Novel plants optimised for lignin, growth, and biofuel production via re-mutagenesis and co-expression analysis
|
|
|
- Pearl Thomas
- 6 years ago
- Views:
Transcription
1 Novel plants optimised for lignin, growth, and biofuel production via re-mutagenesis and co-expression analysis Investigators In the Division of Plant Sciences, College of Life Sciences, University of Dundee: Claire Halpin, Professor and Deputy Head of Division Gordon Simpson, Lecturer Christopher McClellan, Postdoctoral researcher Katarzyna Rataj, Postdoctoral researcher Abdellah Barakate, Postdoctoral researcher Carinna Baird, Technician Sean Hackett, Technician (now left) External Collaborators: Wout Boerjan, Professor, University of Ghent, Belgium Simon McQueen-Mason, Professor, University of York, UK Abstract The objectives of this project are to find novel genes that will be useful for optimising plant biomass to facilitate biofuel production. These include genes influencing saccharification (the release of sugars from biomass), genes involved in the synthesis or control of lignin, and genes that restore normal growth to lignin mutants. Several approaches are being used towards this goal. Firstly, mutagenic screens to find mutants which improve the saccharification potential of existing monolignol biosynthesis mutants have been performed. After re-screening mutants initially identified as having increased saccharification, seven mutants have been isolated which significantly increase the release of sugars. Many of these mutants have reduced lignin levels, and one has already been through mutation mapping by next generation sequencing (NGS) while others are being prepared for mapping. The mapped mutant provides evidence that it is possible to stack two genes, one a gene of lignin biosynthesis and the other an identified gene on an interconnected pathway, to optimise sugar release. In another screen, 43 mutants have been identified that rescue the reduced size phenotype of a lignin biosnthesis mutant. A large portion of these restored expression mutants still contain significant reductions in lignin and a corresponding increase in sugar release upon saccharification. One of these mutants has been selected for mutation mapping to identify the causal mutation. A second approach used to identify novel genes involved in lignin synthesis and regulation is co-expression analysis. Over 65 mutants in genes identified as co-expressed with lignin biosynthesis genes have been analysed. Two of these mutants have been shown to contain reduced lignin and elevated sugar release in the saccharification assay, one of which has been rescued by over-expression of the targeted gene. Protein complex purification and identification has also been used to identify interacting partners of lignin biosynthesis proteins. A known lignin biosynthesis enzyme and a protein identified from the co-expression analysis have been used as baits for tagged purification strategies. Further work involves mapping more mutants from the largescale screens and characterization of the gene isolated by co-expression analysis. Introduction The aim of this project is to identify novel genes involved in limiting the sugar release from plant cell walls, particularly genes involved in lignin deposition by screening for plants that have improved saccharification properties or improved
2 growth characteristics. Novel genes will be identified through mutagenesis of existing lignin-defective mutants, as well as screening plants defective in genes that are co-expressed with known lignin biosynthetic genes. Further analysis of ligninrelated proteins will be undertaken by isolating interaction partners of known monolignol biosynthesis enzymes and these interaction partners will be identified through liquid chromatography/mass spectroscopy analysis. Genes identified with these strategies will be good candidates for manipulation in crops used in biofuel production. Background Many recent advances have shown the potential for improving biofuel production. Since the phenolic polymer lignin coats is cross-linked to other cell wall components, the reduction or alteration of this compound has long been a target for improving sugar release from cell walls. It has recently been shown that disrupting the lignin biosynthesis gene CAD1 causes improved saccharification in Brachypodium distachyon [1]. In addition to disrupting known lignin genes, the engineering of plants to produce altered lignin structure has proved effective at increasing saccharification. Arabidopsis plants engineered to incorporate hydroxybenzaldehydes into lignin rather than the usual hydroxycinnamates have shown improvements in saccharification [2]. Additionally, an engineered enzyme which methylates lignin precursors, making it more difficult for lignin polymerization to occur, increases the digestion efficiency of cell walls in plants expressing this enzyme [3]. Enzymes outside of lignin biosynthesis are possible targets as well, since overexpression of a BAHD acyltransferase increases p-coumarate incorporation into cell wall polysaccharides and improves saccharification [4]. However, more research is needed to obtain the necessary range and specificity of mutants for saccharification engineering. To find novel genes involved in lignin biosynthesis, we are using mutagenic screens of known lignin mutants, an approach that so far, is unique and promises to yield genes not known to be involved in lignin biosynthesis. We are also using co-expression analysis on specific pathways and processes using publicly available, high-quality data sets from microarray expression experiments. Previously, co-expression experiments have been used successfully to explore novel genes involved in some cell wall pathways and processes [5,6]. Recently, it has been found that two monolignol biosynthesis proteins, C3H and C4H interact at the endoplasmic reticulum membrane, and may interact with two other monolignol biosynthesis proteins, HCT and 4CL [7,8]. However, protein interactions beyond the known monolignol biosynthesis proteins are not well studied. We are using protein complex purification to potentially identify novel gene products involved in lignin biosynthesis. Results Mutant Screening Two main approaches have been used in the model plant Arabidopsis thaliana to find novel genes involved in improved biofuel production. The first approach, a forward genetics approach, uses genetic screening to identify mutants which are more desirable for bioethanol production. Two mutagenic screens have been performed. In both cases, a mutant, which has a defect in one of the monolignol biosynthesis genes, was mutagenised with the aim of finding new mutations which improve the original line s ability to be converted to bioethanol.
3 One mutagenic screen involves screening for saccharification improvement in a mutant which already has a small improvement in this phenotype. The mutant used for this screen is ref3, which is mutated in the C4H gene in Arabidopsis. This allele has small reductions in lignin and no visible plant growth phenotypes [9]. The mutant also has an increase of sugar release upon saccharification. It was hypothesized that further increases in saccharification potential could be found without affecting plant growth. To find such plants, a high-throughput saccharification assay was used to screen a mutagenized ref3 population [10]. This population was generated by treating ref3 with ethyl methanesulfonate (EMS), a mutagen, and stem material from each member of the M 2 screening population was collected. Stem material was then ground to powder for use in the saccharification screen. 20,000 M 2 plants were grown for the screen, but plant death and a lack of sufficient stem material for some plants led to only 10,000 plants being used in the screen. Using this screen, 26 potential high saccharification and 30 potential low saccharification mutants were identified. These potential mutants were then rescreened in replicate using the same saccharification assay. Out of the 26 potential high saccharification mutants, seven were determined to be significantly different from the background upon re-screening (Figure 1). Figure 1 ref3 enhancer mutants show increases in sugar release upon saccharification. Arabidopsis plants were grown for twelve weeks, and stems were collected and ground to powder. Powder was used in the highthroughput saccharification assay developed by Gomez et al. [10]. Each data point represents four technical replicates of four biological replicates. Error bars represent standard error of the mean. Since saccharification can be affected by changes in lignin levels, the acetyl bromide assay was used to assess lignin in these mutants (Figure 2). Many of the mutants also displayed reductions in lignin level.
4 Figure 2 ref3-3 enhancer mutants show reductions in lignin content. Arabidopsis plants were grown for twelve weeks and stems were ground to powder. Powder was extracted four times with 80% ethanol and once with chloroform:methanol (2:1 v/v). Extracted powder was used in the acetyl bromide assay [11]. Data represents three technical replicates of two biological replicates. Error bars represent standard error of the mean. Due to its increase in saccharification and reduction in lignin, the mutant was chosen for further analysis. Stem sections from the mutant were stained with a reagent to detect lignin (Figure 3). Interestingly, the shows a marked decrease in phloroglucinol staining, an indication of decreased lignin content in the xylem of mutant stems. The enhancer mutant also displays a collapsed xylem phenotype (Figure 3), a phenotype indicative of cell wall compositional defects. Figure 3 Lignin staining in stem sections of the ref3 enhancer Stems from six-week old Arabidopsis plants were collected and sliced into 100 m sections. Sections were stained with 0.67% (w/v) phloroglucinol and imaged with a light microscope. To find the gene responsible for the high saccharification phenotype of the mutant, a next generation sequencing (NGS) approach was used. The MutMap process, developed by Abe et al. [12] was used. In this approach, there is no need to cross the desired mutant to a different genetic background. Instead, the mutant
5 was crossed back to the wild type and F 2 plants from this cross were used to generate a mapping population. Plants that were homozygous for the ref3 allele were selected and used in the high-throughput saccharification assay (Figure 4). Figure 4 A mapping population generated by a high-throughput saccharification assay. Arabidopsis plants were grown for twelve weeks, and stems were collected and ground to powder. Powder was used in the high-throughput saccharification assay developed by Gomez et al. [10]. Each data point represents four technical replicates. Error bars represent standard error of the mean. Plants that exceeded the cut-off were then selected for the mapping population. Genomic DNA was isolated from these plants (n=16), pooled, and used for sequencing on an Illumina HiSeq2000 platform. The resulting sequences were aligned with the Arabidopsis reference sequence and polymorphisms were detected using VarScan. The resulting single nucleotide polymorphisms (SNPs) were plotted based on the frequency of the mutation in the sequencing pool. This index was plotted for all SNPs over the genome. Two peaks were identified, one on the bottom arm of chromosome 2, and the other on a different chromosome. The peak on chromosome 2 can be explained by the selection of the ref3 allele, so the peak on the other chromosome was determined to be responsible for the improved saccharification phenotype. A mapping window was defined by at least four consecutive SNPs with an index value 0.9. A total of 13 protein coding changes were found within the mapping window. Of these, there was one nonsense mutation, within the gene X. Subsequent Sanger sequencing of the and mutants also identified stop codon mutations within gene X, demonstrating that mutating this gene is responsible for a high saccharification phenotype when combined with ref3. Another screen to find new genes which improve the saccharification properties of plants involved the mutagenesis of the founder mutant, which contains a mutation in a gene encoding a lignin biosynthesis gene in Arabidopsis thaliana. This mutant has
6 decreased lignin amounts and releases more sugar upon saccharification of stem tissue, but has a plant growth defect [13]. The founder mutant was treated with EMS and 80,000 plants from the resulting M 2 generation were screened for increased plant size. 71 mutants were identified originally, but upon re-screening, only 43 of these potential suppressors rescued the plant size defect of the founder mutant. For these 43 mutants, the stem weight of each mutant was quantified (Figure 5). Figure 5 Stem weights of suppressor mutants isolated from a screen for increased plant size. Arabidopsis plants were grown for twelve weeks and stems collected. Each data point represents the mean of eight to ten stems, with error bars representing standard error of the mean. While not all mutants were rescued back to the size of the wild type, every mutant identified has an increase in average stem weight that is statistically different from the founder mutant, with the exception of two (mutants 73-3 and in Figure 1). Many of the mutant lines isolated still show reductions in lignin content when assayed by the acetyl bromide method (Figure 6) [11]. Founder Figure 6 Acetyl bromide lignin content of several founder mutant suppressors. Arabidopsis stems were collected after twelve weeks of growth and ground to powder. Powder was extracted four times with 80% ethanol and once with 2:1 chloroform: methanol. Extracted stem powder was treated with 25% acetyl bromide at 70 C for 30 minutes, and absorbance at 280 nm measured to quantify lignin [11].
