Use of Spike-ins for Sample Tracking in Agilent Array CGH
|
|
|
- Rodger Carter
- 5 years ago
- Views:
Transcription
1 Use of Spike-ins for Sample Tracking in Agilent Array CGH Application Note Authors Srirangan Sampath, Ph.D., FACMG and Heather Klein PreventionGenetics Iman Kishawi, Ph.D. Agilent Technologies, Inc. Abstract This Application Note describes a method that allows sample tracking through introduction of DNA spike-ins to human DNA samples before wet lab handling. Adding the spike-ins to samples ensures the identity of the sample throughout the process, and reduces sample mix-up in high-throughput DNA microarrays. In principle, this method can be applied to many other molecular genetic methodologies including NGS. DNA spike-ins contain one or more PCR-generated amplicons that are approximately 400 bp in length. The amplicons are designed to span sequences of unique and nondisease associated regions in the human genome. Agilent CytoGenomics software identifi es sample identity by detecting the signal intensity of spike-in probes in individual arrays.
2 Introduction Many laboratories process multiple samples on any given day, and human error can contribute to mistakes in the processing and reporting of results for samples. Sample mix-up errors constitute the most serious pre-analytic mistakes 1-. In addition, the chances of a specimen mix-up during the analysis process increases with the number of experimental steps (multistep or multiday experiments). DNA microarray applications, such as array comparative genomic hybridization (acgh), are a multistep process. The wet lab portion includes restriction-digestion (for arrays containing SNP probes), fluorescent-labeling of genomic DNA (gdna), cleanup and quantification of labeled gdna, and hybridization of labeled gdna to microarray slides. Sample mix-up at any of these stages can lead to erroneous interpretation of results. Human error can also occur with data analysis if sample attributes are associated with the wrong array. A spike-in is a calibrated reference standard made up of one or more DNA amplicons of known concentn and identity. Amplicons are added in large excess to each DNA sample being tested. Fluorescent labeling of the spiked amplicon generates a bright fluorescent signal after hybridizing to probes that correspond to the genomic loci of spike-ins. Agilent CytoGenomics software v4 or higher reports the identity of the spike-in in each sample. This reporting is done by calculating the log of the sample signal to the reference signal at any given probe. The software identifies the identity of the spike-in based on a default log equal to, or greater than one. A default threshold of one corresponds to two times more in a spiked DNA sample versus a reference DNA sample where no spike-ins were added. The spike-in identity is entered into the CytoGenomics software by the user, and the software reports if the spike-in identity does or does not match the user-inputted sample attribute. Methods Reagents and supplies needed Spike-in identity The 50 unique amplicons (Table 1) are PCR-generated using the forward and reverse primers shown in the table. The probes corresponding to each amplicon (Table 1) are printed on the CGH arrays. The software recognizes and identifies them by their probe names. Individual probes that hybridize to the spike-ins are printed 5 times on the array. The data presented in Figure 1 show the average log of spike-ins from replicated probes. Spike-in probes are part of the HD Agilent probe database. Spike-in controls can be added to any array by selecting the spike-in probe group in SureDesign, and replicating it 5 times depending on available space on the array. Material Vendor Part number KAPA HS MasterMix Kapa Biosystems KK1606 Oligonucleotide primer mixes IDT Custom (5 nmole DNA oligo) Nanodrop Thermo Scientific Human Reference Genomic DNA Agilent European Male X TE buffer IDT N/A Eppendorf Mastercycler ProS Eppendorf Agilent TapeStation 00 Agilent Technologies G964AA Agilent D1000 ScreenTape Agilent Technologies
3 Table 1. Forward and reverse primers for each of the 50 DNA amplicons that are used as spike-ins. The expected sizes for each amplicon, and the Agilent probe names are provided. Band Probe_name FP_seq RP_seq Amplicon size 1q.1 A_16_P5405 CATTCAATGTTACCTTCCAA AGCCACTCATTTTTAACACC 487 A_16_P AATGTACAAACTGGAAACC TTTCTCTCTCACAGGAATAA 4 A_16_P TTCTCCTTAAGCTGGTTTAA GGAAAACTGGTTCAGTAGAC 500 q1. A_18_P15801 TTATCTCTTGCCTTCTTTTC AGACAATAGACTGCAAATCC 50 A_16_P CCTACCACCTTGTGAAGA GCACTGTACACTGAAAATTG 45 q1. A_16_P CCAAGATGCTTAGAAAAATG GGCATCAAATCTCTGGTT 444 A_16_P AGCTTCAGGTAAGTTTTTGG ACACAGAAGCATCTGTTCTT 451 A_18_P TTACTCCCAAAGTCTATTGC TGTAACCAGTGAGATAAGGC 475 4q1. A_18_P GGTTGTGGGTTTACATTCAA AAGGGTGACCTAGAGCACA 48 A_18_P GAATCTGAAGACAGTCCAGT TTTAACAACCCTTCACTTC 516 A_16_P GAACTCAGTGCATGAACAAG TCTGAGGACAGAGCTGTATC 5q14. A_18_P GCTGTTCTCCAAACACTAAT TTACATCGAGCTAACCTGTC 41 A_18_P1567 TTCATTCAAGAGCACAGAAA ACAAACTAGGGCAAAACAG 49 A_16_P75066 GTCATTTAAATTTTCTTGGC TAATCTAGGTGGTGGTTACC 487 6q14. A_18_P15761 CAGAAGCTCTTTGGTTTAAT GCTGCTTTACAAAATACTGC 41 A_18_P CCTTGGGTTCAATGAAACCT GGCACAGAAGAAAGGGACAG 95 A_14_P GAGCCCACCCTATAAACATT CAGATCTTGGCAAGACTCAG 44 8q. A_16_P CCTTTAGGAATTACCCAGTA TTAGGAAGCAGTAATGAGG 465 A_16_P GTTGACATGCTGGGAAATC TGCCCATACAGCTGTTCTCT 64 A_16_P AGTTGTTTCAGAAGGCCTCT AGTGACAGGAATGAGGCTG 66 9q1. A_18_P GATGTGCACACATGGACC AGACATCTGCTCAGGGAGTT 47 A_16_P CAATCTAAAATCAAGGAAAT ATACAATGTGACAGATGCTA 55 A_16_P TTGTTTGCCTAGAAAACAGG TCACTTTTTCAATCCCACTT 45 10q5.1 A_16_P GGATATTTCTGTCCATTGAT CACAGTGGTATTAAACAAGG 47 A_18_P GGCATTGTCTCTTCTTGCTA TAACAGAACCCAAACTCGTG 47 A_16_P TCATAATTACCATCATAGCA AAATCAGCGATATTAATAGC q.1 A_16_P GGCAAAAGAGTGAGACTC CCTTCCCTTAATAGCCTAAG 468 A_16_P AGATAGCCTGCTAGGATTTG GTGATGGATACCCCATTTAC 41 A_18_P CCATTACTTCATTTCTGCCT TTGGTCACAGAACATTGAAT q1. A_16_P TTATTTCCTTGCTATAGTGG GAAACAACTCAGAAAGCTAG 64 A_16_P TGGAACAAGAACTTGTGAGC GCCAAAATGCATAATTTCA 471 A_18_P TCTCTGTGCATCACCCTATT GGCAAAGTTCAAGTCTCTGG q1.1 A_18_P GATTCCCGTCTGTGTGGTCT TCTTATTTTTGACTCTGAGATTGTGG 51 A_18_P10640 TGGTGCCTAAGAAAGACATTCA TTCCTTCCCAAAGTGGCTTA 87 A_16_P TCCCAAAGAATTGTCAATGG TTTGGTTTCCCATCAAGGTC 00 16q1 A_18_P AACATTGAGGATGGAGCAG CAAAAATGGGGTCTTATCTG 41 A_18_P14805 TGTCAATGTAAAAATGCAAG ACAGCAAAGGTCTTCTATTC 46 A_16_P GTAACAGAGGGCTAGTCTTG AGTTTGAGTGGTGACTGATT q1. A_18_P18458 GCCTTTATTGATTCCTGCTT TGGGGCTCACCTAAAAGTT 5 A_18_P GGTGTCTCATCAGCAGGG GATTCCTATTTTGCTCAGGC 459 A_18_P CCTTAGGTCCACATGACATG GTGCAGTCAAGCTTGAAGTC 45 19q1 A_16_P AAGTTTCCTTTTGATTAGGT ACTAGGAGAAAATATTTGCC 4 A_16_P AACAAACAAACAAAAAACGC CACAGCTTCTTCATCTGACC 41 A_18_P10145 GAAGAGGTCACATATGTGAG ATTACCAAACAAGATTGTTG 97 0q1 A_18_P GCATTCCAGCTTAAAGGA ACATTTAAAAGATGGTGCAA 408 A_18_P GCATAGCCTTGAAAAAATCC TGTGGAATGGATAAATTGCC 48 A_16_P TTCATGGACTCAATAGTCCC TGAGCTTGAAGAGGAAAGAG 4 1q1. A_16_P CCTGTCAGCAGGTAGAAGGA TCTCATGAAGTAGGCAGAGGAA 56 A_16_P TTTTGCAAGTAAAGCAATTTTAGG GCTCTGCTGGACACCTTGAT 99 A_18_P CAAATTGATGAAATCCTTATTTGC AAACCATCAAATACAGGAAAAGAAA 86
4 Figure 1. Spike-in M is assigned three probes. Three different amplicons can be spiked into a DNA sample. CytoGenomics software can identify the identity of the three amplicons as Spike-in M. 4
5 Spike-in genen DNA spike-ins are amplicons generated from PCR of human genomic sequences using a reference human genome (for example, Agilent European Male). Tables and provide the conditions of the PCR reactions. The amplification efficiency differs for each of the 50 amplicons, and may need some level of PCR optimization in the hands of users. The PCR efficiency and identity of the amplicons were checked by running -µl aliquots of all amplicons on an Agilent TapeStation 00 using Agilent D1000 ScreenTape. The quality and identity of each amplicon were confirmed by the presence of a correct sized single band. The PCR products were purified using the Qiagen QIAquick PCR Purification Kit (catalog # 8106) using the manufacturer s protocol. Table. Setup of PCR reactions. Reagents Volume/rxn (µl) KAPA HS MasterMix 0.5 Amplicon F/R Primers, 5 µm.75 Template, 5 ng/µl 6 Total reaction volume 0 Table. PCR Conditions are listed below (7 cycles). Conditions Time 95 C minutes 95 C 0 seconds 55 C 0 seconds 7 C 1 minute 7 C 5 minutes A B C D E F G H Depending on the amount of spike-in needed, each of the 50 amplicons may be amplified in replicates of four or five PCR reactions. Replicates from each amplicon were pooled before Qiagen PCR Purification. Pooled amplicons can be eluted with 0 µl of elution buffer. The purified amplicons were checked again on the TapeStation 00, using the D1000 ScreenTape to ensure proper cleanup and size (Figure ). Figure. To ensure unique amplification of the target region, PCR amplicons are evaluated on the Agilent TapeStation 00 using the Agilent D1000 ScreenTape. Each spike-in is given a unique identifier (in this case, alphabets spike-in A-H for eight amplicons). 5
