Protocol S1: Supporting Information
|
|
|
- Margaret Perkins
- 6 years ago
- Views:
Transcription
1 Protocol S1: Supporting Information Basis for the specificity of the kinase domain of Abl for peptide substrates The crystal structures reported in this work were obtained using two different ATP analog-peptide conjugates that differed in the sequence of the peptide constituent (see Experimental Procedures and Figure S1). The analysis of the interactions between the peptides and the kinase is complicated by the fact that the protein adopts different conformations in the different crystal structures. However, in the structures solved using the sub-optimal peptide sequence (Structures 1 and 2, Table 1) the peptide is observed to bind in the same manner. We are therefore confident that the binding mode is an intrinsic property of the peptide sequence and not simply a consequence of the local conformation of the kinase. In all three cases, the central portion of the peptide flanking the tyrosine derivative is ordered and binds to the C-terminal lobe of the kinase. The overall mode of binding is the same in the three structures, with the formation of an anti-parallel β-sheet between the peptide and the C-terminal portion of the activation loop. This mode of 1
2 binding was first demonstrated for substrates of the receptor tyrosine kinase Irk [1]. The sequences of both peptides contain a glutamate at P-3 and an isoleucine at P-1. The glutamate residue forms an ion pair with Lys 404 in the C-terminal lobe of the kinase, while the isoleucine sidechain makes hydrophobic contacts with Trp 405 and Pro 402. There are clear differences between the binding of the optimal- and sub-optimal peptides (Figure S1). In the case of the optimal peptide, the sidechains of proline (P+3) and phenylalanine (P+4) fit snugly into a hydrophobic pocket in the C-lobe of the kinase. The peptide with the sub-optimal sequence follows a slightly different path in this region of the structure which positions the sidechain of phenylalanine (P+3) in the same pocket, burying considerably less hydrophobic surface area. This may account for the differences in the efficiencies with which the two peptides are phosphorylated by Abl [2]. The observation that the peptide backbone adjusts depending on the location of the phenylalanine sidechain (P+3 in the sub-optimal peptide and P+4 in the optimal peptide) suggests that hydrophobic interactions may dominate the binding of the substrate peptides. The relative paucity of sequence-specific interactions is consistent with experimental data reporting that Abl is a relatively promiscuous kinase. Given that these 2
3 two peptide-atp conjugates inhibit Abl with a very similar potency (0.95 µm and 1.2 µm for the optimal and sub-optimal conjugates, respectively), it is likely that the peptide interactions make only a minor energetic contribution to the affinity of these compounds. The variation in the path followed by the peptide backbone of the two different substrate peptides indicates that a change in substrate sequence may result in a somewhat altered mode of binding. This emphasizes that care must be taken in attributing kinase specificity to interactions observed in a single crystal structure. Presence of Nucleotide and Mg 2+ in targeted molecular dynamics simulations Nucleotide and Mg 2+ were not included in most simulations. Two simulations of transition 2 (Figure 1d) with ATP Mg 2+ or ADP Mg 2+ bound indicated that although the general features of the transitions are similar to those seen in simulations without nucleotide, increased forces are required to break the Asp-Mg 2+ bond. It is not clear whether the DFG flip occurs in the presence of bound nucleotide or in transient states when nucleotide is released, and our calculations do not address this issue. 3
4 Restraint sets and force constants in targeted molecular dynamics simulations Multiple simulations were carried out using restraint sets that included the entire kinase domain, the kinase domain except residues in the activation loop, the entire activation loop (residues ), five residues including the DFG motif ( 379 VADFG 383 ) or four residues including only the aspartate and phenylalanine residues of the DFG motif ( 379 VADF 382 ). The conformation of the activation loop is significantly different in the active, the Src-like inactive and in the imatinib-bound conformations (Figure 1). The inclusion of the entire activation loop in the restraint leads to collisions with the N-lobe of the kinase, causing unreasonably large structural excursions. Most likely, the activation loop has to first unfold, and then refold in both transitions 1 and 2 (Figure 1d), and our restraint strategy does not capture such a stepwise process. Restricting the restraint set to just the DFG motif and the adjacent residues allows the transition to occur with minimal adjustment to the rest of the structure (Figure S2a). These considerations lead us to focus most of our analysis on simulations in which the restraint force was applied to the four residue segment 379 VADF
5 We carried out simulations with force constants ranging from 10 kcal mol -1 Å -2 ( pn atom -1 Å -1 ) to 0.01 kcal mol -1 Å -2 (976 pn atom -1 Å -1 ). Using equation (1), we can estimate the extent to which the simulation structure can lag behind the driving force due to thermal energy by calculating the value of that corresponds to 1 k B T of energy. For 58 atoms in the restraint set (N=58), D DT is ~0.1 Å when k is 10 kcal mol -1 Å -2, and the transition proceeds smoothly because the simulation tracks the restraint closely. For a force constant of 0.15 kcal mol -1 Å -2, D DT is ~0.8 Å, and the transition is discontinuous because the simulated structure lags behind the target until the forces build up to the point where a transition occurs (Figure S2b). When the force constant is reduced below 0.15 kcal mol -1 Å -2, the transition no longer occurs in ps. The simulations with the lowest force constant for which DFG flips are seen to occur (0.15 kcal mol -1 Å -2 ) correspond to very high driving forces (Figure S2b). For example, in one such simulation the DFG motif does not begin to flip until the force builds up to approximately 2000 pn. This value of the force is much larger than the 5
6 experimentally measured force of ~160 pn required to pull biotin away from streptavidin, or the stall forces of the strongest known biological motors (50-65 pn) [3,4]. The large forces in the simulation are probably a consequence of the short trajectory lengths, which do not allow full relaxation of the rest of the protein structure. Further Analysis of Imatinib-Resistance Mutations in Abl In addition to the mutations discussed in the main text, two other imatinib resistance mutations appear to preferentially destabilize the Src-like inactive structure of Abl (molecule B). Mutation of Phe 359 to cysteine, valine or aspartate (F359C/V/D) has been seen in CML patients who have ceased to respond to imatinib [5], and the F359C mutation was also isolated in the in-vitro screen [6]. The C β atom of Phe 359 makes peripheral contact with imatinib in the drug complex (Figure S4a, center), and the remainder of the sidechain points out into solvent. In the active structure of Abl the phenyl ring of Phe 359 packs against Leu 387 in the activation loop (Figure S4a, left) such that mutations at this position might be expected to interfere with the active conformation and to sensitize the protein to imatinib rather than cause resistance. In 6