7 Furthermore, when tested for saccharification, five of the six lines above showed significant differences from the wild type, with the exception being line 47-1 (Figure 7). Founder Figure 7 Saccharification analysis of founder mutant suppressors. Stems were collected from twelve-week old Arabidopsis plants and ground into powder. Powder was then used in high-throughput saccharification analysis as in Gomez et al. [10]. # One of the lines presented in figures 3 and 4 is now being used in mutation mapping. Each line has been crossed to the wild type, and plants from the resulting F 2 generation that exhibit the rescued plant size defect have been selected for the mapping population. DNA from the mapping population will be isolated and a next generation sequencing approach will be used to identify the causal mutation, as developed by Abe et al. [12]. Co-expression analysis and protein interactions The second main approach used to identify novel genes involved in lignin production was a reverse genetics approach, gene co-expression analysis. In order to retrieve potential candidates involved in lignification, co-expression analysis with known monolignol biosynthetic genes was performed with three data sets: ACT [14], CressExpress [15] and Ray (Raymond Wightman and Simon Turner- personal communication). A total of 255 genes were retrieved, several of which were selected for further analysis. T-DNA insertion lines were ordered for the selected genes and these lines were tested for acetyl bromide lignin [11] and several showed reduced lignin levels. A saccharification test on these mutants identified two mutants showing increased sugar release. The Gb mutant was selected for further analysis. To this end, plants overexpressing the Gb gene in the mutant background were prepared. The recovery of wild-type features in these plants was tested to confirm that detected developmental defects and reduced lignin levels are a consequence of the Gb mutation. Gb overexpressing plants were compared to wild type (Col-0) plants, the original Gb9 knock-out mutant and the weak Mx12 knock-down mutant, which shows partial reduction of Gb transcript expression. Phenotypic analysis revealed that wild-type plant growth can be restored in plants with overexpressed Gb:GFP or Gb:HBH introduced (8).
8 Figure 8 Overexpression of tagged Gb proteins results in rescue of stunted growth. Plants overexpressing Gb:GFP and Gb:HBH proteins were grown alongside wild type (Col-0) and Gb9 and Mx12 mutant controls. A reduced growth phenotype can be observed only in Gb9 plants. Transgenic plants were also scored for their height (Figure 9a) and total weight (Figure 9b). Recovery of the wild type features was observed in independent transgenic lines. Figure 9 Overexpression of tagged Gb proteins results in rescue of wild-type growth. Plants overexpressing Gb:GFP and Gb:HBH proteins were grown alongside wild type (Col-0) and Gb9 and Mx12 mutant controls. Reduced growth phenotype can
9 be observed only in Gb9 plants. (a) Final plant height was measured. (b) Dry stem weight of transgenic plants. Gb9 mutants show reduced lignin levels and increased saccharification. To confirm that these characteristics are due to specific gene disruption levels of lignin in Gb9 plants with introduced Gb:GFP overexpressing transgenes were tested. Lignin was detected using phloroglucinol reagent (Figure 10). Figure 10 Lignin staining in stem sections of plants overexpressing Gb:GFP transgene compared to wild type (Col-0) and Gb9 and Mx12 mutant controls. Stems from five-week old Arabidopsis plants were collected and sliced into 100 m sections. Sections were stained with phloroglucinol reagent. Stem staining revealed that lignin reduction was largely restored in complementation lines. Additionally, collapsed vessels that are present in Gb9 mutant plants could not be detected in plants overexpressing the Gb:GFP transgene. Reduced lignin levels in Gb9 mutants are linked with increased sugar release from these plants. To test if restored lignin levels and vessel element structure also leads to restored sugar release, a saccharification assay was performed. This revealed that increased lignin levels observed in Gb:GFP and Gb:HBH overexpressing complementation lines results in decreased sugar release, which indicates that the increased saccharification observed in Gb9 mutants is directly linked with the decreased lignin level seen in the mutant. Analysis of protein complexes formed by known lignin biosynthesis enzymes could reveal new proteins involved in the lignification process. In order to identify protein interactions occurring in vivo, potential protein complexes in the lignin biosynthesis pathway are being purified directly from plants, including complexes involving the Gb protein. Presence of the tagged proteins in transgenic lines was confirmed using western blotting with tag-specific antibodies. The presence of GFP-tagged proteins could be detected in selected transgenic lines. These transgenic lines were used for protein purification. In order to stabilise protein complexes present in the plants, a protein cross-linking step was first performed. This cross-linked material was used in protein purification. GFP-tagged proteins can be purified using GFP trap, which consists of a small peptide that specifically binds GFP [16]. To purify complexes formed by the GFP-tagged
10 proteins, GFP trap fused with agarose beads was used. Presence of GFP-tagged proteins in the protein purification samples was confirmed by western blot. Additional proteins co-purifying with tagged proteins were visualized with Coomassie blue stain and were subsequently identified using using liquid chromatography/mass spectrometry (LC/MS). In order to analyse protein interactions present in lignifying tissue, large scale protein purification of GFP-tagged protein was performed using Arabidopsis stems. Additional proteins co-purifying with tagged Gb:GFP and CCR1:GFP proteins were detected. LC/MS analysis of proteins co-purifying with Gb:GFP revealed the presence of cell wall related proteins, xyloglucan hydrolase 24 being the most abundant. However, no interactions between Gb and core components of lignin pathway could be detected in Arabidopsis stem samples. This analysis reveals additional previously unstudied proteins that could be involved in cell wall formation in Arabidopsis. Progress Several mutants have been identified in Arabidopis thaliana that improve the saccharification potential of the plant. These mutants have been quantified for lignin levels, and found to have decreased lignin. One mutant has had its causal mutation mapped and the mutated gene was identified. Another screen has identified a mutant which has rescued growth but maintains reduced lignin levels and increased saccharification. Co-expression analysis has identified a gene, that when mutated, contains less lignin and releases more sugars upon saccharification. Transformation of this mutant with an overexpression rescue construct confirms that these phenotypes are the result of the studied gene. The protein produced from this gene has also been used for protein complex purification, along with previously studied lignin biosynthesis enzymes. Initial results indicate that the Gb protein may interact with some cell wall components, suggesting a role in cell wall architecture. Future Plans For the remainder of the project, the focus for the mutant screening portion of the project will be on identifying the causal mutations behind the observed phenotypes. Two more mutants from the ref3 enhancer screen will be mapped using NGS, one of high saccharification, the other of low saccharification. The mutant from the suppressor screen of the lignin mutant with impaired growth will also be mapped with the same procedure. Transgenic Arabidopsis overexpressing the rice Gb protein will be analysed to test common function of homologues form different plants. Additionally, an antibody against Arabidopsis Gb protein is being prepared. This antibody will facilitate future studies of the Gb protein. The specificity of this antibody and its ability to recognise Gb homologues from different organisms will also be tested. Publications Patents: Modified Plants filed May Application number GB Papers: Two papers are in preparation, one on the Gb mutant (in collaboration with Wout Boerjan) and one on the stacking of mutations in the large-scale mutant screen.
