ChIP-seq/Functional Genomics/Epigenomics. CBSU/3CPG/CVG Next-Gen Sequencing Workshop. Josh Waterfall. March 31, 2010
|
|
|
- Gordon Gilbert
- 6 years ago
- Views:
Transcription
1 ChIP-seq/Functional Genomics/Epigenomics CBSU/3CPG/CVG Next-Gen Sequencing Workshop Josh Waterfall March 31, 2010
2 Outline Introduction to ChIP-seq Control data sets Peak/enriched region identification Related functional genomics assays Useful web resources
3 Chromatin ImmunoPrecipitation (ChIP-seq) Park, P.J., Nat. Genetics (2009) 10:
4 Advantages over array-based methods No cross-hybridization background Lower end of sensitivity largely dependent on just sequencing depth More linear, quantitative assay Unmappable portion of genome is distinct from, and much smaller than, repeat masked portion No limit based on probe locations Needs less starting material Higher resolution
5 Important Issues: Appropriate Controls Identifying Enriched Regions
6 Controls Sequencing is such high-sensitivity that signals invisible in any other assay are now apparent. Need rigorous controls to be confident of enrichment. Input DNA has non-random pattern (open chromatin shears more easily) - Sono-seq is an actual assay. Mock-IP controls for more steps in ChIP protocol than input DNA but not antibody cross-reactivity. Different antibodies (to different epitopes) in separate experiments, or ChIP after target protein has been depleted (or in cell-line without tagged protein), help control for cross-reactivity. To characterize new antibody, IP and mass-spec everything that comes down to verify only expected binding partners are seen.
7 Background rate is non-uniform Auerbach R K et al. PNAS 2009;106:
8 Identifying enriched regions Identifying narrow ChIP peaks is very different from: identifying broadly enriched regions in ChIP. identifying narrow peaks from other assays. Many well-worked out programs for identifying narrow ChIP peaks (e.g. from sequence specific binding factors). The best programs can exploit strand-specific patterns, local versus global background levels, mappability, etc. Other assays also result in localized peaks but without same strand pattern. Most work on broad regions (e.g. particular histone modifications or RNA polymerase) is based on sliding windows (either of fixed length or fixed read count).
9 Strand specific pattern for localized peaks in ChIP-seq Park, P.J., Nat. Genetics (2009) 10:
10 Peak Detection Software Is strand information used? If fragment size is used, is it defined by user or estimated from data? (How) is control data incorporated? Is background defined locally or globally? (How) are unmappable regions treated? ChIP-seq peak calling software contest on Pepke, S., Wold, B., Mortazavi, A. (2009) Nat. Methods 6:S22-S32
11 Assessing saturation and significance Saturation is dependent on both ChIP efficiency (globally) and factor:dna affinity (locally) Significance depends on both ChIP and control read counts Park, P.J., Nat. Genetics (2009) 10:
12 ChIP-seq read density is quantitative measure of binding level Jothi, R., et. al. (2008) Nucleic Acids Res. 36:
13 Solutions to GC bias Improve Illumina protocol (especially gel purification) Use single molecule sequencing technology Quail, MA, et. al. (2008) Nat. Methods 5: Goren, A, et. al. (2009) Nat. Methods 7:47-49
14 Fragmenting DNA AFA (Adaptive Focused Acoustics)/Covaris enzyme (MNase, NEB Fragmentase) Nebulization Sonication Quality Evaluation with Agilent Bioanalyzer Quail, MA, et. al. (2008) Nat. Methods 5:
15 Fragmenting DNA Open chromatin fragments more easily than closed chromatin - ChIP for factors associated with transcriptional repression or heterochromatin can be more difficult than for factors associated with transcriptional activation. Grewal, SI, Elgin, SCR (2007) Nature 447:
16 Related Functional Genomic Assays Chromatin ImmunoPrecipitation (ChIP) DNAse I Hypersensitivity Formaldehyde Assisted Isolation of Regulatory Elements DNA Methylation (bisulfite, affinity, or restriction enzyme based) Cap Analysis of Gene Expression (CAGE) Genomic Run On (GRO-seq) Many more Each can couple to next-gen sequencing and entails analysis/identification of enriched regions
17
18 Formaldehyde Assisted Isolation of Regulatory Elements (FAIRE) Gaulton, K.J., et. al. (2010) Nat. Genetics 42:
19 Micrococcal Nuclease digestion (MNase-seq) Weiner A et al. Genome Res. 2010;20:90-100
20 DNA methylation 3 broad classes of assays: Enzyme based methylation sensitive restriction enzymes Affinity based antibodies or other medna binding proteins used in ChIP-like experiment Bisulfite based NaHSO 3 deaminates unmethylated cytosines to uracils but does not affect 5-methylcytosine. Reduced Representation based on enzymatic digest or hybridization enrichment common for cost efficiency in large, sparsely methylated (e.g. mammalian) genomes. Read alignments done to in silico bisulfite-converted genome. Direct sequencing of 5th base (future technologies)
21 Cap Analysis of Gene Expression (CAGE)
22 Genomic Run On (GRO-seq) Core, LJ, Waterfall, JJ, Lis, JT (2008) Science 322:1845-8
23 Web resources for analyzing, viewing, sharing, and collecting genomics data UCSC Genome Browser ( Galaxy ( NCBI GEO (
24 The UCSC Genome Browser
25 Uploading data to UCSC
26 Uploading data to UCSC
27 Uploading data to UCSC
28 Modifying display settings
29 Modifying display settings
30 Using publicly available tracks
31 Using publicly available tracks
32 Galaxy
33 NCBI Gene Expression Omnibus (GEO)
34 ChIP-seq exercise Use MACS software to call peaks in STAT1 ChIP-seq experiment in human HeLA S3 cells after inteferon-γ stimulation. Analyze diagnostics of run and upload data to UCSC genome browser to look at results. Office hour: Friday, April 2 1pm - 2pm 102 Weill conference room
Complete Sample to Analysis Solutions for DNA Methylation Discovery using Next Generation Sequencing
 Complete Sample to Analysis Solutions for DNA Methylation Discovery using Next Generation Sequencing SureSelect Human/Mouse Methyl-Seq Kyeong Jeong PhD February 5, 2013 CAG EMEAI DGG/GSD/GFO Agilent Restricted
Complete Sample to Analysis Solutions for DNA Methylation Discovery using Next Generation Sequencing SureSelect Human/Mouse Methyl-Seq Kyeong Jeong PhD February 5, 2013 CAG EMEAI DGG/GSD/GFO Agilent Restricted
Impact of gdna Integrity on the Outcome of DNA Methylation Studies
 Impact of gdna Integrity on the Outcome of DNA Methylation Studies Application Note Nucleic Acid Analysis Authors Emily Putnam, Keith Booher, and Xueguang Sun Zymo Research Corporation, Irvine, CA, USA
Impact of gdna Integrity on the Outcome of DNA Methylation Studies Application Note Nucleic Acid Analysis Authors Emily Putnam, Keith Booher, and Xueguang Sun Zymo Research Corporation, Irvine, CA, USA
