Supplemental figure 1
|
|
|
- Winifred Barber
- 6 years ago
- Views:
Transcription
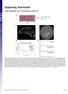
1 Supplemental figure Copy number ratio relative to beta actin Series1 Series2 Series3 0 1 PC3 BP 2 DU145 BP 3 PC3 CMV 4 DU145 CMV Supplemental figure 1. qpcr to quantify the relative copy number of TMEM135 CCDC67 breakpoint and pcmv vector sequence in the genome of transformed cancer cells. One microgram of genome DNA of PC3 BP (PC3 cells transformed with pcmv TMEM135 int13 CCDC67 int9 ), or DU145 BP (DU145 cells transformed with pcmv TMEM135 int13 CCDC67 int9 ), or PC3 CMV (PC3 cells transformed with pcmvscript) or DU145 CMV (DU145 cells transformed with pcmvscript) was quantified for actin or TMEM135 CCDC67 breakpoint through qpcr using the primers listed in Supplemental table 2. The copy numbers of BP and actin were fitted with standard curves generated with serial titrations of known copy numbers of BP and actin, respectively. The BP/ actin ratios were plotted. The CMV/β actin ratios were plotted similarly.
2 Supplemental figure 2 A Virus TMEM intron 13 EGFP tk CCDC67 intron bp 561 bp Genome CMV IE94 Splice acceptor Splice donor TMEM135 int 13 CCDC67 int bp 1234 bp Genome CMV IE94 TMEM intron 13 EGFP tk CCDC67 intron bp 1211 bp Pre integration Pre integration Post integration Post integration Supplemental figure 2A. Schematic diagram for the detection of TMEM135 int13 EGFP tk CCDC67 int9 integration into TMEM135 CCDC67 breakpoint in the PC3 cell genome. Arrows indicate the primer position for PCR. Putative integration sites that generated mutations are indicated by yellow stars. The PCR products obtained from xenografted PC3 cells that contain TMEM135 CCDC67 breakpoint before virus treatment were used as reference control. PCR products obtained after viral (Ad TC) infections were sequenced. The positions of mutations due to DNA integration were detected through Sanger s sequencing.
3 B PC3 BP + Ad TC PC3 CMV + Ad TC DU145 BP + Ad TC DU145 CMV + Ad TC Cas9 D10A RFP 10 µm 10 µm 10 µm 10 µm EGFP tk 10 µm 10 µm 10 µm 10 µm Supplemental figure 2(B) Expression of Cas9 D10A and HSV1 tk in PC3 or DU145 tumors that contain TMEM135 CCDC67 breakpoint (PC3 BP and DU145 BP, respectively) and their control counterparts (PC3 CMV and DU145 CMV). The xenografted tumors were treated with pad5 Cas9 D10A grna TMEM135int13 grna CCDC67int9 and pad TMEM135 int13 EGFP tk CCDC67 int9 (Ad TC). Green arrows indicate unstained mouse stromal cells. Selected images were shown.
4 C HUH7 + Ad MF + G HEP3B + Ad MF + G HUH7 + Ad MF + PBS HUH7 + Ad TC + G Cas9 D10A RFP 10 µm 10 µm 10 µm 10 µm EGFP tk 10 µm 10 µm 10 µm 10 µm Supplemental figure 2C. Expression of Cas9 D10A and HSV1-tk in HUH7 or HEP3B tumors treated with Ad-TC or pad5-cas9 D10A -grna MAN2A1int13 -grna FERint14 /pad-man2a1 int13 -EGFP-tk-FER int14 (Ad-MF). Green arrows indicate unstained mouse stromal cells. Selected images were shown.
5 D Tumor: PC3 CMV DU145 CMV HUH7 Virus: Ad TC Ad TC Ad TC 10 µm 10 µm 10 µm Tumor: PC3 BP DU145 BP HUH7 Virus: Ad TC Ad TC Ad MF 10 µm 10 µm 10 µm Supplemental figure 2D. Genome therapy induced apoptosis of xenografted cancers that contain fusion gene breakpoints. Terminal deoxynucleotidyl transferase (TdT) dutp Nick-End Labeling (TUNEL) assays were performed on the PC3 BP, DU145 BP, PC3 CMV, DU145 CMV, or HUH7 xenografted cancers treated with either Ad-TC or Ad-MF.
6 Supplemental figure 3 A B TMEM135 CCDC67 GBP 1600 bp TMEM135 CCDC67 BP 500 bp MAN2A1 FER FER MAN2A1 FER 100 bp 113 Kd Genome Β actin PC3 BP DU145 BP PC3 CMV DU145 CMV 200 bp Β actin PC3 BP DU145 BP PC3 CMV DU145 CMV 100 bp Β actin HUH7 HEP3B 100 bp GAPDH HUH7 HEP3B 33 Kd Supplemental figure 3: Full view of figures 2B and 5B. (A) Full view of figure 2B. (B) Full view of figure 5B.
7 Supplemental Table 1. Chromosome breakpoint dependent cancer cell killing by Ganciclovir Samples Treatment % Apoptosis PC3 BP Ad-TC+Gan* 20.6%+1.4 PC3 CMV Ad-TC+Gan* 1.5%+0.2 DU145 BP Ad-TC+Gan* 19.8%+0.7 DU145 CMV Ad-TC+Gan* 1.2%+1.1 HUH7 Ad-MF+Gan** 26.9%+1.3 HUH7 Ad-TC+Gan* 2.6%+0.41 HEP3B Ad-MF-Gan** 2.8%+0.16 *- Treatment of Adeno+Gan includes AD5-Cas9 D10A -grna TMEM135int13 -grna CCDC67int9 and AD-TMEM135 int13 - EGFP-tk-CCDC67 int9 at 10 multiplicity of infection and Ganciclovir at 10µg/ml. **- Treatment of Adeno+Gan includes AD5-Cas9 D10A -grna MAN2A1int13 -grna FERint14 and AD-MAN2A1 int13 - EGFP-tk-FER int14 at 10 multiplicity of infection and Ganciclovir at 10µg/ml. BP=pCMV-TMEM135 int13 -CCDC67 int9. CMV=pCMVscript 1
8 Supplemental table 2. Primer sequences for PCR and RT-PCR Samples Forward primer/reverse primer Genome BP PCR GCCCATATATGGAGTTCCGCG/TCTGGCAAGCTATCTAACCCC RNA BP RT-PCR Genome -actin PCR RNA -actin RT-PCR Pre-integration 5 end Pre-integration 3 end CMV-EGFP PCR HSV1-tk-CMV PCR AGCACAGAGACCCAGAAGGTC/AGGAGGAGGAGGAGGAGAAAG TCTTTGCACTTTCTGCATGTCCCC/GTCCATCACGATGCCAGTGGTAC ATGATGATATCGCCGCGCTC/CACGATGGAGGGGAAGACG GCCCATATATGGAGTTCCGCG/AGGCAAAGAGCTCAGTGAGTG TGCCTCATTGGTAATGTTAGCTC/GGCGAATTGGGTACACTTACC ACTCACGGGGATTTCCAAGTC/AAGTCGTGCTGCTTCATGTGG TGTTCTAGCCAAGAGGCTGAG/GGCGAATTGGGTACACTTACC 2
9 Supplemental table 3 On- and off-target sequences Fusion gene grna target Sequences Chr. Position** TMEM135-CCDC67 On target CACTCACTGAGCTCTTTGCC Human Off-target 1 CACTGACTGAGCTCTCTGAC Human Off-target 2 CACTCACTGTCCTCTTTGCC Human Off-target 3 AACTCAGCGAGCTCTTTGCC Human Off-target 4 CACTCACTGAGATCTGTGCC Human Off-target 5 GACTCACTGAACTCTTTGGC Human Off-target 6 CCCTGAATGAGCTCTTTGCC Mouse Off-target 1 CACTGACTCAGTTCTTTGCC Mouse Off-target 2 CACTCCATCAGCTCTTTGCC Mouse Off-target 3 CACTCACTGGCCCCTTTGCC Mouse Off-target 4 TACCCACTGAGCTCTTTCCC Mouse Off-target 5 CACTCACTGAGCACTGTGTC Mouse Off-target 6 CACTGACTGAGTCCTTTGCC MAN2A1-FER On target TAGCATTAAGGGCCCCCTAA Human Off-target 1 TAGCACTGAAGGCCCCCTAA Human Off-target 2 TAGCATTAAGGGCCCACTTG Human Off-target 3 TAGCACTGAGGGCCCCCAAA Human Off-target 4 TAGTATTCAGGGCCCACTAA Human Off-target 5 TGGGATTAGGGGCCCCCTAA Mouse Off-target 1 TAGCTTTAAGTGCCTCCTAA Mouse Off-target 2 TACCATTAAGTGCCCCCAAA Mouse Off-target 3 TGGCATTAAGGGCCCATTAA Mouse Off-target 4 TTGCATTCAGGGTCCCCTAA Mouse Off-target 5 TAGCATTAAGTGCCCTCTTA ** Alignment to GRCh38.p7 primary assembly database for human genome or to GRCm38.p4 C57BL/6J for mouse genome. Chr Chromosome. 3