11 Talk: Claire Halpin at UKPlantSci 2013, Dundee Spinning straw into gold: Barley, biofuels and biosequestration April 2013 Talk: Claire Halpin at Monogram Network Meeting Improving barley straw for bioenergy March 2012 Talk: Chris McClellan at UK PlantSci 2012, Norwich, UK: Improving plants for the production of 2 nd generation biofuels. 18 April 2012 Poster: Kasia Rataj at 23 rd International Conference on Arabidopsis Research, Vienna, Austria: Optimization of lignin content in Arabidopsis to facilitate biofuel production 3-7 July 2012 Poster: Chris McClellan at UK PlantSci 2012, Norwich, UK: Genetic screening for improved saccharification properties in Arabidopsis thaliana April 2012 References 1. Bouvier d Yvoire, M., O. Bouchabke-Coussa, W. Voorend, S. Antelme, L. Cézard, F. Legée, P. Lebris, S. Legay, C. Whitehead, S.J. McQueen-Mason, L.D. Gomez, L. Jouanin, C. Lapierre, and R. Sibout, Disrupting the cinnamyl alcohol dehydrogenase 1 gene (BdCAD1) leads to altered lignification and improved saccharification in Brachypodium distachyon, Plant J., 73, , Eudes, A., A. George, P. Mukerjee, J.S. Kim, B. Pollet, P.I. Benke, F. Yang, P. Mitra, L. Sun, Ö.P. Çetinkol, S. Chabout, G. Mouille, L. Soubigou-Taconnat, S. Balzergue, S. Singh, B.H. Holmes, A. Mukhopadhyay, J.D. Keasling, B.A. Simmons, C. Lapierre, J. Ralph, and D. Loqué, Biosynthesis and incorporation of side-chain-truncated lignin monomers to reduce lignin polymerization and enhance saccharification, Plant Biotechnol. J., 10, , Zhang, K., M.-W. Bhuiya, J.R. Pazo, Y. Miao, H. Kim, J. Ralph, and C.-J. Liu, An engineered monolignol 4-O-methyltransferase depresses lignin biosynthesis and confers novel metabolic capability in Arabidopsis, Plant Cell, 24, , Bartley, L.E., M.L. Peck, S.-R. Kim, B. Ebert, C. Manisseri, D.M. Chiniquy, R. Sykes, L. Gao, C. Rautengarten, M.E. Vega-Sánchez, P.I. Benke, P.E. Canlas, P. Cao, S. Brewer, F. Lin, W.L. Smith, X. Zhang, J.D. Keasling, R.E. Jentoff, S.B. Foster, J. Zhou, A. Ziebell, G. An, H.V. Scheller, and P.C. Ronald, Overexpression of a BAHD acyltransferase, OsAt10, alters rice cell wall hydroxycinnamic acid content and saccharification, Plant Physiol., 161, , Brown, D.M., L.A. Zeef, J. Ellis, R. Goodacre, and S.R. Turner, Identification of novel genes in Arabidopsis involved in secondary cell wall formation using expression profiling and reverse genetics, Plant Cell, 17, , Vanholme, R., V. Storme, B. Vanholme, L. Sundin, J.H. Christensen, G. Goeminne, C. Halpin, A. Rohde, K. Moreel, and W. Boerjan, A systems biology view of responses to lignin biosynthesis perturbations in Arabidopsis thaliana, Plant Cell, 24, , Chen, H.C., Q. Li, C.M. Shuford, J. Liu, D.C. Muddiman, R.R. Sederoff, and V.L. Chiang, Membrane protein complexes catalyze both 4- and 3-hydroxylation of cinnamic acid derivatives in monolignol biosynthesis, Proc. Natl. Acad. Sci. USA, 108, , Bassard, J.-E., L. Richert, J. Geerinck, H. Renault, F. Duval, P. Ullmann, M. Schmitt, E. Meyer, J. Mutterer, W. Boerjan, G. De Jaeger, Y. Mely, A. Goossens, and D. Werck-Reichhart, Proteinprotein and protein-membrane associations in the lignin pathway, Plant Cell, 24, , Schilmiller, A.L., J. Stout, J.K. Weng, J. Humphreys, M.O. Ruegger, and C. Chapple, Mutations in the cinnamate 4-hydroxylase gene impact metabolism, growth, and development in Arabidopsis, Plant J., 60, , Gomez, L.D., C. Whitehead, A. Barakate, C. Halpin, and S.J. McQueen-Mason, Automated saccharification assay for determination of digestibility in plant materials, Biotechnology for Biofuels, 3, 23, Chang, X.F., R. Chandra, T. Berleth, and R.P. Beatson, Rapid, microscale, acetyl bromidebased method for high-throughput determination of lignin content in Arabidopsis thaliana, J. Agric. Food Chem., 56, , 2008.
12 12. Abe, A., S. Kosugi, K. Yoshida, S. Natsume, H. Takagi, H. Kanzaki, H. Matsumura, K. Yoshida, C. Mitsuoka, M. Tamiru, H. Innan, L. Cano, S. Kamoun, and R. Terauchi, Genome sequencing reveals agronomically important loci in rice using MutMap, Nat. Biotechnol., 30, , Mir Derikvand, M., J.B. Sierra, K. Ruel, B. Pollet, C.T. Do, J. Thévenin, D. Buffard, L. Jouanin, and C. Lapierre, Redirection of the phenylpropanoid pathway to feruloyl malate in Arabidopsis mutants deficient for cinnamoyl-coa reductase 1, Planta, 227, , Manfield, I.W., C.H. Jen, J.W. Pinney, I. Michalopolous, J.R. Bradford, P.M. Gilmartin, & D.R. Westhead, Arabidopsis co-expression tool (ACT): web server tools for microarray-based gene expression analysis, Nucleic Acids Res., 34, W504-W509, Srinivasasainagendra, V., G.P. Page, T. Mehta, I. Coulibaly, and A.E. Loraine, CressExpress: A tool for large-scale mining of expression data from Arabidopsis. Plant Physiol., 147, , Rothbauer, U., K. Zolghader, S. Muyldermans, A. Schepers, M.C. Cardoso, and H. Leonhardt, A versatile nanotrap for biochemical and functional studies with fluorescent fusion proteins, Mol. Cell. Proteomics, 7, , Contacts Claire Halpin: c.halpin@dundee.ac.uk Gordon Simpson: g.g.simpson@dundee.ac.uk
Biomass conversion: opportunities in lignin management. Clint Chapple Department of Biochemistry Purdue University
 Biomass conversion: opportunities in lignin management Clint Chapple Department of Biochemistry Purdue University Targets for biomass improvement Yield Water and nutrient use efficiency Pest and pathogen
Biomass conversion: opportunities in lignin management Clint Chapple Department of Biochemistry Purdue University Targets for biomass improvement Yield Water and nutrient use efficiency Pest and pathogen
Enhancers mutations that make the original mutant phenotype more extreme. Suppressors mutations that make the original mutant phenotype less extreme
 Interactomics and Proteomics 1. Interactomics The field of interactomics is concerned with interactions between genes or proteins. They can be genetic interactions, in which two genes are involved in the
Interactomics and Proteomics 1. Interactomics The field of interactomics is concerned with interactions between genes or proteins. They can be genetic interactions, in which two genes are involved in the
Gene expression in the lignin biosynthesis pathway during soybean seed development
 Gene expression in the lignin biosynthesis pathway during soybean seed development A. Baldoni, E.V.R. Von Pinho, J.S. Fernandes, V.M. Abreu and M.L.M. Carvalho Laboratório Central de Sementes, Departamento
Gene expression in the lignin biosynthesis pathway during soybean seed development A. Baldoni, E.V.R. Von Pinho, J.S. Fernandes, V.M. Abreu and M.L.M. Carvalho Laboratório Central de Sementes, Departamento
Lecture 2: Biology Basics Continued
 Lecture 2: Biology Basics Continued Central Dogma DNA: The Code of Life The structure and the four genomic letters code for all living organisms Adenine, Guanine, Thymine, and Cytosine which pair A-T and
Lecture 2: Biology Basics Continued Central Dogma DNA: The Code of Life The structure and the four genomic letters code for all living organisms Adenine, Guanine, Thymine, and Cytosine which pair A-T and
F 11/23 Happy Thanksgiving! 8 M 11/26 Gene identification in the genomic era Bamshad et al. Nature Reviews Genetics 12: , 2011
 3 rd Edition 4 th Edition Lecture Day Date Topic Reading Problems Reading Problems 1 M 11/5 Complementation testing reveals that genes are distinct entities Ch. 7 224-232 2 W 11/7 One gene makes one protein
3 rd Edition 4 th Edition Lecture Day Date Topic Reading Problems Reading Problems 1 M 11/5 Complementation testing reveals that genes are distinct entities Ch. 7 224-232 2 W 11/7 One gene makes one protein
Lignin Management: Optimizing Yield in Lignin Modified Plants
 Lignin Management: Optimizing Yield in Lignin Modified Plants Investigators Clint Chapple (CC), Professor, Department of Biochemistry, Purdue University Whitney Dolan, graduate student, Department of Biochemistry,
