Lecture 11: Gene Prediction
|
|
|
- Silas Henderson
- 6 years ago
- Views:
Transcription
1 Lecture 11: Gene Prediction Study Chapter
2 Gene: A sequence of nucleotides coding for protein Gene Prediction Problem: Determine the beginning and end positions of genes in a genome
3 Where are the genes in the genome? aatgcatgcggctatgctaatgcatgcggctatgctaagctgggatccgatgacaatgcatgcggctatgcta atgcatgcggctatgcaagctgggatccgatgactatgctaagctgggatccgatgacaatgcatgcggcta tgctaatgaatggtcttgggatttaccttggaatgctaagctgggatccgatgacaatgcatgcggctatgcta atgaatggtcttgggatttaccttggaatatgctaatgcatgcggctatgctaagctgggatccgatgacaatg catgcggctatgctaatgcatgcggctatgcaagctgggatccgatgactatgctaagctgcggctatgctaa tgcatgcggctatgctaagctgggatccgatgacaatgcatgcggctatgctaatgcatgcggctatgcaag ctgggatcctgcggctatgctaatgaatggtcttgggatttaccttggaatgctaagctgggatccgatgaca Gene! atgcatgcggctatgctaatgaatggtcttgggatttaccttggaatatgctaatgcatgcggctatgctaagc tgggaatgcatgcggctatgctaagctgggatccgatgacaatgcatgcggctatgctaatgcatgcggcta tgcaagctgggatccgatgactatgctaagctgcggctatgctaatgcatgcggctatgctaagctcatgcgg ctatgctaagctgggaatgcatgcggctatgctaagctgggatccgatgacaatgcatgcggctatgctaat gcatgcggctatgcaagctgggatccgatgactatgctaagctgcggctatgctaatgcatgcggctatgcta agctcggctatgctaatgaatggtcttgggatttaccttggaatgctaagctgggatccgatgacaatgcatg cggctatgctaatgaatggtcttgggatttaccttggaatatgctaatgcatgcggctatgctaagctgggaat gcatgcggctatgctaagctgggatccgatgacaatgcatgcggctatgctaatgcatgcggctatgcaagc tgggatccgatgactatgctaagctgcggctatgctaatgcatgcggctatgctaagctcatgcgg 3
4 Exons and Introns In prokaryotes, one-cell organisms without a cell nucleus (e.g. bacteria), a gene is a single sequence In eukaryotes, the gene is a combination of coding segments (exons) that are interrupted by non-coding segments (introns) This makes computational gene prediction in eukaryotes even more difficult 5 cap intron1 intron2 exon1 exon2 exon3 mrna transcription splicing translation 3 poly-a tail What are the hints that a gene is nearby? 4
5 Splicing Signals Exon 1 GT AG Exon 2 GT AG Exon 3 The genome provides subtle hints where exon/intron boundaries might occur Dinucleotide sequences GT and AG flank an exon >95% of the time With one or two additional flanking sequences cover 99.9% of the cases How helpful is this? 5
6 Two Approaches to Gene Prediction Statistical: coding segments (exons) have typical sequences on either end and use different subwords than non-coding segments (introns). Similarity-based: many human genes are similar to genes in mice, chicken, or even bacteria. Therefore, already known mouse, chicken, and bacterial genes may help to find human genes. 6
7 Genetic Code and Stop Codons TAA, TAG and TGA correspond to 3 Stop codons that (together with Start codon ATG) delineate Open Reading Frames 7
8 Six Frames in a DNA Sequence CTGCAGACGAAACCTCTTGATGTAGTTGGCCTGACACCGACAATAATGAAGACTACCGTCTTACTAACAC CTGCAGACGAAACCTCTTGATGTAGTTGGCCTGACACCGACAATAATGAAGACTACCGTCTTACTAACAC CTGCAGACGAAACCTCTTGATGTAGTTGGCCTGACACCGACAATAATGAAGACTACCGTCTTACTAACAC CTGCAGACGAAACCTCTTGATGTAGTTGGCCTGACACCGACAATAATGAAGACTACCGTCTTACTAACAC GACGTCTGCTTTGGAGAACTACATCAACCGGACTGTGGCTGTTATTACTTCTGATGGCAGAATGATTGTG GACGTCTGCTTTGGAGAACTACATCAACCGGACTGTGGCTGTTATTACTTCTGATGGCAGAATGATTGTG GACGTCTGCTTTGGAGAACTACATCAACCGGACTGTGGCTGTTATTACTTCTGATGGCAGAATGATTGTG GACGTCTGCTTTGGAGAACTACATCAACCGGACTGTGGCTGTTATTACTTCTGATGGCAGAATGATTGTG stop codons TAA, TAG, TGA start codons - ATG 8
9 Open Reading Frames (ORFs) Detect potential coding regions by looking at ORFs A genome of length n is comprised of (n/3) codons Stop codons break genome into segments between consecutive Stop codons The subsegments of these that start from the Start codon (ATG) are ORFs ORFs in different frames may overlap ATG TGA Genomic Sequence Open reading frame 9
10 Long versus Short ORFs Long open reading frames may be a gene At random, we should expect one stop codon every (64/3) ~= 21 codons However, genes are usually much longer than this A basic approach is to scan for ORFs whose length exceeds certain threshold This is an incomplete strategy because some genes (e.g. some neural and immune system genes) are relatively short 10
11 Testing ORFs: Codon Usage Create a 64-element hash table and count the frequencies of codons in an ORF Amino acids typically have more than one codon, but in nature certain codons are more in use Uneven use of the codons may characterize a real gene This compensates for pitfalls of the ORF length test 11
12 Codon Usage in the Human Genome T C A G TTT Phe 46 TCT Ser 19 TAT Tyr 44 TGT Cys 46 T C A G TTC Phe 54 TCC Ser 22 TAC Tyr 56 TGC Cys 54 TTA Leu 8 TCA Ser 15 TAA Stop 30 TGA Stop 47 TTG Leu 13 TCG Ser 5 TAG Stop 24 TGG Trp 100 CTT Leu 13 CCT Pro 29 CAT His 42 CGT Arg 8 CTC Leu 20 CCC Pro 32 CAC His 58 CGC Arg 18 CTA Leu 7 CCA Pro 28 CAA Gln 27 CGA Arg 11 CTG Leu 40 CCG Pro 11 CAG Gln 73 CGG Arg 20 ATT Ile 36 ACT Thr 25 AAT Asn 47 AGT Ser 15 ATC Ile 47 ACC Thr 36 AAC Asn 53 AGC Ser 24 ATA Ile 17 ACA Thr 28 AAA Lys 43 AGA Arg 21 ATG Met 100 ACG Thr 11 AAG Lys 57 AGG Arg 21 GTT Val 18 GCT Ala 27 GAT Asp 46 GGT Gly 16 GTC Val 24 GCC Ala 40 GAC Asp 54 GGC Gly 34 GTA Val 12 GCA Ala 23 GAA Glu 42 GGA Gly 25 GTG Val 46 GCG Ala 11 GAG Glu 58 GGG Gly 25 12
13 Codon Usage in the Mouse Genome T C A G TTT Phe 39 TCT Ser 16 TAT Tyr 41 TGT Cys 45 T C A G TTC Phe 61 TCC Ser 21 TAC Tyr 59 TGC Cys 55 TTA Leu 7 TCA Ser 21 TAA Stop 27 TGA Stop 27 TTG Leu 10 TCG Ser 6 TAG Stop 45 TGG Trp 100 CTT Leu 13 CCT Pro 26 CAT His 35 CGT Arg 6 CTC Leu 21 CCC Pro 30 CAC His 65 CGC Arg 13 CTA Leu 14 CCA Pro 34 CAA Gln 24 CGA Arg 10 CTG Leu 35 CCG Pro 10 CAG Gln 76 CGG Arg 13 ATT Ile 27 ACT Thr 24 AAT Asn 41 AGT Ser 13 ATC Ile 43 ACC Thr 28 AAC Asn 59 AGC Ser 23 ATA Ile 29 ACA Thr 40 AAA Lys 58 AGA Arg 34 ATG Met 100 ACG Thr 8 AAG Lys 42 AGG Arg 25 GTT Val 14 GCT Ala 23 GAT Asp 39 GGT Gly 17 GTC Val 24 GCC Ala 36 GAC Asp 61 GGC Gly 32 GTA Val 19 GCA Ala 30 GAA Glu 49 GGA Gly 29 GTG Val 43 GCG Ala 11 GAG Glu 51 GGG Gly 15 13
14 Codon Usage and Likelihood Ratio An ORF is more believable than another if it has more likely codons Do sliding window calculations to find ORFs that have the likely codon usage Allows for higher precision in identifying true ORFs; much better than merely testing for length. However, average vertebrate exon length is 130 nucleotides, which is often too small to produce reliable peaks in the likelihood ratio Further improvement: in-frame hexamer count (frequencies of pairs of consecutive codons) 14
15 Similarity-Based Approach Genes in different organisms are similar The similarity-based approach uses known genes in one genome to predict (unknown) genes in another genome Problem: Given a known gene and an unannotated genome sequence, find a set of substrings of the genomic sequence whose concatenation best fits the gene An alignment problem! 15
16 Chaining Local Alignments Locate codon substrings that match a given protein subsequence, putative (candidate) exons Define a putative exon as a 3-tuple (l, r, w) l = left starting position r = right ending position w = weight based on some scoring function w(# of amino acid s matched, codon freq, ) Look for a maximum chain of substrings Chain: a set of non-overlapping nonadjacent intervals. 16
17 Exon Chaining Problem Locate the beginning and end of each interval (2n points) Find the best path 17
