The Wnt-5a-derived hexapeptide Foxy-5 inhibits breast cancer. metastasis in vivo by targeting cell motility
|
|
|
- Randall Spencer
- 5 years ago
- Views:
Transcription
1 SUPPLEMENTARY MATERIAL: The Wnt-5a-derived hexapeptide Foxy-5 inhibits breast cancer metastasis in vivo by targeting cell motility Annette Säfholm, Johanna Tuomela, Jeanette Rosenkvist, Janna Dejmek, Pirkko Härkönen and Tommy Andersson* Supplementary Methods: Reverse transcription-polymerase chain reaction (RT-PCR) Total mrna from T47D, MDA-MB-468 and 4T1 cells was obtained using TRIzol Reagent according to the manufacturer's protocol. Isolated total RNA was reversed transcribed using Superscript II RNase H-reverse transcriptase (Invitrogen, Carlsbad, CA), per the manufacturer's instructions. To control for DNA contamination, the reverse transcription reaction was performed without the reverse transcriptase enzyme. The RT reaction products and 1μg of control genomic DNA extracted from the 4T1 cells was then subjected to PCR using Taq DNA polymerase (Invitrogen, Carlsbad, CA). The following primer sets were used for PCR: -actin F (5'-TTC AAC ACC CCA GCC ATG TA-3'), -actin R (5'-TTG CCA ATG GTG ATG ACC TG-3'), corresponding to nucleotides and of the human -actin sequence; human Ror2 F (5'-CCC 1
2 TGG TGC TTT ACG CAG AA-3') and human Ror2 R (5'-CAT GAA CCT CAC CGC AGA CA-3'), corresponding to nucleotides and of the human Ror2 sequence, mouse Ror2 F (5'-CCC TGC GAA CGT GTT CTT TT-3') and mouse Ror2 R (5'-GAA AAC GGC ACA CGG AAA AC-3'), corresponding to nucleotides and of the mouse Ror2 sequence. The resulting PCR products were separated by electrophoresis on 1.5% agarose gels and visualized by ethidium bromide staining. The gels shown (Supplementary Fig. 1) are representative of four independent experiments. Fluorometric assay for caspase-3 activity Caspase-3 activity was analyzed by fluorescence spectrometry using the fluorogenic peptide DEVD-amc as a substrate (Upstate Biotech. Inc., Lake Placid, NY). The 4T1 cells were allowed to adhere for 24h in 6-well plates. The cells were then incubated for 18h in the presence or absence of 100μM of the indicated peptide, 0.4μg/ml rwnt-5a or 100nM taxol (positive control). To complement the results from the Hoechst staining, both adherent and floating cells were lysed in caspase lysis buffer (10mM Tris-HCl, 10mM NaH 2 PO 4 /Na 2 HPO 4 [ph 7.5], 130mM NaCl, 1% Triton-X-100, and 10mM NaPP i ). 50μl lysates were added to the reaction wells, together with 200μl HEPES buffer (20mM HEPES [ph 7.5], 10% glycerol, and 2mM DTT) and 3μl of DEVD-amc (dissolved in DMSO). The reaction mixture was incubated at 37ºC for 1h, and thereafter analyzed for amc fluorescence in a FLUOstar plate reader (BMG Lab Technologies, Germany) at excitation and emission wavelengths of 390 and 460nm, respectively. The protein content of lysates was determined and adjusted for, using the Coomassie Plus Protein Assay. The 2
3 fluorescence data were corrected by subtracting background values and duplicates of each sample were used to calculate a mean value for each separate experiment, which was calculated as percent of control. The experiment was independently repeated 5 times, Supplementary Fig. 2A. Hoechst staining 4T1 cells were plated on cover glasses and incubated for 24h in the presence or absence of 100μM of the indicated peptide, 0.4μg/ml rwnt-5a or 100nM taxol (positive control). The cells were then fixed with 4% paraformaldehyde for 15 min, stained in the dark with Hoechst stain (Hoechst 34580, Molecular Probes, Eugene, OR) for 10min, washed with PBS and mounted with Dako Cytomation fluorescent mounting medium. The samples were examined and photographed in a Nikon Eclipse 800 microscope using a 60X oil immersion objective. 300 cells were counted from each experiment. The experiment was independently repeated 6 times, and the accumulated results are given in Supplementary Fig. 2C. Viable cell count We seeded 5,000 4T1 cells and allowed them to adhere in 96-well plates for 24h. The cells were incubated for 18h with serum-free RPMI medium in the presence or absence of 100μM of the indicated peptide or 0.4μg/ml rwnt-5a. The cells were subsequently detached with trypsin and the number of viable cells was determined by trypan blue exclusion. Triplicates of each sample were analyzed and a mean value was calculated. The experiment was repeated independently 6 times, Supplemental Fig. 2D. 3
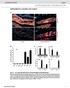
4 Thymidine incorporation An optimal cell density (data not shown) were seeded and allowed to adhere for 24h in 24-well plates. The cells were incubated for 18h with serum-free RPMI medium containing 0.25Ci tritium-labeled thymidine/well in the presence or absence of 100μM of the indicated peptide or 0.4g/ml rwnt-5a. Thereafter the cells were washed with PBS, incubated for 30min with 10% trichloroacetic acid and lysed with 1M NaOH for another 30min. The lysates were transferred to tubes containing scintillation liquid and the tritium activities were counted in a beta-counter. Scintillation liquid without lysate was used as a blank. Triplicates of each sample were analyzed and a mean value was calculated. The experiments were independently repeated 6 times, Supplementary Fig. 2E. As a control, the 4T1 cells were also incubated with 100M Foxy-5 for a week (tritium-labeled thymidine was included during the last 18h), and no difference was found compared to control cells, Supplementary Fig. 2F. Supplementary Figures: Supplementary Figure 1. The western blot to the left shows Ror2 protein expression whereas the Ror2 mrna expression is shown to the right in 4T1, MDA-MB-468 and T47D breast cancer cell-lines. DNA from 4T1 cells (DNA-4T1) was used as positive control for the mouse Ror2. -actin served as loading and PCR control. The experiments were repeated independently at least four times. 4
5 Supplementary Figure 2. In vitro restoration of Wnt-5a signaling has no effect on apoptosis or proliferation of 4T1 breast cancer cells. A) Caspase-3 activity in 4T1 cells incubated for 18h with 0.4μg/ml rwnt5a (W5a), 100μM of Foxy-5 (F5), the nonformylated control peptide (nfp) or the formylated control peptide (fcp). 100nM Taxol (Tax) was used as a positive control in Supplementary Fig. 2A-C. B) Representative Western blot to detect PARP cleavage in 4T1 cells incubated for 18h with either 100μM of the indicated peptides or 0.4μg/ml rwnt5a. The membrane was reprobed for -actin to confirm equal loading. C) Hoechst staining of 4T1 cells incubated with either 0.4μg/ml rwnt-5a or 100μM of the indicated peptides for 24h. D) 4T1 cells were incubated for 18h with 0.4μg/ml rwnt5a or 100μM of the indicated peptides, after which viable cells were counted by trypan blue exclusion. E) Thymidine incorporation in 4T1 cells stimulated with 0.4μg/ml rwnt-5a or 100μM of the described peptides for 18h. F) Thymidine incorporation in the 4T1 cells incubated for 1 week with 100μM F5 peptide. The results in are given as means ± SEM (n = 5-6). Supplementary Figure 3. The dose-response of Foxy-5 on lung metastasis in normal BALB/c mice. The mice were treated with PBS alone or with 5, 20, 40 or 160μg Foxy-5. The effects of Foxy-5 are expressed as percent of PBS treated controls of 17 mice treated with PBS alone, 9 mice treated with 5μg F5, 9 mice treated with 20μg F5, 6 mice treated with 40μg F5, and 7 mice treated with 160μg F5.. 5
6
7
8
Add 5µl of 3N NaOH to DNA sample (final concentration 0.3N NaOH).