Introduction to ChIP Seq data analyses. Acknowledgement: slides taken from Dr. H
 Introduction to ChIP Seq data analyses Acknowledgement: slides taken from Dr. H Wu @Emory ChIP seq: Chromatin ImmunoPrecipitation it ti + sequencing Same biological motivation as ChIP chip: measure specific
Introduction to ChIP Seq data analyses Acknowledgement: slides taken from Dr. H Wu @Emory ChIP seq: Chromatin ImmunoPrecipitation it ti + sequencing Same biological motivation as ChIP chip: measure specific
ChIP-Seq Data Analysis. J Fass UCD Genome Center Bioinformatics Core Wednesday 15 June 2015
 ChIP-Seq Data Analysis J Fass UCD Genome Center Bioinformatics Core Wednesday 15 June 2015 What s the Question? Where do Transcription Factors (TFs) bind genomic DNA 1? (Where do other things bind DNA
ChIP-Seq Data Analysis J Fass UCD Genome Center Bioinformatics Core Wednesday 15 June 2015 What s the Question? Where do Transcription Factors (TFs) bind genomic DNA 1? (Where do other things bind DNA
The ChIP-Seq project. Giovanna Ambrosini, Philipp Bucher. April 19, 2010 Lausanne. EPFL-SV Bucher Group
 The ChIP-Seq project Giovanna Ambrosini, Philipp Bucher EPFL-SV Bucher Group April 19, 2010 Lausanne Overview Focus on technical aspects Description of applications (C programs) Where to find binaries,
The ChIP-Seq project Giovanna Ambrosini, Philipp Bucher EPFL-SV Bucher Group April 19, 2010 Lausanne Overview Focus on technical aspects Description of applications (C programs) Where to find binaries,
Introduction to genome biology
 Introduction to genome biology Lisa Stubbs We ve found most genes; but what about the rest of the genome? Genome size* 12 Mb 95 Mb 170 Mb 1500 Mb 2700 Mb 3200 Mb #coding genes ~7000 ~20000 ~14000 ~26000
Introduction to genome biology Lisa Stubbs We ve found most genes; but what about the rest of the genome? Genome size* 12 Mb 95 Mb 170 Mb 1500 Mb 2700 Mb 3200 Mb #coding genes ~7000 ~20000 ~14000 ~26000
Introductory Next Gen Workshop
 Introductory Next Gen Workshop http://www.illumina.ucr.edu/ http://www.genomics.ucr.edu/ Workshop Objectives Workshop aimed at those who are new to Illumina sequencing and will provide: - a basic overview
Introductory Next Gen Workshop http://www.illumina.ucr.edu/ http://www.genomics.ucr.edu/ Workshop Objectives Workshop aimed at those who are new to Illumina sequencing and will provide: - a basic overview
Assay Standards Working Group Nov 2012 Assay Standards Working Group Recommendations, November 2012
 Assay Standards Working Group Recommendations, November 2012 Contents Assay Standards Working Group Recommendations, August 2012... 1 Contents... 1 Introduction... 2 1: Reference Epigenome Criteria...
Assay Standards Working Group Recommendations, November 2012 Contents Assay Standards Working Group Recommendations, August 2012... 1 Contents... 1 Introduction... 2 1: Reference Epigenome Criteria...
Data and Metadata Models Recommendations Version 1.2 Developed by the IHEC Metadata Standards Workgroup
 Data and Metadata Models Recommendations Version 1.2 Developed by the IHEC Metadata Standards Workgroup 1. Introduction The data produced by IHEC is illustrated in Figure 1. Figure 1. The space of epigenomic
Data and Metadata Models Recommendations Version 1.2 Developed by the IHEC Metadata Standards Workgroup 1. Introduction The data produced by IHEC is illustrated in Figure 1. Figure 1. The space of epigenomic
Identifying and mitigating bias in next-generation sequencing methods for chromatin biology
 STUDY DESIGNS Identifying and mitigating bias in next-generation sequencing methods for chromatin biology Clifford A. Meyer and X. Shirley Liu Abstract Next-generation sequencing (NGS) technologies have
STUDY DESIGNS Identifying and mitigating bias in next-generation sequencing methods for chromatin biology Clifford A. Meyer and X. Shirley Liu Abstract Next-generation sequencing (NGS) technologies have
Bioinformatics of Transcriptional Regulation
 Bioinformatics of Transcriptional Regulation Carl Herrmann IPMB & DKFZ c.herrmann@dkfz.de Wechselwirkung von Maßnahmen und Auswirkungen Einflussmöglichkeiten in einem Dialog From genes to active compounds
Bioinformatics of Transcriptional Regulation Carl Herrmann IPMB & DKFZ c.herrmann@dkfz.de Wechselwirkung von Maßnahmen und Auswirkungen Einflussmöglichkeiten in einem Dialog From genes to active compounds
Gene Expression Technology
 Gene Expression Technology Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Gene expression Gene expression is the process by which information from a gene
Gene Expression Technology Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Gene expression Gene expression is the process by which information from a gene
Green Center Computational Core ChIP- Seq Pipeline, Just a Click Away
 Green Center Computational Core ChIP- Seq Pipeline, Just a Click Away Venkat Malladi Computational Biologist Computational Core Cecil H. and Ida Green Center for Reproductive Biology Science Introduc
Green Center Computational Core ChIP- Seq Pipeline, Just a Click Away Venkat Malladi Computational Biologist Computational Core Cecil H. and Ida Green Center for Reproductive Biology Science Introduc
PeakSeq enables systematic scoring of ChIP-seq experiments relative to controls
 PeakSeq enables systematic scoring of ChIP-seq experiments relative to controls Joel Rozowsky, Ghia Euskirchen 2, Raymond K Auerbach 3, Zhengdong D Zhang, Theodore Gibson, Robert Bjornson 4, Nicholas Carriero
PeakSeq enables systematic scoring of ChIP-seq experiments relative to controls Joel Rozowsky, Ghia Euskirchen 2, Raymond K Auerbach 3, Zhengdong D Zhang, Theodore Gibson, Robert Bjornson 4, Nicholas Carriero
DNA. bioinformatics. epigenetics methylation structural variation. custom. assembly. gene. tumor-normal. mendelian. BS-seq. prediction.
 Epigenomics T TM activation SNP target ncrna validation metagenomics genetics private RRBS-seq de novo trio RIP-seq exome mendelian comparative genomics DNA NGS ChIP-seq bioinformatics assembly tumor-normal
Epigenomics T TM activation SNP target ncrna validation metagenomics genetics private RRBS-seq de novo trio RIP-seq exome mendelian comparative genomics DNA NGS ChIP-seq bioinformatics assembly tumor-normal
What we ll do today. Types of stem cells. Do engineered ips and ES cells have. What genes are special in stem cells?
 Do engineered ips and ES cells have similar molecular signatures? What we ll do today Research questions in stem cell biology Comparing expression and epigenetics in stem cells asuring gene expression
Do engineered ips and ES cells have similar molecular signatures? What we ll do today Research questions in stem cell biology Comparing expression and epigenetics in stem cells asuring gene expression
Do engineered ips and ES cells have similar molecular signatures?
