LECTURE PRESENTATIONS
|
|
|
- Randolph Terry
- 5 years ago
- Views:
Transcription
1 LECTURE PRESENTATIONS For CAMPBELL BIOLOGY, NINTH EDITION Jane B. Reece, Lisa A. Urry, Michael L. Cain, Steven A. Wasserman, Peter V. Minorsky, Robert B. Jackson Chapter 20 Biotechnology Lectures by Erin Barley Kathleen Fitzpatrick
2 Overview: The DNA Toolbox Sequencing of the genomes of more than 7,000 species was under way in 2010 DNA sequencing has depended on advances in technology, starting with making recombinant DNA In recombinant DNA, nucleotide sequences from two different sources, often two species, are combined in vitro into the same DNA molecule
3 Methods for making recombinant DNA are central to genetic engineering, the direct manipulation of genes for practical purposes DNA technology has revolutionized biotechnology, the manipulation of organisms or their genetic components to make useful products An example of DNA technology is the microarray, a measurement of gene expression of thousands of different genes
4 Figure 20.1
5 Concept 20.1: DNA cloning yields multiple copies of a gene or other DNA segment To work directly with specific genes, scientists prepare well-defined segments of DNA in identical copies, a process called DNA cloning
6 DNA Cloning and Its Applications: A Preview Most methods for cloning pieces of DNA in the laboratory share general features, such as the use of bacteria and their plasmids Plasmids are small circular DNA molecules that replicate separately from the bacterial chromosome Cloned genes are useful for making copies of a particular gene and producing a protein product
7 Gene cloning involves using bacteria to make multiple copies of a gene Foreign DNA is inserted into a plasmid, and the recombinant plasmid is inserted into a bacterial cell Reproduction in the bacterial cell results in cloning of the plasmid including the foreign DNA This results in the production of multiple copies of a single gene
8 Figure 20.2 Bacterium 1 Gene inserted into plasmid Cell containing gene of interest Bacterial chromosome Plasmid Recombinant DNA (plasmid) 2 Gene of interest Plasmid put into bacterial cell DNA of chromosome ( foreign DNA) Recombinant bacterium 3 Host cell grown in culture to form a clone of cells containing the cloned gene of interest Gene of interest Copies of gene Protein expressed from gene of interest Protein harvested Basic research on gene 4 Basic research and various applications Basic research on protein Gene for pest resistance inserted into plants Gene used to alter bacteria for cleaning up toxic waste Protein dissolves blood clots in heart attack therapy Human growth hormone treats stunted growth
9 Using Restriction Enzymes to Make Recombinant DNA Bacterial restriction enzymes cut DNA molecules at specific DNA sequences called restriction sites A restriction enzyme usually makes many cuts, yielding restriction fragments The most useful restriction enzymes cut DNA in a staggered way, producing fragments with sticky ends.
10 Sticky ends can bond with complementary sticky ends of other fragments DNA ligase is an enzyme that seals the bonds between restriction fragments
11 Figure Restriction site 1 DNA 5 Restriction enzyme cuts sugar-phosphate backbones GAATTC CTTAAG Sticky end DNA fragment added from another molecule cut by same enzyme. Base pairing occurs G AATT C C TTAA G G AATT C C TTAA G 5 3 One possible combination DNA ligase seals strands Recombinant DNA molecule 5
12 Cloning a Eukaryotic Gene in a Bacterial Plasmid In gene cloning, the original plasmid is called a cloning vector A cloning vector is a DNA molecule that can carry foreign DNA into a host cell and replicate there
13 Producing Clones of Cells Carrying Recombinant Plasmids Several steps are required to clone the hummingbird β-globin gene in a bacterial plasmid The hummingbird genomic DNA and a bacterial plasmid are isolated Both are cut with the same restriction enzyme The fragments are mixed, and DNA ligase is added to bond the fragment sticky ends
14 Some recombinant plasmids now contain hummingbird DNA The DNA mixture is added to bacteria that have been genetically engineered to accept it The bacteria are plated on a type of agar that selects for the bacteria with recombinant plasmids This results in the cloning of many hummingbird DNA fragments, including the β-globin gene
15 Figure 20.4 TECHNIQUE Bacterial plasmid amp R gene lacz gene Restriction site Hummingbird cell Sticky ends Gene of interest Hummingbird DNA fragments Recombinant plasmids Nonrecombinant plasmid Bacteria carrying plasmids RESULTS Colony carrying nonrecombinant plasmid with intact lacz gene Colony carrying recombinant plasmid with disrupted lacz gene One of many bacterial clones
16 Storing Cloned Genes in DNA Libraries A genomic library that is made using bacteria is the collection of recombinant vector clones produced by cloning DNA fragments from an entire genome A genomic library that is made using bacteriophages is stored as a collection of phage clones
17 Figure 20.5 Foreign genome Cut with restriction enzymes into either small or large Bacterial artificial fragments fragments chromosome (BAC) Large insert with many genes Recombinant plasmids (b) BAC clone Plasmid clone (a) Plasmid library (c) Storing genome libraries
18 A bacterial artificial chromosome (BAC) is a large plasmid that has been trimmed down and can carry a large DNA insert BACs are another type of vector used in DNA library construction
19 A complementary DNA (cdna) library is made by cloning DNA made in vitro by reverse transcription of all the mrna produced by a particular cell A cdna library represents only part of the genome only the subset of genes transcribed into mrna in the original cells
20 Figure DNA in nucleus mrnas in cytoplasm mrna Reverse transcriptase Poly-A tail 3 DNA strand 3 5 Primer A A A A A A T T T T T A A A A A A T T T T T DNA polymerase cdna 3 5
21 Screening a Library for Clones Carrying a Gene of Interest A clone carrying the gene of interest can be identified with a nucleic acid probe having a sequence complementary to the gene This process is called nucleic acid hybridization
22 A probe can be synthesized that is complementary to the gene of interest For example, if the desired gene is 5 CTCAT CACCGGC 3 Then we would synthesize this probe 3 G A G T A G T G G C C G 5
23 The DNA probe can be used to screen a large number of clones simultaneously for the gene of interest Once identified, the clone carrying the gene of interest can be cultured
24 Figure 20.7 TECHNIQUE Multiwell plates holding library clones Radioactively labeled probe molecules Gene of interest 5 3 CTCATCACCGGC GAGTAGTGGCCG 3 5 Probe DNA Single- Film stranded DNA from cell Nylon membrane Location of DNA with the complementary sequence Nylon membrane
25 Expressing Cloned Eukaryotic Genes After a gene has been cloned, its protein product can be produced in larger amounts for research Cloned genes can be expressed as protein in either bacterial or eukaryotic cells
26 Bacterial Expression Systems Several technical difficulties hinder expression of cloned eukaryotic genes in bacterial host cells To overcome differences in promoters and other DNA control sequences, scientists usually employ an expression vector, a cloning vector that contains a highly active bacterial promoter
27 Eukaryotic Cloning and Expression Systems Molecular biologists can avoid eukaryote-bacterial incompatibility issues by using eukaryotic cells, such as yeasts, as hosts for cloning and expressing genes Even yeasts may not possess the proteins required to modify expressed mammalian proteins properly In such cases, cultured mammalian or insect cells may be used to express and study proteins
28 One method of introducing recombinant DNA into eukaryotic cells is electroporation, applying a brief electrical pulse to create temporary holes in plasma membranes Alternatively, scientists can inject DNA into cells using microscopically thin needles Once inside the cell, the DNA is incorporated into the cell s DNA by natural genetic recombination
29 Cross-Species Gene Expression and Evolutionary Ancestry The remarkable ability of bacteria to express some eukaryotic proteins underscores the shared evolutionary ancestry of living species For example, Pax-6 is a gene that directs formation of a vertebrate eye; the same gene in flies directs the formation of an insect eye (which is quite different from the vertebrate eye) The Pax-6 genes in flies and vertebrates can substitute for each other
30 Amplifying DNA in Vitro: The Polymerase Chain Reaction (PCR) The polymerase chain reaction, PCR, can produce many copies of a specific target segment of DNA A three-step cycle heating, cooling, and replication brings about a chain reaction that produces an exponentially growing population of identical DNA molecules The key to PCR is an unusual, heat-stable DNA polymerase called Taq polymerase.
