : info@genetixbiotech.com
Lecture #1. Introduction to microarray technology
 Lecture #1 Introduction to microarray technology Outline General purpose Microarray assay concept Basic microarray experimental process cdna/two channel arrays Oligonucleotide arrays Exon arrays Comparing
Lecture #1 Introduction to microarray technology Outline General purpose Microarray assay concept Basic microarray experimental process cdna/two channel arrays Oligonucleotide arrays Exon arrays Comparing
Fluorescent in-situ Hybridization
 Fluorescent in-situ Hybridization Presented for: Presented by: Date: 2 Definition In situ hybridization is the method of localizing/ detecting specific nucleotide sequences in morphologically preserved
Fluorescent in-situ Hybridization Presented for: Presented by: Date: 2 Definition In situ hybridization is the method of localizing/ detecting specific nucleotide sequences in morphologically preserved
with Cell Surface-Compatible Universal Cell Capture Kit
 MAN-10066-01 with Cell Surface-Compatible Universal Cell Capture Kit In this workflow, cells are collected and then bound to the Universal Cell Capture Beads; then the RNA and Protein samples are prepared
MAN-10066-01 with Cell Surface-Compatible Universal Cell Capture Kit In this workflow, cells are collected and then bound to the Universal Cell Capture Beads; then the RNA and Protein samples are prepared
Recent technology allow production of microarrays composed of 70-mers (essentially a hybrid of the two techniques)
 Microarrays and Transcript Profiling Gene expression patterns are traditionally studied using Northern blots (DNA-RNA hybridization assays). This approach involves separation of total or polya + RNA on
Microarrays and Transcript Profiling Gene expression patterns are traditionally studied using Northern blots (DNA-RNA hybridization assays). This approach involves separation of total or polya + RNA on
For Research Use Only Ver
 INSTRUCTION MANUAL Quick-RNA Miniprep Plus Kit Catalog Nos. R1057T, R1057, & R1058 Highlights Purify high-quality total RNA (including small/micro RNAs) from any cells, tissues, blood and biological fluids.
INSTRUCTION MANUAL Quick-RNA Miniprep Plus Kit Catalog Nos. R1057T, R1057, & R1058 Highlights Purify high-quality total RNA (including small/micro RNAs) from any cells, tissues, blood and biological fluids.
Gene Expression Profiling and Validation Using Agilent SurePrint G3 Gene Expression Arrays
 Gene Expression Profiling and Validation Using Agilent SurePrint G3 Gene Expression Arrays Application Note Authors Bahram Arezi, Nilanjan Guha and Anne Bergstrom Lucas Agilent Technologies Inc. Santa
Gene Expression Profiling and Validation Using Agilent SurePrint G3 Gene Expression Arrays Application Note Authors Bahram Arezi, Nilanjan Guha and Anne Bergstrom Lucas Agilent Technologies Inc. Santa
Introduction to Bioinformatics and Gene Expression Technologies
 Introduction to Bioinformatics and Gene Expression Technologies Utah State University Fall 2017 Statistical Bioinformatics (Biomedical Big Data) Notes 1 1 Vocabulary Gene: hereditary DNA sequence at a
Introduction to Bioinformatics and Gene Expression Technologies Utah State University Fall 2017 Statistical Bioinformatics (Biomedical Big Data) Notes 1 1 Vocabulary Gene: hereditary DNA sequence at a
EECS730: Introduction to Bioinformatics
 EECS730: Introduction to Bioinformatics Lecture 14: Microarray Some slides were adapted from Dr. Luke Huan (University of Kansas), Dr. Shaojie Zhang (University of Central Florida), and Dr. Dong Xu and
EECS730: Introduction to Bioinformatics Lecture 14: Microarray Some slides were adapted from Dr. Luke Huan (University of Kansas), Dr. Shaojie Zhang (University of Central Florida), and Dr. Dong Xu and
truxtrac FFPE DNA microtube Kit Column Purification (25)