 Do engineered ips and ES cells have similar molecular signatures? Comparing expression and epigenetics in stem cells George Bell, Ph.D. Bioinformatics and Research Computing 2012 Spring Lecture Series
Do engineered ips and ES cells have similar molecular signatures? Comparing expression and epigenetics in stem cells George Bell, Ph.D. Bioinformatics and Research Computing 2012 Spring Lecture Series
B&DA Committee Bioinformatics and Data Analysis. PAG January 2016
 B&DA Committee Bioinformatics and Data Analysis PAG January 2016 B&DA progress so far Aim: to define standard bfx pipelines for FAANG data Group has skyped many times We have a Wiki: hosted by EBI Sub-groups
B&DA Committee Bioinformatics and Data Analysis PAG January 2016 B&DA progress so far Aim: to define standard bfx pipelines for FAANG data Group has skyped many times We have a Wiki: hosted by EBI Sub-groups
ab ChIP Kit Magnetic One-Step
 ab156907 ChIP Kit Magnetic One-Step Instructions for Use For selective enrichment of a chromatin fraction containing specific DNA sequences in a high throughput format using chromatin isolated from various
ab156907 ChIP Kit Magnetic One-Step Instructions for Use For selective enrichment of a chromatin fraction containing specific DNA sequences in a high throughput format using chromatin isolated from various
Regulation of eukaryotic transcription:
 Promoter definition by mass genome annotation data: in silico primer extension EMBNET course Bioinformatics of transcriptional regulation Jan 28 2008 Christoph Schmid Regulation of eukaryotic transcription:
Promoter definition by mass genome annotation data: in silico primer extension EMBNET course Bioinformatics of transcriptional regulation Jan 28 2008 Christoph Schmid Regulation of eukaryotic transcription:
Le proteine regolative variano nei vari tipi cellulari e in funzione degli stimoli ambientali
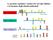 Le proteine regolative variano nei vari tipi cellulari e in funzione degli stimoli ambientali Tipo cellulare 1 Tipo cellulare 2 Tipo cellulare 3 DNA-protein Crosslink Lisi Frammentazione Immunopurificazione
Le proteine regolative variano nei vari tipi cellulari e in funzione degli stimoli ambientali Tipo cellulare 1 Tipo cellulare 2 Tipo cellulare 3 DNA-protein Crosslink Lisi Frammentazione Immunopurificazione
PIP-seq. Cells. Permanganate ChIP-Seq
 PIP-seq ells Formaldehyde Permanganate 5 Harvest Lyse Sonicate First dapter Ligation 3 3 5 hip Elute Reverse rosslinks Piperidine cleavage 5 3 3 5 Primer Extension Second dapter Ligation 5 3 3 5 Deep Sequencing
PIP-seq ells Formaldehyde Permanganate 5 Harvest Lyse Sonicate First dapter Ligation 3 3 5 hip Elute Reverse rosslinks Piperidine cleavage 5 3 3 5 Primer Extension Second dapter Ligation 5 3 3 5 Deep Sequencing
Bioinformatics Advice on Experimental Design
 Bioinformatics Advice on Experimental Design Where do I start? Please refer to the following guide to better plan your experiments for good statistical analysis, best suited for your research needs. Statistics
Bioinformatics Advice on Experimental Design Where do I start? Please refer to the following guide to better plan your experiments for good statistical analysis, best suited for your research needs. Statistics
Microarrays: since we use probes we obviously must know the sequences we are looking at!
 These background are needed: 1. - Basic Molecular Biology & Genetics DNA replication Transcription Post-transcriptional RNA processing Translation Post-translational protein modification Gene expression
These background are needed: 1. - Basic Molecular Biology & Genetics DNA replication Transcription Post-transcriptional RNA processing Translation Post-translational protein modification Gene expression
ANTIBODY PERFORMANCE COMPARISON. Histone Modifications
 ANTIBODY PERFORMANCE COMPARISON Histone Modifications COVER IMAGE: Methylation of cytosine bases in regions called CpG islands is a hallmark of transcriptionally repressed heterochromatin. These methylated
ANTIBODY PERFORMANCE COMPARISON Histone Modifications COVER IMAGE: Methylation of cytosine bases in regions called CpG islands is a hallmark of transcriptionally repressed heterochromatin. These methylated
Whole Transcriptome Analysis of Illumina RNA- Seq Data. Ryan Peters Field Application Specialist
 Whole Transcriptome Analysis of Illumina RNA- Seq Data Ryan Peters Field Application Specialist Partek GS in your NGS Pipeline Your Start-to-Finish Solution for Analysis of Next Generation Sequencing Data
Whole Transcriptome Analysis of Illumina RNA- Seq Data Ryan Peters Field Application Specialist Partek GS in your NGS Pipeline Your Start-to-Finish Solution for Analysis of Next Generation Sequencing Data
1. Cross-linking and cell harvesting
 ChIP is a powerful tool that allows the specific matching of proteins or histone modifications to regions of the genome. Chromatin is isolated and antibodies to the antigen of interest are used to determine
ChIP is a powerful tool that allows the specific matching of proteins or histone modifications to regions of the genome. Chromatin is isolated and antibodies to the antigen of interest are used to determine
Introduction Bioo Scientific
 Next Generation Sequencing Catalog 2014-2015 Introduction Bioo Scientific Bioo Scientific is a global life science company headquartered in Austin, TX, committed to providing innovative products and superior
Next Generation Sequencing Catalog 2014-2015 Introduction Bioo Scientific Bioo Scientific is a global life science company headquartered in Austin, TX, committed to providing innovative products and superior
qpcr Quantitative PCR or Real-time PCR Gives a measurement of PCR product at end of each cycle real time
 qpcr qpcr Quantitative PCR or Real-time PCR Gives a measurement of PCR product at end of each cycle real time Differs from endpoint PCR gel on last cycle Used to determines relative amount of template
qpcr qpcr Quantitative PCR or Real-time PCR Gives a measurement of PCR product at end of each cycle real time Differs from endpoint PCR gel on last cycle Used to determines relative amount of template
The ENCODE Encyclopedia. & Variant Annotation Using RegulomeDB and HaploReg
 The ENCODE Encyclopedia & Variant Annotation Using RegulomeDB and HaploReg Jill E. Moore Weng Lab University of Massachusetts Medical School October 10, 2015 Where s the Encyclopedia? ENCODE: Encyclopedia
The ENCODE Encyclopedia & Variant Annotation Using RegulomeDB and HaploReg Jill E. Moore Weng Lab University of Massachusetts Medical School October 10, 2015 Where s the Encyclopedia? ENCODE: Encyclopedia
Intracellular receptors specify complex patterns of gene expression that are cell and gene
 SUPPLEMENTAL RESULTS AND DISCUSSION Some HPr-1AR ARE-containing Genes Are Unresponsive to Androgen Intracellular receptors specify complex patterns of gene expression that are cell and gene specific. For
SUPPLEMENTAL RESULTS AND DISCUSSION Some HPr-1AR ARE-containing Genes Are Unresponsive to Androgen Intracellular receptors specify complex patterns of gene expression that are cell and gene specific. For
Division Ave. High School AP Biology
 Control of Eukaryotic Genes 2007-2008 The BIG Questions n How are genes turned on & off in eukaryotes? n How do cells with the same genes differentiate to perform completely different, specialized functions?
Control of Eukaryotic Genes 2007-2008 The BIG Questions n How are genes turned on & off in eukaryotes? n How do cells with the same genes differentiate to perform completely different, specialized functions?