 Bisulfite Treatment of DNA Dilute DNA sample to 2µg DNA in 50µl ddh 2 O. Add 5µl of 3N NaOH to DNA sample (final concentration 0.3N NaOH). Incubate in a 37ºC water bath for 30 minutes. To 55µl samples
Bisulfite Treatment of DNA Dilute DNA sample to 2µg DNA in 50µl ddh 2 O. Add 5µl of 3N NaOH to DNA sample (final concentration 0.3N NaOH). Incubate in a 37ºC water bath for 30 minutes. To 55µl samples
Supplemental Information. Human Senataxin Resolves RNA/DNA Hybrids. Formed at Transcriptional Pause Sites. to Promote Xrn2-Dependent Termination
 Supplemental Information Molecular Cell, Volume 42 Human Senataxin Resolves RNA/DNA Hybrids Formed at Transcriptional Pause Sites to Promote Xrn2-Dependent Termination Konstantina Skourti-Stathaki, Nicholas
Supplemental Information Molecular Cell, Volume 42 Human Senataxin Resolves RNA/DNA Hybrids Formed at Transcriptional Pause Sites to Promote Xrn2-Dependent Termination Konstantina Skourti-Stathaki, Nicholas
Supplementary Figures
 Supplementary Figures Supplementary Fig. 1 Characterization of GSCs. a. Immunostaining of primary GSC spheres from GSC lines. Nestin (neural progenitor marker, red), TLX (green). Merged images of nestin,
Supplementary Figures Supplementary Fig. 1 Characterization of GSCs. a. Immunostaining of primary GSC spheres from GSC lines. Nestin (neural progenitor marker, red), TLX (green). Merged images of nestin,
Electronic Supplementary Information
 Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2017 Electronic Supplementary Information Dissecting binding of a β-barrel outer membrane
Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2017 Electronic Supplementary Information Dissecting binding of a β-barrel outer membrane
Anti-Pim-1 (Cat#3247), anti-met (Cat#3127), anti-ron (Cat#2654), Anti-EGFR
 Supplementary Methods Antibodies Anti-Pim-1 (Cat#3247), anti-met (Cat#3127), anti-ron (Cat#2654), Anti-EGFR (Cat#2646), anti-igf1r (Cat#3018), anti-insr (Cat#3020), anti-akt (pan, Cat#4691), anti-phospho-akt
Supplementary Methods Antibodies Anti-Pim-1 (Cat#3247), anti-met (Cat#3127), anti-ron (Cat#2654), Anti-EGFR (Cat#2646), anti-igf1r (Cat#3018), anti-insr (Cat#3020), anti-akt (pan, Cat#4691), anti-phospho-akt
Supporting Information
 Supporting Information Transfection of DNA Cages into Mammalian Cells Email: a.turberfield@physics.ox.ac.uk Table of Contents Supporting Figure 1 DNA tetrahedra used in transfection experiments 2 Supporting
Supporting Information Transfection of DNA Cages into Mammalian Cells Email: a.turberfield@physics.ox.ac.uk Table of Contents Supporting Figure 1 DNA tetrahedra used in transfection experiments 2 Supporting
Supplementary. Table 1: Oligonucleotides and Plasmids. complementary to positions from 77 of the SRα '- GCT CTA GAG AAC TTG AAG TAC AGA CTG C
 Supplementary Table 1: Oligonucleotides and Plasmids 913954 5'- GCT CTA GAG AAC TTG AAG TAC AGA CTG C 913955 5'- CCC AAG CTT ACA GTG TGG CCA TTC TGC TG 223396 5'- CGA CGC GTA CAG TGT GGC CAT TCT GCT G
Supplementary Table 1: Oligonucleotides and Plasmids 913954 5'- GCT CTA GAG AAC TTG AAG TAC AGA CTG C 913955 5'- CCC AAG CTT ACA GTG TGG CCA TTC TGC TG 223396 5'- CGA CGC GTA CAG TGT GGC CAT TCT GCT G
Hes6. PPARα. PPARγ HNF4 CD36
 SUPPLEMENTARY INFORMATION Supplementary Table Positions and Sequences of ChIP primers -63 AGGTCACTGCCA -79 AGGTCTGCTGTG Hes6-0067 GGGCAaAGTTCA ACOT -395 GGGGCAgAGTTCA PPARα -309 GGCTCAaAGTTCAaGTTCA CPTa
SUPPLEMENTARY INFORMATION Supplementary Table Positions and Sequences of ChIP primers -63 AGGTCACTGCCA -79 AGGTCTGCTGTG Hes6-0067 GGGCAaAGTTCA ACOT -395 GGGGCAgAGTTCA PPARα -309 GGCTCAaAGTTCAaGTTCA CPTa
Supplement 1: Sequences of Capture Probes. Capture probes were /5AmMC6/CTG TAG GTG CGG GTG GAC GTA GTC
 Supplementary Appendixes Supplement 1: Sequences of Capture Probes. Capture probes were /5AmMC6/CTG TAG GTG CGG GTG GAC GTA GTC ACG TAG CTC CGG CTG GA-3 for vimentin, /5AmMC6/TCC CTC GCG CGT GGC TTC CGC
Supplementary Appendixes Supplement 1: Sequences of Capture Probes. Capture probes were /5AmMC6/CTG TAG GTG CGG GTG GAC GTA GTC ACG TAG CTC CGG CTG GA-3 for vimentin, /5AmMC6/TCC CTC GCG CGT GGC TTC CGC
Figure S1. Characterization of the irx9l-1 mutant. (A) Diagram of the Arabidopsis IRX9L gene drawn based on information from TAIR (the Arabidopsis
 1 2 3 4 5 6 7 8 9 10 11 12 Figure S1. Characterization of the irx9l-1 mutant. (A) Diagram of the Arabidopsis IRX9L gene drawn based on information from TAIR (the Arabidopsis Information Research). Exons
1 2 3 4 5 6 7 8 9 10 11 12 Figure S1. Characterization of the irx9l-1 mutant. (A) Diagram of the Arabidopsis IRX9L gene drawn based on information from TAIR (the Arabidopsis Information Research). Exons
PCR analysis was performed to show the presence and the integrity of the var1csa and var-
 Supplementary information: Methods: Table S1: Primer Name Nucleotide sequence (5-3 ) DBL3-F tcc ccg cgg agt gaa aca tca tgt gac tg DBL3-R gac tag ttt ctt tca ata aat cac tcg c DBL5-F cgc cct agg tgc ttc
Supplementary information: Methods: Table S1: Primer Name Nucleotide sequence (5-3 ) DBL3-F tcc ccg cgg agt gaa aca tca tgt gac tg DBL3-R gac tag ttt ctt tca ata aat cac tcg c DBL5-F cgc cct agg tgc ttc