 PROTOCOL truxtrac FFPE DNA microtube Kit Column Purification (25) Adaptive Focused Acoustics (AFA) -based DNA extraction and purification from Formalin-Fixed, Paraffin-Embedded Tissue using columns Product
PROTOCOL truxtrac FFPE DNA microtube Kit Column Purification (25) Adaptive Focused Acoustics (AFA) -based DNA extraction and purification from Formalin-Fixed, Paraffin-Embedded Tissue using columns Product
Methods of Biomaterials Testing Lesson 3-5. Biochemical Methods - Molecular Biology -
 Methods of Biomaterials Testing Lesson 3-5 Biochemical Methods - Molecular Biology - Chromosomes in the Cell Nucleus DNA in the Chromosome Deoxyribonucleic Acid (DNA) DNA has double-helix structure The
Methods of Biomaterials Testing Lesson 3-5 Biochemical Methods - Molecular Biology - Chromosomes in the Cell Nucleus DNA in the Chromosome Deoxyribonucleic Acid (DNA) DNA has double-helix structure The
Outline. Array platform considerations: Comparison between the technologies available in microarrays
 Microarray overview Outline Array platform considerations: Comparison between the technologies available in microarrays Differences in array fabrication Differences in array organization Applications of
Microarray overview Outline Array platform considerations: Comparison between the technologies available in microarrays Differences in array fabrication Differences in array organization Applications of
Non-Organic-Based Isolation of Mammalian microrna using Norgen s microrna Purification Kit
 Application Note 13 RNA Sample Preparation Non-Organic-Based Isolation of Mammalian microrna using Norgen s microrna Purification Kit B. Lam, PhD 1, P. Roberts, MSc 1 Y. Haj-Ahmad, M.Sc., Ph.D 1,2 1 Norgen
Application Note 13 RNA Sample Preparation Non-Organic-Based Isolation of Mammalian microrna using Norgen s microrna Purification Kit B. Lam, PhD 1, P. Roberts, MSc 1 Y. Haj-Ahmad, M.Sc., Ph.D 1,2 1 Norgen
IQFISH on Dako Omnis. Panel for Lung Cancer. Dako FAST RESULTS. ALK, ROS1, RET and MET IQFISH. Dako Omnis. Agilent Pathology Solutions
 PRODUCT INFORMATION Dako Omnis ALK, ROS1, RET and MET IQFISH Dako Agilent Pathology Solutions IQFISH on Dako Omnis Panel for Lung Cancer FAST RESULTS Fast, high-quality FISH Integrated into your IHC workflow
PRODUCT INFORMATION Dako Omnis ALK, ROS1, RET and MET IQFISH Dako Agilent Pathology Solutions IQFISH on Dako Omnis Panel for Lung Cancer FAST RESULTS Fast, high-quality FISH Integrated into your IHC workflow
qpcr Quantitative PCR or Real-time PCR Gives a measurement of PCR product at end of each cycle real time
 qpcr qpcr Quantitative PCR or Real-time PCR Gives a measurement of PCR product at end of each cycle real time Differs from endpoint PCR gel on last cycle Used to determines relative amount of template
qpcr qpcr Quantitative PCR or Real-time PCR Gives a measurement of PCR product at end of each cycle real time Differs from endpoint PCR gel on last cycle Used to determines relative amount of template
T7-Based RNA Amplification Protocol (in progress)
 T7-Based RNA Amplification Protocol (in progress) Jacqueline Ann Lopez (modifications) Amy Cash & Justen Andrews INTRODUCTION T7 RNA Amplification, a technique originally developed in the laboratory of
T7-Based RNA Amplification Protocol (in progress) Jacqueline Ann Lopez (modifications) Amy Cash & Justen Andrews INTRODUCTION T7 RNA Amplification, a technique originally developed in the laboratory of
in-situ PCR Presented for: Presented by: Date:
 in-situ PCR Presented for: Presented by: Date: 2 in situ Hybridization - Definition in situ PCR is a method in which the polymerase chain reaction actually takes place in the cell on a slide, and the product
in-situ PCR Presented for: Presented by: Date: 2 in situ Hybridization - Definition in situ PCR is a method in which the polymerase chain reaction actually takes place in the cell on a slide, and the product