Lignin Management: Optimizing Yield in Lignin Modified Plants Investigators Clint Chapple (CC), Professor, Department of Biochemistry, Purdue University Whitney Dolan, graduate student, Department of Biochemistry,
Trasposable elements: Uses of P elements Problem set B at the end
 Trasposable elements: Uses of P elements Problem set B at the end P-elements have revolutionized the way Drosophila geneticists conduct their research. Here, we will discuss just a few of the approaches
Trasposable elements: Uses of P elements Problem set B at the end P-elements have revolutionized the way Drosophila geneticists conduct their research. Here, we will discuss just a few of the approaches
From Biomass to Biofuels. Joint BioEnergy Institute Emeryville, CA
 From Biomass to Biofuels Joint BioEnergy Institute Emeryville, CA JBEI at a glance Six Partners Lawrence Berkeley Nat l Lab Sandia Nat l Lab Lawrence Livermore Nat l Lab UC Berkeley UC Davis Carnegie Institute
From Biomass to Biofuels Joint BioEnergy Institute Emeryville, CA JBEI at a glance Six Partners Lawrence Berkeley Nat l Lab Sandia Nat l Lab Lawrence Livermore Nat l Lab UC Berkeley UC Davis Carnegie Institute
LATE-PCR. Linear-After-The-Exponential
 LATE-PCR Linear-After-The-Exponential A Patented Invention of the Laboratory of Human Genetics and Reproductive Biology Lab. Director: Lawrence J. Wangh, Ph.D. Department of Biology, Brandeis University,
LATE-PCR Linear-After-The-Exponential A Patented Invention of the Laboratory of Human Genetics and Reproductive Biology Lab. Director: Lawrence J. Wangh, Ph.D. Department of Biology, Brandeis University,
Genetic Engineering for Biofuels Production
 Genetic Engineering for Biofuels Production WSE 573 Spring 2013 Greeley Beck INTRODUCTION Alternative transportation fuels are needed in the United States because of oil supply insecurity, oil price increases,
Genetic Engineering for Biofuels Production WSE 573 Spring 2013 Greeley Beck INTRODUCTION Alternative transportation fuels are needed in the United States because of oil supply insecurity, oil price increases,
Lecture 25 (11/15/17)
 Lecture 25 (11/15/17) Reading: Ch9; 328-332 Ch25; 990-995, 1005-1012 Problems: Ch9 (study-guide: applying); 1,2 Ch9 (study-guide: facts); 7,8 Ch25 (text); 1-3,5-7,9,10,13-15 Ch25 (study-guide: applying);
Lecture 25 (11/15/17) Reading: Ch9; 328-332 Ch25; 990-995, 1005-1012 Problems: Ch9 (study-guide: applying); 1,2 Ch9 (study-guide: facts); 7,8 Ch25 (text); 1-3,5-7,9,10,13-15 Ch25 (study-guide: applying);
The Two-Hybrid System
 Encyclopedic Reference of Genomics and Proteomics in Molecular Medicine The Two-Hybrid System Carolina Vollert & Peter Uetz Institut für Genetik Forschungszentrum Karlsruhe PO Box 3640 D-76021 Karlsruhe
Encyclopedic Reference of Genomics and Proteomics in Molecular Medicine The Two-Hybrid System Carolina Vollert & Peter Uetz Institut für Genetik Forschungszentrum Karlsruhe PO Box 3640 D-76021 Karlsruhe
Lecture 2-3: Using Mutants to study Biological processes
 Lecture 2-3: Using Mutants to study Biological processes Objectives: 1. Why use mutants? 2. How are mutants isolated? 3. What important genetic analyses must be done immediately after a genetic screen
Lecture 2-3: Using Mutants to study Biological processes Objectives: 1. Why use mutants? 2. How are mutants isolated? 3. What important genetic analyses must be done immediately after a genetic screen
Genetics - Problem Drill 19: Dissection of Gene Function: Mutational Analysis of Model Organisms
 Genetics - Problem Drill 19: Dissection of Gene Function: Mutational Analysis of Model Organisms No. 1 of 10 1. The mouse gene knockout is based on. (A) Homologous recombination (B) Site-specific recombination
Genetics - Problem Drill 19: Dissection of Gene Function: Mutational Analysis of Model Organisms No. 1 of 10 1. The mouse gene knockout is based on. (A) Homologous recombination (B) Site-specific recombination
AP Biology. Scoring Guidelines
 2017 AP Biology Scoring Guidelines College Board, Advanced Placement Program, AP, AP Central, and the acorn logo are registered trademarks of the College Board. AP Central is the official online home for
2017 AP Biology Scoring Guidelines College Board, Advanced Placement Program, AP, AP Central, and the acorn logo are registered trademarks of the College Board. AP Central is the official online home for
Recombinant DNA Technology. The Role of Recombinant DNA Technology in Biotechnology. yeast. Biotechnology. Recombinant DNA technology.
 PowerPoint Lecture Presentations prepared by Mindy Miller-Kittrell, North Carolina State University C H A P T E R 8 Recombinant DNA Technology The Role of Recombinant DNA Technology in Biotechnology Biotechnology?
PowerPoint Lecture Presentations prepared by Mindy Miller-Kittrell, North Carolina State University C H A P T E R 8 Recombinant DNA Technology The Role of Recombinant DNA Technology in Biotechnology Biotechnology?
Gene Expression Technology
 Gene Expression Technology Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Gene expression Gene expression is the process by which information from a gene
Gene Expression Technology Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Gene expression Gene expression is the process by which information from a gene
Biotechnology. Chapter 20. Biology Eighth Edition Neil Campbell and Jane Reece. PowerPoint Lecture Presentations for
 Chapter 20 Biotechnology PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from Joan Sharp Copyright
Chapter 20 Biotechnology PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from Joan Sharp Copyright
I (ALLEN 1965; GIBSON 1970). In Tetrahymena pyriformis, syngen 1, a genetically
 SHORT PAPERS MUTANT SELECTION IN TETRAHYMENA PYRIFORMIS PETER S. CARLSON' Department of Biology, Wesleyan University, Middletown, Connecticut 06457 Received June 18, 1971 NTRACLONAL variation is a well
SHORT PAPERS MUTANT SELECTION IN TETRAHYMENA PYRIFORMIS PETER S. CARLSON' Department of Biology, Wesleyan University, Middletown, Connecticut 06457 Received June 18, 1971 NTRACLONAL variation is a well
Link between iron homeostasis, oxidative mutagenesis and antibiotic resistance evolution
 Thesis of the PhD dissertation Link between iron homeostasis, oxidative mutagenesis and antibiotic resistance evolution Orsolya Katinka Méhi Supervisor: Dr. Csaba Pál, senior research associate PhD School
Thesis of the PhD dissertation Link between iron homeostasis, oxidative mutagenesis and antibiotic resistance evolution Orsolya Katinka Méhi Supervisor: Dr. Csaba Pál, senior research associate PhD School
When plants are grown in elevated carbon dioxide concentrations, which genes control the stimulation of metabolism and growth?
 When plants are grown in elevated carbon dioxide concentrations, which genes control the stimulation of metabolism and growth? Ryan Boyd Andrew Leakey Overveiw! Introduction! Approach! Materials and Methods!
When plants are grown in elevated carbon dioxide concentrations, which genes control the stimulation of metabolism and growth? Ryan Boyd Andrew Leakey Overveiw! Introduction! Approach! Materials and Methods!
Alternative Cleavage and Polyadenylation of RNA
 Developmental Cell 18 Supplemental Information The Spen Family Protein FPA Controls Alternative Cleavage and Polyadenylation of RNA Csaba Hornyik, Lionel C. Terzi, and Gordon G. Simpson Figure S1, related
Developmental Cell 18 Supplemental Information The Spen Family Protein FPA Controls Alternative Cleavage and Polyadenylation of RNA Csaba Hornyik, Lionel C. Terzi, and Gordon G. Simpson Figure S1, related
AP Biology Gene Expression/Biotechnology REVIEW
 AP Biology Gene Expression/Biotechnology REVIEW Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Gene expression can be a. regulated before transcription.
AP Biology Gene Expression/Biotechnology REVIEW Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Gene expression can be a. regulated before transcription.
