SUPPLEMENTARY INFORMATION
|
|
|
- Flora Shaw
- 5 years ago
- Views:
Transcription
1 doi:138/nature10532 a Human b Platypus Density Ensembl protein coding Ensembl lincrna New exons (protein coding) Intergenic multi exonic loci Density c Total per base read coverage (log2 transform) d Total per base read coverage (log2 transform) Sequence conservation (PhastCons) Sequence conservation (PhastCons) Intron 5' Exon 3' Intron Intron 5' Exon 3' Intron Position along the gene Position along the gene Supplementary Figure 1. Expression level and sequence conservation of newly annotated regions in the human and platypus genomes. a, b Density plot of the total per-base read coverage (all samples combined) for four categories: exons of Ensemblannotated protein-coding genes (black) and large intervening noncoding RNA (lincrna) genes (green), new exons added to previously known (Ensembl) protein-coding genes (blue), and exons of intergenic multi-exonic transcribed loci (red). The read coverage was log2-transformed (an offset of 1 was added to preserve null values). c, d Variation of the mean PhastCons score around exon boundaries, for the same categories of exons. The PhastCons score was computed for 100 positions, divided into two 50 bp segments centered on the exon boundaries, and was averaged over all exons in a class. Only internal exons with a length of at least 50 bp and with flanking introns larger than 25 bp were considered for this analysis. 1
2 RESEARCH SUPPLEMENTARY INFORMATION brain chimp M 4 bonobo F 2 chimp M 5 chimp M 2 chimp M 3 human M 4 human M 5 human M 3 kidney cerebellum heart < 0.9 < 0.7 < 0.5 liver chicken M 2 testis 0.1 platypus M 2 platypus M 3 opossum M 2 Supplementary Figure 2. Mammalian gene expression phylogenies. Neighbor-joining trees based on distance matrices comprising all pairwise expression level distances (1 rho, Spearman s correlation coefficient) for the six different organs. Trees are drawn to the same scale (indicated by the scale bar). Bootstrap values (i.e., proportions of replicate trees that have the branching pattern as in the majority-rule consensus tree shown) are indicated by circles at the corresponding nodes: > 0.9 (white fill), < 0.9 (yellow), < 0.7 (red). Mammalian species names are color-coded according to the three major mammalian lineages: monotremes (i.e., platypus; light blue), marsupials (opossum; dark blue), and eutherians (mice and primates; black). 2
3 RESEARCH brain cerebellum heart < 0.9 < 0.7 < 0.5 chimp M 4 bonobo F 2 chimp M 5 chimp M 2 chimp M 3 human M 4 human M 5 human M 3 kidney liver testis 10 chicken M 2 platypus M 2 platypus M 3 opossum M 2 Supplementary Figure 3. Mammalian gene expression phylogenies. Neighbor-joining trees based on distance matrices comprising all pairwise expression level Euclidean distances for the six different organs. Trees are drawn to the same scale (indicated by the scale bar). Bootstrap values (i.e., proportions of replicate trees that have the branching pattern as in the majority-rule consensus tree shown) are indicated by circles at the corresponding nodes: > 0.9 (white fill), < 0.9 (yellow), < 0.7 (red). Mammalian species names are color-coded according to the three major mammalian lineages: monotremes (i.e., platypus; light blue), marsupials (opossum; dark blue), and eutherians (mice and primates; black). 3
4 RESEARCH SUPPLEMENTARY INFORMATION Supplementary Figure 4. Sex-biased expression of the vitellogenin egg yolk genes in platypus and chicken livers. a, Median gene expression levels (based on read counts) of genes in livers of male (M) and female (F) chicken and platypus animals (X-axis) are plotted against expression differences (shown as log2 fold-changes) between males and females (Y-axis). Top and bottom boxes contain genes with zero expression in females or males, respectively. Vitellogenin genes or gene fragments are marked in green: in the chicken plot, the green plus sign indicates the VIT2 gene (further illustrated in subpanel b), the more central green circle indicates the VIT1 gene, and the less central green circle represents VIT3 (VIT2 and VIT3 have significantly higher expression levels in females, see also Supplementary Table 41); in the platypus plot, the plus sign indicates the contig containing the major part of the predicted platypus VIT gene, whereas the three green circles represent contigs containing the remaining platypus VIT exons. b, Ensembl annotation of chicken VIT2 and a major part of platypus VIT and the corresponding read coverage in females and males. 4
5 RESEARCH brain human M 5 human M 3 chimp M 4 bonobo F 2 chimp M 2 chimp M 5 chimp M 3 human M 4 kidney cerebellum liver heart testis < 0.9 < 0.7 < Supplementary Figure 5. Primate gene expression phylogenies. Neighbor-joining trees based on distance matrices comprising all pairwise expression level distances (1 rho, Spearman s correlation coefficient) for the six different organs. Note that the underlying gene expression values were calculated based on constitutive orthologous exons (from the 13, orthologous genes) that are perfectly aligned (no gaps permitted) between the six primate species (see Supplementary Note section 1.6 for details). Thus, we rule out any gene expression variation that might arise when gene expression levels are computed for potentially different sequences (e.g., as a result of annotation or biological/genomic differences) among 1 1 orthologs from different species. Bootstrap values (i.e., proportions of replicate trees that have the branching pattern as in the majority-rule consensus tree shown) are indicated by circles at the corresponding nodes: > 0.9 (white fill), < 0.9 (yellow), < 0.7 (red). 5
6 RESEARCH SUPPLEMENTARY INFORMATION a ape b platypus Supplementary Figure 6. Comparisons of s among mammalian lineages. a, b, Bootstrap pseudosamples (1,000) were generated that each comprise 5,636 sets of orthologues from one randomly sampled individual per species or group of species (in the case of apes). For each of these pseudosamples, Neighbor-joining trees of two species and the outgroup (chicken) were constructed; the total length of the branch (or branches) from the last common ancestral node to the respective terminal nodes was assessed for each of these trees. 95% confidence intervals are shown. Statistical significance was assessed by counting the number of trees in which one or the other species/lineage had longer s: [] indicates two-tailed P < (i.e., one lineage with longer branches in more than 97.5% of replicates; after Bonferroni correction for the six tests performed for the six tissues in each comparison). To factor out within-species differences, similar analyses were performed that measure s from the last common ancestral node of a given species-pair to the last common ancestral node of individuals for each of the species in the pair (i.e., terminal branches were not considered in this analysis; Supplementary Fig. 6). 6
7 RESEARCH human macaque gorilla opossum br cb ht kd lv orangutan platypus br ht kd lv Supplementary Figure 7. Comparisons of s among mammalian lineages. Bootstrap pseudosamples (1,000) were generated that each comprise 5,636 sets of orthologues from one randomly sampled individual per species. For each of these pseudosamples, Neighbor-joining trees of two species and the outgroup (chicken) were constructed. To factor out within-species branch length differences, the total length of the branch (or branches) from the last common ancestral node of the given species pair to the most recent common ancestral nodes of individuals for each species in the pair, respectively, was assessed for each of these trees. 95% confidence intervals are shown. Statistical significance was assessed by counting the number of trees in which one or the other species/lineage had longer s: [] indicates two-tailed P < (i.e., one lineage with longer branches in more than 97.5% of replicates; after Bonferroni correction for the six tests performed for the six tissues in each comparison). 7