18 Exon Chaining Problem: Formulation Exon Chaining Problem: Given a set of putative exons, find a maximum set of nonoverlapping putative exons Input: a set of weighted intervals (putative exons) Output: A maximum chain of intervals from this set Would a greedy algorithm solve this problem? 18
19 Exon Chaining Problem: Graph Representation A greedy solution takes the highest scoring chain first, followed by largest one left in the uncovered range, and so on until none can be 21 taken, the score is the sum of all taken chains A dynamic programming solution can do better in O(n) time, by constructing a graph 19
20 Exon Chaining Algorithm ExonChaining (G, n) //Graph, number of intervals 1 for i to 2n 2 s i 0 3 for i 1 to 2n 4 if vertex v i in G corresponds to right end of the interval I 5 j index of vertex for left end of the interval I 6 w weight of the interval I 7 s j max {s j + w, s i-1 } 8 else 9 s i s i-1 10 return s 2n 20
21 Exon Chaining: Deficiencies Poor definition of the putative exon endpoints Optimal interval chain may not correspond to a valid alignment We enforce genomic order but not protein order First interval may correspond to a suffix, whereas second interval may correspond to a prefix Combination of such intervals is not a valid alignment 21
22 Spliced Alignment Mikhail Gelfand and colleagues proposed a spliced alignment approach of using a protein within one genome to reconstruct the exon-intron structure of a (related) gene in another genome. Begins by selecting either all putative exons between potential acceptor and donor sites or by finding all substrings similar to the target protein (as in the Exon Chaining Problem). This set is further filtered in a such a way that attempt to retain all true exons, but allow some false ones. (Many false positives, but no false negatives) 22
23 Spliced Alignment Problem: Formulation Goal: Find a chain of blocks in a genomic sequence that best fits a target sequence Input: Genomic sequences G, target sequence T, and a set of candidate exons B. Output: A chain of exons Γ such that the global alignment score between Γ* and T is maximum among all chains of blocks from B. Γ* - concatenation of all exons from chain Γ 23
24 Lewis Carroll Example 24
25 Lewis Carroll Example 25
26 Lewis Carroll Example 26
27 Exon Chaining vs Spliced Alignment In Spliced Alignment, every path spells out a string obtained by concatenating the labels of its vertices. The overall path gives an optimal alignment score between concatenated labels (blocks) and target sequence Defines weight as the sum of vertex weights rather than as the sum of edge weights as in Spliced Alignment Exon Chaining assumes the positions and weights of exons are pre-defined How to solve using Dynamic Programming? 27
28 Spliced Alignment: Idea Compute the best alignment between i-prefix of genomic sequence G and j-prefix of target T: S(i,j) But what is i-prefix of G? There may be a few i-prefixes of G depending on which block B we are in. Compute the best alignment between i-prefix of genomic sequence G and j-prefix of target T under the assumption that the alignment uses the block B at position i, S(i,j,B) Two cases to consider, block B starts at i, or it does not 28
29 Spliced Alignment Recurrence If i is not the starting vertex of block B: S(i 1, j,b) σ S(i, j,b) = max S(i, j 1,B) σ S(i 1, j 1,B) +δ(g i,t j ) Recall σ was the cost of an indel If i is the starting vertex of block B: S(i 1, j,b) σ S(i, j,b) = max max all blocks B' preceding B S(end(B'), j 1,B') δ(g i,t j ) max all blocks B' preceding B S(end(B'), j,b') σ 29
30 Spliced Alignment Solution After computing the three-dimensional table S(i, j, B), the score of the optimal spliced alignment is: max all blocks B S(end(B), length(t), B) 30
31 A Modern Approach Sequence novel transcripts Identify gene(s) with similar protein Align transcript sequence to genome Locus of alignment identifies location and intron/exon structure 34
Disease and selection in the human genome 3
 Disease and selection in the human genome 3 Ka/Ks revisited Please sit in row K or forward RBFD: human populations, adaptation and immunity Neandertal Museum, Mettman Germany Sequence genome Measure expression
Disease and selection in the human genome 3 Ka/Ks revisited Please sit in row K or forward RBFD: human populations, adaptation and immunity Neandertal Museum, Mettman Germany Sequence genome Measure expression
Lecture 10, 20/2/2002: The process of solution development - The CODEHOP strategy for automatic design of consensus-degenerate primers for PCR
 Lecture 10, 20/2/2002: The process of solution development - The CODEHOP strategy for automatic design of consensus-degenerate primers for PCR 1 The problem We wish to clone a yet unknown gene from a known
Lecture 10, 20/2/2002: The process of solution development - The CODEHOP strategy for automatic design of consensus-degenerate primers for PCR 1 The problem We wish to clone a yet unknown gene from a known
NAME:... MODEL ANSWER... STUDENT NUMBER:... Maximum marks: 50. Internal Examiner: Hugh Murrell, Computer Science, UKZN
 COMP710, Bioinformatics with Julia, Test One, Thursday the 20 th of April, 2017, 09h30-11h30 1 NAME:...... MODEL ANSWER... STUDENT NUMBER:...... Maximum marks: 50 Internal Examiner: Hugh Murrell, Computer
COMP710, Bioinformatics with Julia, Test One, Thursday the 20 th of April, 2017, 09h30-11h30 1 NAME:...... MODEL ANSWER... STUDENT NUMBER:...... Maximum marks: 50 Internal Examiner: Hugh Murrell, Computer
ORFs and genes. Please sit in row K or forward
 ORFs and genes Please sit in row K or forward https://www.flickr.com/photos/teseum/3231682806/in/photostream/ Question: why do some strains of Vibrio cause cholera and others don t? Methods Mechanisms
ORFs and genes Please sit in row K or forward https://www.flickr.com/photos/teseum/3231682806/in/photostream/ Question: why do some strains of Vibrio cause cholera and others don t? Methods Mechanisms
Materials Protein synthesis kit. This kit consists of 24 amino acids, 24 transfer RNAs, four messenger RNAs and one ribosome (see below).
 Protein Synthesis Instructions The purpose of today s lab is to: Understand how a cell manufactures proteins from amino acids, using information stored in the genetic code. Assemble models of four very
Protein Synthesis Instructions The purpose of today s lab is to: Understand how a cell manufactures proteins from amino acids, using information stored in the genetic code. Assemble models of four very
Lecture 19A. DNA computing
 Lecture 19A. DNA computing What exactly is DNA (deoxyribonucleic acid)? DNA is the material that contains codes for the many physical characteristics of every living creature. Your cells use different
Lecture 19A. DNA computing What exactly is DNA (deoxyribonucleic acid)? DNA is the material that contains codes for the many physical characteristics of every living creature. Your cells use different
Homework. A bit about the nature of the atoms of interest. Project. The role of electronega<vity
 Homework Why cited articles are especially useful. citeulike science citation index When cutting and pasting less is more. Project Your protein: I will mail these out this weekend If you haven t gotten
Homework Why cited articles are especially useful. citeulike science citation index When cutting and pasting less is more. Project Your protein: I will mail these out this weekend If you haven t gotten
G+C content. 1 Introduction. 2 Chromosomes Topology & Counts. 3 Genome size. 4 Replichores and gene orientation. 5 Chirochores.