6 The purified amplicons were quantified on a Nanodrop. When pools of 4 5 replicate amplicons were eluted in 0 µl of elution buffer, the concentn was usually > 100 ng/µl; hence, the spike-in stock solution was normalized to 40 ng/µl in 1X TE buffer (ph 8). A spike-in working solution was made from the stock solution by diluting it ~1:,000 with 1X TE buffer (ph 8). At this dilution, the spike-in probe fluorescent signal was significantly above background noise in an acgh experiment. The spike-in working solution dilution can be determined empirically by performing microarray experiments at different dilutions. Aliquots of the spike-in stock (40 ng/µl) can be stored at 0 C indefinitely. Working solution (40 ng/µl diluted at ~1:,000 with 1X TE buffer) can be stored at 4 C for a week without appreciable degradation. For optimum performance, avoid repeated freeze-thaw cycles. During the analysis setup in the CytoGenomics software, the user needs to assign a spike-in identifier to an array (or sample). The spike-in can be assigned from the drop down menu, or as part of the sample attribute file with a column header Spike-In. After data analysis, the CytoGenomics software reports the observed and expected value of each spike-in. Spike-in identity in each sample is reported as match or no match based on the user inputted spike-in identifier in the software (Figure ). Additionally, the software reports the log corresponding to each spike-in. Note that based on this procedure, 50 spike-ins can be generated using one spike-in amplicon per sample. This number can go much higher if combinations of spike-in amplicons are added to each sample. This experiment describes using one amplicon per sample. For each experiment, two microliters of DNA spike-in working solution are added to the test sample gdna before labeling with Cy5 (red signal). A different DNA spike-in is used for each test sample. Spike-ins are not added to the reference DNA (labeled with Cy). Consequently, probes corresponding to spike-ins have a high red signal and low green signal. Using spike-ins in an acgh experiment Spike-in identifiers can be associated with the amplicon probe name in the CytoGenomics software. To identify a sample, one or more amplicons can be added to the sample before a wet lab procedure. An identifier can represent one probe name if one PCR amplicon is spiked into the sample. An identifier can represent more than one probe if multiple PCR amplicons are spiked into the sample. For example, Spike-in A represents a single probe, and spike-in M represents more than one probe (Figure 1). Two microliters of spike-in control is added to DNA sample before processing. Figure. Agilent CytoGenomics software identifies the identity of the spike-in controls. 6
7 Spike-in analysis gprocessedsignal and rprocessedsignal values from each acgh experiment were used for plotting. Four plots are shown in Figure 4, each with a different spike-in added to the test DNA sample. For each sample, the log of rprocessedsignal/gprocessedsignal for all 50 spikes (in trilpicate) are plotted. Each of the 50 spike-ins is represented by a different color. There is very little variation between triplicate values for all spike-ins. As can be seen in Figure 4, the log values differ for individual spike-ins despite the fact that the amplicons are diluted to the same concentn of 40 ng/µl. Log A Index Log 6 B Index C 6 D 5 Log 1 Log Index Index Figure 4. Log is plotted for all 50 amplicons. Greater than 1 log value represents the added DNA spike. 7
8 Conclusion In conclusion, 50 unique amplicons can be used as spike-ins for sample tracking. The spike-ins can be one or more amplicons that are introduced into DNA samples before wet lab processing. The log of spike-ins should be equal to or higher than 1, under the dilution conditions described. The data are highly reproducible, and give the user confidence that there was no mix-up of samples during wet lab or analysis processing. References 1. Douglas, J. A.; Boehnke, M.; Lange, K. A. Multipoint method for detecting genotyping errors and mutations in sibling-pair linkage data. Am. J. Hum. Genet. 000, 66(4), doi: /0861. [PMID: ]. Jun, G.; et al. Detecting and estimating contamination of human DNA samples in sequencing and array-based genotype data. Am. J. Hum. Genet. 01, 91(5), doi: /j. ajhg [PMID: 106]. Burrell, A.; Foy, C.; Burns, M. Applicability of Three Alternative Instruments for Food Authenticity Analysis:GMO Identification. Biotechnology Research International 011, vol. 011, Article ID 88, 8 pages, doi: /011/88 For more information and to learn more Find an Agilent customer center in your country U.S. and Canada agilent_inquiries@agilent.com Asia Pacifi c adinquiry_aplsca@agilent.com Europe info_agilent@agilent.com For Research Use Only. Not for use in diagnostic procedures. This information is subject to change without notice. PR Agilent Technologies, Inc., 016 Published in the USA, March 15, EN
High-Resolution Oligonucleotide- Based acgh Analysis of Single Cells in Under 24 Hours
 High-Resolution Oligonucleotide- Based acgh Analysis of Single Cells in Under 24 Hours Application Note Authors Paula Costa and Anniek De Witte Agilent Technologies, Inc. Santa Clara, CA USA Abstract As
High-Resolution Oligonucleotide- Based acgh Analysis of Single Cells in Under 24 Hours Application Note Authors Paula Costa and Anniek De Witte Agilent Technologies, Inc. Santa Clara, CA USA Abstract As
Single Cell Copy Number Screening Using the GenetiSure Pre-Screen Kit
 Single Cell Copy Number Screening Using the GenetiSure Pre-Screen Kit Uncover Aneuploidies in Embryos Application Note Authors Paula Costa Natalia Novoradovskaya Scott Basehore Stephanie Fulmer-Smentek
Single Cell Copy Number Screening Using the GenetiSure Pre-Screen Kit Uncover Aneuploidies in Embryos Application Note Authors Paula Costa Natalia Novoradovskaya Scott Basehore Stephanie Fulmer-Smentek
SEE WHAT S BEYOND. HaloPlex NGS Target Enrichment SureFISH Probes SurePrint CGH+SNP Microarrays
 SEE WHAT S BEYOND THE ARRAY HaloPlex NGS Target Enrichment SureFISH Probes SurePrint CGH+SNP Microarrays ONE COMPANY, LIMITLESS APPLICATIONS A LEADER IN MOLECULAR ANALYSIS & LIFE SCIENCE RESEARCH Agilent
SEE WHAT S BEYOND THE ARRAY HaloPlex NGS Target Enrichment SureFISH Probes SurePrint CGH+SNP Microarrays ONE COMPANY, LIMITLESS APPLICATIONS A LEADER IN MOLECULAR ANALYSIS & LIFE SCIENCE RESEARCH Agilent
The Agilent Total RNA Isolation Kit
 Better RNA purity. Better data. Without DNase treatment. The Agilent Total RNA Isolation Kit Now you can isolate highly purified, intact RNA without DNase treatment Introducing the Agilent Total RNA Isolation
Better RNA purity. Better data. Without DNase treatment. The Agilent Total RNA Isolation Kit Now you can isolate highly purified, intact RNA without DNase treatment Introducing the Agilent Total RNA Isolation
Agilent RNA Spike-In Kit
 Agilent RNA Spike-In Kit Agilent RNA Spike-In Kit Product Number 5188-5279 Introduction The Agilent RNA Spike-In Kit was developed to provide positive controls for monitoring the microarray workflow from
Agilent RNA Spike-In Kit Agilent RNA Spike-In Kit Product Number 5188-5279 Introduction The Agilent RNA Spike-In Kit was developed to provide positive controls for monitoring the microarray workflow from
Labeling Protocol for mytags Immortal Libraries
 5840 Interface Drive, Suite 101 Ann Arbor MI 48103 1 (734) 998 0751 techsupport@arborbiosci.com Labeling Protocol for mytags Immortal Libraries March 2018 Version 1.5 Contents Reagents and Equipment...