8 RESEARCH SUPPLEMENTARY INFORMATION TEX13B (testis) testis expressed 13B SHBG (testis) sex hormone-binding globulin MYL6B (heart) myosin, light chain 6B, alkali, smooth muscle FAM185A (brain) family with sequence similarity 185, member A ALX4 (cerbellum) ALX homeobox 4 A4GALT (cerebellum) alpha 1,4-galactosyltransferase max-min / log2(rpkm) SPO11 (testis) SPO11 meiotic protein covalently bound to DSB homolog (S. cerevisiae) ZNF512 (brain) zinc finger protein 512 ATRX (testis) alpha thalassemia/mental retardation syndrome X-linked (RAD54 homolog, S. cerevisiae) chicken platypus opossum macaque orangutan gorilla chimpanzee bonobo human Supplementary Figure 8. Genes with shifts in optimal expression values (individual tissues). Examples of genes that evolved new optimal expression levels in a given mammalian lineage or species (as detected using a phylogenetic maximum likelihood procedure see main text and Supplementary Note) are shown (details regarding these genes and all other statistically significant expression changes in mammals are provided in Supplementary Tables 9-24): TEX13B (ENSG ), specifically expressed in spermatogonia (i.e., pre-meiotic cells), was significantly upregulated in human testis; SHBG (ENSG ), a key gene in testosterone metabolism evolved high expression in chimpanzee and bonobo testes; MYL6B (ENSG ), encoding a myosin light chain subunit, became upregulated in gorilla heart; FAM185A (ENSG ), a gene with unknown function evolved significantly higher expression in orangutan brain; A4GALT (ENSG ; involved in glycolipid metabolism) and ALX4 (ENSG ; previously implicated in human cranial phenotypes) shifted to lower expression in great ape cerebellum; SPO11 (ENSG ; key for double strand formation during meiotic recombination) evolved high expression in testes; ZNF512 (ENSG ; zinc finger protein 512, a potential transcription factor) shifted to high expression in opossum brain; and ATRX (ENSG ; various molecular functions associated with e.g. chromosome segregation; involved in mental disorders) became downregulated in therian testis. Information about all genes was derived from the OMIM database ( omim)
9 RESEARCH Mitogen-activated protein kinase 7 interacting protein 3 (X-linked) chicken platypus opossum macaque orangutan gorilla chimpanzee bonobo human max-min / log2(rpkm) GNAI3 (therians): guanine nucleotide binding protein (G protein), alpha inhibiting activity polypeptide 3 brain ce rebellum heart kidney liver testis Supplementary Figure 9. Genes with expression level shifts (multiple tissues). Examples of genes that evolved new optimal expression levels in multiple therian tissues are shown (details regarding these genes and all other statistically significant expression changes are provided in Supplementary Tables 9-24). MAP3K7IP3 (an X-linked gene; ENSG ) evolved significantly lower expression in therian heart and kidney and tends to have lower expression in eutherian/therian liver, testis, brain and cerebellum. GNAI3 (a Gi alpha-like protein; ENSG ) became significantly downregulated in the brain and cerebellum and tends to show reduced expression in the other therian tissues as well. 9
1.15 Influence of total read coverage on the detection of transcribed proteincoding. 2 Supplementary Note Tables Supplementary Note Figures 40
 doi:10.1038/nature10532 Contents 1 Supporting text 1 1.1 Biological sample collection and RNA extraction... 1 1.2 RNA sequencing... 1 1.3 Initial read mapping... 2 1.4 Refinement of genomic annotations...
doi:10.1038/nature10532 Contents 1 Supporting text 1 1.1 Biological sample collection and RNA extraction... 1 1.2 RNA sequencing... 1 1.3 Initial read mapping... 2 1.4 Refinement of genomic annotations...
SUPPLEMENTARY INFORMATION
 18Pura Brain Liver 2 kb 9Mtmr Heart Brain 1.4 kb 15Ppara 1.5 kb IPTpcc 1.3 kb GAPDH 1.5 kb IPGpr1 2 kb GAPDH 1.5 kb 17Tcte 13Elov12 Lung Thymus 6 kb 5 kb 17Tbcc Kidney Embryo 4.5 kb 3.7 kb GAPDH 1.5 kb
18Pura Brain Liver 2 kb 9Mtmr Heart Brain 1.4 kb 15Ppara 1.5 kb IPTpcc 1.3 kb GAPDH 1.5 kb IPGpr1 2 kb GAPDH 1.5 kb 17Tcte 13Elov12 Lung Thymus 6 kb 5 kb 17Tbcc Kidney Embryo 4.5 kb 3.7 kb GAPDH 1.5 kb
mrna Sequencing Quality Control (V6)
 mrna Sequencing Quality Control (V6) Notes: the following analyses are based on 8 adult brains sequenced in USC and Yale 1. Error Rates The error rates of each sequencing cycle are reported for 120 tiles
mrna Sequencing Quality Control (V6) Notes: the following analyses are based on 8 adult brains sequenced in USC and Yale 1. Error Rates The error rates of each sequencing cycle are reported for 120 tiles
Figure S1 Correlation in size of analogous introns in mouse and teleost Piccolo genes. Mouse intron size was plotted against teleost intron size for t
 Figure S1 Correlation in size of analogous introns in mouse and teleost Piccolo genes. Mouse intron size was plotted against teleost intron size for the pcloa genes of zebrafish, green spotted puffer (listed
Figure S1 Correlation in size of analogous introns in mouse and teleost Piccolo genes. Mouse intron size was plotted against teleost intron size for the pcloa genes of zebrafish, green spotted puffer (listed
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature09944 Supplementary Figure 1. Establishing DNA sequence similarity thresholds for phylum and genus levels Sequence similarity distributions of pairwise alignments of 40 universal single
doi:10.1038/nature09944 Supplementary Figure 1. Establishing DNA sequence similarity thresholds for phylum and genus levels Sequence similarity distributions of pairwise alignments of 40 universal single
GENETICS - CLUTCH CH.15 GENOMES AND GENOMICS.
 !! www.clutchprep.com CONCEPT: OVERVIEW OF GENOMICS Genomics is the study of genomes in their entirety Bioinformatics is the analysis of the information content of genomes - Genes, regulatory sequences,
!! www.clutchprep.com CONCEPT: OVERVIEW OF GENOMICS Genomics is the study of genomes in their entirety Bioinformatics is the analysis of the information content of genomes - Genes, regulatory sequences,
How does the human genome stack up? Genomic Size. Genome Size. Number of Genes. Eukaryotic genomes are generally larger.
 How does the human genome stack up? Organism Human (Homo sapiens) Laboratory mouse (M. musculus) Mustard weed (A. thaliana) Roundworm (C. elegans) Fruit fly (D. melanogaster) Yeast (S. cerevisiae) Bacterium
How does the human genome stack up? Organism Human (Homo sapiens) Laboratory mouse (M. musculus) Mustard weed (A. thaliana) Roundworm (C. elegans) Fruit fly (D. melanogaster) Yeast (S. cerevisiae) Bacterium
Nature Genetics: doi: /ng.3556 INTEGRATED SUPPLEMENTARY FIGURE TEMPLATE. Supplementary Figure 1
 INTEGRATED SUPPLEMENTARY FIGURE TEMPLATE Supplementary Figure 1 REF6 expression in transgenic lines. (a,b) Expression of REF6 in REF6-HA ref6 and REF6ΔZnF-HA ref6 plants detected by RT qpcr (a) and immunoblot
INTEGRATED SUPPLEMENTARY FIGURE TEMPLATE Supplementary Figure 1 REF6 expression in transgenic lines. (a,b) Expression of REF6 in REF6-HA ref6 and REF6ΔZnF-HA ref6 plants detected by RT qpcr (a) and immunoblot
Outline. Computational Genomics and Molecular Biology. Genes Encode Proteins. DNA forms a double stranded helix. DNA replication T C A G
 Computational Genomics and Molecular Biology Dannie Durand Fall 2004 Lecture 1 Genes Encode Proteins DNA forms a double stranded helix GTGCACCTGACTCCTGAG A gene is a DNA sequence V H L T P E A protein
Computational Genomics and Molecular Biology Dannie Durand Fall 2004 Lecture 1 Genes Encode Proteins DNA forms a double stranded helix GTGCACCTGACTCCTGAG A gene is a DNA sequence V H L T P E A protein
Nature Genetics: doi: /ng Supplementary Figure 1. Neighbor-joining tree of the 183 wild, cultivated, and weedy rice accessions.