10 Supplemental table 4 On- and off-target sequencing primers for Illumina HiSeq2500 Fusion gene Genome sequencing primer Sequence Chr Position** TMEM135-CCDC67 On target CCCTGTTTTACATATGAGGAAAC Human Off-target 1 CCCACAAAAAGGGTACATGCC Human Off-target 2 TTTGGATTCAGCACAGTGGCC Human Off-target 3 AGGAGGACCATGCCATTTCCC Human Off-target 4 AGAGGCTCCAGACGCATTGTG Human Off-target 5a TCAGTGCCCTGCTCAAAACAG Human Off-target 5b CTCAGTCAGTATCCCTGCCAG Human Off-target 6a TGGGAAGGAATTGGAGGGAAG Human Off-target 6b GCTGTCATCTACAGCATCCTG Mouse Off-target 1 CAGGGAAGCTGCTTTAGAATG Mouse Off-target 2a TGAAATCCCTGACCCCCAATC Mouse Off-target 2b CAGACTGGACAAGGTGCTGCG Mouse Off-target 3a CACAGAGTATGCTAGGGGAAG Mouse Off-target 3b GTGAATGCCCTCTCTCTTTGC Mouse Off-target 4 AGGGTATTGTGGAGGTCACAG Mouse Off-target 5 CTTACTACTTAAGCCGCCCAG Mouse Off-target 6 ATCATCTAAGGGGAGTCTTGG EGFP-tk CAGCTCCTCGCCCTTGCTCAC N/A N/A MAN2A1-FER On target GACAGTCTGGCAGAGTTATGC Human Off-target 1a GAGCACGGGAGGCAAATAAAC Human Off-target 1b GTGAAGGGCACACTCTTCCAG Human Off-target 2 GGAGGAGGTATTGGAGGGTTG Human Off-target 3 AAGGCATCACTCACCACACTG Human Off-target 4a TCTATGATGTAGCCTCAGCTC Human Off-target 4b GTGGCATTTGCTTACCCTGGA Human Off-target 5a ATGGGGCTGTATGTGAAGAGG Human Off-target 5b CTGCATCTTCACAGGGTCGTC Mouse Off-target 1a AGAGCTTTCAGGTGTGCTGTC Mouse Off-target 1b TTGCCCACCTGCGACTAGAGA Mouse Off-target 2a TGGGTTGATCTAGCTGATGGAG Mouse Off-target 2b CTCAACCAACCTCTAATACTGTAC Mouse Off-target 3a GTCTGCTCCCTAAACGAGATG Mouse Off-target 3b CCAAACCAAAGAGTGGTGGAG Mouse Off-target 4a AAGGGATGCTACAGCCTGTCC Mouse Off-target 4b TGGGCACGGAGATAGGTTGTC Mouse Off-target 5a GCTTCTGTTAGGGCTTTCATGC Mouse Off-target 5b CACAGCATAGCCAAACTTATGG EGFP-tk CAGCTCCTCGCCCTTGCTCAC N/A N/A TMEM135-CCDC67 BP Primer 1 GCCTCATTGGTAATGTTAGCTC Primer 2 AGCATGGCACACTCAGTGAAC MAN2A1-FER BP Primer 1 CTCCTGACCCCGTGATCCACCT Primer 2 AAACCATAATCATGCTGACTG ** Alignment to GRCh38.p7 primary assembly database for human genome or to GRCm38.p4 C57BL/6J for mouse genome. Chr chromosome. N/A Not applicable; Chr Chromosome. 4
11 Supplemental table 5 Quantification of on and off targets reads of TMEM135 CCDC67 and MAN2A1 FER genome therapy in vitro and in vivo Therapy target Target cells total reads On target Off targets Off/On TMEM135 CCDC67 DU145 BP in vitro <0.1% PC3 BP in vitro <0.1% DU145 BP tumor <0.1% PC3 BP tumor <0.1% MAN2A1 FER HUH7 in vitro % HUH7 tumor % DU145 BP DU145 cell line harboring TMEM135 CCDC67 breakpoint. PC3 BP PC3 cell line harboring TMEM135 CCDC67 breakpoint. DU145 BP in vitro DU145 BP cell culture treated with Ad TC. PC3 BP in vitro PC3 BP cell culture treated with Ad TC. DU145 BP tumor DU145 BP xenografted tumor treated with Ad TC. PC3 BP tumor PC3 BP xenografted tumor treated with Ad TC. HUH7 in vitro HUH7 cell culture treated with Ad MF. HUH7 tumor HUH7 xenografted tumor treated with Ad MF. On target Pair end reads mapped to the correct on target genome sequence at one end and EGFP tk sequence at the paired end. Off target Pair end reads mapped to the off target genome sequence at one end and EGFP tk sequence at the paired end. Off/On: % Off target rate. Total reads Total number of reads including unmapped and mapped reads. 5
12 Supplemental table 6 Quantification of genomic EPGP tk reads in cells without fusion gene breakpoint Therapy viruses Target cells Total reads gegfp tk reads Mapped reads % Off Ad TC DU145 CMV in vitro % PC3 CMV in vitro % DU145 CMV tumor <0.1% PC3 CMV tumor <0.1% mliver (DU145 BP) <0.1% mliver (PC3 BP) <0.1% Ad MF HEP3B in vitro <0.1% HEP3B tumor % mliver (HUH7) <0.1% Ad TC AD5 Cas9 D10A grna TMEM135int13 grna CCDC67int9 and AD TMEM135 int13 EGFP tk CCDC67 int9. Ad MF AD5 Cas9 D10A grna MAN2A1int13 grna FERint14 and AD MAN2A1 int13 EGFP tk FER int14. gegfp tk reads Pair end reads mapped to correct on target or off target genome sequence at one end and EGFP tk sequence at the paired end. Mapped reads Pair end reads mapped to the correct genome sequence adjacent to the sequencing primers. Total reads Total number of reads including unmapped and mapped reads. mliver mouse liver cells from animals xenografted with cancer cells, and treated with recombinant adenoviruses. % Off gegfp tk/mapped reads DU145 CMV DU145 cells harboring pcmvscript. PC3 CMV PC3 cells harboring pcmvscript. DU145 CMV in vitro DU145 CMV cell culture treated with Ad TC. PC3 CMV in vitro PC3 CMV cell culture treated with Ad TC. DU145 CMV tumor DU145 CMV xenografted tumor treated with Ad TC. PC3 CMV tumor PC3 CMV xenografted tumor treated with Ad TC. mliver (DU145 BP) mliver from mice xenografted with DU145 BP cells and treated with Ad TC. mliver PC3 BP) mliver from mice xenografted with PC3 BP cells and treated with Ad TC. HEP3B in vitro HEP3B cell culture treated with Ad MF HEP3B tumor HEP3B xenografted tumor treated with Ad MF. mliver (HUH7) mliver from mice xenografted with HUH7 cells and treated with Ad MF. 6
13 Supplemental table 7 Integration rates of EGFP tk in genome therapy in vitro and in vivo Therapy target Samples genome EGFP tk reads BP reads Integration rates TMEM135 CCDC67 DU145 BP in vitro % DU145 BP tumor % PC3 BP in vitro % PC3 BP tumor % MAN2A FER HUH7 in vitro % HUH7 tumor % DU145 BP in vitro DU145 BP cell culture treated with Ad TC. DU145 BP tumor DU145 BP xenografted tumor treated with Ad TC. PC3 BP in vitro PC3 BP cell culture treated with Ad TC. PC3 BP tumor PC3 BP xenografted tumor treated with Ad TC. HUH7 in vitro HUH7 cell culture treated with Ad MF. HUH7 tumor HUH7 xenografted tumor treated with Ad MF. genome EGFP tk reads Pair end reads mapped to the correct on target genome sequence at one end and EGFP tk sequence at the paired end. BP reads Pair end reads mapped to the left side of the chromosome breakpoint at one end and the right side of the chromosome breakpoint at the paired end, or any mapped read containing the chromosomal breakpoint. Integration rate genome EGFP tk reads/bp reads 7
Inhibition of HSV-1 Replication by Gene Editing Strategy. Running Title: Targeting HSV-1 Infection by CRISPR/Cas9
 SUPPLEMENTARY MATERIALS 2/4/16 Inhibition of HSV-1 Replication by Gene Editing Strategy Running Title: Targeting HSV-1 Infection by CRISPR/Cas9 Pamela C. Roehm 1,2,3*, Masoud Shekarabi 1, Hassen Wollebo
SUPPLEMENTARY MATERIALS 2/4/16 Inhibition of HSV-1 Replication by Gene Editing Strategy Running Title: Targeting HSV-1 Infection by CRISPR/Cas9 Pamela C. Roehm 1,2,3*, Masoud Shekarabi 1, Hassen Wollebo
Fig. S1. eif6 expression in HEK293 transfected with shrna against eif6 or pcmv-eif6 vector.