5840 Interface Drive, Suite 101 Ann Arbor MI 48103 1 (734) 998 0751 techsupport@arborbiosci.com Labeling Protocol for mytags Immortal Libraries March 2018 Version 1.5 Contents Reagents and Equipment...
Gene Expression Profiling of Prokaryotic Samples using Low Input Quick Amp WT Kit
 Gene Expression Profiling of Prokaryotic Samples using Low Input Quick Amp WT Kit Application Note Authors Nilanjan Guha and Becky Mullinax Abstract Agilent s Low Input Quick Amp Labeling WT (LIQA WT)
Gene Expression Profiling of Prokaryotic Samples using Low Input Quick Amp WT Kit Application Note Authors Nilanjan Guha and Becky Mullinax Abstract Agilent s Low Input Quick Amp Labeling WT (LIQA WT)
Implementation of Automated Sample Quality Control in Whole Exome Sequencing
 Journal of Life Sciences 11 (2017) 261-268 doi: 10.17265/1934-7391/2017.06.001 D DAVID PUBLISHING Implementation of Automated Sample Quality Control in Whole Exome Sequencing Elisa Viering 1, Jana Molitor
Journal of Life Sciences 11 (2017) 261-268 doi: 10.17265/1934-7391/2017.06.001 D DAVID PUBLISHING Implementation of Automated Sample Quality Control in Whole Exome Sequencing Elisa Viering 1, Jana Molitor
SURESELECTXT LOW INPUT TARGET ENRICHMENT
 SURESELECTXT LOW INPUT TARGET ENRICHMENT Low Input FFPE Optimized Streamlined Workflow SureSelect XT Low Input What is it? SureSelect XT Low Input is a low-input, FFPE-optimized library preparation kit.
SURESELECTXT LOW INPUT TARGET ENRICHMENT Low Input FFPE Optimized Streamlined Workflow SureSelect XT Low Input What is it? SureSelect XT Low Input is a low-input, FFPE-optimized library preparation kit.
Hybridization capture of DNA libraries using xgen Lockdown Probes and Reagents
 Hybridization capture of DNA libraries using xgen Lockdown Probes and Reagents For use with: llumina TruSeq adapter ligated libraries xgen Universal Blockers TS Mix (Catalog # 1075474, 1075475, 1075476)
Hybridization capture of DNA libraries using xgen Lockdown Probes and Reagents For use with: llumina TruSeq adapter ligated libraries xgen Universal Blockers TS Mix (Catalog # 1075474, 1075475, 1075476)
Complete Success Begins with Sample Quality Control. Agilent 4150 and 4200 TapeStation Systems
 Complete Success Begins with Sample Quality Control Agilent 4150 and 4200 TapeStation Systems Complete Success Begins with Sample Quality Control Agilent TapeStation systems are automated electrophoresis
Complete Success Begins with Sample Quality Control Agilent 4150 and 4200 TapeStation Systems Complete Success Begins with Sample Quality Control Agilent TapeStation systems are automated electrophoresis
SureSelect XT HS. Target Enrichment
 SureSelect XT HS Target Enrichment What Is It? SureSelect XT HS joins the SureSelect library preparation reagent family as Agilent s highest sensitivity hybrid capture-based library prep and target enrichment
SureSelect XT HS Target Enrichment What Is It? SureSelect XT HS joins the SureSelect library preparation reagent family as Agilent s highest sensitivity hybrid capture-based library prep and target enrichment
Evaluating the Agilent 4200 TapeStation System for High Throughput Sequencing Quality Control
 Evaluating the Agilent TapeStation System for High Throughput Sequencing Quality Control Application Note Nucleic Acid Analysis Authors Elisa Viering, Laura-Jane Behl, Stefan Pinkert, and Stephan Wolf
Evaluating the Agilent TapeStation System for High Throughput Sequencing Quality Control Application Note Nucleic Acid Analysis Authors Elisa Viering, Laura-Jane Behl, Stefan Pinkert, and Stephan Wolf
Detection of Rare Variants in Degraded FFPE Samples Using the HaloPlex Target Enrichment System
 Detection of Rare Variants in Degraded FFPE Samples Using the HaloPlex Target Enrichment System Application Note Author Linus Forsmark Henrik Johansson Agilent Technologies Inc. Santa Clara, CA USA Abstract
Detection of Rare Variants in Degraded FFPE Samples Using the HaloPlex Target Enrichment System Application Note Author Linus Forsmark Henrik Johansson Agilent Technologies Inc. Santa Clara, CA USA Abstract
Agilent SurePrint G3 Human Catalog CGH Microarrays
 Agilent SurePrint G3 Human Catalog CGH Microarrays Product Note The new Agilent 1M CGH array combines the excellent probe design and performance characteristics of the Agilent acgh platforms with a million
Agilent SurePrint G3 Human Catalog CGH Microarrays Product Note The new Agilent 1M CGH array combines the excellent probe design and performance characteristics of the Agilent acgh platforms with a million
Get to Know Your DNA. Every Single Fragment.