 Supplementary Figure 1 Neighbor-joining tree of the 183 wild, cultivated, and weedy rice accessions. Relationships of cultivated and wild rice correspond to previously observed relationships 40. Wild rice
Supplementary Figure 1 Neighbor-joining tree of the 183 wild, cultivated, and weedy rice accessions. Relationships of cultivated and wild rice correspond to previously observed relationships 40. Wild rice
SUPPLEMENTARY INFORMATION
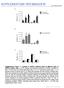 SUPPLEMENTARY INFORMATION doi:10.1038/nature12474 Supplementary Figure 1 Analysis of mtdna mutation loads in different types of mtdna mutator mice. a, PCR, cloning, and sequencing analysis of mtdna mutations
SUPPLEMENTARY INFORMATION doi:10.1038/nature12474 Supplementary Figure 1 Analysis of mtdna mutation loads in different types of mtdna mutator mice. a, PCR, cloning, and sequencing analysis of mtdna mutations
Supplementary Figure 1 An overview of pirna biogenesis during fetal mouse reprogramming. (a) (b)
 Supplementary Figure 1 An overview of pirna biogenesis during fetal mouse reprogramming. (a) A schematic overview of the production and amplification of a single pirna from a transposon transcript. The
Supplementary Figure 1 An overview of pirna biogenesis during fetal mouse reprogramming. (a) A schematic overview of the production and amplification of a single pirna from a transposon transcript. The
Higher Unit 1: DNA and the Genome Topic 1.1 The Structure and Organisation of DNA
 Higher Unit : DNA and the Genome Topic. The Structure and Organisation of DNA. Which of the following diagrams shows the correct structure of DNA? 2. A section of double stranded DNA was found to have
Higher Unit : DNA and the Genome Topic. The Structure and Organisation of DNA. Which of the following diagrams shows the correct structure of DNA? 2. A section of double stranded DNA was found to have
The genome of Leishmania panamensis: insights into genomics of the L. (Viannia) subgenus.
 SUPPLEMENTARY INFORMATION The genome of Leishmania panamensis: insights into genomics of the L. (Viannia) subgenus. Alejandro Llanes, Carlos Mario Restrepo, Gina Del Vecchio, Franklin José Anguizola, Ricardo
SUPPLEMENTARY INFORMATION The genome of Leishmania panamensis: insights into genomics of the L. (Viannia) subgenus. Alejandro Llanes, Carlos Mario Restrepo, Gina Del Vecchio, Franklin José Anguizola, Ricardo
Answer: Sequence overlap is required to align the sequenced segments relative to each other.
 14 Genomes and Genomics WORKING WITH THE FIGURES 1. Based on Figure 14-2, why must the DNA fragments sequenced overlap in order to obtain a genome sequence? Answer: Sequence overlap is required to align
14 Genomes and Genomics WORKING WITH THE FIGURES 1. Based on Figure 14-2, why must the DNA fragments sequenced overlap in order to obtain a genome sequence? Answer: Sequence overlap is required to align
Molecular Genetics of Disease and the Human Genome Project
 9 Molecular Genetics of Disease and the Human Genome Project Fig. 1. The 23 chromosomes in the human genome. There are 22 autosomes (chromosomes 1 to 22) and two sex chromosomes (X and Y). Females inherit
9 Molecular Genetics of Disease and the Human Genome Project Fig. 1. The 23 chromosomes in the human genome. There are 22 autosomes (chromosomes 1 to 22) and two sex chromosomes (X and Y). Females inherit
Guided tour to Ensembl
 Guided tour to Ensembl Introduction Introduction to the Ensembl project Walk-through of the browser Variations and Functional Genomics Comparative Genomics BioMart Ensembl Genome browser http://www.ensembl.org
Guided tour to Ensembl Introduction Introduction to the Ensembl project Walk-through of the browser Variations and Functional Genomics Comparative Genomics BioMart Ensembl Genome browser http://www.ensembl.org
DNA Function: Information Transmission
 DNA Function: Information Transmission DNA is called the code of life. What does it code for? *the information ( code ) to make proteins! Why are proteins so important? Nearly every function of a living
DNA Function: Information Transmission DNA is called the code of life. What does it code for? *the information ( code ) to make proteins! Why are proteins so important? Nearly every function of a living
CS 262 Lecture 14 Notes Human Genome Diversity, Coalescence and Haplotypes
 CS 262 Lecture 14 Notes Human Genome Diversity, Coalescence and Haplotypes Coalescence Scribe: Alex Wells 2/18/16 Whenever you observe two sequences that are similar, there is actually a single individual
CS 262 Lecture 14 Notes Human Genome Diversity, Coalescence and Haplotypes Coalescence Scribe: Alex Wells 2/18/16 Whenever you observe two sequences that are similar, there is actually a single individual
Nature Methods: doi: /nmeth Supplementary Figure 1. Construction of a sensitive TetR mediated auxotrophic off-switch.
 Supplementary Figure 1 Construction of a sensitive TetR mediated auxotrophic off-switch. A Production of the Tet repressor in yeast when conjugated to either the LexA4 or LexA8 promoter DNA binding sequences.
Supplementary Figure 1 Construction of a sensitive TetR mediated auxotrophic off-switch. A Production of the Tet repressor in yeast when conjugated to either the LexA4 or LexA8 promoter DNA binding sequences.