 1 Introduction 2 Chromosomes Topology & Counts 3 Genome size 4 Replichores and gene orientation 5 Chirochores 6 7 Codon usage 121 marc.bailly-bechet@univ-lyon1.fr Bacterial genome structures Introduction
1 Introduction 2 Chromosomes Topology & Counts 3 Genome size 4 Replichores and gene orientation 5 Chirochores 6 7 Codon usage 121 marc.bailly-bechet@univ-lyon1.fr Bacterial genome structures Introduction
Codon Bias with PRISM. 2IM24/25, Fall 2007
 Codon Bias with PRISM 2IM24/25, Fall 2007 from RNA to protein mrna vs. trna aminoacid trna anticodon mrna codon codon-anticodon matching Watson-Crick base pairing A U and C G binding first two nucleotide
Codon Bias with PRISM 2IM24/25, Fall 2007 from RNA to protein mrna vs. trna aminoacid trna anticodon mrna codon codon-anticodon matching Watson-Crick base pairing A U and C G binding first two nucleotide
Primer Design Workshop. École d'été en géné-que des champignons 2012 Dr. Will Hintz University of Victoria
 Primer Design Workshop École d'été en géné-que des champignons 2012 Dr. Will Hintz University of Victoria Scenario You have discovered the presence of a novel endophy5c organism living inside the cells
Primer Design Workshop École d'été en géné-que des champignons 2012 Dr. Will Hintz University of Victoria Scenario You have discovered the presence of a novel endophy5c organism living inside the cells
Protein Structure Analysis
 BINF 731 Protein Structure Analysis http://binf.gmu.edu/vaisman/binf731/ Iosif Vaisman COMPUTATIONAL BIOLOGY COMPUTATIONAL STRUCTURAL BIOLOGY COMPUTATIONAL MOLECULAR BIOLOGY BIOINFORMATICS STRUCTURAL BIOINFORMATICS
BINF 731 Protein Structure Analysis http://binf.gmu.edu/vaisman/binf731/ Iosif Vaisman COMPUTATIONAL BIOLOGY COMPUTATIONAL STRUCTURAL BIOLOGY COMPUTATIONAL MOLECULAR BIOLOGY BIOINFORMATICS STRUCTURAL BIOINFORMATICS
Figure S1. Characterization of the irx9l-1 mutant. (A) Diagram of the Arabidopsis IRX9L gene drawn based on information from TAIR (the Arabidopsis
 1 2 3 4 5 6 7 8 9 10 11 12 Figure S1. Characterization of the irx9l-1 mutant. (A) Diagram of the Arabidopsis IRX9L gene drawn based on information from TAIR (the Arabidopsis Information Research). Exons
1 2 3 4 5 6 7 8 9 10 11 12 Figure S1. Characterization of the irx9l-1 mutant. (A) Diagram of the Arabidopsis IRX9L gene drawn based on information from TAIR (the Arabidopsis Information Research). Exons
Project 07/111 Final Report October 31, Project Title: Cloning and expression of porcine complement C3d for enhanced vaccines
 Project 07/111 Final Report October 31, 2007. Project Title: Cloning and expression of porcine complement C3d for enhanced vaccines Project Leader: Dr Douglas C. Hodgins (519-824-4120 Ex 54758, fax 519-824-5930)
Project 07/111 Final Report October 31, 2007. Project Title: Cloning and expression of porcine complement C3d for enhanced vaccines Project Leader: Dr Douglas C. Hodgins (519-824-4120 Ex 54758, fax 519-824-5930)
PGRP negatively regulates NOD-mediated cytokine production in rainbow trout liver cells
 Supplementary Information for: PGRP negatively regulates NOD-mediated cytokine production in rainbow trout liver cells Ju Hye Jang 1, Hyun Kim 2, Mi Jung Jang 2, Ju Hyun Cho 1,2,* 1 Research Institute
Supplementary Information for: PGRP negatively regulates NOD-mediated cytokine production in rainbow trout liver cells Ju Hye Jang 1, Hyun Kim 2, Mi Jung Jang 2, Ju Hyun Cho 1,2,* 1 Research Institute
Supporting information for Biochemistry, 1995, 34(34), , DOI: /bi00034a013
 Supporting information for Biochemistry, 1995, 34(34), 10807 10815, DOI: 10.1021/bi00034a013 LESNIK 10807-1081 Terms & Conditions Electronic Supporting Information files are available without a subscription
Supporting information for Biochemistry, 1995, 34(34), 10807 10815, DOI: 10.1021/bi00034a013 LESNIK 10807-1081 Terms & Conditions Electronic Supporting Information files are available without a subscription
for Programmed Chemo-enzymatic Synthesis of Antigenic Oligosaccharides
 Supporting Information Design of α-transglucosidases of Controlled Specificity for Programmed Chemo-enzymatic Synthesis of Antigenic Oligosaccharides Elise Champion ±,,,, Isabelle André ±,,, Claire Moulis
Supporting Information Design of α-transglucosidases of Controlled Specificity for Programmed Chemo-enzymatic Synthesis of Antigenic Oligosaccharides Elise Champion ±,,,, Isabelle André ±,,, Claire Moulis
National PHL TB DST Reference Center PSQ Reporting Language Table of Contents
 PSQ Reporting Language Table of Contents Document Page Number PSQ for Rifampin 2-6 Comparison table for rpob Codon Numbering 2 rpob mutation list (new numbering system) 3-5 rpob interpretations 6 PSQ for
PSQ Reporting Language Table of Contents Document Page Number PSQ for Rifampin 2-6 Comparison table for rpob Codon Numbering 2 rpob mutation list (new numbering system) 3-5 rpob interpretations 6 PSQ for
Electronic Supplementary Information
 Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2017 Electronic Supplementary Information Dissecting binding of a β-barrel outer membrane
Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2017 Electronic Supplementary Information Dissecting binding of a β-barrel outer membrane
Det matematisk-naturvitenskapelige fakultet
 UNIVERSITETET I OSLO Det matematisk-naturvitenskapelige fakultet Exam in: MBV4010 Arbeidsmetoder i molekylærbiologi og biokjemi I MBV4010 Methods in molecular biology and biochemistry I Day of exam: Friday
UNIVERSITETET I OSLO Det matematisk-naturvitenskapelige fakultet Exam in: MBV4010 Arbeidsmetoder i molekylærbiologi og biokjemi I MBV4010 Methods in molecular biology and biochemistry I Day of exam: Friday
Supplementary. Table 1: Oligonucleotides and Plasmids. complementary to positions from 77 of the SRα '- GCT CTA GAG AAC TTG AAG TAC AGA CTG C
 Supplementary Table 1: Oligonucleotides and Plasmids 913954 5'- GCT CTA GAG AAC TTG AAG TAC AGA CTG C 913955 5'- CCC AAG CTT ACA GTG TGG CCA TTC TGC TG 223396 5'- CGA CGC GTA CAG TGT GGC CAT TCT GCT G
Supplementary Table 1: Oligonucleotides and Plasmids 913954 5'- GCT CTA GAG AAC TTG AAG TAC AGA CTG C 913955 5'- CCC AAG CTT ACA GTG TGG CCA TTC TGC TG 223396 5'- CGA CGC GTA CAG TGT GGC CAT TCT GCT G
Genomic Sequence Analysis using Electron-Ion Interaction
 University of Aizu, Graduation Thesis. March, 25 s1985 1 Genomic Sequence Analysis using Electron-Ion Interaction Potential Masumi Kobayashi s1985 Supervised by Hiroshi Toyoizumi Abstract This paper proposes
University of Aizu, Graduation Thesis. March, 25 s1985 1 Genomic Sequence Analysis using Electron-Ion Interaction Potential Masumi Kobayashi s1985 Supervised by Hiroshi Toyoizumi Abstract This paper proposes
Hes6. PPARα. PPARγ HNF4 CD36
 SUPPLEMENTARY INFORMATION Supplementary Table Positions and Sequences of ChIP primers -63 AGGTCACTGCCA -79 AGGTCTGCTGTG Hes6-0067 GGGCAaAGTTCA ACOT -395 GGGGCAgAGTTCA PPARα -309 GGCTCAaAGTTCAaGTTCA CPTa
SUPPLEMENTARY INFORMATION Supplementary Table Positions and Sequences of ChIP primers -63 AGGTCACTGCCA -79 AGGTCTGCTGTG Hes6-0067 GGGCAaAGTTCA ACOT -395 GGGGCAgAGTTCA PPARα -309 GGCTCAaAGTTCAaGTTCA CPTa
Arabidopsis actin depolymerizing factor AtADF4 mediates defense signal transduction triggered by the Pseudomonas syringae effector AvrPphB
 Arabidopsis actin depolymerizing factor mediates defense signal transduction triggered by the Pseudomonas syringae effector AvrPphB Files in this Data Supplement: Supplemental Table S1 Supplemental Table
Arabidopsis actin depolymerizing factor mediates defense signal transduction triggered by the Pseudomonas syringae effector AvrPphB Files in this Data Supplement: Supplemental Table S1 Supplemental Table
Supplemental Data Supplemental Figure 1.
