Challenges to measuring intracellular Ca 2+ Calmodulin: nature s Ca 2+ sensor
|
|
|
- Rhoda Cain
- 6 years ago
- Views:
Transcription
1 Calcium Signals in Biological Systems Lecture 3 (2/9/0) Measuring intracellular Ca 2+ signals II: Genetically encoded Ca 2+ sensors Henry M. Colecraft, Ph.D. Challenges to measuring intracellular Ca Diversity of Ca 2+ signals Interesting Ca 2+ signals span wide temporal and spatial ranges. 2. Selectivity of indicator for Ca 2+ Cells have mm concentrations of Mg 2+ vs. nm-µm for Ca Delivery of indicator to cells Plasma membrane poses a formidable barrier 4. Fidelity of reporter 5. Targetability to distinct intracellular sites heterogenous distribution of [Ca 2+ ] in organelles and cytosol. Calmodulin: nature s Ca 2+ sensor Calmodulin is the major Ca 2+ binding protein in the cytosol. Structurally, has two lobes with four EF hands that bind Ca 2+. In Ca2+ free state (apocalmodulin), molecule is stretched out. Upon binding Ca 2+, molecule undergoes conformational change to activate downstream effectors.
2 Ca 2+ - and substrate-dependent conformational changes in calmodulin Green fluorescent protein (GFP) Aequoria Victoria Conformational change in calmodulin upon binding Ca 2+ and substrate brings the N- and C- terminal lobes in closer proximity compared to apocalmodulin. GFP crystal structure Expression of GFP in mammalian cells Expression of GFP in other organisms results in fluorescence. Hence, the gene encodes all the information required for posttranslational synthesis of the chromophore. GFP is widely used as a reporter gene to monitor gene expression. In-frame fusion of the gene for GFP to that of other proteins permits tracking of the subcellular localization of proteins. GFP β 2a -GFP β 4 -GFP Mutagenesis of GFP Several mutations have been engineered into wild-type GFP to maximize use in mammalian cells. F99S, M153T, V13A improved folding of the molecule at 37 C. Altering the sequence of GFP to reflect mammalian codon usage and use of strong promoters permitted improved expression levels in mammalian cells. Importantly, targeted mutations permitted the generation of fluorescent proteins with unique spectral properties.
3 Fluorescence spectra of XFP proteins A. Jablonski diagram Fluorescence B. Excitation and emission spectra Distinct XFPs have different brightness levels and unique excitation and emission spectra. Spectral overlap between emission and excitation spectra of specific XFP pairs permits design of fluorescence resonance energy transfer (FRET)-based assays. Fluorescence resonance energy transfer (FRET) FRET occurs when the following two conditions are met: (1) The emission spectrum of a fluorophore, termed the donor (D), overlaps with the excitation spectrum of a second fluorophore, termed the acceptor (A). (2) D and A are in close proximity. FRET does not involve emission and absorption of photons, but reflects the exchange of energy between two oscillating dipoles with similar resonance frequency. Theory of FRET The extent of energy transfer is a function of the distance between donor and acceptor (r) and the spectral overlap between them. The spectral overlap is usually described in terms of the Förster distance, R o. The rate of energy transfer (k T ) is given by, k T 1 R = τ r D where τ D is the lifetime of the donor in the absence of acceptor. The efficiency of energy transfer for a single donor-acceptor pair is given by, E = R Ro o + When r = R o, the rate of energy transfer equals the rate of emission of the donor, and efficiency of energy transfer is 0.5. R o is typically on the order of the size of biological macromolecules (30 0 Å), making FRET a powerful technique for measuring distances between proteins. o r
4 Strategy for cameleons Construct a gene encoding calmodulin and M13 sandwiched by donor and acceptor XFPs. In Ca 2+ -free state, CaM and M13 are disordered and stretched out, and XFPs are relatively far apart. Exciting the donor results in mostly donor emission. Upon binding Ca 2+, CaM binds to M13 decreasing the distance between donor and acceptor XFPs, thereby increasing FRET between them. Now, exciting the donor results in emission mostly from the acceptor XFP. Design of cameleons for bacterial expression Properties of cameleons in vitro Cameleon-1 consists CaM and M13 sandwiched by BFP and GFP. Protein expressed and purified from bacteria and spectral properties examined in vitro. Emission spectra of cameleon-1 measured following excitation at 380 nm. Two-humped emission spectra at 0 Ca 2+ significant resting FRET between BFP and GFP in the intact molecule. Polyhistidine tag permits column purification of protein from bacteria. Optimized splice sequences for different genes found by trial and error. CaM linked to M13 by glycine linker. Increasing Ca2+ resulted in increase in emission fluorescence intensity at 510 nm, and a decrease at 445 nm, i.e. increased FRET.
5 Ca 2+ titration curves of cameleon-1 Emission ratio change of cameleon-1 showed complicated dependence on Ca 2+ concentration. [ Ca 2+ ] = K ' d R R min Rmax R 1 n where, K d is the apparent dissociation constant, R max and R min are emission ratios in 0 and saturating Ca 2+, respectively, and n is the Hill coefficient. For wild-type cameleon-1 (open circles) the titration curve is biphasic with two apparent dissociation constants of 70 nm and 11 µm, and Hill coefficients of 1 and 1.8, respectively. Ca 2+ titration curves of cameleon-1 Mutation E104Q markedly reduces Ca 2+ affinity of the third Ca 2+ -binding loop of calmodulin. Applying this mutation to cameleon-1 (solid circles) eliminates the high affinity component of the wild-type response (K d = 4.4 µm; n = 0.7). Mutating the first Ca 2+ binding loop (E31Q; open triangles) further decreased the affinity of the low-affinity component of cameleon-1, while having no effect on on the high-affinity component (K d = 83 nm and 700 µm; n = 1.5 and 0.87). Mutations offer opportunities to engineer cameleons with different affinities extending extending the range of Ca 2+ concentrations that can be measured. Kinetic properties of cameleon-1 (E104Q) Improved properties of cameleon-2 Stopped-flow measurements of acceptor emission after rapid mixing with Ca 2+ revealed exponential relaxation k obs = = ( kon[ Ca ] + k τ Calculated K d is on the same order as obtained from fits to the titration curve. Relatively slow kinetics make cameleon-1 inappropriate for tracking rapidly changing Ca 2+ concentrations. off ) Cameleon-1 was relatively ineffective as a reporter of [Ca 2+ ] inside mammalian cells because of the dimness of BFP. Second generation cameleon, yellow cameleon-2 was generated to resolve this issue. Yellow cameleon-2 contains ECFP and EYFP as donor and acceptor fluorophores, respectively. Emission spectra of yellow cameleon-2 in the absence and presence of Ca 2+ are shown (excited at 432 nm).
6 Design of cameleons for cellular expression Cameleon-reported Ca 2+ changes in HeLa cells Yellow cameleon-2 (74 kda) expressed in HeLa cells is present throughout the cytosol and excluded from the nucleus. Application of histamine induced oscillatory increases in emission ratio, indicating oscillations in [Ca 2+ ] cyt, which ultimately desensitized. ATP induced a reversible increase in [Ca 2+ ] cyt. R max determined by elevating extracellular Ca 2+ to 20 mm in the presence of Ca 2+ ionophore, ionomycin. R min determined by clamping extracellular Ca 2+ to 0 with EGTA, and chelating intracellular Ca 2+ with BAPTA-AM. Targeting cameleons to intracellular organelles: nucleus Targeting cameleons to intracellular organelles: ER/SR Cameleons containing a nuclear localization signal target specifically to the nucleus when expressed in HeLa cells. Unique advantage of genetically encoded Ca 2+ sensors is this ability to potentially target any subcellular location. Cameleons containing ER targeting and retrieval signals, are targeted to the ER when expressed in HELA cells. Resting emission ratio is high, reflecting the high Ca 2+ concentration in the ER. Stimuli which increase cytosolic [Ca 2+ ] (e.g. histamine and ATP) decrease ER Ca 2+, demonstrating the ER as the source of Ca 2+ signals in these paradigms. Resting emission ratio is identical to R max. This indicates that yellow cameleon-3er is saturated at rest. Hence, estimates of ER Ca 2+ concentration using this cameleon are unreliable.