Lecture 12. Genomics. Mapping. Definition Species sequencing ESTs. Why? Types of mapping Markers p & Types
 Lecture 12 Reading Lecture 12: p. 335-338, 346-353 Lecture 13: p. 358-371 Genomics Definition Species sequencing ESTs Mapping Why? Types of mapping Markers p.335-338 & 346-353 Types 222 omics Interpreting
Lecture 12 Reading Lecture 12: p. 335-338, 346-353 Lecture 13: p. 358-371 Genomics Definition Species sequencing ESTs Mapping Why? Types of mapping Markers p.335-338 & 346-353 Types 222 omics Interpreting
Evolutionary dynamics and tissue specificity of human long noncoding RNAs in six mammals
 Research Evolutionary dynamics and tissue specificity of human long noncoding RNAs in six mammals Stefan Washietl, 1 Manolis Kellis, 1,2,5,6 and Manuel Garber 2,3,4,5,6 1 Computer Science and Artificial
Research Evolutionary dynamics and tissue specificity of human long noncoding RNAs in six mammals Stefan Washietl, 1 Manolis Kellis, 1,2,5,6 and Manuel Garber 2,3,4,5,6 1 Computer Science and Artificial
TECH NOTE Pushing the Limit: A Complete Solution for Generating Stranded RNA Seq Libraries from Picogram Inputs of Total Mammalian RNA
 TECH NOTE Pushing the Limit: A Complete Solution for Generating Stranded RNA Seq Libraries from Picogram Inputs of Total Mammalian RNA Stranded, Illumina ready library construction in
TECH NOTE Pushing the Limit: A Complete Solution for Generating Stranded RNA Seq Libraries from Picogram Inputs of Total Mammalian RNA Stranded, Illumina ready library construction in
Cufflinks Scripture Both. GENCODE/ UCSC/ RefSeq 0.18% 0.92% Scripture
 Cabili_SuppFig_1 a 1.9.8.7.6.5.4.3.2.1 Partially compatible % Partially recovered RefSeq coding Fully compatible % Fully recovered RefSeq coding Cufflinks Scripture Both b 18.5% GENCODE/ UCSC/ RefSeq Cufflinks
Cabili_SuppFig_1 a 1.9.8.7.6.5.4.3.2.1 Partially compatible % Partially recovered RefSeq coding Fully compatible % Fully recovered RefSeq coding Cufflinks Scripture Both b 18.5% GENCODE/ UCSC/ RefSeq Cufflinks
Unit 1: DNA and the Genome. Sub-Topic (1.3) Gene Expression
 Unit 1: DNA and the Genome Sub-Topic (1.3) Gene Expression Unit 1: DNA and the Genome Sub-Topic (1.3) Gene Expression On completion of this subtopic I will be able to State the meanings of the terms genotype,
Unit 1: DNA and the Genome Sub-Topic (1.3) Gene Expression Unit 1: DNA and the Genome Sub-Topic (1.3) Gene Expression On completion of this subtopic I will be able to State the meanings of the terms genotype,
I. Gene Expression Figure 1: Central Dogma of Molecular Biology
 I. Gene Expression Figure 1: Central Dogma of Molecular Biology Central Dogma: Gene Expression: RNA Structure RNA nucleotides contain the pentose sugar Ribose instead of deoxyribose. Contain the bases
I. Gene Expression Figure 1: Central Dogma of Molecular Biology Central Dogma: Gene Expression: RNA Structure RNA nucleotides contain the pentose sugar Ribose instead of deoxyribose. Contain the bases
Transcription. The sugar molecule found in RNA is ribose, rather than the deoxyribose found in DNA.
 Transcription RNA (ribonucleic acid) is a key intermediary between a DNA sequence and a polypeptide. RNA is an informational polynucleotide similar to DNA, but it differs from DNA in three ways: RNA generally
Transcription RNA (ribonucleic acid) is a key intermediary between a DNA sequence and a polypeptide. RNA is an informational polynucleotide similar to DNA, but it differs from DNA in three ways: RNA generally
Lecture for Wednesday. Dr. Prince BIOL 1408
 Lecture for Wednesday Dr. Prince BIOL 1408 THE FLOW OF GENETIC INFORMATION FROM DNA TO RNA TO PROTEIN Copyright 2009 Pearson Education, Inc. Genes are expressed as proteins A gene is a segment of DNA that
Lecture for Wednesday Dr. Prince BIOL 1408 THE FLOW OF GENETIC INFORMATION FROM DNA TO RNA TO PROTEIN Copyright 2009 Pearson Education, Inc. Genes are expressed as proteins A gene is a segment of DNA that
Course Information. Introduction to Algorithms in Computational Biology Lecture 1. Relations to Some Other Courses
 Course Information Introduction to Algorithms in Computational Biology Lecture 1 Meetings: Lecture, by Dan Geiger: Mondays 16:30 18:30, Taub 4. Tutorial, by Ydo Wexler: Tuesdays 10:30 11:30, Taub 2. Grade:
Course Information Introduction to Algorithms in Computational Biology Lecture 1 Meetings: Lecture, by Dan Geiger: Mondays 16:30 18:30, Taub 4. Tutorial, by Ydo Wexler: Tuesdays 10:30 11:30, Taub 2. Grade:
Transcription and Translation. DANILO V. ROGAYAN JR. Faculty, Department of Natural Sciences
 Transcription and Translation DANILO V. ROGAYAN JR. Faculty, Department of Natural Sciences Protein Structure Made up of amino acids Polypeptide- string of amino acids 20 amino acids are arranged in different
Transcription and Translation DANILO V. ROGAYAN JR. Faculty, Department of Natural Sciences Protein Structure Made up of amino acids Polypeptide- string of amino acids 20 amino acids are arranged in different
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature17405 Supplementary Information 1 Determining a suitable lower size-cutoff for sequence alignments to the nuclear genome Analyses of nuclear DNA sequences from archaic genomes have until
doi:10.1038/nature17405 Supplementary Information 1 Determining a suitable lower size-cutoff for sequence alignments to the nuclear genome Analyses of nuclear DNA sequences from archaic genomes have until
Lecture Summary: Regulation of transcription. General mechanisms-what are the major regulatory points?
 BCH 401G Lecture 37 Andres Lecture Summary: Regulation of transcription. General mechanisms-what are the major regulatory points? RNA processing: Capping, polyadenylation, splicing. Why process mammalian
BCH 401G Lecture 37 Andres Lecture Summary: Regulation of transcription. General mechanisms-what are the major regulatory points? RNA processing: Capping, polyadenylation, splicing. Why process mammalian
Physical Anthropology 1 Milner-Rose
 Physical Anthropology 1 Milner-Rose Chapter 3 Genetics: Reproducing Life and Producing Variation Our Origins By Clark Spencer Larsen Natural Selection operates on the levels of the 1. living, behaving
Physical Anthropology 1 Milner-Rose Chapter 3 Genetics: Reproducing Life and Producing Variation Our Origins By Clark Spencer Larsen Natural Selection operates on the levels of the 1. living, behaving
Introduction to Algorithms in Computational Biology Lecture 1
 Introduction to Algorithms in Computational Biology Lecture 1 Background Readings: The first three chapters (pages 1-31) in Genetics in Medicine, Nussbaum et al., 2001. This class has been edited from
Introduction to Algorithms in Computational Biology Lecture 1 Background Readings: The first three chapters (pages 1-31) in Genetics in Medicine, Nussbaum et al., 2001. This class has been edited from
MODULE 5: TRANSLATION
 MODULE 5: TRANSLATION Lesson Plan: CARINA ENDRES HOWELL, LEOCADIA PALIULIS Title Translation Objectives Determine the codons for specific amino acids and identify reading frames by looking at the Base
MODULE 5: TRANSLATION Lesson Plan: CARINA ENDRES HOWELL, LEOCADIA PALIULIS Title Translation Objectives Determine the codons for specific amino acids and identify reading frames by looking at the Base
CHAPTER 21 LECTURE SLIDES
 CHAPTER 21 LECTURE SLIDES Prepared by Brenda Leady University of Toledo To run the animations you must be in Slideshow View. Use the buttons on the animation to play, pause, and turn audio/text on or off.
CHAPTER 21 LECTURE SLIDES Prepared by Brenda Leady University of Toledo To run the animations you must be in Slideshow View. Use the buttons on the animation to play, pause, and turn audio/text on or off.