Using mutants to clone genes
 Using mutants to clone genes Objectives 1. What is positional cloning? 2. What is insertional tagging? 3. How can one confirm that the gene cloned is the same one that is mutated to give the phenotype
Using mutants to clone genes Objectives 1. What is positional cloning? 2. What is insertional tagging? 3. How can one confirm that the gene cloned is the same one that is mutated to give the phenotype
Synthetic Biology for
 Synthetic Biology for Plasmids and DNA Digestion Plasmids Plasmids are small DNA molecules that are separate from chromosomal DNA They are most commonly found as double stranded, circular DNA Typical plasmids
Synthetic Biology for Plasmids and DNA Digestion Plasmids Plasmids are small DNA molecules that are separate from chromosomal DNA They are most commonly found as double stranded, circular DNA Typical plasmids
Lecture 3 Mutagens and Mutagenesis. 1. Mutagens A. Physical and Chemical mutagens B. Transposons and retrotransposons C. T-DNA
 Lecture 3 Mutagens and Mutagenesis 1. Mutagens A. Physical and Chemical mutagens B. Transposons and retrotransposons C. T-DNA 2. Mutagenesis A. Screen B. Selection C. Lethal mutations Read: 508-514 Figs:
Lecture 3 Mutagens and Mutagenesis 1. Mutagens A. Physical and Chemical mutagens B. Transposons and retrotransposons C. T-DNA 2. Mutagenesis A. Screen B. Selection C. Lethal mutations Read: 508-514 Figs:
Activation of a Floral Homeotic Gene in Arabidopsis
 Activation of a Floral Homeotic Gene in Arabidopsis Busch, M.A., Bomblies, K., Weigel, D. kuleuven-kortrijk.be Simon Olthof Louie Christodoulidis 1 In Arabidopsis flowers, appearance is based, in part,
Activation of a Floral Homeotic Gene in Arabidopsis Busch, M.A., Bomblies, K., Weigel, D. kuleuven-kortrijk.be Simon Olthof Louie Christodoulidis 1 In Arabidopsis flowers, appearance is based, in part,
Contents... vii. List of Figures... xii. List of Tables... xiv. Abbreviatons... xv. Summary... xvii. 1. Introduction In vitro evolution...
 vii Contents Contents... vii List of Figures... xii List of Tables... xiv Abbreviatons... xv Summary... xvii 1. Introduction...1 1.1 In vitro evolution... 1 1.2 Phage Display Technology... 3 1.3 Cell surface
vii Contents Contents... vii List of Figures... xii List of Tables... xiv Abbreviatons... xv Summary... xvii 1. Introduction...1 1.1 In vitro evolution... 1 1.2 Phage Display Technology... 3 1.3 Cell surface
Chapter 9 Genetic Engineering
 Chapter 9 Genetic Engineering Biotechnology: use of microbes to make a protein product Recombinant DNA Technology: Insertion or modification of genes to produce desired proteins Genetic engineering: manipulation
Chapter 9 Genetic Engineering Biotechnology: use of microbes to make a protein product Recombinant DNA Technology: Insertion or modification of genes to produce desired proteins Genetic engineering: manipulation
SuperScript IV Reverse Transcriptase as a better alternative to AMV-based enzymes
 WHITE PAPER SuperScript IV Reverse Transcriptase SuperScript IV Reverse Transcriptase as a better alternative to AMV-based enzymes Abstract Reverse transcriptases (RTs) from avian myeloblastosis virus
WHITE PAPER SuperScript IV Reverse Transcriptase SuperScript IV Reverse Transcriptase as a better alternative to AMV-based enzymes Abstract Reverse transcriptases (RTs) from avian myeloblastosis virus
Two possible explanations for dominant mutations Haploinsufficiency vs Dominant negative
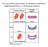 Two possible explanations for dominant mutations Haploinsufficiency vs Dominant negative Incomplete dominance for flower colour in Mirabilis jalapa R 1 R 1 R 2 R 2 R 1 R 2 R 1 Red pigment R 2 No pigment
Two possible explanations for dominant mutations Haploinsufficiency vs Dominant negative Incomplete dominance for flower colour in Mirabilis jalapa R 1 R 1 R 2 R 2 R 1 R 2 R 1 Red pigment R 2 No pigment
Supplementary Figure S1. Immunodetection of full-length XA21 and the XA21 C-terminal cleavage product.
 Supplementary Information Supplementary Figure S1. Immunodetection of full-length XA21 and the XA21 C-terminal cleavage product. Total protein extracted from Kitaake wild type and rice plants carrying
Supplementary Information Supplementary Figure S1. Immunodetection of full-length XA21 and the XA21 C-terminal cleavage product. Total protein extracted from Kitaake wild type and rice plants carrying
Synthetic Biology. Sustainable Energy. Therapeutics Industrial Enzymes. Agriculture. Accelerating Discoveries, Expanding Possibilities. Design.
 Synthetic Biology Accelerating Discoveries, Expanding Possibilities Sustainable Energy Therapeutics Industrial Enzymes Agriculture Design Build Generate Solutions to Advance Synthetic Biology Research
Synthetic Biology Accelerating Discoveries, Expanding Possibilities Sustainable Energy Therapeutics Industrial Enzymes Agriculture Design Build Generate Solutions to Advance Synthetic Biology Research
Lecture #1. Introduction to microarray technology
 Lecture #1 Introduction to microarray technology Outline General purpose Microarray assay concept Basic microarray experimental process cdna/two channel arrays Oligonucleotide arrays Exon arrays Comparing
Lecture #1 Introduction to microarray technology Outline General purpose Microarray assay concept Basic microarray experimental process cdna/two channel arrays Oligonucleotide arrays Exon arrays Comparing
Intragenic suppressors: true revertants
 Suppressor Genetics: intragenic and informational suppressors We have covered one approach used in genetics to study gene function: classical forward genetics where an investigator selects or screens for
Suppressor Genetics: intragenic and informational suppressors We have covered one approach used in genetics to study gene function: classical forward genetics where an investigator selects or screens for
Biochemistry study of the molecular basis of life
 Biochemistry : An Introduction Biochemistry study of the molecular basis of life n Study of the chemistry of living organisms Studies organic molecules & organic reactions in living organisms n Living
Biochemistry : An Introduction Biochemistry study of the molecular basis of life n Study of the chemistry of living organisms Studies organic molecules & organic reactions in living organisms n Living
3 Designing Primers for Site-Directed Mutagenesis
 3 Designing Primers for Site-Directed Mutagenesis 3.1 Learning Objectives During the next two labs you will learn the basics of site-directed mutagenesis: you will design primers for the mutants you designed
3 Designing Primers for Site-Directed Mutagenesis 3.1 Learning Objectives During the next two labs you will learn the basics of site-directed mutagenesis: you will design primers for the mutants you designed
Supplemental Figure 1. Mutation in NLA Causes Increased Pi Uptake Activity and
 Supplemental Figure 1. Mutation in NLA Causes Increased Pi Uptake Activity and PHT1 Protein Amounts. (A) Shoot morphology of 19-day-old nla mutants under Pi-sufficient conditions. (B) [ 33 P]Pi uptake
Supplemental Figure 1. Mutation in NLA Causes Increased Pi Uptake Activity and PHT1 Protein Amounts. (A) Shoot morphology of 19-day-old nla mutants under Pi-sufficient conditions. (B) [ 33 P]Pi uptake
Construction of plant complementation vector and generation of transgenic plants
 MATERIAL S AND METHODS Plant materials and growth conditions Arabidopsis ecotype Columbia (Col0) was used for this study. SALK_072009, SALK_076309, and SALK_027645 were obtained from the Arabidopsis Biological
MATERIAL S AND METHODS Plant materials and growth conditions Arabidopsis ecotype Columbia (Col0) was used for this study. SALK_072009, SALK_076309, and SALK_027645 were obtained from the Arabidopsis Biological
Introduction to Assay Development
 Introduction to Assay Development A poorly designed assay can derail a drug discovery program before it gets off the ground. While traditional bench-top assays are suitable for basic research and target
Introduction to Assay Development A poorly designed assay can derail a drug discovery program before it gets off the ground. While traditional bench-top assays are suitable for basic research and target
Simultaneous Suppression of Multiple Genes by Single Transgenes. Down-Regulation of Three Unrelated Lignin Biosynthetic Genes in Tobacco 1
 Simultaneous Suppression of Multiple Genes by Single Transgenes. Down-Regulation of Three Unrelated Lignin Biosynthetic Genes in Tobacco 1 James C. Abbott 2, Abdellah Barakate, Gaelle Pinçon, Michel Legrand,
Simultaneous Suppression of Multiple Genes by Single Transgenes. Down-Regulation of Three Unrelated Lignin Biosynthetic Genes in Tobacco 1 James C. Abbott 2, Abdellah Barakate, Gaelle Pinçon, Michel Legrand,
University of Groningen. Genomic Wake-Up Call Samol, Marta
 University of Groningen Genomic Wake-Up Call Samol, Marta IMPORTANT NOTE: You are advised to consult the publisher's version (publisher's PDF) if you wish to cite from it. Please check the document version
University of Groningen Genomic Wake-Up Call Samol, Marta IMPORTANT NOTE: You are advised to consult the publisher's version (publisher's PDF) if you wish to cite from it. Please check the document version
Solutions to Quiz II
 MIT Department of Biology 7.014 Introductory Biology, Spring 2005 Solutions to 7.014 Quiz II Class Average = 79 Median = 82 Grade Range % A 90-100 27 B 75-89 37 C 59 74 25 D 41 58 7 F 0 40 2 Question 1
MIT Department of Biology 7.014 Introductory Biology, Spring 2005 Solutions to 7.014 Quiz II Class Average = 79 Median = 82 Grade Range % A 90-100 27 B 75-89 37 C 59 74 25 D 41 58 7 F 0 40 2 Question 1
Orflo Application Brief 3 /2017 GFP Transfection Efficiency Monitoring with Orflo s Moxi GO Next Generation Flow Cytometer. Introduction/Background
 Introduction/Background Cell transfection and transduction refer to an array of techniques used to introduce foreign genetic material, or cloning vectors, into cell genomes. The application of these methods
Introduction/Background Cell transfection and transduction refer to an array of techniques used to introduce foreign genetic material, or cloning vectors, into cell genomes. The application of these methods
Bio 311 Learning Objectives
 Bio 311 Learning Objectives This document outlines the learning objectives for Biol 311 (Principles of Genetics). Biol 311 is part of the BioCore within the Department of Biological Sciences; therefore,
Bio 311 Learning Objectives This document outlines the learning objectives for Biol 311 (Principles of Genetics). Biol 311 is part of the BioCore within the Department of Biological Sciences; therefore,
Supplementary Text. eqtl mapping in the Bay x Sha recombinant population.