7 Measuring ER/SR Ca 2+ with cameleon-4er GFP-based detection of protein association and dissociation Lower affinity yellow cameleon-4er is not saturated at resting ER [Ca 2+ ]. ER [Ca 2+ ] concentration estimated as µm. Expression of yellow cameleon-2 as two separate pieces resulted in distribution of both proteins in the cytosol and nucleus. Stimuli that caused changes in Ca 2+ increased emission ratio in both the cytosolic and nuclear compartments. Hence, GFP-based FRET can be used to monitor protein-protein interactions in cells.
The Green Fluorescent Protein. w.chem.uwec.edu/chem412_s99/ppt/green.ppt
 The Green Fluorescent Protein w.chem.uwec.edu/chem412_s99/ppt/green.ppt www.chem.uwec.edu/chem412_s99/ppt/green.ppt Protein (gene) is from a jellyfish: Aequorea victoria www.chem.uwec.edu/chem412_s99/ppt/green.ppt
The Green Fluorescent Protein w.chem.uwec.edu/chem412_s99/ppt/green.ppt www.chem.uwec.edu/chem412_s99/ppt/green.ppt Protein (gene) is from a jellyfish: Aequorea victoria www.chem.uwec.edu/chem412_s99/ppt/green.ppt
SUPPLEMENTARY MATERIAL
 SUPPLEMENTARY MATERIAL Materials and Methods Circular dichroism (CD) spectroscopy. Far ultraviolet (UV) CD spectra of apo- and holo- CaM and the CaM mutants were recorded on a Jasco J-715 spectropolarimeter
SUPPLEMENTARY MATERIAL Materials and Methods Circular dichroism (CD) spectroscopy. Far ultraviolet (UV) CD spectra of apo- and holo- CaM and the CaM mutants were recorded on a Jasco J-715 spectropolarimeter
Fluorescent protein. Origin of fluorescent proteins. Structural and biophysical analysis of FPs. Application in molecular biology and biotechnology
 Origin of fluorescent proteins types of FPs mechanism of fluorescence Fluorescent protein Structural and biophysical analysis of FPs Application in molecular biology and biotechnology Variety of organisms
Origin of fluorescent proteins types of FPs mechanism of fluorescence Fluorescent protein Structural and biophysical analysis of FPs Application in molecular biology and biotechnology Variety of organisms
F* techniques: FRAP, FLIP, FRET, FLIM,
 F* techniques: FRAP, FLIP, FRET, FLIM, FCS Antonia Göhler March 2015 Fluorescence explained in the Bohr model Absorption of light (blue) causes an electron to move to a higher energy orbit. After a particular
F* techniques: FRAP, FLIP, FRET, FLIM, FCS Antonia Göhler March 2015 Fluorescence explained in the Bohr model Absorption of light (blue) causes an electron to move to a higher energy orbit. After a particular
20.320, notes for 9/13
 20.320 Notes Page 1 20.320, notes for 9/13 Thursday, September 13, 2012 9:36 AM Coming Assignments 1st part of design project coming up, assigned 9/14 (tomorrow) and due 9/21. Last time We covered ITC
20.320 Notes Page 1 20.320, notes for 9/13 Thursday, September 13, 2012 9:36 AM Coming Assignments 1st part of design project coming up, assigned 9/14 (tomorrow) and due 9/21. Last time We covered ITC
Chapter One. Construction of a Fluorescent α5 Subunit. Elucidation of the unique contribution of the α5 subunit is complicated by several factors
 4 Chapter One Construction of a Fluorescent α5 Subunit The significance of the α5 containing nachr receptor (α5* receptor) has been a challenging question for researchers since its characterization by
4 Chapter One Construction of a Fluorescent α5 Subunit The significance of the α5 containing nachr receptor (α5* receptor) has been a challenging question for researchers since its characterization by
The most extensively used technique for tissue analysis is light microscopy.
 Fluorescence Theory Quantum yield Wavelength shift Ligand interactions Membrane interactions Using quenchning effects Fluorescence in-vivo Localization Distance measurements FRET The most extensively used
Fluorescence Theory Quantum yield Wavelength shift Ligand interactions Membrane interactions Using quenchning effects Fluorescence in-vivo Localization Distance measurements FRET The most extensively used
Introduction to Fluorescence Jablonski Diagram
 ntroduction to Fluorescence Jablonski Diagram Excited Singlet Manifold S1 internal conversion S2 k -isc k isc Excited riplet Manifold 1 S0 k nr k k' f nr fluorescence k p phosphorescence Singlet round
ntroduction to Fluorescence Jablonski Diagram Excited Singlet Manifold S1 internal conversion S2 k -isc k isc Excited riplet Manifold 1 S0 k nr k k' f nr fluorescence k p phosphorescence Singlet round
Welcome! openmicberkeley.wordpress.com. Open Berkeley
 Welcome! openmicberkeley.wordpress.com Agenda Jen Lee: Introduction to FRET Marla Feller: Using FRET sensors to look at time resolved measurements Becky Lamason: Using FRET to determine if a bacterial
Welcome! openmicberkeley.wordpress.com Agenda Jen Lee: Introduction to FRET Marla Feller: Using FRET sensors to look at time resolved measurements Becky Lamason: Using FRET to determine if a bacterial
Supplementary Figure 1. Design of linker truncation library between Nluc and either Venus or mneongreen
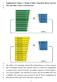 1 2 3 Supplementary Figure 1. Design of linker truncation library between Nluc and either Venus or mneongreen 4 5 6 7 8 9 10 The cdna of C-terminally deleted FPs (mneongreen or Venus) mutants and N-terminally
1 2 3 Supplementary Figure 1. Design of linker truncation library between Nluc and either Venus or mneongreen 4 5 6 7 8 9 10 The cdna of C-terminally deleted FPs (mneongreen or Venus) mutants and N-terminally
Practice Problems 5. Location of LSA-GFP fluorescence
 Life Sciences 1a Practice Problems 5 1. Soluble proteins that are normally secreted from the cell into the extracellular environment must go through a series of steps referred to as the secretory pathway.
Life Sciences 1a Practice Problems 5 1. Soluble proteins that are normally secreted from the cell into the extracellular environment must go through a series of steps referred to as the secretory pathway.