Unit 2: Biological basis of life, heredity, and genetics
 Unit 2: Biological basis of life, heredity, and genetics 1 Issues with Darwin's Evolutionary Theory??? 2 Cells - General Composition Organelles - substructures in the cell which do different things involved
Unit 2: Biological basis of life, heredity, and genetics 1 Issues with Darwin's Evolutionary Theory??? 2 Cells - General Composition Organelles - substructures in the cell which do different things involved
CHAPTERS , 17: Eukaryotic Genetics
 CHAPTERS 14.1 14.6, 17: Eukaryotic Genetics 1. Review the levels of DNA packing within the eukaryote nucleus. Label each level. (A similar diagram is on pg 188 of your textbook.) 2. How do the coding regions
CHAPTERS 14.1 14.6, 17: Eukaryotic Genetics 1. Review the levels of DNA packing within the eukaryote nucleus. Label each level. (A similar diagram is on pg 188 of your textbook.) 2. How do the coding regions
30 Gene expression: Transcription
 30 Gene expression: Transcription Gene structure. o Exons coding region of DNA. o Introns non-coding region of DNA. o Introns are interspersed between exons of a single gene. o Promoter region helps enzymes
30 Gene expression: Transcription Gene structure. o Exons coding region of DNA. o Introns non-coding region of DNA. o Introns are interspersed between exons of a single gene. o Promoter region helps enzymes
Hominoid-Specific De Novo Protein-Coding Genes Originating from Long Non-Coding RNAs
 Hominoid-Specific De Novo Protein-Coding Genes Originating from Long Non-Coding RNAs Chen Xie 1., Yong E. Zhang 2., Jia-Yu Chen 3., Chu-Jun Liu 3, Wei-Zhen Zhou 1, Ying Li 3, Mao Zhang 3, Rongli Zhang
Hominoid-Specific De Novo Protein-Coding Genes Originating from Long Non-Coding RNAs Chen Xie 1., Yong E. Zhang 2., Jia-Yu Chen 3., Chu-Jun Liu 3, Wei-Zhen Zhou 1, Ying Li 3, Mao Zhang 3, Rongli Zhang
FINDING THE PAIN GENE How do geneticists connect a specific gene with a specific phenotype?
 FINDING THE PAIN GENE How do geneticists connect a specific gene with a specific phenotype? 1 Linkage & Recombination HUH? What? Why? Who cares? How? Multiple choice question. Each colored line represents
FINDING THE PAIN GENE How do geneticists connect a specific gene with a specific phenotype? 1 Linkage & Recombination HUH? What? Why? Who cares? How? Multiple choice question. Each colored line represents
Result Tables The Result Table, which indicates chromosomal positions and annotated gene names, promoter regions and CpG islands, is the best way for
 Result Tables The Result Table, which indicates chromosomal positions and annotated gene names, promoter regions and CpG islands, is the best way for you to discover methylation changes at specific genomic
Result Tables The Result Table, which indicates chromosomal positions and annotated gene names, promoter regions and CpG islands, is the best way for you to discover methylation changes at specific genomic
Temperature sensitive gene expression: Two rules.
 32 Research Notes Dros. Inf. Serv. 92 (2009) Temperature sensitive gene expression: Two rules. Gupta, Anand P. Johnson C. Smith University, Department of Science and Mathematics, 100 Beatties Ford Road,
32 Research Notes Dros. Inf. Serv. 92 (2009) Temperature sensitive gene expression: Two rules. Gupta, Anand P. Johnson C. Smith University, Department of Science and Mathematics, 100 Beatties Ford Road,
COMPETITOR NAMES: TEAM NAME: TEAM NUMBER:
 COMPETITOR NAMES: TEAM NAME: TEAM NUMBER: Section 1:Crosses In a fictional species of mice, with species name Mus SciOlyian, fur color is controlled by a single autosomal gene. The allele for brown fur
COMPETITOR NAMES: TEAM NAME: TEAM NUMBER: Section 1:Crosses In a fictional species of mice, with species name Mus SciOlyian, fur color is controlled by a single autosomal gene. The allele for brown fur
A primer on the structure and function of genes
 A primer on the structure and function of genes What is the definition of a gene? GENE: the genetic element which is transmitted from parent to offspring during the process of reproduction that influences
A primer on the structure and function of genes What is the definition of a gene? GENE: the genetic element which is transmitted from parent to offspring during the process of reproduction that influences
Review of Protein (one or more polypeptide) A polypeptide is a long chain of..
 Gene expression Review of Protein (one or more polypeptide) A polypeptide is a long chain of.. In a protein, the sequence of amino acid determines its which determines the protein s A protein with an enzymatic
Gene expression Review of Protein (one or more polypeptide) A polypeptide is a long chain of.. In a protein, the sequence of amino acid determines its which determines the protein s A protein with an enzymatic
Chimp Chunk 3-14 Annotation by Matthew Kwong, Ruth Howe, and Hao Yang
 Chimp Chunk 3-14 Annotation by Matthew Kwong, Ruth Howe, and Hao Yang Ruth Howe Bio 434W April 1, 2010 INTRODUCTION De novo annotation is the process by which a finished genomic sequence is searched for
Chimp Chunk 3-14 Annotation by Matthew Kwong, Ruth Howe, and Hao Yang Ruth Howe Bio 434W April 1, 2010 INTRODUCTION De novo annotation is the process by which a finished genomic sequence is searched for
Differential Gene Expression
 Biology 4361 Developmental Biology Differential Gene Expression June 19, 2008 Differential Gene Expression Overview Chromatin structure Gene anatomy RNA processing and protein production Initiating transcription:
Biology 4361 Developmental Biology Differential Gene Expression June 19, 2008 Differential Gene Expression Overview Chromatin structure Gene anatomy RNA processing and protein production Initiating transcription:
Supplemental Data. Zhou et al. (2016). Plant Cell /tpc
 Supplemental Figure 1. Confirmation of mutant mapping results. (A) Complementation assay with stably transformed genomic fragments (ComN-N) (2 kb upstream of TSS and 1.5 kb downstream of TES) and CaMV
Supplemental Figure 1. Confirmation of mutant mapping results. (A) Complementation assay with stably transformed genomic fragments (ComN-N) (2 kb upstream of TSS and 1.5 kb downstream of TES) and CaMV
Genomes summary. Bacterial genome sizes
 Genomes summary 1. >930 bacterial genomes sequenced. 2. Circular. Genes densely packed. 3. 2-10 Mbases, 470-7,000 genes 4. Genomes of >200 eukaryotes (45 higher ) sequenced. 5. Linear chromosomes 6. On
Genomes summary 1. >930 bacterial genomes sequenced. 2. Circular. Genes densely packed. 3. 2-10 Mbases, 470-7,000 genes 4. Genomes of >200 eukaryotes (45 higher ) sequenced. 5. Linear chromosomes 6. On
7.03, 2005, Lecture 20 EUKARYOTIC GENES AND GENOMES I
 7.03, 2005, Lecture 20 EUKARYOTIC GENES AND GENOMES I For the last several lectures we have been looking at how one can manipulate prokaryotic genomes and how prokaryotic genes are regulated. In the next
7.03, 2005, Lecture 20 EUKARYOTIC GENES AND GENOMES I For the last several lectures we have been looking at how one can manipulate prokaryotic genomes and how prokaryotic genes are regulated. In the next
Identification of individual motifs on the genome scale. Some slides are from Mayukh Bhaowal
 Identification of individual motifs on the genome scale Some slides are from Mayukh Bhaowal Two papers Nature 423, 241-254 (15 May 2003) Sequencing and comparison of yeast species to identify genes and
Identification of individual motifs on the genome scale Some slides are from Mayukh Bhaowal Two papers Nature 423, 241-254 (15 May 2003) Sequencing and comparison of yeast species to identify genes and
SUPPLEMENTARY FIGURES AND TABLES. Exploration of mirna families for hypotheses generation