 Fig. S1. eif6 expression in HEK293 transfected with shrna against eif6 or pcmv-eif6 vector. (a) Western blotting analysis and (b) qpcr analysis of eif6 expression in HEK293 T cells transfected with either
Fig. S1. eif6 expression in HEK293 transfected with shrna against eif6 or pcmv-eif6 vector. (a) Western blotting analysis and (b) qpcr analysis of eif6 expression in HEK293 T cells transfected with either
Rassf1a -/- Sav1 +/- mice. Supplementary Figures:
 1 Supplementary Figures: Figure S1: Genotyping of Rassf1a and Sav1 mutant mice. A. Rassf1a knockout mice lack exon 1 of the Rassf1 gene. A diagram and typical PCR genotyping of wildtype and Rassf1a -/-
1 Supplementary Figures: Figure S1: Genotyping of Rassf1a and Sav1 mutant mice. A. Rassf1a knockout mice lack exon 1 of the Rassf1 gene. A diagram and typical PCR genotyping of wildtype and Rassf1a -/-
Nature Biotechnology: doi: /nbt Supplementary Figure 1. In vitro validation of OTC sgrnas and donor template.
 Supplementary Figure 1 In vitro validation of OTC sgrnas and donor template. (a) In vitro validation of sgrnas targeted to OTC in the MC57G mouse cell line by transient transfection followed by 4-day puromycin
Supplementary Figure 1 In vitro validation of OTC sgrnas and donor template. (a) In vitro validation of sgrnas targeted to OTC in the MC57G mouse cell line by transient transfection followed by 4-day puromycin
Supplemental Data. Na Xu et al. (2016). Plant Cell /tpc
 Supplemental Figure 1. The weak fluorescence phenotype is not caused by the mutation in At3g60240. (A) A mutation mapped to the gene At3g60240. Map-based cloning strategy was used to map the mutated site
Supplemental Figure 1. The weak fluorescence phenotype is not caused by the mutation in At3g60240. (A) A mutation mapped to the gene At3g60240. Map-based cloning strategy was used to map the mutated site
Figure S1. Unrearranged locus. Rearranged locus. Concordant read pairs. Region1. Region2. Cluster of discordant read pairs, bundle
 Figure S1 a Unrearranged locus Rearranged locus Concordant read pairs Region1 Concordant read pairs Cluster of discordant read pairs, bundle Region2 Concordant read pairs b Physical coverage 5 4 3 2 1
Figure S1 a Unrearranged locus Rearranged locus Concordant read pairs Region1 Concordant read pairs Cluster of discordant read pairs, bundle Region2 Concordant read pairs b Physical coverage 5 4 3 2 1
Nature Biotechnology: doi: /nbt Supplementary Figure 1
 Supplementary Figure 1 Generation of the AARE-Gene system construct. (a) Position and sequence alignment of AAREs extracted from human Trb3, Chop or Atf3 promoters. AARE core sequences are boxed in grey.
Supplementary Figure 1 Generation of the AARE-Gene system construct. (a) Position and sequence alignment of AAREs extracted from human Trb3, Chop or Atf3 promoters. AARE core sequences are boxed in grey.
CELLTECHGEN For Research Only. Construction of sgrna expression vector for Lenti-virus system (Example: Lenti-U6 sgrna-ef1 -Puro vector)
 Construction of sgrna expression vector for Lenti-virus system (Example: Lenti-U6 sgrna-ef1 -Puro vector) Catalog number: CTG-CAS9-11 Introduction The vector Lenti-U6 sgrna-ef1 -Puro is designed for expression
Construction of sgrna expression vector for Lenti-virus system (Example: Lenti-U6 sgrna-ef1 -Puro vector) Catalog number: CTG-CAS9-11 Introduction The vector Lenti-U6 sgrna-ef1 -Puro is designed for expression
CELLTECHGEN For Research Only. Construction of all-in-one vector for Lenti-virus system (Example: Lenti-EF1 -Cas9-EGFP-U6 sgrna vector)
 Construction of all-in-one vector for Lenti-virus system (Example: Lenti-EF1 -Cas9-EGFP-U6 sgrna vector) Catalog number: CTG-CAS9-18 Introduction The vector Lenti-EF1 -Cas9-EGFP-U6 sgrna is designed for
Construction of all-in-one vector for Lenti-virus system (Example: Lenti-EF1 -Cas9-EGFP-U6 sgrna vector) Catalog number: CTG-CAS9-18 Introduction The vector Lenti-EF1 -Cas9-EGFP-U6 sgrna is designed for
Supplementary Figure 1 Activities of ABEs using extended sgrnas in HEK293T cells.
 Supplementary Figure 1 Activities of ABEs using extended sgrnas in HEK293T cells. Base editing efficiencies of ABEs with extended sgrnas at Site 18 (a), Site 19 (b), the HBB-E2 site (c), and the HBB-E3
Supplementary Figure 1 Activities of ABEs using extended sgrnas in HEK293T cells. Base editing efficiencies of ABEs with extended sgrnas at Site 18 (a), Site 19 (b), the HBB-E2 site (c), and the HBB-E3
Genome Sequence Assembly
 Genome Sequence Assembly Learning Goals: Introduce the field of bioinformatics Familiarize the student with performing sequence alignments Understand the assembly process in genome sequencing Introduction:
Genome Sequence Assembly Learning Goals: Introduce the field of bioinformatics Familiarize the student with performing sequence alignments Understand the assembly process in genome sequencing Introduction:
Supplemental Data. Cui et al. (2012). Plant Cell /tpc a b c d. Stem UBC32 ACTIN
 A Root Stem Leaf Flower Silique Senescence leaf B a b c d UBC32 ACTIN C * Supplemental Figure 1. Expression Pattern and Protein Sequence of UBC32 Homologues in Yeast, Human, and Arabidopsis. (A) Expression
A Root Stem Leaf Flower Silique Senescence leaf B a b c d UBC32 ACTIN C * Supplemental Figure 1. Expression Pattern and Protein Sequence of UBC32 Homologues in Yeast, Human, and Arabidopsis. (A) Expression
Fig. S1. Effect of p120-catenin overexpression on the interaction of SCUBE2 with E-cadherin. The expression plasmid encoding FLAG.
