Electronic Supplementary Information
|
|
|
- Roderick Todd
- 6 years ago
- Views:
Transcription
1 Electronic Supplementary Information Ultrasensitive and Selective DNA Detection Based on Nicking Endonuclease Assisted Signal Amplification and Its Application in Cancer Cells Detection Sai Bi, Jilei Zhang, Shusheng Zhang* Key Laboratory of Eco-chemical Engineering, Ministry of Education, College of Chemistry and Molecular Engineering, Qingdao University of Science and Technology, Qingdao , P. R. China * Corresponding author. Tel.: ; Fax: shushzhang@126.com 1
2 EXPERIMENTAL SECTION Chemicals All of oligonucleotides were custom-ordered from SBS Genetech Co. Ltd. (China), and their sequences are listed in Table S1. NEase (Nt.AlwI, 10 units) was obtained from New England Biolabs. Tri(2-carboxyethyl)phosphine hydrochloride (TCEP, 98%) was purchased from Alfa Aesar (Massachusetts). H 2 O 2 with analytical grade was purchased from Shanghai Chemical Reagent Company (Shanghai,China). Luminol standard powder and hemin was ordered from Sigma-Aldrich. A luminol stock solution ( M) was prepared by dissolving it in 0.1 M NaOH solution and storing it in dark. A hemin stock solution ( M) was prepared in DMSO and stored in dark at -20 o C. Double-distilled, deionized water was used throughout the experiments. All the chemicals employed were of analytical reagent grade and were used without further purification. Table S1. DNA Sequence Used in This Work oligonucleotides sequences a name 1 5'-CCT CCC GGT GTT CGA TCC-3' 1 5'-CCT CCC GGT GTT CGC TCC-3' 1 5'-GGA GGG CCA CAA GCG AGG-3' 2 5'-CCC TTA AGT GTC CC-3' 5'-CAC AGG GTT GGG CGG GAT GGG TGG ATC GAA CAC CGG GAG 3 GTT GGG TAG GGC GGG TTG GGA ATT-3' 5'-SH-TTT TTT ACA CAG GGT AGG GTT GGG CGG GAT GGG GGA 4 TCG AAC ACC GGG AGG GGG TAG GGC GGG TTG GGT ACC CTG TGT TTT TTT-SH-3 5 5'-CCA CCG AAC TAT CCT CCC GGT GTT CGA TCC-3' 5'-AAC ACC GGG AGG ATA GTT CGG TGG CTG TTC AGG GTC TCC 6 TCC CGG TG-3' a The NEase recognition site of the target is underlined. Oligonucleotides 1 has a single mismatch at different positions of the NEase recoginition site, and 1 is a noncomplementary target DNA sequence. The mismatched positions are highlightened in italic. 2
3 Apparatus.The CL measurements were performed by using a BPCL ultraweak luminescence analyzer (Institute of Biophysics Academic Sinica, Beijing, China). The sample cell was a 2-mL quartz cuvette. Transmission electron microscopy (TEM) images were carried on a JEM-2000EX/ASID2. Magnetic Fe 3 O 4 -Au core-shell nanoparticles (Fe 3 O were obtained from Shaanxi Lifegen Co., Ltd. Preparation of MB-Fe 3 O Conjugates. The oligonucleotide-fe 3 O conjugates were synthesized by adding ~10 µl of the commercialized Fe 3 O solution (~ 5 mg/ml) to 300 µl of M dithiol-modified molecular beacon (MB). Thiol-modified DNA was activated with TCEP (10 mm) for 1 h at room temperature before attaching to Fe 3 O After shaking gently for 16 h at room temperature, the MB-Fe 3 O conjugates were aged in the solution (0.3 M NaCl, 10 mm Tris-acetate, ph 8.2) for another 48 h. Excess reagents were removed by magnetic field. Following removal of the supernatant, the resulting precipitate was washed with 300 µl of 10 mm ph 7.4 phosphate buffer containing 0.01 M NaCl, recentrifuged, and then redispersed in 10 mm ph 7.4 phosphate buffer containing 0.01 M NaCl for further use. Cancer Cell Culture. Ramos cells (target cells) and CEM cells (control cells) were cultured in cell flasks separately according to the instructions from the American Type Culture Collection. The cell line was grown to 90% confluence in RPMI 1640 medium supplemented with 10% (v/v) fetal bovine serum (FBS) and 100 IU/mL penicillin-streptomycin at 37 o C, followed by culturing cells in a humidified atmoshpere with 5% CO 2. The cell density was counted on a hemocytomer prior to each experiment. Then, a 1.0 ml suspension of ~ cells dispersed in RPMI 1640 cell media buffer was centrifuged at 3500 rpm for 5 min and washed with phosphate-buffered saline (18.6 mm phosphate, 4.2 mm KCl, and mm NaCl) five times and resuspended in 1.0 3
4 ml cell media buffer. CL Measurements. The detection of DNA (Scheme 1A in the main text) was performed in 300 µl of solution containing 2 (10 nm), 3 (10 nm) and different concentrations of target DNA 1 in 2 µl of NEBuffer. For the improved strategy (Scheme 1B in the main text), 200 µl of MB-Fe 3 O conjugates was reacted with 100 µl of target DNA 1 with different concentrations. Upon incubation at 58 o C for 5 min, 1 µl of NEase was added to the solution. After standing for 2 h at 37 o C, the resulting mixtures were heated at 80 o C for 20 min to deactivate the NEase, followed by the addition of hemin ( M), luminol ( M). The CL reaction was triggered by injecting 100 µl of H 2 O 2 ( M) with a syringe through a septum after the CL analyzer began to record at 10 s. The kinetics of CL signals between 0 to 30 s were recorded and the peak heights of the emission curves were measured by means of a photon counting unit. For cancer cell assays (Scheme 2 in the main text), the Ramos cells aptamer 5 (10 nm) and its complementary oligonucleotides 6 (10 nm) were previouly hybridized in 300 µl of 10 mm phosphate buffer containing 0.01 M NaCl (ph 7.4) at 37 o C for 1 h to form the duplex structure (II). 10 µl of target cells of different concentrations was added to the solution containing duplex structure (II) and MB-Fe 3 O conjugates followed by incubating the mixture for 1 h at 37 o C. The following procedures were the same as DNA detection. 4
5 Supplementary Results and Discussion Sensitivity of the Original Strategy of DNA Amplified Detection. Fig. S1 The calibration curve of peak height verus the concentration of target DNA from to M obtained by the strategy of Scheme 1A in the main text. To test the feasibility of our system for DNA detection, we first prepared various concentrations of a 18-mer single-stranded target DNA that contained the recognition sequence (-GATCC-) of a NEase (Nt.AlwI). The quasi-circular structure I, with a linker sequence complementary to that of the target DNA was then added to each of these solutions. Fig. S1 shows the calibration curve of peak height verus the concentration of target DNA. As the concentrations of the target DNA increased, the CL signals were enhanced. A linear range from to M was achieved with an equation of I = C (I is the CL intensity; C represents the concentration of target DNA, M; n = 7, R = ) and a detection limit of M. 5
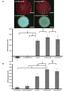
6 Selectivity of the Improved Strategy of DNA Amplified Detection. Fig. S2 CL signals of luminol-h 2 O 2 system catalyzed by HRP-mimicking DNAzyme formed by NEase assisted strand cleavage and signal amplification with different target DNA. (a) blank, (b) noncomplementary sequences, (c) single-base mismatched sequences, and (d) complementary sequences. All the concentrations of target DNA are M. To evaluate the selectivity of the detection system, the effect of base mismatch was studied in the presence of M target DNA with single-base mismatched DNA sequences and noncomplementary sequences. Fig. S2 shows that CL signals were significantly weaker than that of the complementary squences which were not significantly different from the blank. We proposed that this significant sequence specificity is attributed to the requirement of full complementary between the target DNA and the linker strand at the NEase recognition site and sufficient complementarity beyond the NEase recognition site to allow efficient hybridization. 6