7 contrast, Phe 359 plays a more striking structural role in the Src-like inactive conformation of Abl. The sidechain makes extensive hydrophobic contacts in the rearranged interface between the activation loop and helix αc (Figure S4a, right). In addition, an amino-aromatic interaction occurs between the phenyl ring of Phe 359 and one of the sidechain amino groups of Arg 386. Tyrosine 253 is a residue in the phosphate binding P-loop that is mutated to either Phe or His in patients. The conformation of the P-loop is important for imatinib binding and several other resistance mutations map to this region. In the case of Tyr 253, calorimetric measurements indicate that imatinib binding is not affected by the mutations (M. Nidanie Henderson, personal communication). In the Src-like inactive structure Tyr 253 forms hydrogen bonds to Asp 363 and Arg 362, and mutation to Phe or His would disrupt these interactions (Figure S4b). 7
8 Kinetic characterization of peptide-atp conjugates The potency of the peptide-atp conjugates as inhibitors of Abl was assessed using a spectrophotometric kinase assay performed with the wild-type kinase domain of Abl[7]. Peptide and ATP substrates were present at 100 µm and 50 µm concentrations, respectively. By assuming that the peptide-atp conjugates are linear competitive inhibitors of both ATP and peptide substrates the IC 50 can be related to the K i by the following procedure: 1) IC 50 = K i,apparent (1 + A/K m (A)) 2) K i,apparent = K i (1 + B/K m (B)) where A and B are the concentrations of ATP and peptide substrates, respectively, used in the assay. The K m (A) and K m (B) values are the K m values for these substrates which were determined using the same spectrophotometric assay used for the IC 50 experiments. The K m values for peptide and ATP are 70 and 21 µm, respectively, under the conditions of the assay. The K i values for the Abl-optimal and sub-optimal peptide-atp conjugates (see Experimental Procedures and Figure S1) were 0.95 µm and 1.2 µm, respectively. 8
9 Supplementary Tables Table S1. Crystallographic Data and Refinement Table S2. Hydrogen bond distances between key residues Supplementary Figures Figure S1. Sequence-specific interactions between the ATP-peptide conjugates and the kinase domain of Abl. A) The structure of the peptide-atp conjugates. B) Interactions between the ATP-peptide conjugate with the optimal peptide sequence and Abl molecule D. C) Interactions between the ATP-peptide conjugate with the suboptimal peptide sequence and Abl molecule B. Figure S2. Choice of restraint set and force constant for TMD. A) When only the residues 379 VADF 382 are restrained, the deviation in the N-lobe is minimal. Grey: molecule B, starting structure; Yellow-orange: simulation final structure B) Restraint force (pn), and r.m.s. distance of the restraint from the target (Å), plotted versus time (picoseconds) for two low force constant (K=0.15 kcal mol -1 Å -2 ) simulations starting with the active structure. Figure S3. Targeted Molecular Dynamics simulations propose a path for DFG flipping. A) Coordination of Asp 381 during the DFG flip, starting from the Src-like inactive structure (molecule B) and following transition path B in backbone torsion angle space. The coordination of Asp 381 is almost identical for transition path A, except that the carbonyl of Asp 381 flips in the other direction B) The sidechain of Phe 382 passing through a hydrophobic pocket in the Src-like molecule for transition path B C) The paths of Asp 381 and Phe 382 in backbone torsion angle space for two trajectories. Figure S4. Molecule B helps explain mutations in the kinase domain of Abl that confer resistance to imatinib. For two of these mutations we have shown the surface for all atoms within 6 Å of the mutated residue in the context of different structures. The 9
10 active structure is shown in pink, the Abl:imatinib complex in green and the Src-like structure (molecule B) in yellow-orange. A) Phe 359 B) Tyr 253 Supplementary References 1. Hubbard SR (1997) Crystal structure of the activated insulin receptor tyrosine kinase in complex with peptide substrate and ATP analog. Embo J 16: Songyang Z, Carraway KL, 3rd, Eck MJ, Harrison SC, Feldman RA, et al. (1995) Catalytic specificity of protein-tyrosine kinases is critical for selective signalling. Nature 373: Isralewitz B, Gao M, Schulten K (2001) Steered molecular dynamics and mechanical functions of proteins. Curr Opin Struct Biol 11: Wang HY, Elston T, Mogilner A, Oster G (1998) Force generation in RNA polymerase. Biophys J 74: Shah NP, Nicoll JM, Nagar B, Gorre ME, Paquette RL, et al. (2002) Multiple BCR- ABL kinase domain mutations confer polyclonal resistance to the tyrosine kinase inhibitor imatinib (STI571) in chronic phase and blast crisis chronic myeloid leukemia. Cancer Cell 2: Azam M, Latek RR, Daley GQ (2003) Mechanisms of autoinhibition and STI- 571/imatinib resistance revealed by mutagenesis of BCR-ABL. Cell 112: Seeliger MA, Young M, Henderson MN, Pellicena P, King DS, et al. (2005) High yield bacterial expression of active c-abl and c-src tyrosine kinases. Protein Sci 14:
SUPPLEMENTAL FIGURE LEGENDS. Figure S1: Homology alignment of DDR2 amino acid sequence. Shown are
 SUPPLEMENTAL FIGURE LEGENDS Figure S1: Homology alignment of DDR2 amino acid sequence. Shown are the amino acid sequences of human DDR2, mouse DDR2 and the closest homologs in zebrafish and C. Elegans.
SUPPLEMENTAL FIGURE LEGENDS Figure S1: Homology alignment of DDR2 amino acid sequence. Shown are the amino acid sequences of human DDR2, mouse DDR2 and the closest homologs in zebrafish and C. Elegans.
Molecular design principles underlying β-strand swapping. in the adhesive dimerization of cadherins
 Supplementary information for: Molecular design principles underlying β-strand swapping in the adhesive dimerization of cadherins Jeremie Vendome 1,2,3,5, Shoshana Posy 1,2,3,5,6, Xiangshu Jin, 1,3 Fabiana
Supplementary information for: Molecular design principles underlying β-strand swapping in the adhesive dimerization of cadherins Jeremie Vendome 1,2,3,5, Shoshana Posy 1,2,3,5,6, Xiangshu Jin, 1,3 Fabiana
Suppl. Figure 1: RCC1 sequence and sequence alignments. (a) Amino acid
 Supplementary Figures Suppl. Figure 1: RCC1 sequence and sequence alignments. (a) Amino acid sequence of Drosophila RCC1. Same colors are for Figure 1 with sequence of β-wedge that interacts with Ran in
Supplementary Figures Suppl. Figure 1: RCC1 sequence and sequence alignments. (a) Amino acid sequence of Drosophila RCC1. Same colors are for Figure 1 with sequence of β-wedge that interacts with Ran in
Structure formation and association of biomolecules. Prof. Dr. Martin Zacharias Lehrstuhl für Molekulardynamik (T38) Technische Universität München
 Structure formation and association of biomolecules Prof. Dr. Martin Zacharias Lehrstuhl für Molekulardynamik (T38) Technische Universität München Motivation Many biomolecules are chemically synthesized
Structure formation and association of biomolecules Prof. Dr. Martin Zacharias Lehrstuhl für Molekulardynamik (T38) Technische Universität München Motivation Many biomolecules are chemically synthesized
Basic concepts of molecular biology
 Basic concepts of molecular biology Gabriella Trucco Email: gabriella.trucco@unimi.it Life The main actors in the chemistry of life are molecules called proteins nucleic acids Proteins: many different
Basic concepts of molecular biology Gabriella Trucco Email: gabriella.trucco@unimi.it Life The main actors in the chemistry of life are molecules called proteins nucleic acids Proteins: many different
Fundamentals of Protein Structure
 Outline Fundamentals of Protein Structure Yu (Julie) Chen and Thomas Funkhouser Princeton University CS597A, Fall 2005 Protein structure Primary Secondary Tertiary Quaternary Forces and factors Levels
Outline Fundamentals of Protein Structure Yu (Julie) Chen and Thomas Funkhouser Princeton University CS597A, Fall 2005 Protein structure Primary Secondary Tertiary Quaternary Forces and factors Levels
6-Foot Mini Toober Activity
 Big Idea The interaction between the substrate and enzyme is highly specific. Even a slight change in shape of either the substrate or the enzyme may alter the efficient and selective ability of the enzyme
Big Idea The interaction between the substrate and enzyme is highly specific. Even a slight change in shape of either the substrate or the enzyme may alter the efficient and selective ability of the enzyme