 Fig. S1. Effect of p120-catenin overexpression on the interaction of SCUBE2 with E-cadherin. The expression plasmid encoding FLAG.SCUBE2, E-cadherin.Myc, or HA.p120-catenin was transfected in a combination
Fig. S1. Effect of p120-catenin overexpression on the interaction of SCUBE2 with E-cadherin. The expression plasmid encoding FLAG.SCUBE2, E-cadherin.Myc, or HA.p120-catenin was transfected in a combination
Supporting Online Material for
 www.sciencemag.org/cgi/content/full/317/5837/477/dc1 Supporting Online Material for AAV Vector Integration Sites in Mouse Hepatocellular Carcinoma Anthony Donsante, Daniel G. Miller, Yi Li, Carole Vogler,
www.sciencemag.org/cgi/content/full/317/5837/477/dc1 Supporting Online Material for AAV Vector Integration Sites in Mouse Hepatocellular Carcinoma Anthony Donsante, Daniel G. Miller, Yi Li, Carole Vogler,
Supplementary Figure 1
 Supplementary Figure 1 Supplementary Fig. 1 shrna mediated knockdown of ZRSR2 in K562 and 293T cells. (a) ZRSR2 transcript levels in stably transduced K562 cells were determined using qrt-pcr. GAPDH was
Supplementary Figure 1 Supplementary Fig. 1 shrna mediated knockdown of ZRSR2 in K562 and 293T cells. (a) ZRSR2 transcript levels in stably transduced K562 cells were determined using qrt-pcr. GAPDH was
Supplementary Figures
 Supplementary Figures Supplementary Figure 1 Experimental schema for the identification of circular RNAs in six normal tissues and seven cancerous tissues. Supplementary Fiure 2 Comparison of human circrnas
Supplementary Figures Supplementary Figure 1 Experimental schema for the identification of circular RNAs in six normal tissues and seven cancerous tissues. Supplementary Fiure 2 Comparison of human circrnas
Student Learning Outcomes (SLOS)
 Student Learning Outcomes (SLOS) KNOWLEDGE AND LEARNING SKILLS USE OF KNOWLEDGE AND LEARNING SKILLS - how to use Annhyb to save and manage sequences - how to use BLAST to compare sequences - how to get
Student Learning Outcomes (SLOS) KNOWLEDGE AND LEARNING SKILLS USE OF KNOWLEDGE AND LEARNING SKILLS - how to use Annhyb to save and manage sequences - how to use BLAST to compare sequences - how to get
8Br-cAMP was purchased from Sigma (St. Louis, MO). Silencer Negative Control sirna #1 and
 1 Supplemental information 2 3 Materials and Methods 4 Reagents and animals 5 8Br-cAMP was purchased from Sigma (St. Louis, MO). Silencer Negative Control sirna #1 and 6 Silencer Select Pre-designed sirna
1 Supplemental information 2 3 Materials and Methods 4 Reagents and animals 5 8Br-cAMP was purchased from Sigma (St. Louis, MO). Silencer Negative Control sirna #1 and 6 Silencer Select Pre-designed sirna
Cell proliferation was measured with Cell Counting Kit-8 (Dojindo Laboratories, Kumamoto, Japan).
 1 2 3 4 5 6 7 8 Supplemental Materials and Methods Cell proliferation assay Cell proliferation was measured with Cell Counting Kit-8 (Dojindo Laboratories, Kumamoto, Japan). GCs were plated at 96-well
1 2 3 4 5 6 7 8 Supplemental Materials and Methods Cell proliferation assay Cell proliferation was measured with Cell Counting Kit-8 (Dojindo Laboratories, Kumamoto, Japan). GCs were plated at 96-well
SUPPLEMENTAL MATERIAL
 SUPPLEMENTAL MATERIAL MATERIALS AND METHODS Generation of TSPOΔ/Δ murine embryonic fibroblasts Embryos were harvested from 13.5-day pregnant TSPOfl/fl mice. After dissection to eviscerate and remove the
SUPPLEMENTAL MATERIAL MATERIALS AND METHODS Generation of TSPOΔ/Δ murine embryonic fibroblasts Embryos were harvested from 13.5-day pregnant TSPOfl/fl mice. After dissection to eviscerate and remove the
PrimePCR Assay Validation Report
 Gene Information Gene Name Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID protein kinase N1 PKN1 Human The protein encoded by this gene belongs to the protein
Gene Information Gene Name Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID protein kinase N1 PKN1 Human The protein encoded by this gene belongs to the protein
Full-length single-cell RNA-seq applied to a viral human. cancer: Applications to HPV expression and splicing analysis. Supplementary Information
 Full-length single-cell RNA-seq applied to a viral human cancer: Applications to HPV expression and splicing analysis in HeLa S3 cells Liang Wu 1, Xiaolong Zhang 1,2, Zhikun Zhao 1,3,4, Ling Wang 5, Bo
Full-length single-cell RNA-seq applied to a viral human cancer: Applications to HPV expression and splicing analysis in HeLa S3 cells Liang Wu 1, Xiaolong Zhang 1,2, Zhikun Zhao 1,3,4, Ling Wang 5, Bo
PrimePCR Assay Validation Report
 Gene Information Gene Name laminin, beta 3 Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID LAMB3 Human The product encoded by this gene is a laminin that
Gene Information Gene Name laminin, beta 3 Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID LAMB3 Human The product encoded by this gene is a laminin that
OmicsLink shrna Clones guaranteed knockdown even in difficult-to-transfect cells
 OmicsLink shrna Clones guaranteed knockdown even in difficult-to-transfect cells OmicsLink shrna clone collections consist of lentiviral, and other mammalian expression vector based small hairpin RNA (shrna)
OmicsLink shrna Clones guaranteed knockdown even in difficult-to-transfect cells OmicsLink shrna clone collections consist of lentiviral, and other mammalian expression vector based small hairpin RNA (shrna)
Supplementary Figures and Tables
 Supplementary Figures and Tables SI Fig. 1. Schematic diagrams of the Gateway-system based T. pseudonana transformation vectors. A-C, for expression and in vivo localization of egfp-tagged SIT genes under
Supplementary Figures and Tables SI Fig. 1. Schematic diagrams of the Gateway-system based T. pseudonana transformation vectors. A-C, for expression and in vivo localization of egfp-tagged SIT genes under
A CRISPR/Cas9 Vector System for Tissue-Specific Gene Disruption in Zebrafish
 Developmental Cell Supplemental Information A CRISPR/Cas9 Vector System for Tissue-Specific Gene Disruption in Zebrafish Julien Ablain, Ellen M. Durand, Song Yang, Yi Zhou, and Leonard I. Zon % larvae
Developmental Cell Supplemental Information A CRISPR/Cas9 Vector System for Tissue-Specific Gene Disruption in Zebrafish Julien Ablain, Ellen M. Durand, Song Yang, Yi Zhou, and Leonard I. Zon % larvae
PrimePCR Assay Validation Report
 Gene Information Gene Name transforming growth factor, beta 1 Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID TGFB1 Human This gene encodes a member of the
Gene Information Gene Name transforming growth factor, beta 1 Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID TGFB1 Human This gene encodes a member of the
Supplementary Fig. 1 related to Fig. 1 Clinical relevance of lncrna candidate
 Supplementary Figure Legends Supplementary Fig. 1 related to Fig. 1 Clinical relevance of lncrna candidate BC041951 in gastric cancer. (A) The flow chart for selected candidate lncrnas in 660 up-regulated
Supplementary Figure Legends Supplementary Fig. 1 related to Fig. 1 Clinical relevance of lncrna candidate BC041951 in gastric cancer. (A) The flow chart for selected candidate lncrnas in 660 up-regulated
Table S1. Primers used in the study
 Table S1. Primers used in the study Primer name Application Sequence I1F16 Genotyping GGCAAGTGAGTGAGTGCCTA I1R11 Genotyping CCCACTCGTATTGACGCTCT V19 Genotyping GGGTCTCAAAGTCAGGGTCA D18Mit184-F Genotyping
Table S1. Primers used in the study Primer name Application Sequence I1F16 Genotyping GGCAAGTGAGTGAGTGCCTA I1R11 Genotyping CCCACTCGTATTGACGCTCT V19 Genotyping GGGTCTCAAAGTCAGGGTCA D18Mit184-F Genotyping
PrimePCR Assay Validation Report
 Gene Information Gene Name Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID Glyceraldehyde-3-phosphate dehydrogenase Gapdh Rat This gene encodes a member of
Gene Information Gene Name Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID Glyceraldehyde-3-phosphate dehydrogenase Gapdh Rat This gene encodes a member of
PrimePCR Assay Validation Report
 Gene Information Gene Name Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID mcf.2 transforming sequence-like Mcf2l Mouse Description Not Available C130040G20Rik,
Gene Information Gene Name Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID mcf.2 transforming sequence-like Mcf2l Mouse Description Not Available C130040G20Rik,
PrimePCR Assay Validation Report
 Gene Information Gene Name Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID protein phosphatase, Mg2+/Mn2+ dependent, 1A PPM1A Human The protein encoded by
Gene Information Gene Name Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID protein phosphatase, Mg2+/Mn2+ dependent, 1A PPM1A Human The protein encoded by
Contact Person: Fax: IACUC Protocol Title: Address: Department: Yes. Primers Name: Concentration: Fragment Size:
 Transgenic Mouse Request Form Please attach a gel photo and a linear map of the construct to this form. The map should indicate locations of: Promoter/ enhancer, splice site, poly A site, and CAP site.