EpiNext High-Sensitivity Bisulfite-Seq Kit (Illumina)
 EpiNext High-Sensitivity Bisulfite-Seq Kit (Illumina) Base Catalog # PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE Uses: The EpiNext High-Sensitivity Bisulfite-Seq Kit is designed to prepare bisulfite-converted
EpiNext High-Sensitivity Bisulfite-Seq Kit (Illumina) Base Catalog # PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE Uses: The EpiNext High-Sensitivity Bisulfite-Seq Kit is designed to prepare bisulfite-converted
Chromatin and Transcription
 Chromatin and Transcription Chromatin Structure Chromatin Represses Transcription Nucleosome Positioning Histone Acetylation Chromatin Remodeling Histone Methylation CHIP Analysis Chromatin and Elongation
Chromatin and Transcription Chromatin Structure Chromatin Represses Transcription Nucleosome Positioning Histone Acetylation Chromatin Remodeling Histone Methylation CHIP Analysis Chromatin and Elongation
Supplementary Table 1: Oligo designs. A list of ATAC-seq oligos used for PCR.
 Ad1_noMX: Ad2.1_TAAGGCGA Ad2.2_CGTACTAG Ad2.3_AGGCAGAA Ad2.4_TCCTGAGC Ad2.5_GGACTCCT Ad2.6_TAGGCATG Ad2.7_CTCTCTAC Ad2.8_CAGAGAGG Ad2.9_GCTACGCT Ad2.10_CGAGGCTG Ad2.11_AAGAGGCA Ad2.12_GTAGAGGA Ad2.13_GTCGTGAT
Ad1_noMX: Ad2.1_TAAGGCGA Ad2.2_CGTACTAG Ad2.3_AGGCAGAA Ad2.4_TCCTGAGC Ad2.5_GGACTCCT Ad2.6_TAGGCATG Ad2.7_CTCTCTAC Ad2.8_CAGAGAGG Ad2.9_GCTACGCT Ad2.10_CGAGGCTG Ad2.11_AAGAGGCA Ad2.12_GTAGAGGA Ad2.13_GTCGTGAT
Gene Regulation Solutions. Microarrays and Next-Generation Sequencing
 Gene Regulation Solutions Microarrays and Next-Generation Sequencing Gene Regulation Solutions The Microarrays Advantage Microarrays Lead the Industry in: Comprehensive Content SurePrint G3 Human Gene
Gene Regulation Solutions Microarrays and Next-Generation Sequencing Gene Regulation Solutions The Microarrays Advantage Microarrays Lead the Industry in: Comprehensive Content SurePrint G3 Human Gene
AP Biology Gene Expression/Biotechnology REVIEW
 AP Biology Gene Expression/Biotechnology REVIEW Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Gene expression can be a. regulated before transcription.
AP Biology Gene Expression/Biotechnology REVIEW Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Gene expression can be a. regulated before transcription.
CHAPTERS , 17: Eukaryotic Genetics
 CHAPTERS 14.1 14.6, 17: Eukaryotic Genetics 1. Review the levels of DNA packing within the eukaryote nucleus. Label each level. (A similar diagram is on pg 188 of your textbook.) 2. How do the coding regions
CHAPTERS 14.1 14.6, 17: Eukaryotic Genetics 1. Review the levels of DNA packing within the eukaryote nucleus. Label each level. (A similar diagram is on pg 188 of your textbook.) 2. How do the coding regions
Enhancers mutations that make the original mutant phenotype more extreme. Suppressors mutations that make the original mutant phenotype less extreme
 Interactomics and Proteomics 1. Interactomics The field of interactomics is concerned with interactions between genes or proteins. They can be genetic interactions, in which two genes are involved in the
Interactomics and Proteomics 1. Interactomics The field of interactomics is concerned with interactions between genes or proteins. They can be genetic interactions, in which two genes are involved in the
CM581A2: NEXT GENERATION SEQUENCING PLATFORMS AND LIBRARY GENERATION
 CM581A2: NEXT GENERATION SEQUENCING PLATFORMS AND LIBRARY GENERATION Fall 2015 Instructors: Coordinator: Carol Wilusz, Associate Professor MIP, CMB Instructor: Dan Sloan, Assistant Professor, Biology,
CM581A2: NEXT GENERATION SEQUENCING PLATFORMS AND LIBRARY GENERATION Fall 2015 Instructors: Coordinator: Carol Wilusz, Associate Professor MIP, CMB Instructor: Dan Sloan, Assistant Professor, Biology,
Non-Organic-Based Isolation of Mammalian microrna using Norgen s microrna Purification Kit
 Application Note 13 RNA Sample Preparation Non-Organic-Based Isolation of Mammalian microrna using Norgen s microrna Purification Kit B. Lam, PhD 1, P. Roberts, MSc 1 Y. Haj-Ahmad, M.Sc., Ph.D 1,2 1 Norgen
Application Note 13 RNA Sample Preparation Non-Organic-Based Isolation of Mammalian microrna using Norgen s microrna Purification Kit B. Lam, PhD 1, P. Roberts, MSc 1 Y. Haj-Ahmad, M.Sc., Ph.D 1,2 1 Norgen
GENE REGULATION slide shows by Kim Foglia modified Slides with blue edges are Kim s
 GENE REGULATION slide shows by Kim Foglia modified Slides with blue edges are Kim s 2007-2008 Bacterial metabolism Bacteria need to respond quickly to changes in their environment STOP GO if they have
GENE REGULATION slide shows by Kim Foglia modified Slides with blue edges are Kim s 2007-2008 Bacterial metabolism Bacteria need to respond quickly to changes in their environment STOP GO if they have
Technical tips Session 5
 Technical tips Session 5 Chromatine Immunoprecipitation (ChIP): This is a powerful in vivo method to quantitate interaction of proteins associated with specific regions of the genome. It involves the immunoprecipitation
Technical tips Session 5 Chromatine Immunoprecipitation (ChIP): This is a powerful in vivo method to quantitate interaction of proteins associated with specific regions of the genome. It involves the immunoprecipitation
Fatchiyah
 Fatchiyah Email: fatchiya@yahoo.co.id RNAs: mrna trna rrna RNAi DNAs: Protein: genome DNA cdna mikro-makro mono-poly single-multi Analysis: Identification human and animal disease Finger printing Sexing
Fatchiyah Email: fatchiya@yahoo.co.id RNAs: mrna trna rrna RNAi DNAs: Protein: genome DNA cdna mikro-makro mono-poly single-multi Analysis: Identification human and animal disease Finger printing Sexing
ab Post-Bisulfite DNA Library Preparation Kit (For Illumina )
 ab185906 Post-Bisulfite DNA Library Preparation Kit (For Illumina ) Instructions for Use For the preparation of a DNA library for various Illumina platform-based bisulfite sequencing (bisulfite-seq) assays.
ab185906 Post-Bisulfite DNA Library Preparation Kit (For Illumina ) Instructions for Use For the preparation of a DNA library for various Illumina platform-based bisulfite sequencing (bisulfite-seq) assays.