 Problem: The GC base pairs are more stable than AT base pairs. Why? 5. Triple-stranded DNA was first observed in 1957. Scientists later discovered that the formation of triplestranded DNA involves a type
Problem: The GC base pairs are more stable than AT base pairs. Why? 5. Triple-stranded DNA was first observed in 1957. Scientists later discovered that the formation of triplestranded DNA involves a type
Antibody-Antigen recognition. Structural Biology Weekend Seminar Annegret Kramer
 Antibody-Antigen recognition Structural Biology Weekend Seminar 10.07.2005 Annegret Kramer Contents Function and structure of antibodies Features of antibody-antigen interfaces Examples of antibody-antigen
Antibody-Antigen recognition Structural Biology Weekend Seminar 10.07.2005 Annegret Kramer Contents Function and structure of antibodies Features of antibody-antigen interfaces Examples of antibody-antigen
Tutorial. Visualize Variants on Protein Structure. Sample to Insight. November 21, 2017
 Visualize Variants on Protein Structure November 21, 2017 Sample to Insight QIAGEN Aarhus Silkeborgvej 2 Prismet 8000 Aarhus C Denmark Telephone: +45 70 22 32 44 www.qiagenbioinformatics.com AdvancedGenomicsSupport@qiagen.com
Visualize Variants on Protein Structure November 21, 2017 Sample to Insight QIAGEN Aarhus Silkeborgvej 2 Prismet 8000 Aarhus C Denmark Telephone: +45 70 22 32 44 www.qiagenbioinformatics.com AdvancedGenomicsSupport@qiagen.com
Supplementary Information for. Structure of human tyrosylprotein sulfotransferase-2 reveals the mechanism of protein tyrosine sulfation reaction
 Supplementary Information for Structure of human tyrosylprotein sulfotransferase-2 reveals the mechanism of protein tyrosine sulfation reaction Takamasa Teramoto, Yukari Fujikawa, Yoshirou Kawaguchi, Katsuhisa
Supplementary Information for Structure of human tyrosylprotein sulfotransferase-2 reveals the mechanism of protein tyrosine sulfation reaction Takamasa Teramoto, Yukari Fujikawa, Yoshirou Kawaguchi, Katsuhisa
1) The penicillin family of antibiotics, discovered by Alexander Fleming in 1928, has the following general structure: O O
 ame: TF ame: LS1a Fall 06 Problem Set #3 Due Friday 10/13 at noon in your TF s drop box on the 2 nd floor of the Science Center All questions including the (*extra*) ones should be turned in 1) The penicillin
ame: TF ame: LS1a Fall 06 Problem Set #3 Due Friday 10/13 at noon in your TF s drop box on the 2 nd floor of the Science Center All questions including the (*extra*) ones should be turned in 1) The penicillin
36. The double bonds in naturally-occuring fatty acids are usually isomers. A. cis B. trans C. both cis and trans D. D- E. L-
 36. The double bonds in naturally-occuring fatty acids are usually isomers. A. cis B. trans C. both cis and trans D. D- E. L- 37. The essential fatty acids are A. palmitic acid B. linoleic acid C. linolenic
36. The double bonds in naturally-occuring fatty acids are usually isomers. A. cis B. trans C. both cis and trans D. D- E. L- 37. The essential fatty acids are A. palmitic acid B. linoleic acid C. linolenic
Comparative Modeling Part 1. Jaroslaw Pillardy Computational Biology Service Unit Cornell Theory Center
 Comparative Modeling Part 1 Jaroslaw Pillardy Computational Biology Service Unit Cornell Theory Center Function is the most important feature of a protein Function is related to structure Structure is
Comparative Modeling Part 1 Jaroslaw Pillardy Computational Biology Service Unit Cornell Theory Center Function is the most important feature of a protein Function is related to structure Structure is
Supplementary Note 1. Enzymatic properties of the purified Syn BVR
 Supplementary Note 1. Enzymatic properties of the purified Syn BVR The expression vector pet15b-syn bvr allowed us to routinely prepare 15 mg of electrophoretically homogenous Syn BVR from 2.5 L of TB-medium
Supplementary Note 1. Enzymatic properties of the purified Syn BVR The expression vector pet15b-syn bvr allowed us to routinely prepare 15 mg of electrophoretically homogenous Syn BVR from 2.5 L of TB-medium
All Rights Reserved. U.S. Patents 6,471,520B1; 5,498,190; 5,916, North Market Street, Suite CC130A, Milwaukee, WI 53202
 Secondary Structure In the previous protein folding activity, you created a hypothetical 15-amino acid protein and learned that basic principles of chemistry determine how each protein spontaneously folds
Secondary Structure In the previous protein folding activity, you created a hypothetical 15-amino acid protein and learned that basic principles of chemistry determine how each protein spontaneously folds
Clamping down on pathogenic bacteria how to shut down a key DNA polymerase complex
 Clamping down on pathogenic bacteria how to shut down a key DNA polymerase complex Bacterial DNA-replication machinery Pathogenic bacteria that are resistant to the current armoury of antibiotics are an
Clamping down on pathogenic bacteria how to shut down a key DNA polymerase complex Bacterial DNA-replication machinery Pathogenic bacteria that are resistant to the current armoury of antibiotics are an
SUPPLEMENTARY INFORMATION
 doi: 10.1038/nature06147 SUPPLEMENTARY INFORMATION Figure S1 The genomic and domain structure of Dscam. The Dscam gene comprises 24 exons, encoding a signal peptide (SP), 10 IgSF domains, 6 fibronectin
doi: 10.1038/nature06147 SUPPLEMENTARY INFORMATION Figure S1 The genomic and domain structure of Dscam. The Dscam gene comprises 24 exons, encoding a signal peptide (SP), 10 IgSF domains, 6 fibronectin
Supplementary Fig. 1. Initial electron density maps for the NOX-D20:mC5a complex obtained after SAD-phasing. (a) Initial experimental electron
 Supplementary Fig. 1. Initial electron density maps for the NOX-D20:mC5a complex obtained after SAD-phasing. (a) Initial experimental electron density map obtained after SAD-phasing and density modification
Supplementary Fig. 1. Initial electron density maps for the NOX-D20:mC5a complex obtained after SAD-phasing. (a) Initial experimental electron density map obtained after SAD-phasing and density modification
Basic concepts of molecular biology
 Basic concepts of molecular biology Gabriella Trucco Email: gabriella.trucco@unimi.it What is life made of? 1665: Robert Hooke discovered that organisms are composed of individual compartments called cells
Basic concepts of molecular biology Gabriella Trucco Email: gabriella.trucco@unimi.it What is life made of? 1665: Robert Hooke discovered that organisms are composed of individual compartments called cells
SUPPLEMENTARY INFORMATION Figures. Supplementary Figure 1 a. Page 1 of 30. Nature Chemical Biology: doi: /nchembio.2528
 SUPPLEMENTARY INFORMATION Figures Supplementary Figure 1 a b c Page 1 of 0 11 Supplementary Figure 1: Biochemical characterisation and binding validation of the reversible USP inhibitor 1. a, Biochemical
SUPPLEMENTARY INFORMATION Figures Supplementary Figure 1 a b c Page 1 of 0 11 Supplementary Figure 1: Biochemical characterisation and binding validation of the reversible USP inhibitor 1. a, Biochemical
Solutions to Problem Set 1
 MIT Department of Biology 7.014 Introductory Biology, Spring 004 Question 1 Solutions to 7.014 Problem Set 1 a) Describe the conditions of the atmosphere on prebiotic earth and how these conditions differ
MIT Department of Biology 7.014 Introductory Biology, Spring 004 Question 1 Solutions to 7.014 Problem Set 1 a) Describe the conditions of the atmosphere on prebiotic earth and how these conditions differ
The Stringent Response
 The Stringent Response When amino acids are limiting a response is triggered to shut down a wide range of biosynthetic processes This process is called the Stringent Response It results in the synthesis
The Stringent Response When amino acids are limiting a response is triggered to shut down a wide range of biosynthetic processes This process is called the Stringent Response It results in the synthesis
5.36 Biochemistry Laboratory Spring 2009