7 Generality of the Improved Strategy of DNA Amplified Detection for Target DNA with Varied Lengths Fig. S3 CL intensities of target DNA sequences of different length containing NEase recognition sites: (a) 18-mer, (b) 36-mer, and (c) 72-mer. All the concentrations of target DNA are M. DNA sequences: 18-mer: 5'-CCT CCC GGT GTT CGA TCC-3'; 36-mer: 5'-GAA CAG CCA CCG AAC TAT CCT CCC GGT GTT CGA TCC-3'; 72-mer: 5'-GCA AAA TCT AAA AGA GCA CCA CCG GGA GGA GAC CCT GAA CAG CCA CCG AAC TAT CCT CCC GGT GTT CGA TCC-3'. From Fig. S3, the improved amplification strategy demonstrates remarkable generality for target DNA with varied lengths. In principle, the length of the dithiol-modified MBs structure could increase with the increase of the base number of the target strand. However, for relatively long-stranded MBs, immobilization and instability become significant challenges. Practically in our sytem, a fixed set of MBs can be readily extended to detection of long single-stranded DNA sequences that contains NEase recognition site. As shown in Fig. S3, CL detection of a range of varied length target DNA sequence (18-mer, 36-mer, and 72-mer) was achieved without the requirement of longer MBs. 7
Electronic Supplementary Information
 Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2017 Electronic Supplementary Information Dissecting binding of a β-barrel outer membrane
Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2017 Electronic Supplementary Information Dissecting binding of a β-barrel outer membrane
Lecture 10, 20/2/2002: The process of solution development - The CODEHOP strategy for automatic design of consensus-degenerate primers for PCR
 Lecture 10, 20/2/2002: The process of solution development - The CODEHOP strategy for automatic design of consensus-degenerate primers for PCR 1 The problem We wish to clone a yet unknown gene from a known
Lecture 10, 20/2/2002: The process of solution development - The CODEHOP strategy for automatic design of consensus-degenerate primers for PCR 1 The problem We wish to clone a yet unknown gene from a known
Supplement 1: Sequences of Capture Probes. Capture probes were /5AmMC6/CTG TAG GTG CGG GTG GAC GTA GTC
 Supplementary Appendixes Supplement 1: Sequences of Capture Probes. Capture probes were /5AmMC6/CTG TAG GTG CGG GTG GAC GTA GTC ACG TAG CTC CGG CTG GA-3 for vimentin, /5AmMC6/TCC CTC GCG CGT GGC TTC CGC
Supplementary Appendixes Supplement 1: Sequences of Capture Probes. Capture probes were /5AmMC6/CTG TAG GTG CGG GTG GAC GTA GTC ACG TAG CTC CGG CTG GA-3 for vimentin, /5AmMC6/TCC CTC GCG CGT GGC TTC CGC
Supplementary. Table 1: Oligonucleotides and Plasmids. complementary to positions from 77 of the SRα '- GCT CTA GAG AAC TTG AAG TAC AGA CTG C
 Supplementary Table 1: Oligonucleotides and Plasmids 913954 5'- GCT CTA GAG AAC TTG AAG TAC AGA CTG C 913955 5'- CCC AAG CTT ACA GTG TGG CCA TTC TGC TG 223396 5'- CGA CGC GTA CAG TGT GGC CAT TCT GCT G
Supplementary Table 1: Oligonucleotides and Plasmids 913954 5'- GCT CTA GAG AAC TTG AAG TAC AGA CTG C 913955 5'- CCC AAG CTT ACA GTG TGG CCA TTC TGC TG 223396 5'- CGA CGC GTA CAG TGT GGC CAT TCT GCT G
Supporting Informations
 Supporting Informations Design of Molecular Beacon as Signaling Probes for ATP Detection in Cancer Cells Based on Chemiluminescence Resonance Energy Transfer Shusheng Zhang*, Yameng Yan, and Sai Bi* Key
Supporting Informations Design of Molecular Beacon as Signaling Probes for ATP Detection in Cancer Cells Based on Chemiluminescence Resonance Energy Transfer Shusheng Zhang*, Yameng Yan, and Sai Bi* Key
Electronic Supplementary Information Sensitive detection of polynucleotide kinase using rolling circle amplification-induced chemiluminescence
 Electronic Supplementary Material (ESI) for Chemical Communications. This journal is The Royal Society of Chemistry 2014 Electronic Supplementary Information Sensitive detection of polynucleotide kinase
Electronic Supplementary Material (ESI) for Chemical Communications. This journal is The Royal Society of Chemistry 2014 Electronic Supplementary Information Sensitive detection of polynucleotide kinase
Electronic Supplementary Information
 Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2014 Electronic Supplementary Information Multiplexed Detection of Lung Cancer Cells at the Single-Molecule
Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2014 Electronic Supplementary Information Multiplexed Detection of Lung Cancer Cells at the Single-Molecule
Supporting information for Biochemistry, 1995, 34(34), , DOI: /bi00034a013
 Supporting information for Biochemistry, 1995, 34(34), 10807 10815, DOI: 10.1021/bi00034a013 LESNIK 10807-1081 Terms & Conditions Electronic Supporting Information files are available without a subscription
Supporting information for Biochemistry, 1995, 34(34), 10807 10815, DOI: 10.1021/bi00034a013 LESNIK 10807-1081 Terms & Conditions Electronic Supporting Information files are available without a subscription
Supporting Information
 Supporting Information Transfection of DNA Cages into Mammalian Cells Email: a.turberfield@physics.ox.ac.uk Table of Contents Supporting Figure 1 DNA tetrahedra used in transfection experiments 2 Supporting
Supporting Information Transfection of DNA Cages into Mammalian Cells Email: a.turberfield@physics.ox.ac.uk Table of Contents Supporting Figure 1 DNA tetrahedra used in transfection experiments 2 Supporting
Hes6. PPARα. PPARγ HNF4 CD36
 SUPPLEMENTARY INFORMATION Supplementary Table Positions and Sequences of ChIP primers -63 AGGTCACTGCCA -79 AGGTCTGCTGTG Hes6-0067 GGGCAaAGTTCA ACOT -395 GGGGCAgAGTTCA PPARα -309 GGCTCAaAGTTCAaGTTCA CPTa
SUPPLEMENTARY INFORMATION Supplementary Table Positions and Sequences of ChIP primers -63 AGGTCACTGCCA -79 AGGTCTGCTGTG Hes6-0067 GGGCAaAGTTCA ACOT -395 GGGGCAgAGTTCA PPARα -309 GGCTCAaAGTTCAaGTTCA CPTa
Gene synthesis by circular assembly amplification
 Gene synthesis by circular assembly amplification Duhee Bang & George M Church Supplementary figures and text: Supplementary Figure 1. Dpo4 gene (1.05kb) construction by various methods. Supplementary
Gene synthesis by circular assembly amplification Duhee Bang & George M Church Supplementary figures and text: Supplementary Figure 1. Dpo4 gene (1.05kb) construction by various methods. Supplementary
Y-chromosomal haplogroup typing Using SBE reaction
 Schematic of multiplex PCR followed by SBE reaction Multiplex PCR Exo SAP purification SBE reaction 5 A 3 ddatp ddgtp 3 T 5 A G 3 T 5 3 5 G C 5 3 3 C 5 ddttp ddctp 5 T 3 T C 3 A 5 3 A 5 5 C 3 3 G 5 3 G
Schematic of multiplex PCR followed by SBE reaction Multiplex PCR Exo SAP purification SBE reaction 5 A 3 ddatp ddgtp 3 T 5 A G 3 T 5 3 5 G C 5 3 3 C 5 ddttp ddctp 5 T 3 T C 3 A 5 3 A 5 5 C 3 3 G 5 3 G
Electronic Supplementary Information
 Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2015 Electronic Supplementary Information Amplified Binding-Induced Homogeneous Assay through Catalytic
Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2015 Electronic Supplementary Information Amplified Binding-Induced Homogeneous Assay through Catalytic
PGRP negatively regulates NOD-mediated cytokine production in rainbow trout liver cells
 Supplementary Information for: PGRP negatively regulates NOD-mediated cytokine production in rainbow trout liver cells Ju Hye Jang 1, Hyun Kim 2, Mi Jung Jang 2, Ju Hyun Cho 1,2,* 1 Research Institute
Supplementary Information for: PGRP negatively regulates NOD-mediated cytokine production in rainbow trout liver cells Ju Hye Jang 1, Hyun Kim 2, Mi Jung Jang 2, Ju Hyun Cho 1,2,* 1 Research Institute
Add 5µl of 3N NaOH to DNA sample (final concentration 0.3N NaOH).