Transgenic Mouse Request Form Please attach a gel photo and a linear map of the construct to this form. The map should indicate locations of: Promoter/ enhancer, splice site, poly A site, and CAP site.
4. Analysing genes II Isolate mutants*
 .. 4. Analysing s II Isolate mutants* Using the mutant to isolate the classify mutants by complementation analysis wild type study phenotype of mutants mutant 1 - use mutant to isolate sequence put individual
.. 4. Analysing s II Isolate mutants* Using the mutant to isolate the classify mutants by complementation analysis wild type study phenotype of mutants mutant 1 - use mutant to isolate sequence put individual
NGS to address ncrna and viruses
 NGS to address ncrna and viruses Introduction & TRON Next generation sequencing transcriptomics ncrnas vrna June 30, 2010 John Castle Institute for Translational Oncology and Immunology (TRON) Mainz, Germany
NGS to address ncrna and viruses Introduction & TRON Next generation sequencing transcriptomics ncrnas vrna June 30, 2010 John Castle Institute for Translational Oncology and Immunology (TRON) Mainz, Germany
Nature Biotechnology doi: /nbt Supplementary Figure 1. Substrate-linked protein evolution components.
 Supplementary Figure 1 Substrate-linked protein evolution components. (a) loxbtr subsites used for the combinational directed-evolution process to generate Brec1. Sequence differences (compared to the
Supplementary Figure 1 Substrate-linked protein evolution components. (a) loxbtr subsites used for the combinational directed-evolution process to generate Brec1. Sequence differences (compared to the
SUPPLEMENTARY INFORMATION
 AS-NMD modulates FLM-dependent thermosensory flowering response in Arabidopsis NATURE PLANTS www.nature.com/natureplants 1 Supplementary Figure 1. Genomic sequence of FLM along with the splice sites. Sequencing
AS-NMD modulates FLM-dependent thermosensory flowering response in Arabidopsis NATURE PLANTS www.nature.com/natureplants 1 Supplementary Figure 1. Genomic sequence of FLM along with the splice sites. Sequencing
Figure S1. Figure S2. Figure S3 HB Anti-FSP27 (COOH-terminal peptide) Ab. Anti-GST-FSP27(45-127) Ab.
 / 36B4 mrna ratio Figure S1 * 2. 1.6 1.2.8 *.4 control TNFα BRL49653 Figure S2 Su bw AT p iw Anti- (COOH-terminal peptide) Ab Blot : Anti-GST-(45-127) Ab β-actin Figure S3 HB2 HW AT BA T Figure S4 A TAG
/ 36B4 mrna ratio Figure S1 * 2. 1.6 1.2.8 *.4 control TNFα BRL49653 Figure S2 Su bw AT p iw Anti- (COOH-terminal peptide) Ab Blot : Anti-GST-(45-127) Ab β-actin Figure S3 HB2 HW AT BA T Figure S4 A TAG
Supplemental Table S1. Experimental details of the qpcr analyses according to the checklist of the MIQE* guidelines.
 Supplemental Table S1, page 1 Supplemental Table S1. Experimental details of the qpcr analyses according to the checklist of the MIQE* guidelines. ITEM TO ERXPERIMENTAL DESIGN Definition of experimental
Supplemental Table S1, page 1 Supplemental Table S1. Experimental details of the qpcr analyses according to the checklist of the MIQE* guidelines. ITEM TO ERXPERIMENTAL DESIGN Definition of experimental
Bi 8 Lecture 5. Ellen Rothenberg 19 January 2016
 Bi 8 Lecture 5 MORE ON HOW WE KNOW WHAT WE KNOW and intro to the protein code Ellen Rothenberg 19 January 2016 SIZE AND PURIFICATION BY SYNTHESIS: BASIS OF EARLY SEQUENCING complex mixture of aborted DNA
Bi 8 Lecture 5 MORE ON HOW WE KNOW WHAT WE KNOW and intro to the protein code Ellen Rothenberg 19 January 2016 SIZE AND PURIFICATION BY SYNTHESIS: BASIS OF EARLY SEQUENCING complex mixture of aborted DNA
Introducing new DNA into the genome requires cloning the donor sequence, delivery of the cloned DNA into the cell, and integration into the genome.
 Key Terms Chapter 32: Genetic Engineering Cloning describes propagation of a DNA sequence by incorporating it into a hybrid construct that can be replicated in a host cell. A cloning vector is a plasmid
Key Terms Chapter 32: Genetic Engineering Cloning describes propagation of a DNA sequence by incorporating it into a hybrid construct that can be replicated in a host cell. A cloning vector is a plasmid
H3K36me3 polyclonal antibody
 H3K36me3 polyclonal antibody Cat. No. C15410192 Type: Polyclonal ChIP-grade/ChIP-seq grade Source: Rabbit Lot #: A1845P Size: 50 µg/32 µl Concentration: 1.6 μg/μl Specificity: Human, mouse, Arabidopsis,
H3K36me3 polyclonal antibody Cat. No. C15410192 Type: Polyclonal ChIP-grade/ChIP-seq grade Source: Rabbit Lot #: A1845P Size: 50 µg/32 µl Concentration: 1.6 μg/μl Specificity: Human, mouse, Arabidopsis,
Ad5 genome. Before filtering. After filtering. Suppl. File 4 PAGE mated reads. Ad5 DNA insertion site 5' 3' 293 unmated reads
 Before filtering 0 20 40 60 80 293 Ad5 DNA insertion site 5' 3' 0 20 60 100 48 000 000 48 100 000 48 200 000 48 300 000 48 400 000 48 500 000 After filtering 48 000 000 48 100 000 48 200 000 48 300 000
Before filtering 0 20 40 60 80 293 Ad5 DNA insertion site 5' 3' 0 20 60 100 48 000 000 48 100 000 48 200 000 48 300 000 48 400 000 48 500 000 After filtering 48 000 000 48 100 000 48 200 000 48 300 000
Construction of plant complementation vector and generation of transgenic plants
 MATERIAL S AND METHODS Plant materials and growth conditions Arabidopsis ecotype Columbia (Col0) was used for this study. SALK_072009, SALK_076309, and SALK_027645 were obtained from the Arabidopsis Biological
MATERIAL S AND METHODS Plant materials and growth conditions Arabidopsis ecotype Columbia (Col0) was used for this study. SALK_072009, SALK_076309, and SALK_027645 were obtained from the Arabidopsis Biological
Supplementary Figure 1. Quantitative RT-PCR experimental validation of CRISPR/Cas9 and sgrnas expression in HEK293A transfected cells.