 HaloPlex HS NGS Target Enrichment System Get to Know Your DNA. Every Single Fragment. High sensitivity detection of rare variants using molecular barcodes How Does Molecular Barcoding Work? HaloPlex HS
HaloPlex HS NGS Target Enrichment System Get to Know Your DNA. Every Single Fragment. High sensitivity detection of rare variants using molecular barcodes How Does Molecular Barcoding Work? HaloPlex HS
Cancer Genetics Solutions
 Cancer Genetics Solutions Cancer Genetics Solutions Pushing the Boundaries in Cancer Genetics Cancer is a formidable foe that presents significant challenges. The complexity of this disease can be daunting
Cancer Genetics Solutions Cancer Genetics Solutions Pushing the Boundaries in Cancer Genetics Cancer is a formidable foe that presents significant challenges. The complexity of this disease can be daunting
HyperCap, an automatable workflow on the Agilent Bravo B
 Automation Note February 2018 HyperCap, an automatable workflow on the Agilent Bravo B 1. OVERVIEW As the demand for next-generation sequencing (NGS) grows, laboratories must adapt to manage increased
Automation Note February 2018 HyperCap, an automatable workflow on the Agilent Bravo B 1. OVERVIEW As the demand for next-generation sequencing (NGS) grows, laboratories must adapt to manage increased
Biomek Automated Genomic Sample Prep Accelerates Research
 APPLICATION OVERVIEW Biomek Automated Genomic Sample Prep Accelerates Research Biomek Automation of the Beckman Coulter Agencourt Genfind v2 Blood & Serum DNA Isolation Kit Introduction The Agencourt Genfind
APPLICATION OVERVIEW Biomek Automated Genomic Sample Prep Accelerates Research Biomek Automation of the Beckman Coulter Agencourt Genfind v2 Blood & Serum DNA Isolation Kit Introduction The Agencourt Genfind
Automation of xgen hybridization capture on the Sciclone G3 NGS Workstation
 next generation sequencing protocol Automation of xgen hybridization capture on the Sciclone G3 NGS Workstation www.idtdna.com For Research Use Only Version 2 Revision history Document version Date released
next generation sequencing protocol Automation of xgen hybridization capture on the Sciclone G3 NGS Workstation www.idtdna.com For Research Use Only Version 2 Revision history Document version Date released
hpsc Genetic Analysis Kit
 qpcr analysis kit for detecting the majority of karyotypic abnormalities reported in human ES and ips cells Catalog #07550 60 Reactions Product Description hpsc Genetic Analysis Kit contains nine primer-probe
qpcr analysis kit for detecting the majority of karyotypic abnormalities reported in human ES and ips cells Catalog #07550 60 Reactions Product Description hpsc Genetic Analysis Kit contains nine primer-probe
scgem Workflow Experimental Design Single cell DNA methylation primer design
 scgem Workflow Experimental Design Single cell DNA methylation primer design The scgem DNA methylation assay uses qpcr to measure digestion of target loci by the methylation sensitive restriction endonuclease
scgem Workflow Experimental Design Single cell DNA methylation primer design The scgem DNA methylation assay uses qpcr to measure digestion of target loci by the methylation sensitive restriction endonuclease
High Resolution Microarray Scanner used to Compare CGH Labeling Methods
 High Resolution Microarray Scanner used to Compare CGH Labeling Methods Jack Coleman 1, Wini Luty 1,Marie-Laure Schneider 2 1 Enzo Life Sciences, Farmingdale, NY USA, 2 Innopsys, Toulouse, France CYTAG
High Resolution Microarray Scanner used to Compare CGH Labeling Methods Jack Coleman 1, Wini Luty 1,Marie-Laure Schneider 2 1 Enzo Life Sciences, Farmingdale, NY USA, 2 Innopsys, Toulouse, France CYTAG
Absolute Human Telomere Length Quantification qpcr Assay Kit (AHTLQ) Catalog # reactions
 Absolute Human Telomere Length Quantification qpcr Assay Kit (AHTLQ) Catalog #8918 100 reactions Product Description Telomeres are repetitive nucleotide elements at the ends of chromosomes that protect
Absolute Human Telomere Length Quantification qpcr Assay Kit (AHTLQ) Catalog #8918 100 reactions Product Description Telomeres are repetitive nucleotide elements at the ends of chromosomes that protect
HELINI Hepatitis B virus [HBV] Real-time PCR Kit (Genotype A to H)
![HELINI Hepatitis B virus [HBV] Real-time PCR Kit (Genotype A to H) HELINI Hepatitis B virus [HBV] Real-time PCR Kit (Genotype A to H)](/thumbs/87/95094490.jpg) HELINI Hepatitis B virus [HBV] Real-time PCR Kit (Genotype A to H) Quantitative In vitro diagnostics Instruction manual Cat. No: 8001-25/50/100 tests Compatible with: Agilent, Bio-Rad, Applied Bio systems
HELINI Hepatitis B virus [HBV] Real-time PCR Kit (Genotype A to H) Quantitative In vitro diagnostics Instruction manual Cat. No: 8001-25/50/100 tests Compatible with: Agilent, Bio-Rad, Applied Bio systems
Magnis NGS Prep System. Your lab s automated library preparation solution for next-generation sequencing
 Magnis NGS Prep System Your lab s automated library preparation solution for next-generation sequencing Magnis NGS Prep System Complete system for NGS preparation One of the challenges with implementing
Magnis NGS Prep System Your lab s automated library preparation solution for next-generation sequencing Magnis NGS Prep System Complete system for NGS preparation One of the challenges with implementing
Automation of the IDT xgen Lockdown Panels on the Sciclone G3 NGS Workstation
 A PPLI C A TION NOT E NGS Library Preparation Sciclone G3 NGSx Workstation Automation of the IDT xgen Lockdown Panels on the Sciclone G3 NGS Workstation Introduction Integrated DNA Technologies (IDT) xgen
A PPLI C A TION NOT E NGS Library Preparation Sciclone G3 NGSx Workstation Automation of the IDT xgen Lockdown Panels on the Sciclone G3 NGS Workstation Introduction Integrated DNA Technologies (IDT) xgen
Wu et al., Determination of genetic identity in therapeutic chimeric states. We used two approaches for identifying potentially suitable deletion loci
 SUPPLEMENTARY METHODS AND DATA General strategy for identifying deletion loci We used two approaches for identifying potentially suitable deletion loci for PDP-FISH analysis. In the first approach, we
SUPPLEMENTARY METHODS AND DATA General strategy for identifying deletion loci We used two approaches for identifying potentially suitable deletion loci for PDP-FISH analysis. In the first approach, we
Protocol for submitting gdna for Illumina 16S rrna gene sequencing, V4 region
 Protocol for submitting gdna for Illumina 16S rrna gene sequencing, V4 region Version: 2018 Summary: This protocol is for the submission of gdna to generate libraries for 16S rrna sequencing, which can
Protocol for submitting gdna for Illumina 16S rrna gene sequencing, V4 region Version: 2018 Summary: This protocol is for the submission of gdna to generate libraries for 16S rrna sequencing, which can
A Recommended Procedure for Real-Time Quantitative TaqMan PCR for Roundup Ready Canola RT73 Monsanto Biotechnology Regulatory Sciences
 Page 1 of 7 Overview Purpose & Scope This procedure describes an event-specific real-time TaqMan PCR method for determination of the relative content of Roundup Ready canola RT73 (hereafter referred to
Page 1 of 7 Overview Purpose & Scope This procedure describes an event-specific real-time TaqMan PCR method for determination of the relative content of Roundup Ready canola RT73 (hereafter referred to
A Systematic Approach to Optimize Real-Time Quantitative RT-qPCR Experiments with the Agilent 2200 TapeStation System
 Systematic pproach to Optimize Real-Time Quantitative RT-qPCR Experiments with the gilent TapeStation System pplication Note Nucleic cid nalysis uthor runkumar Padmanaban gilent Technologies, Inc. angalore,
Systematic pproach to Optimize Real-Time Quantitative RT-qPCR Experiments with the gilent TapeStation System pplication Note Nucleic cid nalysis uthor runkumar Padmanaban gilent Technologies, Inc. angalore,
HALOPLEX PCR TARGET ENRICHMENT & LIBRARY PREPARATION PROTOCOL. Version 1.0.3, April 2011 For research use only
 HALOPLEX PCR TARGET ENRICHMENT & LIBRARY PREPARATION PROTOCOL Version 1.0.3, April 2011 For research use only HALOPLEX PCR TARGET ENRICHMENT & LIBRARY PREPARATION PROTOCOL Halo Genomics AB, 2011 No part
HALOPLEX PCR TARGET ENRICHMENT & LIBRARY PREPARATION PROTOCOL Version 1.0.3, April 2011 For research use only HALOPLEX PCR TARGET ENRICHMENT & LIBRARY PREPARATION PROTOCOL Halo Genomics AB, 2011 No part