31 Figure 20.8 TECHNIQUE 5 3 Target sequence Genomic DNA Denaturation Annealing Cycle 1 yields 2 molecules Primers 3 Extension New nucleotides Cycle 2 yields 4 molecules Cycle 3 yields 8 molecules; 2 molecules (in white boxes) match target sequence
32 Concept 20.2: DNA technology allows us to study the sequence, expression, and function of a gene DNA cloning allows researchers to Compare genes and alleles between individuals Locate gene expression in a body Determine the role of a gene in an organism Several techniques are used to analyze the DNA of genes
33 Gel Electrophoresis and Southern Blotting One indirect method of rapidly analyzing and comparing genomes is gel electrophoresis This technique uses a gel as a molecular sieve to separate nucleic acids or proteins by size, electrical charge, and other properties A current is applied that causes charged molecules to move through the gel Molecules are sorted into bands by their size
34 Figure 20.9 TECHNIQUE 1 Mixture of DNA molecules of different sizes Power source Cathode Wells Anode Gel 2 Longer molecules Power source Shorter molecules RESULTS
35 In restriction fragment analysis, DNA fragments produced by restriction enzyme digestion of a DNA molecule are sorted by gel electrophoresis Restriction fragment analysis can be used to compare two different DNA molecules, such as two alleles for a gene, if the nucleotide difference alters a restriction site
36 Variations in DNA sequence are called polymorphisms Sequence changes that alter restriction sites are called RFLPs (restriction fragment length polymorphisms)
37 Figure Normal -globin allele Normal allele Sickle-cell allele 175 bp Sickle-cell mutant -globin allele 376 bp 201 bp Large fragment DdeI DdeI DdeI DdeI Large fragment DdeI DdeI DdeI (a) DdeI restriction sites in normal and sickle-cell alleles of the -globin gene Large fragment 201 bp 175 bp 376 bp (b) Electrophoresis of restriction fragments from normal and sickle-cell alleles
38 A technique called Southern blotting combines gel electrophoresis of DNA fragments with nucleic acid hybridization Specific DNA fragments can be identified by Southern blotting, using labeled probes that hybridize to the DNA immobilized on a blot of gel
39 Figure TECHNIQUE DNA restriction enzyme Restriction fragments I II III Nitrocellulose membrane (blot) Heavy weight Gel Sponge I Normal -globin allele 1 II Sickle-cell allele Preparation of restriction fragments III Heterozygote 2 Gel electrophoresis Alkaline solution Paper towels 3 DNA transfer (blotting) Probe base-pairs I II III with fragments I II III Radioactively labeled probe for -globin gene 4 Nitrocellulose blot Hybridization with labeled probe Fragment from sickle-cell -globin allele Fragment from normal - globin allele Film over blot 5 Probe detection
40 DNA Sequencing Relatively short DNA fragments can be sequenced by the dideoxy chain termination method, the first automated method to be employed Modified nucleotides called dideoxyribonucleotides (ddntp) attach to synthesized DNA strands of different lengths Each type of ddntp is tagged with a distinct fluorescent label that identifies the nucleotide at the end of each DNA fragment The DNA sequence can be read from the resulting spectrogram
41 Figure TECHNIQUE DNA (template strand) 5 C T G A C T T C G A C A 3 A 5 3 C T G A C T T C G A C A A Primer T 3 G T T 5 DNA (template strand) ddg ddc C T T G G T T T T Shortest Direction of movement of strands DNA polymerase dda G C T G T T Deoxyribonucleotides Dideoxyribonucleotides (fluorescently tagged) datp dctp dttp dgtp P P P OH G Labeled strands dda A G C T G T T ddg A A G C T G T T ddt G A A G C T G T T Longest labeled strand ddc T G A A G C T G T T ddatp ddctp ddttp ddgtp P P P dda C T G A A G C T G T T H G ddg 3 A C T G A A G C T G T T 5 Longest Detector RESULTS Laser Last nucleotide of longest labeled strand Last nucleotide of shortest labeled strand Shortest labeled strand G A C T G A A G C
42 Analyzing Gene Expression Nucleic acid probes can hybridize with mrnas transcribed from a gene Probes can be used to identify where or when a gene is transcribed in an organism
43 Studying the Expression of Single Genes Changes in the expression of a gene during embryonic development can be tested using Northern blotting Reverse transcriptase-polymerase chain reaction Both methods are used to compare mrna from different developmental stages
44 Northern blotting combines gel electrophoresis of mrna followed by hybridization with a probe on a membrane Identification of mrna at a particular developmental stage suggests protein function at that stage
45 Reverse transcriptase-polymerase chain reaction (RT-PCR) is quicker and more sensitive because it requires less mrna than Northern blotting Reverse transcriptase is added to mrna to make cdna, which serves as a template for PCR amplification of the gene of interest The products are run on a gel and the mrna of interest is identified
46 Figure TECHNIQUE 1 cdna synthesis mrnas cdnas 2 PCR amplification Primers 3 Gel electrophoresis -globin gene RESULTS Embryonic stages
47 In situ hybridization uses fluorescent dyes attached to probes to identify the location of specific mrnas in place in the intact organism
48 Figure m
49 Studying the Expression of Interacting Groups of Genes Automation has allowed scientists to measure the expression of thousands of genes at one time using DNA microarray assays DNA microarray assays compare patterns of gene expression in different tissues, at different times, or under different conditions
50 Figure TECHNIQUE 1 Isolate mrna. Tissue sample 2 Make cdna by reverse transcription, using fluorescently labeled nucleotides. mrna molecules 3 Apply the cdna mixture to a microarray, a different gene in each spot. The cdna hybridizes with any complementary DNA on the microarray. Labeled cdna molecules (single strands) DNA microarray DNA fragments representing a specific gene 4 Rinse off excess cdna; scan microarray for fluorescence. Each fluorescent spot (yellow) represents a gene expressed in the tissue sample. DNA microarray with 2,400 human genes
51 Determining Gene Function One way to determine function is to disable the gene and observe the consequences Using in vitro mutagenesis, mutations are introduced into a cloned gene, altering or destroying its function When the mutated gene is returned to the cell, the normal gene s function might be determined by examining the mutant s phenotype
52 Gene expression can also be silenced using RNA interference (RNAi) Synthetic double-stranded RNA molecules matching the sequence of a particular gene are used to break down or block the gene s mrna
53 In humans, researchers analyze the genomes of many people with a certain genetic condition to try to find nucleotide changes specific to the condition Genetic markers called SNPs (single nucleotide polymorphisms) occur on average every base pairs SNPs can be detected by PCR, and any SNP shared by people affected with a disorder but not among unaffected people may pinpoint the location of the disease-causing gene
54 Figure DNA SNP T Normal allele C Disease-causing allele
55 Concept 20.3: Cloning organisms may lead to production of stem cells for research and other applications Organismal cloning produces one or more organisms genetically identical to the parent that donated the single cell
56 Cloning Plants: Single-Cell Cultures One experimental approach for testing genomic equivalence is to see whether a differentiated cell can generate a whole organism A totipotent cell is one that can generate a complete new organism Plant cloning is used extensively in agriculture
57 Figure Cross section of carrot root 2-mg fragments Fragments were cultured in nutrient medium; stirring caused single cells to shear off into the liquid. Single cells free in suspension began to divide. Embryonic plant developed from a cultured single cell. Plantlet was cultured on agar medium. Later it was planted in soil. Adult plant
58 Cloning Animals: Nuclear Transplantation In nuclear transplantation, the nucleus of an unfertilized egg cell or zygote is replaced with the nucleus of a differentiated cell Experiments with frog embryos have shown that a transplanted nucleus can often support normal development of the egg However, the older the donor nucleus, the lower the percentage of normally developing tadpoles
59 Figure EXPERIMENT Frog embryo Frog egg cell Frog tadpole UV RESULTS Less differentiated cell Donor nucleus transplanted Enucleated egg cell Egg with donor nucleus activated to begin development Fully differentiated (intestinal) cell Donor nucleus transplanted Most develop into tadpoles. Most stop developing before tadpole stage.
60 Reproductive Cloning of Mammals In 1997, Scottish researchers announced the birth of Dolly, a lamb cloned from an adult sheep by nuclear transplantation from a differentiated mammary cell Dolly s premature death in 2003, as well as her arthritis, led to speculation that her cells were not as healthy as those of a normal sheep, possibly reflecting incomplete reprogramming of the original transplanted nucleus
61 Figure TECHNIQUE Mammary cell donor Egg cell donor 1 Cultured mammary cells 3 Egg cell from ovary Cells fused 2 Nucleus removed 4 5 Grown in culture Implanted in uterus of a third sheep Nucleus from mammary cell Early embryo 6 Embryonic development RESULTS Surrogate mother Lamb ( Dolly ) genetically identical to mammary cell donor
62 Since 1997, cloning has been demonstrated in many mammals, including mice, cats, cows, horses, mules, pigs, and dogs CC (for Carbon Copy) was the first cat cloned; however, CC differed somewhat from her female parent Cloned animals do not always look or behave exactly the same
63 Figure 20.20
64 Problems Associated with Animal Cloning In most nuclear transplantation studies, only a small percentage of cloned embryos have developed normally to birth, and many cloned animals exhibit defects Many epigenetic changes, such as acetylation of histones or methylation of DNA, must be reversed in the nucleus from a donor animal in order for genes to be expressed or repressed appropriately for early stages of development
65 Stem Cells of Animals A stem cell is a relatively unspecialized cell that can reproduce itself indefinitely and differentiate into specialized cells of one or more types Stem cells isolated from early embryos at the blastocyst stage are called embryonic stem (ES) cells; these are able to differentiate into all cell types The adult body also has stem cells, which replace nonreproducing specialized cells
66 Figure Embryonic stem cells Adult stem cells Cultured stem cells Cells generating all embryonic cell types Cells generating some cell types Different culture conditions Different types of differentiated cells Liver cells Nerve cells Blood cells
67 Researchers can transform skin cells into ES cells by using viruses to introduce stem cell master regulatory genes These transformed cells are called ips cells (induced pluripotent cells) These cells can be used to treat some diseases and to replace nonfunctional tissues
68 Figure Remove skin cells from patient. 2 Reprogram skin cells so the cells become induced pluripotent stem (ips) cells. Patient with damaged heart tissue or other disease 3 Treat ips cells so that they differentiate into a specific cell type. 4 Return cells to patient, where they can repair damaged tissue.
69 Concept 20.4: The practical applications of DNA technology affect our lives in many ways Many fields benefit from DNA technology and genetic engineering
70 Medical Applications One benefit of DNA technology is identification of human genes in which mutation plays a role in genetic diseases
71 Diagnosis and Treatment of Diseases Scientists can diagnose many human genetic disorders using PCR and sequence-specific primers, then sequencing the amplified product to look for the disease-causing mutation SNPs may be associated with a disease-causing mutation SNPs may also be correlated with increased risks for conditions such as heart disease or certain types of cancer
72 Human Gene Therapy Gene therapy is the alteration of an afflicted individual s genes Gene therapy holds great potential for treating disorders traceable to a single defective gene Vectors are used for delivery of genes into specific types of cells, for example bone marrow Gene therapy provokes both technical and ethical questions
73 Figure Cloned gene 1 Insert RNA version of normal allele into retrovirus. Viral RNA Retrovirus capsid 2 Let retrovirus infect bone marrow cells that have been removed from the patient and cultured. 3 Viral DNA carrying the normal allele inserts into chromosome. Bone marrow cell from patient 4 Inject engineered cells into patient. Bone marrow
74 Pharmaceutical Products Advances in DNA technology and genetic research are important to the development of new drugs to treat diseases
75 Synthesis of Small Molecules for Use as Drugs The drug imatinib is a small molecule that inhibits overexpression of a specific leukemia-causing receptor Pharmaceutical products that are proteins can be synthesized on a large scale
76 Protein Production in Cell Cultures Host cells in culture can be engineered to secrete a protein as it is made, simplifying the task of purifying it This is useful for the production of insulin, human growth hormones, and vaccines
77 Protein Production by Pharm Animals Transgenic animals are made by introducing genes from one species into the genome of another animal Transgenic animals are pharmaceutical factories, producers of large amounts of otherwise rare substances for medical use
78 Figure 20.24
79 Forensic Evidence and Genetic Profiles An individual s unique DNA sequence, or genetic profile, can be obtained by analysis of tissue or body fluids DNA testing can identify individuals with a high degree of certainty Genetic profiles can be analyzed using RFLP analysis by Southern blotting
80 Even more sensitive is the use of genetic markers called short tandem repeats (STRs), which are variations in the number of repeats of specific DNA sequences PCR and gel electrophoresis are used to amplify and then identify STRs of different lengths The probability that two people who are not identical twins have the same STR markers is exceptionally small
81 Figure (a) This photo shows Washington just before his release in 2001, after 17 years in prison. Source of sample STR marker 1 STR marker 2 STR marker 3 Semen on victim 17,19 13,16 12,12 Earl Washington 16,18 14,15 11,12 Kenneth Tinsley 17,19 13,16 12,12 (b) These and other STR data exonerated Washington and led Tinsley to plead guilty to the murder.