 Bisulfite Treatment of DNA Dilute DNA sample to 2µg DNA in 50µl ddh 2 O. Add 5µl of 3N NaOH to DNA sample (final concentration 0.3N NaOH). Incubate in a 37ºC water bath for 30 minutes. To 55µl samples
Bisulfite Treatment of DNA Dilute DNA sample to 2µg DNA in 50µl ddh 2 O. Add 5µl of 3N NaOH to DNA sample (final concentration 0.3N NaOH). Incubate in a 37ºC water bath for 30 minutes. To 55µl samples
Supplementary Figure 1A A404 Cells +/- Retinoic Acid
 Supplementary Figure 1A A44 Cells +/- Retinoic Acid 1 1 H3 Lys4 di-methylation SM-actin VEC cfos (-) RA (+) RA 14 1 1 8 6 4 H3 Lys79 di-methylation SM-actin VEC cfos (-) RA (+) RA Supplementary Figure
Supplementary Figure 1A A44 Cells +/- Retinoic Acid 1 1 H3 Lys4 di-methylation SM-actin VEC cfos (-) RA (+) RA 14 1 1 8 6 4 H3 Lys79 di-methylation SM-actin VEC cfos (-) RA (+) RA Supplementary Figure
Supplemental Data Supplemental Figure 1.
 Supplemental Data Supplemental Figure 1. Silique arrangement in the wild-type, jhs, and complemented lines. Wild-type (WT) (A), the jhs1 mutant (B,C), and the jhs1 mutant complemented with JHS1 (Com) (D)
Supplemental Data Supplemental Figure 1. Silique arrangement in the wild-type, jhs, and complemented lines. Wild-type (WT) (A), the jhs1 mutant (B,C), and the jhs1 mutant complemented with JHS1 (Com) (D)
Arabidopsis actin depolymerizing factor AtADF4 mediates defense signal transduction triggered by the Pseudomonas syringae effector AvrPphB
 Arabidopsis actin depolymerizing factor mediates defense signal transduction triggered by the Pseudomonas syringae effector AvrPphB Files in this Data Supplement: Supplemental Table S1 Supplemental Table
Arabidopsis actin depolymerizing factor mediates defense signal transduction triggered by the Pseudomonas syringae effector AvrPphB Files in this Data Supplement: Supplemental Table S1 Supplemental Table
Figure S1. Characterization of the irx9l-1 mutant. (A) Diagram of the Arabidopsis IRX9L gene drawn based on information from TAIR (the Arabidopsis
 1 2 3 4 5 6 7 8 9 10 11 12 Figure S1. Characterization of the irx9l-1 mutant. (A) Diagram of the Arabidopsis IRX9L gene drawn based on information from TAIR (the Arabidopsis Information Research). Exons
1 2 3 4 5 6 7 8 9 10 11 12 Figure S1. Characterization of the irx9l-1 mutant. (A) Diagram of the Arabidopsis IRX9L gene drawn based on information from TAIR (the Arabidopsis Information Research). Exons
Supplementary Information. Construction of Lasso Peptide Fusion Proteins
 Supplementary Information Construction of Lasso Peptide Fusion Proteins Chuhan Zong 1, Mikhail O. Maksimov 2, A. James Link 2,3 * Departments of 1 Chemistry, 2 Chemical and Biological Engineering, and
Supplementary Information Construction of Lasso Peptide Fusion Proteins Chuhan Zong 1, Mikhail O. Maksimov 2, A. James Link 2,3 * Departments of 1 Chemistry, 2 Chemical and Biological Engineering, and
SUPPORTING INFORMATION
 SUPPORTING INFORMATION Investigation of the Biosynthesis of the Lasso Peptide Chaxapeptin Using an E. coli-based Production System Helena Martin-Gómez, Uwe Linne, Fernando Albericio, Judit Tulla-Puche,*
SUPPORTING INFORMATION Investigation of the Biosynthesis of the Lasso Peptide Chaxapeptin Using an E. coli-based Production System Helena Martin-Gómez, Uwe Linne, Fernando Albericio, Judit Tulla-Puche,*
Quantitative reverse-transcription PCR. Transcript levels of flgs, flgr, flia and flha were
 1 Supplemental methods 2 3 4 5 6 7 8 9 1 11 12 13 14 15 16 17 18 19 21 22 23 Quantitative reverse-transcription PCR. Transcript levels of flgs, flgr, flia and flha were monitored by quantitative reverse-transcription
1 Supplemental methods 2 3 4 5 6 7 8 9 1 11 12 13 14 15 16 17 18 19 21 22 23 Quantitative reverse-transcription PCR. Transcript levels of flgs, flgr, flia and flha were monitored by quantitative reverse-transcription
NAME:... MODEL ANSWER... STUDENT NUMBER:... Maximum marks: 50. Internal Examiner: Hugh Murrell, Computer Science, UKZN
 COMP710, Bioinformatics with Julia, Test One, Thursday the 20 th of April, 2017, 09h30-11h30 1 NAME:...... MODEL ANSWER... STUDENT NUMBER:...... Maximum marks: 50 Internal Examiner: Hugh Murrell, Computer
COMP710, Bioinformatics with Julia, Test One, Thursday the 20 th of April, 2017, 09h30-11h30 1 NAME:...... MODEL ANSWER... STUDENT NUMBER:...... Maximum marks: 50 Internal Examiner: Hugh Murrell, Computer
ΔPDD1 x ΔPDD1. ΔPDD1 x wild type. 70 kd Pdd1. Pdd3
 Supplemental Fig. S1 ΔPDD1 x wild type ΔPDD1 x ΔPDD1 70 kd Pdd1 50 kd 37 kd Pdd3 Supplemental Fig. S1. ΔPDD1 strains express no detectable Pdd1 protein. Western blot analysis of whole-protein extracts
Supplemental Fig. S1 ΔPDD1 x wild type ΔPDD1 x ΔPDD1 70 kd Pdd1 50 kd 37 kd Pdd3 Supplemental Fig. S1. ΔPDD1 strains express no detectable Pdd1 protein. Western blot analysis of whole-protein extracts
MacBlunt PCR Cloning Kit Manual
 MacBlunt PCR Cloning Kit Manual Shipping and Storage MacBlunt PCR Cloning Kits are shipped on dry ice. Each kit contains a box with cloning reagents and an attached bag with Eco-Blue Competent Cells (optional).
MacBlunt PCR Cloning Kit Manual Shipping and Storage MacBlunt PCR Cloning Kits are shipped on dry ice. Each kit contains a box with cloning reagents and an attached bag with Eco-Blue Competent Cells (optional).
Supplemental Data. mir156-regulated SPL Transcription. Factors Define an Endogenous Flowering. Pathway in Arabidopsis thaliana
 Cell, Volume 138 Supplemental Data mir156-regulated SPL Transcription Factors Define an Endogenous Flowering Pathway in Arabidopsis thaliana Jia-Wei Wang, Benjamin Czech, and Detlef Weigel Table S1. Interaction
Cell, Volume 138 Supplemental Data mir156-regulated SPL Transcription Factors Define an Endogenous Flowering Pathway in Arabidopsis thaliana Jia-Wei Wang, Benjamin Czech, and Detlef Weigel Table S1. Interaction
RPA-AB RPA-C Supplemental Figure S1: SDS-PAGE stained with Coomassie Blue after protein purification.