TECHNICAL BULLETIN. SeqPlex DNA Amplification Kit for use with high throughput sequencing technologies. Catalog Number SEQX Storage Temperature 20 C
 SeqPlex DNA Amplification Kit for use with high throughput sequencing technologies Catalog Number SEQX Storage Temperature 20 C TECHNICAL BULLETIN Product Description The SeqPlex DNA Amplification Kit
SeqPlex DNA Amplification Kit for use with high throughput sequencing technologies Catalog Number SEQX Storage Temperature 20 C TECHNICAL BULLETIN Product Description The SeqPlex DNA Amplification Kit
Absolute Mouse Telomere Length Quantification qpcr Assay Kit (AMTLQ) Catalog #M reactions
 Absolute Mouse Telomere Length Quantification qpcr Assay Kit (AMTLQ) Catalog #M8918 100 reactions Product Description Telomeres are repetitive nucleotide elements at the ends of chromosomes that protect
Absolute Mouse Telomere Length Quantification qpcr Assay Kit (AMTLQ) Catalog #M8918 100 reactions Product Description Telomeres are repetitive nucleotide elements at the ends of chromosomes that protect
Executive Summary. clinical supply services
 clinical supply services case study Development and NDA-level validation of quantitative polymerase chain reaction (qpcr) procedure for detection and quantification of residual E.coli genomic DNA Executive
clinical supply services case study Development and NDA-level validation of quantitative polymerase chain reaction (qpcr) procedure for detection and quantification of residual E.coli genomic DNA Executive
Performance of the Agilent D1000 and the Agilent High Sensitivity D1000 ScreenTape Assay for the Agilent 4200 TapeStation System
 Performance of the gilent D and the gilent High Sensitivity D ScreenTape ssay for the gilent 42 TapeStation System Technical Overview Introduction The gilent 42 TapeStation system is an automated system
Performance of the gilent D and the gilent High Sensitivity D ScreenTape ssay for the gilent 42 TapeStation System Technical Overview Introduction The gilent 42 TapeStation system is an automated system
Comparison of ExoSAP-IT and ExoSAP-IT Express reagents to alternative PCR cleanup methods
 WHITE PAPER ExoSAP-IT PCR cleanup reagents Comparison of ExoSAP-IT and ExoSAP-IT Express reagents to alternative PCR cleanup methods Abstract Here we present superior workflow advantages of enzymatic PCR
WHITE PAPER ExoSAP-IT PCR cleanup reagents Comparison of ExoSAP-IT and ExoSAP-IT Express reagents to alternative PCR cleanup methods Abstract Here we present superior workflow advantages of enzymatic PCR
GreenMasterMix (2X) b i o s c i e n c e. G E N A X X O N b i o s c i e n c e. High ROX (500nM)
 G:\products\productflyer\pcr\polymerasen\hotstart\manu_m3052_green_en.docx GreenMasterMix (2) High RO (500nM) qpcr master mix with fluorescence dye and passive reference dye Contact & Technical support
G:\products\productflyer\pcr\polymerasen\hotstart\manu_m3052_green_en.docx GreenMasterMix (2) High RO (500nM) qpcr master mix with fluorescence dye and passive reference dye Contact & Technical support
QUANTITATIVE RT-PCR PROTOCOL (SYBR Green I) (Last Revised: April, 2007)
 QUANTITATIVE RT-PCR PROTOCOL (SYBR Green I) (Last Revised: April, 007) Please contact Center for Plant Genomics (CPG) facility manager Hailing Jin (hljin@iastate.edu) regarding questions or corrections.
QUANTITATIVE RT-PCR PROTOCOL (SYBR Green I) (Last Revised: April, 007) Please contact Center for Plant Genomics (CPG) facility manager Hailing Jin (hljin@iastate.edu) regarding questions or corrections.
Analysis of DNA sequence copy number in archival formalin-fixed, paraffinembedded. Dione K. Bailey Anniek De Witte October 9, 2007
 Analysis of DNA sequence copy number in archival formalin-fixed, paraffinembedded (FFPE) tumor samples by oligo array CGH Dione K. Bailey Anniek De Witte October 9, 2007 Agenda Oligo acgh Review Problems
Analysis of DNA sequence copy number in archival formalin-fixed, paraffinembedded (FFPE) tumor samples by oligo array CGH Dione K. Bailey Anniek De Witte October 9, 2007 Agenda Oligo acgh Review Problems
AmpF STR NGM PCR Amplification Kit - Overview
 AmpF STR NGM PCR Amplification Kit - Overview The most advanced STR kits optimized for analysis of forensic casework and database samples in Europe Data Quality Worth Sharing! 9/7/2011 Life Technologies
AmpF STR NGM PCR Amplification Kit - Overview The most advanced STR kits optimized for analysis of forensic casework and database samples in Europe Data Quality Worth Sharing! 9/7/2011 Life Technologies
Event COT102 Cotton. Real-time, Event-specific Polymerase Chain Reaction Method
 Event COT102 Cotton Real-time, Event-specific Polymerase Chain Reaction Method Syngenta Crop Protection, LLC 3054 East Cornwallis Road PO Box 12257 Research Triangle Park, NC 27709-2257, USA For technical
Event COT102 Cotton Real-time, Event-specific Polymerase Chain Reaction Method Syngenta Crop Protection, LLC 3054 East Cornwallis Road PO Box 12257 Research Triangle Park, NC 27709-2257, USA For technical
Roche Molecular Biochemicals Technical Note No. LC 12/2000
 Roche Molecular Biochemicals Technical Note No. LC 12/2000 LightCycler Absolute Quantification with External Standards and an Internal Control 1. General Introduction Purpose of this Note Overview of Method
Roche Molecular Biochemicals Technical Note No. LC 12/2000 LightCycler Absolute Quantification with External Standards and an Internal Control 1. General Introduction Purpose of this Note Overview of Method
CytoChip Oligo Summary Protocol
 FOR RESEARCH USE ONLY ILLUMINA PROPRIETARY October 2014 Page 1 of 15 Sample Preparation Sample Preparation Materials Table 1 Starting materials for sample preparation Starting materials Amount DNA (unsheared,
FOR RESEARCH USE ONLY ILLUMINA PROPRIETARY October 2014 Page 1 of 15 Sample Preparation Sample Preparation Materials Table 1 Starting materials for sample preparation Starting materials Amount DNA (unsheared,
Copy Number Analysis of Archival FFPE Tumor Samples by Oligo Array CGH
 GENOMICS INFORMATICS PROTEOMICS METABOLOMICS A T C T G A T C C T T C T G AAC GGAAC T AAT T T C AA G AAT C T G AT C C T T G A A C T A C C T T C C AAG G T G Copy Number Analysis of Archival FFPE Tumor Samples
GENOMICS INFORMATICS PROTEOMICS METABOLOMICS A T C T G A T C C T T C T G AAC GGAAC T AAT T T C AA G AAT C T G AT C C T T G A A C T A C C T T C C AAG G T G Copy Number Analysis of Archival FFPE Tumor Samples
Detecting copy-neutral LOH in cancer using Agilent SurePrint G3 Cancer CGH+SNP Microarrays
 Detecting copy-neutral LOH in cancer using Agilent SurePrint G3 Cancer CGH+SNP Microarrays Application Note Authors Paula Costa Anniek De Witte Jayati Ghosh Agilent Technologies, Inc. Santa Clara, CA USA
Detecting copy-neutral LOH in cancer using Agilent SurePrint G3 Cancer CGH+SNP Microarrays Application Note Authors Paula Costa Anniek De Witte Jayati Ghosh Agilent Technologies, Inc. Santa Clara, CA USA
Complete protocol in 110 minutes Enzymatic fragmentation without sonication One-step fragmentation/tagging to save time
 Molecular Cloning Laboratories Manual Version 1.2 Product name: MCNext UT DNA Sample Prep Kit Cat #: MCUDS-4, MCUDS-24, MCUDS-96 Description: This protocol explains how to prepare up to 96 pooled indexed
Molecular Cloning Laboratories Manual Version 1.2 Product name: MCNext UT DNA Sample Prep Kit Cat #: MCUDS-4, MCUDS-24, MCUDS-96 Description: This protocol explains how to prepare up to 96 pooled indexed
Reference Gene Panel. Human - Short Assays Version 1 - April 2016 Probe protocol. User manual. Order #:
 User manual Reference Gene Panel Human - Short Assays Version 1 - April 2016 Probe protocol Order #: Human Reference Gene Panel Probe FAM - Short Assays : A104P Reference Gene Panel - Human Short Assays
User manual Reference Gene Panel Human - Short Assays Version 1 - April 2016 Probe protocol Order #: Human Reference Gene Panel Probe FAM - Short Assays : A104P Reference Gene Panel - Human Short Assays
NUCLEIC ACID PURIFICATION KITS FAST SIMPLE QUALITY CONVENIENT REPRODUCIBLE
 NUCLEIC ACID PURIFICATION KITS FAST SIMPLE QUALITY CONVENIENT REPRODUCIBLE COMPANY PROFILE Since its founding in 1998,, Inc. has been at the forefront of nucleic acid purification by offering products
NUCLEIC ACID PURIFICATION KITS FAST SIMPLE QUALITY CONVENIENT REPRODUCIBLE COMPANY PROFILE Since its founding in 1998,, Inc. has been at the forefront of nucleic acid purification by offering products
Impact of gdna Integrity on the Outcome of DNA Methylation Studies