82 Environmental Cleanup Genetic engineering can be used to modify the metabolism of microorganisms Some modified microorganisms can be used to extract minerals from the environment or degrade potentially toxic waste materials
83 Agricultural Applications DNA technology is being used to improve agricultural productivity and food quality Genetic engineering of transgenic animals speeds up the selective breeding process Beneficial genes can be transferred between varieties or species
84 Agricultural scientists have endowed a number of crop plants with genes for desirable traits The Ti plasmid is the most commonly used vector for introducing new genes into plant cells Genetic engineering in plants has been used to transfer many useful genes including those for herbicide resistance, increased resistance to pests, increased resistance to salinity, and improved nutritional value of crops
85 Figure TECHNIQUE Agrobacterium tumefaciens Ti plasmid Site where restriction enzyme cuts DNA with the gene of interest Recombinant Ti plasmid T DNA RESULTS Plant with new trait
86 Safety and Ethical Questions Raised by DNA Technology Potential benefits of genetic engineering must be weighed against potential hazards of creating harmful products or procedures Guidelines are in place in the United States and other countries to ensure safe practices for recombinant DNA technology
87 Most public concern about possible hazards centers on genetically modified (GM) organisms used as food Some are concerned about the creation of super weeds from the transfer of genes from GM crops to their wild relatives Other worries include the possibility that transgenic protein products might cause allergic reactions
88 As biotechnology continues to change, so does its use in agriculture, industry, and medicine National agencies and international organizations strive to set guidelines for safe and ethical practices in the use of biotechnology
89 Figure 20.UN G CTTAA 5 5 AATTC G 3 Sticky end 3 5
90 Figure 20.UN04 Cloning vector DNA fragments from genomic DNA or cdna or copy of DNA obtained by PCR Mix and ligate Recombinant DNA plasmids
91 Figure 20.UN TCCATGAATTCTAAAGCGCTTATGAATTCACGGC AGGTACTTAAGATTTCGCGAATACTTAAGTGCCG Aardvark DNA 3 5 Plasmid
92 Figure 20.UN06
93 Figure 20.UN07
94 Figure 20.UN08
95 CH 21 Genome and Evolution
96 Overview: Reading the Leaves from the Tree of Life Complete genome sequences exist for a human, chimpanzee, E. coli, brewer s yeast, corn, fruit fly, house mouse, rhesus macaque, and other organisms Comparisons of genomes among organisms provide information about the evolutionary history of genes and taxonomic groups
97 Genomics is the study of whole sets of genes and their interactions Bioinformatics is the application of computational methods to the storage and analysis of biological data
98 Figure 21.1
99 Concept 21.1: New approaches have accelerated the pace of genome sequencing The most ambitious mapping project to date has been the sequencing of the human genome Officially begun as the Human Genome Project in 1990, the sequencing was largely completed by 2003 The project had three stages Genetic (or linkage) mapping Physical mapping DNA sequencing
100 Three-Stage Approach to Genome Sequencing A linkage map (genetic map) maps the location of several thousand genetic markers on each chromosome A genetic marker is a gene or other identifiable DNA sequence Recombination frequencies are used to determine the order and relative distances between genetic markers
101 Figure Chromosome bands Cytogenetic map Genes located by FISH 1 Linkage mapping Genetic markers 2 Physical mapping Overlapping fragments 3 DNA sequencing
102 A physical map expresses the distance between genetic markers, usually as the number of base pairs along the DNA It is constructed by cutting a DNA molecule into many short fragments and arranging them in order by identifying overlaps
103 Sequencing machines are used to determine the complete nucleotide sequence of each chromosome A complete haploid set of human chromosomes consists of 3.2 billion base pairs
104 Whole-Genome Shotgun Approach to Genome Sequencing The whole-genome shotgun approach was developed by J. Craig Venter in 1992 This approach skips genetic and physical mapping and sequences random DNA fragments directly Powerful computer programs are used to order fragments into a continuous sequence
105 Figure Cut the DNA into overlapping fragments short enough for sequencing. 2 Clone the fragments in plasmid or phage vectors. 3 Sequence each fragment. 4 Order the sequences into one overall sequence with computer software.
106 Both the three-stage process and the wholegenome shotgun approach were used for the Human Genome Project and for genome sequencing of other organisms At first many scientists were skeptical about the whole-genome shotgun approach, but it is now widely used as the sequencing method of choice The development of newer sequencing techniques has resulted in massive increases in speed and decreases in cost
107 Technological advances have also facilitated metagenomics, in which DNA from a group of species (a metagenome) is collected from an environmental sample and sequenced This technique has been used on microbial communities, allowing the sequencing of DNA of mixed populations, and eliminating the need to culture species in the lab
108 Concept 21.2 Scientists use bioinformatics to analyze genomes and their functions The Human Genome Project established databases and refined analytical software to make data available on the Internet This has accelerated progress in DNA sequence analysis
109 Concept 21.3 Genomes vary in size, number of genes, and gene density By early 2010, over 1,200 genomes were completely sequenced, including 1,000 bacteria, 80 archaea, and 124 eukaryotes Sequencing of over 5,500 genomes and over 200 metagenomes is currently in progress
110 Genome Size Genomes of most bacteria and archaea range from 1 to 6 million base pairs (Mb); genomes of eukaryotes are usually larger Most plants and animals have genomes greater than 100 Mb; humans have 3,000 Mb Within each domain there is no systematic relationship between genome size and phenotype
111 Table 21.1
112 Number of Genes Free-living bacteria and archaea have 1,500 to 7,500 genes Unicellular fungi have from about 5,000 genes and multicellular eukaryotes up to at least 40,000 genes
113 Number of genes is not correlated to genome size For example, it is estimated that the nematode C. elegans has 100 Mb and 20,000 genes, while Drosophila has 165 Mb and 13,700 genes Vertebrate genomes can produce more than one polypeptide per gene because of alternative splicing of RNA transcripts
114 Gene Density and Noncoding DNA Humans and other mammals have the lowest gene density, or number of genes, in a given length of DNA Multicellular eukaryotes have many introns within genes and noncoding DNA between genes
115 Concept 21.4: Multicellular eukaryotes have much noncoding DNA and many multigene families The bulk of most eukaryotic genomes neither encodes proteins nor functional RNAs Much evidence indicates that noncoding DNA (previously called junk DNA ) plays important roles in the cell For example, genomes of humans, rats, and mice show high sequence conservation for about 500 noncoding regions
116 Sequencing of the human genome reveals that 98.5% does not code for proteins, rrnas, or trnas About a quarter of the human genome codes for introns and gene-related regulatory sequences
117 Intergenic DNA is noncoding DNA found between genes Pseudogenes are former genes that have accumulated mutations and are nonfunctional Repetitive DNA is present in multiple copies in the genome About three-fourths of repetitive DNA is made up of transposable elements and sequences related to them
118 Figure 21.7 Exons (1.5%) Introns (5%) L1 sequences (17%) Alu elements (10%) Repetitive DNA that includes transposable elements and related sequences (44%) Simple sequence DNA (3%) Repetitive DNA unrelated to transposable elements (14%) Regulatory sequences ( 20%) Unique noncoding DNA (15%) Large-segment duplications (5 6%)
119 Transposable Elements and Related Sequences The first evidence for mobile DNA segments came from geneticist Barbara McClintock s breeding experiments with Indian corn McClintock identified changes in the color of corn kernels that made sense only by postulating that some genetic elements move from other genome locations into the genes for kernel color These transposable elements move from one site to another in a cell s DNA; they are present in both prokaryotes and eukaryotes
120 Figure 21.8
121 Movement of Transposons and Retrotransposons Eukaryotic transposable elements are of two types Transposons, which move by means of a DNA intermediate Retrotransposons, which move by means of an RNA intermediate
122 Figure 21.9 Transposon New copy of transposon DNA of genome Transposon is copied Insertion Mobile transposon
123 Figure Retrotransposon New copy of retrotransposon Formation of a single-stranded RNA intermediate RNA Insertion Reverse transcriptase
124 Sequences Related to Transposable Elements Multiple copies of transposable elements and related sequences are scattered throughout the eukaryotic genome In primates, a large portion of transposable element related DNA consists of a family of similar sequences called Alu elements Many Alu elements are transcribed into RNA molecules; however their function, if any, is unknown
125 The human genome also contains many sequences of a type of retrotransposon called LINE-1 (L1) L1 sequences have a low rate of transposition and may help regulate gene expression
126 Other Repetitive DNA, Including Simple Sequence DNA About 15% of the human genome consists of duplication of long sequences of DNA from one location to another In contrast, simple sequence DNA contains many copies of tandemly repeated short sequences
127 A series of repeating units of 2 to 5 nucleotides is called a short tandem repeat (STR) The repeat number for STRs can vary among sites (within a genome) or individuals Simple sequence DNA is common in centromeres and telomeres, where it probably plays structural roles in the chromosome