B. ADM: C. D. Apoptosis: 1.68% 2.99% 1.31% Figure.S1,Li et al. number. invaded cells. HuH7 BxPC-3 DLD-1.
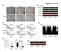 A. - Figure.S1,Li et al. B. : - + - + - + E-cadherin CK19 α-sma vimentin β -actin C. D. Apoptosis: 1.68% 2.99% 1.31% - : - + - + - + Apoptosis: 48.33% 45.32% 44.59% E. invaded cells number 400 300 200
A. - Figure.S1,Li et al. B. : - + - + - + E-cadherin CK19 α-sma vimentin β -actin C. D. Apoptosis: 1.68% 2.99% 1.31% - : - + - + - + Apoptosis: 48.33% 45.32% 44.59% E. invaded cells number 400 300 200
Supplementary Figure 1. Localization of MST1 in RPE cells. Proliferating or ciliated HA- MST1 expressing RPE cells (see Fig. 5b for establishment of
 Supplementary Figure 1. Localization of MST1 in RPE cells. Proliferating or ciliated HA- MST1 expressing RPE cells (see Fig. 5b for establishment of the cell line) were immunostained for HA, acetylated
Supplementary Figure 1. Localization of MST1 in RPE cells. Proliferating or ciliated HA- MST1 expressing RPE cells (see Fig. 5b for establishment of the cell line) were immunostained for HA, acetylated
An engineered tryptophan zipper-type peptide as a molecular recognition scaffold
 SUPPLEMENTARY MATERIAL An engineered tryptophan zipper-type peptide as a molecular recognition scaffold Zihao Cheng and Robert E. Campbell* Supplementary Methods Library construction for FRET-based screening
SUPPLEMENTARY MATERIAL An engineered tryptophan zipper-type peptide as a molecular recognition scaffold Zihao Cheng and Robert E. Campbell* Supplementary Methods Library construction for FRET-based screening
Y-chromosomal haplogroup typing Using SBE reaction
 Schematic of multiplex PCR followed by SBE reaction Multiplex PCR Exo SAP purification SBE reaction 5 A 3 ddatp ddgtp 3 T 5 A G 3 T 5 3 5 G C 5 3 3 C 5 ddttp ddctp 5 T 3 T C 3 A 5 3 A 5 5 C 3 3 G 5 3 G
Schematic of multiplex PCR followed by SBE reaction Multiplex PCR Exo SAP purification SBE reaction 5 A 3 ddatp ddgtp 3 T 5 A G 3 T 5 3 5 G C 5 3 3 C 5 ddttp ddctp 5 T 3 T C 3 A 5 3 A 5 5 C 3 3 G 5 3 G
Quantitative reverse-transcription PCR. Transcript levels of flgs, flgr, flia and flha were
 1 Supplemental methods 2 3 4 5 6 7 8 9 1 11 12 13 14 15 16 17 18 19 21 22 23 Quantitative reverse-transcription PCR. Transcript levels of flgs, flgr, flia and flha were monitored by quantitative reverse-transcription
1 Supplemental methods 2 3 4 5 6 7 8 9 1 11 12 13 14 15 16 17 18 19 21 22 23 Quantitative reverse-transcription PCR. Transcript levels of flgs, flgr, flia and flha were monitored by quantitative reverse-transcription
Supplemental Data Supplemental Figure 1.
 Supplemental Data Supplemental Figure 1. Silique arrangement in the wild-type, jhs, and complemented lines. Wild-type (WT) (A), the jhs1 mutant (B,C), and the jhs1 mutant complemented with JHS1 (Com) (D)
Supplemental Data Supplemental Figure 1. Silique arrangement in the wild-type, jhs, and complemented lines. Wild-type (WT) (A), the jhs1 mutant (B,C), and the jhs1 mutant complemented with JHS1 (Com) (D)
Cat. # Product Size DS130 DynaExpress TA PCR Cloning Kit (ptakn-2) 20 reactions Box 1 (-20 ) ptakn-2 Vector, linearized 20 µl (50 ng/µl) 1
 Product Name: Kit Component TA PCR Cloning Kit (ptakn-2) Cat. # Product Size DS130 TA PCR Cloning Kit (ptakn-2) 20 reactions Box 1 (-20 ) ptakn-2 Vector, linearized 20 µl (50 ng/µl) 1 2 Ligation Buffer
Product Name: Kit Component TA PCR Cloning Kit (ptakn-2) Cat. # Product Size DS130 TA PCR Cloning Kit (ptakn-2) 20 reactions Box 1 (-20 ) ptakn-2 Vector, linearized 20 µl (50 ng/µl) 1 2 Ligation Buffer
Supplementary Methods Quantitative RT-PCR. For mrna, total RNA was prepared using TRIzol reagent (Invitrogen) and genomic DNA was eliminated with TURB
 Supplementary Methods Quantitative RT-PCR. For mrna, total RNA was prepared using TRIzol reagent (Invitrogen) and genomic DNA was eliminated with TURBO DNA-free Kit (Ambion). One µg of total RNA was reverse
Supplementary Methods Quantitative RT-PCR. For mrna, total RNA was prepared using TRIzol reagent (Invitrogen) and genomic DNA was eliminated with TURBO DNA-free Kit (Ambion). One µg of total RNA was reverse
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature12119 SUPPLEMENTARY FIGURES AND LEGENDS pre-let-7a- 1 +14U pre-let-7a- 1 Ddx3x Dhx30 Dis3l2 Elavl1 Ggt5 Hnrnph 2 Osbpl5 Puf60 Rnpc3 Rpl7 Sf3b3 Sf3b4 Tia1 Triobp U2af1 U2af2 1 6 2 4 3
doi:10.1038/nature12119 SUPPLEMENTARY FIGURES AND LEGENDS pre-let-7a- 1 +14U pre-let-7a- 1 Ddx3x Dhx30 Dis3l2 Elavl1 Ggt5 Hnrnph 2 Osbpl5 Puf60 Rnpc3 Rpl7 Sf3b3 Sf3b4 Tia1 Triobp U2af1 U2af2 1 6 2 4 3
strain devoid of the aox1 gene [1]. Thus, the identification of AOX1 in the intracellular
![strain devoid of the aox1 gene [1]. Thus, the identification of AOX1 in the intracellular strain devoid of the aox1 gene [1]. Thus, the identification of AOX1 in the intracellular](/thumbs/72/66361212.jpg) Additional file 2 Identification of AOX1 in P. pastoris GS115 with a Mut s phenotype Results and Discussion The HBsAg producing strain was originally identified as a Mut s (methanol utilization slow) strain
Additional file 2 Identification of AOX1 in P. pastoris GS115 with a Mut s phenotype Results and Discussion The HBsAg producing strain was originally identified as a Mut s (methanol utilization slow) strain
PGRP negatively regulates NOD-mediated cytokine production in rainbow trout liver cells
 Supplementary Information for: PGRP negatively regulates NOD-mediated cytokine production in rainbow trout liver cells Ju Hye Jang 1, Hyun Kim 2, Mi Jung Jang 2, Ju Hyun Cho 1,2,* 1 Research Institute
Supplementary Information for: PGRP negatively regulates NOD-mediated cytokine production in rainbow trout liver cells Ju Hye Jang 1, Hyun Kim 2, Mi Jung Jang 2, Ju Hyun Cho 1,2,* 1 Research Institute
Supporting Information
 Supporting Information Table S1. Oligonucleotide sequences used in this work Oligo DNA A B C D CpG-A CpG-B CpG-C CpG-D Sequence 5 ACA TTC CTA AGT CTG AAA CAT TAC AGC TTG CTA CAC GAG AAG AGC CGC CAT AGT
Supporting Information Table S1. Oligonucleotide sequences used in this work Oligo DNA A B C D CpG-A CpG-B CpG-C CpG-D Sequence 5 ACA TTC CTA AGT CTG AAA CAT TAC AGC TTG CTA CAC GAG AAG AGC CGC CAT AGT
Supplementary Figure 1A A404 Cells +/- Retinoic Acid
 Supplementary Figure 1A A44 Cells +/- Retinoic Acid 1 1 H3 Lys4 di-methylation SM-actin VEC cfos (-) RA (+) RA 14 1 1 8 6 4 H3 Lys79 di-methylation SM-actin VEC cfos (-) RA (+) RA Supplementary Figure
Supplementary Figure 1A A44 Cells +/- Retinoic Acid 1 1 H3 Lys4 di-methylation SM-actin VEC cfos (-) RA (+) RA 14 1 1 8 6 4 H3 Lys79 di-methylation SM-actin VEC cfos (-) RA (+) RA Supplementary Figure
Lecture 10, 20/2/2002: The process of solution development - The CODEHOP strategy for automatic design of consensus-degenerate primers for PCR
 Lecture 10, 20/2/2002: The process of solution development - The CODEHOP strategy for automatic design of consensus-degenerate primers for PCR 1 The problem We wish to clone a yet unknown gene from a known
Lecture 10, 20/2/2002: The process of solution development - The CODEHOP strategy for automatic design of consensus-degenerate primers for PCR 1 The problem We wish to clone a yet unknown gene from a known
Supplemental Table 1. Mutant ADAMTS3 alleles detected in HEK293T clone 4C2. WT CCTGTCACTTTGGTTGATAGC MVLLSLWLIAAALVEVR
 Supplemental Dataset Supplemental Table 1. Mutant ADAMTS3 alleles detected in HEK293T clone 4C2. DNA sequence Amino acid sequence WT CCTGTCACTTTGGTTGATAGC MVLLSLWLIAAALVEVR Allele 1 CCTGTC------------------GATAGC
Supplemental Dataset Supplemental Table 1. Mutant ADAMTS3 alleles detected in HEK293T clone 4C2. DNA sequence Amino acid sequence WT CCTGTCACTTTGGTTGATAGC MVLLSLWLIAAALVEVR Allele 1 CCTGTC------------------GATAGC
Disease and selection in the human genome 3
 Disease and selection in the human genome 3 Ka/Ks revisited Please sit in row K or forward RBFD: human populations, adaptation and immunity Neandertal Museum, Mettman Germany Sequence genome Measure expression
Disease and selection in the human genome 3 Ka/Ks revisited Please sit in row K or forward RBFD: human populations, adaptation and immunity Neandertal Museum, Mettman Germany Sequence genome Measure expression
RPA-AB RPA-C Supplemental Figure S1: SDS-PAGE stained with Coomassie Blue after protein purification.