 Supplementary Figure 1. Quantitative RT-PCR experimental validation of CRISPR/Cas9 and sgrnas expression in HEK293A transfected cells. HEK293A cells were transfected with the indicated combinations of
Supplementary Figure 1. Quantitative RT-PCR experimental validation of CRISPR/Cas9 and sgrnas expression in HEK293A transfected cells. HEK293A cells were transfected with the indicated combinations of
GC GC - AAV2/8-4E10
 SUPPLEMENTARY INFORMATION doi:.38/nature660 a Luciferase Expression (Photons/Sec) c f 12 11 11 GC - AAV2/8-Luciferase 9 GC - AAV2/8-Luciferase 0 8 16 24 32 40 48 56 64 Weeks Post-AAV Injection Human IgG
SUPPLEMENTARY INFORMATION doi:.38/nature660 a Luciferase Expression (Photons/Sec) c f 12 11 11 GC - AAV2/8-Luciferase 9 GC - AAV2/8-Luciferase 0 8 16 24 32 40 48 56 64 Weeks Post-AAV Injection Human IgG
Utility of the dual-specificity protein kinase TTK as a therapeutic target
 Utility of the dual-specificity protein kinase TTK as a therapeutic target for intrahepatic spread of liver cancer Ruoyu Miao, 1,2* Yan Wu, 2* Haohai Zhang, 1 Huandi Zhou, 3 Xiaofeng Sun, 2 Eva Csizmadia,
Utility of the dual-specificity protein kinase TTK as a therapeutic target for intrahepatic spread of liver cancer Ruoyu Miao, 1,2* Yan Wu, 2* Haohai Zhang, 1 Huandi Zhou, 3 Xiaofeng Sun, 2 Eva Csizmadia,
Supplementary Information
 Supplementary Information MLL histone methylases regulate expression of HDLR- in presence of estrogen and control plasma cholesterol in vivo Khairul I. Ansari 1, Sahba Kasiri 1, Imran Hussain 1, Samara
Supplementary Information MLL histone methylases regulate expression of HDLR- in presence of estrogen and control plasma cholesterol in vivo Khairul I. Ansari 1, Sahba Kasiri 1, Imran Hussain 1, Samara
FIG S1: Calibration curves of standards for HPLC detection of Dopachrome (Dopac), Dopamine (DA) and Homovanillic acid (HVA), showing area of peak vs
 FIG S1: Calibration curves of standards for HPLC detection of Dopachrome (Dopac), Dopamine (DA) and Homovanillic acid (HVA), showing area of peak vs amount of standard analysed. Each point represents the
FIG S1: Calibration curves of standards for HPLC detection of Dopachrome (Dopac), Dopamine (DA) and Homovanillic acid (HVA), showing area of peak vs amount of standard analysed. Each point represents the
beta-galactosidase (A420) ctrl
 0.6 beta-galactosidase (A420) 0.4 0.2 ctrl 2A 22A 35B 36B 58A 64A 68E 75C 86Fa 86Fb 102F Fig. S1. Levels of induced expression at various attp sites. The induced β-galactosidase activity was measured from
0.6 beta-galactosidase (A420) 0.4 0.2 ctrl 2A 22A 35B 36B 58A 64A 68E 75C 86Fa 86Fb 102F Fig. S1. Levels of induced expression at various attp sites. The induced β-galactosidase activity was measured from
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10163 Supplementary Table 1 Efficiency of vector construction. Process wells recovered efficiency (%) Recombineering* 480 461 96 Intermediate plasmids 461 381 83 Recombineering efficiency
doi:10.1038/nature10163 Supplementary Table 1 Efficiency of vector construction. Process wells recovered efficiency (%) Recombineering* 480 461 96 Intermediate plasmids 461 381 83 Recombineering efficiency
PrimePCR Assay Validation Report
 Gene Information Gene Name Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID caspase 9, apoptosis-related cysteine peptidase CASP9 Human This gene encodes a
Gene Information Gene Name Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID caspase 9, apoptosis-related cysteine peptidase CASP9 Human This gene encodes a
Novel methods for RNA and DNA- Seq analysis using SMART Technology. Andrew Farmer, D. Phil. Vice President, R&D Clontech Laboratories, Inc.
 Novel methods for RNA and DNA- Seq analysis using SMART Technology Andrew Farmer, D. Phil. Vice President, R&D Clontech Laboratories, Inc. Agenda Enabling Single Cell RNA-Seq using SMART Technology SMART
Novel methods for RNA and DNA- Seq analysis using SMART Technology Andrew Farmer, D. Phil. Vice President, R&D Clontech Laboratories, Inc. Agenda Enabling Single Cell RNA-Seq using SMART Technology SMART
CRISPR RNA-guided activation of endogenous human genes
 CRISPR RNA-guided activation of endogenous human genes Morgan L Maeder, Samantha J Linder, Vincent M Cascio, Yanfang Fu, Quan H Ho, J Keith Joung Supplementary Figure 1 Comparison of VEGF activation induced
CRISPR RNA-guided activation of endogenous human genes Morgan L Maeder, Samantha J Linder, Vincent M Cascio, Yanfang Fu, Quan H Ho, J Keith Joung Supplementary Figure 1 Comparison of VEGF activation induced
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb3240 Supplementary Figure 1 GBM cell lines display similar levels of p100 to p52 processing but respond differentially to TWEAK-induced TERT expression according to TERT promoter mutation
DOI: 10.1038/ncb3240 Supplementary Figure 1 GBM cell lines display similar levels of p100 to p52 processing but respond differentially to TWEAK-induced TERT expression according to TERT promoter mutation
Relative Quantification: Data Management & Analysis Settings
 Relative Quantification: Data Management & Analysis Settings Ray Meng, Ph.D. International Field Applications Specialist Gene Expression Division Bio-Rad Laboratories Gene Expression AMPLIFICATION Assay
Relative Quantification: Data Management & Analysis Settings Ray Meng, Ph.D. International Field Applications Specialist Gene Expression Division Bio-Rad Laboratories Gene Expression AMPLIFICATION Assay
Supplementary Figure 1. Nature Structural & Molecular Biology: doi: /nsmb.3494
 Supplementary Figure 1 Pol structure-function analysis (a) Inactivating polymerase and helicase mutations do not alter the stability of Pol. Flag epitopes were introduced using CRISPR/Cas9 gene targeting
Supplementary Figure 1 Pol structure-function analysis (a) Inactivating polymerase and helicase mutations do not alter the stability of Pol. Flag epitopes were introduced using CRISPR/Cas9 gene targeting
Cancer cells that survive radiation therapy acquire HIF-1 activity and translocate toward tumor blood vessels Supplementary Information
 Cancer cells that survive radiation therapy acquire HIF-1 activity and translocate toward tumor blood vessels Supplementary Information 1. Supplementary Figure S1-S10: Pages 2-11 2. Supplementary References:
Cancer cells that survive radiation therapy acquire HIF-1 activity and translocate toward tumor blood vessels Supplementary Information 1. Supplementary Figure S1-S10: Pages 2-11 2. Supplementary References:
Chapter 20 Recombinant DNA Technology. Copyright 2009 Pearson Education, Inc.
 Chapter 20 Recombinant DNA Technology Copyright 2009 Pearson Education, Inc. 20.1 Recombinant DNA Technology Began with Two Key Tools: Restriction Enzymes and DNA Cloning Vectors Recombinant DNA refers
Chapter 20 Recombinant DNA Technology Copyright 2009 Pearson Education, Inc. 20.1 Recombinant DNA Technology Began with Two Key Tools: Restriction Enzymes and DNA Cloning Vectors Recombinant DNA refers
12/31/16. I. Manipulating DNA (9.1) A. Scientists use several techniques to manipulate DNA. 1. DNA is a very large molecule
 I. Manipulating DNA (9.1) A. Scientists use several techniques to manipulate DNA 1. DNA is a very large molecule 3. Led to many biotechnology applications- genetic engineering, DNA fingerprinting, cloning,
I. Manipulating DNA (9.1) A. Scientists use several techniques to manipulate DNA 1. DNA is a very large molecule 3. Led to many biotechnology applications- genetic engineering, DNA fingerprinting, cloning,
Taking Advantage of Long RNA-Seq Reads. Vince Magrini Pacific Biosciences User Group Meeting September 18, 2013
 Taking Advantage of Long RNA-Seq Reads Vince Magrini Pacific Biosciences User Group Meeting September 18, 2013 Overview Proof-of-Principle SMART-cDNA Synthesis PB-SBL size distributions Gene Annotation
Taking Advantage of Long RNA-Seq Reads Vince Magrini Pacific Biosciences User Group Meeting September 18, 2013 Overview Proof-of-Principle SMART-cDNA Synthesis PB-SBL size distributions Gene Annotation
Genetic tests are available for hundreds of disorders. DNA testing can pinpoint the exact genetic basis of a disorder.
 Human DNA Analysis Human DNA Analysis There are roughly 6 billion base pairs in your DNA. Biologists search the human genome using sequences of DNA bases. Genetic tests are available for hundreds of disorders.
Human DNA Analysis Human DNA Analysis There are roughly 6 billion base pairs in your DNA. Biologists search the human genome using sequences of DNA bases. Genetic tests are available for hundreds of disorders.
SUPPLEMENTARY INFORMATION
 6 Relative levels of ma3 RNA 5 4 3 2 1 LN MG SG PG Spl Thy ma3 ß actin Lymph node Mammary gland Prostate gland Salivary gland Spleen Thymus MMTV target tissues Fig. S1: MMTV target tissues express ma3.
6 Relative levels of ma3 RNA 5 4 3 2 1 LN MG SG PG Spl Thy ma3 ß actin Lymph node Mammary gland Prostate gland Salivary gland Spleen Thymus MMTV target tissues Fig. S1: MMTV target tissues express ma3.
INNOVAPI WP3 Activity 3 Health Parameters
 Harmonization of methods for measuring viral load and bio-markers of aging Stage1 : Quality check of extracted RNA in Torino A qpcr on 18S gene showed that the extracted samples also contain genomic DNA.
Harmonization of methods for measuring viral load and bio-markers of aging Stage1 : Quality check of extracted RNA in Torino A qpcr on 18S gene showed that the extracted samples also contain genomic DNA.