 MIT OpenCourseWare http://ocw.mit.edu 5.36 Biochemistry Laboratory Spring 2009 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms. Laboratory Manual for URIECA
MIT OpenCourseWare http://ocw.mit.edu 5.36 Biochemistry Laboratory Spring 2009 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms. Laboratory Manual for URIECA
From mechanism to medicne
 From mechanism to medicne a look at proteins and drug design Chem 342 δ δ δ+ M 2009 δ+ δ+ δ M Drug Design - an Iterative Approach @ DSU Structural Analysis of Receptor Structural Analysis of Ligand-Receptor
From mechanism to medicne a look at proteins and drug design Chem 342 δ δ δ+ M 2009 δ+ δ+ δ M Drug Design - an Iterative Approach @ DSU Structural Analysis of Receptor Structural Analysis of Ligand-Receptor
Algorithms in Bioinformatics ONE Transcription Translation
 Algorithms in Bioinformatics ONE Transcription Translation Sami Khuri Department of Computer Science San José State University sami.khuri@sjsu.edu Biology Review DNA RNA Proteins Central Dogma Transcription
Algorithms in Bioinformatics ONE Transcription Translation Sami Khuri Department of Computer Science San José State University sami.khuri@sjsu.edu Biology Review DNA RNA Proteins Central Dogma Transcription
Protein-Ligand Analysis with SID
 Protein-Ligand Analysis with SID A new post-simulation analysis tool Dmitry Lupyan, PhD Understanding molecular processes with MD Drug discovery and design Protein-protein interactions + Protein-DNA interactions
Protein-Ligand Analysis with SID A new post-simulation analysis tool Dmitry Lupyan, PhD Understanding molecular processes with MD Drug discovery and design Protein-protein interactions + Protein-DNA interactions
Turning λ Cro into a
 Turning λ Cro into a 3 Transcriptional Activator Figure by MIT pencourseware. Fred Bushman and Mark Ptashne Cell (1988) 54:191-197 Presented by atalie Kuldell for 0.90 February 4th, 009 Small patch of
Turning λ Cro into a 3 Transcriptional Activator Figure by MIT pencourseware. Fred Bushman and Mark Ptashne Cell (1988) 54:191-197 Presented by atalie Kuldell for 0.90 February 4th, 009 Small patch of
Nucleic acid and protein Flow of genetic information
 Nucleic acid and protein Flow of genetic information References: Glick, BR and JJ Pasternak, 2003, Molecular Biotechnology: Principles and Applications of Recombinant DNA, ASM Press, Washington DC, pages.
Nucleic acid and protein Flow of genetic information References: Glick, BR and JJ Pasternak, 2003, Molecular Biotechnology: Principles and Applications of Recombinant DNA, ASM Press, Washington DC, pages.
Bioinformatics. ONE Introduction to Biology. Sami Khuri Department of Computer Science San José State University Biology/CS 123A Fall 2012
 Bioinformatics ONE Introduction to Biology Sami Khuri Department of Computer Science San José State University Biology/CS 123A Fall 2012 Biology Review DNA RNA Proteins Central Dogma Transcription Translation
Bioinformatics ONE Introduction to Biology Sami Khuri Department of Computer Science San José State University Biology/CS 123A Fall 2012 Biology Review DNA RNA Proteins Central Dogma Transcription Translation
STRUCTURAL BIOLOGY. α/β structures Closed barrels Open twisted sheets Horseshoe folds
 STRUCTURAL BIOLOGY α/β structures Closed barrels Open twisted sheets Horseshoe folds The α/β domains Most frequent domain structures are α/β domains: A central parallel or mixed β sheet Surrounded by α
STRUCTURAL BIOLOGY α/β structures Closed barrels Open twisted sheets Horseshoe folds The α/β domains Most frequent domain structures are α/β domains: A central parallel or mixed β sheet Surrounded by α
Not The Real Exam Just for Fun! Chemistry 391 Fall 2018
 Not The Real Exam Just for Fun! Chemistry 391 Fall 2018 Do not open the exam until ready to begin! Rules of the Game: This is a take-home Exam. The exam is due on Thursday, October 11 th at 9 AM. Otherwise,
Not The Real Exam Just for Fun! Chemistry 391 Fall 2018 Do not open the exam until ready to begin! Rules of the Game: This is a take-home Exam. The exam is due on Thursday, October 11 th at 9 AM. Otherwise,
RNA does not adopt the classic B-DNA helix conformation when it forms a self-complementary double helix
 Reason: RNA has ribose sugar ring, with a hydroxyl group (OH) If RNA in B-from conformation there would be unfavorable steric contact between the hydroxyl group, base, and phosphate backbone. RNA structure
Reason: RNA has ribose sugar ring, with a hydroxyl group (OH) If RNA in B-from conformation there would be unfavorable steric contact between the hydroxyl group, base, and phosphate backbone. RNA structure
Protein Folding Problem I400: Introduction to Bioinformatics
 Protein Folding Problem I400: Introduction to Bioinformatics November 29, 2004 Protein biomolecule, macromolecule more than 50% of the dry weight of cells is proteins polymer of amino acids connected into
Protein Folding Problem I400: Introduction to Bioinformatics November 29, 2004 Protein biomolecule, macromolecule more than 50% of the dry weight of cells is proteins polymer of amino acids connected into
SUPPLEMENTARY INFORMATION
 Supplementary Table 1. Crystallographic statistics CRM1-SNUPN complex Space group P6 4 22 a=b=250.4, c=190.4 Data collection statistics: CRM1-selenomethionine SNUPN MAD data Peak Inflection Remote Native
Supplementary Table 1. Crystallographic statistics CRM1-SNUPN complex Space group P6 4 22 a=b=250.4, c=190.4 Data collection statistics: CRM1-selenomethionine SNUPN MAD data Peak Inflection Remote Native
BIRKBECK COLLEGE (University of London)
 BIRKBECK COLLEGE (University of London) SCHOOL OF BIOLOGICAL SCIENCES M.Sc. EXAMINATION FOR INTERNAL STUDENTS ON: Postgraduate Certificate in Principles of Protein Structure MSc Structural Molecular Biology
BIRKBECK COLLEGE (University of London) SCHOOL OF BIOLOGICAL SCIENCES M.Sc. EXAMINATION FOR INTERNAL STUDENTS ON: Postgraduate Certificate in Principles of Protein Structure MSc Structural Molecular Biology
Protein Structure Prediction by Constraint Logic Programming
 MPRI C2-19 Protein Structure Prediction by Constraint Logic Programming François Fages, Constraint Programming Group, INRIA Rocquencourt mailto:francois.fages@inria.fr http://contraintes.inria.fr/ Molecules
MPRI C2-19 Protein Structure Prediction by Constraint Logic Programming François Fages, Constraint Programming Group, INRIA Rocquencourt mailto:francois.fages@inria.fr http://contraintes.inria.fr/ Molecules
HEK293T. Fig. 1 in the
 Supplementary Information Supplementary Figure 1 Zinc uptake assay of hzip4 and hzip4-δecd transiently expressed in HEK293T cells. The results of one representative e experiment are shown in Fig. 1 in
Supplementary Information Supplementary Figure 1 Zinc uptake assay of hzip4 and hzip4-δecd transiently expressed in HEK293T cells. The results of one representative e experiment are shown in Fig. 1 in
Solutions to 7.02 Quiz II 10/27/05
 Solutions to 7.02 Quiz II 10/27/05 Class Average = 83 Standard Deviation = 9 Range Grade % 87-100 A 43 74-86 B 39 55-73 C 17 > 54 D 1 Question 1 (56 points) While studying deep sea bacteria, you discover
Solutions to 7.02 Quiz II 10/27/05 Class Average = 83 Standard Deviation = 9 Range Grade % 87-100 A 43 74-86 B 39 55-73 C 17 > 54 D 1 Question 1 (56 points) While studying deep sea bacteria, you discover
human Cdc45 Figure 1c. (c)
 1 Details of the refined crystallographic model of human Cdc45 and comparison of its active-site region with that of bacterial RecJ. (a) Stereo view of a representative example of the final 2F o -F c electron
1 Details of the refined crystallographic model of human Cdc45 and comparison of its active-site region with that of bacterial RecJ. (a) Stereo view of a representative example of the final 2F o -F c electron
2018 Protein Modeling Exam Key
 2018 Protein Modeling Exam Key Multiple Choice: 1. Which of the following amino acids has a negative charge at ph 7? a. Gln b. Glu c. Ser d. Cys 2. Which of the following is an example of secondary structure?