 SUPPLEMENTARY FIGURES AND TABLES Exploration of mirna families for hypotheses generation Timothy K. K. Kamanu, Aleksandar Radovanovic, John A. C. Archer, and Vladimir B. Bajic King Abdullah University
SUPPLEMENTARY FIGURES AND TABLES Exploration of mirna families for hypotheses generation Timothy K. K. Kamanu, Aleksandar Radovanovic, John A. C. Archer, and Vladimir B. Bajic King Abdullah University
Supplementary Fig. S1. Building a training set of cardiac enhancers. (A-E) Empirical validation of candidate enhancers containing matches to Twi and
 Supplementary Fig. S1. Building a training set of cardiac enhancers. (A-E) Empirical validation of candidate enhancers containing matches to Twi and Tin TFBS motifs and located in the flanking or intronic
Supplementary Fig. S1. Building a training set of cardiac enhancers. (A-E) Empirical validation of candidate enhancers containing matches to Twi and Tin TFBS motifs and located in the flanking or intronic
Annotating the Genome (H)
 Annotating the Genome (H) Annotation principles (H1) What is annotation? In general: annotation = explanatory note* What could be useful as an annotation of a DNA sequence? an amino acid sequence? What
Annotating the Genome (H) Annotation principles (H1) What is annotation? In general: annotation = explanatory note* What could be useful as an annotation of a DNA sequence? an amino acid sequence? What
Wednesday, November 22, 17. Exons and Introns
 Exons and Introns Introns and Exons Exons: coded regions of DNA that get transcribed and translated into proteins make up 5% of the genome Introns and Exons Introns: non-coded regions of DNA Must be removed
Exons and Introns Introns and Exons Exons: coded regions of DNA that get transcribed and translated into proteins make up 5% of the genome Introns and Exons Introns: non-coded regions of DNA Must be removed
Mammalian non-cg methylations are conserved and cell-type specific and may have been involved in the evolution of transposon elements
 Mammalian non-cg methylations are conserved and cell-type specific and may have been involved in the evolution of transposon elements Weilong Guo, Michael Zhang, Hong Wu Supplementary Figures Fig. S1-S16
Mammalian non-cg methylations are conserved and cell-type specific and may have been involved in the evolution of transposon elements Weilong Guo, Michael Zhang, Hong Wu Supplementary Figures Fig. S1-S16
The genetic code of gene regulatory elements
 The genetic code of gene regulatory elements Ivan Ovcharenko Computational Biology Branch National Center for Biotechnology Information National Institutes of Health October 23, 2008 Outline Gene deserts
The genetic code of gene regulatory elements Ivan Ovcharenko Computational Biology Branch National Center for Biotechnology Information National Institutes of Health October 23, 2008 Outline Gene deserts
CHAPTER 21 GENOMES AND THEIR EVOLUTION
 GENETICS DATE CHAPTER 21 GENOMES AND THEIR EVOLUTION COURSE 213 AP BIOLOGY 1 Comparisons of genomes provide information about the evolutionary history of genes and taxonomic groups Genomics - study of
GENETICS DATE CHAPTER 21 GENOMES AND THEIR EVOLUTION COURSE 213 AP BIOLOGY 1 Comparisons of genomes provide information about the evolutionary history of genes and taxonomic groups Genomics - study of
M1 - Biochemistry. Nucleic Acid Structure II/Transcription I
 M1 - Biochemistry Nucleic Acid Structure II/Transcription I PH Ratz, PhD (Resources: Lehninger et al., 5th ed., Chapters 8, 24 & 26) 1 Nucleic Acid Structure II/Transcription I Learning Objectives: 1.
M1 - Biochemistry Nucleic Acid Structure II/Transcription I PH Ratz, PhD (Resources: Lehninger et al., 5th ed., Chapters 8, 24 & 26) 1 Nucleic Acid Structure II/Transcription I Learning Objectives: 1.
BLASTing through the kingdom of life
 Information for teachers Description: In this activity, students copy unknown DNA sequences and use them to search GenBank, the database of nucleotide sequences at the National Center for Biotechnology
Information for teachers Description: In this activity, students copy unknown DNA sequences and use them to search GenBank, the database of nucleotide sequences at the National Center for Biotechnology
Differences between prokaryotes & eukaryotes. Gene function
 GENE REGULATION Differences between prokaryotes & eukaryotes Gene function Description of Prokaryotic Chromosome and E.coli Review Differences between Prokaryotic & Eukaryotic Chromosomes Four differences
GENE REGULATION Differences between prokaryotes & eukaryotes Gene function Description of Prokaryotic Chromosome and E.coli Review Differences between Prokaryotic & Eukaryotic Chromosomes Four differences
Figure 1. FasterDB SEARCH PAGE corresponding to human WNK1 gene. In the search page, gene searching, in the mouse or human genome, can be done: 1- By
 1 2 3 Figure 1. FasterD SERCH PGE corresponding to human WNK1 gene. In the search page, gene searching, in the mouse or human genome, can be done: 1- y keywords (ENSEML ID, HUGO gene name, synonyms or
1 2 3 Figure 1. FasterD SERCH PGE corresponding to human WNK1 gene. In the search page, gene searching, in the mouse or human genome, can be done: 1- y keywords (ENSEML ID, HUGO gene name, synonyms or
Transcription is the first stage of gene expression
 Transcription is the first stage of gene expression RNA synthesis is catalyzed by RNA polymerase, which pries the DNA strands apart and hooks together the RNA nucleotides The RNA is complementary to the
Transcription is the first stage of gene expression RNA synthesis is catalyzed by RNA polymerase, which pries the DNA strands apart and hooks together the RNA nucleotides The RNA is complementary to the
Enzyme that uses RNA as a template to synthesize a complementary DNA
 Biology 105: Introduction to Genetics PRACTICE FINAL EXAM 2006 Part I: Definitions Homology: Comparison of two or more protein or DNA sequence to ascertain similarities in sequences. If two genes have
Biology 105: Introduction to Genetics PRACTICE FINAL EXAM 2006 Part I: Definitions Homology: Comparison of two or more protein or DNA sequence to ascertain similarities in sequences. If two genes have
The Structure of Proteins The Structure of Proteins. How Proteins are Made: Genetic Transcription, Translation, and Regulation
 How Proteins are Made: Genetic, Translation, and Regulation PLAY The Structure of Proteins 14.1 The Structure of Proteins Proteins - polymer amino acids - monomers Linked together with peptide bonds A
How Proteins are Made: Genetic, Translation, and Regulation PLAY The Structure of Proteins 14.1 The Structure of Proteins Proteins - polymer amino acids - monomers Linked together with peptide bonds A
Mapping long-range promoter contacts in human cells with high-resolution capture Hi-C
 CORRECTION NOTICE Nat. Genet. 47, 598 606 (2015) Mapping long-range promoter contacts in human cells with high-resolution capture Hi-C Borbala Mifsud, Filipe Tavares-Cadete, Alice N Young, Robert Sugar,
CORRECTION NOTICE Nat. Genet. 47, 598 606 (2015) Mapping long-range promoter contacts in human cells with high-resolution capture Hi-C Borbala Mifsud, Filipe Tavares-Cadete, Alice N Young, Robert Sugar,
Genomic resources for canine cancer research. Jessica Alföldi (for Kerstin Lindblad-Toh) Broad Institute of MIT and Harvard
 Genomic resources for canine cancer research Jessica Alföldi (for Kerstin Lindblad-Toh) Broad Institute of MIT and Harvard Dog genome assembly CanFam 3.1 Comparison of most recent assembly versions Sequence
Genomic resources for canine cancer research Jessica Alföldi (for Kerstin Lindblad-Toh) Broad Institute of MIT and Harvard Dog genome assembly CanFam 3.1 Comparison of most recent assembly versions Sequence
FINDING THE PAIN GENE How do geneticists connect a specific gene with a specific phenotype?