Activation of a Floral Homeotic Gene in Arabidopsis
 Activation of a Floral Homeotic Gene in Arabidopsis By Maximiliam A. Busch, Kirsten Bomblies, and Detlef Weigel Presentation by Lis Garrett and Andrea Stevenson http://ucsdnews.ucsd.edu/archive/graphics/images/image5.jpg
Activation of a Floral Homeotic Gene in Arabidopsis By Maximiliam A. Busch, Kirsten Bomblies, and Detlef Weigel Presentation by Lis Garrett and Andrea Stevenson http://ucsdnews.ucsd.edu/archive/graphics/images/image5.jpg
DNA Microarray Technology
 CHAPTER 1 DNA Microarray Technology All living organisms are composed of cells. As a functional unit, each cell can make copies of itself, and this process depends on a proper replication of the genetic
CHAPTER 1 DNA Microarray Technology All living organisms are composed of cells. As a functional unit, each cell can make copies of itself, and this process depends on a proper replication of the genetic
Chapter 24: Promoters and Enhancers
 Chapter 24: Promoters and Enhancers A typical gene transcribed by RNA polymerase II has a promoter that usually extends upstream from the site where transcription is initiated the (#1) of transcription
Chapter 24: Promoters and Enhancers A typical gene transcribed by RNA polymerase II has a promoter that usually extends upstream from the site where transcription is initiated the (#1) of transcription
#9005. SimpleChIP Plus Enzymatic Chromatin IP Kit (Magnetic Beads) n 1 Kit. (30 Immunoprecipitations) Store at RT, 4 C, and 20 C
 Store at RT, 4 C, and 20 C #9005 SimpleChIP Plus Enzymatic Chromatin IP Kit (Magnetic Beads) 3 n 1 Kit (30 Immunoprecipitations) rev. 09/05/17 Orders n 877-616-CELL (2355) orders@cellsignal.com Support
Store at RT, 4 C, and 20 C #9005 SimpleChIP Plus Enzymatic Chromatin IP Kit (Magnetic Beads) 3 n 1 Kit (30 Immunoprecipitations) rev. 09/05/17 Orders n 877-616-CELL (2355) orders@cellsignal.com Support
Lecture Four. Molecular Approaches I: Nucleic Acids
 Lecture Four. Molecular Approaches I: Nucleic Acids I. Recombinant DNA and Gene Cloning Recombinant DNA is DNA that has been created artificially. DNA from two or more sources is incorporated into a single
Lecture Four. Molecular Approaches I: Nucleic Acids I. Recombinant DNA and Gene Cloning Recombinant DNA is DNA that has been created artificially. DNA from two or more sources is incorporated into a single
Targeted Sequencing Using Droplet-Based Microfluidics. Keith Brown Director, Sales
 Targeted Sequencing Using Droplet-Based Microfluidics Keith Brown Director, Sales brownk@raindancetech.com Who we are: is a Provider of Microdroplet-based Solutions The Company s RainStorm TM Technology
Targeted Sequencing Using Droplet-Based Microfluidics Keith Brown Director, Sales brownk@raindancetech.com Who we are: is a Provider of Microdroplet-based Solutions The Company s RainStorm TM Technology
Agilent Genomic Workbench 7.0
 Agilent Genomic Workbench 7.0 Product Overview Guide Agilent Technologies Notices Agilent Technologies, Inc. 2012, 2015 No part of this manual may be reproduced in any form or by any means (including electronic
Agilent Genomic Workbench 7.0 Product Overview Guide Agilent Technologies Notices Agilent Technologies, Inc. 2012, 2015 No part of this manual may be reproduced in any form or by any means (including electronic
Transcriptional Regulation in Eukaryotes
 Transcriptional Regulation in Eukaryotes Concepts, Strategies, and Techniques Michael Carey Stephen T. Smale COLD SPRING HARBOR LABORATORY PRESS NEW YORK 2000 Cold Spring Harbor Laboratory Press, 0-87969-537-4
Transcriptional Regulation in Eukaryotes Concepts, Strategies, and Techniques Michael Carey Stephen T. Smale COLD SPRING HARBOR LABORATORY PRESS NEW YORK 2000 Cold Spring Harbor Laboratory Press, 0-87969-537-4
Quick and easy 7 minute protocol to select for 300 bp, 200 bp, 150 bp, 100 bp, 50 bp DNA fragments or perform a double size selection
 INSTRUCTION MANUAL Select-a-Size DNA Clean & Concentrator Catalog No. D4080 Highlights Quick and easy 7 minute protocol to select for 300 bp, 200 bp, 150 bp, 100 bp, 50 bp DNA fragments or perform a double
INSTRUCTION MANUAL Select-a-Size DNA Clean & Concentrator Catalog No. D4080 Highlights Quick and easy 7 minute protocol to select for 300 bp, 200 bp, 150 bp, 100 bp, 50 bp DNA fragments or perform a double
KAPA HiFi HotStart Uracil+ ReadyMix PCR Kit
 ReadyMix PCR Kit KR043 v4.7 Product Description KAPA HiFi HotStart is a novel B-family DNA polymerase, engineered to have increased affinity for DNA, without the need for accessory proteins or DNA-binding
ReadyMix PCR Kit KR043 v4.7 Product Description KAPA HiFi HotStart is a novel B-family DNA polymerase, engineered to have increased affinity for DNA, without the need for accessory proteins or DNA-binding
Unit 6: Molecular Genetics & DNA Technology Guided Reading Questions (100 pts total)
 Name: AP Biology Biology, Campbell and Reece, 7th Edition Adapted from chapter reading guides originally created by Lynn Miriello Chapter 16 The Molecular Basis of Inheritance Unit 6: Molecular Genetics
Name: AP Biology Biology, Campbell and Reece, 7th Edition Adapted from chapter reading guides originally created by Lynn Miriello Chapter 16 The Molecular Basis of Inheritance Unit 6: Molecular Genetics
ACCEL-NGS 2S DNA LIBRARY KITS
 ACCEL-NGS 2S DNA LIBRARY KITS Accel-NGS 2S DNA Library Kits produce high quality libraries with an all-inclusive, easy-to-use format. The kits contain all reagents necessary to build high complexity libraries
ACCEL-NGS 2S DNA LIBRARY KITS Accel-NGS 2S DNA Library Kits produce high quality libraries with an all-inclusive, easy-to-use format. The kits contain all reagents necessary to build high complexity libraries
# SimpleChIP ChIP-seq Multiplex Oligos for Illumina (Dual Index Primers) 1 Kit (96 assays) Store at -20 C
 Store at -20 C #47538 SimpleChIP ChIP-seq Multiplex Oligos (Dual Index Primers) 1 Kit (96 assays) New 10/17 Support: +1-978-867-2388 (.S.) www.cellsignal.com/support Orders: 877-616-2355 (.S.) orders@cellsignal.com
Store at -20 C #47538 SimpleChIP ChIP-seq Multiplex Oligos (Dual Index Primers) 1 Kit (96 assays) New 10/17 Support: +1-978-867-2388 (.S.) www.cellsignal.com/support Orders: 877-616-2355 (.S.) orders@cellsignal.com
Performance characteristics of the High Sensitivity DNA kit for the Agilent 2100 Bioanalyzer