 RPA-AB RPA-C (a) (b) (c) (d) (e) (f) Supplemental Figure S: SDS-PAGE stained with Coomassie Blue after protein purification. (a) RPA; (b) RPA-AB; (c) RPA-CDE; (d) RPA-CDE core; (e) RPA-DE; and (f) RPA-C
RPA-AB RPA-C (a) (b) (c) (d) (e) (f) Supplemental Figure S: SDS-PAGE stained with Coomassie Blue after protein purification. (a) RPA; (b) RPA-AB; (c) RPA-CDE; (d) RPA-CDE core; (e) RPA-DE; and (f) RPA-C
Supplemental Information. Human Senataxin Resolves RNA/DNA Hybrids. Formed at Transcriptional Pause Sites. to Promote Xrn2-Dependent Termination
 Supplemental Information Molecular Cell, Volume 42 Human Senataxin Resolves RNA/DNA Hybrids Formed at Transcriptional Pause Sites to Promote Xrn2-Dependent Termination Konstantina Skourti-Stathaki, Nicholas
Supplemental Information Molecular Cell, Volume 42 Human Senataxin Resolves RNA/DNA Hybrids Formed at Transcriptional Pause Sites to Promote Xrn2-Dependent Termination Konstantina Skourti-Stathaki, Nicholas
Supporting Online Information
 Supporting Online Information Isolation of Human Genomic DNA Sequences with Expanded Nucleobase Selectivity Preeti Rathi, Sara Maurer, Grzegorz Kubik and Daniel Summerer* Department of Chemistry and Chemical
Supporting Online Information Isolation of Human Genomic DNA Sequences with Expanded Nucleobase Selectivity Preeti Rathi, Sara Maurer, Grzegorz Kubik and Daniel Summerer* Department of Chemistry and Chemical
PCR analysis was performed to show the presence and the integrity of the var1csa and var-
 Supplementary information: Methods: Table S1: Primer Name Nucleotide sequence (5-3 ) DBL3-F tcc ccg cgg agt gaa aca tca tgt gac tg DBL3-R gac tag ttt ctt tca ata aat cac tcg c DBL5-F cgc cct agg tgc ttc
Supplementary information: Methods: Table S1: Primer Name Nucleotide sequence (5-3 ) DBL3-F tcc ccg cgg agt gaa aca tca tgt gac tg DBL3-R gac tag ttt ctt tca ata aat cac tcg c DBL5-F cgc cct agg tgc ttc
Supporting Information
 Supporting Information Table S1. Oligonucleotide sequences used in this work Oligo DNA A B C D CpG-A CpG-B CpG-C CpG-D Sequence 5 ACA TTC CTA AGT CTG AAA CAT TAC AGC TTG CTA CAC GAG AAG AGC CGC CAT AGT
Supporting Information Table S1. Oligonucleotide sequences used in this work Oligo DNA A B C D CpG-A CpG-B CpG-C CpG-D Sequence 5 ACA TTC CTA AGT CTG AAA CAT TAC AGC TTG CTA CAC GAG AAG AGC CGC CAT AGT
A high efficient electrochemiluminescence resonance energy. transfer system in one nanostructure: its application for
 Supporting Information for A high efficient electrochemiluminescence resonance energy transfer system in one nanostructure: its application for ultrasensitive detection of microrna in cancer cells Zhaoyang
Supporting Information for A high efficient electrochemiluminescence resonance energy transfer system in one nanostructure: its application for ultrasensitive detection of microrna in cancer cells Zhaoyang
SUPPLEMENTARY INFORMATION
 1. RNA/DNA sequences used in this study 2. Height and stiffness measurements on hybridized molecules 3. Stiffness maps at varying concentrations of target DNA 4. Stiffness measurements on RNA/DNA hybrids.
1. RNA/DNA sequences used in this study 2. Height and stiffness measurements on hybridized molecules 3. Stiffness maps at varying concentrations of target DNA 4. Stiffness measurements on RNA/DNA hybrids.
Supplemental Table 1. Mutant ADAMTS3 alleles detected in HEK293T clone 4C2. WT CCTGTCACTTTGGTTGATAGC MVLLSLWLIAAALVEVR
 Supplemental Dataset Supplemental Table 1. Mutant ADAMTS3 alleles detected in HEK293T clone 4C2. DNA sequence Amino acid sequence WT CCTGTCACTTTGGTTGATAGC MVLLSLWLIAAALVEVR Allele 1 CCTGTC------------------GATAGC
Supplemental Dataset Supplemental Table 1. Mutant ADAMTS3 alleles detected in HEK293T clone 4C2. DNA sequence Amino acid sequence WT CCTGTCACTTTGGTTGATAGC MVLLSLWLIAAALVEVR Allele 1 CCTGTC------------------GATAGC
Disease and selection in the human genome 3
 Disease and selection in the human genome 3 Ka/Ks revisited Please sit in row K or forward RBFD: human populations, adaptation and immunity Neandertal Museum, Mettman Germany Sequence genome Measure expression
Disease and selection in the human genome 3 Ka/Ks revisited Please sit in row K or forward RBFD: human populations, adaptation and immunity Neandertal Museum, Mettman Germany Sequence genome Measure expression
Anti-Pim-1 (Cat#3247), anti-met (Cat#3127), anti-ron (Cat#2654), Anti-EGFR
 Supplementary Methods Antibodies Anti-Pim-1 (Cat#3247), anti-met (Cat#3127), anti-ron (Cat#2654), Anti-EGFR (Cat#2646), anti-igf1r (Cat#3018), anti-insr (Cat#3020), anti-akt (pan, Cat#4691), anti-phospho-akt
Supplementary Methods Antibodies Anti-Pim-1 (Cat#3247), anti-met (Cat#3127), anti-ron (Cat#2654), Anti-EGFR (Cat#2646), anti-igf1r (Cat#3018), anti-insr (Cat#3020), anti-akt (pan, Cat#4691), anti-phospho-akt
Sequencing of DNA lesions facilitated by site-specific excision via base. excision repair DNA glycosylases yielding ligatable gaps
 Supporting information Sequencing of DNA lesions facilitated by site-specific excision via base excision repair DNA glycosylases yielding ligatable gaps Jan Riedl, Aaron M. Fleming, and Cynthia J. Burrows*
Supporting information Sequencing of DNA lesions facilitated by site-specific excision via base excision repair DNA glycosylases yielding ligatable gaps Jan Riedl, Aaron M. Fleming, and Cynthia J. Burrows*
Supporting Information. Trifluoroacetophenone-Linked Nucleotides and DNA for Studying of DNA-protein Interactions by 19 F NMR Spectroscopy
 Supporting Information Trifluoroacetophenone-Linked Nucleotides and DNA for Studying of DNA-protein Interactions by 19 F NMR Spectroscopy Agata Olszewska, Radek Pohl and Michal Hocek # * Institute of Organic
Supporting Information Trifluoroacetophenone-Linked Nucleotides and DNA for Studying of DNA-protein Interactions by 19 F NMR Spectroscopy Agata Olszewska, Radek Pohl and Michal Hocek # * Institute of Organic
hcd1tg/hj1tg/ ApoE-/- hcd1tg/hj1tg/ ApoE+/+
 ApoE+/+ ApoE-/- ApoE-/- H&E (1x) Supplementary Figure 1. No obvious pathology is observed in the colon of diseased ApoE-/me. Colon samples were fixed in 1% formalin and laid out in Swiss rolls for paraffin
ApoE+/+ ApoE-/- ApoE-/- H&E (1x) Supplementary Figure 1. No obvious pathology is observed in the colon of diseased ApoE-/me. Colon samples were fixed in 1% formalin and laid out in Swiss rolls for paraffin
Supporting Information. Copyright Wiley-VCH Verlag GmbH & Co. KGaA, Weinheim, 2006
 Supporting Information Copyright Wiley-VCH Verlag GmbH & Co. KGaA, 69451 Weinheim, 2006 Copyright Wiley-VCH Verlag GmbH & Co. KGaA, 69451 Weinheim, 2006 Supporting Information for Expanding the Genetic
Supporting Information Copyright Wiley-VCH Verlag GmbH & Co. KGaA, 69451 Weinheim, 2006 Copyright Wiley-VCH Verlag GmbH & Co. KGaA, 69451 Weinheim, 2006 Supporting Information for Expanding the Genetic
Materials Protein synthesis kit. This kit consists of 24 amino acids, 24 transfer RNAs, four messenger RNAs and one ribosome (see below).