Welcome to the NGS webinar series
 Welcome to the NGS webinar series Webinar 1 NGS: Introduction to technology, and applications NGS Technology Webinar 2 Targeted NGS for Cancer Research NGS in cancer Webinar 3 NGS: Data analysis for genetic
Welcome to the NGS webinar series Webinar 1 NGS: Introduction to technology, and applications NGS Technology Webinar 2 Targeted NGS for Cancer Research NGS in cancer Webinar 3 NGS: Data analysis for genetic
Please purchase PDFcamp Printer on to remove this watermark. DNA microarray
 DNA microarray Example of an approximately 40,000 probe spotted oligo microarray with enlarged inset to show detail. A DNA microarray is a multiplex technology used in molecular biology. It consists of
DNA microarray Example of an approximately 40,000 probe spotted oligo microarray with enlarged inset to show detail. A DNA microarray is a multiplex technology used in molecular biology. It consists of
ID3EAL Spike-in RNA Kit. Principle, Workflow and Protocol
 ID3EAL Spike-in RNA Kit Principle, Workflow and Protocol Table of Contents Content Page Working Principle 3 Kit Contents & Storage 4 Additional Equipment and Compatibility 5 Additional Equipment and Compatibility
ID3EAL Spike-in RNA Kit Principle, Workflow and Protocol Table of Contents Content Page Working Principle 3 Kit Contents & Storage 4 Additional Equipment and Compatibility 5 Additional Equipment and Compatibility
Data Sheet. GeneChip Human Genome U133 Arrays
 GeneChip Human Genome Arrays AFFYMETRIX PRODUCT FAMILY > ARRAYS > Data Sheet GeneChip Human Genome U133 Arrays The Most Comprehensive Coverage of the Human Genome in Two Flexible Formats: Single-array
GeneChip Human Genome Arrays AFFYMETRIX PRODUCT FAMILY > ARRAYS > Data Sheet GeneChip Human Genome U133 Arrays The Most Comprehensive Coverage of the Human Genome in Two Flexible Formats: Single-array
CATS RNA-seq v2 library preparation protocol on RNA from FFPE-samples
 CATS RNA-seq v2 library preparation protocol on RNA from FFPE-samples INTRODUCTION The following protocol offers a streamlined solution for whole transcriptome sequencing studies from human/ mouse/rat
CATS RNA-seq v2 library preparation protocol on RNA from FFPE-samples INTRODUCTION The following protocol offers a streamlined solution for whole transcriptome sequencing studies from human/ mouse/rat
High-Resolution Oligonucleotide- Based acgh Analysis of Single Cells in Under 24 Hours
 High-Resolution Oligonucleotide- Based acgh Analysis of Single Cells in Under 24 Hours Application Note Authors Paula Costa and Anniek De Witte Agilent Technologies, Inc. Santa Clara, CA USA Abstract As
High-Resolution Oligonucleotide- Based acgh Analysis of Single Cells in Under 24 Hours Application Note Authors Paula Costa and Anniek De Witte Agilent Technologies, Inc. Santa Clara, CA USA Abstract As
HIGH SCHOOL STUDENT SCIENCE WEEK. St. Paul s Hospital Vancouver, BC
 HIGH SCHOOL STUDENT SCIENCE WEEK St. Paul s Hospital Vancouver, BC Sponsors 2 AGENDA Location: UBC James Hogg Research Centre (JHRC), St. Paul s Hospital, Room 166 Burrard Building, 1081 Burrard Street,
HIGH SCHOOL STUDENT SCIENCE WEEK St. Paul s Hospital Vancouver, BC Sponsors 2 AGENDA Location: UBC James Hogg Research Centre (JHRC), St. Paul s Hospital, Room 166 Burrard Building, 1081 Burrard Street,
GeneChip Eukaryotic Small Sample Target Labeling Assay Version II *
 GENE EXPRESSION MONITORING TECHNICAL NOTE GeneChip Eukaryotic Small Sample Target Labeling Assay Version II * Introduction There is an overwhelming and continuing demand for a well characterized protocol
GENE EXPRESSION MONITORING TECHNICAL NOTE GeneChip Eukaryotic Small Sample Target Labeling Assay Version II * Introduction There is an overwhelming and continuing demand for a well characterized protocol