Chapter 4. the biological community to assay for protein-protein interactions. FRET describes the
 31 Chapter 4 Determination of nachr stoichiometry using Normalized Försters Resonance Energy Transfer (NFRET) Försters resonance energy transfer (FRET) has become a technique widely used in the biological
31 Chapter 4 Determination of nachr stoichiometry using Normalized Försters Resonance Energy Transfer (NFRET) Försters resonance energy transfer (FRET) has become a technique widely used in the biological
FRET and FRET based Microscopy Techniques
 Big Question: We can see rafts in Model Membranes (GUVs or Supported Lipid Bilayers, LM), but how to study in cells? Do rafts really exist in cells? Are they static large structures? Are they small transient
Big Question: We can see rafts in Model Membranes (GUVs or Supported Lipid Bilayers, LM), but how to study in cells? Do rafts really exist in cells? Are they static large structures? Are they small transient
Using Quantum Dots in Fluorescence Resonance Energy Transfer Studies
 p.1/31 Using Quantum Dots in Fluorescence Resonance Energy Transfer Studies Rajarshi Guha Pennsylvania State University p.2/31 Introduction Using organic fluorophores as labels A brief overview of fluorescence
p.1/31 Using Quantum Dots in Fluorescence Resonance Energy Transfer Studies Rajarshi Guha Pennsylvania State University p.2/31 Introduction Using organic fluorophores as labels A brief overview of fluorescence
Intracellular localization and trafficking of proteins or How (and why) to find a needle in a haystack
 Intracellular localization and trafficking of proteins or How (and why) to find a needle in a haystack :: The Structure of a Cell :: Relative sizes SUBUNITS 0.2 mm (200 μm) minimum resolvable by unaided
Intracellular localization and trafficking of proteins or How (and why) to find a needle in a haystack :: The Structure of a Cell :: Relative sizes SUBUNITS 0.2 mm (200 μm) minimum resolvable by unaided
Reminder: absorption. OD = A = - log (I / I 0 ) = ε (λ) c x. I = I ε(λ) c x. Definitions. Fluorescence quenching and FRET.
 Reminder: absorption Special fluorescence applications I 0 I Fluorescence quenching and FRET Miklós Nyitrai; 24 th of Februry 2011. substance OD = A = - log (I / I 0 ) = ε (λ) c x optical density I = I
Reminder: absorption Special fluorescence applications I 0 I Fluorescence quenching and FRET Miklós Nyitrai; 24 th of Februry 2011. substance OD = A = - log (I / I 0 ) = ε (λ) c x optical density I = I
Real-Time PCR Principles and Applications
 Real-Time PCR Principles and Applications Dr Esam Ibraheem Azhar (BSc, MSc, Ph.D Molecular Medical Virology) Asst. Prof. Medical Laboratory Technology Department Objectives Real-Time PCR Principles and
Real-Time PCR Principles and Applications Dr Esam Ibraheem Azhar (BSc, MSc, Ph.D Molecular Medical Virology) Asst. Prof. Medical Laboratory Technology Department Objectives Real-Time PCR Principles and
Supplementary Figure 1. The normalized absorption and emission spectra of 605QD
 1..8 65Q Absorbance 65Q Emission Cy5 Absorbance Cy5 Emission 1..8 Extiction Absorption Coefficient.6.4.2. 45 5 55 6 65 7 75 8 Wavelength (nm).6.4.2. Fluorescence Emission Intensity Supplementary Figure
1..8 65Q Absorbance 65Q Emission Cy5 Absorbance Cy5 Emission 1..8 Extiction Absorption Coefficient.6.4.2. 45 5 55 6 65 7 75 8 Wavelength (nm).6.4.2. Fluorescence Emission Intensity Supplementary Figure
Contact Details. Dr Alexander Galkin. Office: MBC Room 186. Tel: (028) Frequency and wavelength.
 Contact Details The electromagnetic spectrum Biological Spectroscopy Dr Alexander Galkin Email: a.galkin@qub.ac.uk Dr Alexander Galkin MSc Biomolecular Function - BBC8045 Office: MBC Room 186 Tel: (028)
Contact Details The electromagnetic spectrum Biological Spectroscopy Dr Alexander Galkin Email: a.galkin@qub.ac.uk Dr Alexander Galkin MSc Biomolecular Function - BBC8045 Office: MBC Room 186 Tel: (028)
FRET measurement between YFP and CFP
 FRET measurement between YFP and CFP EYFP and ECFP function as a donor-acceptor pair for fluorescence resonance energy transfer (FRET), in which excitation of the donor (cyan) molecule leads to emission
FRET measurement between YFP and CFP EYFP and ECFP function as a donor-acceptor pair for fluorescence resonance energy transfer (FRET), in which excitation of the donor (cyan) molecule leads to emission
Supplementary information. for the paper:
 Supplementary information for the paper: Screening for protein-protein interactions using Förster resonance energy transfer (FRET) and fluorescence lifetime imaging microscopy (FLIM) Anca Margineanu 1*,
Supplementary information for the paper: Screening for protein-protein interactions using Förster resonance energy transfer (FRET) and fluorescence lifetime imaging microscopy (FLIM) Anca Margineanu 1*,
SUPPLEMENTARY INFORMATION. Supplementary Figures 1-8
 SUPPLEMENTARY INFORMATION Supplementary Figures 1-8 Supplementary Figure 1. TFAM residues contacting the DNA minor groove (A) TFAM contacts on nonspecific DNA. Leu58, Ile81, Asn163, Pro178, and Leu182
SUPPLEMENTARY INFORMATION Supplementary Figures 1-8 Supplementary Figure 1. TFAM residues contacting the DNA minor groove (A) TFAM contacts on nonspecific DNA. Leu58, Ile81, Asn163, Pro178, and Leu182
Spontaneous network activity visualized by ultrasensitive Ca 2+ indicators, yellow Cameleon-Nano
 nature methods Spontaneous network activity visualized by ultrasensitive Ca 2+ indicators, yellow Cameleon-Nano Kazuki Horikawa, Yoshiyuki Yamada, Tomoki Matsuda, Kentarou Kobayashi, Mitsuhiro Hashimoto,
nature methods Spontaneous network activity visualized by ultrasensitive Ca 2+ indicators, yellow Cameleon-Nano Kazuki Horikawa, Yoshiyuki Yamada, Tomoki Matsuda, Kentarou Kobayashi, Mitsuhiro Hashimoto,
Lecture 25 (11/15/17)
 Lecture 25 (11/15/17) Reading: Ch9; 328-332 Ch25; 990-995, 1005-1012 Problems: Ch9 (study-guide: applying); 1,2 Ch9 (study-guide: facts); 7,8 Ch25 (text); 1-3,5-7,9,10,13-15 Ch25 (study-guide: applying);
Lecture 25 (11/15/17) Reading: Ch9; 328-332 Ch25; 990-995, 1005-1012 Problems: Ch9 (study-guide: applying); 1,2 Ch9 (study-guide: facts); 7,8 Ch25 (text); 1-3,5-7,9,10,13-15 Ch25 (study-guide: applying);
FRET from Multiple Pathways in Fluorophore Labeled DNA
 Supporting Information for FRET from Multiple Pathways in Fluorophore Labeled DNA Joseph S. Melinger 1,*, Ani Khachatrian 1, Mario G. Ancona 1, Susan Buckhout-White 3, Ellen R. Goldman 3, Christopher M.
Supporting Information for FRET from Multiple Pathways in Fluorophore Labeled DNA Joseph S. Melinger 1,*, Ani Khachatrian 1, Mario G. Ancona 1, Susan Buckhout-White 3, Ellen R. Goldman 3, Christopher M.
Chapter 4 Fluorescence Resonance Energy Transfer (FRET) by Minor Groove-Associated Cyanine-Polyamide Conjugates
 Chapter 4 Fluorescence Resonance Energy Transfer (FRET) by Minor Groove-Associated Cyanine-Polyamide Conjugates The work described in this chapter was accomplished in collaboration with V. Rucker (Dervan
Chapter 4 Fluorescence Resonance Energy Transfer (FRET) by Minor Groove-Associated Cyanine-Polyamide Conjugates The work described in this chapter was accomplished in collaboration with V. Rucker (Dervan
Experiment 3: Fluorescence Spectroscopy I (continued) All life appears to be nurtured by the excitation of electrons by light in photosynthesis.