 FINDING THE PAIN GENE How do geneticists connect a specific gene with a specific phenotype? 1 Linkage & Recombination HUH? What? Why? Who cares? How? Multiple choice question. Each colored line represents
FINDING THE PAIN GENE How do geneticists connect a specific gene with a specific phenotype? 1 Linkage & Recombination HUH? What? Why? Who cares? How? Multiple choice question. Each colored line represents
CS313 Exercise 1 Cover Page Fall 2017
 CS313 Exercise 1 Cover Page Fall 2017 Due by the start of class on Monday, September 18, 2017. Name(s): In the TIME column, please estimate the time you spent on the parts of this exercise. Please try
CS313 Exercise 1 Cover Page Fall 2017 Due by the start of class on Monday, September 18, 2017. Name(s): In the TIME column, please estimate the time you spent on the parts of this exercise. Please try
03-511/711 Computational Genomics and Molecular Biology, Fall
 03-511/711 Computational Genomics and Molecular Biology, Fall 2010 1 Study questions These study problems are intended to help you to review for the final exam. This is not an exhaustive list of the topics
03-511/711 Computational Genomics and Molecular Biology, Fall 2010 1 Study questions These study problems are intended to help you to review for the final exam. This is not an exhaustive list of the topics
Chapter 13. From DNA to Protein
 Chapter 13 From DNA to Protein Proteins All proteins consist of polypeptide chains A linear sequence of amino acids Each chain corresponds to the nucleotide base sequenceof a gene The Path From Genes to
Chapter 13 From DNA to Protein Proteins All proteins consist of polypeptide chains A linear sequence of amino acids Each chain corresponds to the nucleotide base sequenceof a gene The Path From Genes to
SUPPLEMENTARY INFORMATION
 doi:1.138/nature11233 Supplementary Figure S1 Sample Flowchart. The ENCODE transcriptome data are obtained from several cell lines which have been cultured in replicates. They were either left intact (whole
doi:1.138/nature11233 Supplementary Figure S1 Sample Flowchart. The ENCODE transcriptome data are obtained from several cell lines which have been cultured in replicates. They were either left intact (whole
T and B cell gene rearrangement October 17, Ram Savan
 T and B cell gene rearrangement October 17, 2016 Ram Savan savanram@uw.edu 441 Lecture #9 Slide 1 of 28 Three lectures on antigen receptors Part 1 (Last Friday): Structural features of the BCR and TCR
T and B cell gene rearrangement October 17, 2016 Ram Savan savanram@uw.edu 441 Lecture #9 Slide 1 of 28 Three lectures on antigen receptors Part 1 (Last Friday): Structural features of the BCR and TCR
Differential Gene Expression
 Biology 4361 Developmental Biology Differential Gene Expression September 28, 2006 Chromatin Structure ~140 bp ~60 bp Transcriptional Regulation: 1. Packing prevents access CH 3 2. Acetylation ( C O )
Biology 4361 Developmental Biology Differential Gene Expression September 28, 2006 Chromatin Structure ~140 bp ~60 bp Transcriptional Regulation: 1. Packing prevents access CH 3 2. Acetylation ( C O )
Key Area 1.3: Gene Expression
 Key Area 1.3: Gene Expression RNA There is a second type of nucleic acid in the cell, called RNA. RNA plays a vital role in the production of protein from the code in the DNA. What is gene expression?
Key Area 1.3: Gene Expression RNA There is a second type of nucleic acid in the cell, called RNA. RNA plays a vital role in the production of protein from the code in the DNA. What is gene expression?
What would this eye color phenomenon be called?
 Name: School: Total Score: / 50 1 1. Which nitrogenous bases present in DNA are purines, and which are pyrimidines? What is the main difference between a purine and a pyrimidine? (2 points) 2. To the right
Name: School: Total Score: / 50 1 1. Which nitrogenous bases present in DNA are purines, and which are pyrimidines? What is the main difference between a purine and a pyrimidine? (2 points) 2. To the right
Solutions will be posted on the web.
 MIT Biology Department 7.012: Introductory Biology - Fall 2004 Instructors: Professor Eric Lander, Professor Robert A. Weinberg, Dr. Claudette Gardel NAME TA SEC 7.012 Problem Set 7 FRIDAY December 3,
MIT Biology Department 7.012: Introductory Biology - Fall 2004 Instructors: Professor Eric Lander, Professor Robert A. Weinberg, Dr. Claudette Gardel NAME TA SEC 7.012 Problem Set 7 FRIDAY December 3,
Nature Genetics: doi: /ng Supplementary Figure 1. ChIP-seq genome browser views of BRM occupancy at previously identified BRM targets.
 Supplementary Figure 1 ChIP-seq genome browser views of BRM occupancy at previously identified BRM targets. Gene structures are shown underneath each panel. Supplementary Figure 2 pref6::ref6-gfp complements
Supplementary Figure 1 ChIP-seq genome browser views of BRM occupancy at previously identified BRM targets. Gene structures are shown underneath each panel. Supplementary Figure 2 pref6::ref6-gfp complements
Predicting Protein Structure and Examining Similarities of Protein Structure by Spectral Analysis Techniques
 Predicting Protein Structure and Examining Similarities of Protein Structure by Spectral Analysis Techniques, Melanie Abeysundera, Krista Collins, Chris Field Department of Mathematics and Statistics Dalhousie
Predicting Protein Structure and Examining Similarities of Protein Structure by Spectral Analysis Techniques, Melanie Abeysundera, Krista Collins, Chris Field Department of Mathematics and Statistics Dalhousie
Solutions will be posted on the web.
 MIT Biology Department 7.012: Introductory Biology - Fall 2004 Instructors: Professor Eric Lander, Professor Robert A. Weinberg, Dr. Claudette Gardel NAME TA SEC 7.012 Problem Set 7 FRIDAY December 3,
MIT Biology Department 7.012: Introductory Biology - Fall 2004 Instructors: Professor Eric Lander, Professor Robert A. Weinberg, Dr. Claudette Gardel NAME TA SEC 7.012 Problem Set 7 FRIDAY December 3,
DNA. Essential Question: How does the structure of the DNA molecule allow it to carry information?
 DNA Essential Question: How does the structure of the DNA molecule allow it to carry information? Fun Website to Explore! http://learn.genetics.utah.edu/content/molecules/ DNA History Griffith Experimented
DNA Essential Question: How does the structure of the DNA molecule allow it to carry information? Fun Website to Explore! http://learn.genetics.utah.edu/content/molecules/ DNA History Griffith Experimented
DNA & Protein Synthesis. The source and the process!
 DNA & Protein Synthesis The source and the process! Agenda I. DNA and Genes II. Protein Synthesis III. The Genetic Code I. DNA & Genes: The beauty of DNA Remember: DNA is a macromolecule that stores information
DNA & Protein Synthesis The source and the process! Agenda I. DNA and Genes II. Protein Synthesis III. The Genetic Code I. DNA & Genes: The beauty of DNA Remember: DNA is a macromolecule that stores information
DNA and Biotechnology Form of DNA Form of DNA Form of DNA Form of DNA Replication of DNA Replication of DNA
 21 DNA and Biotechnology DNA and Biotechnology OUTLINE: Replication of DNA Gene Expression Mutations Regulating Gene Activity Genetic Engineering Genomics DNA (deoxyribonucleic acid) Double-stranded molecule
21 DNA and Biotechnology DNA and Biotechnology OUTLINE: Replication of DNA Gene Expression Mutations Regulating Gene Activity Genetic Engineering Genomics DNA (deoxyribonucleic acid) Double-stranded molecule
Molecular Genetics. The flow of genetic information from DNA. DNA Replication. Two kinds of nucleic acids in cells: DNA and RNA.