 Supplemental Data Supplemental Figure 1. Silique arrangement in the wild-type, jhs, and complemented lines. Wild-type (WT) (A), the jhs1 mutant (B,C), and the jhs1 mutant complemented with JHS1 (Com) (D)
Supplemental Data Supplemental Figure 1. Silique arrangement in the wild-type, jhs, and complemented lines. Wild-type (WT) (A), the jhs1 mutant (B,C), and the jhs1 mutant complemented with JHS1 (Com) (D)
PROTEIN SYNTHESIS Study Guide
 PART A. Read the following: PROTEIN SYNTHESIS Study Guide Protein synthesis is the process used by the body to make proteins. The first step of protein synthesis is called Transcription. It occurs in the
PART A. Read the following: PROTEIN SYNTHESIS Study Guide Protein synthesis is the process used by the body to make proteins. The first step of protein synthesis is called Transcription. It occurs in the
 www.lessonplansinc.com Topic: Gene Mutations WS Summary: Students will learn about frame shift mutations and base substitution mutations. Goals & Objectives: Students will be able to demonstrate how mutations
www.lessonplansinc.com Topic: Gene Mutations WS Summary: Students will learn about frame shift mutations and base substitution mutations. Goals & Objectives: Students will be able to demonstrate how mutations
Supplementary Figure 1A A404 Cells +/- Retinoic Acid
 Supplementary Figure 1A A44 Cells +/- Retinoic Acid 1 1 H3 Lys4 di-methylation SM-actin VEC cfos (-) RA (+) RA 14 1 1 8 6 4 H3 Lys79 di-methylation SM-actin VEC cfos (-) RA (+) RA Supplementary Figure
Supplementary Figure 1A A44 Cells +/- Retinoic Acid 1 1 H3 Lys4 di-methylation SM-actin VEC cfos (-) RA (+) RA 14 1 1 8 6 4 H3 Lys79 di-methylation SM-actin VEC cfos (-) RA (+) RA Supplementary Figure
Outline. 1. Introduction. 2. Exon Chaining Problem. 3. Spliced Alignment. 4. Gene Prediction Tools
 Outline 1. Introduction 2. Exon Chaining Problem 3. Spliced Alignment 4. Gene Prediction Tools Section 1: Introduction Similarity-Based Approach to Gene Prediction Some genomes may be well-studied, with
Outline 1. Introduction 2. Exon Chaining Problem 3. Spliced Alignment 4. Gene Prediction Tools Section 1: Introduction Similarity-Based Approach to Gene Prediction Some genomes may be well-studied, with
Dierks Supplementary Fig. S1
 Dierks Supplementary Fig. S1 ITK SYK PH TH K42R wt K42R (kinase deficient) R29C E42K Y323F R29C E42K Y323F (reduced phospholipid binding) (enhanced phospholipid binding) (reduced Cbl binding) E42K Y323F
Dierks Supplementary Fig. S1 ITK SYK PH TH K42R wt K42R (kinase deficient) R29C E42K Y323F R29C E42K Y323F (reduced phospholipid binding) (enhanced phospholipid binding) (reduced Cbl binding) E42K Y323F
Supplemental Data. mir156-regulated SPL Transcription. Factors Define an Endogenous Flowering. Pathway in Arabidopsis thaliana
 Cell, Volume 138 Supplemental Data mir156-regulated SPL Transcription Factors Define an Endogenous Flowering Pathway in Arabidopsis thaliana Jia-Wei Wang, Benjamin Czech, and Detlef Weigel Table S1. Interaction
Cell, Volume 138 Supplemental Data mir156-regulated SPL Transcription Factors Define an Endogenous Flowering Pathway in Arabidopsis thaliana Jia-Wei Wang, Benjamin Czech, and Detlef Weigel Table S1. Interaction
Lezione 10. Bioinformatica. Mauro Ceccanti e Alberto Paoluzzi
 Lezione 10 Bioinformatica Mauro Ceccanti e Alberto Paoluzzi Dip. Informatica e Automazione Università Roma Tre Dip. Medicina Clinica Università La Sapienza Lezione 10: Sintesi proteica Synthesis of proteins
Lezione 10 Bioinformatica Mauro Ceccanti e Alberto Paoluzzi Dip. Informatica e Automazione Università Roma Tre Dip. Medicina Clinica Università La Sapienza Lezione 10: Sintesi proteica Synthesis of proteins
SAY IT WITH DNA: Protein Synthesis Activity by Larry Flammer
 TEACHER S GUIDE SAY IT WITH DNA: Protein Synthesis Activity by Larry Flammer SYNOPSIS This activity uses the metaphor of decoding a secret message for the Protein Synthesis process. Students teach themselves
TEACHER S GUIDE SAY IT WITH DNA: Protein Synthesis Activity by Larry Flammer SYNOPSIS This activity uses the metaphor of decoding a secret message for the Protein Synthesis process. Students teach themselves
Expression of Recombinant Proteins
 Expression of Recombinant Proteins Uses of Cloned Genes sequencing reagents (eg, probes) protein production insufficient natural quantities modify/mutagenesis library screening Expression Vector Features
Expression of Recombinant Proteins Uses of Cloned Genes sequencing reagents (eg, probes) protein production insufficient natural quantities modify/mutagenesis library screening Expression Vector Features
Introduction to Bioinformatics Dr. Robert Moss
 Introduction to Bioinformatics Dr. Robert Moss Bioinformatics is about searching biological databases, comparing sequences, looking at protein structures, and more generally, asking biological questions
Introduction to Bioinformatics Dr. Robert Moss Bioinformatics is about searching biological databases, comparing sequences, looking at protein structures, and more generally, asking biological questions
PCR analysis was performed to show the presence and the integrity of the var1csa and var-
 Supplementary information: Methods: Table S1: Primer Name Nucleotide sequence (5-3 ) DBL3-F tcc ccg cgg agt gaa aca tca tgt gac tg DBL3-R gac tag ttt ctt tca ata aat cac tcg c DBL5-F cgc cct agg tgc ttc
Supplementary information: Methods: Table S1: Primer Name Nucleotide sequence (5-3 ) DBL3-F tcc ccg cgg agt gaa aca tca tgt gac tg DBL3-R gac tag ttt ctt tca ata aat cac tcg c DBL5-F cgc cct agg tgc ttc
DNA sentences. How are proteins coded for by DNA? Materials. Teacher instructions. Student instructions. Reflection
 DNA sentences How are proteins coded for by DNA? Deoxyribonucleic acid (DNA) is the molecule of life. DNA is one of the most recognizable nucleic acids, a double-stranded helix. The process by which DNA
DNA sentences How are proteins coded for by DNA? Deoxyribonucleic acid (DNA) is the molecule of life. DNA is one of the most recognizable nucleic acids, a double-stranded helix. The process by which DNA
Supplemental Data. Bennett et al. (2010). Plant Cell /tpc
 BRN1 ---------MSSSNGGVPPGFRFHPTDEELLHYYLKKKISYEKFEMEVIKEVDLNKIEPWDLQDRCKIGSTPQNEWYFFSHKDRKYPTGS 81 BRN2 --------MGSSSNGGVPPGFRFHPTDEELLHYYLKKKISYQKFEMEVIREVDLNKLEPWDLQERCKIGSTPQNEWYFFSHKDRKYPTGS 82 SMB
BRN1 ---------MSSSNGGVPPGFRFHPTDEELLHYYLKKKISYEKFEMEVIKEVDLNKIEPWDLQDRCKIGSTPQNEWYFFSHKDRKYPTGS 81 BRN2 --------MGSSSNGGVPPGFRFHPTDEELLHYYLKKKISYQKFEMEVIREVDLNKLEPWDLQERCKIGSTPQNEWYFFSHKDRKYPTGS 82 SMB
Supporting Information
 Supporting Information Table S1. Oligonucleotide sequences used in this work Oligo DNA A B C D CpG-A CpG-B CpG-C CpG-D Sequence 5 ACA TTC CTA AGT CTG AAA CAT TAC AGC TTG CTA CAC GAG AAG AGC CGC CAT AGT
