42 fl organelles = 34.5 fl (1) 3.5X X 0.93 = 78,000 (2)
|
|
|
- Rhoda Gilmore
- 6 years ago
- Views:
Transcription
1 SUPPLEMENTAL DATA Supplementary Experimental Procedures Fluorescence Microscopy - A Zeiss Axiovert 200M microscope equipped with a Zeiss 100x Plan- Apochromat (1.40 NA) DIC objective and Hamamatsu Orca 2 CCD camera was used to capture fluorescence images of yeast cells expressing Cdc34-GFP (Invitrogen) (1). Data was collected and analyzed with Zeiss Axiovision 4.6. Images were cropped and adjustments to brightness were made within Axiovision 4.6. Zeiss Axiovision 4.6 with quantification module was used to measure mean pixel intensity. Circles of interest were drawn within the nucleus or cytoplasm. Background light was measured by selecting a large circle of interest outside of each cell. Mean pixel intensities (arbitrary units) were calculated by subtracting background from nuclear and cytoplasmic values. Estimating the nuclear concentration of yeast Cdc34 - To estimate the nuclear concentration of yeast Cdc34 in haploid yeast cells, the number of Cdc34 molecules per cell was estimated by serially diluting known quantities of purified, bacterially-expressed Cdc into an extract of CDC34-myc cells in which the genomic locus that encoded Cdc34 was modified by addition of sequences that encoded a myc epitope tag (RJD982). The tagged Cdc34 migrated slower than untagged bacterially-derived Cdc34-270, which allowed us to distinguish the two species by SDS-PAGE. By immunoblotting with anti-cdc34, scanning the film, and quantifying the resulting file with NIH Image, we estimated the endogenous protein to be present at 78,000 molecules/cell. The ratio of nuclear and cytoplasmic Cdc34 was estimated by fluorescence microscopy of cells in which the genomic copy of CDC34 was tagged with sequences that encode GFP. The average value for this ratio from 20 cells was 3.5 ± 0.6. The average total volume of a haploid yeast cell was estimated at 42 fl (2,3). The total volume of the following organelles was also estimated: endoplasmic reticulum 0.7 fl, mitochondria 2.3 fl, vacuole 1.9 fl, and nucleus 2.6 fl (3). Given these values, the volume of cytoplasm was estimated from equation 1: 42 fl organelles = 34.5 fl (1) Given these values, equation 2 derives the number of molecules in the nucleus and in the cytoplasm: 3.5X X 0.93 = 78,000 (2) 1
2 where 0.07 is the fractional volume of the nucleus and 0.93 is the fractional volume of the cytoplasm. Therefore X = 66,383, and the number of molecules of Cdc34 in the nucleus is 16,264. This was then divided by the respective volumes of the cytoplasm and nucleus and Avogadro s number to arrive at an estimate of ~ 10μM for the nuclear and 2.9 M for the cytoplasmic concentration of yeast Cdc34. GST pull-down binding assay 2 nmole of either GST alone or GST-HACT were incubated in the presence of 30 ul of glutathione-sepharose 4B resin at 4 C for 1 hour. The beads were briefly incubated twice with wash buffer containing 30 mm Tris, ph 7.5, 100 mm NaCl, 0.1 % IgePal, 1 mm DTT and 5 % glycerol. Beads were then incubated with either Cul1 Rbx1 (80, 40, 20, 10, or 5 pmole) or a buffer only control at 4 C for 1 hour. The beads were washed twice with 1 ml of wash buffer and followed by the addition of 100 ul of 1X SDS-PAGE buffer. These reactions were boiled for 5 minutes and then used for analysis by SDS-PAGE. The amount of Cul1 Rbx1 bound to the beads was determined by Western blot analysis with a monoclonal Cul1 antibody (Zymogen) and by comparison with known amounts of Cul1 Rbx1. Quantitative Western analysis was performed using an Alexa Fluor 680 goat anti-mouse IgG antibody (Molecular Probes) and the Odyssey infrared imaging system (Li-Cor Biosciences). Neddylation of Cul1+Rbx1 and Cul1 Cdc34-WT and Cul1 Cdc fusions nm Cul1+Rbx1 or Cul1 fusion proteins were incubated in the presence of 125 nm Nedd8 E1 (4), 0.5 μm Ubc12, 1 μm Nedd8 and 2 mm ATP at C for 20 minutes. Neddylation was confirmed by Western blot analysis using a monoclonal Cul1 antibody (data not shown). 2
3 REFERENCES 1. Huh, W. K., Falvo, J. V., Gerke, L. C., Carroll, A. S., Howson, R. W., Weissman, J. S., and O'Shea, E. K. (2003) Nature 425, Jorgensen, P., Nishikawa, J. L., Breitkreutz, B. J., and Tyers, M. (2002) Science 297, Perktold, A., Zechmann, B., Daum, G., and Zellnig, G. (2007) FEMS Yeast Res 7, Huang, D. T., Schulman, B. A., and Raymond, J. D. (2005) Expression, Purification, and Characterization of the E1 for Human NEDD8, the Heterodimeric APPBP1-UBA3 Complex. in Methods in Enzymology, Academic Press. pp
4 Supplementary Figure Legends Fig S1. Multiple sequence alignment of yeast Cdc34, human Cdc34a and Cdc34b, yeast Ubc8 and yeast Rad6 amino acid sequences. Residues encompassing the ubiquitin conjugating catalytic domain are shaded in pink. The human acidic tail residues and the yeast ones that are aligned to them are shaded purple. The residues in the yeast tail that extend beyond the human Cdc34 tail sequences are shaded green. Fig S2. Yeast Cdc and Cdc have comparable activities when assayed with SCF and Sic1. Reactions containing 1.6 μm yeast E1, 100 nm yeast SCF, 1.2 μm 32 P-labeled Sic1, 100 nm yeast Cdc or Cdc34-270, and 150 μm ubiquitin were incubated for 2, 4, 6, 8 and 10 minutes prior to quenching with reducing SDS-PAGE buffer. Note that the Cdc34 concentration was within the k cat /K m range for both constructs which would maximize any potential differences in activity between the two Cdc34 proteins. The percentage of Sic1 substrate that was converted to product is given at the top of the gel for each time point. The differences observed between Cdc and Cdc are well within experimental error. Fig S3. (A) Anti-Cdc34 Western blot of normalized extracts (12.5 g/lane) from yeast strains sustained by either Cdc34-mycHIS6 protein (RJD982; lanes 1, 3-6) or un-tagged Cdc34 protein (RJD379; lane 2). Lanes 3-6 contain RJD982 extract mixed with either 1 ng (lane 3), 2 ng (lane 4), 4 ng (lane 5), or 8 ng (lane 6) of recombinant Cdc protein. Lanes 7 and 8 contain only recombinant Cdc protein (2 and 4 ng, respectively). (B) The Intensity of Cdc34- GFP fluorescence inside the nucleus (red circle nearest the center of the cell) and inside the cytoplasm (red circle adjacent to the cell wall) was determined. Left: representative DIC images for wild type cells expressing Cdc34-GFP from the endogenous locus. Middle: Cdc34-GFP localizes to the nucleus. Right: Red circles define areas for which mean pixel intensity was calculated. White bars: 2 m. Fig S4. Graph plotting the rate of Sic1 ubiquitylation against the concentration of Cdc34 acidic tail truncation proteins. The data were fit to the Michaelis-Menten equation using non-linear curve fitting (Prism). (A) Cdc (B) Cdc (C) Cdc (D) Cdc (E) Cdc Each graphical data point represents the mean of duplicate data values from 2 independent experiments and the error bars are the standard deviation. 4