128 CH 18 Gene Regulation
129 Overview: Conducting the Genetic Orchestra Prokaryotes and eukaryotes alter gene expression in response to their changing environment In multicellular eukaryotes, gene expression regulates development and is responsible for differences in cell types RNA molecules play many roles in regulating gene expression in eukaryotes
130 Concept 18.1: Bacteria often respond to environmental change by regulating transcription Natural selection has favored bacteria that produce only the products needed by that cell A cell can regulate the production of enzymes by feedback inhibition or by gene regulation Gene expression in bacteria is controlled by the operon model
131 Figure 18.2 Feedback inhibition Precursor trpe gene Enzyme 1 trpd gene Regulation of gene expression Enzyme 2 trpc gene Enzyme 3 trpb gene trpa gene Tryptophan (a) Regulation of enzyme activity (b) Regulation of enzyme production
132 Operons: The Basic Concept A cluster of functionally related genes can be under coordinated control by a single on-off switch The regulatory switch is a segment of DNA called an operator usually positioned within the promoter An operon is the entire stretch of DNA that includes the operator, the promoter, and the genes that they control
133 The operon can be switched off by a protein repressor The repressor prevents gene transcription by binding to the operator and blocking RNA polymerase The repressor is the product of a separate regulatory gene
134 The repressor can be in an active or inactive form, depending on the presence of other molecules A corepressor is a molecule that cooperates with a repressor protein to switch an operon off For example, E. coli can synthesize the amino acid tryptophan
135 By default the trp operon is on and the genes for tryptophan synthesis are transcribed When tryptophan is present, it binds to the trp repressor protein, which turns the operon off The repressor is active only in the presence of its corepressor tryptophan; thus the trp operon is turned off (repressed) if tryptophan levels are high
136 Figure 18.3 Promoter Promoter trp operon Genes of operon DNA Regulatory gene mrna 5 trpr 3 RNA polymerase Operator mrna 5 Start codon trpe trpd trpc trpb trpa Stop codon E D C B A Protein Inactive repressor (a) Tryptophan absent, repressor inactive, operon on Polypeptide subunits that make up enzymes for tryptophan synthesis DNA No RNA made mrna Protein Active repressor Tryptophan (corepressor) (b) Tryptophan present, repressor active, operon off
137 Repressible and Inducible Operons: Two Types of Negative Gene Regulation A repressible operon is one that is usually on; binding of a repressor to the operator shuts off transcription The trp operon is a repressible operon An inducible operon is one that is usually off; a molecule called an inducer inactivates the repressor and turns on transcription
138 The lac operon is an inducible operon and contains genes that code for enzymes used in the hydrolysis and metabolism of lactose By itself, the lac repressor is active and switches the lac operon off A molecule called an inducer inactivates the repressor to turn the lac operon on
139 Figure 18.4 Regulatory gene Promoter Operator DNA laci lacz mrna 5 3 RNA polymerase No RNA made Protein Active repressor (a) Lactose absent, repressor active, operon off lac operon DNA laci lacz lacy laca mrna 5 RNA polymerase 3 mrna 5 Protein -Galactosidase Permease Transacetylase Allolactose (inducer) Inactive repressor (b) Lactose present, repressor inactive, operon on
140 Inducible enzymes usually function in catabolic pathways; their synthesis is induced by a chemical signal Repressible enzymes usually function in anabolic pathways; their synthesis is repressed by high levels of the end product Regulation of the trp and lac operons involves negative control of genes because operons are switched off by the active form of the repressor
141 Positive Gene Regulation Some operons are also subject to positive control through a stimulatory protein, such as catabolite activator protein (CAP), an activator of transcription When glucose (a preferred food source of E. coli) is scarce, CAP is activated by binding with cyclic AMP (camp) Activated CAP attaches to the promoter of the lac operon and increases the affinity of RNA polymerase, thus accelerating transcription
142 When glucose levels increase, CAP detaches from the lac operon, and transcription returns to a normal rate CAP helps regulate other operons that encode enzymes used in catabolic pathways
143 Figure 18.5 Promoter DNA laci lacz CAP-binding site camp Active CAP RNA Operator polymerase binds and transcribes Inactive CAP Allolactose Inactive lac repressor (a) Lactose present, glucose scarce (camp level high): abundant lac mrna synthesized Promoter DNA laci lacz CAP-binding site Inactive CAP Operator RNA polymerase less likely to bind Inactive lac repressor (b) Lactose present, glucose present (camp level low): little lac mrna synthesized
Biotechnology. Chapter 20. Biology Eighth Edition Neil Campbell and Jane Reece. PowerPoint Lecture Presentations for
 Chapter 20 Biotechnology PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from Joan Sharp Copyright
Chapter 20 Biotechnology PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from Joan Sharp Copyright
Chapter 20 Biotechnology
 Chapter 20 Biotechnology Manipulation of DNA In 2007, the first entire human genome had been sequenced. The ability to sequence an organisms genomes were made possible by advances in biotechnology, (the
Chapter 20 Biotechnology Manipulation of DNA In 2007, the first entire human genome had been sequenced. The ability to sequence an organisms genomes were made possible by advances in biotechnology, (the
Biotechnology. Chapter 20. Biology Eighth Edition Neil Campbell and Jane Reece. PowerPoint Lecture Presentations for
 Chapter 20 Biotechnology PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from Joan Sharp Copyright
Chapter 20 Biotechnology PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from Joan Sharp Copyright
Biotechnology. Chapter 20. Biology Eighth Edition Neil Campbell and Jane Reece. PowerPoint Lecture Presentations for
 The image cannot be displayed. Your computer may not have enough memory to open the image, or the image may have been corrupted. Restart your computer, and then open the file again. If the red x still
The image cannot be displayed. Your computer may not have enough memory to open the image, or the image may have been corrupted. Restart your computer, and then open the file again. If the red x still
Chapter 20: Biotechnology
 Chapter 20: Biotechnology 1. DNA Sequencing 2. DNA Cloning 3. Studying Gene Expression 4. Manipulating Genomes 5. herapeutic & Diagnostic echniques 1. DNA Sequencing Chapter Reading pp. 409-412 DNA Sequencing
Chapter 20: Biotechnology 1. DNA Sequencing 2. DNA Cloning 3. Studying Gene Expression 4. Manipulating Genomes 5. herapeutic & Diagnostic echniques 1. DNA Sequencing Chapter Reading pp. 409-412 DNA Sequencing
LECTURE PRESENTATIONS
 LECURE PRESENAIONS For CAMPBELL BIOLOY, NINH EDIION Jane B. Reece, Lisa A. Urry, Michael L. Cain, Steven A. Wasserman, Peter V. Minorsky, Robert B. Jackson Chapter 20 Biotechnology Lectures by Erin Barley
LECURE PRESENAIONS For CAMPBELL BIOLOY, NINH EDIION Jane B. Reece, Lisa A. Urry, Michael L. Cain, Steven A. Wasserman, Peter V. Minorsky, Robert B. Jackson Chapter 20 Biotechnology Lectures by Erin Barley
Overview: The DNA Toolbox
 Overview: The DNA Toolbox Sequencing of the genomes of more than 7,000 species was under way in 2010 DNA sequencing has depended on advances in technology, starting with making recombinant DNA In recombinant
Overview: The DNA Toolbox Sequencing of the genomes of more than 7,000 species was under way in 2010 DNA sequencing has depended on advances in technology, starting with making recombinant DNA In recombinant
Biotechnology. Chapter 20. Biology Eighth Edition Neil Campbell and Jane Reece. PowerPoint Lecture Presentations for
 Chapter 20 Biotechnology PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from Joan Sharp Copyright
Chapter 20 Biotechnology PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from Joan Sharp Copyright
Overview: The DNA Toolbox
 Overview: The DNA Toolbox Sequencing of the genomes of more than 7,000 species was under way in 2010 DNA sequencing has depended on advances in technology, starting with making recombinant DNA In recombinant
Overview: The DNA Toolbox Sequencing of the genomes of more than 7,000 species was under way in 2010 DNA sequencing has depended on advances in technology, starting with making recombinant DNA In recombinant
CHAPTER 20 DNA TECHNOLOGY AND GENOMICS. Section A: DNA Cloning
 Section A: DNA Cloning 1. DNA technology makes it possible to clone genes for basic research and commercial applications: an overview 2. Restriction enzymes are used to make recombinant DNA 3. Genes can
Section A: DNA Cloning 1. DNA technology makes it possible to clone genes for basic research and commercial applications: an overview 2. Restriction enzymes are used to make recombinant DNA 3. Genes can
Regulation of Gene Expression
 Chapter 18 Regulation of Gene Expression Edited by Shawn Lester PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley
Chapter 18 Regulation of Gene Expression Edited by Shawn Lester PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley
Biotechnology. Chapter 20. Biology Eighth Edition Neil Campbell and Jane Reece. PowerPoint Lecture Presentations for
 Chapter 20 Biotechnology PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from Joan Sharp Copyright
Chapter 20 Biotechnology PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from Joan Sharp Copyright
AP Biology Gene Expression/Biotechnology REVIEW
 AP Biology Gene Expression/Biotechnology REVIEW Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Gene expression can be a. regulated before transcription.
AP Biology Gene Expression/Biotechnology REVIEW Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Gene expression can be a. regulated before transcription.