 Protein Synthesis Instructions The purpose of today s lab is to: Understand how a cell manufactures proteins from amino acids, using information stored in the genetic code. Assemble models of four very
Protein Synthesis Instructions The purpose of today s lab is to: Understand how a cell manufactures proteins from amino acids, using information stored in the genetic code. Assemble models of four very
Supplemental material
 Supplemental material Diversity of O-antigen repeat-unit structures can account for the substantial sequence variation of Wzx translocases Yaoqin Hong and Peter R. Reeves School of Molecular Bioscience,
Supplemental material Diversity of O-antigen repeat-unit structures can account for the substantial sequence variation of Wzx translocases Yaoqin Hong and Peter R. Reeves School of Molecular Bioscience,
Nongenetic Reprogramming of the Ligand Specificity. of Growth Factor Receptors by Bispecific DNA Aptamers
 Supporting Information For Nongenetic Reprogramming of the Ligand Specificity of Growth Factor Receptors by Bispecific DNA Aptamers Ryosuke Ueki,* Saki Atsuta, Ayaka Ueki and Shinsuke Sando* Department
Supporting Information For Nongenetic Reprogramming of the Ligand Specificity of Growth Factor Receptors by Bispecific DNA Aptamers Ryosuke Ueki,* Saki Atsuta, Ayaka Ueki and Shinsuke Sando* Department
Sensitive SERS Detection of DNA and Lysozyme Based on. Polymerase Assisted Cross Strand-Displacement Amplification
 Supplementary Information Sensitive SERS Detection of DNA and Lysozyme Based on Polymerase Assisted Cross Strand-Displacement Amplification Ying Li, Chengcun Lei, Yan Zeng, Xiaoting Ji and Shusheng Zhang*
Supplementary Information Sensitive SERS Detection of DNA and Lysozyme Based on Polymerase Assisted Cross Strand-Displacement Amplification Ying Li, Chengcun Lei, Yan Zeng, Xiaoting Ji and Shusheng Zhang*
Project 07/111 Final Report October 31, Project Title: Cloning and expression of porcine complement C3d for enhanced vaccines
 Project 07/111 Final Report October 31, 2007. Project Title: Cloning and expression of porcine complement C3d for enhanced vaccines Project Leader: Dr Douglas C. Hodgins (519-824-4120 Ex 54758, fax 519-824-5930)
Project 07/111 Final Report October 31, 2007. Project Title: Cloning and expression of porcine complement C3d for enhanced vaccines Project Leader: Dr Douglas C. Hodgins (519-824-4120 Ex 54758, fax 519-824-5930)
Supporting Information
 Supporting Information Wiley-VCH 2007 69451 Weinheim, Germany Site-Specific Control of Distances between Gold Nanoparticles using Phosphorothioate Anchors on DNA and a Short Bifunctional Molecular Fastener
Supporting Information Wiley-VCH 2007 69451 Weinheim, Germany Site-Specific Control of Distances between Gold Nanoparticles using Phosphorothioate Anchors on DNA and a Short Bifunctional Molecular Fastener
Supplemental Data. Bennett et al. (2010). Plant Cell /tpc
 BRN1 ---------MSSSNGGVPPGFRFHPTDEELLHYYLKKKISYEKFEMEVIKEVDLNKIEPWDLQDRCKIGSTPQNEWYFFSHKDRKYPTGS 81 BRN2 --------MGSSSNGGVPPGFRFHPTDEELLHYYLKKKISYQKFEMEVIREVDLNKLEPWDLQERCKIGSTPQNEWYFFSHKDRKYPTGS 82 SMB
BRN1 ---------MSSSNGGVPPGFRFHPTDEELLHYYLKKKISYEKFEMEVIKEVDLNKIEPWDLQDRCKIGSTPQNEWYFFSHKDRKYPTGS 81 BRN2 --------MGSSSNGGVPPGFRFHPTDEELLHYYLKKKISYQKFEMEVIREVDLNKLEPWDLQERCKIGSTPQNEWYFFSHKDRKYPTGS 82 SMB
Modular Design of Ultrahigh-Intensity Nanoparticle Probe for Cancer Cell Imaging and Rapid Visual Detection of Nucleic Acid
 Supporting Information for Modular Design of Ultrahigh-Intensity Nanoparticle Probe for Cancer Cell Imaging and Rapid Visual Detection of Nucleic Acid Hua Zhong, Rumin Zhang, Hui Zhang, Shusheng Zhang*
Supporting Information for Modular Design of Ultrahigh-Intensity Nanoparticle Probe for Cancer Cell Imaging and Rapid Visual Detection of Nucleic Acid Hua Zhong, Rumin Zhang, Hui Zhang, Shusheng Zhang*
Converting rabbit hybridoma into recombinant antibodies with effective transient production in an optimized human expression system
 Converting rabbit hybridoma into recombinant antibodies with effective transient production in an optimized human expression system Dr. Tim Welsink Molecular Biology Transient Gene Expression OUTLINE Short
Converting rabbit hybridoma into recombinant antibodies with effective transient production in an optimized human expression system Dr. Tim Welsink Molecular Biology Transient Gene Expression OUTLINE Short
Amplified Analysis of DNA by the Autonomous Assembly of Polymers Consisting of DNAzyme Wires
 Supporting information for the paper: Amplified Analysis of DNA by the Autonomous Assembly of Polymers Consisting of DNAzyme Wires Fuan Wang, Johann Elbaz, Ron Orbach, Nimrod Magen and Itamar Willner*
Supporting information for the paper: Amplified Analysis of DNA by the Autonomous Assembly of Polymers Consisting of DNAzyme Wires Fuan Wang, Johann Elbaz, Ron Orbach, Nimrod Magen and Itamar Willner*
Table S1. Bacterial strains (Related to Results and Experimental Procedures)
 Table S1. Bacterial strains (Related to Results and Experimental Procedures) Strain number Relevant genotype Source or reference 1045 AB1157 Graham Walker (Donnelly and Walker, 1989) 2458 3084 (MG1655)
Table S1. Bacterial strains (Related to Results and Experimental Procedures) Strain number Relevant genotype Source or reference 1045 AB1157 Graham Walker (Donnelly and Walker, 1989) 2458 3084 (MG1655)
Electronic Supplementary Information
 Electronic Supplementary Material (ESI) for Chemical Communications. This journal is The Royal Society of Chemistry 2015 Electronic Supplementary Information Photocatalytic Oxidation of TMB with the Double
Electronic Supplementary Material (ESI) for Chemical Communications. This journal is The Royal Society of Chemistry 2015 Electronic Supplementary Information Photocatalytic Oxidation of TMB with the Double
DNA Tetrahedron-Based Molecular Beacon for Tumor-Related
 Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2015 Electronic Supporting Information (ESI) DNA Tetrahedron-Based Molecular Beacon for Tumor-Related
Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2015 Electronic Supporting Information (ESI) DNA Tetrahedron-Based Molecular Beacon for Tumor-Related
Specific Detection of Cancer Cells through Aggregation- Induced Emission of a Light-Up Bioprobe
 Electronic Supplementary Material (ESI) for Chemical Communications. This journal is The Royal Society of Chemistry 2017 Supporting Information Specific Detection of Cancer Cells through Aggregation- Induced
Electronic Supplementary Material (ESI) for Chemical Communications. This journal is The Royal Society of Chemistry 2017 Supporting Information Specific Detection of Cancer Cells through Aggregation- Induced
Supporting Information
 Supporting Information Ultrasensitive Electrochemiluminescence Biosensor for MicroRNA Detection by 3D DNA Walking Machine Based Target Conversion and Distance-Controllable Signal Quenching and Enhancing
Supporting Information Ultrasensitive Electrochemiluminescence Biosensor for MicroRNA Detection by 3D DNA Walking Machine Based Target Conversion and Distance-Controllable Signal Quenching and Enhancing
ORFs and genes. Please sit in row K or forward