5 Fig S5. The human acidic tail domain is sufficient for binding to human SCF. (A) 2 nmole of either GST or a GST fusion of the human acidic C-terminal tail domain (GST-HACT) were immobilized on glutathione beads. Beads were then incubated in the presence of either 80 (lanes 2 and 8), 40 (lanes 3 and 9), 20 (lanes 4 and 10), 10 (lanes 5 and 11) or 5 (lanes 6 and 12) pmole of human Cul1 Rbx1 or in the presence of buffer as a negative control (lanes 1 and 7). Samples were then prepared as noted in the Experimental Procedures section above. To demonstrate even loading of GST and GST-HACT, 10 ul of each sample was loaded onto a 16 % SDS-PAGE gel and stained with coomassie blue Safe Stain (Invitrogen). Because Cul1 is expressed as two fragments, the NTD and CTD, these two fragments co migrate by SDS-PAGE and only one band is visible at the apparent MW of ~ 40 kda. (B) Western blot of a 2-fold dilution series of Cul1 Rbx1. Intensities of the individual bands were quantified using Li-Cor Biosciences software and used to create a standard curve. The standard curve was used to quantify the amount of protein bound to the glutathione-sepharose beads and to generate the plot in Figure S4C. (C) Quantification of the GST pull-down experiment demonstrated specific binding between GST- HACT and Cul1 Rbx1 in a dose dependent manner. The statistical significance of the greater binding of Cul1 Rbx1 to GST-HACT than to GST alone was demonstrated using the unpaired Student s t test with Welch s correction (P-value < 0.01). Fig S6. Neddylation of both Cul1 Cdc34-WT and Cul1 Cdc fusion proteins has a stimulatory effect similar to the un-fused proteins. Di-ubiquitin synthesis assays comparing (A) wild-type and (B) Cdc in the presence of either Cul1+Rbx1 or Cul1 Nedd8 +Rbx1 or comparing (A) Cul1 Cdc34-WT and Cul1 Nedd8 Cdc34-WT or (B) Cul1 Cdc and Cul1 Nedd8 Cdc Reactions containing either 300 nm Cul1+Rbx1 and 300 nm Cdc34 or containing 300 nm Cul1 Cdc34 fusion, 0.7 μm human E1, 6 μm 32 P-labeled K48R ubiquitin, and 50 μm D77 ubiquitin were incubated at C for the specified times and quenched with SDS-PAGE buffer. Note that neddylated Cul1 Cdc34-WT migrated at approximately the same apparent MW as the E1~Ub and was therefore obscured from view, whereas neddylated Cul1 Cdc34-190~Ub was easily identified between the un-neddylated Cul1 Cdc34-190~Ub species and E1~Ub (from this we were able to estimate the efficiency of the neddylation reaction at approximately 50 %). Also note that thioester formation for the Cul1 Cdc34-WT and Cul1 Nedd8 Cdc34-WT fusions was abnormally low in this experiment which resulted in the lower than normal activity of this protein. 5
6 Table S1. Statistics for the bioinformatics analysis of the yeast proteome. Window Percent ORFs Rand1 Rand2 Rand3 Average Window: number of residues within the search window. Percent: minimum percentage of residues within the window that must be acidic. ORFs: the number of yeast Open Reading Frames with at least one stretch of residues that satisfy the previous two conditions. Rand1, Rand2, and Rand3: Expected number of ORFs if amino acid sequences are shuffled from three 6
7 independent randomizations. Average: the average expected value from Rand1, Rand2, and Rand3. : the standard deviation for the expected values from Rand1, Rand2 and Rand3. 7
8 8
9 9
10 10
11 11
12 12
13 13
SUPPLEMENTAL MATERIAL. Supplemental Methods:
 SUPPLEMENTAL MATERIAL Supplemental Methods: Immunoprecipitation- As we described but with some modifications [22]. As part of another ongoing project, lysate from human umbilical vein endothelial cells
SUPPLEMENTAL MATERIAL Supplemental Methods: Immunoprecipitation- As we described but with some modifications [22]. As part of another ongoing project, lysate from human umbilical vein endothelial cells
Figure S1. USP-46 is expressed in several tissues including the nervous system
 Supplemental Figure legends Figure S1. USP-46 is expressed in several tissues including the nervous system Transgenic animals expressing a transcriptional reporter (P::GFP) were imaged using epifluorescence
Supplemental Figure legends Figure S1. USP-46 is expressed in several tissues including the nervous system Transgenic animals expressing a transcriptional reporter (P::GFP) were imaged using epifluorescence
pt7ht vector and over-expressed in E. coli as inclusion bodies. Cells were lysed in 6 M
 Supplementary Methods MIG6 production, purification, inhibition, and kinase assays MIG6 segment 1 (30mer, residues 334 364) peptide was synthesized using standard solid-phase peptide synthesis as described
Supplementary Methods MIG6 production, purification, inhibition, and kinase assays MIG6 segment 1 (30mer, residues 334 364) peptide was synthesized using standard solid-phase peptide synthesis as described
Viral RNAi suppressor reversibly binds sirna to. outcompete Dicer and RISC via multiple-turnover
 Supplementary Data Viral RNAi suppressor reversibly binds sirna to outcompete Dicer and RISC via multiple-turnover Renata A. Rawlings 1,2, Vishalakshi Krishnan 2 and Nils G. Walter 2 * 1 Biophysics and
Supplementary Data Viral RNAi suppressor reversibly binds sirna to outcompete Dicer and RISC via multiple-turnover Renata A. Rawlings 1,2, Vishalakshi Krishnan 2 and Nils G. Walter 2 * 1 Biophysics and
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION Supplementary figures Supplementary Figure 1: Suv39h1, but not Suv39h2, promotes HP1α sumoylation in vivo. In vivo HP1α sumoylation assay. Top: experimental scheme. Middle: we
SUPPLEMENTARY INFORMATION Supplementary figures Supplementary Figure 1: Suv39h1, but not Suv39h2, promotes HP1α sumoylation in vivo. In vivo HP1α sumoylation assay. Top: experimental scheme. Middle: we
Fig. S1. CrgA intracellular levels in M. tuberculosis. Ten and twenty micrograms of
 Supplementary data Fig. S1. CrgA intracellular levels in M. tuberculosis. Ten and twenty micrograms of cell free protein lysates from WT M. tuberculosis (Rv) together with various known concentrations
Supplementary data Fig. S1. CrgA intracellular levels in M. tuberculosis. Ten and twenty micrograms of cell free protein lysates from WT M. tuberculosis (Rv) together with various known concentrations
Supplementary Information (Ha, et. al) Supplementary Figures Supplementary Fig. S1
 Supplementary Information (Ha, et. al) Supplementary Figures Supplementary Fig. S1 a His-ORMDL3 ~ 17 His-ORMDL3 GST-ORMDL3 - + - + IPTG GST-ORMDL3 ~ b Integrated Density (ORMDL3/ -actin) 0.4 0.3 0.2 0.1
Supplementary Information (Ha, et. al) Supplementary Figures Supplementary Fig. S1 a His-ORMDL3 ~ 17 His-ORMDL3 GST-ORMDL3 - + - + IPTG GST-ORMDL3 ~ b Integrated Density (ORMDL3/ -actin) 0.4 0.3 0.2 0.1
Coleman et al., Supplementary Figure 1
 Coleman et al., Supplementary Figure 1 BrdU Merge G1 Early S Mid S Supplementary Figure 1. Sequential destruction of CRL4 Cdt2 targets during the G1/S transition. HCT116 cells were synchronized by sequential
Coleman et al., Supplementary Figure 1 BrdU Merge G1 Early S Mid S Supplementary Figure 1. Sequential destruction of CRL4 Cdt2 targets during the G1/S transition. HCT116 cells were synchronized by sequential
Figure legends for supplement
 Figure legends for supplement Supplemental Figure 1 Characterization of purified and recombinant proteins Relevant fractions related the final stage of the purification protocol(bingham et al., 1998; Toba
Figure legends for supplement Supplemental Figure 1 Characterization of purified and recombinant proteins Relevant fractions related the final stage of the purification protocol(bingham et al., 1998; Toba
The molecular basis of lysine 48 ubiquitin chain synthesis by Ube2K
 Supplementary Information The molecular basis of lysine 48 ubiquitin chain synthesis by Adam J. Middleton, Catherine L. Day* Department of Biochemistry, Otago School of Medical Sciences, University of
Supplementary Information The molecular basis of lysine 48 ubiquitin chain synthesis by Adam J. Middleton, Catherine L. Day* Department of Biochemistry, Otago School of Medical Sciences, University of
Supplementary Figure S1 Purification of deubiquitinases HEK293 cells were transfected with the indicated DUB-expressing plasmids.