 Performance characteristics of the High Sensitivity DNA kit for the Agilent 2100 Bioanalyzer Technical Note 10 Measured conc. [ng/µl] 1 Y intercept = 0.09 r 2 = 0.993 0.1 0.1 1 10 Reference concentration
Performance characteristics of the High Sensitivity DNA kit for the Agilent 2100 Bioanalyzer Technical Note 10 Measured conc. [ng/µl] 1 Y intercept = 0.09 r 2 = 0.993 0.1 0.1 1 10 Reference concentration
DNA Function: Information Transmission
 DNA Function: Information Transmission DNA is called the code of life. What does it code for? *the information ( code ) to make proteins! Why are proteins so important? Nearly every function of a living
DNA Function: Information Transmission DNA is called the code of life. What does it code for? *the information ( code ) to make proteins! Why are proteins so important? Nearly every function of a living
Tech Note. Using the ncounter Analysis System with FFPE Samples for Gene Expression Analysis. ncounter Gene Expression. Molecules That Count
 ncounter Gene Expression Tech Note Using the ncounter Analysis System with FFPE Samples for Gene Expression Analysis Introduction For the past several decades, pathologists have kept samples obtained from
ncounter Gene Expression Tech Note Using the ncounter Analysis System with FFPE Samples for Gene Expression Analysis Introduction For the past several decades, pathologists have kept samples obtained from
Microbial Metabolism Systems Microbiology
 1 Microbial Metabolism Systems Microbiology Ching-Tsan Huang ( 黃慶璨 ) Office: Agronomy Hall, Room 111 Tel: (02) 33664454 E-mail: cthuang@ntu.edu.tw MIT OCW Systems Microbiology aims to integrate basic biological
1 Microbial Metabolism Systems Microbiology Ching-Tsan Huang ( 黃慶璨 ) Office: Agronomy Hall, Room 111 Tel: (02) 33664454 E-mail: cthuang@ntu.edu.tw MIT OCW Systems Microbiology aims to integrate basic biological
Considerations for Illumina library preparation. Henriette O Geen June 20, 2014 UCD Genome Center
 Considerations for Illumina library preparation Henriette O Geen June 20, 2014 UCD Genome Center Diversity of applications De novo genome Sequencing ranscriptome Expression Splice Isoform bundance Genotyping
Considerations for Illumina library preparation Henriette O Geen June 20, 2014 UCD Genome Center Diversity of applications De novo genome Sequencing ranscriptome Expression Splice Isoform bundance Genotyping
Methods of Biomaterials Testing Lesson 3-5. Biochemical Methods - Molecular Biology -
 Methods of Biomaterials Testing Lesson 3-5 Biochemical Methods - Molecular Biology - Chromosomes in the Cell Nucleus DNA in the Chromosome Deoxyribonucleic Acid (DNA) DNA has double-helix structure The
Methods of Biomaterials Testing Lesson 3-5 Biochemical Methods - Molecular Biology - Chromosomes in the Cell Nucleus DNA in the Chromosome Deoxyribonucleic Acid (DNA) DNA has double-helix structure The
Recombinant DNA: Basics and Advanced Applications
 Recombinant DNA: Basics and Advanced Applications 2015/2016 Code: 42895 ECTS Credits: 9 Degree Type Year Semester 4313794 Biochemistry, Molecular Biology and Biomedicine OT 0 A Contact Name: Antonio Casamayor
Recombinant DNA: Basics and Advanced Applications 2015/2016 Code: 42895 ECTS Credits: 9 Degree Type Year Semester 4313794 Biochemistry, Molecular Biology and Biomedicine OT 0 A Contact Name: Antonio Casamayor
EUKARYOTIC GENE CONTROL
 EUKARYOTIC GENE CONTROL THE BIG QUESTIONS How are genes turned on and off? How do cells with the same DNA/ genes differentiate to perform completely different and specialized functions? GENE EXPRESSION
EUKARYOTIC GENE CONTROL THE BIG QUESTIONS How are genes turned on and off? How do cells with the same DNA/ genes differentiate to perform completely different and specialized functions? GENE EXPRESSION
Functional Genomics Overview RORY STARK PRINCIPAL BIOINFORMATICS ANALYST CRUK CAMBRIDGE INSTITUTE 18 SEPTEMBER 2017
 Functional Genomics Overview RORY STARK PRINCIPAL BIOINFORMATICS ANALYST CRUK CAMBRIDGE INSTITUTE 18 SEPTEMBER 2017 Agenda What is Functional Genomics? RNA Transcription/Gene Expression Measuring Gene
Functional Genomics Overview RORY STARK PRINCIPAL BIOINFORMATICS ANALYST CRUK CAMBRIDGE INSTITUTE 18 SEPTEMBER 2017 Agenda What is Functional Genomics? RNA Transcription/Gene Expression Measuring Gene
NEBNext. for Ion Torrent LIBRARY PREPARATION KITS
 NEBNext for Ion Torrent LIBRARY PREPARATION KITS NEBNEXT PRODUCTS FOR ION TORRENT Table of Contents 3 General Introduction 4 5 6 6 7 8 DNA Library Preparation Workflow Product Selection Product Details
NEBNext for Ion Torrent LIBRARY PREPARATION KITS NEBNEXT PRODUCTS FOR ION TORRENT Table of Contents 3 General Introduction 4 5 6 6 7 8 DNA Library Preparation Workflow Product Selection Product Details
Ensembl Funcgen: A Database and API for Epigenomics and Gene Regulation Data.
 Ensembl Funcgen: A Database and API for Epigenomics and Gene Regulation Data. Nathan Johnson Ensembl Regulation EBI is an Outstation of the European Molecular Biology Laboratory.! Workshop Overview http://www.ebi.ac.uk/~njohnson/courses/23.05.2013-
Ensembl Funcgen: A Database and API for Epigenomics and Gene Regulation Data. Nathan Johnson Ensembl Regulation EBI is an Outstation of the European Molecular Biology Laboratory.! Workshop Overview http://www.ebi.ac.uk/~njohnson/courses/23.05.2013-
NEBNext Magnesium RNA Fragmentation Module
 SAMPLE PREPARATION NEBNext Magnesium RNA Fragmentation Module Instruction Manual NEB #E6150S 200 reactions NEBNext Magnesium RNA Fragmentation Module Table of Contents: Description....2 Applications....2
SAMPLE PREPARATION NEBNext Magnesium RNA Fragmentation Module Instruction Manual NEB #E6150S 200 reactions NEBNext Magnesium RNA Fragmentation Module Table of Contents: Description....2 Applications....2
Microarray Industry Products
 Via Nicaragua, 12-14 00040 Pomezia (Roma) Phone: +39 06 91601628 Fax: +39 06 91612477 info@lifelinelab.com www.lifelinelab.com Microarray Industry Products Page 10 NBT / BCPIP Chromogenic phosphatase
Via Nicaragua, 12-14 00040 Pomezia (Roma) Phone: +39 06 91601628 Fax: +39 06 91612477 info@lifelinelab.com www.lifelinelab.com Microarray Industry Products Page 10 NBT / BCPIP Chromogenic phosphatase
Method development for DNA methylation on a single cell level