2018 Protein Modeling Exam Key Multiple Choice: 1. Which of the following amino acids has a negative charge at ph 7? a. Gln b. Glu c. Ser d. Cys 2. Which of the following is an example of secondary structure?
Programme Good morning and summary of last week Levels of Protein Structure - I Levels of Protein Structure - II
 Programme 8.00-8.10 Good morning and summary of last week 8.10-8.30 Levels of Protein Structure - I 8.30-9.00 Levels of Protein Structure - II 9.00-9.15 Break 9.15-11.15 Exercise: Building a protein model
Programme 8.00-8.10 Good morning and summary of last week 8.10-8.30 Levels of Protein Structure - I 8.30-9.00 Levels of Protein Structure - II 9.00-9.15 Break 9.15-11.15 Exercise: Building a protein model
BMB/Bi/Ch 170 Fall 2017 Problem Set 1: Proteins I
 BMB/Bi/Ch 170 Fall 2017 Problem Set 1: Proteins I Please use ray-tracing feature for all the images you are submitting. Use either the Ray button on the right side of the command window in PyMOL or variations
BMB/Bi/Ch 170 Fall 2017 Problem Set 1: Proteins I Please use ray-tracing feature for all the images you are submitting. Use either the Ray button on the right side of the command window in PyMOL or variations
Structural Bioinformatics (C3210) DNA and RNA Structure
 Structural Bioinformatics (C3210) DNA and RNA Structure Importance of DNA/RNA 3D Structure Nucleic acids are essential materials found in all living organisms. Their main function is to maintain and transmit
Structural Bioinformatics (C3210) DNA and RNA Structure Importance of DNA/RNA 3D Structure Nucleic acids are essential materials found in all living organisms. Their main function is to maintain and transmit
Dr. R. Sankar, BSE 631 (2018)
 Pauling, Corey and Branson Diffraction of DNA http://www.nature.com/scitable/topicpage/dna-is-a-structure-that-encodes-biological-6493050 In short, stereochemistry is important in determining which helices
Pauling, Corey and Branson Diffraction of DNA http://www.nature.com/scitable/topicpage/dna-is-a-structure-that-encodes-biological-6493050 In short, stereochemistry is important in determining which helices
Introduction to Proteins
 Introduction to Proteins Lecture 4 Module I: Molecular Structure & Metabolism Molecular Cell Biology Core Course (GSND5200) Matthew Neiditch - Room E450U ICPH matthew.neiditch@umdnj.edu What is a protein?
Introduction to Proteins Lecture 4 Module I: Molecular Structure & Metabolism Molecular Cell Biology Core Course (GSND5200) Matthew Neiditch - Room E450U ICPH matthew.neiditch@umdnj.edu What is a protein?
Zool 3200: Cell Biology Exam 3 3/6/15
 Name: Trask Zool 3200: Cell Biology Exam 3 3/6/15 Answer each of the following questions in the space provided; circle the correct answer or answers for each multiple choice question and circle either
Name: Trask Zool 3200: Cell Biology Exam 3 3/6/15 Answer each of the following questions in the space provided; circle the correct answer or answers for each multiple choice question and circle either
Supplementary Information. Single-molecule analysis reveals multi-state folding of a guanine. riboswitch
 Supplementary Information Single-molecule analysis reveals multi-state folding of a guanine riboswitch Vishnu Chandra 1,4,#, Zain Hannan 1,5,#, Huizhong Xu 2,# and Maumita Mandal 1,2,3,6* Department of
Supplementary Information Single-molecule analysis reveals multi-state folding of a guanine riboswitch Vishnu Chandra 1,4,#, Zain Hannan 1,5,#, Huizhong Xu 2,# and Maumita Mandal 1,2,3,6* Department of
Drug DNA interaction. Modeling DNA ligand interaction of intercalating ligands
 Drug DNA interaction DNA as carrier of genetic information is a major target for drug interaction because of the ability to interfere with transcription (gene expression and protein synthesis) and DNA
Drug DNA interaction DNA as carrier of genetic information is a major target for drug interaction because of the ability to interfere with transcription (gene expression and protein synthesis) and DNA
The mechanism(s) of protein folding. What is meant by mechanism. Experimental approaches
 The mechanism(s) of protein folding What is meant by mechanism Computational approaches Experimental approaches Questions: What events occur and in what time sequence when a protein folds Is there a specified
The mechanism(s) of protein folding What is meant by mechanism Computational approaches Experimental approaches Questions: What events occur and in what time sequence when a protein folds Is there a specified
Chapter 14 Regulation of Transcription
 Chapter 14 Regulation of Transcription Cis-acting sequences Distance-independent cis-acting elements Dissecting regulatory elements Transcription factors Overview transcriptional regulation Transcription
Chapter 14 Regulation of Transcription Cis-acting sequences Distance-independent cis-acting elements Dissecting regulatory elements Transcription factors Overview transcriptional regulation Transcription
Protein-Protein Interactions I
 Biochemistry 412 Protein-Protein Interactions I March 11, 2008 Macromolecular Recognition by Proteins Protein folding is a process governed by intramolecular recognition. Protein-protein association is
Biochemistry 412 Protein-Protein Interactions I March 11, 2008 Macromolecular Recognition by Proteins Protein folding is a process governed by intramolecular recognition. Protein-protein association is
Structure and Function of the First Full-Length Murein Peptide Ligase (Mpl) Cell Wall Recycling Protein
 Paper Presentation PLoS ONE 2011 Structure and Function of the First Full-Length Murein Peptide Ligase (Mpl) Cell Wall Recycling Protein Debanu Das, Mireille Herve, Julie Feuerhelm, etc. and Dominique
Paper Presentation PLoS ONE 2011 Structure and Function of the First Full-Length Murein Peptide Ligase (Mpl) Cell Wall Recycling Protein Debanu Das, Mireille Herve, Julie Feuerhelm, etc. and Dominique
Supplementary Table 1: List of CH3 domain interface residues in the first chain (A) and
 Supplementary Tables Supplementary Table 1: List of CH3 domain interface residues in the first chain (A) and their side chain contacting residues in the second chain (B) a Interface Res. in Contacting
Supplementary Tables Supplementary Table 1: List of CH3 domain interface residues in the first chain (A) and their side chain contacting residues in the second chain (B) a Interface Res. in Contacting
X-ray structures of fructosyl peptide oxidases revealing residues responsible for gating oxygen access in the oxidative half reaction
 X-ray structures of fructosyl peptide oxidases revealing residues responsible for gating oxygen access in the oxidative half reaction Tomohisa Shimasaki 1, Hiromi Yoshida 2, Shigehiro Kamitori 2 & Koji
X-ray structures of fructosyl peptide oxidases revealing residues responsible for gating oxygen access in the oxidative half reaction Tomohisa Shimasaki 1, Hiromi Yoshida 2, Shigehiro Kamitori 2 & Koji
Amino Acids and Proteins
 Various Functions of Proteins SB203 Amino Acids and Proteins Jirundon Yuvaniyama, Ph.D. Department of Biochemistry Faculty of Science Mahidol University Enzymes Transport proteins utrient and storage proteins
Various Functions of Proteins SB203 Amino Acids and Proteins Jirundon Yuvaniyama, Ph.D. Department of Biochemistry Faculty of Science Mahidol University Enzymes Transport proteins utrient and storage proteins
Τάσος Οικονόµου ιαλεξη 8. Kινηση, λειτουργια, ελεγχος.