 RPA-AB RPA-C (a) (b) (c) (d) (e) (f) Supplemental Figure S: SDS-PAGE stained with Coomassie Blue after protein purification. (a) RPA; (b) RPA-AB; (c) RPA-CDE; (d) RPA-CDE core; (e) RPA-DE; and (f) RPA-C
RPA-AB RPA-C (a) (b) (c) (d) (e) (f) Supplemental Figure S: SDS-PAGE stained with Coomassie Blue after protein purification. (a) RPA; (b) RPA-AB; (c) RPA-CDE; (d) RPA-CDE core; (e) RPA-DE; and (f) RPA-C
FMF NIRCA PROTOCOL STEP 1.
 FMF NIRCA PROTOCOL STEP 1. After you have isolated patient s DNA and DNA from a healthy donor (wild type), you perform a nested PCR. The primers used to amplify exon 2 and exon 10 of the mefv gene are
FMF NIRCA PROTOCOL STEP 1. After you have isolated patient s DNA and DNA from a healthy donor (wild type), you perform a nested PCR. The primers used to amplify exon 2 and exon 10 of the mefv gene are
Electronic Supplementary Information
 Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2014 Electronic Supplementary Information Multiplexed Detection of Lung Cancer Cells at the Single-Molecule
Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2014 Electronic Supplementary Information Multiplexed Detection of Lung Cancer Cells at the Single-Molecule
Proteasome Activity Fluorometric Assay Kit II (Cat. # J4120)
 Proteasome Activity Fluorometric Assay Kit II (Cat. # J4120) Each supplied substrate is sufficient for use in 250 X 100 µl reactions to monitor the chymotrypsin-like (Suc-LLVY- AMC), trypsin-like (Boc-LRR-AMC)
Proteasome Activity Fluorometric Assay Kit II (Cat. # J4120) Each supplied substrate is sufficient for use in 250 X 100 µl reactions to monitor the chymotrypsin-like (Suc-LLVY- AMC), trypsin-like (Boc-LRR-AMC)
Nongenetic Reprogramming of the Ligand Specificity. of Growth Factor Receptors by Bispecific DNA Aptamers
 Supporting Information For Nongenetic Reprogramming of the Ligand Specificity of Growth Factor Receptors by Bispecific DNA Aptamers Ryosuke Ueki,* Saki Atsuta, Ayaka Ueki and Shinsuke Sando* Department
Supporting Information For Nongenetic Reprogramming of the Ligand Specificity of Growth Factor Receptors by Bispecific DNA Aptamers Ryosuke Ueki,* Saki Atsuta, Ayaka Ueki and Shinsuke Sando* Department
Overexpression Normal expression Overexpression Normal expression. 26 (21.1%) N (%) P-value a N (%)
 SUPPLEMENTARY TABLES Table S1. Alteration of ZNF322A protein expression levels in relation to clinicopathological parameters in 123 Asian and 74 Caucasian lung cancer patients. Asian patients Caucasian
SUPPLEMENTARY TABLES Table S1. Alteration of ZNF322A protein expression levels in relation to clinicopathological parameters in 123 Asian and 74 Caucasian lung cancer patients. Asian patients Caucasian
MacBlunt PCR Cloning Kit Manual
 MacBlunt PCR Cloning Kit Manual Shipping and Storage MacBlunt PCR Cloning Kits are shipped on dry ice. Each kit contains a box with cloning reagents and an attached bag with Eco-Blue Competent Cells (optional).
MacBlunt PCR Cloning Kit Manual Shipping and Storage MacBlunt PCR Cloning Kits are shipped on dry ice. Each kit contains a box with cloning reagents and an attached bag with Eco-Blue Competent Cells (optional).
RNA was isolated using NucleoSpin RNA II (Macherey-Nagel, Bethlehem, PA) according to the
 Supplementary Methods RT-PCR and real-time PCR analysis RNA was isolated using NucleoSpin RNA II (Macherey-Nagel, Bethlehem, PA) according to the manufacturer s protocol and quantified by measuring the
Supplementary Methods RT-PCR and real-time PCR analysis RNA was isolated using NucleoSpin RNA II (Macherey-Nagel, Bethlehem, PA) according to the manufacturer s protocol and quantified by measuring the
Materials Protein synthesis kit. This kit consists of 24 amino acids, 24 transfer RNAs, four messenger RNAs and one ribosome (see below).
 Protein Synthesis Instructions The purpose of today s lab is to: Understand how a cell manufactures proteins from amino acids, using information stored in the genetic code. Assemble models of four very
Protein Synthesis Instructions The purpose of today s lab is to: Understand how a cell manufactures proteins from amino acids, using information stored in the genetic code. Assemble models of four very
Arabidopsis actin depolymerizing factor AtADF4 mediates defense signal transduction triggered by the Pseudomonas syringae effector AvrPphB
 Arabidopsis actin depolymerizing factor mediates defense signal transduction triggered by the Pseudomonas syringae effector AvrPphB Files in this Data Supplement: Supplemental Table S1 Supplemental Table
Arabidopsis actin depolymerizing factor mediates defense signal transduction triggered by the Pseudomonas syringae effector AvrPphB Files in this Data Supplement: Supplemental Table S1 Supplemental Table
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb3240 Supplementary Figure 1 GBM cell lines display similar levels of p100 to p52 processing but respond differentially to TWEAK-induced TERT expression according to TERT promoter mutation
DOI: 10.1038/ncb3240 Supplementary Figure 1 GBM cell lines display similar levels of p100 to p52 processing but respond differentially to TWEAK-induced TERT expression according to TERT promoter mutation
CytoScan Fluoro Assay
 089PR G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name CytoScan Fluoro Assay (Cat. # 786 211) think proteins! think G-Biosciences www.gbiosciences.com
089PR G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name CytoScan Fluoro Assay (Cat. # 786 211) think proteins! think G-Biosciences www.gbiosciences.com
Luo et al. Supplemental Figures and Materials and Methods
 Luo et al. Supplemental Figures and Materials and Methods The supplemental figures demonstrate that nuclear NFAT is situated at PODs, overexpressed PML does not increase NFAT nuclear localization, and
Luo et al. Supplemental Figures and Materials and Methods The supplemental figures demonstrate that nuclear NFAT is situated at PODs, overexpressed PML does not increase NFAT nuclear localization, and
Nanotip for single-step preparation of DNA for qpcr analysis
 Supplementary information Nanotip for single-step preparation of DNA for qpcr analysis Dinesh Kalyanasundaram, a Jong-Hoon Kim, a Gareth Fotouhi, a Hyun-Boo Lee, a Morgan Hiraiwa, a Kieseok Oh, b Kyong-Hoon
Supplementary information Nanotip for single-step preparation of DNA for qpcr analysis Dinesh Kalyanasundaram, a Jong-Hoon Kim, a Gareth Fotouhi, a Hyun-Boo Lee, a Morgan Hiraiwa, a Kieseok Oh, b Kyong-Hoon
Supporting Information. Copyright Wiley-VCH Verlag GmbH & Co. KGaA, Weinheim, 2006
 Supporting Information Copyright Wiley-VCH Verlag GmbH & Co. KGaA, 69451 Weinheim, 2006 Copyright Wiley-VCH Verlag GmbH & Co. KGaA, 69451 Weinheim, 2006 Supporting Information for Expanding the Genetic