 Impact of gdna Integrity on the Outcome of DNA Methylation Studies Application Note Nucleic Acid Analysis Authors Emily Putnam, Keith Booher, and Xueguang Sun Zymo Research Corporation, Irvine, CA, USA
Impact of gdna Integrity on the Outcome of DNA Methylation Studies Application Note Nucleic Acid Analysis Authors Emily Putnam, Keith Booher, and Xueguang Sun Zymo Research Corporation, Irvine, CA, USA
High Performance Labeling Kits for cdna and Oligo Microarrays
 Agilent in Life Sciences > Genomics > Proteomics > Drug Discovery > Development > QA/QC High Performance Labeling Kits for cdna and Oligo Microarrays Get more from one experiment Agilent has perfected
Agilent in Life Sciences > Genomics > Proteomics > Drug Discovery > Development > QA/QC High Performance Labeling Kits for cdna and Oligo Microarrays Get more from one experiment Agilent has perfected
Functional Genomics Research Stream. Research Meeting: June 19, 2012 SYBR Green qpcr, Research Update
 Functional Genomics Research Stream Research Meeting: June 19, 2012 SYBR Green qpcr, Research Update Updates Alternate Lab Meeting Fridays 11:30-1:00 WEL 4.224 Welcome to attend either one Lab Log thanks
Functional Genomics Research Stream Research Meeting: June 19, 2012 SYBR Green qpcr, Research Update Updates Alternate Lab Meeting Fridays 11:30-1:00 WEL 4.224 Welcome to attend either one Lab Log thanks
Fully Automated Library Quantification for Illumina Sequencing on the NGS STAR
 Fully Automated Library Quantification for Illumina Sequencing on the NGS STAR Introduction Hamilton Robotics, an industry leader in liquid handling and laboratory automation equipment, has partnered with
Fully Automated Library Quantification for Illumina Sequencing on the NGS STAR Introduction Hamilton Robotics, an industry leader in liquid handling and laboratory automation equipment, has partnered with
Agilent SureCycler 8800
 Agilent SureCycler 8800 CHOOSING THE RIGHT THERMAL CYCLER HAS NEVER BEEN EASIER SURELY BETTER PCR FUNCTIONALITY FOR YOUR LAB Remote access experiment from across the room or across town hardware purchase
Agilent SureCycler 8800 CHOOSING THE RIGHT THERMAL CYCLER HAS NEVER BEEN EASIER SURELY BETTER PCR FUNCTIONALITY FOR YOUR LAB Remote access experiment from across the room or across town hardware purchase
Archaea V3-5 16S rrna Amplicon Sequencing
 Archaea V3-5 16S rrna Amplicon Sequencing Standard Protocol Version 1.0 Skill Prerequisites: DNA handling, gel electrophoresis, DNA concentration measurement, polymerase chain reaction (PCR) Introduction
Archaea V3-5 16S rrna Amplicon Sequencing Standard Protocol Version 1.0 Skill Prerequisites: DNA handling, gel electrophoresis, DNA concentration measurement, polymerase chain reaction (PCR) Introduction
Relative Mouse Telomere Length Quantification qpcr Assay Kit (RMTLQ) Catalog #M reactions
 Relative Mouse Telomere Length Quantification qpcr Assay Kit (RMTLQ) Catalog #M8908 100 reactions Product Description Telomeres are repetitive nucleotide elements at the ends of chromosomes that protect
Relative Mouse Telomere Length Quantification qpcr Assay Kit (RMTLQ) Catalog #M8908 100 reactions Product Description Telomeres are repetitive nucleotide elements at the ends of chromosomes that protect
Updated August well High-multiplexing Tagmentation and Amplification Robyn Tanny August Company Kit Catalog Number.
 96-well High-multiplexing Tagmentation and Amplification Robyn Tanny August 2015 This protocol uses the following purchased reagents: Company Kit Catalog Number Illumina Nextera DNA Sample Preparation
96-well High-multiplexing Tagmentation and Amplification Robyn Tanny August 2015 This protocol uses the following purchased reagents: Company Kit Catalog Number Illumina Nextera DNA Sample Preparation
Cold Fusion Cloning Kit. Cat. #s MC100A-1, MC101A-1. User Manual
 Fusion Cloning technology Cold Fusion Cloning Kit Store the master mixture and positive controls at -20 C Store the competent cells at -80 C. (ver. 120909) A limited-use label license covers this product.
Fusion Cloning technology Cold Fusion Cloning Kit Store the master mixture and positive controls at -20 C Store the competent cells at -80 C. (ver. 120909) A limited-use label license covers this product.
High specificity and sensitivity for fungal DNA allows reliable quantification in a background of nonfungal. Contents
 INSTRUCTION MANUAL Femto Fungal DNA Quantification Kit Catalog No. E2007 Highlights Accurately and reproducibly quantify as little as 20 fg of fungal DNA from 1 µl of sample using realtime PCR. High specificity
INSTRUCTION MANUAL Femto Fungal DNA Quantification Kit Catalog No. E2007 Highlights Accurately and reproducibly quantify as little as 20 fg of fungal DNA from 1 µl of sample using realtime PCR. High specificity
HELINI White spot Syndrome virus [WSSV] Real-time PCR Kit
![HELINI White spot Syndrome virus [WSSV] Real-time PCR Kit HELINI White spot Syndrome virus [WSSV] Real-time PCR Kit](/thumbs/72/67570787.jpg) HELINI White spot Syndrome virus [WSSV] Real-time PCR Kit Instruction manual Cat. No: 6001-25/50/100 tests Compatible with: Agilent, Bio-Rad, Roche, Applied Bio systems [ABI], Rotor-gene, Cepheid, Bioer,
HELINI White spot Syndrome virus [WSSV] Real-time PCR Kit Instruction manual Cat. No: 6001-25/50/100 tests Compatible with: Agilent, Bio-Rad, Roche, Applied Bio systems [ABI], Rotor-gene, Cepheid, Bioer,
Forensic News. FAS Corner April The Maximizing Data Quality Series Part 2: DNA Quantitation
 The Maximizing Data Quality Series Part 2: DNA Quantitation (Editor's Note: This is part of an ongoing series of articles focusing on maximizing data quality throughout the HID workflow. Part 1 on DNA
The Maximizing Data Quality Series Part 2: DNA Quantitation (Editor's Note: This is part of an ongoing series of articles focusing on maximizing data quality throughout the HID workflow. Part 1 on DNA
QIAGEN Supplementary Protocol
 Triplex to 5-plex real-time PCR analysis using the QuantiFast Pathogen PCR +IC Kit on the Rotor-Gene Q This protocol describes how to use the QuantiFast Pathogen PCR +IC Kit to perform real-time PCR analysis
Triplex to 5-plex real-time PCR analysis using the QuantiFast Pathogen PCR +IC Kit on the Rotor-Gene Q This protocol describes how to use the QuantiFast Pathogen PCR +IC Kit to perform real-time PCR analysis
Cat. # RR391A. For Research Use. Probe qpcr Mix. Product Manual. v201610da
 Cat. # RR391A For Research Use Probe qpcr Mix Product Manual Table of Contents I. Introduction... 3 II. Principle... 3 III. Components... 5 IV. Materials Required but not Provided... 5 V. Storage... 5
Cat. # RR391A For Research Use Probe qpcr Mix Product Manual Table of Contents I. Introduction... 3 II. Principle... 3 III. Components... 5 IV. Materials Required but not Provided... 5 V. Storage... 5
Precipio, Inc. Instructions for Use. PIK3CA Exon 9 Mutation Analysis using ICE COLD-PCR for Detection with High Resolution Melting
 Precipio, Inc. Instructions for Use PIK3CA Exon 9 Mutation Analysis using ICE COLD-PCR for Detection with High Resolution Melting Table of Contents Manufacturer 2 Reagent Preparation 2 Kit Components and
Precipio, Inc. Instructions for Use PIK3CA Exon 9 Mutation Analysis using ICE COLD-PCR for Detection with High Resolution Melting Table of Contents Manufacturer 2 Reagent Preparation 2 Kit Components and
Efficient Method for Isolation of High Quality Concentrated Cellular RNA with Extremely Low Levels of Genomic DNA Contamination Application
 Efficient Method for Isolation of High Quality Concentrated Cellular RNA with Extremely Low Levels of Genomic DNA Contamination Application Gene Expression Authors Ilgar Abbaszade, Claudia Robbins, John
Efficient Method for Isolation of High Quality Concentrated Cellular RNA with Extremely Low Levels of Genomic DNA Contamination Application Gene Expression Authors Ilgar Abbaszade, Claudia Robbins, John
PAXgene Blood DNA PAXgene Blood DNA Tube (IVD) for clinical use PAXgene Blood DNA System (RUO) for research use. Explore more at
 PAXgene Blood DNA PAXgene Blood DNA Tube (IVD) for clinical use PAXgene Blood DNA System (RUO) for research use Explore more at www.preanalytix.com Situation The composition, amount, quality and integrity
PAXgene Blood DNA PAXgene Blood DNA Tube (IVD) for clinical use PAXgene Blood DNA System (RUO) for research use Explore more at www.preanalytix.com Situation The composition, amount, quality and integrity
HaloPlex HS. Get to Know Your DNA. Every Single Fragment. Kevin Poon, Ph.D.