 Supplementary Text eqtl mapping in the Bay x Sha recombinant population. Expression levels for 24,576 traits (Gene-specific Sequence Tags: GSTs, CATMA array version 2) was measured in RNA extracted from
Supplementary Text eqtl mapping in the Bay x Sha recombinant population. Expression levels for 24,576 traits (Gene-specific Sequence Tags: GSTs, CATMA array version 2) was measured in RNA extracted from
pgbkt7 Anti- Myc AH109 strain (KDa) 50
 pgbkt7 (KDa) 50 37 Anti- Myc AH109 strain Supplementary Figure 1. Protein expression of CRN and TDR in yeast. To analyse the protein expression of CRNKD and TDRKD, total proteins extracted from yeast culture
pgbkt7 (KDa) 50 37 Anti- Myc AH109 strain Supplementary Figure 1. Protein expression of CRN and TDR in yeast. To analyse the protein expression of CRNKD and TDRKD, total proteins extracted from yeast culture
Recent technology allow production of microarrays composed of 70-mers (essentially a hybrid of the two techniques)
 Microarrays and Transcript Profiling Gene expression patterns are traditionally studied using Northern blots (DNA-RNA hybridization assays). This approach involves separation of total or polya + RNA on
Microarrays and Transcript Profiling Gene expression patterns are traditionally studied using Northern blots (DNA-RNA hybridization assays). This approach involves separation of total or polya + RNA on
DNA Structure & the Genome. Bio160 General Biology
 DNA Structure & the Genome Bio160 General Biology Lecture Outline I. DNA A nucleic acid II. Chromosome Structure III. Chromosomes and Genes IV. DNA vs. RNA I. DNA A Nucleic Acid Structure of DNA: Remember:
DNA Structure & the Genome Bio160 General Biology Lecture Outline I. DNA A nucleic acid II. Chromosome Structure III. Chromosomes and Genes IV. DNA vs. RNA I. DNA A Nucleic Acid Structure of DNA: Remember:
Mathematical Biosciences
 Mathematical Biosciences 228 (2010) 78 89 Contents lists available at ScienceDirect Mathematical Biosciences journal homepage: www.elsevier.com/locate/mbs Mathematical modeling of monolignol biosynthesis
Mathematical Biosciences 228 (2010) 78 89 Contents lists available at ScienceDirect Mathematical Biosciences journal homepage: www.elsevier.com/locate/mbs Mathematical modeling of monolignol biosynthesis
Functional Genomics in Plants
 Functional Genomics in Plants Jeffrey L Bennetzen, Purdue University, West Lafayette, Indiana, USA Functional genomics refers to a suite of genetic technologies that will contribute to a comprehensive
Functional Genomics in Plants Jeffrey L Bennetzen, Purdue University, West Lafayette, Indiana, USA Functional genomics refers to a suite of genetic technologies that will contribute to a comprehensive
Chapter 15 Gene Technologies and Human Applications
 Chapter Outline Chapter 15 Gene Technologies and Human Applications Section 1: The Human Genome KEY IDEAS > Why is the Human Genome Project so important? > How do genomics and gene technologies affect
Chapter Outline Chapter 15 Gene Technologies and Human Applications Section 1: The Human Genome KEY IDEAS > Why is the Human Genome Project so important? > How do genomics and gene technologies affect
Nature Protocols: doi: /nprot Supplementary Figure 1. Efficient genome editing in human pluripotent stem cells.
 Supplementary Figure 1 Efficient genome editing in human pluripotent stem cells. Our genome editing protocol was highly efficient in an additional pluripotent human cell line (CHB10) and at several different
Supplementary Figure 1 Efficient genome editing in human pluripotent stem cells. Our genome editing protocol was highly efficient in an additional pluripotent human cell line (CHB10) and at several different
Lecture 8: Sequencing and SNP. Sept 15, 2006
 Lecture 8: Sequencing and SNP Sept 15, 2006 Announcements Random questioning during literature discussion sessions starts next week for real! Schedule changes Moved QTL lecture up Removed landscape genetics
Lecture 8: Sequencing and SNP Sept 15, 2006 Announcements Random questioning during literature discussion sessions starts next week for real! Schedule changes Moved QTL lecture up Removed landscape genetics
Learning Objectives :
 Learning Objectives : Understand the basic differences between genomic and cdna libraries Understand how genomic libraries are constructed Understand the purpose for having overlapping DNA fragments in
Learning Objectives : Understand the basic differences between genomic and cdna libraries Understand how genomic libraries are constructed Understand the purpose for having overlapping DNA fragments in
CHAPTER 20 DNA TECHNOLOGY AND GENOMICS. Section A: DNA Cloning
 Section A: DNA Cloning 1. DNA technology makes it possible to clone genes for basic research and commercial applications: an overview 2. Restriction enzymes are used to make recombinant DNA 3. Genes can
Section A: DNA Cloning 1. DNA technology makes it possible to clone genes for basic research and commercial applications: an overview 2. Restriction enzymes are used to make recombinant DNA 3. Genes can
Protocol for tissue-specific gene disruption in zebrafish
 Protocol for tissue-specific gene disruption in zebrafish Overview This protocol describes a method to inactivate genes in zebrafish in a tissue-specific manner. It can be used to analyze mosaic loss-of-function
Protocol for tissue-specific gene disruption in zebrafish Overview This protocol describes a method to inactivate genes in zebrafish in a tissue-specific manner. It can be used to analyze mosaic loss-of-function
Molecular Markers CRITFC Genetics Workshop December 9, 2014
 Molecular Markers CRITFC Genetics Workshop December 9, 2014 Molecular Markers Tools that allow us to collect information about an individual, a population, or a species Application in fisheries mating
Molecular Markers CRITFC Genetics Workshop December 9, 2014 Molecular Markers Tools that allow us to collect information about an individual, a population, or a species Application in fisheries mating
CHAPTER 4A MAKING SURE YOU VE GOT A RECOMBINANT PLASMID. CHAPTER 4A STUDENT GUIDE 2013 Amgen Foundation. All rights reserved.
 CHAPTER 4A MAKING SURE YOU VE GOT A RECOMBINANT PLASMID 55 INTRODUCTION When biologists clone a gene in order to produce human insulin, they create a recombinant plasmid that has the human insulin gene.
CHAPTER 4A MAKING SURE YOU VE GOT A RECOMBINANT PLASMID 55 INTRODUCTION When biologists clone a gene in order to produce human insulin, they create a recombinant plasmid that has the human insulin gene.
A subclass of HSP70s regulate development and abiotic stress responses in Arabidopsis thaliana
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 Journal of Plant Research A subclass of HSP70s regulate development and abiotic stress responses in Arabidopsis thaliana Linna Leng 1 Qianqian Liang
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 Journal of Plant Research A subclass of HSP70s regulate development and abiotic stress responses in Arabidopsis thaliana Linna Leng 1 Qianqian Liang
The GeneEditor TM in vitro Mutagenesis System: Site- Directed Mutagenesis Using Altered Beta-Lactamase Specificity
 Promega Notes Magazine Number 62, 1997, p. 02 The GeneEditor TM in vitro Mutagenesis System: Site- Directed Mutagenesis Using Altered Beta-Lactamase Specificity By Christine Andrews and Scott Lesley Promega
Promega Notes Magazine Number 62, 1997, p. 02 The GeneEditor TM in vitro Mutagenesis System: Site- Directed Mutagenesis Using Altered Beta-Lactamase Specificity By Christine Andrews and Scott Lesley Promega
CRISPR/Cas9 Genome Editing: Transfection Methods
 CRISPR/ Genome Editing: Transfection Methods For over 20 years Mirus Bio has developed and manufactured high performance transfection products and technologies. That expertise is now being applied to the
CRISPR/ Genome Editing: Transfection Methods For over 20 years Mirus Bio has developed and manufactured high performance transfection products and technologies. That expertise is now being applied to the
Impact of Retinoic acid induced-1 (Rai1) on Regulators of Metabolism and Adipogenesis
 Impact of Retinoic acid induced-1 (Rai1) on Regulators of Metabolism and Adipogenesis The mammalian system undergoes ~24 hour cycles known as circadian rhythms that temporally orchestrate metabolism, behavior,
Impact of Retinoic acid induced-1 (Rai1) on Regulators of Metabolism and Adipogenesis The mammalian system undergoes ~24 hour cycles known as circadian rhythms that temporally orchestrate metabolism, behavior,
Total Test Questions: 71 Levels: Grades Units of Credit: 1.0 STANDARD 1 STUDENTS WILL INVESTIGATE THE PAST, PRESENT, AND FUTURE APPLICATIONS OF
 DESCRIPTION Biotechnology is designed to create an awareness of career possibilities in the field of biotechnology. Students are introduced to diagnostic and therapeutic laboratory procedures that support
DESCRIPTION Biotechnology is designed to create an awareness of career possibilities in the field of biotechnology. Students are introduced to diagnostic and therapeutic laboratory procedures that support
Genome research in eukaryotes
 Functional Genomics Genome and EST sequencing can tell us how many POTENTIAL genes are present in the genome Proteomics can tell us about proteins and their interactions The goal of functional genomics
Functional Genomics Genome and EST sequencing can tell us how many POTENTIAL genes are present in the genome Proteomics can tell us about proteins and their interactions The goal of functional genomics
MIT Department of Biology 7.013: Introductory Biology - Spring 2005 Instructors: Professor Hazel Sive, Professor Tyler Jacks, Dr.