PrimePCR Assay Validation Report
 Gene Information Gene Name collagen, type IV, alpha 1 Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID COL4A1 Human This gene encodes the major type IV alpha
Gene Information Gene Name collagen, type IV, alpha 1 Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID COL4A1 Human This gene encodes the major type IV alpha
Roche Molecular Biochemicals Technical Note No. LC 10/2000
 Roche Molecular Biochemicals Technical Note No. LC 10/2000 LightCycler Overview of LightCycler Quantification Methods 1. General Introduction Introduction Content Definitions This Technical Note will introduce
Roche Molecular Biochemicals Technical Note No. LC 10/2000 LightCycler Overview of LightCycler Quantification Methods 1. General Introduction Introduction Content Definitions This Technical Note will introduce
Absolute Mouse Telomere Length Quantification qpcr Assay Kit (AMTLQ) Catalog #M reactions
 Absolute Mouse Telomere Length Quantification qpcr Assay Kit (AMTLQ) Catalog #M8918 100 reactions Product Description Telomeres are repetitive nucleotide elements at the ends of chromosomes that protect
Absolute Mouse Telomere Length Quantification qpcr Assay Kit (AMTLQ) Catalog #M8918 100 reactions Product Description Telomeres are repetitive nucleotide elements at the ends of chromosomes that protect
PrimePCR Assay Validation Report
 Gene Information Gene Name minichromosome maintenance complex component 8 Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID MCM8 Human The protein encoded by
Gene Information Gene Name minichromosome maintenance complex component 8 Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID MCM8 Human The protein encoded by
Revised: RG-RV2 by Fukuhara et al.
 Supplemental Figure 1 The generation of Spns2 conditional knockout mice. (A) Schematic representation of the wild type Spns2 locus (Spns2 + ), the targeted allele, the floxed allele (Spns2 f ) and the
Supplemental Figure 1 The generation of Spns2 conditional knockout mice. (A) Schematic representation of the wild type Spns2 locus (Spns2 + ), the targeted allele, the floxed allele (Spns2 f ) and the
Percent survival. Supplementary fig. S3 A.
 Supplementary fig. S3 A. B. 100 Percent survival 80 60 40 20 Ml 0 0 100 C. Fig. S3 Comparison of leukaemia incidence rate in the triple targeted chimaeric mice and germline-transmission translocator mice
Supplementary fig. S3 A. B. 100 Percent survival 80 60 40 20 Ml 0 0 100 C. Fig. S3 Comparison of leukaemia incidence rate in the triple targeted chimaeric mice and germline-transmission translocator mice
Supplementary Methods
 Supplementary Methods Reverse transcribed Quantitative PCR. Total RNA was isolated from bone marrow derived macrophages using RNeasy Mini Kit (Qiagen), DNase-treated (Promega RQ1), and reverse transcribed
Supplementary Methods Reverse transcribed Quantitative PCR. Total RNA was isolated from bone marrow derived macrophages using RNeasy Mini Kit (Qiagen), DNase-treated (Promega RQ1), and reverse transcribed
Supplemental Materials. Matrix Proteases Contribute to Progression of Pelvic Organ Prolapse in Mice and Humans
 Supplemental Materials Matrix Proteases Contribute to Progression of Pelvic Organ Prolapse in Mice and Humans Madhusudhan Budatha, Shayzreen Roshanravan, Qian Zheng, Cecilia Weislander, Shelby L. Chapman,
Supplemental Materials Matrix Proteases Contribute to Progression of Pelvic Organ Prolapse in Mice and Humans Madhusudhan Budatha, Shayzreen Roshanravan, Qian Zheng, Cecilia Weislander, Shelby L. Chapman,
PrimePCR Assay Validation Report
 Gene Information Gene Name SRY (sex determining region Y)-box 6 Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID SOX6 Human This gene encodes a member of the
Gene Information Gene Name SRY (sex determining region Y)-box 6 Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID SOX6 Human This gene encodes a member of the
Theoretical cloning project
 Theoretical cloning project Needed to get credits Make it up yourself, don't copy Possible to do in groups of 2-4 students If you need help or an idea, ask! If you have no idea what to clone, I can give
Theoretical cloning project Needed to get credits Make it up yourself, don't copy Possible to do in groups of 2-4 students If you need help or an idea, ask! If you have no idea what to clone, I can give
Supplementary Figure 1. NORAD expression in mouse (A) and dog (B). The black boxes indicate the position of the regions alignable to the 12 repeat
 Supplementary Figure 1. NORAD expression in mouse (A) and dog (B). The black boxes indicate the position of the regions alignable to the 12 repeat units in the human genome. Annotated transposable elements
Supplementary Figure 1. NORAD expression in mouse (A) and dog (B). The black boxes indicate the position of the regions alignable to the 12 repeat units in the human genome. Annotated transposable elements
The Biotechnology Toolbox
 Chapter 15 The Biotechnology Toolbox Cutting and Pasting DNA Cutting DNA Restriction endonuclease or restriction enzymes Cellular protection mechanism for infected foreign DNA Recognition and cutting specific
Chapter 15 The Biotechnology Toolbox Cutting and Pasting DNA Cutting DNA Restriction endonuclease or restriction enzymes Cellular protection mechanism for infected foreign DNA Recognition and cutting specific
7.012 Problem Set 5. Question 1
 Name Section 7.012 Problem Set 5 Question 1 While studying the problem of infertility, you attempt to isolate a hypothetical rabbit gene that accounts for the prolific reproduction of rabbits. After much
Name Section 7.012 Problem Set 5 Question 1 While studying the problem of infertility, you attempt to isolate a hypothetical rabbit gene that accounts for the prolific reproduction of rabbits. After much
Supporting Information
 Supporting Information Zorde Khvalevsky et al..73/pnas.337.9 mm sikrsg Molecule PG Matrix ± mm right sc left retained in OER E T= days T= days (iver) T= days (PS) Fig. S. OER characteristics. () iagram
Supporting Information Zorde Khvalevsky et al..73/pnas.337.9 mm sikrsg Molecule PG Matrix ± mm right sc left retained in OER E T= days T= days (iver) T= days (PS) Fig. S. OER characteristics. () iagram
TRANSGENIC ANIMALS. transient. stable. - Two methods to produce transgenic animals:
 Only for teaching purposes - not for reproduction or sale CELL TRANSFECTION transient stable TRANSGENIC ANIMALS - Two methods to produce transgenic animals: 1- DNA microinjection 2- embryonic stem cell-mediated
Only for teaching purposes - not for reproduction or sale CELL TRANSFECTION transient stable TRANSGENIC ANIMALS - Two methods to produce transgenic animals: 1- DNA microinjection 2- embryonic stem cell-mediated
TRIM31 is recruited to mitochondria after infection with SeV.