 Experiment 3: Fluorescence Spectroscopy I (continued) Last week: Part I.A: Introduction to steady state spectra Today: Part 1.B: Fluorescence Quenching and the Stern-Volmer Relation Prelab Lecture 9feb17
Experiment 3: Fluorescence Spectroscopy I (continued) Last week: Part I.A: Introduction to steady state spectra Today: Part 1.B: Fluorescence Quenching and the Stern-Volmer Relation Prelab Lecture 9feb17
PLA2 domain in parvoviruses The enzymatic features of viral PLA2
 PLA2 domain in parvoviruses Sequence analysis of 21 Pavovirinae members revealed that they contain an 80 amino acid conserved region in their VP1up. 20 amino acids out of the 80 were fully conserved and
PLA2 domain in parvoviruses Sequence analysis of 21 Pavovirinae members revealed that they contain an 80 amino acid conserved region in their VP1up. 20 amino acids out of the 80 were fully conserved and
Fluorescence quenching, Fluorescence anisotropy, Fluorescence resonance energy transfer (FRET)
 Fluorescence quenching, Fluorescence anisotropy, Fluorescence resonance energy transfer (FRET) Timescale of fluorescence processes The excited electron decay possibilities k f k ph k q k t k ic Biophysics
Fluorescence quenching, Fluorescence anisotropy, Fluorescence resonance energy transfer (FRET) Timescale of fluorescence processes The excited electron decay possibilities k f k ph k q k t k ic Biophysics
Comparison of LANCE Ultra TR-FRET to PerkinElmer s Classical LANCE TR-FRET Platform for Kinase Applications
 Comparison of LANCE Ultra TR-FRET to PerkinElmer s Classical LANCE TR-FRET Platform for Kinase Applications LANCE ULTRA TR-FRET TECHNOLOGY A P P L I C A T I O N N O T E Introduction Protein kinases play
Comparison of LANCE Ultra TR-FRET to PerkinElmer s Classical LANCE TR-FRET Platform for Kinase Applications LANCE ULTRA TR-FRET TECHNOLOGY A P P L I C A T I O N N O T E Introduction Protein kinases play
LanthaScreen Terbium-Labeled TR-FRET Anti-Epitope Tag Reagents - User Guide
 LanthaScreen Terbium-Labeled TR-FRET Anti-Epitope Tag Reagents - User Guide Cat. nos. PV3568, PV3569, PV4216, PV4217 Shipping Condition: Frozen Storage: -20 C Lit. no. PV421X.pps Rev. Date: 30 Aug 2005
LanthaScreen Terbium-Labeled TR-FRET Anti-Epitope Tag Reagents - User Guide Cat. nos. PV3568, PV3569, PV4216, PV4217 Shipping Condition: Frozen Storage: -20 C Lit. no. PV421X.pps Rev. Date: 30 Aug 2005
Supplementary Materials: A Toolbox of Genetically Encoded FRET-Based Biosensors for Rapid L-Lysine Analysis
 S1 of S9 Supplementary Materials: A Toolbox of Genetically Encoded FRET-Based Biosensors for Rapid L-Lysine Analysis Victoria Steffen, Julia Otten, Susann Engelmann, Andreas Radek, Michael Limberg, Bernd
S1 of S9 Supplementary Materials: A Toolbox of Genetically Encoded FRET-Based Biosensors for Rapid L-Lysine Analysis Victoria Steffen, Julia Otten, Susann Engelmann, Andreas Radek, Michael Limberg, Bernd
Supplementary Information. A superfolding Spinach2 reveals the dynamic nature of. trinucleotide repeat RNA
 Supplementary Information A superfolding Spinach2 reveals the dynamic nature of trinucleotide repeat RNA Rita L. Strack 1, Matthew D. Disney 2 & Samie R. Jaffrey 1 1 Department of Pharmacology, Weill Medical
Supplementary Information A superfolding Spinach2 reveals the dynamic nature of trinucleotide repeat RNA Rita L. Strack 1, Matthew D. Disney 2 & Samie R. Jaffrey 1 1 Department of Pharmacology, Weill Medical
Gene Expression: From Genes to Proteins
 The Flow of Genetic Information Gene Expression: From Genes to Proteins Chapter 9 Central Dogma in Molecular Biology molecule Gene 1 Strand to be transcribed Gene 2 Gene 3 strand Codon : Polymerase transcribes
The Flow of Genetic Information Gene Expression: From Genes to Proteins Chapter 9 Central Dogma in Molecular Biology molecule Gene 1 Strand to be transcribed Gene 2 Gene 3 strand Codon : Polymerase transcribes
Design. Construction. Characterization
 Design Construction Characterization DNA mrna (messenger) A C C transcription translation C A C protein His A T G C T A C G Plasmids replicon copy number incompatibility selection marker origin of replication
Design Construction Characterization DNA mrna (messenger) A C C transcription translation C A C protein His A T G C T A C G Plasmids replicon copy number incompatibility selection marker origin of replication
Confocal Microscopy Analyzes Cells
 Choosing Filters for Fluorescence A Laurin Publication Photonic Solutions for Biotechnology and Medicine November 2002 Confocal Microscopy Analyzes Cells Reprinted from the November 2002 issue of Biophotonics
Choosing Filters for Fluorescence A Laurin Publication Photonic Solutions for Biotechnology and Medicine November 2002 Confocal Microscopy Analyzes Cells Reprinted from the November 2002 issue of Biophotonics
BoTest A/E Botulinum Neurotoxin Detection Kit- Detailed Description
 Figure 1. BoTest A/E Reporter. (A) BioSentinel s BoTest A/E Reporter. CFP and YFP are connected by SNAP-25 (amino acids 141-206). The cleavage sites for each BoNT sero-type are indicated. (B) A FRET assay
Figure 1. BoTest A/E Reporter. (A) BioSentinel s BoTest A/E Reporter. CFP and YFP are connected by SNAP-25 (amino acids 141-206). The cleavage sites for each BoNT sero-type are indicated. (B) A FRET assay
TECHNICAL BULLETIN. In Vitro Bacterial Split Fluorescent Protein Fold n Glow Solubility Assay Kits
 In Vitro Bacterial Split Fluorescent Protein Fold n Glow Solubility Assay Kits Catalog Numbers APPA001 In Vitro Bacterial Split GFP "Fold 'n' Glow" Solubility Assay Kit (Green) APPA008 In Vitro Bacterial
In Vitro Bacterial Split Fluorescent Protein Fold n Glow Solubility Assay Kits Catalog Numbers APPA001 In Vitro Bacterial Split GFP "Fold 'n' Glow" Solubility Assay Kit (Green) APPA008 In Vitro Bacterial
Biochimica et Biophysica Acta
 Biochimica et Biophysica Acta 1833 (2013) 1787 1797 Contents lists available at SciVerse ScienceDirect Biochimica et Biophysica Acta journal homepage: www.elsevier.com/locate/bbamcr Genetically encoded
Biochimica et Biophysica Acta 1833 (2013) 1787 1797 Contents lists available at SciVerse ScienceDirect Biochimica et Biophysica Acta journal homepage: www.elsevier.com/locate/bbamcr Genetically encoded
Monitoring Protein:Protein Interactions in Living Cells Using NanoBRET Technology. Danette L. Daniels, Ph.D. Group Leader, Functional Proteomics
 Monitoring Protein:Protein Interactions in Living Cells Using NanoBRET Technology Danette L. Daniels, Ph.D. Group Leader, Functional Proteomics Proteomics and studying dynamic interactions AP-MS Proteomics
Monitoring Protein:Protein Interactions in Living Cells Using NanoBRET Technology Danette L. Daniels, Ph.D. Group Leader, Functional Proteomics Proteomics and studying dynamic interactions AP-MS Proteomics
SUPPLEMENTARY INFORMATION