Supporting Information Copyright Wiley-VCH Verlag GmbH & Co. KGaA, 69451 Weinheim, 2006 Copyright Wiley-VCH Verlag GmbH & Co. KGaA, 69451 Weinheim, 2006 Supporting Information for Expanding the Genetic
Electronic Supplementary Information Sensitive detection of polynucleotide kinase using rolling circle amplification-induced chemiluminescence
 Electronic Supplementary Material (ESI) for Chemical Communications. This journal is The Royal Society of Chemistry 2014 Electronic Supplementary Information Sensitive detection of polynucleotide kinase
Electronic Supplementary Material (ESI) for Chemical Communications. This journal is The Royal Society of Chemistry 2014 Electronic Supplementary Information Sensitive detection of polynucleotide kinase
Wet Lab Tutorial: Genelet Circuits
 Wet Lab Tutorial: Genelet Circuits DNA 17 This tutorial will introduce the in vitro transcriptional circuits toolkit. The tutorial will focus on the design, synthesis, and experimental testing of a synthetic
Wet Lab Tutorial: Genelet Circuits DNA 17 This tutorial will introduce the in vitro transcriptional circuits toolkit. The tutorial will focus on the design, synthesis, and experimental testing of a synthetic
Supplementary Information: Materials and Methods. Immunoblot and immunoprecipitation. Cells were washed in phosphate buffered
 Supplementary Information: Materials and Methods Immunoblot and immunoprecipitation. Cells were washed in phosphate buffered saline (PBS) and lysed in TNN lysis buffer (50mM Tris at ph 8.0, 120mM NaCl
Supplementary Information: Materials and Methods Immunoblot and immunoprecipitation. Cells were washed in phosphate buffered saline (PBS) and lysed in TNN lysis buffer (50mM Tris at ph 8.0, 120mM NaCl
SUPPLEMENTARY INFORMATION
 doi: 10.1038/nature07182 SUPPLEMENTAL FIGURES AND TABLES Fig. S1. myf5-expressing cells give rise to brown fat depots and skeletal muscle (a) Perirenal BAT from control (cre negative) and myf5-cre:r26r3-yfp
doi: 10.1038/nature07182 SUPPLEMENTAL FIGURES AND TABLES Fig. S1. myf5-expressing cells give rise to brown fat depots and skeletal muscle (a) Perirenal BAT from control (cre negative) and myf5-cre:r26r3-yfp
hcd1tg/hj1tg/ ApoE-/- hcd1tg/hj1tg/ ApoE+/+
 ApoE+/+ ApoE-/- ApoE-/- H&E (1x) Supplementary Figure 1. No obvious pathology is observed in the colon of diseased ApoE-/me. Colon samples were fixed in 1% formalin and laid out in Swiss rolls for paraffin
ApoE+/+ ApoE-/- ApoE-/- H&E (1x) Supplementary Figure 1. No obvious pathology is observed in the colon of diseased ApoE-/me. Colon samples were fixed in 1% formalin and laid out in Swiss rolls for paraffin
Supplemental Materials and Methods
 Supplemental Materials and Methods Antibodies: Anti-SRF (cat# Sc-335) and anti-igf1r (sc-712) (Santa Cruz Biotech), and anti- ADAM-10 (14-6211) were from e-bioscience, anti-ku70 (cat# MS-329-P) (Labvision),
Supplemental Materials and Methods Antibodies: Anti-SRF (cat# Sc-335) and anti-igf1r (sc-712) (Santa Cruz Biotech), and anti- ADAM-10 (14-6211) were from e-bioscience, anti-ku70 (cat# MS-329-P) (Labvision),
Gene Expression Profiling Applicable for Cells Cultivated in Channel-Slides
 Gene Expression Profiling Applicable for Cells Cultivated in Channel-Slides Gene expression profiling provides information about the transcriptome of a living cell. The DNA is transcribed into mrna, when
Gene Expression Profiling Applicable for Cells Cultivated in Channel-Slides Gene expression profiling provides information about the transcriptome of a living cell. The DNA is transcribed into mrna, when
Cdc42 Activation Assay Kit
 A helping hand for your research Product Manual Configuration-specific Monoclonal Antibody Based Cdc42 Activation Assay Kit Catalog Number: 80701 20 assays 1 Table of Content Product Description 3 Assay
A helping hand for your research Product Manual Configuration-specific Monoclonal Antibody Based Cdc42 Activation Assay Kit Catalog Number: 80701 20 assays 1 Table of Content Product Description 3 Assay
Supplemental Material S1, Zhang et al. (2009)
 Supplemental Material S1, Zhang et al. (2009) Cloning of PS small RNA fragments. PS RNA labeled with [γ 32 P]-ATP (NEN) of a length from 30 to 90 nucleotides were isolated from the gel after denaturing
Supplemental Material S1, Zhang et al. (2009) Cloning of PS small RNA fragments. PS RNA labeled with [γ 32 P]-ATP (NEN) of a length from 30 to 90 nucleotides were isolated from the gel after denaturing
Supporting Online Information
 Supporting Online Information Isolation of Human Genomic DNA Sequences with Expanded Nucleobase Selectivity Preeti Rathi, Sara Maurer, Grzegorz Kubik and Daniel Summerer* Department of Chemistry and Chemical
Supporting Online Information Isolation of Human Genomic DNA Sequences with Expanded Nucleobase Selectivity Preeti Rathi, Sara Maurer, Grzegorz Kubik and Daniel Summerer* Department of Chemistry and Chemical
Supplementary Information
 Supplementary Information A general solution for opening double-stranded DNA for isothermal amplification Gangyi Chen, Juan Dong, Yi Yuan, Na Li, Xin Huang, Xin Cui* and Zhuo Tang* Supplementary Materials
Supplementary Information A general solution for opening double-stranded DNA for isothermal amplification Gangyi Chen, Juan Dong, Yi Yuan, Na Li, Xin Huang, Xin Cui* and Zhuo Tang* Supplementary Materials
ORFs and genes. Please sit in row K or forward
 ORFs and genes Please sit in row K or forward https://www.flickr.com/photos/teseum/3231682806/in/photostream/ Question: why do some strains of Vibrio cause cholera and others don t? Methods Mechanisms
ORFs and genes Please sit in row K or forward https://www.flickr.com/photos/teseum/3231682806/in/photostream/ Question: why do some strains of Vibrio cause cholera and others don t? Methods Mechanisms
ΔPDD1 x ΔPDD1. ΔPDD1 x wild type. 70 kd Pdd1. Pdd3
 Supplemental Fig. S1 ΔPDD1 x wild type ΔPDD1 x ΔPDD1 70 kd Pdd1 50 kd 37 kd Pdd3 Supplemental Fig. S1. ΔPDD1 strains express no detectable Pdd1 protein. Western blot analysis of whole-protein extracts
Supplemental Fig. S1 ΔPDD1 x wild type ΔPDD1 x ΔPDD1 70 kd Pdd1 50 kd 37 kd Pdd3 Supplemental Fig. S1. ΔPDD1 strains express no detectable Pdd1 protein. Western blot analysis of whole-protein extracts
Supporting Information
 Supporting Information Barderas et al. 10.1073/pnas.0801221105 SI Text: Docking of gastrin to Constructed scfv Models Interactive predocking of the 4-WL-5 motif into the central pocket observed in the
Supporting Information Barderas et al. 10.1073/pnas.0801221105 SI Text: Docking of gastrin to Constructed scfv Models Interactive predocking of the 4-WL-5 motif into the central pocket observed in the
Supporting information for Biochemistry, 1995, 34(34), , DOI: /bi00034a013