 Supplementary Figure S1 Purification of deubiquitinases HEK293 cells were transfected with the indicated DUB-expressing plasmids. The cells were harvested 72 h after transfection. FLAG-tagged deubiquitinases
Supplementary Figure S1 Purification of deubiquitinases HEK293 cells were transfected with the indicated DUB-expressing plasmids. The cells were harvested 72 h after transfection. FLAG-tagged deubiquitinases
Supplementary Figure 1. α-synuclein is truncated in PD and LBD brains. Nature Structural & Molecular Biology: doi: /nsmb.
 Supplementary Figure 1 α-synuclein is truncated in PD and LBD brains. (a) Specificity of anti-n103 antibody. Anti-N103 antibody was coated on an ELISA plate and different concentrations of full-length
Supplementary Figure 1 α-synuclein is truncated in PD and LBD brains. (a) Specificity of anti-n103 antibody. Anti-N103 antibody was coated on an ELISA plate and different concentrations of full-length
PDIP46 (DNA polymerase δ interacting protein 46) is an activating factor for human DNA polymerase δ
 PDIP46 (DNA polymerase δ interacting protein 46) is an activating factor for human DNA polymerase δ Supplementary Material Figure S1. PDIP46 is associated with Pol isolated by immunoaffinity chromatography.
PDIP46 (DNA polymerase δ interacting protein 46) is an activating factor for human DNA polymerase δ Supplementary Material Figure S1. PDIP46 is associated with Pol isolated by immunoaffinity chromatography.
T H E J O U R N A L O F C E L L B I O L O G Y
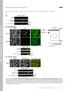 T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Nakajima and Tanoue, http://www.jcb.org/cgi/content/full/jcb.201104118/dc1 Figure S1. DLD-1 cells exhibit the characteristic morphology
T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Nakajima and Tanoue, http://www.jcb.org/cgi/content/full/jcb.201104118/dc1 Figure S1. DLD-1 cells exhibit the characteristic morphology
Supplemental Data. Farmer et al. (2010) Plant Cell /tpc
 Supplemental Figure 1. Amino acid sequence comparison of RAD23 proteins. Identical and similar residues are shown in the black and gray boxes, respectively. Dots denote gaps. The sequence of plant Ub is
Supplemental Figure 1. Amino acid sequence comparison of RAD23 proteins. Identical and similar residues are shown in the black and gray boxes, respectively. Dots denote gaps. The sequence of plant Ub is
Hossain_Supplemental Figure 1
 Hossain_Supplemental Figure 1 GFP-PACT GFP-PACT Motif I GFP-PACT Motif II A. MG132 (1µM) GFP Tubulin GFP-PACT Pericentrin GFP-PACT GFP-PACT Pericentrin Fig. S1. Expression and localization of Orc1 PACT
Hossain_Supplemental Figure 1 GFP-PACT GFP-PACT Motif I GFP-PACT Motif II A. MG132 (1µM) GFP Tubulin GFP-PACT Pericentrin GFP-PACT GFP-PACT Pericentrin Fig. S1. Expression and localization of Orc1 PACT
JCB. Supplemental material THE JOURNAL OF CELL BIOLOGY. Hong et al.,
 Supplemental material JCB Hong et al., http://www.jcb.org/cgi/content/full/jcb.201412127/dc1 THE JOURNAL OF CELL BIOLOGY Figure S1. Analysis of purified proteins by SDS-PAGE and pull-down assays. (A) Coomassie-stained
Supplemental material JCB Hong et al., http://www.jcb.org/cgi/content/full/jcb.201412127/dc1 THE JOURNAL OF CELL BIOLOGY Figure S1. Analysis of purified proteins by SDS-PAGE and pull-down assays. (A) Coomassie-stained
Supplementary data. sienigma. F-Enigma F-EnigmaSM. a-p53
 Supplementary data Supplemental Figure 1 A sienigma #2 sienigma sicontrol a-enigma - + ++ - - - - - - + ++ - - - - - - ++ B sienigma F-Enigma F-EnigmaSM a-flag HLK3 cells - - - + ++ + ++ - + - + + - -
Supplementary data Supplemental Figure 1 A sienigma #2 sienigma sicontrol a-enigma - + ++ - - - - - - + ++ - - - - - - ++ B sienigma F-Enigma F-EnigmaSM a-flag HLK3 cells - - - + ++ + ++ - + - + + - -
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature12119 SUPPLEMENTARY FIGURES AND LEGENDS pre-let-7a- 1 +14U pre-let-7a- 1 Ddx3x Dhx30 Dis3l2 Elavl1 Ggt5 Hnrnph 2 Osbpl5 Puf60 Rnpc3 Rpl7 Sf3b3 Sf3b4 Tia1 Triobp U2af1 U2af2 1 6 2 4 3
doi:10.1038/nature12119 SUPPLEMENTARY FIGURES AND LEGENDS pre-let-7a- 1 +14U pre-let-7a- 1 Ddx3x Dhx30 Dis3l2 Elavl1 Ggt5 Hnrnph 2 Osbpl5 Puf60 Rnpc3 Rpl7 Sf3b3 Sf3b4 Tia1 Triobp U2af1 U2af2 1 6 2 4 3
Polyclonal ARHGAP25 antibody was prepared from rabbit serum after intracutaneous
 Preparation and purification of polyclonal antibodies Polyclonal ARHGAP25 antibody was prepared from rabbit serum after intracutaneous injections of glutathione S-transferase-ARHGAP25-(509-619) (GST-coiled
Preparation and purification of polyclonal antibodies Polyclonal ARHGAP25 antibody was prepared from rabbit serum after intracutaneous injections of glutathione S-transferase-ARHGAP25-(509-619) (GST-coiled
Supplemental Materials and Methods
 Supplemental Materials and Methods Co-immunoprecipitation (Co-IP) assay Cells were lysed with NETN buffer (20 mm Tris-HCl, ph 8.0, 0 mm NaCl, 1 mm EDT, 0.5% Nonidet P-40) containing 50 mm β-glycerophosphate,
Supplemental Materials and Methods Co-immunoprecipitation (Co-IP) assay Cells were lysed with NETN buffer (20 mm Tris-HCl, ph 8.0, 0 mm NaCl, 1 mm EDT, 0.5% Nonidet P-40) containing 50 mm β-glycerophosphate,
Nature Structural & Molecular Biology: doi: /nsmb.1583
 Acetylation by GCN5 regulates CDC6 phosphorylation in the S-phase of the cell cycle Roberta Paolinelli 1,2, Ramiro Mendoza-Maldonado 2, Anna Cereseto 1 and Mauro Giacca 2 1 Molecular Biology Laboratory,
Acetylation by GCN5 regulates CDC6 phosphorylation in the S-phase of the cell cycle Roberta Paolinelli 1,2, Ramiro Mendoza-Maldonado 2, Anna Cereseto 1 and Mauro Giacca 2 1 Molecular Biology Laboratory,
Supplementary Fig. 1 Identification of Nedd4 as an IRS-2-associated protein in camp-treated FRTL-5 cells.