Gene Expression. Chapters 11 & 12: Gene Conrtrol and DNA Technology. Cloning. Honors Biology Fig
 Chapters & : Conrtrol and Technology Honors Biology 0 Cloning Produced by asexual reproduction and so it is genetically identical to the parent st large cloned mammal: Dolly the sheep Animals that are
Chapters & : Conrtrol and Technology Honors Biology 0 Cloning Produced by asexual reproduction and so it is genetically identical to the parent st large cloned mammal: Dolly the sheep Animals that are
BIOLOGY. DNA Tools and Biotechnology CAMPBELL. Reece Urry Cain Wasserman Minorsky Jackson
 CAMPBELL BIOLOY TENTH EDITION Reece Urry Cain Wasserman Minorsky Jackson 20 DNA Tools and Biotechnology Lecture Presentation by Nicole Tunbridge and Kathleen Fitzpatrick The DNA Toolbox Recently the genome
CAMPBELL BIOLOY TENTH EDITION Reece Urry Cain Wasserman Minorsky Jackson 20 DNA Tools and Biotechnology Lecture Presentation by Nicole Tunbridge and Kathleen Fitzpatrick The DNA Toolbox Recently the genome
Unit 8: Genomics Guided Reading Questions (150 pts total)
 Name: AP Biology Biology, Campbell and Reece, 7th Edition Adapted from chapter reading guides originally created by Lynn Miriello Chapter 18 The Genetics of Viruses and Bacteria Unit 8: Genomics Guided
Name: AP Biology Biology, Campbell and Reece, 7th Edition Adapted from chapter reading guides originally created by Lynn Miriello Chapter 18 The Genetics of Viruses and Bacteria Unit 8: Genomics Guided
GENE EXPRESSSION. Promoter sequence where RNA polymerase binds. Operator sequence that acts as a switch (yellow) OPERON
 GENE EXPRESSSION 1 GENE REGULATION IN PROKARYOTES Bacteria can turn genes on or off depending on their environment Prokaryotes have operons clusters of related genes and regulatory sequences Promoter sequence
GENE EXPRESSSION 1 GENE REGULATION IN PROKARYOTES Bacteria can turn genes on or off depending on their environment Prokaryotes have operons clusters of related genes and regulatory sequences Promoter sequence
Chapter 11. How Genes Are Controlled. Lectures by Edward J. Zalisko
 Chapter 11 How Genes Are Controlled PowerPoint Lectures for Campbell Essential Biology, Fifth Edition, and Campbell Essential Biology with Physiology, Fourth Edition Eric J. Simon, Jean L. Dickey, and
Chapter 11 How Genes Are Controlled PowerPoint Lectures for Campbell Essential Biology, Fifth Edition, and Campbell Essential Biology with Physiology, Fourth Edition Eric J. Simon, Jean L. Dickey, and
Recombinant DNA recombinant DNA DNA cloning gene cloning
 DNA Technology Recombinant DNA In recombinant DNA, DNA from two different sources, often two species, are combined into the same DNA molecule. DNA cloning permits production of multiple copies of a specific
DNA Technology Recombinant DNA In recombinant DNA, DNA from two different sources, often two species, are combined into the same DNA molecule. DNA cloning permits production of multiple copies of a specific
CHAPTER 21 GENOMES AND THEIR EVOLUTION
 GENETICS DATE CHAPTER 21 GENOMES AND THEIR EVOLUTION COURSE 213 AP BIOLOGY 1 Comparisons of genomes provide information about the evolutionary history of genes and taxonomic groups Genomics - study of
GENETICS DATE CHAPTER 21 GENOMES AND THEIR EVOLUTION COURSE 213 AP BIOLOGY 1 Comparisons of genomes provide information about the evolutionary history of genes and taxonomic groups Genomics - study of
Regulation of Gene Expression
 CAMPBELL BIOLOGY IN FOCUS URRY CAIN WASSERMAN MINORSKY REECE 15 Regulation of Gene Expression Lecture Presentations by Kathleen Fitzpatrick and Nicole Tunbridge, Simon Fraser University SECOND EDITION
CAMPBELL BIOLOGY IN FOCUS URRY CAIN WASSERMAN MINORSKY REECE 15 Regulation of Gene Expression Lecture Presentations by Kathleen Fitzpatrick and Nicole Tunbridge, Simon Fraser University SECOND EDITION
Chapter 20: Biotechnology. 1. DNA Sequencing 3/26/2017. DNA Sequencing. 2. DNA Cloning. 3. Studying Gene Expression. 4. Manipulating Genomes
 hapter 0: Biotechnology 1. DN Sequencing. DN loning 3. Studying ene Expression 4. Manipulating enomes 5. herapeutic & Diagnostic echniques 1. DN Sequencing hapter Reading pp. 409-41 EHNIQUE DN (template
hapter 0: Biotechnology 1. DN Sequencing. DN loning 3. Studying ene Expression 4. Manipulating enomes 5. herapeutic & Diagnostic echniques 1. DN Sequencing hapter Reading pp. 409-41 EHNIQUE DN (template
Biotechnology Chapter 20
 Biotechnology Chapter 20 DNA Cloning DNA Cloning AKA Plasmid-based transformation or molecular cloning First off-let s sum up what happens. A plasmid is taken from a bacteria A gene is inserted into the
Biotechnology Chapter 20 DNA Cloning DNA Cloning AKA Plasmid-based transformation or molecular cloning First off-let s sum up what happens. A plasmid is taken from a bacteria A gene is inserted into the
Chapter 20: Biotechnology
 Name Period The AP Biology exam has reached into this chapter for essay questions on a regular basis over the past 15 years. Student responses show that biotechnology is a difficult topic. This chapter
Name Period The AP Biology exam has reached into this chapter for essay questions on a regular basis over the past 15 years. Student responses show that biotechnology is a difficult topic. This chapter
Campbell Biology 10. Chapter 19 DNA Biotechnology. A Global Approach. Chul-Su Yang, Ph.D., Lecture on General Biology 2
 Lecture on General Biology 2 Campbell Biology 10 A Global Approach th edition Chapter 19 DNA Biotechnology Chul-Su Yang, Ph.D., chulsuyang@hanyang.ac.kr Infection Biology Lab., Dept. of Molecular & Life
Lecture on General Biology 2 Campbell Biology 10 A Global Approach th edition Chapter 19 DNA Biotechnology Chul-Su Yang, Ph.D., chulsuyang@hanyang.ac.kr Infection Biology Lab., Dept. of Molecular & Life
Chapter 12 DNA Technology and Genomics
 Chapter 12 DNA Technology and Genomics PowerPoint Lectures for Campbell Biology: Concepts & Connections, Seventh Edition Reece, Taylor, Simon, and Dickey 2012 Pearson Education, Inc. Lecture by Edward
Chapter 12 DNA Technology and Genomics PowerPoint Lectures for Campbell Biology: Concepts & Connections, Seventh Edition Reece, Taylor, Simon, and Dickey 2012 Pearson Education, Inc. Lecture by Edward
-Is the process of manipulating genes and genomes
 Genetic Engineering -Is the process of manipulating genes and genomes Biotechnology -Is the process of manipulating organisms or their components for the purpose of making useful products Restriction Enzymes
Genetic Engineering -Is the process of manipulating genes and genomes Biotechnology -Is the process of manipulating organisms or their components for the purpose of making useful products Restriction Enzymes
DNA Technology and Genomics
 Chapter 20 DNA echnology and enomics PowerPoint Lectures for Biology, Seventh Edition Neil Campbell and Jane Reece Lectures by Chris Romero Overview: Understanding and Manipulating enomes One of the greatest
Chapter 20 DNA echnology and enomics PowerPoint Lectures for Biology, Seventh Edition Neil Campbell and Jane Reece Lectures by Chris Romero Overview: Understanding and Manipulating enomes One of the greatest
Regulation of Gene Expression
 Slide 1 Chapter 18 Regulation of Gene Expression PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions
Slide 1 Chapter 18 Regulation of Gene Expression PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions
Regulation of metabolic pathways
 Regulation of metabolic pathways Bacterial control of gene expression Operon: cluster of related genes with on/off switch Three Parts: 1. Promoter where RNA polymerase attaches 2. Operator on/off, controls
Regulation of metabolic pathways Bacterial control of gene expression Operon: cluster of related genes with on/off switch Three Parts: 1. Promoter where RNA polymerase attaches 2. Operator on/off, controls
Biotechnolog y and DNA Technology
 PowerPoint Lecture Presentations prepared by Bradley W. Christian, McLennan Community College C H A P T E R 9 Biotechnolog y and DNA Technology Introduction to Biotechnology Biotechnology: the use of microorganisms,
PowerPoint Lecture Presentations prepared by Bradley W. Christian, McLennan Community College C H A P T E R 9 Biotechnolog y and DNA Technology Introduction to Biotechnology Biotechnology: the use of microorganisms,
AP Biology Day 34. Monday, November 14, 2016
 AP Biology Day 34 Monday, November 14, 2016 Essen%al knowledge standards 3.A.1: DNA, and in some cases RNA, is the primary source of heritable informa%on 3.A.1.e: Gene%c engineering techniques can manipulate
AP Biology Day 34 Monday, November 14, 2016 Essen%al knowledge standards 3.A.1: DNA, and in some cases RNA, is the primary source of heritable informa%on 3.A.1.e: Gene%c engineering techniques can manipulate
2054, Chap. 14, page 1
 2054, Chap. 14, page 1 I. Recombinant DNA technology (Chapter 14) A. recombinant DNA technology = collection of methods used to perform genetic engineering 1. genetic engineering = deliberate modification
2054, Chap. 14, page 1 I. Recombinant DNA technology (Chapter 14) A. recombinant DNA technology = collection of methods used to perform genetic engineering 1. genetic engineering = deliberate modification
7.1 Techniques for Producing and Analyzing DNA. SBI4U Ms. Ho-Lau
 7.1 Techniques for Producing and Analyzing DNA SBI4U Ms. Ho-Lau What is Biotechnology? From Merriam-Webster: the manipulation of living organisms or their components to produce useful usually commercial
7.1 Techniques for Producing and Analyzing DNA SBI4U Ms. Ho-Lau What is Biotechnology? From Merriam-Webster: the manipulation of living organisms or their components to produce useful usually commercial
Chapter 10 Genetic Engineering: A Revolution in Molecular Biology
 Chapter 10 Genetic Engineering: A Revolution in Molecular Biology Genetic Engineering Direct, deliberate modification of an organism s genome bioengineering Biotechnology use of an organism s biochemical
Chapter 10 Genetic Engineering: A Revolution in Molecular Biology Genetic Engineering Direct, deliberate modification of an organism s genome bioengineering Biotechnology use of an organism s biochemical
Recombinant DNA. Lesson Overview. Lesson Overview Recombinant DNA
 Lesson Overview 15.2 Finding Genes In 1987, Douglas Prasher, a biologist at Woods Hole Oceanographic Institute in Massachusetts, wanted to find a specific gene in a jellyfish that codes for a molecule
Lesson Overview 15.2 Finding Genes In 1987, Douglas Prasher, a biologist at Woods Hole Oceanographic Institute in Massachusetts, wanted to find a specific gene in a jellyfish that codes for a molecule
Name AP Biology Mrs. Laux Take home test #11 on Chapters 14, 15, and 17 DUE: MONDAY, DECEMBER 21, 2009
 MULTIPLE CHOICE QUESTIONS 1. Inducible genes are usually actively transcribed when: A. the molecule degraded by the enzyme(s) is present in the cell. B. repressor molecules bind to the promoter. C. lactose
MULTIPLE CHOICE QUESTIONS 1. Inducible genes are usually actively transcribed when: A. the molecule degraded by the enzyme(s) is present in the cell. B. repressor molecules bind to the promoter. C. lactose
DNA Technology. B. Using Bacteria to Clone Genes: Overview:
 DNA Technology A. Basic Vocabulary: is DNA from 2 different sources that is combined. is the direct manipulation of genes for practical purposes. literally means or in a test tube or flask. is the manipulation
DNA Technology A. Basic Vocabulary: is DNA from 2 different sources that is combined. is the direct manipulation of genes for practical purposes. literally means or in a test tube or flask. is the manipulation
Chapter 20 Recombinant DNA Technology. Copyright 2009 Pearson Education, Inc.