 Τάσος Οικονόµου ιαλεξη 8 Kινηση, λειτουργια, ελεγχος http://ecoserver.imbb.forth.gr/bio321.htm εν ξεχνω. Cell The peptide bond Polypeptides are stabilized by: 1. Covalent bonds= amide bond 2. Noncovalent,
Τάσος Οικονόµου ιαλεξη 8 Kινηση, λειτουργια, ελεγχος http://ecoserver.imbb.forth.gr/bio321.htm εν ξεχνω. Cell The peptide bond Polypeptides are stabilized by: 1. Covalent bonds= amide bond 2. Noncovalent,
11 questions for a total of 120 points
 Your Name: BYS 201, Final Exam, May 3, 2010 11 questions for a total of 120 points 1. 25 points Take a close look at these tables of amino acids. Some of them are hydrophilic, some hydrophobic, some positive
Your Name: BYS 201, Final Exam, May 3, 2010 11 questions for a total of 120 points 1. 25 points Take a close look at these tables of amino acids. Some of them are hydrophilic, some hydrophobic, some positive
Examination I PHRM 836 Biochemistry for Pharmaceutical Sciences II September 29, 2015
 Examination I PHRM 836 Biochemistry for Pharmaceutical Sciences II September 29, 2015 PHRM 836 Exam I - 1 Name: Instructions 1. Check your exam to make certain that it has 9 pages including this cover
Examination I PHRM 836 Biochemistry for Pharmaceutical Sciences II September 29, 2015 PHRM 836 Exam I - 1 Name: Instructions 1. Check your exam to make certain that it has 9 pages including this cover
Structural Bioinformatics (C3210) Conformational Analysis Protein Folding Protein Structure Prediction
 Structural Bioinformatics (C3210) Conformational Analysis Protein Folding Protein Structure Prediction Conformational Analysis 2 Conformational Analysis Properties of molecules depend on their three-dimensional
Structural Bioinformatics (C3210) Conformational Analysis Protein Folding Protein Structure Prediction Conformational Analysis 2 Conformational Analysis Properties of molecules depend on their three-dimensional
Hmwk # 8 : DNA-Binding Proteins : Part II
 The purpose of this exercise is : Hmwk # 8 : DNA-Binding Proteins : Part II 1). to examine the case of a tandem head-to-tail homodimer binding to DNA 2). to view a Zn finger motif 3). to consider the case
The purpose of this exercise is : Hmwk # 8 : DNA-Binding Proteins : Part II 1). to examine the case of a tandem head-to-tail homodimer binding to DNA 2). to view a Zn finger motif 3). to consider the case
SUPPLEMENTARY INFORMATION
 Structure of a tyrosyl-trna synthetase splicing factor bound to a group I intron RNA Paul J. Paukstelis 1, Jui-Hui Chen 2, Elaine Chase 2, Alan M. Lambowitz 1,*, and Barbara L. Golden 2,*,. 1 Institute
Structure of a tyrosyl-trna synthetase splicing factor bound to a group I intron RNA Paul J. Paukstelis 1, Jui-Hui Chen 2, Elaine Chase 2, Alan M. Lambowitz 1,*, and Barbara L. Golden 2,*,. 1 Institute
Materials Protein synthesis kit. This kit consists of 24 amino acids, 24 transfer RNAs, four messenger RNAs and one ribosome (see below).
 Protein Synthesis Instructions The purpose of today s lab is to: Understand how a cell manufactures proteins from amino acids, using information stored in the genetic code. Assemble models of four very
Protein Synthesis Instructions The purpose of today s lab is to: Understand how a cell manufactures proteins from amino acids, using information stored in the genetic code. Assemble models of four very
Proteins Higher Order Structures
 Proteins Higher Order Structures Dr. Mohammad Alsenaidy Department of Pharmaceutics College of Pharmacy King Saud University Office: AA 101 msenaidy@ksu.edu.sa Previously on PHT 426!! Protein Structures
Proteins Higher Order Structures Dr. Mohammad Alsenaidy Department of Pharmaceutics College of Pharmacy King Saud University Office: AA 101 msenaidy@ksu.edu.sa Previously on PHT 426!! Protein Structures
Supplementary Fig. S1. SAMHD1c has a more potent dntpase activity than. SAMHD1c. Purified recombinant SAMHD1c and SAMHD1c proteins (with
 Supplementary Fig. S1. SAMHD1c has a more potent dntpase activity than SAMHD1c. Purified recombinant SAMHD1c and SAMHD1c proteins (with concentration of 800nM) were incubated with 1mM dgtp for the indicated
Supplementary Fig. S1. SAMHD1c has a more potent dntpase activity than SAMHD1c. Purified recombinant SAMHD1c and SAMHD1c proteins (with concentration of 800nM) were incubated with 1mM dgtp for the indicated
Conformational changes in IgE contribute to its. uniquely slow dissociation rate from receptor FcεRI
 Conformational changes in IgE contribute to its uniquely slow dissociation rate from receptor FcεRI M.D. Holdom, A.M. Davies, J.E. Nettleship, S.C. Bagby, B. Dhaliwal, E. Girardi, J. Hunt, H.J. Gould,
Conformational changes in IgE contribute to its uniquely slow dissociation rate from receptor FcεRI M.D. Holdom, A.M. Davies, J.E. Nettleship, S.C. Bagby, B. Dhaliwal, E. Girardi, J. Hunt, H.J. Gould,
LIST OF ACRONYMS & ABBREVIATIONS
 LIST OF ACRONYMS & ABBREVIATIONS ALA ARG ASN ATD CRD CYS GLN GLU GLY GPCR HIS hstr ILE LEU LYS MET mglur1 NHDC PDB PHE PRO SER T1R2 T1R3 TMD TRP TYR THR 7-TM VFTM ZnSO 4 Alanine, A Arginine, R Asparagines,
LIST OF ACRONYMS & ABBREVIATIONS ALA ARG ASN ATD CRD CYS GLN GLU GLY GPCR HIS hstr ILE LEU LYS MET mglur1 NHDC PDB PHE PRO SER T1R2 T1R3 TMD TRP TYR THR 7-TM VFTM ZnSO 4 Alanine, A Arginine, R Asparagines,
Unit 1. DNA and the Genome
 Unit 1 DNA and the Genome Gene Expression Key Area 3 Vocabulary 1: Transcription Translation Phenotype RNA (mrna, trna, rrna) Codon Anticodon Ribosome RNA polymerase RNA splicing Introns Extrons Gene Expression
Unit 1 DNA and the Genome Gene Expression Key Area 3 Vocabulary 1: Transcription Translation Phenotype RNA (mrna, trna, rrna) Codon Anticodon Ribosome RNA polymerase RNA splicing Introns Extrons Gene Expression
Student Name Biochem. 461 Exam 1 Key, September 23, 2010
 Student Name Biochem. 461 Exam 1 Key, September 23, 2010 (100 POINTS TOTAL) ANSWER ALL QUESTIONS IN THE SPACES PROVIDED [10 pts] We will learn in Chapter 28 that ribonuclosides and deoxyribonucleosides
Student Name Biochem. 461 Exam 1 Key, September 23, 2010 (100 POINTS TOTAL) ANSWER ALL QUESTIONS IN THE SPACES PROVIDED [10 pts] We will learn in Chapter 28 that ribonuclosides and deoxyribonucleosides
Protein Structure and Function! Lecture 4: ph, pka and pi!