Supporting Information Table S1. Oligonucleotide sequences used in this work Oligo DNA A B C D CpG-A CpG-B CpG-C CpG-D Sequence 5 ACA TTC CTA AGT CTG AAA CAT TAC AGC TTG CTA CAC GAG AAG AGC CGC CAT AGT
Supplementary Information. Construction of Lasso Peptide Fusion Proteins
 Supplementary Information Construction of Lasso Peptide Fusion Proteins Chuhan Zong 1, Mikhail O. Maksimov 2, A. James Link 2,3 * Departments of 1 Chemistry, 2 Chemical and Biological Engineering, and
Supplementary Information Construction of Lasso Peptide Fusion Proteins Chuhan Zong 1, Mikhail O. Maksimov 2, A. James Link 2,3 * Departments of 1 Chemistry, 2 Chemical and Biological Engineering, and
ΔPDD1 x ΔPDD1. ΔPDD1 x wild type. 70 kd Pdd1. Pdd3
 Supplemental Fig. S1 ΔPDD1 x wild type ΔPDD1 x ΔPDD1 70 kd Pdd1 50 kd 37 kd Pdd3 Supplemental Fig. S1. ΔPDD1 strains express no detectable Pdd1 protein. Western blot analysis of whole-protein extracts
Supplemental Fig. S1 ΔPDD1 x wild type ΔPDD1 x ΔPDD1 70 kd Pdd1 50 kd 37 kd Pdd3 Supplemental Fig. S1. ΔPDD1 strains express no detectable Pdd1 protein. Western blot analysis of whole-protein extracts
Creation of A Caspese-3 Sensing System Using A Combination of Split- GFP and Split-Intein
 Supplementary Information Creation of A Caspese-3 Sensing System Using A Combination of Split- GFP and Split-Intein Seiji Sakamoto,* Mika Terauchi, Anna Hugo, Tanner Kim, Yasuyuki Araki and Takehiko Wada*
Supplementary Information Creation of A Caspese-3 Sensing System Using A Combination of Split- GFP and Split-Intein Seiji Sakamoto,* Mika Terauchi, Anna Hugo, Tanner Kim, Yasuyuki Araki and Takehiko Wada*
Supplement 1: Sequences of Capture Probes. Capture probes were /5AmMC6/CTG TAG GTG CGG GTG GAC GTA GTC
 Supplementary Appendixes Supplement 1: Sequences of Capture Probes. Capture probes were /5AmMC6/CTG TAG GTG CGG GTG GAC GTA GTC ACG TAG CTC CGG CTG GA-3 for vimentin, /5AmMC6/TCC CTC GCG CGT GGC TTC CGC
Supplementary Appendixes Supplement 1: Sequences of Capture Probes. Capture probes were /5AmMC6/CTG TAG GTG CGG GTG GAC GTA GTC ACG TAG CTC CGG CTG GA-3 for vimentin, /5AmMC6/TCC CTC GCG CGT GGC TTC CGC
Y-chromosomal haplogroup typing Using SBE reaction
 Schematic of multiplex PCR followed by SBE reaction Multiplex PCR Exo SAP purification SBE reaction 5 A 3 ddatp ddgtp 3 T 5 A G 3 T 5 3 5 G C 5 3 3 C 5 ddttp ddctp 5 T 3 T C 3 A 5 3 A 5 5 C 3 3 G 5 3 G
Schematic of multiplex PCR followed by SBE reaction Multiplex PCR Exo SAP purification SBE reaction 5 A 3 ddatp ddgtp 3 T 5 A G 3 T 5 3 5 G C 5 3 3 C 5 ddttp ddctp 5 T 3 T C 3 A 5 3 A 5 5 C 3 3 G 5 3 G
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/10/494/eaan6284/dc1 Supplementary Materials for Activation of master virulence regulator PhoP in acidic ph requires the Salmonella-specific protein UgtL Jeongjoon
www.sciencesignaling.org/cgi/content/full/10/494/eaan6284/dc1 Supplementary Materials for Activation of master virulence regulator PhoP in acidic ph requires the Salmonella-specific protein UgtL Jeongjoon
Thr Gly Tyr. Gly Lys Asn
 Your unique body characteristics (traits), such as hair color or blood type, are determined by the proteins your body produces. Proteins are the building blocks of life - in fact, about 45% of the human
Your unique body characteristics (traits), such as hair color or blood type, are determined by the proteins your body produces. Proteins are the building blocks of life - in fact, about 45% of the human
Multiplexing Genome-scale Engineering
 Multiplexing Genome-scale Engineering Harris Wang, Ph.D. Department of Systems Biology Department of Pathology & Cell Biology http://wanglab.c2b2.columbia.edu Rise of Genomics An Expanding Toolbox Esvelt
Multiplexing Genome-scale Engineering Harris Wang, Ph.D. Department of Systems Biology Department of Pathology & Cell Biology http://wanglab.c2b2.columbia.edu Rise of Genomics An Expanding Toolbox Esvelt
FROM DNA TO GENETIC GENEALOGY Stephen P. Morse
 1. GENES, CHROMOSOMES, AND DNA Chromosomes FROM DNA TO GENETIC GENEALOGY Stephen P. Morse (steve@stevemorse.org) Every human cell = 46 chromosomes (1 to 22 in pairs, 2 sex chromosomes) Male: sex chromosomes
1. GENES, CHROMOSOMES, AND DNA Chromosomes FROM DNA TO GENETIC GENEALOGY Stephen P. Morse (steve@stevemorse.org) Every human cell = 46 chromosomes (1 to 22 in pairs, 2 sex chromosomes) Male: sex chromosomes
INTRODUCTION TO THE MOLECULAR GENETICS OF THE COLOR MUTATIONS IN ROCK POCKET MICE
 The Making of the The Fittest: Making of the Fittest Natural Selection Natural and Adaptation Selection and Adaptation Educator Materials TEACHER MATERIALS INTRODUCTION TO THE MOLECULAR GENETICS OF THE
The Making of the The Fittest: Making of the Fittest Natural Selection Natural and Adaptation Selection and Adaptation Educator Materials TEACHER MATERIALS INTRODUCTION TO THE MOLECULAR GENETICS OF THE
Supporting Information
 Supporting Information Barderas et al. 10.1073/pnas.0801221105 SI Text: Docking of gastrin to Constructed scfv Models Interactive predocking of the 4-WL-5 motif into the central pocket observed in the
Supporting Information Barderas et al. 10.1073/pnas.0801221105 SI Text: Docking of gastrin to Constructed scfv Models Interactive predocking of the 4-WL-5 motif into the central pocket observed in the
Table S1. Bacterial strains (Related to Results and Experimental Procedures)
 Table S1. Bacterial strains (Related to Results and Experimental Procedures) Strain number Relevant genotype Source or reference 1045 AB1157 Graham Walker (Donnelly and Walker, 1989) 2458 3084 (MG1655)
Table S1. Bacterial strains (Related to Results and Experimental Procedures) Strain number Relevant genotype Source or reference 1045 AB1157 Graham Walker (Donnelly and Walker, 1989) 2458 3084 (MG1655)
Additional Table A1. Accession numbers of resource records for all rhodopsin sequences downloaded from NCBI. Species common name
 1 2 3 Additional Table A1. Accession numbers of resource records for all rhodopsin sequences downloaded from NCBI. Species common name Scientific name Accession number Accession number (introns) Codons
1 2 3 Additional Table A1. Accession numbers of resource records for all rhodopsin sequences downloaded from NCBI. Species common name Scientific name Accession number Accession number (introns) Codons
BIOSTAT516 Statistical Methods in Genetic Epidemiology Autumn 2005 Handout1, prepared by Kathleen Kerr and Stephanie Monks
 Rationale of Genetic Studies Some goals of genetic studies include: to identify the genetic causes of phenotypic variation develop genetic tests o benefits to individuals and to society are still uncertain
Rationale of Genetic Studies Some goals of genetic studies include: to identify the genetic causes of phenotypic variation develop genetic tests o benefits to individuals and to society are still uncertain