 doi: 10.1038/nature08627 Supplementary Figure 1. DNA sequences used to construct nucleosomes in this work. a, DNA sequences containing the 601 positioning sequence (blue)24 with a PstI restriction site
doi: 10.1038/nature08627 Supplementary Figure 1. DNA sequences used to construct nucleosomes in this work. a, DNA sequences containing the 601 positioning sequence (blue)24 with a PstI restriction site
Special Techniques 1. Mark Scott FILM Facility
 Special Techniques 1 Mark Scott FILM Facility SPECIAL TECHNIQUES Multi-photon microscopy Second Harmonic Generation FRAP FRET FLIM In-vivo imaging TWO-PHOTON MICROSCOPY Alternative to confocal and deconvolution
Special Techniques 1 Mark Scott FILM Facility SPECIAL TECHNIQUES Multi-photon microscopy Second Harmonic Generation FRAP FRET FLIM In-vivo imaging TWO-PHOTON MICROSCOPY Alternative to confocal and deconvolution
Translation BIT 220 Chapter 13
 Translation BIT 220 Chapter 13 Making protein from mrna Most genes encode for proteins -some make RNA as end product Proteins -Monomer Amino Acid 20 amino acids -peptides -polypeptides -Structure of Amino
Translation BIT 220 Chapter 13 Making protein from mrna Most genes encode for proteins -some make RNA as end product Proteins -Monomer Amino Acid 20 amino acids -peptides -polypeptides -Structure of Amino
FLIM Fluorescence Lifetime IMaging
 FLIM Fluorescence Lifetime IMaging Fluorescence lifetime t I(t) = F0 exp( ) τ 1 τ = k f + k nr k nr = k IC + k ISC + k bl Batiaens et al, Trends in Cell Biology, 1999 τ τ = fluorescence lifetime (~ns to
FLIM Fluorescence Lifetime IMaging Fluorescence lifetime t I(t) = F0 exp( ) τ 1 τ = k f + k nr k nr = k IC + k ISC + k bl Batiaens et al, Trends in Cell Biology, 1999 τ τ = fluorescence lifetime (~ns to
Chapter 3.5. Protein Synthesis
 Chapter 3.5 Protein Synthesis Summary of Protein Synthesis How chemical Information is transfer during protein synthesis DNA mrna protein transcription the step from DNA to mrna occurs in the nucleus where
Chapter 3.5 Protein Synthesis Summary of Protein Synthesis How chemical Information is transfer during protein synthesis DNA mrna protein transcription the step from DNA to mrna occurs in the nucleus where
Metabolism BIOL 3702: Chapter 10
 Metabolism BIOL 3702: Chapter 10 Introduction to Metabolism u Metabolism is the sum total of all the chemical reactions occurring in a cell u Two major parts of metabolism: v Catabolism Ø Large, more complex
Metabolism BIOL 3702: Chapter 10 Introduction to Metabolism u Metabolism is the sum total of all the chemical reactions occurring in a cell u Two major parts of metabolism: v Catabolism Ø Large, more complex
Concept review: Fluorescence
 16 Concept review: Fluorescence Some definitions: Chromophore. The structural feature of a molecule responsible for the absorption of UV or visible light. Fluorophore. A chromophore that remits an absorbed
16 Concept review: Fluorescence Some definitions: Chromophore. The structural feature of a molecule responsible for the absorption of UV or visible light. Fluorophore. A chromophore that remits an absorbed
Biophysics of Macromolecules
 Biophysics of Macromolecules Lecture 18: In vivo Methods Braun/Lipfert SS 2015 How to create methods to probe macromolecules in vivo? 6. July 2015 Crowding alters Biochemical Equilibria Excluded volume
Biophysics of Macromolecules Lecture 18: In vivo Methods Braun/Lipfert SS 2015 How to create methods to probe macromolecules in vivo? 6. July 2015 Crowding alters Biochemical Equilibria Excluded volume
Application of Quantum Mechanics to Biology
 Application of Quantum Mechanics to Biology How can we apply quantum mechanics to biology? Polymers of nucleotides and amino acids - millions of atoms bounded into a large molecule Visual System Must turn
Application of Quantum Mechanics to Biology How can we apply quantum mechanics to biology? Polymers of nucleotides and amino acids - millions of atoms bounded into a large molecule Visual System Must turn
Where Are The Protein-synthesizing Instructions Stored On A Dna Molecule
 Where Are The Protein-synthesizing Instructions Stored On A Dna Molecule DNA contains all the information a cell needs in order to make certain proteins. Where are the protein-synthesizing instructions
Where Are The Protein-synthesizing Instructions Stored On A Dna Molecule DNA contains all the information a cell needs in order to make certain proteins. Where are the protein-synthesizing instructions
Supplementary Figure 1. FRET probe labeling locations in the Cas9-RNA-DNA complex.
 Supplementary Figure 1. FRET probe labeling locations in the Cas9-RNA-DNA complex. (a) Cy3 and Cy5 labeling locations shown in the crystal structure of Cas9-RNA bound to a cognate DNA target (PDB ID: 4UN3)
Supplementary Figure 1. FRET probe labeling locations in the Cas9-RNA-DNA complex. (a) Cy3 and Cy5 labeling locations shown in the crystal structure of Cas9-RNA bound to a cognate DNA target (PDB ID: 4UN3)
Masayoshi Honda, Jeehae Park, Robert A. Pugh, Taekjip Ha, and Maria Spies
 Molecular Cell, Volume 35 Supplemental Data Single-Molecule Analysis Reveals Differential Effect of ssdna-binding Proteins on DNA Translocation by XPD Helicase Masayoshi Honda, Jeehae Park, Robert A. Pugh,
Molecular Cell, Volume 35 Supplemental Data Single-Molecule Analysis Reveals Differential Effect of ssdna-binding Proteins on DNA Translocation by XPD Helicase Masayoshi Honda, Jeehae Park, Robert A. Pugh,
LIGAND BINDING LABORATORIES
 LIGAND BINDING LABORATORIES Laboratory Page Aims of laboratory 1 Introduction to the ligand binding studies 1 Direct methods of measuring ligand binding 3 EXPERIMENT 1 An Equilibrium Dialysis Study of
LIGAND BINDING LABORATORIES Laboratory Page Aims of laboratory 1 Introduction to the ligand binding studies 1 Direct methods of measuring ligand binding 3 EXPERIMENT 1 An Equilibrium Dialysis Study of
Chapters 31-32: Ribonucleic Acid (RNA)
 Chapters 31-32: Ribonucleic Acid (RNA) Short segments from the transcription, processing and translation sections of each chapter Slide 1 RNA In comparison with DNA RNA utilizes uracil in place of thymine
Chapters 31-32: Ribonucleic Acid (RNA) Short segments from the transcription, processing and translation sections of each chapter Slide 1 RNA In comparison with DNA RNA utilizes uracil in place of thymine
Tracking Cellular Protein Localization and Movement in Cells with a Flexible Fluorescent Labeling Technology. Chad Zimprich January 2015
 Tracking Cellular Protein Localization and Movement in Cells with a Flexible Fluorescent Labeling Technology Chad Zimprich January 2015 Presentation verview HaloTag Fusion Technology Design Functionality
Tracking Cellular Protein Localization and Movement in Cells with a Flexible Fluorescent Labeling Technology Chad Zimprich January 2015 Presentation verview HaloTag Fusion Technology Design Functionality
FLUORESCENCE. Matyas Molnar and Dirk Pacholsky