 ORFs and genes Please sit in row K or forward https://www.flickr.com/photos/teseum/3231682806/in/photostream/ Question: why do some strains of Vibrio cause cholera and others don t? Methods Mechanisms
ORFs and genes Please sit in row K or forward https://www.flickr.com/photos/teseum/3231682806/in/photostream/ Question: why do some strains of Vibrio cause cholera and others don t? Methods Mechanisms
strain devoid of the aox1 gene [1]. Thus, the identification of AOX1 in the intracellular
![strain devoid of the aox1 gene [1]. Thus, the identification of AOX1 in the intracellular strain devoid of the aox1 gene [1]. Thus, the identification of AOX1 in the intracellular](/thumbs/72/66361212.jpg) Additional file 2 Identification of AOX1 in P. pastoris GS115 with a Mut s phenotype Results and Discussion The HBsAg producing strain was originally identified as a Mut s (methanol utilization slow) strain
Additional file 2 Identification of AOX1 in P. pastoris GS115 with a Mut s phenotype Results and Discussion The HBsAg producing strain was originally identified as a Mut s (methanol utilization slow) strain
Glutathione (GSH)-Decorated Magnetic Nanoparticles for Binding Glutathione-S-transferase (GST) Fusion Protein and Manipulating Live Cells
 Glutathione (GSH)-Decorated Magnetic Nanoparticles for Binding Glutathione-S-transferase (GST) Fusion Protein and Manipulating Live Cells Yue Pan, Marcus J. C. Long, Xinming Li, Junfeng Shi, Lizbeth Hedstrom,
Glutathione (GSH)-Decorated Magnetic Nanoparticles for Binding Glutathione-S-transferase (GST) Fusion Protein and Manipulating Live Cells Yue Pan, Marcus J. C. Long, Xinming Li, Junfeng Shi, Lizbeth Hedstrom,
SAY IT WITH DNA: Protein Synthesis Activity by Larry Flammer
 TEACHER S GUIDE SAY IT WITH DNA: Protein Synthesis Activity by Larry Flammer SYNOPSIS This activity uses the metaphor of decoding a secret message for the Protein Synthesis process. Students teach themselves
TEACHER S GUIDE SAY IT WITH DNA: Protein Synthesis Activity by Larry Flammer SYNOPSIS This activity uses the metaphor of decoding a secret message for the Protein Synthesis process. Students teach themselves
II 0.95 DM2 (RPP1) DM3 (At3g61540) b
 Table S2. F 2 Segregation Ratios at 16 C, Related to Figure 2 Cross n c Phenotype Model e 2 Locus A Locus B Normal F 1 -like Enhanced d Uk-1/Uk-3 149 64 36 49 DM2 (RPP1) DM1 (SSI4) a Bla-1/Hh-0 F 3 111
Table S2. F 2 Segregation Ratios at 16 C, Related to Figure 2 Cross n c Phenotype Model e 2 Locus A Locus B Normal F 1 -like Enhanced d Uk-1/Uk-3 149 64 36 49 DM2 (RPP1) DM1 (SSI4) a Bla-1/Hh-0 F 3 111
Cat. # Product Size DS130 DynaExpress TA PCR Cloning Kit (ptakn-2) 20 reactions Box 1 (-20 ) ptakn-2 Vector, linearized 20 µl (50 ng/µl) 1
 Product Name: Kit Component TA PCR Cloning Kit (ptakn-2) Cat. # Product Size DS130 TA PCR Cloning Kit (ptakn-2) 20 reactions Box 1 (-20 ) ptakn-2 Vector, linearized 20 µl (50 ng/µl) 1 2 Ligation Buffer
Product Name: Kit Component TA PCR Cloning Kit (ptakn-2) Cat. # Product Size DS130 TA PCR Cloning Kit (ptakn-2) 20 reactions Box 1 (-20 ) ptakn-2 Vector, linearized 20 µl (50 ng/µl) 1 2 Ligation Buffer
An enzyme-free DNA walker that moves on the surface of. functionalized magnetic microparticles and its biosensing analysis
 Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2017 Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry
Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2017 Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry
Legends for supplementary figures 1-3
 High throughput resistance profiling of Plasmodium falciparum infections based on custom dual indexing and Illumina next generation sequencing-technology Sidsel Nag 1,2 *, Marlene D. Dalgaard 3, Poul-Erik
High throughput resistance profiling of Plasmodium falciparum infections based on custom dual indexing and Illumina next generation sequencing-technology Sidsel Nag 1,2 *, Marlene D. Dalgaard 3, Poul-Erik
An engineered tryptophan zipper-type peptide as a molecular recognition scaffold
 SUPPLEMENTARY MATERIAL An engineered tryptophan zipper-type peptide as a molecular recognition scaffold Zihao Cheng and Robert E. Campbell* Supplementary Methods Library construction for FRET-based screening
SUPPLEMENTARY MATERIAL An engineered tryptophan zipper-type peptide as a molecular recognition scaffold Zihao Cheng and Robert E. Campbell* Supplementary Methods Library construction for FRET-based screening
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/10/494/eaan6284/dc1 Supplementary Materials for Activation of master virulence regulator PhoP in acidic ph requires the Salmonella-specific protein UgtL Jeongjoon
www.sciencesignaling.org/cgi/content/full/10/494/eaan6284/dc1 Supplementary Materials for Activation of master virulence regulator PhoP in acidic ph requires the Salmonella-specific protein UgtL Jeongjoon
Dierks Supplementary Fig. S1
 Dierks Supplementary Fig. S1 ITK SYK PH TH K42R wt K42R (kinase deficient) R29C E42K Y323F R29C E42K Y323F (reduced phospholipid binding) (enhanced phospholipid binding) (reduced Cbl binding) E42K Y323F
Dierks Supplementary Fig. S1 ITK SYK PH TH K42R wt K42R (kinase deficient) R29C E42K Y323F R29C E42K Y323F (reduced phospholipid binding) (enhanced phospholipid binding) (reduced Cbl binding) E42K Y323F
evaluated with UAS CLB eliciting UAS CIT -N Libraries increase in the
 Supplementary Figures Supplementary Figure 1: Promoter scaffold library assemblies. Many ensembless of libraries were evaluated in this work. As a legend, the box outline color in top half of the figure
Supplementary Figures Supplementary Figure 1: Promoter scaffold library assemblies. Many ensembless of libraries were evaluated in this work. As a legend, the box outline color in top half of the figure
Supplementary Information
 Supplementary Information A general solution for opening double-stranded DNA for isothermal amplification Gangyi Chen, Juan Dong, Yi Yuan, Na Li, Xin Huang, Xin Cui* and Zhuo Tang* Supplementary Materials
Supplementary Information A general solution for opening double-stranded DNA for isothermal amplification Gangyi Chen, Juan Dong, Yi Yuan, Na Li, Xin Huang, Xin Cui* and Zhuo Tang* Supplementary Materials
Supplemental information for. Fabrication of Satellite Silver-Gold Nanoclusters Linked by DNA Hybridization
 Supplemental information for Fabrication of Satellite Silver-Gold Nanoclusters Linked by DNA Hybridization Suchetan Pal, Jaswinder Sharma, Hao Yan and Yan Liu Biodesign Institute, Arizona State University,
Supplemental information for Fabrication of Satellite Silver-Gold Nanoclusters Linked by DNA Hybridization Suchetan Pal, Jaswinder Sharma, Hao Yan and Yan Liu Biodesign Institute, Arizona State University,
Universal Split Spinach Aptamer (USSA) for Nucleic Acid Analysis and DNA Computation
 Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2017 Supporting materials Universal Split Spinach Aptamer (USSA) for Nucleic Acid Analysis and DNA Computation
Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2017 Supporting materials Universal Split Spinach Aptamer (USSA) for Nucleic Acid Analysis and DNA Computation
Multiplexing Genome-scale Engineering
 Multiplexing Genome-scale Engineering Harris Wang, Ph.D. Department of Systems Biology Department of Pathology & Cell Biology http://wanglab.c2b2.columbia.edu Rise of Genomics An Expanding Toolbox Esvelt