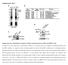 Supplementary Fig. 1 Supplementary Fig. 1 Identification of Nedd4 as an IRS-2-associated protein in camp-treated FRTL-5 cells. (a) FRTL-5 cells were treated with 1 mm dibutyryl camp for 24 h, and the lysates
Supplementary Fig. 1 Supplementary Fig. 1 Identification of Nedd4 as an IRS-2-associated protein in camp-treated FRTL-5 cells. (a) FRTL-5 cells were treated with 1 mm dibutyryl camp for 24 h, and the lysates
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb3363 Supplementary Figure 1 Several WNTs bind to the extracellular domains of PKD1. (a) HEK293T cells were co-transfected with indicated plasmids. Flag-tagged proteins were immunoprecipiated
DOI: 10.1038/ncb3363 Supplementary Figure 1 Several WNTs bind to the extracellular domains of PKD1. (a) HEK293T cells were co-transfected with indicated plasmids. Flag-tagged proteins were immunoprecipiated
supplementary information
 DOI: 10.1038/ncb2172 Figure S1 p53 regulates cellular NADPH and lipid levels via inhibition of G6PD. (a) U2OS cells stably expressing p53 shrna or a control shrna were transfected with control sirna or
DOI: 10.1038/ncb2172 Figure S1 p53 regulates cellular NADPH and lipid levels via inhibition of G6PD. (a) U2OS cells stably expressing p53 shrna or a control shrna were transfected with control sirna or
This is the author's accepted version of the manuscript.
 This is the author's accepted version of the manuscript. The definitive version is published in Nature Communications Online Edition: 2015/4/16 (Japan time), doi:10.1038/ncomms7780. The final version published
This is the author's accepted version of the manuscript. The definitive version is published in Nature Communications Online Edition: 2015/4/16 (Japan time), doi:10.1038/ncomms7780. The final version published
Expanded View Figures
 EMO reports rystal structure of Mis18 Yippee-like domain Lakxmi Subramanian et al Expanded View Figures Figure EV1. Structural characterization of the N-terminal Yippee-like globular domain of spmis18.
EMO reports rystal structure of Mis18 Yippee-like domain Lakxmi Subramanian et al Expanded View Figures Figure EV1. Structural characterization of the N-terminal Yippee-like globular domain of spmis18.
Supplementary methods
 Supplementary methods Cell culture, infection, transfection, and RNA interference HEK293 cells and its derivatives were grown in DMEM supplemented with 10% FBS. Various constructs were introduced into
Supplementary methods Cell culture, infection, transfection, and RNA interference HEK293 cells and its derivatives were grown in DMEM supplemented with 10% FBS. Various constructs were introduced into
Supplementary Figure 1. GST pull-down analysis of the interaction of GST-cIAP1 (A, B), GSTcIAP1
 Legends Supplementary Figure 1. GST pull-down analysis of the interaction of GST- (A, B), GST mutants (B) or GST- (C) with indicated proteins. A, B, Cell lysate from untransfected HeLa cells were loaded
Legends Supplementary Figure 1. GST pull-down analysis of the interaction of GST- (A, B), GST mutants (B) or GST- (C) with indicated proteins. A, B, Cell lysate from untransfected HeLa cells were loaded
Supplementary information, Figure S1A ShHTL7 interacted with MAX2 but not another F-box protein COI1.
 GR24 (μm) 0 20 0 20 GST-ShHTL7 anti-gst His-MAX2 His-COI1 PVDF staining Supplementary information, Figure S1A ShHTL7 interacted with MAX2 but not another F-box protein COI1. Pull-down assays using GST-ShHTL7
GR24 (μm) 0 20 0 20 GST-ShHTL7 anti-gst His-MAX2 His-COI1 PVDF staining Supplementary information, Figure S1A ShHTL7 interacted with MAX2 but not another F-box protein COI1. Pull-down assays using GST-ShHTL7
Nature Structural & Molecular Biology: doi: /nsmb.3308
 Supplementary Figure 1 Analysis of CED-3 autoactivation and CED-4-induced CED-3 activation. (a) Repeat experiments of Fig. 1a. (b) Repeat experiments of Fig. 1b. (c) Quantitative analysis of three independent
Supplementary Figure 1 Analysis of CED-3 autoactivation and CED-4-induced CED-3 activation. (a) Repeat experiments of Fig. 1a. (b) Repeat experiments of Fig. 1b. (c) Quantitative analysis of three independent
Rapid and sensitive determination of recombinant protein expression
 APPLIAION NOE Pro-Detect Rapid assays Rapid and sensitive determination of recombinant protein expression Introduction Recombinant protein expression and purification is a multistep process that includes:
APPLIAION NOE Pro-Detect Rapid assays Rapid and sensitive determination of recombinant protein expression Introduction Recombinant protein expression and purification is a multistep process that includes:
Supplemental Information
 Supplemental Information ATP-dependent unwinding of U4/U6 snrnas by the Brr2 helicase requires the C-terminus of Prp8 Corina Maeder 1,3, Alan K. Kutach 1,2,3, and Christine Guthrie 1 1 Department of Biochemistry
Supplemental Information ATP-dependent unwinding of U4/U6 snrnas by the Brr2 helicase requires the C-terminus of Prp8 Corina Maeder 1,3, Alan K. Kutach 1,2,3, and Christine Guthrie 1 1 Department of Biochemistry
Supplementary information
 Supplementary information The E3 ligase RNF8 regulates KU80 removal and NHEJ repair Lin Feng 1, Junjie Chen 1 1 Department of Experimental Radiation Oncology, The University of Texas M. D. Anderson Cancer
Supplementary information The E3 ligase RNF8 regulates KU80 removal and NHEJ repair Lin Feng 1, Junjie Chen 1 1 Department of Experimental Radiation Oncology, The University of Texas M. D. Anderson Cancer
Single-molecule imaging of DNA curtains reveals intrinsic energy landscapes for nucleosome deposition
 SUPPLEMENTARY INFORMATION Single-molecule imaging of DNA curtains reveals intrinsic energy landscapes for nucleosome deposition Mari-Liis Visnapuu 1 and Eric C. Greene 1 1 Department of Biochemistry &
SUPPLEMENTARY INFORMATION Single-molecule imaging of DNA curtains reveals intrinsic energy landscapes for nucleosome deposition Mari-Liis Visnapuu 1 and Eric C. Greene 1 1 Department of Biochemistry &
T H E J O U R N A L O F C E L L B I O L O G Y
 T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Craft et al., http://www.jcb.org/cgi/content/full/jcb.201409036/dc1 Figure S1. GFP -tubulin interacts with endogenous -tubulin. Ciliary
T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Craft et al., http://www.jcb.org/cgi/content/full/jcb.201409036/dc1 Figure S1. GFP -tubulin interacts with endogenous -tubulin. Ciliary
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Detection of MCM-subunit SUMOylation under normal growth conditions. a. Sumoylated forms of MCM subunits show differential shifts when SUMO is attached to differently sized tags.
Supplementary Figure 1 Detection of MCM-subunit SUMOylation under normal growth conditions. a. Sumoylated forms of MCM subunits show differential shifts when SUMO is attached to differently sized tags.
Isolation of the recombinant middle and head + middle modules.
 Supplementary Figure 1 Isolation of the recombinant middle and head + middle modules. (a) Scheme illustrating the multi-step purification protocol for the reconstituted middle module. Extract from infected
Supplementary Figure 1 Isolation of the recombinant middle and head + middle modules. (a) Scheme illustrating the multi-step purification protocol for the reconstituted middle module. Extract from infected
Glutathione Resin. (Cat. # , , , ) think proteins! think G-Biosciences
 191PR 05 G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name Glutathione Resin (Cat. # 786 280, 786 310, 786 311, 786 312) think proteins! think
191PR 05 G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name Glutathione Resin (Cat. # 786 280, 786 310, 786 311, 786 312) think proteins! think
SUPPLEMENTARY INFORMATION
 doi: 10.1038/nature06721 SUPPLEMENTARY INFORMATION. Supplemental Figure Legends Supplemental Figure 1 The distribution of hatx-1[82q] in Cos7 cells. Cos7 cells are co-transfected with hatx-1[82q]-gfp (green)
doi: 10.1038/nature06721 SUPPLEMENTARY INFORMATION. Supplemental Figure Legends Supplemental Figure 1 The distribution of hatx-1[82q] in Cos7 cells. Cos7 cells are co-transfected with hatx-1[82q]-gfp (green)
indicated numbers of pups at day of life (DOL) 10, or embryonic day (ED) B. Male mice of
 SUPPLEMENTRY FIGURE LEGENDS Figure S1. USP44 loss leads to chromosome missegregation.. Genotypes obtained from the indicated numbers of pups at day of life (DOL) 10, or embryonic day (ED) 13.5.. Male mice
SUPPLEMENTRY FIGURE LEGENDS Figure S1. USP44 loss leads to chromosome missegregation.. Genotypes obtained from the indicated numbers of pups at day of life (DOL) 10, or embryonic day (ED) 13.5.. Male mice
supplementary information
 supplementary information Figure S1 (a) Alignment of the Sgf11 amino acid sequences from S. cerevisiae, Yarrowia lipolytica and S. pombe with Sgf73 sequences from S. cerevisiae, S. pombe and Candida albicans.
supplementary information Figure S1 (a) Alignment of the Sgf11 amino acid sequences from S. cerevisiae, Yarrowia lipolytica and S. pombe with Sgf73 sequences from S. cerevisiae, S. pombe and Candida albicans.