 Supporting information for Biochemistry, 1995, 34(34), 10807 10815, DOI: 10.1021/bi00034a013 LESNIK 10807-1081 Terms & Conditions Electronic Supporting Information files are available without a subscription
Supporting information for Biochemistry, 1995, 34(34), 10807 10815, DOI: 10.1021/bi00034a013 LESNIK 10807-1081 Terms & Conditions Electronic Supporting Information files are available without a subscription
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/10/494/eaan6284/dc1 Supplementary Materials for Activation of master virulence regulator PhoP in acidic ph requires the Salmonella-specific protein UgtL Jeongjoon
www.sciencesignaling.org/cgi/content/full/10/494/eaan6284/dc1 Supplementary Materials for Activation of master virulence regulator PhoP in acidic ph requires the Salmonella-specific protein UgtL Jeongjoon
Glutathione (GSH)-Decorated Magnetic Nanoparticles for Binding Glutathione-S-transferase (GST) Fusion Protein and Manipulating Live Cells
 Glutathione (GSH)-Decorated Magnetic Nanoparticles for Binding Glutathione-S-transferase (GST) Fusion Protein and Manipulating Live Cells Yue Pan, Marcus J. C. Long, Xinming Li, Junfeng Shi, Lizbeth Hedstrom,
Glutathione (GSH)-Decorated Magnetic Nanoparticles for Binding Glutathione-S-transferase (GST) Fusion Protein and Manipulating Live Cells Yue Pan, Marcus J. C. Long, Xinming Li, Junfeng Shi, Lizbeth Hedstrom,
SUPPLEMENTAL DATA SUPPLEMENTAL FIGURE LEGENDS
 SUPPLEMENTAL DATA SUPPLEMENTAL FIGURE LEGENDS SUPPLEMENTAL FIGURE S1. Identification of BmCREC. (A) Amino acid sequences of BmCREC show the peptides identified in LC-MS/MS analysis (marked by red letters
SUPPLEMENTAL DATA SUPPLEMENTAL FIGURE LEGENDS SUPPLEMENTAL FIGURE S1. Identification of BmCREC. (A) Amino acid sequences of BmCREC show the peptides identified in LC-MS/MS analysis (marked by red letters
SensoLyte Homogeneous AFC Caspase-3/7 Assay Kit
 SensoLyte Homogeneous AFC Caspase-3/7 Assay Kit Catalog # 71114 Unit Size Kit Size 1 Kit 500 Assays (96-well) or 1250 Assays (384-well) This kit is optimized to detect caspase-3/7 activity in cell culture
SensoLyte Homogeneous AFC Caspase-3/7 Assay Kit Catalog # 71114 Unit Size Kit Size 1 Kit 500 Assays (96-well) or 1250 Assays (384-well) This kit is optimized to detect caspase-3/7 activity in cell culture
A netlike rolling circle nucleic acid amplification technique
 Electronic Supplementary Material (ESI) for Analyst. This journal is The Royal Society of Chemistry 2014 Supplementary Information A netlike rolling circle nucleic acid amplification technique Xiaoli Zhu,
Electronic Supplementary Material (ESI) for Analyst. This journal is The Royal Society of Chemistry 2014 Supplementary Information A netlike rolling circle nucleic acid amplification technique Xiaoli Zhu,
Annex 6.5 CDC real-time rubella RT-PCR protocol targeting rubella p150 gene (154 nt region)
 Annex 6.5 CDC real-time rubella RT-PCR protocol targeting rubella p150 gene (154 nt region) NOTE: This document is intended to provide basic test method details and is not an SOP. Laboratories need to
Annex 6.5 CDC real-time rubella RT-PCR protocol targeting rubella p150 gene (154 nt region) NOTE: This document is intended to provide basic test method details and is not an SOP. Laboratories need to
SUPPORTING INFORMATION FILE
 Intrinsic and extrinsic connections of Tet3 dioxygenase with CXXC zinc finger modules Nan Liu, Mengxi Wang, Wen Deng, Christine S. Schmidt, Weihua Qin, Heinrich Leonhardt and Fabio Spada Department of
Intrinsic and extrinsic connections of Tet3 dioxygenase with CXXC zinc finger modules Nan Liu, Mengxi Wang, Wen Deng, Christine S. Schmidt, Weihua Qin, Heinrich Leonhardt and Fabio Spada Department of
Supplemental Data. mir156-regulated SPL Transcription. Factors Define an Endogenous Flowering. Pathway in Arabidopsis thaliana
 Cell, Volume 138 Supplemental Data mir156-regulated SPL Transcription Factors Define an Endogenous Flowering Pathway in Arabidopsis thaliana Jia-Wei Wang, Benjamin Czech, and Detlef Weigel Table S1. Interaction
Cell, Volume 138 Supplemental Data mir156-regulated SPL Transcription Factors Define an Endogenous Flowering Pathway in Arabidopsis thaliana Jia-Wei Wang, Benjamin Czech, and Detlef Weigel Table S1. Interaction
Dierks Supplementary Fig. S1
 Dierks Supplementary Fig. S1 ITK SYK PH TH K42R wt K42R (kinase deficient) R29C E42K Y323F R29C E42K Y323F (reduced phospholipid binding) (enhanced phospholipid binding) (reduced Cbl binding) E42K Y323F
Dierks Supplementary Fig. S1 ITK SYK PH TH K42R wt K42R (kinase deficient) R29C E42K Y323F R29C E42K Y323F (reduced phospholipid binding) (enhanced phospholipid binding) (reduced Cbl binding) E42K Y323F
A green method of staining DNA in polyacrylamide gel electrophoresis based on fluorescent copper nanoclusters synthesized in situ
 Electronic Supplementary Material A green method of staining DNA in polyacrylamide gel electrophoresis based on fluorescent copper nanoclusters synthesized in situ Xiaoli Zhu 1, Hai Shi 1, Yalan Shen 1,
Electronic Supplementary Material A green method of staining DNA in polyacrylamide gel electrophoresis based on fluorescent copper nanoclusters synthesized in situ Xiaoli Zhu 1, Hai Shi 1, Yalan Shen 1,
Transfection of CRISPR/Cas9 Nuclease NLS ribonucleoprotein (RNP) into adherent mammalian cells using Lipofectamine RNAiMAX
 Transfection of CRISPR/Cas9 Nuclease NLS ribonucleoprotein (RNP) into adherent mammalian cells using Lipofectamine RNAiMAX INTRODUCTION The CRISPR/Cas genome editing system consists of a single guide RNA
Transfection of CRISPR/Cas9 Nuclease NLS ribonucleoprotein (RNP) into adherent mammalian cells using Lipofectamine RNAiMAX INTRODUCTION The CRISPR/Cas genome editing system consists of a single guide RNA
Sexing Bovine Preimplantation Embryos by PCR
 Sexing Bovine Preimplantation Embryos by PCR Katherine E.M. Hendricks 1, Leydson F. Martins 2, Justin M. Fear 1 and Peter J. Hansen 1 1 Dept. of Animal Sciences, University of Florida and Department of
Sexing Bovine Preimplantation Embryos by PCR Katherine E.M. Hendricks 1, Leydson F. Martins 2, Justin M. Fear 1 and Peter J. Hansen 1 1 Dept. of Animal Sciences, University of Florida and Department of
Converting rabbit hybridoma into recombinant antibodies with effective transient production in an optimized human expression system
 Converting rabbit hybridoma into recombinant antibodies with effective transient production in an optimized human expression system Dr. Tim Welsink Molecular Biology Transient Gene Expression OUTLINE Short
Converting rabbit hybridoma into recombinant antibodies with effective transient production in an optimized human expression system Dr. Tim Welsink Molecular Biology Transient Gene Expression OUTLINE Short