 Chapter 20 Recombinant DNA Technology Copyright 2009 Pearson Education, Inc. 20.1 Recombinant DNA Technology Began with Two Key Tools: Restriction Enzymes and DNA Cloning Vectors Recombinant DNA refers
Chapter 20 Recombinant DNA Technology Copyright 2009 Pearson Education, Inc. 20.1 Recombinant DNA Technology Began with Two Key Tools: Restriction Enzymes and DNA Cloning Vectors Recombinant DNA refers
2. Outline the levels of DNA packing in the eukaryotic nucleus below next to the diagram provided.
 AP Biology Reading Packet 6- Molecular Genetics Part 2 Name Chapter 19: Eukaryotic Genomes 1. Define the following terms: a. Euchromatin b. Heterochromatin c. Nucleosome 2. Outline the levels of DNA packing
AP Biology Reading Packet 6- Molecular Genetics Part 2 Name Chapter 19: Eukaryotic Genomes 1. Define the following terms: a. Euchromatin b. Heterochromatin c. Nucleosome 2. Outline the levels of DNA packing
BIOLOGY. Chapter 16 GenesExpression
 BIOLOGY Chapter 16 GenesExpression CAMPBELL BIOLOGY TENTH EDITION Reece Urry Cain Wasserman Minorsky Jackson 18 Gene Expression 2014 Pearson Education, Inc. Figure 16.1 Differential Gene Expression results
BIOLOGY Chapter 16 GenesExpression CAMPBELL BIOLOGY TENTH EDITION Reece Urry Cain Wasserman Minorsky Jackson 18 Gene Expression 2014 Pearson Education, Inc. Figure 16.1 Differential Gene Expression results
DNA Technology and Genomics
 DNA Technology and Genomics I. DNA cloning permits production of many copies of a specific gene or other DNA segment. A. To study a particular gene, scientists needed to develop methods to isolate the
DNA Technology and Genomics I. DNA cloning permits production of many copies of a specific gene or other DNA segment. A. To study a particular gene, scientists needed to develop methods to isolate the
Biotechnology and DNA Technology
 11/27/2017 PowerPoint Lecture Presentations prepared by Bradley W. Christian, McLennan Community College CHAPTER 9 Biotechnology and DNA Technology Introduction to Biotechnology Learning Objectives Compare
11/27/2017 PowerPoint Lecture Presentations prepared by Bradley W. Christian, McLennan Community College CHAPTER 9 Biotechnology and DNA Technology Introduction to Biotechnology Learning Objectives Compare
Edexcel (B) Biology A-level
 Edexcel (B) Biology A-level Topic 7: Modern Genetics Notes Using Gene Sequencing Genome = all of an organism s DNA, including mitochondrial/chloroplast DNA. Polymerase chain reaction (PCR) is used to amplify
Edexcel (B) Biology A-level Topic 7: Modern Genetics Notes Using Gene Sequencing Genome = all of an organism s DNA, including mitochondrial/chloroplast DNA. Polymerase chain reaction (PCR) is used to amplify
Chapter 12. DNA Technology. Lectures by Edward J. Zalisko
 Chapter 12 DNA Technology PowerPoint Lectures for Campbell Essential Biology, Fifth Edition, and Campbell Essential Biology with Physiology, Fourth Edition Eric J. Simon, Jean L. Dickey, and Jane B. Reece
Chapter 12 DNA Technology PowerPoint Lectures for Campbell Essential Biology, Fifth Edition, and Campbell Essential Biology with Physiology, Fourth Edition Eric J. Simon, Jean L. Dickey, and Jane B. Reece
Molecular Genetics Techniques. BIT 220 Chapter 20
 Molecular Genetics Techniques BIT 220 Chapter 20 What is Cloning? Recombinant DNA technologies 1. Producing Recombinant DNA molecule Incorporate gene of interest into plasmid (cloning vector) 2. Recombinant
Molecular Genetics Techniques BIT 220 Chapter 20 What is Cloning? Recombinant DNA technologies 1. Producing Recombinant DNA molecule Incorporate gene of interest into plasmid (cloning vector) 2. Recombinant
2014 Pearson Education, Inc. CH 8: Recombinant DNA Technology
 CH 8: Recombinant DNA Technology Biotechnology the use of microorganisms to make practical products Recombinant DNA = DNA from 2 different sources What is Recombinant DNA Technology? modifying genomes
CH 8: Recombinant DNA Technology Biotechnology the use of microorganisms to make practical products Recombinant DNA = DNA from 2 different sources What is Recombinant DNA Technology? modifying genomes
Recombinant DNA Technology. The Role of Recombinant DNA Technology in Biotechnology. yeast. Biotechnology. Recombinant DNA technology.
 PowerPoint Lecture Presentations prepared by Mindy Miller-Kittrell, North Carolina State University C H A P T E R 8 Recombinant DNA Technology The Role of Recombinant DNA Technology in Biotechnology Biotechnology?
PowerPoint Lecture Presentations prepared by Mindy Miller-Kittrell, North Carolina State University C H A P T E R 8 Recombinant DNA Technology The Role of Recombinant DNA Technology in Biotechnology Biotechnology?
Unit 6: Molecular Genetics & DNA Technology Guided Reading Questions (100 pts total)
 Name: AP Biology Biology, Campbell and Reece, 7th Edition Adapted from chapter reading guides originally created by Lynn Miriello Chapter 16 The Molecular Basis of Inheritance Unit 6: Molecular Genetics
Name: AP Biology Biology, Campbell and Reece, 7th Edition Adapted from chapter reading guides originally created by Lynn Miriello Chapter 16 The Molecular Basis of Inheritance Unit 6: Molecular Genetics
A Lot of Cutting and Pasting Going on Here Recombinant DNA and Biotechnology
 A Lot of Cutting and Pasting Going on Here Recombinant DNA and Biotechnology How Are Large DNA Molecules Analyzed? Naturally occurring enzymes that cleave and repair DNA are used in the laboratory to manipulate
A Lot of Cutting and Pasting Going on Here Recombinant DNA and Biotechnology How Are Large DNA Molecules Analyzed? Naturally occurring enzymes that cleave and repair DNA are used in the laboratory to manipulate
4/26/2015. Cut DNA either: Cut DNA either:
 Ch.20 Enzymes that cut DNA at specific sequences (restriction sites) resulting in segments of DNA (restriction fragments) Typically 4-8 bp in length & often palindromic Isolated from bacteria (Hundreds
Ch.20 Enzymes that cut DNA at specific sequences (restriction sites) resulting in segments of DNA (restriction fragments) Typically 4-8 bp in length & often palindromic Isolated from bacteria (Hundreds
Chapter 18: Regulation of Gene Expression. 1. Gene Regulation in Bacteria 2. Gene Regulation in Eukaryotes 3. Gene Regulation & Cancer
 Chapter 18: Regulation of Gene Expression 1. Gene Regulation in Bacteria 2. Gene Regulation in Eukaryotes 3. Gene Regulation & Cancer Gene Regulation Gene regulation refers to all aspects of controlling
Chapter 18: Regulation of Gene Expression 1. Gene Regulation in Bacteria 2. Gene Regulation in Eukaryotes 3. Gene Regulation & Cancer Gene Regulation Gene regulation refers to all aspects of controlling
CH 8: Recombinant DNA Technology
 CH 8: Recombinant DNA Technology Biotechnology the use of microorganisms to make practical products Recombinant DNA = DNA from 2 different sources What is Recombinant DNA Technology? modifying genomes
CH 8: Recombinant DNA Technology Biotechnology the use of microorganisms to make practical products Recombinant DNA = DNA from 2 different sources What is Recombinant DNA Technology? modifying genomes
Biotechnology DNA technology
 Biotechnology Biotechnology is the manipulation of organisms or their components to make useful products The applications of DNA technology affect everything from agriculture, to criminal law, to medical
Biotechnology Biotechnology is the manipulation of organisms or their components to make useful products The applications of DNA technology affect everything from agriculture, to criminal law, to medical
Virus- infectious particle consisting of nucleic acid packaged in a protein coat.