 Supplementary Figure 1 TRIM31 is recruited to mitochondria after infection with SeV. (a) Confocal microscopy of TRIM31-GFP transfected into HEK293T cells for 24 h followed with SeV infection for 6 h. MitoTracker
Supplementary Figure 1 TRIM31 is recruited to mitochondria after infection with SeV. (a) Confocal microscopy of TRIM31-GFP transfected into HEK293T cells for 24 h followed with SeV infection for 6 h. MitoTracker
Pei et al. Supplementary Figure S1
 Pei et al. Supplementary Figure S1 C H-CUL9: + + + + + Myc-ROC1: - - + + + U2OS/pcDN3 U2OS/H-CUL9 U2OS/ + H-CUL9 IP: -H -myc input -H -myc 1 2 3 4 5 H-CUL9 Myc-ROC1 -H -H -H -H H-CUL9: wt RR myc-roc1:
Pei et al. Supplementary Figure S1 C H-CUL9: + + + + + Myc-ROC1: - - + + + U2OS/pcDN3 U2OS/H-CUL9 U2OS/ + H-CUL9 IP: -H -myc input -H -myc 1 2 3 4 5 H-CUL9 Myc-ROC1 -H -H -H -H H-CUL9: wt RR myc-roc1:
scgem Workflow Experimental Design Single cell DNA methylation primer design
 scgem Workflow Experimental Design Single cell DNA methylation primer design The scgem DNA methylation assay uses qpcr to measure digestion of target loci by the methylation sensitive restriction endonuclease
scgem Workflow Experimental Design Single cell DNA methylation primer design The scgem DNA methylation assay uses qpcr to measure digestion of target loci by the methylation sensitive restriction endonuclease
PrimePCR Assay Validation Report
 Gene Information Gene Name keratin 3 Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID KRT3 Human The protein encoded by this gene is a member of the keratin
Gene Information Gene Name keratin 3 Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID KRT3 Human The protein encoded by this gene is a member of the keratin
PrimePCR Assay Validation Report
 Gene Information Gene Name keratin associated protein 9-2 Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID KRTAP9-2 Human This protein is a member of the keratin-associated
Gene Information Gene Name keratin associated protein 9-2 Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID KRTAP9-2 Human This protein is a member of the keratin-associated
TITLE: Development of Targeted Sindbis Virus Vectors for Potential^Application to Breast Cancer Therapy
 .^. AD Award Number: DAMD17-99-1-9232 TITLE: Development of Targeted Sindbis Virus Vectors for Potential^Application to Breast Cancer Therapy PRINCIPAL INVESTIGATOR: Lesia K. Dropulic, M.D. CONTRACTING
.^. AD Award Number: DAMD17-99-1-9232 TITLE: Development of Targeted Sindbis Virus Vectors for Potential^Application to Breast Cancer Therapy PRINCIPAL INVESTIGATOR: Lesia K. Dropulic, M.D. CONTRACTING
PrimePCR Assay Validation Report
 Gene Information Gene Name glutathione S-transferase alpha 1 Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID GSTA1 Human Cytosolic and membrane-bound forms
Gene Information Gene Name glutathione S-transferase alpha 1 Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID GSTA1 Human Cytosolic and membrane-bound forms
PrimePCR Assay Validation Report
 Gene Information Gene Name Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID glyceraldehyde-3-phosphate dehydrogenase GAPDH Human The product of this gene catalyzes
Gene Information Gene Name Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID glyceraldehyde-3-phosphate dehydrogenase GAPDH Human The product of this gene catalyzes
PrimePCR Assay Validation Report
 Gene Information Gene Name integrin, alpha 1 Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID ITGA1 Human This gene encodes the alpha 1 subunit of integrin
Gene Information Gene Name integrin, alpha 1 Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID ITGA1 Human This gene encodes the alpha 1 subunit of integrin
Supplementary Information
 Supplementary Information Super-resolution imaging of fluorescently labeled, endogenous RNA Polymerase II in living cells with CRISPR/Cas9-mediated gene editing Won-Ki Cho 1, Namrata Jayanth 1, Susan Mullen
Supplementary Information Super-resolution imaging of fluorescently labeled, endogenous RNA Polymerase II in living cells with CRISPR/Cas9-mediated gene editing Won-Ki Cho 1, Namrata Jayanth 1, Susan Mullen
Supplementary Information
 Supplementary Information Vidigal and Ventura a wt locus 5 region 3 region CCTCTGCCACTGCGAGGGCGTCCAATGGTGCTTG(...)AACAGGTGGAATATCCCTACTCTA predicted deletion clone 1 clone 2 clone 3 CCTCTGCCACTGCGAGGGCGTC-AGGTGGAATATCCCTACTCTA
Supplementary Information Vidigal and Ventura a wt locus 5 region 3 region CCTCTGCCACTGCGAGGGCGTCCAATGGTGCTTG(...)AACAGGTGGAATATCCCTACTCTA predicted deletion clone 1 clone 2 clone 3 CCTCTGCCACTGCGAGGGCGTC-AGGTGGAATATCCCTACTCTA
resequencing storage SNP ncrna metagenomics private trio de novo exome ncrna RNA DNA bioinformatics RNA-seq comparative genomics
 RNA Sequencing T TM variation genetics validation SNP ncrna metagenomics private trio de novo exome mendelian ChIP-seq RNA DNA bioinformatics custom target high-throughput resequencing storage ncrna comparative
RNA Sequencing T TM variation genetics validation SNP ncrna metagenomics private trio de novo exome mendelian ChIP-seq RNA DNA bioinformatics custom target high-throughput resequencing storage ncrna comparative
Figure S1: NUN preparation yields nascent, unadenylated RNA with a different profile from Total RNA.
 Summary of Supplemental Information Figure S1: NUN preparation yields nascent, unadenylated RNA with a different profile from Total RNA. Figure S2: rrna removal procedure is effective for clearing out
Summary of Supplemental Information Figure S1: NUN preparation yields nascent, unadenylated RNA with a different profile from Total RNA. Figure S2: rrna removal procedure is effective for clearing out
Relative Mouse Telomere Length Quantification qpcr Assay Kit (RMTLQ) Catalog #M reactions
 Relative Mouse Telomere Length Quantification qpcr Assay Kit (RMTLQ) Catalog #M8908 100 reactions Product Description Telomeres are repetitive nucleotide elements at the ends of chromosomes that protect
Relative Mouse Telomere Length Quantification qpcr Assay Kit (RMTLQ) Catalog #M8908 100 reactions Product Description Telomeres are repetitive nucleotide elements at the ends of chromosomes that protect
Supplemental Figure 1: CIP2A and SET levels are increased in some. primary human pancreatic cancer samples. (A) CIP2A mrna levels were
 Supplemental Figure Legends and Figures: Supplemental Figure 1: CIP2A and SET levels are increased in some primary human pancreatic cancer samples. (A) CIP2A mrna levels were measured in 3 benign (non-cancer)
Supplemental Figure Legends and Figures: Supplemental Figure 1: CIP2A and SET levels are increased in some primary human pancreatic cancer samples. (A) CIP2A mrna levels were measured in 3 benign (non-cancer)
PrimePCR Assay Validation Report
 Gene Information Gene Name keratin 5 Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID KRT5 Human The protein encoded by this gene is a member of the keratin
Gene Information Gene Name keratin 5 Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID KRT5 Human The protein encoded by this gene is a member of the keratin
Supplementary Figure 1. Phenotype and genotype of cultured and transplanted S1 KCST (A) Brightfield and mcherry fluorescence images of the spheres
 Supplementary Figure 1. Phenotype and genotype of cultured and transplanted S1 KCST (A) Brightfield and mcherry fluorescence images of the spheres generated from the CD133-positive cells infected with
Supplementary Figure 1. Phenotype and genotype of cultured and transplanted S1 KCST (A) Brightfield and mcherry fluorescence images of the spheres generated from the CD133-positive cells infected with
monoclonal antibody. (a) The specificity of the anti-rhbdd1 monoclonal antibody was examined in
 Supplementary information Supplementary figures Supplementary Figure 1 Determination of the s pecificity of in-house anti-rhbdd1 mouse monoclonal antibody. (a) The specificity of the anti-rhbdd1 monoclonal
Supplementary information Supplementary figures Supplementary Figure 1 Determination of the s pecificity of in-house anti-rhbdd1 mouse monoclonal antibody. (a) The specificity of the anti-rhbdd1 monoclonal
a KYSE270-CON KYSE270-Id1
 a KYSE27-CON KYSE27- shcon shcon sh b Human Mouse CD31 Relative MVD 3.5 3 2.5 2 1.5 1.5 *** *** c KYSE15 KYSE27 sirna (nm) 5 1 Id2 Id2 sirna 5 1 sirna (nm) 5 1 Id2 sirna 5 1 Id2 [h] (pg per ml) d 3 2 1
a KYSE27-CON KYSE27- shcon shcon sh b Human Mouse CD31 Relative MVD 3.5 3 2.5 2 1.5 1.5 *** *** c KYSE15 KYSE27 sirna (nm) 5 1 Id2 Id2 sirna 5 1 sirna (nm) 5 1 Id2 sirna 5 1 Id2 [h] (pg per ml) d 3 2 1
Single Cell Genomics
 Single Cell Genomics Application Cost Platform/Protoc ol Note Single cell 3 mrna-seq cell lysis/rt/library prep $2460/Sample 10X Genomics Chromium 500-10,000 cells/sample Single cell 5 V(D)J mrna-seq cell
Single Cell Genomics Application Cost Platform/Protoc ol Note Single cell 3 mrna-seq cell lysis/rt/library prep $2460/Sample 10X Genomics Chromium 500-10,000 cells/sample Single cell 5 V(D)J mrna-seq cell
PrimePCR Assay Validation Report
 Gene Information Gene Name Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID sema domain, immunoglobulin domain (Ig), short basic domain, secreted, (semaphorin)
Gene Information Gene Name Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID sema domain, immunoglobulin domain (Ig), short basic domain, secreted, (semaphorin)