 Method development for DNA methylation on a single cell level The project is carried out in TATAA Biocenter, Gothenburg, Sweden/ Secondment at Babraham institute, Cambridge Dr. Bentolhoda Fereydouni, Marie
Method development for DNA methylation on a single cell level The project is carried out in TATAA Biocenter, Gothenburg, Sweden/ Secondment at Babraham institute, Cambridge Dr. Bentolhoda Fereydouni, Marie
Introduction to Bioinformatics and Gene Expression Technologies
 Introduction to Bioinformatics and Gene Expression Technologies Utah State University Fall 2017 Statistical Bioinformatics (Biomedical Big Data) Notes 1 1 Vocabulary Gene: hereditary DNA sequence at a
Introduction to Bioinformatics and Gene Expression Technologies Utah State University Fall 2017 Statistical Bioinformatics (Biomedical Big Data) Notes 1 1 Vocabulary Gene: hereditary DNA sequence at a
Myers Lab ChIP-seq Protocol v Modified January 10, 2014
 Myers Lab ChIP-seq Protocol V011014 1 Contact information: Dr. Florencia Pauli Behn HudsonAlpha Institute for Biotechnology 601 Genome Way Huntsville, AL 35806 Telephone: 256-327-5229 Email: fpauli@hudsonalpha.org
Myers Lab ChIP-seq Protocol V011014 1 Contact information: Dr. Florencia Pauli Behn HudsonAlpha Institute for Biotechnology 601 Genome Way Huntsville, AL 35806 Telephone: 256-327-5229 Email: fpauli@hudsonalpha.org
M Keramatipour 2. M Keramatipour 1. M Keramatipour 4. M Keramatipour 3. M Keramatipour 5. M Keramatipour
 Molecular Cloning Methods Mohammad Keramatipour MD, PhD keramatipour@tums.ac.ir Outline DNA recombinant technology DNA cloning co Cell based PCR PCR-based Some application of DNA cloning Genomic libraries
Molecular Cloning Methods Mohammad Keramatipour MD, PhD keramatipour@tums.ac.ir Outline DNA recombinant technology DNA cloning co Cell based PCR PCR-based Some application of DNA cloning Genomic libraries
Multi-omics in biology: integration of omics techniques
 31/07/17 Летняя школа по биоинформатике 2017 Multi-omics in biology: integration of omics techniques Konstantin Okonechnikov Division of Pediatric Neurooncology German Cancer Research Center (DKFZ) 2 Short
31/07/17 Летняя школа по биоинформатике 2017 Multi-omics in biology: integration of omics techniques Konstantin Okonechnikov Division of Pediatric Neurooncology German Cancer Research Center (DKFZ) 2 Short
Mentor: Dr. Bino John
 MAT Shiksha Mantri Mentor: Dr. Bino John Model-based Analysis y of Tiling-arrays g y for ChIP-chip X. Shirley Liu et al., PNAS (2006) vol. 103 no. 33 12457 12462 Tiling Arrays Subtype of microarray chips
MAT Shiksha Mantri Mentor: Dr. Bino John Model-based Analysis y of Tiling-arrays g y for ChIP-chip X. Shirley Liu et al., PNAS (2006) vol. 103 no. 33 12457 12462 Tiling Arrays Subtype of microarray chips
ncounter Elements TagSets General Purpose Reagents PACKAGE INSERT Molecules That Count NanoString Technologies, Inc.
 ncounter Elements TagSets General Purpose Reagents PACKAGE INSERT NanoString Technologies, Inc. 530 Fairview Ave N Suite 2000 Seattle, Washington 98109 www.nanostring.com Tel: 206.378.6266 888.358.6266
ncounter Elements TagSets General Purpose Reagents PACKAGE INSERT NanoString Technologies, Inc. 530 Fairview Ave N Suite 2000 Seattle, Washington 98109 www.nanostring.com Tel: 206.378.6266 888.358.6266
Instruction Manual Version DNA Elution Module. Cat. No. C (mc-magme-002)
 Instruction Manual Version 2-01.14 DNA Elution Module Cat. No. C01010120 (mc-magme-002) diagenode headquarters Diagenode s.a. BELGIUM EUROPE LIEGE SCIENCE PARK Rue Bois Saint-Jean, 3 4102 Seraing - Belgium
Instruction Manual Version 2-01.14 DNA Elution Module Cat. No. C01010120 (mc-magme-002) diagenode headquarters Diagenode s.a. BELGIUM EUROPE LIEGE SCIENCE PARK Rue Bois Saint-Jean, 3 4102 Seraing - Belgium
Application Note. Genomics. Abstract. Authors. Kirill Gromadski Ruediger Salowsky Susanne Glueck Agilent Technologies Waldbronn, Germany
 Improving sample quality for target enrichment and next-gen sequencing with the Agilent High Sensitivity DNA Kit and the Agilent SureSelect Target Enrichment Platform Application Note Genomics Authors
Improving sample quality for target enrichment and next-gen sequencing with the Agilent High Sensitivity DNA Kit and the Agilent SureSelect Target Enrichment Platform Application Note Genomics Authors
Genetics Biology 331 Exam 3B Spring 2015
 Genetics Biology 331 Exam 3B Spring 2015 MULTIPLE CHOICE. Choose the one alternative that best completes the statement or answers the question. 1) DNA methylation may be a significant mode of genetic regulation
Genetics Biology 331 Exam 3B Spring 2015 MULTIPLE CHOICE. Choose the one alternative that best completes the statement or answers the question. 1) DNA methylation may be a significant mode of genetic regulation
Recombinant DNA: Basics and Advanced Applications
 Recombinant DNA: Basics and Advanced Applications 2014/2015 Code: 42895 ECTS Credits: 9 Degree Type Year Semester 4313794 Bioquímica, Biologia Molecular i Biomedicina OT 0 A Contact Name: Antonio Casamayor
Recombinant DNA: Basics and Advanced Applications 2014/2015 Code: 42895 ECTS Credits: 9 Degree Type Year Semester 4313794 Bioquímica, Biologia Molecular i Biomedicina OT 0 A Contact Name: Antonio Casamayor
Introduction to Microarray Data Analysis and Gene Networks. Alvis Brazma European Bioinformatics Institute
 Introduction to Microarray Data Analysis and Gene Networks Alvis Brazma European Bioinformatics Institute A brief outline of this course What is gene expression, why it s important Microarrays and how
Introduction to Microarray Data Analysis and Gene Networks Alvis Brazma European Bioinformatics Institute A brief outline of this course What is gene expression, why it s important Microarrays and how
Make the protein through the genetic dogma process.
 Make the protein through the genetic dogma process. Coding Strand 5 AGCAATCATGGATTGGGTACATTTGTAACTGT 3 Template Strand mrna Protein Complete the table. DNA strand DNA s strand G mrna A C U G T A T Amino
Make the protein through the genetic dogma process. Coding Strand 5 AGCAATCATGGATTGGGTACATTTGTAACTGT 3 Template Strand mrna Protein Complete the table. DNA strand DNA s strand G mrna A C U G T A T Amino
What happens after DNA Replication??? Transcription, translation, gene expression/protein synthesis!!!!