 Chapter 19 Virus- infectious particle consisting of nucleic acid packaged in a protein coat. Most scientists consider viruses non-living because they cannot reproduce or carry out metabolic activities
Chapter 19 Virus- infectious particle consisting of nucleic acid packaged in a protein coat. Most scientists consider viruses non-living because they cannot reproduce or carry out metabolic activities
Guided Notes Unit 5: Molecular Genetics
 Name: Date: Block: Chapter 8: From DNA to Protein I. Concept 8.4: Transcription a. Central Dogma of Molecular Biology i. Information flows in one direction: ii. How? Guided Notes Unit 5: Molecular Genetics
Name: Date: Block: Chapter 8: From DNA to Protein I. Concept 8.4: Transcription a. Central Dogma of Molecular Biology i. Information flows in one direction: ii. How? Guided Notes Unit 5: Molecular Genetics
13-1 Changing the Living World
 13-1 Changing the Living World In the past, variation was limited to the variations already in nature or random variations that resulted from mutations. Now, scientists can change DNA and swap genes from
13-1 Changing the Living World In the past, variation was limited to the variations already in nature or random variations that resulted from mutations. Now, scientists can change DNA and swap genes from
Cloning and Genetic Engineering
 Cloning and Genetic Engineering Bởi: OpenStaxCollege Biotechnology is the use of artificial methods to modify the genetic material of living organisms or cells to produce novel compounds or to perform
Cloning and Genetic Engineering Bởi: OpenStaxCollege Biotechnology is the use of artificial methods to modify the genetic material of living organisms or cells to produce novel compounds or to perform
Name Biol Group Number. ALE 11. The Genetics of Viruses, Control of Gene Expression, and Recombinant DNA Technology
 Name Biol 211 - Group Number ALE 11. The Genetics of Viruses, Control of Gene Expression, and Recombinant DNA Technology Chapter 19: The Genetics of Viruses (pp. 381-395, Biology by Campbell/Reece, 8 th
Name Biol 211 - Group Number ALE 11. The Genetics of Viruses, Control of Gene Expression, and Recombinant DNA Technology Chapter 19: The Genetics of Viruses (pp. 381-395, Biology by Campbell/Reece, 8 th
Concept 13.1 Recombinant DNA Can Be Made in the Laboratory
 13 Biotechnology Concept 13.1 Recombinant DNA Can Be Made in the Laboratory It is possible to modify organisms with genes from other, distantly related organisms. Recombinant DNA is a DNA molecule made
13 Biotechnology Concept 13.1 Recombinant DNA Can Be Made in the Laboratory It is possible to modify organisms with genes from other, distantly related organisms. Recombinant DNA is a DNA molecule made
Chapter 9 Genetic Engineering
 Chapter 9 Genetic Engineering Biotechnology: use of microbes to make a protein product Recombinant DNA Technology: Insertion or modification of genes to produce desired proteins Genetic engineering: manipulation
Chapter 9 Genetic Engineering Biotechnology: use of microbes to make a protein product Recombinant DNA Technology: Insertion or modification of genes to produce desired proteins Genetic engineering: manipulation
Genetics Part 2B. AP Biology. Repressible Operon. Bacterial control of gene expression Operon: cluster of related genes with on/off switch
 Regulation of metabolic pathways Bacterial control of gene expression Operon: cluster of related genes with on/off switch Genetics Part 2B Three Parts: 1. Promoter where RNA polymerase attaches 2. Operator
Regulation of metabolic pathways Bacterial control of gene expression Operon: cluster of related genes with on/off switch Genetics Part 2B Three Parts: 1. Promoter where RNA polymerase attaches 2. Operator
GENE REGULATION IN PROKARYOTES
 GENE REGULATION IN PROKARYOTES Prepared by Brenda Leady, University of Toledo Copyright (c) The McGraw-Hill Companies, Inc. Permission required for reproduction or display. 1 Gene regulation refers to
GENE REGULATION IN PROKARYOTES Prepared by Brenda Leady, University of Toledo Copyright (c) The McGraw-Hill Companies, Inc. Permission required for reproduction or display. 1 Gene regulation refers to
BIOLOGY - CLUTCH CH.20 - BIOTECHNOLOGY.
 !! www.clutchprep.com CONCEPT: DNA CLONING DNA cloning is a technique that inserts a foreign gene into a living host to replicate the gene and produce gene products. Transformation the process by which
!! www.clutchprep.com CONCEPT: DNA CLONING DNA cloning is a technique that inserts a foreign gene into a living host to replicate the gene and produce gene products. Transformation the process by which
Bio 101 Sample questions: Chapter 10
 Bio 101 Sample questions: Chapter 10 1. Which of the following is NOT needed for DNA replication? A. nucleotides B. ribosomes C. Enzymes (like polymerases) D. DNA E. all of the above are needed 2 The information
Bio 101 Sample questions: Chapter 10 1. Which of the following is NOT needed for DNA replication? A. nucleotides B. ribosomes C. Enzymes (like polymerases) D. DNA E. all of the above are needed 2 The information
Biosc10 schedule reminders
 Biosc10 schedule reminders Review of molecular biology basics DNA Is each person s DNA the same, or unique? What does DNA look like? What are the three parts of each DNA nucleotide Which DNA bases pair,
Biosc10 schedule reminders Review of molecular biology basics DNA Is each person s DNA the same, or unique? What does DNA look like? What are the three parts of each DNA nucleotide Which DNA bases pair,
Chapter 20 DNA Technology & Genomics. If we can, should we?
 Chapter 20 DNA Technology & Genomics If we can, should we? Biotechnology Genetic manipulation of organisms or their components to make useful products Humans have been doing this for 1,000s of years plant
Chapter 20 DNA Technology & Genomics If we can, should we? Biotechnology Genetic manipulation of organisms or their components to make useful products Humans have been doing this for 1,000s of years plant
BIOTECHNOLOGY. Sticky & blunt ends. Restriction endonucleases. Gene cloning an overview. DNA isolation & restriction
 BIOTECHNOLOGY RECOMBINANT DNA TECHNOLOGY Recombinant DNA technology involves sticking together bits of DNA from different sources. Made possible because DNA & the genetic code are universal. 2004 Biology
BIOTECHNOLOGY RECOMBINANT DNA TECHNOLOGY Recombinant DNA technology involves sticking together bits of DNA from different sources. Made possible because DNA & the genetic code are universal. 2004 Biology
The Biotechnology Toolbox
 Chapter 15 The Biotechnology Toolbox Cutting and Pasting DNA Cutting DNA Restriction endonuclease or restriction enzymes Cellular protection mechanism for infected foreign DNA Recognition and cutting specific
Chapter 15 The Biotechnology Toolbox Cutting and Pasting DNA Cutting DNA Restriction endonuclease or restriction enzymes Cellular protection mechanism for infected foreign DNA Recognition and cutting specific
Unit 6: Molecular Genetics & DNA Technology Guided Reading Questions (100 pts total)
 AP Biology Biology, Campbell and Reece, 10th Edition Adapted from chapter reading guides originally created by Lynn Miriello Name: Chapter 16 The Molecular Basis of Inheritance Concept 16.1 DNA is the
AP Biology Biology, Campbell and Reece, 10th Edition Adapted from chapter reading guides originally created by Lynn Miriello Name: Chapter 16 The Molecular Basis of Inheritance Concept 16.1 DNA is the
CHAPTERS 16 & 17: DNA Technology
 CHAPTERS 16 & 17: DNA Technology 1. What is the function of restriction enzymes in bacteria? 2. How do bacteria protect their DNA from the effects of the restriction enzymes? 3. How do biologists make
CHAPTERS 16 & 17: DNA Technology 1. What is the function of restriction enzymes in bacteria? 2. How do bacteria protect their DNA from the effects of the restriction enzymes? 3. How do biologists make
Chapter 15 Gene Technologies and Human Applications
 Chapter Outline Chapter 15 Gene Technologies and Human Applications Section 1: The Human Genome KEY IDEAS > Why is the Human Genome Project so important? > How do genomics and gene technologies affect
Chapter Outline Chapter 15 Gene Technologies and Human Applications Section 1: The Human Genome KEY IDEAS > Why is the Human Genome Project so important? > How do genomics and gene technologies affect
Genetics Lecture 21 Recombinant DNA
 Genetics Lecture 21 Recombinant DNA Recombinant DNA In 1971, a paper published by Kathleen Danna and Daniel Nathans marked the beginning of the recombinant DNA era. The paper described the isolation of
Genetics Lecture 21 Recombinant DNA Recombinant DNA In 1971, a paper published by Kathleen Danna and Daniel Nathans marked the beginning of the recombinant DNA era. The paper described the isolation of
Biology 201 (Genetics) Exam #3 120 points 20 November Read the question carefully before answering. Think before you write.
 Name KEY Section Biology 201 (Genetics) Exam #3 120 points 20 November 2006 Read the question carefully before answering. Think before you write. You will have up to 50 minutes to take this exam. After
Name KEY Section Biology 201 (Genetics) Exam #3 120 points 20 November 2006 Read the question carefully before answering. Think before you write. You will have up to 50 minutes to take this exam. After
CHAPTER 9 DNA Technologies
 CHAPTER 9 DNA Technologies Recombinant DNA Artificially created DNA that combines sequences that do not occur together in the nature Basis of much of the modern molecular biology Molecular cloning of genes
CHAPTER 9 DNA Technologies Recombinant DNA Artificially created DNA that combines sequences that do not occur together in the nature Basis of much of the modern molecular biology Molecular cloning of genes
2 Gene Technologies in Our Lives
 CHAPTER 15 2 Gene Technologies in Our Lives SECTION Gene Technologies and Human Applications KEY IDEAS As you read this section, keep these questions in mind: For what purposes are genes and proteins manipulated?
CHAPTER 15 2 Gene Technologies in Our Lives SECTION Gene Technologies and Human Applications KEY IDEAS As you read this section, keep these questions in mind: For what purposes are genes and proteins manipulated?