Supplementary Fig. 1. Schematic structure of TRAIP and RAP80. The prey line below TRAIP indicates bait and the two lines above RAP80 highlight the
 Supplementary Fig. 1. Schematic structure of TRAIP and RAP80. The prey line below TRAIP indicates bait and the two lines above RAP80 highlight the prey clones identified in the yeast two hybrid screen.
Supplementary Fig. 1. Schematic structure of TRAIP and RAP80. The prey line below TRAIP indicates bait and the two lines above RAP80 highlight the prey clones identified in the yeast two hybrid screen.
Supplementary Information
 Supplementary Information Supplementary Figure 1. Elevated Smurf1 expression is associated with the progression of colorectal cancer. (a) The tumor microarray analysis with anti-smurf1 antibody. (b) Box
Supplementary Information Supplementary Figure 1. Elevated Smurf1 expression is associated with the progression of colorectal cancer. (a) The tumor microarray analysis with anti-smurf1 antibody. (b) Box
Supplementary Materials and Methods
 Supplementary Materials and Methods sirna sequences used in this study The sequences of Stealth Select RNAi for ALK and FLOT-1 were as follows: ALK sense no.1 (ALK): 5 -AAUACUGACAGCCACAGGCAAUGUC-3 ; ALK
Supplementary Materials and Methods sirna sequences used in this study The sequences of Stealth Select RNAi for ALK and FLOT-1 were as follows: ALK sense no.1 (ALK): 5 -AAUACUGACAGCCACAGGCAAUGUC-3 ; ALK
Quantifying small numbers of antibodies with a near-universal protein-dna chimera
 Quantifying small numbers of antibodies with a near-universal protein-dna chimera Ian Burbulis, Kumiko Yamaguchi, Richard Yu, Orna Resnekov & Roger Brent Supplementary figures and text: Supplementary figure
Quantifying small numbers of antibodies with a near-universal protein-dna chimera Ian Burbulis, Kumiko Yamaguchi, Richard Yu, Orna Resnekov & Roger Brent Supplementary figures and text: Supplementary figure
SUPPLEMENTARY INFORMATION
 a 14 12 Densitometry (AU) 1 8 6 4 2 t b 16 NMHC-IIA GAPDH NMHC-IIB Densitometry (AU) 14 12 1 8 6 4 2 1 nm 1 nm 1 nm 1 nm sirna 1 nm 1 nm Figure S1 S4 Quantification of protein levels. (a) The microtubule
a 14 12 Densitometry (AU) 1 8 6 4 2 t b 16 NMHC-IIA GAPDH NMHC-IIB Densitometry (AU) 14 12 1 8 6 4 2 1 nm 1 nm 1 nm 1 nm sirna 1 nm 1 nm Figure S1 S4 Quantification of protein levels. (a) The microtubule
Western blot troubleshooting guide
 Specializing in Secondary Antibodies and Conjugates Western blot troubleshooting guide Optimize your Western blotting with Jackson ImmunoResearch Secondary antibodies Troubleshooting for better blots Western
Specializing in Secondary Antibodies and Conjugates Western blot troubleshooting guide Optimize your Western blotting with Jackson ImmunoResearch Secondary antibodies Troubleshooting for better blots Western
Technical Note Detection of post-immunoprecipitation proteins by Western blot using the Quick Western Kit IRDye 680RD
 Technical Note Detection of post-immunoprecipitation proteins by Western blot using the Quick Western Kit IRDye 680RD Developed for: Aerius, Odyssey Classic, Odyssey CLx and Odyssey Sa Imaging Systems
Technical Note Detection of post-immunoprecipitation proteins by Western blot using the Quick Western Kit IRDye 680RD Developed for: Aerius, Odyssey Classic, Odyssey CLx and Odyssey Sa Imaging Systems
Fig. S1. Effect of p120-catenin overexpression on the interaction of SCUBE2 with E-cadherin. The expression plasmid encoding FLAG.
 Fig. S1. Effect of p120-catenin overexpression on the interaction of SCUBE2 with E-cadherin. The expression plasmid encoding FLAG.SCUBE2, E-cadherin.Myc, or HA.p120-catenin was transfected in a combination
Fig. S1. Effect of p120-catenin overexpression on the interaction of SCUBE2 with E-cadherin. The expression plasmid encoding FLAG.SCUBE2, E-cadherin.Myc, or HA.p120-catenin was transfected in a combination
Solutions to 7.02 Quiz II 10/27/05
 Solutions to 7.02 Quiz II 10/27/05 Class Average = 83 Standard Deviation = 9 Range Grade % 87-100 A 43 74-86 B 39 55-73 C 17 > 54 D 1 Question 1 (56 points) While studying deep sea bacteria, you discover
Solutions to 7.02 Quiz II 10/27/05 Class Average = 83 Standard Deviation = 9 Range Grade % 87-100 A 43 74-86 B 39 55-73 C 17 > 54 D 1 Question 1 (56 points) While studying deep sea bacteria, you discover
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION Dynamic Phosphorylation of HP1 Regulates Mitotic Progression in Human Cells Supplementary Figures Supplementary Figure 1. NDR1 interacts with HP1. (a) Immunoprecipitation using
SUPPLEMENTARY INFORMATION Dynamic Phosphorylation of HP1 Regulates Mitotic Progression in Human Cells Supplementary Figures Supplementary Figure 1. NDR1 interacts with HP1. (a) Immunoprecipitation using
SUPPLEMENTARY INFORMATION
 (Supplementary Methods and Materials) GST pull-down assay GST-fusion proteins Fe65 365-533, and Fe65 538-700 were expressed in BL21 bacterial cells and purified with glutathione-agarose beads (Sigma).
(Supplementary Methods and Materials) GST pull-down assay GST-fusion proteins Fe65 365-533, and Fe65 538-700 were expressed in BL21 bacterial cells and purified with glutathione-agarose beads (Sigma).