II 0.95 DM2 (RPP1) DM3 (At3g61540) b
 Table S2. F 2 Segregation Ratios at 16 C, Related to Figure 2 Cross n c Phenotype Model e 2 Locus A Locus B Normal F 1 -like Enhanced d Uk-1/Uk-3 149 64 36 49 DM2 (RPP1) DM1 (SSI4) a Bla-1/Hh-0 F 3 111
Table S2. F 2 Segregation Ratios at 16 C, Related to Figure 2 Cross n c Phenotype Model e 2 Locus A Locus B Normal F 1 -like Enhanced d Uk-1/Uk-3 149 64 36 49 DM2 (RPP1) DM1 (SSI4) a Bla-1/Hh-0 F 3 111
Overexpression Normal expression Overexpression Normal expression. 26 (21.1%) N (%) P-value a N (%)
 SUPPLEMENTARY TABLES Table S1. Alteration of ZNF322A protein expression levels in relation to clinicopathological parameters in 123 Asian and 74 Caucasian lung cancer patients. Asian patients Caucasian
SUPPLEMENTARY TABLES Table S1. Alteration of ZNF322A protein expression levels in relation to clinicopathological parameters in 123 Asian and 74 Caucasian lung cancer patients. Asian patients Caucasian
Sampling Random Bioinformatics Puzzles using Adaptive Probability Distributions
 Sampling Random Bioinformatics Puzzles using Adaptive Probability Distributions Christian Theil Have 1, Emil Vincent Appel 1, Jette Bork-Jensen 1, and Ole Torp Lassen 2 1 Novo Nordisk Foundation Center
Sampling Random Bioinformatics Puzzles using Adaptive Probability Distributions Christian Theil Have 1, Emil Vincent Appel 1, Jette Bork-Jensen 1, and Ole Torp Lassen 2 1 Novo Nordisk Foundation Center
A Circular Code in the Protein Coding Genes of Mitochondria
 J. theor. Biol. (1997) 189, 273 290 A Circular Code in the Protein Coding Genes of Mitochondria DIDIER G. ARQUE` S* AND CHRISTIAN J. MICHEL *Equipe de Biologie The orique, Universite de Marne la Valle
J. theor. Biol. (1997) 189, 273 290 A Circular Code in the Protein Coding Genes of Mitochondria DIDIER G. ARQUE` S* AND CHRISTIAN J. MICHEL *Equipe de Biologie The orique, Universite de Marne la Valle
SUPPORTING INFORMATION
 SUPPORTING INFORMATION Investigation of the Biosynthesis of the Lasso Peptide Chaxapeptin Using an E. coli-based Production System Helena Martin-Gómez, Uwe Linne, Fernando Albericio, Judit Tulla-Puche,*
SUPPORTING INFORMATION Investigation of the Biosynthesis of the Lasso Peptide Chaxapeptin Using an E. coli-based Production System Helena Martin-Gómez, Uwe Linne, Fernando Albericio, Judit Tulla-Puche,*
Supplemental material
 Supplemental material Diversity of O-antigen repeat-unit structures can account for the substantial sequence variation of Wzx translocases Yaoqin Hong and Peter R. Reeves School of Molecular Bioscience,
Supplemental material Diversity of O-antigen repeat-unit structures can account for the substantial sequence variation of Wzx translocases Yaoqin Hong and Peter R. Reeves School of Molecular Bioscience,
Supporting Online Information
 Supporting Online Information Isolation of Human Genomic DNA Sequences with Expanded Nucleobase Selectivity Preeti Rathi, Sara Maurer, Grzegorz Kubik and Daniel Summerer* Department of Chemistry and Chemical
Supporting Online Information Isolation of Human Genomic DNA Sequences with Expanded Nucleobase Selectivity Preeti Rathi, Sara Maurer, Grzegorz Kubik and Daniel Summerer* Department of Chemistry and Chemical
SUPPLEMENTAL MATERIAL GENOTYPING WITH MULTIPLEXING TARGETED RESEQUENCING
 SUPPLEMENTAL MATERIAL GENOTYPING WITH MULTIPLEXING TARGETED RESEQUENCING All of the patients and control subjects were sequenced and genotyped in the same way. Shotgun libraries of approximately 250 bp
SUPPLEMENTAL MATERIAL GENOTYPING WITH MULTIPLEXING TARGETED RESEQUENCING All of the patients and control subjects were sequenced and genotyped in the same way. Shotgun libraries of approximately 250 bp
Nucleic Acids Research
 Volume 10 Number 1 1982 VoLume 10 Number 11982 Nucleic Acids Research Nucleic Acids Research A convenient and adaptable package of DNA sequence analysis programs for microcomputers James Pustell and Fotis
Volume 10 Number 1 1982 VoLume 10 Number 11982 Nucleic Acids Research Nucleic Acids Research A convenient and adaptable package of DNA sequence analysis programs for microcomputers James Pustell and Fotis
Supporting Information. Copyright Wiley-VCH Verlag GmbH & Co. KGaA, Weinheim, 2006
 Supporting Information Copyright Wiley-VCH Verlag GmbH & Co. KGaA, 69451 Weinheim, 2006 Copyright Wiley-VCH Verlag GmbH & Co. KGaA, 69451 Weinheim, 2006 Supporting Information for Expanding the Genetic
Supporting Information Copyright Wiley-VCH Verlag GmbH & Co. KGaA, 69451 Weinheim, 2006 Copyright Wiley-VCH Verlag GmbH & Co. KGaA, 69451 Weinheim, 2006 Supporting Information for Expanding the Genetic
2
 1 2 3 4 5 6 7 Supplemental Table 1. Magnaporthe oryzae strains generated in this study. Strain background Genotype Strain name Description Guy-11 H1:RFP H1:RFP Strain expressing Histone H1- encoding gene
1 2 3 4 5 6 7 Supplemental Table 1. Magnaporthe oryzae strains generated in this study. Strain background Genotype Strain name Description Guy-11 H1:RFP H1:RFP Strain expressing Histone H1- encoding gene
Add 5µl of 3N NaOH to DNA sample (final concentration 0.3N NaOH).
 Bisulfite Treatment of DNA Dilute DNA sample to 2µg DNA in 50µl ddh 2 O. Add 5µl of 3N NaOH to DNA sample (final concentration 0.3N NaOH). Incubate in a 37ºC water bath for 30 minutes. To 55µl samples
Bisulfite Treatment of DNA Dilute DNA sample to 2µg DNA in 50µl ddh 2 O. Add 5µl of 3N NaOH to DNA sample (final concentration 0.3N NaOH). Incubate in a 37ºC water bath for 30 minutes. To 55µl samples
Supplemental Table 1. Mutant ADAMTS3 alleles detected in HEK293T clone 4C2. WT CCTGTCACTTTGGTTGATAGC MVLLSLWLIAAALVEVR
 Supplemental Dataset Supplemental Table 1. Mutant ADAMTS3 alleles detected in HEK293T clone 4C2. DNA sequence Amino acid sequence WT CCTGTCACTTTGGTTGATAGC MVLLSLWLIAAALVEVR Allele 1 CCTGTC------------------GATAGC
Supplemental Dataset Supplemental Table 1. Mutant ADAMTS3 alleles detected in HEK293T clone 4C2. DNA sequence Amino acid sequence WT CCTGTCACTTTGGTTGATAGC MVLLSLWLIAAALVEVR Allele 1 CCTGTC------------------GATAGC
Supplemental Information. Human Senataxin Resolves RNA/DNA Hybrids. Formed at Transcriptional Pause Sites. to Promote Xrn2-Dependent Termination
 Supplemental Information Molecular Cell, Volume 42 Human Senataxin Resolves RNA/DNA Hybrids Formed at Transcriptional Pause Sites to Promote Xrn2-Dependent Termination Konstantina Skourti-Stathaki, Nicholas
Supplemental Information Molecular Cell, Volume 42 Human Senataxin Resolves RNA/DNA Hybrids Formed at Transcriptional Pause Sites to Promote Xrn2-Dependent Termination Konstantina Skourti-Stathaki, Nicholas
RPA-AB RPA-C Supplemental Figure S1: SDS-PAGE stained with Coomassie Blue after protein purification.