Multiplexing Genome-scale Engineering Harris Wang, Ph.D. Department of Systems Biology Department of Pathology & Cell Biology http://wanglab.c2b2.columbia.edu Rise of Genomics An Expanding Toolbox Esvelt
S4B fluorescence (AU)
 A S4B fluorescence (AU) S4B fluorescence (AU) dsbb csgba csgd dsbb csgba bcsa 5000 * NS NS 4000 * 3000 2000 1000 0 ΔcsgBAΔbcsA ΔcsgDΔdsbBΔbcsA ΔcsgBA ΔdsbBΔcsgBA ΔcsgDΔdsbB B -1000 4000 * * NS 3500 * 3000
A S4B fluorescence (AU) S4B fluorescence (AU) dsbb csgba csgd dsbb csgba bcsa 5000 * NS NS 4000 * 3000 2000 1000 0 ΔcsgBAΔbcsA ΔcsgDΔdsbBΔbcsA ΔcsgBA ΔdsbBΔcsgBA ΔcsgDΔdsbB B -1000 4000 * * NS 3500 * 3000
Supporting Information
 Supporting Information Multiplexed Instrument-Free Bar-Chart SpinChip Integrated with Nanoparticle-Mediated Magnetic Aptasensors for Visual Quantitative Detection of Multiple Pathogens Xiaofeng Wei, Wan
Supporting Information Multiplexed Instrument-Free Bar-Chart SpinChip Integrated with Nanoparticle-Mediated Magnetic Aptasensors for Visual Quantitative Detection of Multiple Pathogens Xiaofeng Wei, Wan
Supplementary Figures
 Supplementary Figures Supplementary Fig. 1 Characterization of GSCs. a. Immunostaining of primary GSC spheres from GSC lines. Nestin (neural progenitor marker, red), TLX (green). Merged images of nestin,
Supplementary Figures Supplementary Fig. 1 Characterization of GSCs. a. Immunostaining of primary GSC spheres from GSC lines. Nestin (neural progenitor marker, red), TLX (green). Merged images of nestin,
A green method of staining DNA in polyacrylamide gel electrophoresis based on fluorescent copper nanoclusters synthesized in situ
 Electronic Supplementary Material A green method of staining DNA in polyacrylamide gel electrophoresis based on fluorescent copper nanoclusters synthesized in situ Xiaoli Zhu 1, Hai Shi 1, Yalan Shen 1,
Electronic Supplementary Material A green method of staining DNA in polyacrylamide gel electrophoresis based on fluorescent copper nanoclusters synthesized in situ Xiaoli Zhu 1, Hai Shi 1, Yalan Shen 1,
Overexpression Normal expression Overexpression Normal expression. 26 (21.1%) N (%) P-value a N (%)
 SUPPLEMENTARY TABLES Table S1. Alteration of ZNF322A protein expression levels in relation to clinicopathological parameters in 123 Asian and 74 Caucasian lung cancer patients. Asian patients Caucasian
SUPPLEMENTARY TABLES Table S1. Alteration of ZNF322A protein expression levels in relation to clinicopathological parameters in 123 Asian and 74 Caucasian lung cancer patients. Asian patients Caucasian
SUPPORTING INFORMATION FILE
 Intrinsic and extrinsic connections of Tet3 dioxygenase with CXXC zinc finger modules Nan Liu, Mengxi Wang, Wen Deng, Christine S. Schmidt, Weihua Qin, Heinrich Leonhardt and Fabio Spada Department of
Intrinsic and extrinsic connections of Tet3 dioxygenase with CXXC zinc finger modules Nan Liu, Mengxi Wang, Wen Deng, Christine S. Schmidt, Weihua Qin, Heinrich Leonhardt and Fabio Spada Department of
DNA sentences. How are proteins coded for by DNA? Materials. Teacher instructions. Student instructions. Reflection
 DNA sentences How are proteins coded for by DNA? Deoxyribonucleic acid (DNA) is the molecule of life. DNA is one of the most recognizable nucleic acids, a double-stranded helix. The process by which DNA
DNA sentences How are proteins coded for by DNA? Deoxyribonucleic acid (DNA) is the molecule of life. DNA is one of the most recognizable nucleic acids, a double-stranded helix. The process by which DNA
NESTED Sequence-based Typing (SBT) protocol for epidemiological typing of Legionella pneumophila directly from clinical samples
 NESTED Sequence-based Typing (SBT) protocol for epidemiological typing of Legionella pneumophila directly from clinical samples VERSION 2.0 SUMMARY This procedure describes the use of nested Sequence-Based
NESTED Sequence-based Typing (SBT) protocol for epidemiological typing of Legionella pneumophila directly from clinical samples VERSION 2.0 SUMMARY This procedure describes the use of nested Sequence-Based
Det matematisk-naturvitenskapelige fakultet
 UNIVERSITETET I OSLO Det matematisk-naturvitenskapelige fakultet Exam in: MBV4010 Arbeidsmetoder i molekylærbiologi og biokjemi I MBV4010 Methods in molecular biology and biochemistry I Day of exam: Friday
UNIVERSITETET I OSLO Det matematisk-naturvitenskapelige fakultet Exam in: MBV4010 Arbeidsmetoder i molekylærbiologi og biokjemi I MBV4010 Methods in molecular biology and biochemistry I Day of exam: Friday
G+C content. 1 Introduction. 2 Chromosomes Topology & Counts. 3 Genome size. 4 Replichores and gene orientation. 5 Chirochores.
 1 Introduction 2 Chromosomes Topology & Counts 3 Genome size 4 Replichores and gene orientation 5 Chirochores 6 7 Codon usage 121 marc.bailly-bechet@univ-lyon1.fr Bacterial genome structures Introduction
1 Introduction 2 Chromosomes Topology & Counts 3 Genome size 4 Replichores and gene orientation 5 Chirochores 6 7 Codon usage 121 marc.bailly-bechet@univ-lyon1.fr Bacterial genome structures Introduction
11th Meeting of the Science Working Group. Lima, Peru, October 2012 SWG-11-JM-11
 11th Meeting of the Science Working Group Lima, Peru, 15-19 October 2012 Russian population genetics studies of jack mackerel in the South Pacific P.K.Afanasiev M.A.Rabchun A.I.Glubokov Introduction. In
11th Meeting of the Science Working Group Lima, Peru, 15-19 October 2012 Russian population genetics studies of jack mackerel in the South Pacific P.K.Afanasiev M.A.Rabchun A.I.Glubokov Introduction. In
Primer Design Workshop. École d'été en géné-que des champignons 2012 Dr. Will Hintz University of Victoria
 Primer Design Workshop École d'été en géné-que des champignons 2012 Dr. Will Hintz University of Victoria Scenario You have discovered the presence of a novel endophy5c organism living inside the cells
Primer Design Workshop École d'été en géné-que des champignons 2012 Dr. Will Hintz University of Victoria Scenario You have discovered the presence of a novel endophy5c organism living inside the cells
DNA Computing Circuits Using Libraries of. DNAzyme Subunits
 SUPPLEMENTARY INFORMATION Supplementary Information for the paper: DNA Computing Circuits Using Libraries of DNAzyme Subunits Johann Elbaz a, Oleg lioubashevski a, Fuan Wang a, Françoise Remacle b, Raphael
SUPPLEMENTARY INFORMATION Supplementary Information for the paper: DNA Computing Circuits Using Libraries of DNAzyme Subunits Johann Elbaz a, Oleg lioubashevski a, Fuan Wang a, Françoise Remacle b, Raphael
Circular bivalent aptamers enable in vivo stability and recognition
 Supporting Information Circular bivalent aptamers enable in vivo stability and recognition Hailan Kuai,, Zilong Zhao,, Liuting Mo, Hui Liu, Xiaoxiao Hu, Ting Fu, Xiaobing Zhang, Weihong Tan*,, Molecular
Supporting Information Circular bivalent aptamers enable in vivo stability and recognition Hailan Kuai,, Zilong Zhao,, Liuting Mo, Hui Liu, Xiaoxiao Hu, Ting Fu, Xiaobing Zhang, Weihong Tan*,, Molecular
Supplementary Figure 1. Localization of MST1 in RPE cells. Proliferating or ciliated HA- MST1 expressing RPE cells (see Fig. 5b for establishment of