Project 07/111 Final Report October 31, Project Title: Cloning and expression of porcine complement C3d for enhanced vaccines
 Project 07/111 Final Report October 31, 2007. Project Title: Cloning and expression of porcine complement C3d for enhanced vaccines Project Leader: Dr Douglas C. Hodgins (519-824-4120 Ex 54758, fax 519-824-5930)
Project 07/111 Final Report October 31, 2007. Project Title: Cloning and expression of porcine complement C3d for enhanced vaccines Project Leader: Dr Douglas C. Hodgins (519-824-4120 Ex 54758, fax 519-824-5930)
Gene Expression Profiling: RNA isolation, crna synthesis and gene expression profiling were
 Gene Expression Profiling: RNA isolation, crna synthesis and gene expression profiling were performed as described by Giordano et al 24. A Unigene cluster was assigned to each of the Affymetrix probe sets
Gene Expression Profiling: RNA isolation, crna synthesis and gene expression profiling were performed as described by Giordano et al 24. A Unigene cluster was assigned to each of the Affymetrix probe sets
SUPPLEMENTAL MATERIALS SIRTUIN 1 PROMOTES HYPEROXIA-INDUCED LUNG EPITHELIAL DEATH INDEPENDENT OF NRF2 ACTIVATION
 SUPPLEMENTAL MATERIALS SIRTUIN PROMOTES HYPEROXIA-INDUCED LUNG EPITHELIAL DEATH INDEPENDENT OF NRF ACTIVATION Haranatha R. Potteti*, Subbiah Rajasekaran*, Senthilkumar B. Rajamohan*, Chandramohan R. Tamatam,
SUPPLEMENTAL MATERIALS SIRTUIN PROMOTES HYPEROXIA-INDUCED LUNG EPITHELIAL DEATH INDEPENDENT OF NRF ACTIVATION Haranatha R. Potteti*, Subbiah Rajasekaran*, Senthilkumar B. Rajamohan*, Chandramohan R. Tamatam,
Supplementary Information (Ha, et. al) Supplementary Figures Supplementary Fig. S1
 Supplementary Information (Ha, et. al) Supplementary Figures Supplementary Fig. S1 a His-ORMDL3 ~ 17 His-ORMDL3 GST-ORMDL3 - + - + IPTG GST-ORMDL3 ~ b Integrated Density (ORMDL3/ -actin) 0.4 0.3 0.2 0.1
Supplementary Information (Ha, et. al) Supplementary Figures Supplementary Fig. S1 a His-ORMDL3 ~ 17 His-ORMDL3 GST-ORMDL3 - + - + IPTG GST-ORMDL3 ~ b Integrated Density (ORMDL3/ -actin) 0.4 0.3 0.2 0.1
Supporting Information
 Supporting Information Tal et al. 10.1073/pnas.0807694106 SI Materials and Methods VSV Infection and Quantification. Infection was carried out by seeding 5 10 5 MEF cells per well in a 6-well plate and
Supporting Information Tal et al. 10.1073/pnas.0807694106 SI Materials and Methods VSV Infection and Quantification. Infection was carried out by seeding 5 10 5 MEF cells per well in a 6-well plate and
DNA Tetrahedron-Based Molecular Beacon for Tumor-Related
 Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2015 Electronic Supporting Information (ESI) DNA Tetrahedron-Based Molecular Beacon for Tumor-Related
Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2015 Electronic Supporting Information (ESI) DNA Tetrahedron-Based Molecular Beacon for Tumor-Related
Emanuela Tumini, Sonia Barroso, Carmen Pérez Calero and Andrés Aguilera
 SUPPLEMENTARY INFORMATION Roles of human POLD1 and POLD3 in genome stability Emanuela Tumini, Sonia Barroso, Carmen Pérez Calero and Andrés Aguilera SUPPLEMENTARY METHODS Cell proliferation After sirna
SUPPLEMENTARY INFORMATION Roles of human POLD1 and POLD3 in genome stability Emanuela Tumini, Sonia Barroso, Carmen Pérez Calero and Andrés Aguilera SUPPLEMENTARY METHODS Cell proliferation After sirna
SOP for Quantitation of the Hepatitis C Virus (HCV) Genome by RT-PCR
 Virus Bank SOP-HCV-001 1. Scope SOP for Quantitation of the Hepatitis C Virus (HCV) Genome by RT-PCR 1.1 This procedure describes a method for quantitation of the HCV genome in HCV-infected cells by RT-PCR.
Virus Bank SOP-HCV-001 1. Scope SOP for Quantitation of the Hepatitis C Virus (HCV) Genome by RT-PCR 1.1 This procedure describes a method for quantitation of the HCV genome in HCV-infected cells by RT-PCR.
Supplementary Information. Construction of Lasso Peptide Fusion Proteins
 Supplementary Information Construction of Lasso Peptide Fusion Proteins Chuhan Zong 1, Mikhail O. Maksimov 2, A. James Link 2,3 * Departments of 1 Chemistry, 2 Chemical and Biological Engineering, and
Supplementary Information Construction of Lasso Peptide Fusion Proteins Chuhan Zong 1, Mikhail O. Maksimov 2, A. James Link 2,3 * Departments of 1 Chemistry, 2 Chemical and Biological Engineering, and
Flow cytometric determination of apoptosis by annexin V/propidium iodide double staining.
 Supplementary materials and methods Flow cytometric determination of apoptosis by annexin V/propidium iodide double staining. Cells were analyzed for phosphatidylserine exposure by an annexin-v FITC/propidium
Supplementary materials and methods Flow cytometric determination of apoptosis by annexin V/propidium iodide double staining. Cells were analyzed for phosphatidylserine exposure by an annexin-v FITC/propidium
RNA Blue REAGENT FOR RAPID ISOLATION OF PURE AND INTACT RNA (Cat. No. R011, R012, R013)
 RNA Blue REAGENT FOR RAPID ISOLATION OF PURE AND INTACT RNA (Cat. No. R011, R012, R013) WARNING: RNA Blue contains phenol and some other toxic components. After contact with skin, wash immediately with
RNA Blue REAGENT FOR RAPID ISOLATION OF PURE AND INTACT RNA (Cat. No. R011, R012, R013) WARNING: RNA Blue contains phenol and some other toxic components. After contact with skin, wash immediately with
Online Data Supplement
 Persistence of Rhinovirus Infection of Lower Airways in Patients with Bronchial Asthma Monika Wo, Marek Sanak, Jerzy Soja, Henryk Olechnowicz,William W. Busse, Andrew Szczeklik Online Data Supplement 39
Persistence of Rhinovirus Infection of Lower Airways in Patients with Bronchial Asthma Monika Wo, Marek Sanak, Jerzy Soja, Henryk Olechnowicz,William W. Busse, Andrew Szczeklik Online Data Supplement 39
Single cell imaging of Bruton's Tyrosine Kinase using an irreversible inhibitor
 SUPPLEMENTARY INFORMATION Single cell imaging of Bruton's Tyrosine Kinase using an irreversible inhibitor Anna Turetsky 1,a, Eunha Kim 1,a, Rainer H. Kohler 1, Miles A. Miller 1, Ralph Weissleder 1,2,
SUPPLEMENTARY INFORMATION Single cell imaging of Bruton's Tyrosine Kinase using an irreversible inhibitor Anna Turetsky 1,a, Eunha Kim 1,a, Rainer H. Kohler 1, Miles A. Miller 1, Ralph Weissleder 1,2,
Chapter 13 Chromatin Structure and its Effects on Transcription
 Chapter 13 Chromatin Structure and its Effects on Transcription Students must be positive that they understand standard PCR. There is a resource on the web for this purpose. Warn them before this class.
Chapter 13 Chromatin Structure and its Effects on Transcription Students must be positive that they understand standard PCR. There is a resource on the web for this purpose. Warn them before this class.