 What happens after DNA Replication??? Transcription, translation, gene expression/protein synthesis!!!! Protein Synthesis/Gene Expression Why do we need to make proteins? To build parts for our body as
What happens after DNA Replication??? Transcription, translation, gene expression/protein synthesis!!!! Protein Synthesis/Gene Expression Why do we need to make proteins? To build parts for our body as
Differential Gene Expression
 Biology 4361 Developmental Biology Differential Gene Expression September 28, 2006 Chromatin Structure ~140 bp ~60 bp Transcriptional Regulation: 1. Packing prevents access CH 3 2. Acetylation ( C O )
Biology 4361 Developmental Biology Differential Gene Expression September 28, 2006 Chromatin Structure ~140 bp ~60 bp Transcriptional Regulation: 1. Packing prevents access CH 3 2. Acetylation ( C O )
ENCODE RBP Antibody Characterization Guidelines
 ENCODE RBP Antibody Characterization Guidelines Approved on November 18, 2016 Background An integral part of the ENCODE Project is to characterize the antibodies used in the experiments. This document
ENCODE RBP Antibody Characterization Guidelines Approved on November 18, 2016 Background An integral part of the ENCODE Project is to characterize the antibodies used in the experiments. This document
Modern Epigenomics. Histone Code
 Modern Epigenomics Histone Code Ting Wang Department of Genetics Center for Genome Sciences and Systems Biology Washington University Dragon Star 2012 Changchun, China July 2, 2012 DNA methylation + Histone
Modern Epigenomics Histone Code Ting Wang Department of Genetics Center for Genome Sciences and Systems Biology Washington University Dragon Star 2012 Changchun, China July 2, 2012 DNA methylation + Histone
TECHNICAL BULLETIN. SeqPlex DNA Amplification Kit for use with high throughput sequencing technologies. Catalog Number SEQX Storage Temperature 20 C
 SeqPlex DNA Amplification Kit for use with high throughput sequencing technologies Catalog Number SEQX Storage Temperature 20 C TECHNICAL BULLETIN Product Description The SeqPlex DNA Amplification Kit
SeqPlex DNA Amplification Kit for use with high throughput sequencing technologies Catalog Number SEQX Storage Temperature 20 C TECHNICAL BULLETIN Product Description The SeqPlex DNA Amplification Kit
3.1.4 DNA Microarray Technology
 3.1.4 DNA Microarray Technology Scientists have discovered that one of the differences between healthy and cancer is which genes are turned on in each. Scientists can compare the gene expression patterns
3.1.4 DNA Microarray Technology Scientists have discovered that one of the differences between healthy and cancer is which genes are turned on in each. Scientists can compare the gene expression patterns
RNA-Sequencing analysis
 RNA-Sequencing analysis Markus Kreuz 25. 04. 2012 Institut für Medizinische Informatik, Statistik und Epidemiologie Content: Biological background Overview transcriptomics RNA-Seq RNA-Seq technology Challenges
RNA-Sequencing analysis Markus Kreuz 25. 04. 2012 Institut für Medizinische Informatik, Statistik und Epidemiologie Content: Biological background Overview transcriptomics RNA-Seq RNA-Seq technology Challenges
Supporting Information
 Supporting Information Ho et al. 1.173/pnas.81288816 SI Methods Sequences of shrna hairpins: Brg shrna #1: ccggcggctcaagaaggaagttgaactcgagttcaacttccttcttgacgnttttg (TRCN71383; Open Biosystems). Brg shrna
Supporting Information Ho et al. 1.173/pnas.81288816 SI Methods Sequences of shrna hairpins: Brg shrna #1: ccggcggctcaagaaggaagttgaactcgagttcaacttccttcttgacgnttttg (TRCN71383; Open Biosystems). Brg shrna
Introduction to the UCSC genome browser
 Introduction to the UCSC genome browser Dominik Beck NHMRC Peter Doherty and CINSW ECR Fellow, Senior Lecturer Lowy Cancer Research Centre, UNSW and Centre for Health Technology, UTS SYDNEY NSW AUSTRALIA
Introduction to the UCSC genome browser Dominik Beck NHMRC Peter Doherty and CINSW ECR Fellow, Senior Lecturer Lowy Cancer Research Centre, UNSW and Centre for Health Technology, UTS SYDNEY NSW AUSTRALIA
Axygen AxyPrep Magnetic Bead Purification Kits. A Corning Brand
 Axygen AxyPrep Magnetic Bead Purification Kits A Corning Brand D Sample Prep Solutions for Genomics Obtaining Pure Nucleic Acids from Your Sample is Precious The purification of high quality DNA is the
Axygen AxyPrep Magnetic Bead Purification Kits A Corning Brand D Sample Prep Solutions for Genomics Obtaining Pure Nucleic Acids from Your Sample is Precious The purification of high quality DNA is the
WORKSHOP. Transcriptional circuitry and the regulatory conformation of the genome. Ofir Hakim Faculty of Life Sciences
 WORKSHOP Transcriptional circuitry and the regulatory conformation of the genome Ofir Hakim Faculty of Life Sciences Chromosome conformation capture (3C) Most GR Binding Sites Are Distant From Regulated
WORKSHOP Transcriptional circuitry and the regulatory conformation of the genome Ofir Hakim Faculty of Life Sciences Chromosome conformation capture (3C) Most GR Binding Sites Are Distant From Regulated
Fig. 16-7a. 5 end Hydrogen bond 3 end. 1 nm. 3.4 nm nm
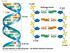 Fig. 16-7a end Hydrogen bond end 1 nm 3.4 nm 0.34 nm (a) Key features of DNA structure end (b) Partial chemical structure end Fig. 16-8 Adenine (A) Thymine (T) Guanine (G) Cytosine (C) Concept 16.2: Many
Fig. 16-7a end Hydrogen bond end 1 nm 3.4 nm 0.34 nm (a) Key features of DNA structure end (b) Partial chemical structure end Fig. 16-8 Adenine (A) Thymine (T) Guanine (G) Cytosine (C) Concept 16.2: Many
A step-by-step guide to ChIP-seq data analysis
 A step-by-step guide to ChIP-seq data analysis December 03, 2014 Xi Chen, Ph.D. EMBL-European Bioinformatics Institute Wellcome Trust Sanger Institute Target audience Wet-lab biologists with no experience
A step-by-step guide to ChIP-seq data analysis December 03, 2014 Xi Chen, Ph.D. EMBL-European Bioinformatics Institute Wellcome Trust Sanger Institute Target audience Wet-lab biologists with no experience
Wednesday, November 22, 17. Exons and Introns
 Exons and Introns Introns and Exons Exons: coded regions of DNA that get transcribed and translated into proteins make up 5% of the genome Introns and Exons Introns: non-coded regions of DNA Must be removed
Exons and Introns Introns and Exons Exons: coded regions of DNA that get transcribed and translated into proteins make up 5% of the genome Introns and Exons Introns: non-coded regions of DNA Must be removed
Gene Expression Profiling and Validation Using Agilent SurePrint G3 Gene Expression Arrays
 Gene Expression Profiling and Validation Using Agilent SurePrint G3 Gene Expression Arrays Application Note Authors Bahram Arezi, Nilanjan Guha and Anne Bergstrom Lucas Agilent Technologies Inc. Santa
Gene Expression Profiling and Validation Using Agilent SurePrint G3 Gene Expression Arrays Application Note Authors Bahram Arezi, Nilanjan Guha and Anne Bergstrom Lucas Agilent Technologies Inc. Santa
EPIGENTEK. EpiQuik In Situ Histone H3-K27 Tri-Methylation Assay Kit. Base Catalog # P-3014T PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE
 EpiQuik In Situ Histone H3-K27 Tri-Methylation Assay Kit Base Catalog # PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE The EpiQuik In Situ Histone H3-K27 Tri-Methylation Assay Kit is suitable for specifically
EpiQuik In Situ Histone H3-K27 Tri-Methylation Assay Kit Base Catalog # PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE The EpiQuik In Situ Histone H3-K27 Tri-Methylation Assay Kit is suitable for specifically