Automated size selection of NEBNext Small RNA libraries with the Sage Pippin Prep
 Automated size selection of NEBNext Small RNA libraries with the Sage Pippin Prep DNA CLONING DNA AMPLIFICATION & PCR EPIGENETICS RNA ANALYSIS LIBRARY PREP FOR NEXT GEN SEQUENCING PROTEIN EXPRESSION &
Automated size selection of NEBNext Small RNA libraries with the Sage Pippin Prep DNA CLONING DNA AMPLIFICATION & PCR EPIGENETICS RNA ANALYSIS LIBRARY PREP FOR NEXT GEN SEQUENCING PROTEIN EXPRESSION &
SNP GENOTYPING WITH iplex REAGENTS AND THE MASSARRAY SYSTEM
 SNP GENOTYPING Accurate, sensitive, flexible MassARRAY System SNP GENOTYPING WITH iplex REAGENTS AND THE MASSARRAY SYSTEM Biomarker validation Routine genetic testing Somatic mutation profiling Up to 400
SNP GENOTYPING Accurate, sensitive, flexible MassARRAY System SNP GENOTYPING WITH iplex REAGENTS AND THE MASSARRAY SYSTEM Biomarker validation Routine genetic testing Somatic mutation profiling Up to 400
36 μl 90 μl 360 μl 20 C
 Package Insert QuantiGene Sample Processing Kit FFPE Tissues About Sample Processing Kits Sample Processing Kits are designed for use with both single plex and multiplex QuantiGene assays for quantification
Package Insert QuantiGene Sample Processing Kit FFPE Tissues About Sample Processing Kits Sample Processing Kits are designed for use with both single plex and multiplex QuantiGene assays for quantification
1 INTENDED USE / INDICATIONS FOR USE TABLE OF CONTENTS 2 SUMMARY OF THE TEST SYSTEM
 www.nanostring.com Package Insert Prosigna Breast Cancer Prognostic Gene Signature Assay 14 LIMITATIONS OF THE PROCEDURES...10 15 EXPECTED VALUES...10 15.1 Distant Recurrence-Free Survival by Risk Categorization...
www.nanostring.com Package Insert Prosigna Breast Cancer Prognostic Gene Signature Assay 14 LIMITATIONS OF THE PROCEDURES...10 15 EXPECTED VALUES...10 15.1 Distant Recurrence-Free Survival by Risk Categorization...
American Society of Cytopathology Core Curriculum in Molecular Biology
 Brought American Society of Cytopathology Core Curriculum in Molecular Biology Brought American Society of Cytopathology Core Curriculum in Molecular Biology Chapter 3 Molecular Techniques Alternatives
Brought American Society of Cytopathology Core Curriculum in Molecular Biology Brought American Society of Cytopathology Core Curriculum in Molecular Biology Chapter 3 Molecular Techniques Alternatives
Determinants of RNA Quality from FFPE Samples
 Determinants of RNA Quality from FFPE Samples Silke von Ahlfen 1, Andreas Missel 1, Klaus Bendrat 2, Martin Schlumpberger 1 * 1 QIAGEN GmbH, Hilden, Germany, 2 Gemeinschaftspraxis für Pathologie Niendorf
Determinants of RNA Quality from FFPE Samples Silke von Ahlfen 1, Andreas Missel 1, Klaus Bendrat 2, Martin Schlumpberger 1 * 1 QIAGEN GmbH, Hilden, Germany, 2 Gemeinschaftspraxis für Pathologie Niendorf
E.Z.N.A. Tissue RNA Kit. R preps R preps
 E.Z.N.A. Tissue RNA Kit R6688-00 5 preps R6688-01 50 preps May 2015 E.Z.N.A. Tissue RNA Kit Table of Contents Introduction and Overview...2 Kit Contents/Storage and Stability...3 Important Notes...4 Homogenization
E.Z.N.A. Tissue RNA Kit R6688-00 5 preps R6688-01 50 preps May 2015 E.Z.N.A. Tissue RNA Kit Table of Contents Introduction and Overview...2 Kit Contents/Storage and Stability...3 Important Notes...4 Homogenization
Real-Time PCR Validations
 Real-Time PCR Validations Relative gene expression Example experiment: I have 2 samples: untreated and treated. Question: What happens to the expression of gene X when I treat the cells? Answer: Expressed
Real-Time PCR Validations Relative gene expression Example experiment: I have 2 samples: untreated and treated. Question: What happens to the expression of gene X when I treat the cells? Answer: Expressed
FOR RESEARCH USE ONLY. NOT FOR HUMAN OR DIAGNOSTIC USE.