 Protein Structure and Function! Lecture 4: ph, pka and pi! Definition of ph and pk a! ph is a measure of the concentration of H +.! + ph = log 10[H ] For a weak acid,! HA #!!"! H + + A!, K a = [H + ][A!
Protein Structure and Function! Lecture 4: ph, pka and pi! Definition of ph and pk a! ph is a measure of the concentration of H +.! + ph = log 10[H ] For a weak acid,! HA #!!"! H + + A!, K a = [H + ][A!
Supplementary Table 1. DNA sequence synthesized to express the Zika virus NS5 protein.
 Supplementary Table 1. DNA sequence synthesized to express the Zika virus NS5 protein. MR766 NS5 sequence 1 ACAGAGAACAGATTGGTGGTGGAGGTGGGACGGGAGAGACTCTGGGAGAGAAGTGGAAAG 61 CTCGTCTGAATCAGATGTCGGCCCTGGAGTTCTACTCTTATAAAAAGTCAGGTATCACTG
Supplementary Table 1. DNA sequence synthesized to express the Zika virus NS5 protein. MR766 NS5 sequence 1 ACAGAGAACAGATTGGTGGTGGAGGTGGGACGGGAGAGACTCTGGGAGAGAAGTGGAAAG 61 CTCGTCTGAATCAGATGTCGGCCCTGGAGTTCTACTCTTATAAAAAGTCAGGTATCACTG
CS 4491/CS 7990 SPECIAL TOPICS IN BIOINFORMATICS
 1 CS 4491/CS 7990 SPECIAL TOPICS IN BIOINFORMATICS * Some contents are adapted from Dr. Jean Gao at UT Arlington Mingon Kang, PhD Computer Science, Kennesaw State University 2 Genetics The discovery of
1 CS 4491/CS 7990 SPECIAL TOPICS IN BIOINFORMATICS * Some contents are adapted from Dr. Jean Gao at UT Arlington Mingon Kang, PhD Computer Science, Kennesaw State University 2 Genetics The discovery of
SUPPLEMENTARY INFORMATION. Supplementary Figures 1-8
 SUPPLEMENTARY INFORMATION Supplementary Figures 1-8 Supplementary Figure 1. TFAM residues contacting the DNA minor groove (A) TFAM contacts on nonspecific DNA. Leu58, Ile81, Asn163, Pro178, and Leu182
SUPPLEMENTARY INFORMATION Supplementary Figures 1-8 Supplementary Figure 1. TFAM residues contacting the DNA minor groove (A) TFAM contacts on nonspecific DNA. Leu58, Ile81, Asn163, Pro178, and Leu182
Protein-Protein Interactions I
 Biochemistry 412 Protein-Protein Interactions I March 23, 2007 Macromolecular Recognition by Proteins Protein folding is a process governed by intramolecular recognition. Protein-protein association is
Biochemistry 412 Protein-Protein Interactions I March 23, 2007 Macromolecular Recognition by Proteins Protein folding is a process governed by intramolecular recognition. Protein-protein association is
 The replication of DNA Kornberg 1957 Meselson and Stahl 1958 Cairns 1963 Okazaki 1968 DNA Replication The driving force for DNA synthesis. The addition of a nucleotide to a growing polynucleotide
The replication of DNA Kornberg 1957 Meselson and Stahl 1958 Cairns 1963 Okazaki 1968 DNA Replication The driving force for DNA synthesis. The addition of a nucleotide to a growing polynucleotide
Supplementary Information Titles
 Supplementary Information Titles Please list each supplementary item and its title or caption, in the order shown below. Note that we do NOT copy edit or otherwise change supplementary information, and
Supplementary Information Titles Please list each supplementary item and its title or caption, in the order shown below. Note that we do NOT copy edit or otherwise change supplementary information, and
Structural Basis for Phosphorylationdependent. Damage Response J. N. Mark Glover Department of Biochemistry, University of Alberta, Edmonton
 Structural Basis for Phosphorylationdependent Signaling in the DNA Damage Response J. N. Mark Glover Department of Biochemistry, University of Alberta, Edmonton Abstract The response of eukaryotic cells
Structural Basis for Phosphorylationdependent Signaling in the DNA Damage Response J. N. Mark Glover Department of Biochemistry, University of Alberta, Edmonton Abstract The response of eukaryotic cells
D-3-Phosphoglycerate Dehydrogenase* Gregory A. Grant, Zhiqin Hu, and Xiao Lan Xu
 THE JOURNAL OF BIOLOGICAL CHEMISTRY Vol. 276, No. 2, Issue of January 12, pp. 1078 1083, 2001 2001 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in U.S.A. Specific Interactions
THE JOURNAL OF BIOLOGICAL CHEMISTRY Vol. 276, No. 2, Issue of January 12, pp. 1078 1083, 2001 2001 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in U.S.A. Specific Interactions
Prediction of Protein-Protein Binding Sites and Epitope Mapping. Copyright 2017 Chemical Computing Group ULC All Rights Reserved.
 Prediction of Protein-Protein Binding Sites and Epitope Mapping Epitope Mapping Antibodies interact with antigens at epitopes Epitope is collection residues on antigen Continuous (sequence) or non-continuous
Prediction of Protein-Protein Binding Sites and Epitope Mapping Epitope Mapping Antibodies interact with antigens at epitopes Epitope is collection residues on antigen Continuous (sequence) or non-continuous
Homework. A bit about the nature of the atoms of interest. Project. The role of electronega<vity
 Homework Why cited articles are especially useful. citeulike science citation index When cutting and pasting less is more. Project Your protein: I will mail these out this weekend If you haven t gotten
Homework Why cited articles are especially useful. citeulike science citation index When cutting and pasting less is more. Project Your protein: I will mail these out this weekend If you haven t gotten
CSE : Computational Issues in Molecular Biology. Lecture 19. Spring 2004
 CSE 397-497: Computational Issues in Molecular Biology Lecture 19 Spring 2004-1- Protein structure Primary structure of protein is determined by number and order of amino acids within polypeptide chain.
CSE 397-497: Computational Issues in Molecular Biology Lecture 19 Spring 2004-1- Protein structure Primary structure of protein is determined by number and order of amino acids within polypeptide chain.
Nucleic Acids, Proteins, and Enzymes
 3 Nucleic Acids, Proteins, and Enzymes Chapter 3 Nucleic Acids, Proteins, and Enzymes Key Concepts 3.1 Nucleic Acids Are Informational Macromolecules 3.2 Proteins Are Polymers with Important Structural
3 Nucleic Acids, Proteins, and Enzymes Chapter 3 Nucleic Acids, Proteins, and Enzymes Key Concepts 3.1 Nucleic Acids Are Informational Macromolecules 3.2 Proteins Are Polymers with Important Structural
Supplementary Information. Synergistic action of RNA polymerases in overcoming the nucleosomal barrier
 Supplementary Information Synergistic action of RNA polymerases in overcoming the nucleosomal barrier Jing Jin, Lu Bai, Daniel S. Johnson, Robert M. Fulbright, Maria L. Kireeva, Mikhail Kashlev, Michelle
Supplementary Information Synergistic action of RNA polymerases in overcoming the nucleosomal barrier Jing Jin, Lu Bai, Daniel S. Johnson, Robert M. Fulbright, Maria L. Kireeva, Mikhail Kashlev, Michelle
Proteins Amides from Amino Acids
 Chapter 26 and Chapter 28 Proteins Amides from Amino Acids Amino acids contain a basic amino group and an acidic carboxyl group Joined as amides between the ¾NH 2 of one amino acid and the ¾CO 2 H to the
Chapter 26 and Chapter 28 Proteins Amides from Amino Acids Amino acids contain a basic amino group and an acidic carboxyl group Joined as amides between the ¾NH 2 of one amino acid and the ¾CO 2 H to the
DNA.notebook March 08, DNA Overview
 DNA Overview Deoxyribonucleic Acid, or DNA, must be able to do 2 things: 1) give instructions for building and maintaining cells. 2) be copied each time a cell divides. DNA is made of subunits called nucleotides
DNA Overview Deoxyribonucleic Acid, or DNA, must be able to do 2 things: 1) give instructions for building and maintaining cells. 2) be copied each time a cell divides. DNA is made of subunits called nucleotides
green B 1 ) into a single unit to model the substrate in this reaction. enzyme
 Teacher Key Objectives You will use the model pieces in the kit to: Simulate enzymatic actions. Explain enzymatic specificity. Investigate two types of enzyme inhibitors used in regulating enzymatic activity.