Figure S1. Verification of ihog Mutation by Protein Immunoblotting Figure S2. Verification of ihog and boi
 Figure S1. Verification of ihog Mutation by Protein Immunoblotting Extracts from S2R+ cells, embryos, and adults were analyzed by immunoprecipitation and immunoblotting with anti-ihog antibody. The Ihog
Figure S1. Verification of ihog Mutation by Protein Immunoblotting Extracts from S2R+ cells, embryos, and adults were analyzed by immunoprecipitation and immunoblotting with anti-ihog antibody. The Ihog
GST Elution Buffer. (Cat. # ) think proteins! think G-Biosciences
 191PR-05 G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name GST Elution Buffer (Cat. #786-541) think proteins! think G-Biosciences www.gbiosciences.com
191PR-05 G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name GST Elution Buffer (Cat. #786-541) think proteins! think G-Biosciences www.gbiosciences.com
Glutathione Resin. (Cat. # , , , ) think proteins! think G-Biosciences
 191PR-05 G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name Glutathione Resin (Cat. # 786-280, 786-310, 786-311, 786-312) think proteins! think
191PR-05 G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name Glutathione Resin (Cat. # 786-280, 786-310, 786-311, 786-312) think proteins! think
Supplemental Information. Pacer Mediates the Function of Class III PI3K. and HOPS Complexes in Autophagosome. Maturation by Engaging Stx17
 Molecular Cell, Volume 65 Supplemental Information Pacer Mediates the Function of Class III PI3K and HOPS Complexes in Autophagosome Maturation by Engaging Stx17 Xiawei Cheng, Xiuling Ma, Xianming Ding,
Molecular Cell, Volume 65 Supplemental Information Pacer Mediates the Function of Class III PI3K and HOPS Complexes in Autophagosome Maturation by Engaging Stx17 Xiawei Cheng, Xiuling Ma, Xianming Ding,
human Cdc45 Figure 1c. (c)
 1 Details of the refined crystallographic model of human Cdc45 and comparison of its active-site region with that of bacterial RecJ. (a) Stereo view of a representative example of the final 2F o -F c electron
1 Details of the refined crystallographic model of human Cdc45 and comparison of its active-site region with that of bacterial RecJ. (a) Stereo view of a representative example of the final 2F o -F c electron
ONE-HOUR Complete IP-Western Kit
 ONE-HOUR Complete IP-Western Kit Technical Manual No. 0218 Version 06192009 I Description.. 1 II Kit Contents.. 2 III Related Products 2 IV Key Features.. 2 V Storage.. 2 VI ONE-HOUR Western Protocol 3
ONE-HOUR Complete IP-Western Kit Technical Manual No. 0218 Version 06192009 I Description.. 1 II Kit Contents.. 2 III Related Products 2 IV Key Features.. 2 V Storage.. 2 VI ONE-HOUR Western Protocol 3
Supplementary Figure 1. APP cleavage assay. HEK293 cells were transfected with various
 Supplementary Figure 1. APP cleavage assay. HEK293 cells were transfected with various GST-tagged N-terminal truncated APP fragments including GST-APP full-length (FL), APP (123-695), APP (189-695), or
Supplementary Figure 1. APP cleavage assay. HEK293 cells were transfected with various GST-tagged N-terminal truncated APP fragments including GST-APP full-length (FL), APP (123-695), APP (189-695), or
Supporting Information
 Electronic Supplementary Material (ESI) for Nanoscale. This journal is The Royal Society of Chemistry 215 Supporting Information Quantitative Description of Thermodynamic and Kinetic Properties of the
Electronic Supplementary Material (ESI) for Nanoscale. This journal is The Royal Society of Chemistry 215 Supporting Information Quantitative Description of Thermodynamic and Kinetic Properties of the
Supplementary Figure S1 Supplementary Figure S2 Supplementary Figure S3. Supplementary Figure S4
 Supplementary Figure S1 Supplementary Figure S2 Supplementary Figure S3 Supplementary Figure S4 Supplementary Figure S5 Supplementary Figure S6 Supplementary Figure S7 Supplementary Figure S8 Supplementary
Supplementary Figure S1 Supplementary Figure S2 Supplementary Figure S3 Supplementary Figure S4 Supplementary Figure S5 Supplementary Figure S6 Supplementary Figure S7 Supplementary Figure S8 Supplementary
Reach greater highs. (and lows ) with new protein ladder choices. Fermentas now sold as. Thermo Scientific
 Introducing Thermo Scientific Pierce Prestained and Unstained s for easy protein molecular weight determination. Fermentas now sold as Thermo Scientific Reach greater highs (and lows ) with new protein
Introducing Thermo Scientific Pierce Prestained and Unstained s for easy protein molecular weight determination. Fermentas now sold as Thermo Scientific Reach greater highs (and lows ) with new protein
Recruitment of Grb2 to surface IgG and IgE provides antigen receptor-intrinsic costimulation to class-switched B cells
 SUPPLEMENTARY FIGURES Recruitment of Grb2 to surface IgG and IgE provides antigen receptor-intrinsic costimulation to class-switched B cells Niklas Engels, Lars Morten König, Christina Heemann, Johannes
SUPPLEMENTARY FIGURES Recruitment of Grb2 to surface IgG and IgE provides antigen receptor-intrinsic costimulation to class-switched B cells Niklas Engels, Lars Morten König, Christina Heemann, Johannes
Supplementary Online Material. Structural mimicry in transcription regulation of human RNA polymerase II by the. DNA helicase RECQL5
 Supplementary Online Material Structural mimicry in transcription regulation of human RNA polymerase II by the DNA helicase RECQL5 Susanne A. Kassube, Martin Jinek, Jie Fang, Susan Tsutakawa and Eva Nogales
Supplementary Online Material Structural mimicry in transcription regulation of human RNA polymerase II by the DNA helicase RECQL5 Susanne A. Kassube, Martin Jinek, Jie Fang, Susan Tsutakawa and Eva Nogales
Supplementary Fig. 1
 a FL (1-2266) NL (1-1190) CL (1191-2266) HA-ICE1: - HA-ICE1: - - - FLAG-ICE2: + + + + FLAG-ELL: + + + + + + IP: anti-ha FLAG-ICE2 HA-ICE1-FL HA-ICE1-NL HA-ICE1-CL FLAG-ICE2 b IP: anti-ha FL (1-2266) NL
a FL (1-2266) NL (1-1190) CL (1191-2266) HA-ICE1: - HA-ICE1: - - - FLAG-ICE2: + + + + FLAG-ELL: + + + + + + IP: anti-ha FLAG-ICE2 HA-ICE1-FL HA-ICE1-NL HA-ICE1-CL FLAG-ICE2 b IP: anti-ha FL (1-2266) NL
Supporting Information
 Supporting Information Stavru et al. 0.073/pnas.357840 SI Materials and Methods Immunofluorescence. For immunofluorescence, cells were fixed for 0 min in 4% (wt/vol) paraformaldehyde (Electron Microscopy
Supporting Information Stavru et al. 0.073/pnas.357840 SI Materials and Methods Immunofluorescence. For immunofluorescence, cells were fixed for 0 min in 4% (wt/vol) paraformaldehyde (Electron Microscopy
Cdc42 Activation Assay Kit
 A helping hand for your research Product Manual Configuration-specific Monoclonal Antibody Based Cdc42 Activation Assay Kit Catalog Number: 80701 20 assays 1 Table of Content Product Description 3 Assay
A helping hand for your research Product Manual Configuration-specific Monoclonal Antibody Based Cdc42 Activation Assay Kit Catalog Number: 80701 20 assays 1 Table of Content Product Description 3 Assay
Supplemental Figure 1 HDA18 has an HDAC domain and therefore has concentration dependent and TSA inhibited histone deacetylase activity.
 Supplemental Figure 1 HDA18 has an HDAC domain and therefore has concentration dependent and TSA inhibited histone deacetylase activity. (A) Amino acid alignment of HDA5, HDA15 and HDA18. The blue line
Supplemental Figure 1 HDA18 has an HDAC domain and therefore has concentration dependent and TSA inhibited histone deacetylase activity. (A) Amino acid alignment of HDA5, HDA15 and HDA18. The blue line
Antibody Profiling on Invitrogen ProtoArray High-Density Protein Microarrays
 1 Antibody Profiling on Invitrogen ProtoArray High-Density Protein Microarrays The application of antibodies in research and development, in vitro/in vivo diagnostics, or therapeutics requires that antibodies
1 Antibody Profiling on Invitrogen ProtoArray High-Density Protein Microarrays The application of antibodies in research and development, in vitro/in vivo diagnostics, or therapeutics requires that antibodies
Supplementary Figure 1
 Supplementary Figure 1 Supplementary Figure 1. Analysis of unique sequences isolated in Sort 14. Unique sequences are listed next to their frequency of occurrence, in parenthesis. Dithiol resistance is
Supplementary Figure 1 Supplementary Figure 1. Analysis of unique sequences isolated in Sort 14. Unique sequences are listed next to their frequency of occurrence, in parenthesis. Dithiol resistance is
Figure S1. Sequence alignments of ATRIP and ATR TopBP1 interacting regions.