 Instruction manual RNA-direct SYBR Green Realtime PCR Master Mix 0810 F0930K RNA-direct SYBR Green Realtime PCR Master Mix Contents QRT-201T QRT-201 0.5mLx2 0.5mLx5 Store at -20 C, protected from light
Instruction manual RNA-direct SYBR Green Realtime PCR Master Mix 0810 F0930K RNA-direct SYBR Green Realtime PCR Master Mix Contents QRT-201T QRT-201 0.5mLx2 0.5mLx5 Store at -20 C, protected from light
IDEAL TM Spike-In RNA Kit. Individual Assays Principle, Workflow and Protocol
 IDEAL TM Spike-In RNA Kit Individual Assays Principle, Workflow and Protocol Table of Contents Content Page Working Principle 3 Kit Contents & Storage 4 Additional Equipment and Compatibility 5 Workflow
IDEAL TM Spike-In RNA Kit Individual Assays Principle, Workflow and Protocol Table of Contents Content Page Working Principle 3 Kit Contents & Storage 4 Additional Equipment and Compatibility 5 Workflow
truxtrac FFPE DNA microtube Kit - Bead Purification (24)
 truxtrac FFPE DNA microtube Kit - Bead Purification (24) Adaptive Focused Acoustics (AFA) -based DNA extraction and purification from Formalin-Fixed, Paraffin-Embedded (FFPE) Tissue using Magnetic Beads
truxtrac FFPE DNA microtube Kit - Bead Purification (24) Adaptive Focused Acoustics (AFA) -based DNA extraction and purification from Formalin-Fixed, Paraffin-Embedded (FFPE) Tissue using Magnetic Beads
Sample Preparation Demonstrated Protocol
 10x Genomics Sample Preparation Demonstrated Protocol DNA Extraction from Fresh Frozen Tissue NOTICES Notices Manual Part Number CG000072 Rev B Legal Notices 2017 10x Genomics, Inc. All rights reserved.
10x Genomics Sample Preparation Demonstrated Protocol DNA Extraction from Fresh Frozen Tissue NOTICES Notices Manual Part Number CG000072 Rev B Legal Notices 2017 10x Genomics, Inc. All rights reserved.
E.Z.N.A. Blood DNA Mini Kit. D preps D preps D preps
 E.Z.N.A. Blood DNA Mini Kit D3392-00 5 preps D3392-01 50 preps D3392-02 200 preps January 2017 E.Z.N.A. Blood DNA Mini Kit Table of Contents Introduction and Overview...2 Illustrated Protocol...3 Kit Contents/Storage
E.Z.N.A. Blood DNA Mini Kit D3392-00 5 preps D3392-01 50 preps D3392-02 200 preps January 2017 E.Z.N.A. Blood DNA Mini Kit Table of Contents Introduction and Overview...2 Illustrated Protocol...3 Kit Contents/Storage
Nature Methods: doi: /nmeth Supplementary Figure 1. Retention of RNA with LabelX.
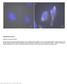 Supplementary Figure 1 Retention of RNA with LabelX. (a) Epi-fluorescence image of single molecule FISH (smfish) against GAPDH on HeLa cells expanded without LabelX treatment. (b) Epi-fluorescence image
Supplementary Figure 1 Retention of RNA with LabelX. (a) Epi-fluorescence image of single molecule FISH (smfish) against GAPDH on HeLa cells expanded without LabelX treatment. (b) Epi-fluorescence image
Performance characteristics of the High Sensitivity DNA kit for the Agilent 2100 Bioanalyzer
 Performance characteristics of the High Sensitivity DNA kit for the Agilent 2100 Bioanalyzer Technical Note 10 Measured conc. [ng/µl] 1 Y intercept = 0.09 r 2 = 0.993 0.1 0.1 1 10 Reference concentration
Performance characteristics of the High Sensitivity DNA kit for the Agilent 2100 Bioanalyzer Technical Note 10 Measured conc. [ng/µl] 1 Y intercept = 0.09 r 2 = 0.993 0.1 0.1 1 10 Reference concentration