 Molecular Genetics DNA Replication Two kinds of nucleic acids in cells: DNA and RNA. DNA function 1: DNA transmits genetic information from parents to offspring. DNA function 2: DNA controls the functions
Molecular Genetics DNA Replication Two kinds of nucleic acids in cells: DNA and RNA. DNA function 1: DNA transmits genetic information from parents to offspring. DNA function 2: DNA controls the functions
DNA. Using DNA to solve crimes
 DNA Using DNA to solve crimes Physical characteristics are inherited from both parents DNA contains all the inherited information for each person DNA is contained in the nucleus of every cell in your body
DNA Using DNA to solve crimes Physical characteristics are inherited from both parents DNA contains all the inherited information for each person DNA is contained in the nucleus of every cell in your body
SUPPLEMENTARY INFORMATION
 Figure S1. lev-9 mutants are resistant to levamisole. The levamisole dose-response curves indicate that lev-9 mutants are partially resistant to levamisole similar to lev-10(kr26) mutants. unc-29(x29)
Figure S1. lev-9 mutants are resistant to levamisole. The levamisole dose-response curves indicate that lev-9 mutants are partially resistant to levamisole similar to lev-10(kr26) mutants. unc-29(x29)
9/19/13. cdna libraries, EST clusters, gene prediction and functional annotation. Biosciences 741: Genomics Fall, 2013 Week 3
 cdna libraries, EST clusters, gene prediction and functional annotation Biosciences 741: Genomics Fall, 2013 Week 3 1 2 3 4 5 6 Figure 2.14 Relationship between gene structure, cdna, and EST sequences
cdna libraries, EST clusters, gene prediction and functional annotation Biosciences 741: Genomics Fall, 2013 Week 3 1 2 3 4 5 6 Figure 2.14 Relationship between gene structure, cdna, and EST sequences
Chromosomal Mutations. 2. Gene Mutations
 12-4 12-4 1. Chromosomal 3. NOT! 2. Gene A genetic mutation is any change in the DNA nucleotide sequence. Mutation is caused by mistakes during DNA replication, as well as mutagens, like certain chemicals
12-4 12-4 1. Chromosomal 3. NOT! 2. Gene A genetic mutation is any change in the DNA nucleotide sequence. Mutation is caused by mistakes during DNA replication, as well as mutagens, like certain chemicals
PrimePCR Assay Validation Report
 Gene Information Gene Name Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID STE20-related kinase adaptor alpha STRADA Human The protein encoded by this gene
Gene Information Gene Name Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID STE20-related kinase adaptor alpha STRADA Human The protein encoded by this gene
BIOLOGY - CLUTCH CH.17 - GENE EXPRESSION.
 !! www.clutchprep.com CONCEPT: GENES Beadle and Tatum develop the one gene one enzyme hypothesis through their work with Neurospora (bread mold). This idea was later revised as the one gene one polypeptide
!! www.clutchprep.com CONCEPT: GENES Beadle and Tatum develop the one gene one enzyme hypothesis through their work with Neurospora (bread mold). This idea was later revised as the one gene one polypeptide
Bio 101 Sample questions: Chapter 10
 Bio 101 Sample questions: Chapter 10 1. Which of the following is NOT needed for DNA replication? A. nucleotides B. ribosomes C. Enzymes (like polymerases) D. DNA E. all of the above are needed 2 The information
Bio 101 Sample questions: Chapter 10 1. Which of the following is NOT needed for DNA replication? A. nucleotides B. ribosomes C. Enzymes (like polymerases) D. DNA E. all of the above are needed 2 The information
Supplementary Figure 1
 Supplementary Figure 1 Advantages of RNA CaptureSeq for profiling genes with alternative splicing or low expression. (a) Schematic figure indicating limitations of qrt PCR for quantifying alternative splicing
Supplementary Figure 1 Advantages of RNA CaptureSeq for profiling genes with alternative splicing or low expression. (a) Schematic figure indicating limitations of qrt PCR for quantifying alternative splicing
The evolution of lncrna repertoires and expression patterns in tetrapods
 doi:10.1038/nature12943 The evolution of lncrna repertoires and expression patterns in tetrapods Anamaria Necsulea 1,2 {, Magali Soumillon 1,2 {, Maria Warnefors 1,2, Angélica Liechti 1,2, Tasman Daish
doi:10.1038/nature12943 The evolution of lncrna repertoires and expression patterns in tetrapods Anamaria Necsulea 1,2 {, Magali Soumillon 1,2 {, Maria Warnefors 1,2, Angélica Liechti 1,2, Tasman Daish
PrimePCR Assay Validation Report
 Gene Information Gene Name SRY (sex determining region Y)-box 6 Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID SOX6 Human This gene encodes a member of the
Gene Information Gene Name SRY (sex determining region Y)-box 6 Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID SOX6 Human This gene encodes a member of the
Genome-Wide Survey of MicroRNA - Transcription Factor Feed-Forward Regulatory Circuits in Human. Supporting Information
 Genome-Wide Survey of MicroRNA - Transcription Factor Feed-Forward Regulatory Circuits in Human Angela Re #, Davide Corá #, Daniela Taverna and Michele Caselle # equal contribution * corresponding author,
Genome-Wide Survey of MicroRNA - Transcription Factor Feed-Forward Regulatory Circuits in Human Angela Re #, Davide Corá #, Daniela Taverna and Michele Caselle # equal contribution * corresponding author,
Systematic evaluation of spliced alignment programs for RNA- seq data
 Systematic evaluation of spliced alignment programs for RNA- seq data Pär G. Engström, Tamara Steijger, Botond Sipos, Gregory R. Grant, André Kahles, RGASP Consortium, Gunnar Rätsch, Nick Goldman, Tim
Systematic evaluation of spliced alignment programs for RNA- seq data Pär G. Engström, Tamara Steijger, Botond Sipos, Gregory R. Grant, André Kahles, RGASP Consortium, Gunnar Rätsch, Nick Goldman, Tim
Selective constraints on noncoding DNA of mammals. Peter Keightley Institute of Evolutionary Biology University of Edinburgh
 Selective constraints on noncoding DNA of mammals Peter Keightley Institute of Evolutionary Biology University of Edinburgh Most mammalian noncoding DNA evolves rapidly Homo-Pan Divergence (%) 1.5 1.25
Selective constraints on noncoding DNA of mammals Peter Keightley Institute of Evolutionary Biology University of Edinburgh Most mammalian noncoding DNA evolves rapidly Homo-Pan Divergence (%) 1.5 1.25
Mate-pair library data improves genome assembly
 De Novo Sequencing on the Ion Torrent PGM APPLICATION NOTE Mate-pair library data improves genome assembly Highly accurate PGM data allows for de Novo Sequencing and Assembly For a draft assembly, generate
De Novo Sequencing on the Ion Torrent PGM APPLICATION NOTE Mate-pair library data improves genome assembly Highly accurate PGM data allows for de Novo Sequencing and Assembly For a draft assembly, generate
Introduction to Genome Biology
 Introduction to Genome Biology Sandrine Dudoit, Wolfgang Huber, Robert Gentleman Bioconductor Short Course 2006 Copyright 2006, all rights reserved Outline Cells, chromosomes, and cell division DNA structure
Introduction to Genome Biology Sandrine Dudoit, Wolfgang Huber, Robert Gentleman Bioconductor Short Course 2006 Copyright 2006, all rights reserved Outline Cells, chromosomes, and cell division DNA structure