 RPA-AB RPA-C (a) (b) (c) (d) (e) (f) Supplemental Figure S: SDS-PAGE stained with Coomassie Blue after protein purification. (a) RPA; (b) RPA-AB; (c) RPA-CDE; (d) RPA-CDE core; (e) RPA-DE; and (f) RPA-C
RPA-AB RPA-C (a) (b) (c) (d) (e) (f) Supplemental Figure S: SDS-PAGE stained with Coomassie Blue after protein purification. (a) RPA; (b) RPA-AB; (c) RPA-CDE; (d) RPA-CDE core; (e) RPA-DE; and (f) RPA-C
hcd1tg/hj1tg/ ApoE-/- hcd1tg/hj1tg/ ApoE+/+
 ApoE+/+ ApoE-/- ApoE-/- H&E (1x) Supplementary Figure 1. No obvious pathology is observed in the colon of diseased ApoE-/me. Colon samples were fixed in 1% formalin and laid out in Swiss rolls for paraffin
ApoE+/+ ApoE-/- ApoE-/- H&E (1x) Supplementary Figure 1. No obvious pathology is observed in the colon of diseased ApoE-/me. Colon samples were fixed in 1% formalin and laid out in Swiss rolls for paraffin
Genomics and Gene Recognition Genes and Blue Genes
 Genomics and Gene Recognition Genes and Blue Genes November 1, 2004 Prokaryotic Gene Structure prokaryotes are simplest free-living organisms studying prokaryotes can give us a sense what is the minimum
Genomics and Gene Recognition Genes and Blue Genes November 1, 2004 Prokaryotic Gene Structure prokaryotes are simplest free-living organisms studying prokaryotes can give us a sense what is the minimum
Evolution of protein coding sequences
 Evolution of protein coding sequences Kinds of nucleo-de subs-tu-ons Given 2 nucleo-de sequences, how their similari-es and differences arose from a common ancestor? We assume A the common ancestor: Single
Evolution of protein coding sequences Kinds of nucleo-de subs-tu-ons Given 2 nucleo-de sequences, how their similari-es and differences arose from a common ancestor? We assume A the common ancestor: Single
Supplementary Figure 1. Localization of MST1 in RPE cells. Proliferating or ciliated HA- MST1 expressing RPE cells (see Fig. 5b for establishment of
 Supplementary Figure 1. Localization of MST1 in RPE cells. Proliferating or ciliated HA- MST1 expressing RPE cells (see Fig. 5b for establishment of the cell line) were immunostained for HA, acetylated
Supplementary Figure 1. Localization of MST1 in RPE cells. Proliferating or ciliated HA- MST1 expressing RPE cells (see Fig. 5b for establishment of the cell line) were immunostained for HA, acetylated
SUPPLEMENTARY INFORMATION
 doi: 10.1038/nature07182 SUPPLEMENTAL FIGURES AND TABLES Fig. S1. myf5-expressing cells give rise to brown fat depots and skeletal muscle (a) Perirenal BAT from control (cre negative) and myf5-cre:r26r3-yfp
doi: 10.1038/nature07182 SUPPLEMENTAL FIGURES AND TABLES Fig. S1. myf5-expressing cells give rise to brown fat depots and skeletal muscle (a) Perirenal BAT from control (cre negative) and myf5-cre:r26r3-yfp
evaluated with UAS CLB eliciting UAS CIT -N Libraries increase in the
 Supplementary Figures Supplementary Figure 1: Promoter scaffold library assemblies. Many ensembless of libraries were evaluated in this work. As a legend, the box outline color in top half of the figure
Supplementary Figures Supplementary Figure 1: Promoter scaffold library assemblies. Many ensembless of libraries were evaluated in this work. As a legend, the box outline color in top half of the figure
strain devoid of the aox1 gene [1]. Thus, the identification of AOX1 in the intracellular
![strain devoid of the aox1 gene [1]. Thus, the identification of AOX1 in the intracellular strain devoid of the aox1 gene [1]. Thus, the identification of AOX1 in the intracellular](/thumbs/72/66361212.jpg) Additional file 2 Identification of AOX1 in P. pastoris GS115 with a Mut s phenotype Results and Discussion The HBsAg producing strain was originally identified as a Mut s (methanol utilization slow) strain
Additional file 2 Identification of AOX1 in P. pastoris GS115 with a Mut s phenotype Results and Discussion The HBsAg producing strain was originally identified as a Mut s (methanol utilization slow) strain
Supporting Information
 Supporting Information Transfection of DNA Cages into Mammalian Cells Email: a.turberfield@physics.ox.ac.uk Table of Contents Supporting Figure 1 DNA tetrahedra used in transfection experiments 2 Supporting
Supporting Information Transfection of DNA Cages into Mammalian Cells Email: a.turberfield@physics.ox.ac.uk Table of Contents Supporting Figure 1 DNA tetrahedra used in transfection experiments 2 Supporting
1. DNA, RNA structure. 2. DNA replication. 3. Transcription, translation
 1. DNA, RNA structure 2. DNA replication 3. Transcription, translation DNA and RNA are polymers of nucleotides DNA is a nucleic acid, made of long chains of nucleotides Nucleotide Phosphate group Nitrogenous
1. DNA, RNA structure 2. DNA replication 3. Transcription, translation DNA and RNA are polymers of nucleotides DNA is a nucleic acid, made of long chains of nucleotides Nucleotide Phosphate group Nitrogenous
Quantitative reverse-transcription PCR. Transcript levels of flgs, flgr, flia and flha were
 1 Supplemental methods 2 3 4 5 6 7 8 9 1 11 12 13 14 15 16 17 18 19 21 22 23 Quantitative reverse-transcription PCR. Transcript levels of flgs, flgr, flia and flha were monitored by quantitative reverse-transcription
1 Supplemental methods 2 3 4 5 6 7 8 9 1 11 12 13 14 15 16 17 18 19 21 22 23 Quantitative reverse-transcription PCR. Transcript levels of flgs, flgr, flia and flha were monitored by quantitative reverse-transcription
Gene Prediction: Statistical Approaches
 Gene Prediction: Statistical Approaches Outline 1. Central Dogma and Codons 2. Discovery of Split Genes 3. Splicing 4. Open Reading Frames 5. Codon Usage 6. Splicing Signals 7. TestCode Section 1: Central
Gene Prediction: Statistical Approaches Outline 1. Central Dogma and Codons 2. Discovery of Split Genes 3. Splicing 4. Open Reading Frames 5. Codon Usage 6. Splicing Signals 7. TestCode Section 1: Central
Complexity of the Ruminococcus flavefaciens FD-1 cellulosome reflects an expansion of family-related protein-protein interactions
 Complexity of the Ruminococcus flavefaciens FD-1 cellulosome reflects an expansion of family-related protein-protein interactions Vered Israeli-Ruimy 1,*, Pedro Bule 2,*, Sadanari Jindou 3, Bareket Dassa
Complexity of the Ruminococcus flavefaciens FD-1 cellulosome reflects an expansion of family-related protein-protein interactions Vered Israeli-Ruimy 1,*, Pedro Bule 2,*, Sadanari Jindou 3, Bareket Dassa
Search for and Analysis of Single Nucleotide Polymorphisms (SNPs) in Rice (Oryza sativa, Oryza rufipogon) and Establishment of SNP Markers
 DNA Research 9, 163 171 (2002) Search for and Analysis of Single Nucleotide Polymorphisms (SNPs) in Rice (Oryza sativa, Oryza rufipogon) and Establishment of SNP Markers Shinobu Nasu, Junko Suzuki, Rieko
DNA Research 9, 163 171 (2002) Search for and Analysis of Single Nucleotide Polymorphisms (SNPs) in Rice (Oryza sativa, Oryza rufipogon) and Establishment of SNP Markers Shinobu Nasu, Junko Suzuki, Rieko
Cover Page. The handle holds various files of this Leiden University dissertation.