 HaloPlex HS Get to Know Your DNA. Every Single Fragment. Kevin Poon, Ph.D. Sr. Global Product Manager Diagnostics & Genomics Group Agilent Technologies For Research Use Only. Not for Use in Diagnostic
HaloPlex HS Get to Know Your DNA. Every Single Fragment. Kevin Poon, Ph.D. Sr. Global Product Manager Diagnostics & Genomics Group Agilent Technologies For Research Use Only. Not for Use in Diagnostic
Amplicon Sequencing Template Preparation
 Amplicon Sequencing Template Preparation The DNA sample preparation procedure for Amplicon Sequencing consists of a simple PCR amplification reaction, but uses special Fusion Primers (Figure 1-1). The
Amplicon Sequencing Template Preparation The DNA sample preparation procedure for Amplicon Sequencing consists of a simple PCR amplification reaction, but uses special Fusion Primers (Figure 1-1). The
PrimePCR Assay Validation Report
 Gene Information Gene Name keratin 78 Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID KRT78 Human This gene is a member of the type II keratin gene family
Gene Information Gene Name keratin 78 Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID KRT78 Human This gene is a member of the type II keratin gene family
Procedure & Checklist - Preparing Asymmetric SMRTbell Templates
 Procedure & Checklist - Preparing Asymmetric SMRTbell Templates Before You Begin In this procedure, PCR products are generated using two rounds of amplification. The first round uses target specific primers
Procedure & Checklist - Preparing Asymmetric SMRTbell Templates Before You Begin In this procedure, PCR products are generated using two rounds of amplification. The first round uses target specific primers
ThruPLEX -FD Prep Kit Instruction Manual. Single Tube Library Preparation for Illumina NGS Platforms
 ThruPLEX -FD Prep Kit Instruction Manual Single Tube Library Preparation for Illumina NGS Platforms Contents Product Description... 2 Kit Contents... 2 Shipping and Storage... 2 Getting Started... 3 Input
ThruPLEX -FD Prep Kit Instruction Manual Single Tube Library Preparation for Illumina NGS Platforms Contents Product Description... 2 Kit Contents... 2 Shipping and Storage... 2 Getting Started... 3 Input
PrimePCR Assay Validation Report
 Gene Information Gene Name collagen, type IV, alpha 1 Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID COL4A1 Human This gene encodes the major type IV alpha
Gene Information Gene Name collagen, type IV, alpha 1 Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID COL4A1 Human This gene encodes the major type IV alpha
Absolute Human Telomere Length and Mitochondrial DNA Copy Number Dual Quantification qpcr Assay Kit (AHDQ) Catalog # reactions
 Absolute Human Telomere Length and Mitochondrial DNA Copy Number Dual Quantification qpcr Assay Kit (AHDQ) Catalog #8958 100 reactions Product Description Telomeres are repetitive nucleotide elements at
Absolute Human Telomere Length and Mitochondrial DNA Copy Number Dual Quantification qpcr Assay Kit (AHDQ) Catalog #8958 100 reactions Product Description Telomeres are repetitive nucleotide elements at
Relative Rat Telomere Length Quantification qpcr Assay Kit (RRTLQ) Catalog #R reactions
 Relative Rat Telomere Length Quantification qpcr Assay Kit (RRTLQ) Catalog #R8908 100 reactions Product Description Telomeres are repetitive nucleotide elements at the ends of chromosomes that protect
Relative Rat Telomere Length Quantification qpcr Assay Kit (RRTLQ) Catalog #R8908 100 reactions Product Description Telomeres are repetitive nucleotide elements at the ends of chromosomes that protect
Guidelines for Developing Robust and Reliable PCR Assays
 Guidelines for Developing Robust and Reliable PCR Assays Leta Steffen, PhD Applications Scientist Promega Corporation Outline 1) PCR reaction components What is in the reaction? How does it affect assay
Guidelines for Developing Robust and Reliable PCR Assays Leta Steffen, PhD Applications Scientist Promega Corporation Outline 1) PCR reaction components What is in the reaction? How does it affect assay
Application Protocol: DNA quantification by using Absolute Quantitative PCR (qpcr)
 REPRODUCTION AND USE This document is protected by copyright and cannot be used or shared without permission from Vortex Biosciences, Inc. Such permission is given on condition that Vortex Biosciences
REPRODUCTION AND USE This document is protected by copyright and cannot be used or shared without permission from Vortex Biosciences, Inc. Such permission is given on condition that Vortex Biosciences
10 RXN 50 RXN 500 RXN
 SeqPlex Enhanced DNA Amplification Kit for use with high throughput sequencing technologies Catalog Number SEQXE Storage Temperature 20 C TECHNICAL BULLETIN Product Description The SeqPlex DNA Amplification
SeqPlex Enhanced DNA Amplification Kit for use with high throughput sequencing technologies Catalog Number SEQXE Storage Temperature 20 C TECHNICAL BULLETIN Product Description The SeqPlex DNA Amplification
PrimePCR Assay Validation Report
 Gene Information Gene Name keratin 3 Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID KRT3 Human The protein encoded by this gene is a member of the keratin
Gene Information Gene Name keratin 3 Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID KRT3 Human The protein encoded by this gene is a member of the keratin
Internal controls for real-time PCR-based pathogen identification
 Internal controls for real-time PCR-based pathogen identification QIAGEN Dr. Lillian Roth Lead Scientist Veterinary R&D Team Global R&D Competence Center Applied Testing QIAGEN webinar June 1, 2011 Introduction
Internal controls for real-time PCR-based pathogen identification QIAGEN Dr. Lillian Roth Lead Scientist Veterinary R&D Team Global R&D Competence Center Applied Testing QIAGEN webinar June 1, 2011 Introduction
Amplicon size (bp) E ± SD
 Development of a probe-based qpcr assay for the detection of constitutional and somatic deletions in the NF1 gene. Application to genetic testing and tumor analysis. Terribas E, Garcia-Linares C, Lázaro
Development of a probe-based qpcr assay for the detection of constitutional and somatic deletions in the NF1 gene. Application to genetic testing and tumor analysis. Terribas E, Garcia-Linares C, Lázaro
Amplicon Library Preparation Method Manual. GS FLX Titanium Series October 2009
 GS FLX Titanium Series 1. Workflow 3. Procedure The procedure to prepare Amplicon libraries is shown in Figure 1. It consists of a PCR amplification, performed using special Fusion Primers for the Genome
GS FLX Titanium Series 1. Workflow 3. Procedure The procedure to prepare Amplicon libraries is shown in Figure 1. It consists of a PCR amplification, performed using special Fusion Primers for the Genome
Surely Better Target Enrichment from Sample to Sequencer
 sureselect TARGET ENRICHMENT solutions Surely Better Target Enrichment from Sample to Sequencer Agilent s market leading SureSelect platform provides a complete portfolio of catalog to custom products,
sureselect TARGET ENRICHMENT solutions Surely Better Target Enrichment from Sample to Sequencer Agilent s market leading SureSelect platform provides a complete portfolio of catalog to custom products,
2014 Hands-On Introductory Workshop on Array CGH
 2014 Hands-On Introductory Workshop on Array CGH UTMDACC, Program in Cytogenetic Technology Agilent Technologies 1 Welcome! Workshop Agenda LABORATORY TRAINING Y3.5721 9:00-9:15 Introduction and Protocol
2014 Hands-On Introductory Workshop on Array CGH UTMDACC, Program in Cytogenetic Technology Agilent Technologies 1 Welcome! Workshop Agenda LABORATORY TRAINING Y3.5721 9:00-9:15 Introduction and Protocol
Getting the Most from Your DNA Analysis from Purification to Downstream Analysis. Eric B. Vincent, Ph.D. February 2013
 Getting the Most from Your DNA Analysis from Purification to Downstream Analysis Eric B. Vincent, Ph.D. February 2013 1 Presentation Outline Genomic Analysis Purification Quantitation Qualification Analysis
Getting the Most from Your DNA Analysis from Purification to Downstream Analysis Eric B. Vincent, Ph.D. February 2013 1 Presentation Outline Genomic Analysis Purification Quantitation Qualification Analysis
General Product Insert. End-Point PCR/RT-PCR Kit Product# EPxxxxx
 3430 Schmon Parkway Thorold, ON, Canada L2V 4Y6 Phone: 866-667-4362 (905) 227-8848 Fax: (905) 227-1061 Email: techsupport@norgenbiotek.com End-Point PCR/RT-PCR Kit Product# EPxxxxx General Product Insert
3430 Schmon Parkway Thorold, ON, Canada L2V 4Y6 Phone: 866-667-4362 (905) 227-8848 Fax: (905) 227-1061 Email: techsupport@norgenbiotek.com End-Point PCR/RT-PCR Kit Product# EPxxxxx General Product Insert
Purpose: To sequence 16S microbial DNA isolated from animal fecal matter and perform compositional analysis of bacterial communities.