Genetic Engineering & Recombinant DNA
 Genetic Engineering & Recombinant DNA Chapter 10 Copyright The McGraw-Hill Companies, Inc) Permission required for reproduction or display. Applications of Genetic Engineering Basic science vs. Applied
Genetic Engineering & Recombinant DNA Chapter 10 Copyright The McGraw-Hill Companies, Inc) Permission required for reproduction or display. Applications of Genetic Engineering Basic science vs. Applied
Section B: The Genetics of Bacteria
 CHAPTER 18 MICROBIAL MODELS: THE GENETICS OF VIRUSES AND BACTERIA Section B: The Genetics of Bacteria 1. The short generation span of bacteria helps them adapt to changing environments 2. Genetic recombination
CHAPTER 18 MICROBIAL MODELS: THE GENETICS OF VIRUSES AND BACTERIA Section B: The Genetics of Bacteria 1. The short generation span of bacteria helps them adapt to changing environments 2. Genetic recombination
Chapter 6 - Molecular Genetic Techniques
 Chapter 6 - Molecular Genetic Techniques Two objects of molecular & genetic technologies For analysis For generation Molecular genetic technologies! For analysis DNA gel electrophoresis Southern blotting
Chapter 6 - Molecular Genetic Techniques Two objects of molecular & genetic technologies For analysis For generation Molecular genetic technologies! For analysis DNA gel electrophoresis Southern blotting
Unit 2: Metabolism and Survival Sub-Topic (2.7) Genetic Control of Metabolism (2.8) Ethical considerations in the use of microorganisms
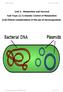 Unit 2: Metabolism and Survival Sub-Topic (2.7) Genetic Control of Metabolism (2.8) Ethical considerations in the use of microorganisms Duncanrig Secondary JHM&MHC 2015 Page 1 of 18 On completion of this
Unit 2: Metabolism and Survival Sub-Topic (2.7) Genetic Control of Metabolism (2.8) Ethical considerations in the use of microorganisms Duncanrig Secondary JHM&MHC 2015 Page 1 of 18 On completion of this
What is Biotechnology? 15.1 What is Biotechnology? Transgenic Biotechnology Transgenic Biotechnology. Biotechnology. Transgenic organism
 What is Biotechnology? 15.1 What is Biotechnology? Biotechnology the use of technology to control biological processes as a means of meeting societal needs Gene therapy Genetic engineering Bioremediation
What is Biotechnology? 15.1 What is Biotechnology? Biotechnology the use of technology to control biological processes as a means of meeting societal needs Gene therapy Genetic engineering Bioremediation
DNA and Biotechnology Form of DNA Form of DNA Form of DNA Form of DNA Replication of DNA Replication of DNA
 21 DNA and Biotechnology DNA and Biotechnology OUTLINE: Replication of DNA Gene Expression Mutations Regulating Gene Activity Genetic Engineering Genomics DNA (deoxyribonucleic acid) Double-stranded molecule
21 DNA and Biotechnology DNA and Biotechnology OUTLINE: Replication of DNA Gene Expression Mutations Regulating Gene Activity Genetic Engineering Genomics DNA (deoxyribonucleic acid) Double-stranded molecule
Regulation of enzyme synthesis
 Regulation of enzyme synthesis The lac operon is an example of an inducible operon - it is normally off, but when a molecule called an inducer is present, the operon turns on. The trp operon is an example
Regulation of enzyme synthesis The lac operon is an example of an inducible operon - it is normally off, but when a molecule called an inducer is present, the operon turns on. The trp operon is an example
NOTES - CH 15 (and 14.3): DNA Technology ( Biotech )
 NOTES - CH 15 (and 14.3): DNA Technology ( Biotech ) Vocabulary Genetic Engineering Gene Recombinant DNA Transgenic Restriction Enzymes Vectors Plasmids Cloning Key Concepts What is genetic engineering?
NOTES - CH 15 (and 14.3): DNA Technology ( Biotech ) Vocabulary Genetic Engineering Gene Recombinant DNA Transgenic Restriction Enzymes Vectors Plasmids Cloning Key Concepts What is genetic engineering?
M I C R O B I O L O G Y WITH DISEASES BY TAXONOMY, THIRD EDITION
 M I C R O B I O L O G Y WITH DISEASES BY TAXONOMY, THIRD EDITION Chapter 7 Microbial Genetics Lecture prepared by Mindy Miller-Kittrell, University of Tennessee, Knoxville The Structure and Replication
M I C R O B I O L O G Y WITH DISEASES BY TAXONOMY, THIRD EDITION Chapter 7 Microbial Genetics Lecture prepared by Mindy Miller-Kittrell, University of Tennessee, Knoxville The Structure and Replication
Chapter 11: Applications of Biotechnology
 Chapter 11: Applications of Biotechnology Lecture Outline Enger, E. D., Ross, F. C., & Bailey, D. B. (2012). Concepts in biology (14th ed.). New York: McGraw- Hill. 11-1 Why Biotechnology Works 11-2 Biotechnology
Chapter 11: Applications of Biotechnology Lecture Outline Enger, E. D., Ross, F. C., & Bailey, D. B. (2012). Concepts in biology (14th ed.). New York: McGraw- Hill. 11-1 Why Biotechnology Works 11-2 Biotechnology
March 15, Genetics_of_Viruses_and_Bacteria_p5.notebook. smallest viruses are smaller than ribosomes. A virulent phage (Lytic)
 Genetics_of_Viruses_and_Bacteria_p5.notebook smallest viruses are smaller than ribosomes Adenovirus Tobacco mosaic virus Bacteriophage Influenza virus envelope is derived from the host cell The capsids
Genetics_of_Viruses_and_Bacteria_p5.notebook smallest viruses are smaller than ribosomes Adenovirus Tobacco mosaic virus Bacteriophage Influenza virus envelope is derived from the host cell The capsids
UNIT III: Genetics Chapter 9 Frontiers of Biotechnology
 UNIT III: Genetics Chapter 9 Frontiers of Biotechnology I. Manipulating DNA (9.1) A. Scientists use several techniques to manipulate DNA 1. DNA is a very large molecule 2. Still to small to see or work
UNIT III: Genetics Chapter 9 Frontiers of Biotechnology I. Manipulating DNA (9.1) A. Scientists use several techniques to manipulate DNA 1. DNA is a very large molecule 2. Still to small to see or work
CHAPTER 08: RECOMBINANT DNA TECHNOLOGY Pearson Education, Inc.
 CHAPTER 08: RECOMBINANT DNA TECHNOLOGY The Role of Recombinant DNA Technology in Biotechnology Biotechnology the use of microorganisms to make practical products Recombinant DNA technology Intentionally
CHAPTER 08: RECOMBINANT DNA TECHNOLOGY The Role of Recombinant DNA Technology in Biotechnology Biotechnology the use of microorganisms to make practical products Recombinant DNA technology Intentionally
Studying the Human Genome. Lesson Overview. Lesson Overview Studying the Human Genome
 Lesson Overview 14.3 Studying the Human Genome THINK ABOUT IT Just a few decades ago, computers were gigantic machines found only in laboratories and universities. Today, many of us carry small, powerful
Lesson Overview 14.3 Studying the Human Genome THINK ABOUT IT Just a few decades ago, computers were gigantic machines found only in laboratories and universities. Today, many of us carry small, powerful
Motivation From Protein to Gene
 MOLECULAR BIOLOGY 2003-4 Topic B Recombinant DNA -principles and tools Construct a library - what for, how Major techniques +principles Bioinformatics - in brief Chapter 7 (MCB) 1 Motivation From Protein
MOLECULAR BIOLOGY 2003-4 Topic B Recombinant DNA -principles and tools Construct a library - what for, how Major techniques +principles Bioinformatics - in brief Chapter 7 (MCB) 1 Motivation From Protein
12/31/16. I. Manipulating DNA (9.1) A. Scientists use several techniques to manipulate DNA. 1. DNA is a very large molecule
 I. Manipulating DNA (9.1) A. Scientists use several techniques to manipulate DNA 1. DNA is a very large molecule 3. Led to many biotechnology applications- genetic engineering, DNA fingerprinting, cloning,
I. Manipulating DNA (9.1) A. Scientists use several techniques to manipulate DNA 1. DNA is a very large molecule 3. Led to many biotechnology applications- genetic engineering, DNA fingerprinting, cloning,
Copyright 2002 Pearson Education, Inc., publishing as Benjamin Cummings
 Here s one thing genetic engineers do: Techniques for gene cloning enable scientists to prepare multiple identical copies of gene-sized pieces of DNA. Cloning means to make copies, in this case, copies
Here s one thing genetic engineers do: Techniques for gene cloning enable scientists to prepare multiple identical copies of gene-sized pieces of DNA. Cloning means to make copies, in this case, copies
BIOLOGY. DNA Tools and Biotechnology CAMPBELL. Reece Urry Cain Wasserman Minorsky Jackson
 CMPBELL BIOLOY ENH EDIION Reece Urry Cain Wasserman Minorsky Jackson 20 DN ools and Biotechnology Lecture Presentation by Nicole unbridge and Kathleen Fitzpatrick he DN oolbox Recently the genome sequences
CMPBELL BIOLOY ENH EDIION Reece Urry Cain Wasserman Minorsky Jackson 20 DN ools and Biotechnology Lecture Presentation by Nicole unbridge and Kathleen Fitzpatrick he DN oolbox Recently the genome sequences
BIOLOGY. DNA Tools and Biotechnology CAMPBELL. Reece Urry Cain Wasserman Minorsky Jackson
 CMPBELL BIOLOY ENH EDIION Reece Urry Cain Wasserman Minorsky Jackson 20 DN ools and Biotechnology Lecture Presentation by Nicole unbridge and Kathleen Fitzpatrick he DN oolbox Recently the genome sequences
CMPBELL BIOLOY ENH EDIION Reece Urry Cain Wasserman Minorsky Jackson 20 DN ools and Biotechnology Lecture Presentation by Nicole unbridge and Kathleen Fitzpatrick he DN oolbox Recently the genome sequences
Enzyme that uses RNA as a template to synthesize a complementary DNA
 Biology 105: Introduction to Genetics PRACTICE FINAL EXAM 2006 Part I: Definitions Homology: Comparison of two or more protein or DNA sequence to ascertain similarities in sequences. If two genes have
Biology 105: Introduction to Genetics PRACTICE FINAL EXAM 2006 Part I: Definitions Homology: Comparison of two or more protein or DNA sequence to ascertain similarities in sequences. If two genes have
Name Per AP: CHAPTER 27: PROKARYOTES (Bacteria) p559,
 AP: CHAPTER 27: PROKARYOTES (Bacteria) p559, 561-564 1. How does the bacterial chromosome compare to a eukaryotic chromosome? 2. What is a plasmid? 3. How fast can bacteria reproduce? 4. What is a bacterial
AP: CHAPTER 27: PROKARYOTES (Bacteria) p559, 561-564 1. How does the bacterial chromosome compare to a eukaryotic chromosome? 2. What is a plasmid? 3. How fast can bacteria reproduce? 4. What is a bacterial
AGRO/ANSC/BIOL/GENE/HORT 305 Fall, 2017 Recombinant DNA Technology (Chpt 20, Genetics by Brooker) Lecture outline: (#14)
 AGRO/ANSC/BIOL/GENE/HORT 305 Fall, 2017 Recombinant DNA Technology (Chpt 20, Genetics by Brooker) Lecture outline: (#14) - RECOMBINANT DNA TECHNOLOGY is the use of in vitro molecular techniques to isolate
AGRO/ANSC/BIOL/GENE/HORT 305 Fall, 2017 Recombinant DNA Technology (Chpt 20, Genetics by Brooker) Lecture outline: (#14) - RECOMBINANT DNA TECHNOLOGY is the use of in vitro molecular techniques to isolate