 FLUORESCENCE Matyas Molnar and Dirk Pacholsky 1 Information This lecture contains images and information from the following internet homepages http://micro.magnet.fsu.edu/primer/index.html http://www.microscopyu.com/
FLUORESCENCE Matyas Molnar and Dirk Pacholsky 1 Information This lecture contains images and information from the following internet homepages http://micro.magnet.fsu.edu/primer/index.html http://www.microscopyu.com/
MOLECULAR GENETICS PROTEIN SYNTHESIS. Molecular Genetics Activity #2 page 1
 AP BIOLOGY MOLECULAR GENETICS ACTIVITY #2 NAME DATE HOUR PROTEIN SYNTHESIS Molecular Genetics Activity #2 page 1 GENETIC CODE PROTEIN SYNTHESIS OVERVIEW Molecular Genetics Activity #2 page 2 PROTEIN SYNTHESIS
AP BIOLOGY MOLECULAR GENETICS ACTIVITY #2 NAME DATE HOUR PROTEIN SYNTHESIS Molecular Genetics Activity #2 page 1 GENETIC CODE PROTEIN SYNTHESIS OVERVIEW Molecular Genetics Activity #2 page 2 PROTEIN SYNTHESIS
Chapter 8 Lecture Outline. Transcription, Translation, and Bioinformatics
 Chapter 8 Lecture Outline Transcription, Translation, and Bioinformatics Replication, Transcription, Translation n Repetitive processes Build polymers of nucleotides or amino acids n All have 3 major steps
Chapter 8 Lecture Outline Transcription, Translation, and Bioinformatics Replication, Transcription, Translation n Repetitive processes Build polymers of nucleotides or amino acids n All have 3 major steps
Introduction. BRET Principle. K. M. Kroeger 1, K. A. Eidne 1, Bernd Hutter 2
 Bioluminescence Resonance Energy Transfer (BRET) as a means of monitoring dynamic receptor-protein interactions in living cells measured on LB 940 Mithras K. M. Kroeger 1, K. A. Eidne 1, Bernd Hutter 2
Bioluminescence Resonance Energy Transfer (BRET) as a means of monitoring dynamic receptor-protein interactions in living cells measured on LB 940 Mithras K. M. Kroeger 1, K. A. Eidne 1, Bernd Hutter 2
More on fluorescence
 More on fluorescence Last class Fluorescence Absorption emission Jablonski diagrams This class More on fluorescence Common fluorophores Jablonski diagrams to spectra Properties of fluorophores Excitation
More on fluorescence Last class Fluorescence Absorption emission Jablonski diagrams This class More on fluorescence Common fluorophores Jablonski diagrams to spectra Properties of fluorophores Excitation
Bioluminescence Resonance Energy Transfer (BRET 2 TM)
 BRET 2 Technology Application Note BRT-001 Bioluminescence Resonance Energy Transfer (BRET 2 TM) Principle, Applications and Products Introduction BRET 2 (Bioluminescence Resonance Energy Transfer) is
BRET 2 Technology Application Note BRT-001 Bioluminescence Resonance Energy Transfer (BRET 2 TM) Principle, Applications and Products Introduction BRET 2 (Bioluminescence Resonance Energy Transfer) is
BIO 315 Lab Exam I. Section #: Name:
 Section #: Name: Also provide this information on the computer grid sheet given to you. (Section # in special code box) BIO 315 Lab Exam I 1. In labeling the parts of a standard compound light microscope
Section #: Name: Also provide this information on the computer grid sheet given to you. (Section # in special code box) BIO 315 Lab Exam I 1. In labeling the parts of a standard compound light microscope
Transcription in Eukaryotes
 Transcription in Eukaryotes Biology I Hayder A Giha Transcription Transcription is a DNA-directed synthesis of RNA, which is the first step in gene expression. Gene expression, is transformation of the
Transcription in Eukaryotes Biology I Hayder A Giha Transcription Transcription is a DNA-directed synthesis of RNA, which is the first step in gene expression. Gene expression, is transformation of the
Text Reference, Campbell v.8, chapter 17 PROTEIN SYNTHESIS
 AP BIOLOGY Text Reference, Campbell v.8, chapter 17 ACTIVITY 1.22 NAME DATE HOUR PROTEIN SYNTHESIS GENETIC CODE PROTEIN SYNTHESIS OVERVIEW PROTEIN SYNTHESIS TRANSCRIPTION PROTEIN SYNTHESIS TRANSLATION
AP BIOLOGY Text Reference, Campbell v.8, chapter 17 ACTIVITY 1.22 NAME DATE HOUR PROTEIN SYNTHESIS GENETIC CODE PROTEIN SYNTHESIS OVERVIEW PROTEIN SYNTHESIS TRANSCRIPTION PROTEIN SYNTHESIS TRANSLATION
Roche Molecular Biochemicals Technical Note No. LC 6/99
 Roche Molecular Biochemicals Technical Note No. LC 6/99 LightCycler Selection of Hybridization Probe Sequences for Use with the LightCycler Olfert Landt and Andreas Nitsche, TIB MOLBIOL, Berlin 1. Introduction
Roche Molecular Biochemicals Technical Note No. LC 6/99 LightCycler Selection of Hybridization Probe Sequences for Use with the LightCycler Olfert Landt and Andreas Nitsche, TIB MOLBIOL, Berlin 1. Introduction
Development of an AlphaLISA MEK1 Kinase Assay Using Full-Length ERK2 Substrate
 TECHNICAL NOTE Development of an AlphaLISA MEK1 Kinase Assay Using Full-Length ERK2 Substrate AlphaLISA Technology Author: Jeanine Hinterneder, PhD PerkinElmer, Inc. Hopkinton, MA Introduction The mitogen-activated
TECHNICAL NOTE Development of an AlphaLISA MEK1 Kinase Assay Using Full-Length ERK2 Substrate AlphaLISA Technology Author: Jeanine Hinterneder, PhD PerkinElmer, Inc. Hopkinton, MA Introduction The mitogen-activated
DNA Transcription. Dr Aliwaini
 DNA Transcription 1 DNA Transcription-Introduction The synthesis of an RNA molecule from DNA is called Transcription. All eukaryotic cells have five major classes of RNA: ribosomal RNA (rrna), messenger
DNA Transcription 1 DNA Transcription-Introduction The synthesis of an RNA molecule from DNA is called Transcription. All eukaryotic cells have five major classes of RNA: ribosomal RNA (rrna), messenger
Gene function at the level of traits Gene function at the molecular level
 Gene expression Gene function at the level of traits Gene function at the molecular level Two levels tied together since the molecular level affects the structure and function of cells which determines
Gene expression Gene function at the level of traits Gene function at the molecular level Two levels tied together since the molecular level affects the structure and function of cells which determines
Fluorescence Light Microscopy for Cell Biology
 Fluorescence Light Microscopy for Cell Biology Why use light microscopy? Traditional questions that light microscopy has addressed: Structure within a cell Locations of specific molecules within a cell
Fluorescence Light Microscopy for Cell Biology Why use light microscopy? Traditional questions that light microscopy has addressed: Structure within a cell Locations of specific molecules within a cell
Monitoring and Optimizing the Lipopolysaccharides-plasmid DNA interaction by FLIM-FRET
 Transactions on Science and Technology Vol. 4, No. 3-3, 342-347, 2017 Monitoring and Optimizing the Lipopolysaccharides-plasmid DNA interaction by FLIM-FRET Nur Syahadatain Abdul Razak 1#, Clarence M.
Transactions on Science and Technology Vol. 4, No. 3-3, 342-347, 2017 Monitoring and Optimizing the Lipopolysaccharides-plasmid DNA interaction by FLIM-FRET Nur Syahadatain Abdul Razak 1#, Clarence M.