 Cover Page The handle http://hdl.handle.net/1887/45045 holds various files of this Leiden University dissertation. Author: Vis, J.K. Title: Algorithms for the description of molecular sequences Issue Date:
Cover Page The handle http://hdl.handle.net/1887/45045 holds various files of this Leiden University dissertation. Author: Vis, J.K. Title: Algorithms for the description of molecular sequences Issue Date:
Biomolecules: lecture 6
 Biomolecules: lecture 6 - to learn the basics on how DNA serves to make RNA = transcription - to learn how the genetic code instructs protein synthesis - to learn the basics on how proteins are synthesized
Biomolecules: lecture 6 - to learn the basics on how DNA serves to make RNA = transcription - to learn how the genetic code instructs protein synthesis - to learn the basics on how proteins are synthesized
Gene synthesis by circular assembly amplification
 Gene synthesis by circular assembly amplification Duhee Bang & George M Church Supplementary figures and text: Supplementary Figure 1. Dpo4 gene (1.05kb) construction by various methods. Supplementary
Gene synthesis by circular assembly amplification Duhee Bang & George M Church Supplementary figures and text: Supplementary Figure 1. Dpo4 gene (1.05kb) construction by various methods. Supplementary
Genes and Proteins. Objectives
 Genes and Proteins Lecture 15 Objectives At the end of this series of lectures, you should be able to: Define terms. Explain the central dogma of molecular biology. Describe the locations, reactants, and
Genes and Proteins Lecture 15 Objectives At the end of this series of lectures, you should be able to: Define terms. Explain the central dogma of molecular biology. Describe the locations, reactants, and
Supplementary Figures
 Supplementary Figures Supplementary Fig. 1 Characterization of GSCs. a. Immunostaining of primary GSC spheres from GSC lines. Nestin (neural progenitor marker, red), TLX (green). Merged images of nestin,
Supplementary Figures Supplementary Fig. 1 Characterization of GSCs. a. Immunostaining of primary GSC spheres from GSC lines. Nestin (neural progenitor marker, red), TLX (green). Merged images of nestin,
Supplemental Data. Distinct Pathways for snorna and mrna Termination
 Molecular Cell, Volume 24 Supplemental Data Distinct Pathways for snorna and mrna Termination Minkyu Kim, Lidia Vasiljeva, Oliver J. Rando, Alexander Zhelkovsky, Claire Moore, and Stephen Buratowski A
Molecular Cell, Volume 24 Supplemental Data Distinct Pathways for snorna and mrna Termination Minkyu Kim, Lidia Vasiljeva, Oliver J. Rando, Alexander Zhelkovsky, Claire Moore, and Stephen Buratowski A
FAT10 and NUB1L bind the VWA domain of Rpn10 and Rpn1 to enable proteasome-mediated proteolysis
 SUPPLEMENTARY INFORMATION FAT10 and NUB1L bind the VWA domain of Rpn10 and Rpn1 to enable proteasome-mediated proteolysis Neha Rani, Annette Aichem, Gunter Schmidtke, Stefan Kreft, and Marcus Groettrup
SUPPLEMENTARY INFORMATION FAT10 and NUB1L bind the VWA domain of Rpn10 and Rpn1 to enable proteasome-mediated proteolysis Neha Rani, Annette Aichem, Gunter Schmidtke, Stefan Kreft, and Marcus Groettrup
11th Meeting of the Science Working Group. Lima, Peru, October 2012 SWG-11-JM-11
 11th Meeting of the Science Working Group Lima, Peru, 15-19 October 2012 Russian population genetics studies of jack mackerel in the South Pacific P.K.Afanasiev M.A.Rabchun A.I.Glubokov Introduction. In
11th Meeting of the Science Working Group Lima, Peru, 15-19 October 2012 Russian population genetics studies of jack mackerel in the South Pacific P.K.Afanasiev M.A.Rabchun A.I.Glubokov Introduction. In
Supplemental Table 1. Primers used for PCR.
 Supplemental Table 1. Primers used for PCR. Gene Type Primer Sequence Genotyping and semi-quantitative RT-PCR F 5 -TTG CCC GAT CAC CAT CTG TA-3 rwa1-1 R 5 -TGT AGC GAT CAA GGC CTG ATC TAA-3 LB 5 -TAG CAT
Supplemental Table 1. Primers used for PCR. Gene Type Primer Sequence Genotyping and semi-quantitative RT-PCR F 5 -TTG CCC GAT CAC CAT CTG TA-3 rwa1-1 R 5 -TGT AGC GAT CAA GGC CTG ATC TAA-3 LB 5 -TAG CAT
BioInformatics and Computational Molecular Biology. Course Website
 BioInformatics and Computational Molecular Biology Course Website http://bioinformatics.uchc.edu What is Bioinformatics Bioinformatics upgrades the information content of biological measurements. Discovery
BioInformatics and Computational Molecular Biology Course Website http://bioinformatics.uchc.edu What is Bioinformatics Bioinformatics upgrades the information content of biological measurements. Discovery
Gene Prediction: Statistical Approaches
 Gene Prediction: Statistical Approaches Outline Codons Discovery of Split Genes Exons and Introns Splicing Open Reading Frames Codon Usage Splicing Signals TestCode Gene Prediction: Computational Challenge
Gene Prediction: Statistical Approaches Outline Codons Discovery of Split Genes Exons and Introns Splicing Open Reading Frames Codon Usage Splicing Signals TestCode Gene Prediction: Computational Challenge
Converting rabbit hybridoma into recombinant antibodies with effective transient production in an optimized human expression system
 Converting rabbit hybridoma into recombinant antibodies with effective transient production in an optimized human expression system Dr. Tim Welsink Molecular Biology Transient Gene Expression OUTLINE Short
Converting rabbit hybridoma into recombinant antibodies with effective transient production in an optimized human expression system Dr. Tim Welsink Molecular Biology Transient Gene Expression OUTLINE Short
Cat. # Product Size DS130 DynaExpress TA PCR Cloning Kit (ptakn-2) 20 reactions Box 1 (-20 ) ptakn-2 Vector, linearized 20 µl (50 ng/µl) 1
 Product Name: Kit Component TA PCR Cloning Kit (ptakn-2) Cat. # Product Size DS130 TA PCR Cloning Kit (ptakn-2) 20 reactions Box 1 (-20 ) ptakn-2 Vector, linearized 20 µl (50 ng/µl) 1 2 Ligation Buffer
Product Name: Kit Component TA PCR Cloning Kit (ptakn-2) Cat. # Product Size DS130 TA PCR Cloning Kit (ptakn-2) 20 reactions Box 1 (-20 ) ptakn-2 Vector, linearized 20 µl (50 ng/µl) 1 2 Ligation Buffer
Legends for supplementary figures 1-3
 High throughput resistance profiling of Plasmodium falciparum infections based on custom dual indexing and Illumina next generation sequencing-technology Sidsel Nag 1,2 *, Marlene D. Dalgaard 3, Poul-Erik
High throughput resistance profiling of Plasmodium falciparum infections based on custom dual indexing and Illumina next generation sequencing-technology Sidsel Nag 1,2 *, Marlene D. Dalgaard 3, Poul-Erik
Gene Prediction: Statistical Approaches
 Gene Prediction: Statistical Approaches Outline Codons Discovery of Split Genes Exons and Introns Splicing Open Reading Frames Codon Usage Splicing Signals TestCode Gene Prediction: Computational Challenge
Gene Prediction: Statistical Approaches Outline Codons Discovery of Split Genes Exons and Introns Splicing Open Reading Frames Codon Usage Splicing Signals TestCode Gene Prediction: Computational Challenge
EECS730: Introduction to Bioinformatics
 EECS730: Introduction to Bioinformatics Lecture 08: Gene finding aatgcatgcggctatgctaatgcatgcggctatgctaagctgggatccgatgacaatgcatgcggctatgctaatgcatgcggc tatgcaagctgggatccgatgactatgctaagctgggatccgatgacaatgcatgcggctatgctaatgaatggtcttgggatt
EECS730: Introduction to Bioinformatics Lecture 08: Gene finding aatgcatgcggctatgctaatgcatgcggctatgctaagctgggatccgatgacaatgcatgcggctatgctaatgcatgcggc tatgcaagctgggatccgatgactatgctaagctgggatccgatgacaatgcatgcggctatgctaatgaatggtcttgggatt
iclicker Question #28B - after lecture Shown below is a diagram of a typical eukaryotic gene which encodes a protein: start codon stop codon 2 3
 Bio 111 Handout for Molecular Biology 4 This handout contains: Today s iclicker Questions Information on Exam 3 Solutions Fall 2008 Exam 3 iclicker Question #28A - before lecture Which of the following
Bio 111 Handout for Molecular Biology 4 This handout contains: Today s iclicker Questions Information on Exam 3 Solutions Fall 2008 Exam 3 iclicker Question #28A - before lecture Which of the following
PCR-based Markers and Cut Flower Longevity in Carnation
 PCRbased Markers and Cut Flower Longevity in Carnation Laura De Benedetti, Luca Braglia, Simona Bruna, Gianluca Burchi *, Antonio Mercuri and Tito Schiva Istituto Sperimentale per la Floricoltura, Corso
PCRbased Markers and Cut Flower Longevity in Carnation Laura De Benedetti, Luca Braglia, Simona Bruna, Gianluca Burchi *, Antonio Mercuri and Tito Schiva Istituto Sperimentale per la Floricoltura, Corso
S4B fluorescence (AU)
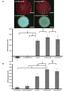 A S4B fluorescence (AU) S4B fluorescence (AU) dsbb csgba csgd dsbb csgba bcsa 5000 * NS NS 4000 * 3000 2000 1000 0 ΔcsgBAΔbcsA ΔcsgDΔdsbBΔbcsA ΔcsgBA ΔdsbBΔcsgBA ΔcsgDΔdsbB B -1000 4000 * * NS 3500 * 3000
A S4B fluorescence (AU) S4B fluorescence (AU) dsbb csgba csgd dsbb csgba bcsa 5000 * NS NS 4000 * 3000 2000 1000 0 ΔcsgBAΔbcsA ΔcsgDΔdsbBΔbcsA ΔcsgBA ΔdsbBΔcsgBA ΔcsgDΔdsbB B -1000 4000 * * NS 3500 * 3000