 Supplementary Figure 1. Localization of MST1 in RPE cells. Proliferating or ciliated HA- MST1 expressing RPE cells (see Fig. 5b for establishment of the cell line) were immunostained for HA, acetylated
Supplementary Figure 1. Localization of MST1 in RPE cells. Proliferating or ciliated HA- MST1 expressing RPE cells (see Fig. 5b for establishment of the cell line) were immunostained for HA, acetylated
Wet Lab Tutorial: Genelet Circuits
 Wet Lab Tutorial: Genelet Circuits DNA 17 This tutorial will introduce the in vitro transcriptional circuits toolkit. The tutorial will focus on the design, synthesis, and experimental testing of a synthetic
Wet Lab Tutorial: Genelet Circuits DNA 17 This tutorial will introduce the in vitro transcriptional circuits toolkit. The tutorial will focus on the design, synthesis, and experimental testing of a synthetic
Electronic Supplementary Information
 Electronic Supplementary Information Highly sensitive detection of telomerase using telomeres-triggered isothermal exponential amplificationbased DNAzyme biosensor Yan Zhang, Li-juan Wang, and Chun-yang
Electronic Supplementary Information Highly sensitive detection of telomerase using telomeres-triggered isothermal exponential amplificationbased DNAzyme biosensor Yan Zhang, Li-juan Wang, and Chun-yang
Electronic Supplementary Information. Transcription Monitoring Using Fused RNA with a Dye-Binding Light-Up Aptamer as a Tag: A Blue Fluorescent RNA
 Electronic Supplementary Information for Transcription Monitoring Using Fused RNA with a Dye-Binding Light-Up Aptamer as a Tag: A Blue Fluorescent RNA Shinsuke Sando,* Atsushi Narita, Masayoshi Hayami,
Electronic Supplementary Information for Transcription Monitoring Using Fused RNA with a Dye-Binding Light-Up Aptamer as a Tag: A Blue Fluorescent RNA Shinsuke Sando,* Atsushi Narita, Masayoshi Hayami,
Electronic supplementary information (ESI) Kinetic Study of DNA Hybridization on DNA-modified Gold Nanoparticles. with Engineered Nano-Interfaces
 Electronic Supplementary Material (ESI) for RSC Advances. This journal is The Royal Society of Chemistry 2015 Electronic supplementary information (ESI) Kinetic Study of DNA Hybridization on DNA-modified
Electronic Supplementary Material (ESI) for RSC Advances. This journal is The Royal Society of Chemistry 2015 Electronic supplementary information (ESI) Kinetic Study of DNA Hybridization on DNA-modified
Supplementary Methods Quantitative RT-PCR. For mrna, total RNA was prepared using TRIzol reagent (Invitrogen) and genomic DNA was eliminated with TURB
 Supplementary Methods Quantitative RT-PCR. For mrna, total RNA was prepared using TRIzol reagent (Invitrogen) and genomic DNA was eliminated with TURBO DNA-free Kit (Ambion). One µg of total RNA was reverse
Supplementary Methods Quantitative RT-PCR. For mrna, total RNA was prepared using TRIzol reagent (Invitrogen) and genomic DNA was eliminated with TURBO DNA-free Kit (Ambion). One µg of total RNA was reverse
Lecture 11: Gene Prediction
 Lecture 11: Gene Prediction Study Chapter 6.11-6.14 1 Gene: A sequence of nucleotides coding for protein Gene Prediction Problem: Determine the beginning and end positions of genes in a genome Where are
Lecture 11: Gene Prediction Study Chapter 6.11-6.14 1 Gene: A sequence of nucleotides coding for protein Gene Prediction Problem: Determine the beginning and end positions of genes in a genome Where are
Supporting Information. Electrochemiluminescence Peptide-Based Biosensor with Hetero-
 Supporting Information Electrochemiluminescence Peptide-Based Biosensor with Hetero- Nanostructures as Co-reaction Accelerator for the Ultra-sensitive Determination of Tryptase Fang-Fang Wu, Ying Zhou,
Supporting Information Electrochemiluminescence Peptide-Based Biosensor with Hetero- Nanostructures as Co-reaction Accelerator for the Ultra-sensitive Determination of Tryptase Fang-Fang Wu, Ying Zhou,
2
 1 2 3 4 5 6 7 Supplemental Table 1. Magnaporthe oryzae strains generated in this study. Strain background Genotype Strain name Description Guy-11 H1:RFP H1:RFP Strain expressing Histone H1- encoding gene
1 2 3 4 5 6 7 Supplemental Table 1. Magnaporthe oryzae strains generated in this study. Strain background Genotype Strain name Description Guy-11 H1:RFP H1:RFP Strain expressing Histone H1- encoding gene
SCIENTIFIC REPORT. regarding the implementation of the project between December December 2014
 SCIENTIFIC REPORT regarding the implementation of the project between December 013 - December 014 Title of the project: DETECTION AND IDENTIFICATION OF BIOMOLECULES OF MEDICAL INTEREST BY USING MAGNETIC
SCIENTIFIC REPORT regarding the implementation of the project between December 013 - December 014 Title of the project: DETECTION AND IDENTIFICATION OF BIOMOLECULES OF MEDICAL INTEREST BY USING MAGNETIC
A netlike rolling circle nucleic acid amplification technique
 Electronic Supplementary Material (ESI) for Analyst. This journal is The Royal Society of Chemistry 2014 Supplementary Information A netlike rolling circle nucleic acid amplification technique Xiaoli Zhu,
Electronic Supplementary Material (ESI) for Analyst. This journal is The Royal Society of Chemistry 2014 Supplementary Information A netlike rolling circle nucleic acid amplification technique Xiaoli Zhu,
Supplemental Information. Target-Mediated Protection of Endogenous. MicroRNAs in C. elegans. Inventory of Supplementary Information
 Developmental Cell, Volume 20 Supplemental Information Target-Mediated Protection of Endogenous MicroRNAs in C. elegans Saibal Chatterjee, Monika Fasler, Ingo Büssing, and Helge Großhans Inventory of Supplementary
Developmental Cell, Volume 20 Supplemental Information Target-Mediated Protection of Endogenous MicroRNAs in C. elegans Saibal Chatterjee, Monika Fasler, Ingo Büssing, and Helge Großhans Inventory of Supplementary
Supplemental Table 1. Primers used for PCR.
 Supplemental Table 1. Primers used for PCR. Gene Type Primer Sequence Genotyping and semi-quantitative RT-PCR F 5 -TTG CCC GAT CAC CAT CTG TA-3 rwa1-1 R 5 -TGT AGC GAT CAA GGC CTG ATC TAA-3 LB 5 -TAG CAT
Supplemental Table 1. Primers used for PCR. Gene Type Primer Sequence Genotyping and semi-quantitative RT-PCR F 5 -TTG CCC GAT CAC CAT CTG TA-3 rwa1-1 R 5 -TGT AGC GAT CAA GGC CTG ATC TAA-3 LB 5 -TAG CAT
Homework. A bit about the nature of the atoms of interest. Project. The role of electronega<vity
 Homework Why cited articles are especially useful. citeulike science citation index When cutting and pasting less is more. Project Your protein: I will mail these out this weekend If you haven t gotten
Homework Why cited articles are especially useful. citeulike science citation index When cutting and pasting less is more. Project Your protein: I will mail these out this weekend If you haven t gotten
for Programmed Chemo-enzymatic Synthesis of Antigenic Oligosaccharides
 Supporting Information Design of α-transglucosidases of Controlled Specificity for Programmed Chemo-enzymatic Synthesis of Antigenic Oligosaccharides Elise Champion ±,,,, Isabelle André ±,,, Claire Moulis
Supporting Information Design of α-transglucosidases of Controlled Specificity for Programmed Chemo-enzymatic Synthesis of Antigenic Oligosaccharides Elise Champion ±,,,, Isabelle André ±,,, Claire Moulis