Teacher Key Objectives You will use the model pieces in the kit to: Simulate enzymatic actions. Explain enzymatic specificity. Investigate two types of enzyme inhibitors used in regulating enzymatic activity.
1/4/18 NUCLEIC ACIDS. Nucleic Acids. Nucleic Acids. ECS129 Instructor: Patrice Koehl
 NUCLEIC ACIDS ECS129 Instructor: Patrice Koehl Nucleic Acids Nucleotides DNA Structure RNA Synthesis Function Secondary structure Tertiary interactions Wobble hypothesis DNA RNA Replication Transcription
NUCLEIC ACIDS ECS129 Instructor: Patrice Koehl Nucleic Acids Nucleotides DNA Structure RNA Synthesis Function Secondary structure Tertiary interactions Wobble hypothesis DNA RNA Replication Transcription
NUCLEIC ACIDS. ECS129 Instructor: Patrice Koehl
 NUCLEIC ACIDS ECS129 Instructor: Patrice Koehl Nucleic Acids Nucleotides DNA Structure RNA Synthesis Function Secondary structure Tertiary interactions Wobble hypothesis DNA RNA Replication Transcription
NUCLEIC ACIDS ECS129 Instructor: Patrice Koehl Nucleic Acids Nucleotides DNA Structure RNA Synthesis Function Secondary structure Tertiary interactions Wobble hypothesis DNA RNA Replication Transcription
Solution Structure of the DNA-binding Domain of GAL4 from Saccharomyces cerevisiae
 Vol. 14, No. 1-4 175 Solution Structure of the DNA-binding Domain of GAL4 from Saccharomyces cerevisiae James D. Baleja, V. Thanabal, Ted Mau, and Gerhard Wagner Department of Biological Chemistry and
Vol. 14, No. 1-4 175 Solution Structure of the DNA-binding Domain of GAL4 from Saccharomyces cerevisiae James D. Baleja, V. Thanabal, Ted Mau, and Gerhard Wagner Department of Biological Chemistry and
Station 1: DNA Structure Use the figure above to answer each of the following questions. 1.This is the subunit that DNA is composed of. 2.
 1. Station 1: DNA Structure Use the figure above to answer each of the following questions. 1.This is the subunit that DNA is composed of. 2.This subunit is composed of what 3 parts? 3.What molecules make
1. Station 1: DNA Structure Use the figure above to answer each of the following questions. 1.This is the subunit that DNA is composed of. 2.This subunit is composed of what 3 parts? 3.What molecules make
Structural basis of a novel PD-L1 nanobody for immune checkpoint blockade
 Structural basis of a novel PD-L1 nanobody for immune checkpoint blockade Supplementary Materials Supplementary methods Table S1-S Figure S1-S 1 1 1 1 1 1 1 1 1 0 1 0 Supplementary Methods Competitive
Structural basis of a novel PD-L1 nanobody for immune checkpoint blockade Supplementary Materials Supplementary methods Table S1-S Figure S1-S 1 1 1 1 1 1 1 1 1 0 1 0 Supplementary Methods Competitive
(Due Sept 9 th ) Problem Set 2
 Problem Set 2 (Due Sept 9 th ) 1. Consider these two polynucleotides: AAGCGT GCACTG a. Draw each molecule in the 2 deoxy form (DNA). SEE BELOW b. What is the sequence of the complementary strand? Write
Problem Set 2 (Due Sept 9 th ) 1. Consider these two polynucleotides: AAGCGT GCACTG a. Draw each molecule in the 2 deoxy form (DNA). SEE BELOW b. What is the sequence of the complementary strand? Write
Find this material useful? You can help our team to keep this site up and bring you even more content consider donating via the link on our site.
 Find this material useful? You can help our team to keep this site up and bring you even more content consider donating via the link on our site. Still having trouble understanding the material? Check
Find this material useful? You can help our team to keep this site up and bring you even more content consider donating via the link on our site. Still having trouble understanding the material? Check
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION A human XRCC4-XLF complex bridges DNA ends. Sara N. Andres 1, Alexandra Vergnes 2, Dejan Ristic 3, Claire Wyman 3, Mauro Modesti 2,4, and Murray Junop 2,4 1 Department of Biochemistry
SUPPLEMENTARY INFORMATION A human XRCC4-XLF complex bridges DNA ends. Sara N. Andres 1, Alexandra Vergnes 2, Dejan Ristic 3, Claire Wyman 3, Mauro Modesti 2,4, and Murray Junop 2,4 1 Department of Biochemistry
Tyrosine Kinase Assay Kit, Red*
 Rh Tyrosine Kinase Assay Kit, Red* 1.0 INTRODUCTION Part # P2882, P2883 Lit. # L0531 Rev. 11/02 Page 1 of 6 The phosphorylation of proteins by protein tyrosine kinases (PTKs) is critical to the normal
Rh Tyrosine Kinase Assay Kit, Red* 1.0 INTRODUCTION Part # P2882, P2883 Lit. # L0531 Rev. 11/02 Page 1 of 6 The phosphorylation of proteins by protein tyrosine kinases (PTKs) is critical to the normal
Examination I PHRM 836 Biochemistry for Pharmaceutical Sciences II September 29, 2015
 Examination I PHRM 836 Biochemistry for Pharmaceutical Sciences II September 29, 2015 PHRM 836 Exam I - 1 Name: Instructions 1. Check your exam to make certain that it has 9 pages including this cover
Examination I PHRM 836 Biochemistry for Pharmaceutical Sciences II September 29, 2015 PHRM 836 Exam I - 1 Name: Instructions 1. Check your exam to make certain that it has 9 pages including this cover
Proteins: Wide range of func2ons. Polypep2des. Amino Acid Monomers
 Proteins: Wide range of func2ons Proteins coded in DNA account for more than 50% of the dry mass of most cells Protein func9ons structural support storage transport cellular communica9ons movement defense
Proteins: Wide range of func2ons Proteins coded in DNA account for more than 50% of the dry mass of most cells Protein func9ons structural support storage transport cellular communica9ons movement defense
Packing of Secondary Structures
 7.88 Lecture Notes - 5 7.24/7.88J/5.48J The Protein Folding and Human Disease Packing of Secondary Structures Packing of Helices against sheets Packing of sheets against sheets Parallel Orthogonal Table:
7.88 Lecture Notes - 5 7.24/7.88J/5.48J The Protein Folding and Human Disease Packing of Secondary Structures Packing of Helices against sheets Packing of sheets against sheets Parallel Orthogonal Table:
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
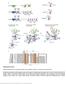 Supplementary Figure 1 Domain architecture and conformational states of the decapping complex, as revealed by structural studies. (a) Domain organization of Schizosaccharomyces pombe (Sp) and Saccharomyces
Supplementary Figure 1 Domain architecture and conformational states of the decapping complex, as revealed by structural studies. (a) Domain organization of Schizosaccharomyces pombe (Sp) and Saccharomyces
Electronic Supplementary Information
 Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 2018 Electronic Supplementary Information Unraveling the effects of amino acid substitutions
Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 2018 Electronic Supplementary Information Unraveling the effects of amino acid substitutions