 MIT Department of Biology 7.01: Introductory Biology - Spring 2005 Instructors: Professor Hazel Sive, Professor Tyler Jacks, Dr. Claudette Gardel iv) Would Xba I be useful for cloning? Why or why not?
MIT Department of Biology 7.01: Introductory Biology - Spring 2005 Instructors: Professor Hazel Sive, Professor Tyler Jacks, Dr. Claudette Gardel iv) Would Xba I be useful for cloning? Why or why not?
Total Test Questions: 66 Levels: Grades Units of Credit: 1.0 STANDARD 2. Demonstrate appropriate use of personal protective devices.
 DESCRIPTION Biotechnology is designed to create an awareness of career possibilities in the field of biotechnology. Students are introduced to diagnostic and therapeutic laboratory procedures that support
DESCRIPTION Biotechnology is designed to create an awareness of career possibilities in the field of biotechnology. Students are introduced to diagnostic and therapeutic laboratory procedures that support
Supplementary Materials
 Supplementary Materials Supplementary Methods Supplementary Discussion Supplementary Figure 1 Calculated frequencies of embryo cells bearing bi-allelic alterations. Targeted indel mutations induced by
Supplementary Materials Supplementary Methods Supplementary Discussion Supplementary Figure 1 Calculated frequencies of embryo cells bearing bi-allelic alterations. Targeted indel mutations induced by
Mutations and Disease
 Mutations and Disease Objectives and lecture plan Describe what are mutations Explain how do we identify mutations Explain how and why mutant proteins lead to disease Describe what kinds of DNA mutations
Mutations and Disease Objectives and lecture plan Describe what are mutations Explain how do we identify mutations Explain how and why mutant proteins lead to disease Describe what kinds of DNA mutations
Bio11 Announcements. Ch 21: DNA Biology and Technology. DNA Functions. DNA and RNA Structure. How do DNA and RNA differ? What are genes?
 Bio11 Announcements TODAY Genetics (review) and quiz (CP #4) Structure and function of DNA Extra credit due today Next week in lab: Case study presentations Following week: Lab Quiz 2 Ch 21: DNA Biology
Bio11 Announcements TODAY Genetics (review) and quiz (CP #4) Structure and function of DNA Extra credit due today Next week in lab: Case study presentations Following week: Lab Quiz 2 Ch 21: DNA Biology
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10163 Supplementary Table 1 Efficiency of vector construction. Process wells recovered efficiency (%) Recombineering* 480 461 96 Intermediate plasmids 461 381 83 Recombineering efficiency
doi:10.1038/nature10163 Supplementary Table 1 Efficiency of vector construction. Process wells recovered efficiency (%) Recombineering* 480 461 96 Intermediate plasmids 461 381 83 Recombineering efficiency
3. human genomics clone genes associated with genetic disorders. 4. many projects generate ordered clones that cover genome
 Lectures 30 and 31 Genome analysis I. Genome analysis A. two general areas 1. structural 2. functional B. genome projects a status report 1. 1 st sequenced: several viral genomes 2. mitochondria and chloroplasts
Lectures 30 and 31 Genome analysis I. Genome analysis A. two general areas 1. structural 2. functional B. genome projects a status report 1. 1 st sequenced: several viral genomes 2. mitochondria and chloroplasts
Non-Organic-Based Isolation of Mammalian microrna using Norgen s microrna Purification Kit
 Application Note 13 RNA Sample Preparation Non-Organic-Based Isolation of Mammalian microrna using Norgen s microrna Purification Kit B. Lam, PhD 1, P. Roberts, MSc 1 Y. Haj-Ahmad, M.Sc., Ph.D 1,2 1 Norgen
Application Note 13 RNA Sample Preparation Non-Organic-Based Isolation of Mammalian microrna using Norgen s microrna Purification Kit B. Lam, PhD 1, P. Roberts, MSc 1 Y. Haj-Ahmad, M.Sc., Ph.D 1,2 1 Norgen
Highly efficient genome engineering in flowering plants ~ Development of a rapid method to knockout genes in Arabidopsis thaliana ~
 Highly efficient genome engineering in flowering plants ~ Development of a rapid method to knockout genes in Arabidopsis thaliana ~ December 5, 2016 Plant biologists at ITbM, Nagoya University have developed
Highly efficient genome engineering in flowering plants ~ Development of a rapid method to knockout genes in Arabidopsis thaliana ~ December 5, 2016 Plant biologists at ITbM, Nagoya University have developed
BIOTECHNOLOGY. Course Syllabus. Section A: Engineering Mathematics. Subject Code: BT. Course Structure. Engineering Mathematics. General Biotechnology
 BIOTECHNOLOGY Subject Code: BT Course Structure Sections/Units Section A Section B Unit 1 Unit 2 Unit 3 Unit 4 Unit 5 Unit 6 Unit 7 Section C Section D Section E Topics Engineering Mathematics General
BIOTECHNOLOGY Subject Code: BT Course Structure Sections/Units Section A Section B Unit 1 Unit 2 Unit 3 Unit 4 Unit 5 Unit 6 Unit 7 Section C Section D Section E Topics Engineering Mathematics General
ENDEXT TM Technology. Wheat Germ Premium Expression Kit. Ver 1.7. CellFree Sciences Co., Ltd.
 ENDEXT TM Technology Wheat Germ Premium Expression Kit Ver 1.7 CellFree Sciences Co., Ltd. 1. Purpose Wheat Germ Premium Expression Kit is a starter kit to ascertain if the Wheat Germ Cell-Free System
ENDEXT TM Technology Wheat Germ Premium Expression Kit Ver 1.7 CellFree Sciences Co., Ltd. 1. Purpose Wheat Germ Premium Expression Kit is a starter kit to ascertain if the Wheat Germ Cell-Free System
Figure 1: E. Coli lysate transfer using liquid handling automation
 Figure 1: E. Coli lysate transfer using liquid handling automation Figure 1 - E. coli lysate transfer using liquid handling automation. Following the manufacturer s procedures, a 96-well plate miniprep
Figure 1: E. Coli lysate transfer using liquid handling automation Figure 1 - E. coli lysate transfer using liquid handling automation. Following the manufacturer s procedures, a 96-well plate miniprep
Advances in analytical biochemistry and systems biology: Proteomics
 Advances in analytical biochemistry and systems biology: Proteomics Brett Boghigian Department of Chemical & Biological Engineering Tufts University July 29, 2005 Proteomics The basics History Current
Advances in analytical biochemistry and systems biology: Proteomics Brett Boghigian Department of Chemical & Biological Engineering Tufts University July 29, 2005 Proteomics The basics History Current
Molecular Biology of the Gene
 Molecular Biology of the Gene : where the genetic information is stored, blueprint for making proteins. RNA: Always involved in protein synthesis Macromolecules (polymers!) Monomers (units): nucleotides
Molecular Biology of the Gene : where the genetic information is stored, blueprint for making proteins. RNA: Always involved in protein synthesis Macromolecules (polymers!) Monomers (units): nucleotides
Metabolomics in Systems Biology
 Metabolomics in Systems Biology Basil J. Nikolau W.M. Keck Metabolomics Research Laboratory Iowa State University February 7, 2008 Outline What is metabolomics? Why is metabolomics important? What are
Metabolomics in Systems Biology Basil J. Nikolau W.M. Keck Metabolomics Research Laboratory Iowa State University February 7, 2008 Outline What is metabolomics? Why is metabolomics important? What are
BIOCHEMISTRY 551: BIOCHEMICAL METHODS SYLLABUS
 BIOCHEMISTRY 551: BIOCHEMICAL METHODS SYLLABUS Course Description: Biochemistry 551 is an integrated lecture, lab and seminar course that covers biochemistry-centered theory and techniques. The course
BIOCHEMISTRY 551: BIOCHEMICAL METHODS SYLLABUS Course Description: Biochemistry 551 is an integrated lecture, lab and seminar course that covers biochemistry-centered theory and techniques. The course
Lecture Four. Molecular Approaches I: Nucleic Acids
 Lecture Four. Molecular Approaches I: Nucleic Acids I. Recombinant DNA and Gene Cloning Recombinant DNA is DNA that has been created artificially. DNA from two or more sources is incorporated into a single
Lecture Four. Molecular Approaches I: Nucleic Acids I. Recombinant DNA and Gene Cloning Recombinant DNA is DNA that has been created artificially. DNA from two or more sources is incorporated into a single
The Molecular Basis of Bacterial Innate Immunity in Arabidopsis thaliana
 The Molecular Basis of Bacterial Innate Immunity in Arabidopsis thaliana Brian Staskawicz Department of Plant and Microbial Biology University of California, Berkeley Rice Model Plant-Pathogen Systems
The Molecular Basis of Bacterial Innate Immunity in Arabidopsis thaliana Brian Staskawicz Department of Plant and Microbial Biology University of California, Berkeley Rice Model Plant-Pathogen Systems
Read the question carefully before answering. Think before you write. If I can not read your handwriting, I will count the question wrong.