FLUORESCENT PEPTIDES. Outstanding Performance and Wide Application Range
 FLUORESCENT PEPTIDES Peptides and amino acids labeled with and Tide Quencher TM We offer peptides and amino acids tagged with fluorescent dyes. They meet highest demands in fluorescence intensity and photo-stability,
FLUORESCENT PEPTIDES Peptides and amino acids labeled with and Tide Quencher TM We offer peptides and amino acids tagged with fluorescent dyes. They meet highest demands in fluorescence intensity and photo-stability,
Supplementary Materials. Signal recognition particle binds to translating ribosomes before emergence of a signal anchor sequence
 Supplementary Materials Signal recognition particle binds to translating ribosomes before emergence of a signal anchor sequence Evan Mercier, Wolf Holtkamp, Marina V. Rodnina, Wolfgang Wintermeyer* Department
Supplementary Materials Signal recognition particle binds to translating ribosomes before emergence of a signal anchor sequence Evan Mercier, Wolf Holtkamp, Marina V. Rodnina, Wolfgang Wintermeyer* Department
5.36 Biochemistry Laboratory Spring 2009
 MIT OpenCourseWare http://ocw.mit.edu 5.36 Biochemistry Laboratory Spring 2009 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms. Laboratory Manual for URIECA
MIT OpenCourseWare http://ocw.mit.edu 5.36 Biochemistry Laboratory Spring 2009 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms. Laboratory Manual for URIECA
Stopped-Flow Fluorescence Polarisation/Anisotropy
 SX SERIES APPLICATION NOTE Stopped-Flow Fluorescence Polarisation/Anisotropy Abstract: Stopped-flow fluorescence polarisation/anisotropy is a highly useful technique for obtaining valuable kinetic information
SX SERIES APPLICATION NOTE Stopped-Flow Fluorescence Polarisation/Anisotropy Abstract: Stopped-flow fluorescence polarisation/anisotropy is a highly useful technique for obtaining valuable kinetic information
Chapter 17. From Gene to Protein
 Chapter 17 From Gene to Protein Overview: The Flow of Genetic Information The information content of DNA is in the form of specific sequences of nucleotides The DNA inherited by an organism leads to specific
Chapter 17 From Gene to Protein Overview: The Flow of Genetic Information The information content of DNA is in the form of specific sequences of nucleotides The DNA inherited by an organism leads to specific
Lab 1: Ensemble Fluorescence Basics
 Lab 1: Ensemble Fluorescence Basics This laboratory module is divided into two sections. The first one is on organic fluorophores, and the second one is on ensemble measurement of FRET (Fluorescence Resonance
Lab 1: Ensemble Fluorescence Basics This laboratory module is divided into two sections. The first one is on organic fluorophores, and the second one is on ensemble measurement of FRET (Fluorescence Resonance
Supporting Information for. Electrical control of Förster energy transfer.
 1 Supporting Information for Electrical control of Förster energy transfer. Klaus Becker 1, John M. Lupton 1*, Josef Müller 1, Andrey. L. Rogach 1, Dmitri V. Talapin, Horst Weller & Jochen Feldmann 1 1
1 Supporting Information for Electrical control of Förster energy transfer. Klaus Becker 1, John M. Lupton 1*, Josef Müller 1, Andrey. L. Rogach 1, Dmitri V. Talapin, Horst Weller & Jochen Feldmann 1 1
Fluorescence Resonance Energy Transfer (FRET) Microscopy
 Applications in Confocal Microscopy Fluorescence Resonance Energy Transfer (FRET) Microscopy Product Info Brochures Confocal Theory Java Tutorials Glossary Applications Image Gallery Resource Links The
Applications in Confocal Microscopy Fluorescence Resonance Energy Transfer (FRET) Microscopy Product Info Brochures Confocal Theory Java Tutorials Glossary Applications Image Gallery Resource Links The
Supporting Information: A GPCR dimerization interface in human cone opsins
 Supporting Information: A GPCR dimerization interface in human cone opsins Beata Jastrzebska *, William D. Comar *, Megan J. Kaliszewski, Kevin C. Skinner, Morgan H. Torcasio, Anthony S. Esway, Hui Jin,
Supporting Information: A GPCR dimerization interface in human cone opsins Beata Jastrzebska *, William D. Comar *, Megan J. Kaliszewski, Kevin C. Skinner, Morgan H. Torcasio, Anthony S. Esway, Hui Jin,
Calcium monitoring with the Mithras LB 940 multimode plate reader
 Calcium monitoring with the Mithras LB 940 multimode plate reader Tanja Scheikl 1, Marit Harzenetter 1, Bernd Hutter 2 Introduction Bivalent Calcium is an intracellular messenger in many eukaryotic signal
Calcium monitoring with the Mithras LB 940 multimode plate reader Tanja Scheikl 1, Marit Harzenetter 1, Bernd Hutter 2 Introduction Bivalent Calcium is an intracellular messenger in many eukaryotic signal
Alexander P. Savitsky A.N. Bach Institute of Biochemistry, Moscow 1 th German-Russian Symposium on Nanobiotechnology
 Alexander P. Savitsky A.N. Bach Institute of Biochemistry, Moscow 1 th German-Russian Symposium on Nanobiotechnology 15-17 ctober 2009 Lichtenwalde (Sachsen) Fluorescent detection systems - high sensitivity
Alexander P. Savitsky A.N. Bach Institute of Biochemistry, Moscow 1 th German-Russian Symposium on Nanobiotechnology 15-17 ctober 2009 Lichtenwalde (Sachsen) Fluorescent detection systems - high sensitivity
Fluorescence Spectroscopy. Student: Marin Cristina Antonia Coordinator:S.l. Preda Liliana
 Fluorescence Spectroscopy Student: Marin Cristina Antonia Coordinator:S.l. Preda Liliana Fluorescence Electron in the ground state is excited to a higher energy state After loss of some energy in vibrational
Fluorescence Spectroscopy Student: Marin Cristina Antonia Coordinator:S.l. Preda Liliana Fluorescence Electron in the ground state is excited to a higher energy state After loss of some energy in vibrational
Multiplexed 3D FRET imaging in deep tissue of live embryos Ming Zhao, Xiaoyang Wan, Yu Li, Weibin Zhou and Leilei Peng
 Scientific Reports Multiplexed 3D FRET imaging in deep tissue of live embryos Ming Zhao, Xiaoyang Wan, Yu Li, Weibin Zhou and Leilei Peng 1 Supplementary figures and notes Supplementary Figure S1 Volumetric
Scientific Reports Multiplexed 3D FRET imaging in deep tissue of live embryos Ming Zhao, Xiaoyang Wan, Yu Li, Weibin Zhou and Leilei Peng 1 Supplementary figures and notes Supplementary Figure S1 Volumetric
Chemical Biology, WS. Part II: Biology
 Chemical Biology, WS Part II: Biology 1 Markus Müller (Dipl. Biologe) Studium 1994-1999 Biologie in Marburg Doktorarbeit (Pflanzenphysiologie) 1999-2003 in Marburg Postdoc 2003-2006 in Umeå, Schweden Seit
Chemical Biology, WS Part II: Biology 1 Markus Müller (Dipl. Biologe) Studium 1994-1999 Biologie in Marburg Doktorarbeit (Pflanzenphysiologie) 1999-2003 in Marburg Postdoc 2003-2006 in Umeå, Schweden Seit
LanthaScreen Terbium Labeled TR-FRET Secondary Antibody Reagents-User Guide
 LanthaScreen Terbium Labeled TR-FRET Secondary Antibody Reagents-User Guide Cat. nos. PV3765, PV3767, PV3769, PV3771, PV3773, PV3775, PV3777, and PV3779 O-062132-r1 US 0405 TABLE OF CONTENTS 1.0 REAGENTS
LanthaScreen Terbium Labeled TR-FRET Secondary Antibody Reagents-User Guide Cat. nos. PV3765, PV3767, PV3769, PV3771, PV3773, PV3775, PV3777, and PV3779 O-062132-r1 US 0405 TABLE OF CONTENTS 1.0 REAGENTS