 Purpose: To sequence 16S microbial DNA isolated from animal fecal matter and perform compositional analysis of bacterial communities. Reference: This protocol was adapted from: A custom and streamlined
Purpose: To sequence 16S microbial DNA isolated from animal fecal matter and perform compositional analysis of bacterial communities. Reference: This protocol was adapted from: A custom and streamlined
User Bulletin. Veriti 96-Well Thermal Cycler AmpFlSTR Kit Validation. Overview
 User Bulletin Veriti 96-Well Thermal Cycler AmpFlSTR Kit Validation June 2009 SUBJECT: AmpFlSTR PCR Amplification Kit validation on the Veriti 96-Well Thermal Cycler with 0.2 ml sample block format In
User Bulletin Veriti 96-Well Thermal Cycler AmpFlSTR Kit Validation June 2009 SUBJECT: AmpFlSTR PCR Amplification Kit validation on the Veriti 96-Well Thermal Cycler with 0.2 ml sample block format In
Premix Ex Taq (Probe qpcr)
 For Research Use Premix Ex Taq (Probe qpcr) Product Manual Table of Contents I. Description... 3 II. Principle... 4 III. Components... 5 IV. Materials Required but not Provided... 5 V. Storage... 5 VI.
For Research Use Premix Ex Taq (Probe qpcr) Product Manual Table of Contents I. Description... 3 II. Principle... 4 III. Components... 5 IV. Materials Required but not Provided... 5 V. Storage... 5 VI.
PrimePCR Assay Validation Report
 Gene Information Gene Name Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID mcf.2 transforming sequence-like Mcf2l Mouse Description Not Available C130040G20Rik,
Gene Information Gene Name Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID mcf.2 transforming sequence-like Mcf2l Mouse Description Not Available C130040G20Rik,
A comparative performance evaluation of illustra Ready-To-Go GenomiPhi V3 and illustra GenomiPhi V2 DNA amplification kits
 GE Healthcare Life Sciences Application note 29-0245-46 AA Nucleic acid sample preparation A comparative performance evaluation of illustra Ready-To-Go GenomiPhi V3 and illustra GenomiPhi V2 DNA amplification
GE Healthcare Life Sciences Application note 29-0245-46 AA Nucleic acid sample preparation A comparative performance evaluation of illustra Ready-To-Go GenomiPhi V3 and illustra GenomiPhi V2 DNA amplification
Accurately and reproducibly quantify as little as 20 fg of bacterial DNA using real-time PCR.
 INSTRUCTION MANUAL Femto Bacterial DNA Quantification Kit Catalog No. E2006 Highlights Accurately and reproducibly quantify as little as 20 fg of bacterial DNA using real-time PCR. High specificity and
INSTRUCTION MANUAL Femto Bacterial DNA Quantification Kit Catalog No. E2006 Highlights Accurately and reproducibly quantify as little as 20 fg of bacterial DNA using real-time PCR. High specificity and
Complementary Technologies for Precision Genetic Analysis
 Complementary NGS, CGH and Workflow Featured Publication Zhu, J. et al. Duplication of C7orf58, WNT16 and FAM3C in an obese female with a t(7;22)(q32.1;q11.2) chromosomal translocation and clinical features
Complementary NGS, CGH and Workflow Featured Publication Zhu, J. et al. Duplication of C7orf58, WNT16 and FAM3C in an obese female with a t(7;22)(q32.1;q11.2) chromosomal translocation and clinical features
QPCR Designed for You. Superior Instruments, Reagents, and Support.
 QPCR Designed for You Superior Instruments, Reagents, and Support. Agilent s portfolio of qpcr products delivers speed, reliability, and above all confidence in your results. Technology Designed for You
QPCR Designed for You Superior Instruments, Reagents, and Support. Agilent s portfolio of qpcr products delivers speed, reliability, and above all confidence in your results. Technology Designed for You
Sanger Sequencing Portfolio
 Molecular Cloning Laboratories Sanger Sequencing Portfolio Significant Cost Savings vs ABI Uncompromised Performance Longer Read Length Longer Shelf Life No Recalibrations Necessary PAGE 1 BrightDye Terminator
Molecular Cloning Laboratories Sanger Sequencing Portfolio Significant Cost Savings vs ABI Uncompromised Performance Longer Read Length Longer Shelf Life No Recalibrations Necessary PAGE 1 BrightDye Terminator
Best Quantification Practices with the Agilent 5200 Fragment Analyzer System
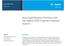 Application Note Genomics Best Quantification Practices with the Agilent 5 Fragment Analyzer System Authors Chava Pocernich, Jolita Uthe, Whitney Pike, and Kit Sum Wong Agilent Technologies, Inc. Abstract
Application Note Genomics Best Quantification Practices with the Agilent 5 Fragment Analyzer System Authors Chava Pocernich, Jolita Uthe, Whitney Pike, and Kit Sum Wong Agilent Technologies, Inc. Abstract
THUNDERBIRD SYBR qpcr Mix
 Instruction manual THUNDERBIRD SYBR qpcr Mix 1304 A4251K THUNDERBIRD SYBR qpcr Mix QPS-201T 1 ml x 1 QPS-201 1.67 ml x 3 Contents [1] Introduction [2] Components [3] Primer design [4] Template DNA [5]
Instruction manual THUNDERBIRD SYBR qpcr Mix 1304 A4251K THUNDERBIRD SYBR qpcr Mix QPS-201T 1 ml x 1 QPS-201 1.67 ml x 3 Contents [1] Introduction [2] Components [3] Primer design [4] Template DNA [5]
T7-Based RNA Amplification Protocol (in progress)
 T7-Based RNA Amplification Protocol (in progress) Jacqueline Ann Lopez (modifications) Amy Cash & Justen Andrews INTRODUCTION T7 RNA Amplification, a technique originally developed in the laboratory of
T7-Based RNA Amplification Protocol (in progress) Jacqueline Ann Lopez (modifications) Amy Cash & Justen Andrews INTRODUCTION T7 RNA Amplification, a technique originally developed in the laboratory of
Brian Williams, Richele Gwirtz Version 1.0, December 29, 2003
 Direct labeling of mouse genomic DNA for microarray hybridization using the Klenow fragment of E. coli DNA polymerase I and Cy3 dctp. This is a detailed version of the protocol published in Genomic DNA
Direct labeling of mouse genomic DNA for microarray hybridization using the Klenow fragment of E. coli DNA polymerase I and Cy3 dctp. This is a detailed version of the protocol published in Genomic DNA