 Name KEY Note Total points added up to only 98 points so everyone received 2 free points to make total points 100. Biology 201 (Genetics) Exam #3 23 November 2004 Read the question carefully before answering.
Name KEY Note Total points added up to only 98 points so everyone received 2 free points to make total points 100. Biology 201 (Genetics) Exam #3 23 November 2004 Read the question carefully before answering.
Supplemental Materials and Methods
 Supplemental Materials and Methods Proteins and reagents Proteins were purified as described previously: RecA, RecQ, and SSB proteins (Harmon and Kowalczykowski 1998); RecF protein (Morimatsu and Kowalczykowski
Supplemental Materials and Methods Proteins and reagents Proteins were purified as described previously: RecA, RecQ, and SSB proteins (Harmon and Kowalczykowski 1998); RecF protein (Morimatsu and Kowalczykowski
MMG 301, Lec. 25 Mutations and Bacteriophage
 MMG 301, Lec. 25 Mutations and Bacteriophage Questions for today: 1. What are mutations and how do they form? 2. How are mutant bacteria used in research? 3. What are the general properties of bacteriophage
MMG 301, Lec. 25 Mutations and Bacteriophage Questions for today: 1. What are mutations and how do they form? 2. How are mutant bacteria used in research? 3. What are the general properties of bacteriophage
Course Overview. Interacting genes. Complementation. Complementation. February 15
 Course Overview Interacting genes http://www.erin.utoronto.ca/~w3bio/bio207/index.htm February 15 Outline Week Topic Chapter 1 Course objectives and Introduction to genetics Ch. 1 & Ch. 2 2 Human Pedigrees
Course Overview Interacting genes http://www.erin.utoronto.ca/~w3bio/bio207/index.htm February 15 Outline Week Topic Chapter 1 Course objectives and Introduction to genetics Ch. 1 & Ch. 2 2 Human Pedigrees
Genomic resources and gene/qtl discovery in cereals
 Genomic resources and gene/qtl discovery in cereals Roberto Tuberosa Dept. of Agroenvironmental Sciences & Technology University of Bologna, Italy The ABDC Congress 1-4 March 2010 Gudalajara, Mexico Outline
Genomic resources and gene/qtl discovery in cereals Roberto Tuberosa Dept. of Agroenvironmental Sciences & Technology University of Bologna, Italy The ABDC Congress 1-4 March 2010 Gudalajara, Mexico Outline
Concepts: What are RFLPs and how do they act like genetic marker loci?
 Restriction Fragment Length Polymorphisms (RFLPs) -1 Readings: Griffiths et al: 7th Edition: Ch. 12 pp. 384-386; Ch.13 pp404-407 8th Edition: pp. 364-366 Assigned Problems: 8th Ch. 11: 32, 34, 38-39 7th
Restriction Fragment Length Polymorphisms (RFLPs) -1 Readings: Griffiths et al: 7th Edition: Ch. 12 pp. 384-386; Ch.13 pp404-407 8th Edition: pp. 364-366 Assigned Problems: 8th Ch. 11: 32, 34, 38-39 7th
ab Methylated DNA Quantification Kit (Colorimetric)
 ab117128 Methylated DNA Quantification Kit (Colorimetric) Instructions for Use For the measurement of global DNA methylation status using DNA isolated from any species such as mammalians, plants, fungi,
ab117128 Methylated DNA Quantification Kit (Colorimetric) Instructions for Use For the measurement of global DNA methylation status using DNA isolated from any species such as mammalians, plants, fungi,
Midterm 1 Results. Midterm 1 Akey/ Fields Median Number of Students. Exam Score
 Midterm 1 Results 10 Midterm 1 Akey/ Fields Median - 69 8 Number of Students 6 4 2 0 21 26 31 36 41 46 51 56 61 66 71 76 81 86 91 96 101 Exam Score Quick review of where we left off Parental type: the
Midterm 1 Results 10 Midterm 1 Akey/ Fields Median - 69 8 Number of Students 6 4 2 0 21 26 31 36 41 46 51 56 61 66 71 76 81 86 91 96 101 Exam Score Quick review of where we left off Parental type: the
Multiple choice questions (numbers in brackets indicate the number of correct answers)
 1 Multiple choice questions (numbers in brackets indicate the number of correct answers) February 1, 2013 1. Ribose is found in Nucleic acids Proteins Lipids RNA DNA (2) 2. Most RNA in cells is transfer
1 Multiple choice questions (numbers in brackets indicate the number of correct answers) February 1, 2013 1. Ribose is found in Nucleic acids Proteins Lipids RNA DNA (2) 2. Most RNA in cells is transfer
Rapid Learning Center Presents. Teach Yourself AP Biology in 24 Hours
 Rapid Learning Center Chemistry :: Biology :: Physics :: Math Rapid Learning Center Presents Teach Yourself AP Biology in 24 Hours 1/35 *AP is a registered trademark of the College Board, which does not
Rapid Learning Center Chemistry :: Biology :: Physics :: Math Rapid Learning Center Presents Teach Yourself AP Biology in 24 Hours 1/35 *AP is a registered trademark of the College Board, which does not
ON-CHIP AMPLIFICATION OF GENOMIC DNA WITH SHORT TANDEM REPEAT AND SINGLE NUCLEOTIDE POLYMORPHISM ANALYSIS
 ON-CHIP AMPLIFICATION OF GENOMIC DNA WITH SHORT TANDEM REPEAT AND SINGLE NUCLEOTIDE POLYMORPHISM ANALYSIS David Canter, Di Wu, Tamara Summers, Jeff Rogers, Karen Menge, Ray Radtkey, and Ron Sosnowski Nanogen,
ON-CHIP AMPLIFICATION OF GENOMIC DNA WITH SHORT TANDEM REPEAT AND SINGLE NUCLEOTIDE POLYMORPHISM ANALYSIS David Canter, Di Wu, Tamara Summers, Jeff Rogers, Karen Menge, Ray Radtkey, and Ron Sosnowski Nanogen,
CHAPTER 9 DNA Technologies
 CHAPTER 9 DNA Technologies Recombinant DNA Artificially created DNA that combines sequences that do not occur together in the nature Basis of much of the modern molecular biology Molecular cloning of genes
CHAPTER 9 DNA Technologies Recombinant DNA Artificially created DNA that combines sequences that do not occur together in the nature Basis of much of the modern molecular biology Molecular cloning of genes
Introduction to Bioinformatics and Gene Expression Technology
 Vocabulary Introduction to Bioinformatics and Gene Expression Technology Utah State University Spring 2014 STAT 5570: Statistical Bioinformatics Notes 1.1 Gene: Genetics: Genome: Genomics: hereditary DNA
Vocabulary Introduction to Bioinformatics and Gene Expression Technology Utah State University Spring 2014 STAT 5570: Statistical Bioinformatics Notes 1.1 Gene: Genetics: Genome: Genomics: hereditary DNA
Do engineered ips and ES cells have similar molecular signatures?
 Do engineered ips and ES cells have similar molecular signatures? Comparing expression and epigenetics in stem cells George Bell, Ph.D. Bioinformatics and Research Computing 2012 Spring Lecture Series
Do engineered ips and ES cells have similar molecular signatures? Comparing expression and epigenetics in stem cells George Bell, Ph.D. Bioinformatics and Research Computing 2012 Spring Lecture Series
Module I: Introduction
 Module I: Introduction 20.109 Lecture 1 3 February, 2011 Introduction to: Module Overview Fundamental concepts and techniques in molecular biology Appreciating nucleic acids (RNA in particular) as more
Module I: Introduction 20.109 Lecture 1 3 February, 2011 Introduction to: Module Overview Fundamental concepts and techniques in molecular biology Appreciating nucleic acids (RNA in particular) as more
Protocol for cloning SEC-based repair templates using Gibson assembly and ccdb negative selection
 Protocol for cloning SEC-based repair templates using Gibson assembly and ccdb negative selection Written by Dan Dickinson (daniel.dickinson@austin.utexas.edu) and last updated January 2018. A version
Protocol for cloning SEC-based repair templates using Gibson assembly and ccdb negative selection Written by Dan Dickinson (daniel.dickinson@austin.utexas.edu) and last updated January 2018. A version
Chapter 20: Biotechnology
 Name Period The AP Biology exam has reached into this chapter for essay questions on a regular basis over the past 15 years. Student responses show that biotechnology is a difficult topic. This chapter
Name Period The AP Biology exam has reached into this chapter for essay questions on a regular basis over the past 15 years. Student responses show that biotechnology is a difficult topic. This chapter