 A H. sapiens 204 TKLQTS--ERANKLAAPSVSH VSPRKNPSVVIKPEACS-PQFGKTSFPTKESFSANMS LP 259 B. taurus 201 TKLQSS--ERANKLAVPTVSH VSPRKSPSVVIKPEACS-PQFGKPSFPTKESFSANKS LP 257 M. musculus 204 TKSQSN--GRTNKPAAPSVSH
A H. sapiens 204 TKLQTS--ERANKLAAPSVSH VSPRKNPSVVIKPEACS-PQFGKTSFPTKESFSANMS LP 259 B. taurus 201 TKLQSS--ERANKLAVPTVSH VSPRKSPSVVIKPEACS-PQFGKPSFPTKESFSANKS LP 257 M. musculus 204 TKSQSN--GRTNKPAAPSVSH
Supplementary Information for. Regulation of Rev1 by the Fanconi Anemia Core Complex
 Supplementary Information for Regulation of Rev1 by the Fanconi Anemia Core Complex Hyungjin Kim, Kailin Yang, Donniphat Dejsuphong, Alan D. D Andrea* *Corresponding Author: Alan D. D Andrea, M.D. Alan_dandrea@dfci.harvard.edu
Supplementary Information for Regulation of Rev1 by the Fanconi Anemia Core Complex Hyungjin Kim, Kailin Yang, Donniphat Dejsuphong, Alan D. D Andrea* *Corresponding Author: Alan D. D Andrea, M.D. Alan_dandrea@dfci.harvard.edu
Supplementary information
 Supplementary information Table of Content: Supplementary Results... 2 Supplementary Figure S1: Experimental validation of AP-MS results by coimmunprecipitation Western blot analysis.... 3 Supplementary
Supplementary information Table of Content: Supplementary Results... 2 Supplementary Figure S1: Experimental validation of AP-MS results by coimmunprecipitation Western blot analysis.... 3 Supplementary
A novel mechanism of post-translational modulation of HMGA functions by the histone chaperone nucleophosmin.
 A novel mechanism of post-translational modulation of HMGA functions by the histone chaperone nucleophosmin. Laura Arnoldo 1,#, Riccardo Sgarra 1,#, Eusebio Chiefari 2, Stefania Iiritano 2, Biagio Arcidiacono
A novel mechanism of post-translational modulation of HMGA functions by the histone chaperone nucleophosmin. Laura Arnoldo 1,#, Riccardo Sgarra 1,#, Eusebio Chiefari 2, Stefania Iiritano 2, Biagio Arcidiacono
T H E J O U R N A L O F C E L L B I O L O G Y
 T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Han et al., http://www.jcb.org/cgi/content/full/jcb.201311007/dc1 Figure S1. SIVA1 interacts with PCNA. (A) HEK293T cells were transiently
T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Han et al., http://www.jcb.org/cgi/content/full/jcb.201311007/dc1 Figure S1. SIVA1 interacts with PCNA. (A) HEK293T cells were transiently
CD93 and dystroglycan cooperation in human endothelial cell adhesion and migration
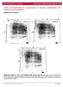 /, Supplementary Advance Publications Materials 2016 CD93 and dystroglycan cooperation in human endothelial cell adhesion and migration Supplementary Materials Supplementary Figure S1: In ECs CD93 silencing
/, Supplementary Advance Publications Materials 2016 CD93 and dystroglycan cooperation in human endothelial cell adhesion and migration Supplementary Materials Supplementary Figure S1: In ECs CD93 silencing
T H E J O U R N A L O F C E L L B I O L O G Y
 Supplemental material Moutin et al., http://www.jcb.org/cgi/content/full/jcb.201110101/dc1 T H E J O U R N A L O F C E L L B I O L O G Y Figure S1. Tagged Homer1a and Homer are functional and display different
Supplemental material Moutin et al., http://www.jcb.org/cgi/content/full/jcb.201110101/dc1 T H E J O U R N A L O F C E L L B I O L O G Y Figure S1. Tagged Homer1a and Homer are functional and display different
ONE-HOUR Western TM Multiplex Kit II
 ONE-HOUR Western TM Multiplex Kit II Technical Manual No. 0256 Version 06192009 I Description... 1 II Kit Contents.. 2 III Related Products 2 IV Key Features. 2 V Storage... 2 VI ONE-HOUR Multiplex Western
ONE-HOUR Western TM Multiplex Kit II Technical Manual No. 0256 Version 06192009 I Description... 1 II Kit Contents.. 2 III Related Products 2 IV Key Features. 2 V Storage... 2 VI ONE-HOUR Multiplex Western
VCP adaptor interactions are exceptionally dynamic and subject to differential modulation by a VCP inhibitor
 VCP adaptor interactions are exceptionally dynamic and subject to differential modulation by a VCP inhibitor Liang Xue 1, Emily E. Blythe 1, Elyse C. Freiberger 2, Jennifer Mamrosh 1, Alexander S. Hebert
VCP adaptor interactions are exceptionally dynamic and subject to differential modulation by a VCP inhibitor Liang Xue 1, Emily E. Blythe 1, Elyse C. Freiberger 2, Jennifer Mamrosh 1, Alexander S. Hebert
Supplementary methods Shoc2 In Vitro Ubiquitination Assay
 Supplementary methods Shoc2 In Vitro Ubiquitination Assay 35 S-labelled Shoc2 was prepared using a TNT quick Coupled transcription/ translation System (Promega) as recommended by manufacturer. For the
Supplementary methods Shoc2 In Vitro Ubiquitination Assay 35 S-labelled Shoc2 was prepared using a TNT quick Coupled transcription/ translation System (Promega) as recommended by manufacturer. For the
Figure S1 is related to Figure 1B, showing more details of outer segment of
 Supplemental Information Supplementary Figure legends and Figures Figure S1. Electron microscopic images in Sema4A +/+ and Sema4A / retinas Figure S1 is related to Figure 1B, showing more details of outer
Supplemental Information Supplementary Figure legends and Figures Figure S1. Electron microscopic images in Sema4A +/+ and Sema4A / retinas Figure S1 is related to Figure 1B, showing more details of outer
Autophagy induction [h]
![Autophagy induction [h] Autophagy induction [h]](/thumbs/80/80564099.jpg) A SD-N [h]: SD-N [h]: 1 2 4 1 2 4 1 2 4 -GFP 1 2 4 percentage GFP-Atg8 [%] 1 9 8 7 6 5 4 3 2 1 1 2 4 Autophagy induction [h] -13xmyc SD-N [h]: 1 2 4 -GFP -13xmyc 1 prape1 mape1 - - GFP - 13xmyc mape1 [%]
A SD-N [h]: SD-N [h]: 1 2 4 1 2 4 1 2 4 -GFP 1 2 4 percentage GFP-Atg8 [%] 1 9 8 7 6 5 4 3 2 1 1 2 4 Autophagy induction [h] -13xmyc SD-N [h]: 1 2 4 -GFP -13xmyc 1 prape1 mape1 - - GFP - 13xmyc mape1 [%]
Figure S1 early log mid log late log stat.