Gα i Activation Assay Kit
 A helping hand for your research Product Manual Configuration-specific Monoclonal Antibody Based Gα i Activation Assay Kit Catalog Number 80301 20 assays NewEast Biosciences, Inc 1 Table of Content Product
A helping hand for your research Product Manual Configuration-specific Monoclonal Antibody Based Gα i Activation Assay Kit Catalog Number 80301 20 assays NewEast Biosciences, Inc 1 Table of Content Product
Supplementary data. sienigma. F-Enigma F-EnigmaSM. a-p53
 Supplementary data Supplemental Figure 1 A sienigma #2 sienigma sicontrol a-enigma - + ++ - - - - - - + ++ - - - - - - ++ B sienigma F-Enigma F-EnigmaSM a-flag HLK3 cells - - - + ++ + ++ - + - + + - -
Supplementary data Supplemental Figure 1 A sienigma #2 sienigma sicontrol a-enigma - + ++ - - - - - - + ++ - - - - - - ++ B sienigma F-Enigma F-EnigmaSM a-flag HLK3 cells - - - + ++ + ++ - + - + + - -
Apoptosis assay: Apoptotic cells were identified by Annexin V-Alexa Fluor 488 and Propidium
 Apoptosis assay: Apoptotic cells were identified by Annexin V-Alexa Fluor 488 and Propidium Iodide (Invitrogen, Carlsbad, CA) staining. Briefly, 2x10 5 cells were washed once in cold PBS and resuspended
Apoptosis assay: Apoptotic cells were identified by Annexin V-Alexa Fluor 488 and Propidium Iodide (Invitrogen, Carlsbad, CA) staining. Briefly, 2x10 5 cells were washed once in cold PBS and resuspended
High-throughput cloning and expression in recalcitrant bacteria
 High-throughput cloning and expression in recalcitrant bacteria Eric R Geertsma & Bert Poolman Supplementary text and figures: Supplementary Figure 1 Frequency of SfiI sites yielding identical 3 extensions
High-throughput cloning and expression in recalcitrant bacteria Eric R Geertsma & Bert Poolman Supplementary text and figures: Supplementary Figure 1 Frequency of SfiI sites yielding identical 3 extensions
Gα 13 Activation Assay Kit
 A helping hand for your research Product Manual Configuration-specific Monoclonal Antibody Based Gα 13 Activation Assay Kit Catalog Number: 80401 20 assays NewEast Biosciences 1 Table of Content Product
A helping hand for your research Product Manual Configuration-specific Monoclonal Antibody Based Gα 13 Activation Assay Kit Catalog Number: 80401 20 assays NewEast Biosciences 1 Table of Content Product
Gene synthesis by circular assembly amplification
 Gene synthesis by circular assembly amplification Duhee Bang & George M Church Supplementary figures and text: Supplementary Figure 1. Dpo4 gene (1.05kb) construction by various methods. Supplementary
Gene synthesis by circular assembly amplification Duhee Bang & George M Church Supplementary figures and text: Supplementary Figure 1. Dpo4 gene (1.05kb) construction by various methods. Supplementary
HiYield TM Genomic DNA Extraction Kit Reagent
 HiYield TM Genomic DNA Extraction Kit Reagent CONTENTS Genomic DNA Extraction Kit Blood Cat.No. YGBE1KR // YGBE 100R Blood Protocol..... 3 Genomic DNA Extraction Kit Cultured Cells /Tissue / Bacterial
HiYield TM Genomic DNA Extraction Kit Reagent CONTENTS Genomic DNA Extraction Kit Blood Cat.No. YGBE1KR // YGBE 100R Blood Protocol..... 3 Genomic DNA Extraction Kit Cultured Cells /Tissue / Bacterial
Total Histone H3 Acetylation Detection Fast Kit (Fluorometric)
 Total Histone H3 Acetylation Detection Fast Kit (Fluorometric) Catalog Number KA1539 48 assays Version: 02 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Intended Use...
Total Histone H3 Acetylation Detection Fast Kit (Fluorometric) Catalog Number KA1539 48 assays Version: 02 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Intended Use...
MonsterScript Reverse Transcriptase
 MonsterScript Reverse Transcriptase Cat. Nos. MSTA5110, and MSTA5124 Connect with Epicentre on our blog (epicentral.blogspot.com), Facebook (facebook.com/epicentrebio), and Twitter (@EpicentreBio). www.epicentre.com
MonsterScript Reverse Transcriptase Cat. Nos. MSTA5110, and MSTA5124 Connect with Epicentre on our blog (epicentral.blogspot.com), Facebook (facebook.com/epicentrebio), and Twitter (@EpicentreBio). www.epicentre.com
Electronic Supplementary Information
 Electronic Supplementary Information An Affinity Capture Involved Enzymatic Assay for Thrombin by Using Peptide Aptamer as Affinity Ligands on Magnetic Beads Qiang Zhao a* Jie Gao b Research Center for
Electronic Supplementary Information An Affinity Capture Involved Enzymatic Assay for Thrombin by Using Peptide Aptamer as Affinity Ligands on Magnetic Beads Qiang Zhao a* Jie Gao b Research Center for
Functional Genomics Research Stream. Research Meeting: June 19, 2012 SYBR Green qpcr, Research Update
 Functional Genomics Research Stream Research Meeting: June 19, 2012 SYBR Green qpcr, Research Update Updates Alternate Lab Meeting Fridays 11:30-1:00 WEL 4.224 Welcome to attend either one Lab Log thanks
Functional Genomics Research Stream Research Meeting: June 19, 2012 SYBR Green qpcr, Research Update Updates Alternate Lab Meeting Fridays 11:30-1:00 WEL 4.224 Welcome to attend either one Lab Log thanks
MOLECULAR CLONING AND SEQUENCING OF FISH MPR 46
 MOLECULAR CLONING AND SEQUENCING OF FISH MPR 46 INTRODUCTION The mammalian MPR 46 (human, mouse, bovine) sequences show extensive homologies. Among the non-mammalian vertebrates, only a partial sequence
MOLECULAR CLONING AND SEQUENCING OF FISH MPR 46 INTRODUCTION The mammalian MPR 46 (human, mouse, bovine) sequences show extensive homologies. Among the non-mammalian vertebrates, only a partial sequence
HCV Genotype Primer Kit
 Instruction Manual for HCV Genotype Primer Kit HCV Genotype Determination Kit for Research Purpose Thoroughly read this instruction manual before use of this kit Background Study of nucleotide sequence
Instruction Manual for HCV Genotype Primer Kit HCV Genotype Determination Kit for Research Purpose Thoroughly read this instruction manual before use of this kit Background Study of nucleotide sequence
Fig. S1. Effect of p120-catenin overexpression on the interaction of SCUBE2 with E-cadherin. The expression plasmid encoding FLAG.
 Fig. S1. Effect of p120-catenin overexpression on the interaction of SCUBE2 with E-cadherin. The expression plasmid encoding FLAG.SCUBE2, E-cadherin.Myc, or HA.p120-catenin was transfected in a combination
Fig. S1. Effect of p120-catenin overexpression on the interaction of SCUBE2 with E-cadherin. The expression plasmid encoding FLAG.SCUBE2, E-cadherin.Myc, or HA.p120-catenin was transfected in a combination
Caspase-3 Assay Kit (Fluorometric)
 Caspase-3 Assay Kit (Fluorometric) Catalog Number KA0741 100 assays Version: 03 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Background... 3 Principle of the Assay...
Caspase-3 Assay Kit (Fluorometric) Catalog Number KA0741 100 assays Version: 03 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Background... 3 Principle of the Assay...
Sequencing of DNA lesions facilitated by site-specific excision via base. excision repair DNA glycosylases yielding ligatable gaps
 Supporting information Sequencing of DNA lesions facilitated by site-specific excision via base excision repair DNA glycosylases yielding ligatable gaps Jan Riedl, Aaron M. Fleming, and Cynthia J. Burrows*
Supporting information Sequencing of DNA lesions facilitated by site-specific excision via base excision repair DNA glycosylases yielding ligatable gaps Jan Riedl, Aaron M. Fleming, and Cynthia J. Burrows*