cell and tissue imaging by fluorescence microscopy
 cell and tissue imaging by fluorescence microscopy Steven NEDELLEC Plateforme Micropicell SFR Santé François Bonamy Nantes 1 A matter of size Limit of resolution 0.15mm aims: building the image of an object
cell and tissue imaging by fluorescence microscopy Steven NEDELLEC Plateforme Micropicell SFR Santé François Bonamy Nantes 1 A matter of size Limit of resolution 0.15mm aims: building the image of an object
E1 LITE - UBE1 Activity Assay Kit. Catalog Number UC105
 - 1 - MANUAL Catalog Number UC105 - 2 - BACKGROUND ABOUT THE ASSAY Conjugation of ubiquitin to a protein substrate involves the coordinated action of three separate proteins, an activating enzyme or E1,
- 1 - MANUAL Catalog Number UC105 - 2 - BACKGROUND ABOUT THE ASSAY Conjugation of ubiquitin to a protein substrate involves the coordinated action of three separate proteins, an activating enzyme or E1,
An Introduction to Fluorescence Resonance Energy Transfer (FRET) Technology and its Application in Bioscience
 W h i t e P a p e r An Introduction to Fluorescence Resonance Energy Transfer (FRET) Technology and its Application in Bioscience Paul Held, Laboratory Manager, Applications Department, BioTek Instruments,
W h i t e P a p e r An Introduction to Fluorescence Resonance Energy Transfer (FRET) Technology and its Application in Bioscience Paul Held, Laboratory Manager, Applications Department, BioTek Instruments,
SUPPLEMENTARY INFORMATION. Tolerance of a knotted near infrared fluorescent protein to random circular permutation
 SUPPLEMENTARY INFORMATION Tolerance of a knotted near infrared fluorescent protein to random circular permutation Naresh Pandey 1,3, Brianna E. Kuypers 2,4, Barbara Nassif 1, Emily E. Thomas 1,3, Razan
SUPPLEMENTARY INFORMATION Tolerance of a knotted near infrared fluorescent protein to random circular permutation Naresh Pandey 1,3, Brianna E. Kuypers 2,4, Barbara Nassif 1, Emily E. Thomas 1,3, Razan
Supplemental Information
 Supplemental Information ATP-dependent unwinding of U4/U6 snrnas by the Brr2 helicase requires the C-terminus of Prp8 Corina Maeder 1,3, Alan K. Kutach 1,2,3, and Christine Guthrie 1 1 Department of Biochemistry
Supplemental Information ATP-dependent unwinding of U4/U6 snrnas by the Brr2 helicase requires the C-terminus of Prp8 Corina Maeder 1,3, Alan K. Kutach 1,2,3, and Christine Guthrie 1 1 Department of Biochemistry
The Phasor approach: Application to FRET analysis and Tissue autofluorescence
 The Phasor approach: Application to FRET analysis and Tissue autofluorescence Laboratory for Fluorescence Dynamics University of California at Irvine lfd lfd Outline Background: Lifetime Intro to Fluorescence
The Phasor approach: Application to FRET analysis and Tissue autofluorescence Laboratory for Fluorescence Dynamics University of California at Irvine lfd lfd Outline Background: Lifetime Intro to Fluorescence
Chapter 14 Regulation of Transcription
 Chapter 14 Regulation of Transcription Cis-acting sequences Distance-independent cis-acting elements Dissecting regulatory elements Transcription factors Overview transcriptional regulation Transcription
Chapter 14 Regulation of Transcription Cis-acting sequences Distance-independent cis-acting elements Dissecting regulatory elements Transcription factors Overview transcriptional regulation Transcription
Genetically encoded fluorescent biosensors
 Genetically encoded fluorescent biosensors - fluorescent proteins attached to an additional protein sequence that make it sensitive to small biomolecules or other physiological intracellular processes
Genetically encoded fluorescent biosensors - fluorescent proteins attached to an additional protein sequence that make it sensitive to small biomolecules or other physiological intracellular processes
Synergy NEO HTS Multi-Mode Microplate Reader and Life Technologies Reagents
 A p p l i c a t i o n G u i d e Synergy NEO HTS Multi-Mode Microplate Reader and Life Technologies Reagents High Performance Multi-Detection Capabilities that Span the Small Molecule Drug Discovery Process
A p p l i c a t i o n G u i d e Synergy NEO HTS Multi-Mode Microplate Reader and Life Technologies Reagents High Performance Multi-Detection Capabilities that Span the Small Molecule Drug Discovery Process
Dear members of the Council,
 Dear members of the Council, Thank you for awarding me the ESC First Contact Initiative Grant, which offered me the opportunity to travel to the University of Arizona to closely collaborate with Professor
Dear members of the Council, Thank you for awarding me the ESC First Contact Initiative Grant, which offered me the opportunity to travel to the University of Arizona to closely collaborate with Professor
Biochemistry. Biochemical Techniques. 18 Spectrofluorimetry
 Description of Module Subject Name Paper Name 12 Module Name/Title 1. Objectives 1.1 To understand technique of Spectrofluorimetry. 1.2 To explain instrumentation design 1.3 What are applications of Spectrofluorimetry?
Description of Module Subject Name Paper Name 12 Module Name/Title 1. Objectives 1.1 To understand technique of Spectrofluorimetry. 1.2 To explain instrumentation design 1.3 What are applications of Spectrofluorimetry?
30 Gene expression: Transcription
 30 Gene expression: Transcription Gene structure. o Exons coding region of DNA. o Introns non-coding region of DNA. o Introns are interspersed between exons of a single gene. o Promoter region helps enzymes
30 Gene expression: Transcription Gene structure. o Exons coding region of DNA. o Introns non-coding region of DNA. o Introns are interspersed between exons of a single gene. o Promoter region helps enzymes
Metabolism. BIOL 3702: Chapter 10. Introduction to Metabolism. Energy and Work. BIOL 3702: Chapter 10 AY Dr. Cooper 1. Metabolism (cont.
 Metabolism BIOL 3702: Chapter 10 Introduction to Metabolism u Metabolism is the sum total of all the chemical reactions occurring in a cell u Two major parts of metabolism: v Catabolism Ø Large, more complex
Metabolism BIOL 3702: Chapter 10 Introduction to Metabolism u Metabolism is the sum total of all the chemical reactions occurring in a cell u Two major parts of metabolism: v Catabolism Ø Large, more complex
HEK293T. Fig. 1 in the
 Supplementary Information Supplementary Figure 1 Zinc uptake assay of hzip4 and hzip4-δecd transiently expressed in HEK293T cells. The results of one representative e experiment are shown in Fig. 1 in
Supplementary Information Supplementary Figure 1 Zinc uptake assay of hzip4 and hzip4-δecd transiently expressed in HEK293T cells. The results of one representative e experiment are shown in Fig. 1 in
Biochemistry 111. Carl Parker x A Braun
 Biochemistry 111 Carl Parker x6368 101A Braun csp@caltech.edu Central Dogma of Molecular Biology DNA-Dependent RNA Polymerase Requires a DNA Template Synthesizes RNA in a 5 to 3 direction Requires ribonucleoside
Biochemistry 111 Carl Parker x6368 101A Braun csp@caltech.edu Central Dogma of Molecular Biology DNA-Dependent RNA Polymerase Requires a DNA Template Synthesizes RNA in a 5 to 3 direction Requires ribonucleoside