 Figure S1 early log mid log late log stat. C D E F Fig. S1. The chromosome-expressed EI localizes to the cell poles independent of the growth phase and growth media. MG1655 (ptsi-mcherry) cells, which
Figure S1 early log mid log late log stat. C D E F Fig. S1. The chromosome-expressed EI localizes to the cell poles independent of the growth phase and growth media. MG1655 (ptsi-mcherry) cells, which
Lecture 25 (11/15/17)
 Lecture 25 (11/15/17) Reading: Ch9; 328-332 Ch25; 990-995, 1005-1012 Problems: Ch9 (study-guide: applying); 1,2 Ch9 (study-guide: facts); 7,8 Ch25 (text); 1-3,5-7,9,10,13-15 Ch25 (study-guide: applying);
Lecture 25 (11/15/17) Reading: Ch9; 328-332 Ch25; 990-995, 1005-1012 Problems: Ch9 (study-guide: applying); 1,2 Ch9 (study-guide: facts); 7,8 Ch25 (text); 1-3,5-7,9,10,13-15 Ch25 (study-guide: applying);
Comparison of different methods for purification analysis of a green fluorescent Strep-tag fusion protein. Application
 Comparison of different methods for purification analysis of a green fluorescent Strep-tag fusion protein Application Petra Sebastian Meike Kuschel Stefan Schmidt Abstract This Application Note describes
Comparison of different methods for purification analysis of a green fluorescent Strep-tag fusion protein Application Petra Sebastian Meike Kuschel Stefan Schmidt Abstract This Application Note describes
Arf6 Activation Assay Kit
 A helping hand for your research Product Manual Configuration-specific Monoclonal Antibody Based Arf6 Activation Assay Kit Catalog Number: 82401 20 assays NewEast Biosciences 1 Table of Content Product
A helping hand for your research Product Manual Configuration-specific Monoclonal Antibody Based Arf6 Activation Assay Kit Catalog Number: 82401 20 assays NewEast Biosciences 1 Table of Content Product
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb2579 Figure S1 Incorporation of heavy isotope-labeled amino acids and enrichment of di-glycine modified peptides. The incorporation of isotopelabeled amino acids in peptides was calculated
DOI: 10.1038/ncb2579 Figure S1 Incorporation of heavy isotope-labeled amino acids and enrichment of di-glycine modified peptides. The incorporation of isotopelabeled amino acids in peptides was calculated
Conformation of the Mineralocorticoid Receptor N- terminal Domain: Evidence for Induced and Stable Structure
 ME-10-0005 Conformation of the Mineralocorticoid Receptor N- terminal Domain: Evidence for Induced and Stable Structure Katharina Fischer 1, Sharon M. Kelly 2, Kate Watt 1, Nicholas C. Price 2 and Iain
ME-10-0005 Conformation of the Mineralocorticoid Receptor N- terminal Domain: Evidence for Induced and Stable Structure Katharina Fischer 1, Sharon M. Kelly 2, Kate Watt 1, Nicholas C. Price 2 and Iain
Four different active promoter genes were chosen, ATXN7L2, PSRC1, CELSR2 and
 SUPPLEMENTARY MATERIALS AND METHODS Chromatin Immunoprecipitation for qpcr analysis Four different active promoter genes were chosen, ATXN7L2, PSRC1, CELSR2 and IL24, all located on chromosome 1. Primer
SUPPLEMENTARY MATERIALS AND METHODS Chromatin Immunoprecipitation for qpcr analysis Four different active promoter genes were chosen, ATXN7L2, PSRC1, CELSR2 and IL24, all located on chromosome 1. Primer
Supplementary Figure 1. Nature Structural & Molecular Biology: doi: /nsmb.3494
 Supplementary Figure 1 Pol structure-function analysis (a) Inactivating polymerase and helicase mutations do not alter the stability of Pol. Flag epitopes were introduced using CRISPR/Cas9 gene targeting
Supplementary Figure 1 Pol structure-function analysis (a) Inactivating polymerase and helicase mutations do not alter the stability of Pol. Flag epitopes were introduced using CRISPR/Cas9 gene targeting
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION Contents: Supplementary Figure 1. Additional structural and binding data for designed tuim peptides. Supplementary Figure 2. Subcellular localization patterns of designed tuim
SUPPLEMENTARY INFORMATION Contents: Supplementary Figure 1. Additional structural and binding data for designed tuim peptides. Supplementary Figure 2. Subcellular localization patterns of designed tuim
HPV E6 oncoprotein targets histone methyltransferases for modulating specific. Chih-Hung Hsu, Kai-Lin Peng, Hua-Ci Jhang, Chia-Hui Lin, Shwu-Yuan Wu,
 1 HPV E oncoprotein targets histone methyltransferases for modulating specific gene transcription 3 5 Chih-Hung Hsu, Kai-Lin Peng, Hua-Ci Jhang, Chia-Hui Lin, Shwu-Yuan Wu, Cheng-Ming Chiang, Sheng-Chung
1 HPV E oncoprotein targets histone methyltransferases for modulating specific gene transcription 3 5 Chih-Hung Hsu, Kai-Lin Peng, Hua-Ci Jhang, Chia-Hui Lin, Shwu-Yuan Wu, Cheng-Ming Chiang, Sheng-Chung
Figure 1: TDP-43 is subject to lysine acetylation within the RNA-binding domain a) QBI-293 cells were transfected with TDP-43 in the presence or
 Figure 1: TDP-43 is subject to lysine acetylation within the RNA-binding domain a) QBI-293 cells were transfected with TDP-43 in the presence or absence of the acetyltransferase CBP and acetylated TDP-43
Figure 1: TDP-43 is subject to lysine acetylation within the RNA-binding domain a) QBI-293 cells were transfected with TDP-43 in the presence or absence of the acetyltransferase CBP and acetylated TDP-43
Analysing protein protein interactions using a GST-fusion protein to pull down the interacting target from the cell lysate Hong Wang and Xin Zeng
 Analysing protein protein interactions using a GST-fusion protein to pull down the interacting target from the cell lysate Hong Wang and Xin Zeng Department of Molecular Genetics, Biochemistry and Microbiology,
Analysing protein protein interactions using a GST-fusion protein to pull down the interacting target from the cell lysate Hong Wang and Xin Zeng Department of Molecular Genetics, Biochemistry and Microbiology,
SUPPLEMENTARY INFORMATION
 The Supplementary Information (SI) Methods Cell culture and transfections H1299, U2OS, 293, HeLa cells were maintained in DMEM medium supplemented with 10% fetal bovine serum. H1299 and 293 cells were
The Supplementary Information (SI) Methods Cell culture and transfections H1299, U2OS, 293, HeLa cells were maintained in DMEM medium supplemented with 10% fetal bovine serum. H1299 and 293 cells were
VERIFY Tagged Antigen. Validation Data
 VERIFY Tagged Antigen Validation Data Antibody Validation Figure 1. Over-expression cell lysate for human STAT3 (NM_139276) was used to test 3 commercial antibodies. Antibody A shows strong antigen binding.
VERIFY Tagged Antigen Validation Data Antibody Validation Figure 1. Over-expression cell lysate for human STAT3 (NM_139276) was used to test 3 commercial antibodies. Antibody A shows strong antigen binding.
Supporting Information
 Supporting Information Deng et al. 10.1073/pnas.1515692112 SI Materials and Methods FPLC. All fusion proteins were expressed and purified through a three-step FPLC purification protocol, as described (20),
Supporting Information Deng et al. 10.1073/pnas.1515692112 SI Materials and Methods FPLC. All fusion proteins were expressed and purified through a three-step FPLC purification protocol, as described (20),
Dolphin-Chemi Plus. Aim: To visualise and evaluate the performance of chemiluminescent immunoblots using Wealtec s Dolphin-Chemi plus image system
 Application Note 03 Dolphin-Chemi plus 8/22/2007 Dolphin-Chemi Plus Aim: To visualise and evaluate the performance of chemiluminescent immunoblots using Wealtec s Dolphin-Chemi plus image system INTRODUCTION
Application Note 03 Dolphin-Chemi plus 8/22/2007 Dolphin-Chemi Plus Aim: To visualise and evaluate the performance of chemiluminescent immunoblots using Wealtec s Dolphin-Chemi plus image system INTRODUCTION
Figure S1, related to Figure 1. Characterization of biosensor behavior in vivo.
 Developmental Cell, Volume 23 Supplemental Information Separase Biosensor Reveals that Cohesin Cleavage Timing Depends on Phosphatase PP2A Cdc55 Regulation Gilad Yaakov, Kurt Thorn, and David O. Morgan
Developmental Cell, Volume 23 Supplemental Information Separase Biosensor Reveals that Cohesin Cleavage Timing Depends on Phosphatase PP2A Cdc55 Regulation Gilad Yaakov, Kurt Thorn, and David O. Morgan
