Investigating the interactions of FCS-304 and other Monoamine Oxidase (MAO) A Inhibitors Using Molecular Docking
|
|
|
- Virginia Matthews
- 6 years ago
- Views:
Transcription
1 Available online at J. Comput. Method. Mol. Design, 2011, 1 (3): ( ISSN : CODEN (USA): JCMMDA Investigating the interactions of FCS-304 and other Monoamine Oxidase (MAO) A Inhibitors Using Molecular Docking Vijay H. Masand a, Devidas T. Mahajan a, Anil M. Manikrao b, Niranjan Mahajan b,pravin N. Khatale b, Rahul D. Jawarkar b*, Taibi Ben Hadda c a Department of Chemistry,Vidyabharati College, Camp, Amaravati, Maharashtra, India b Department of Pharmaceutical Chemistry, Sahyadri College of Pharmacy, Methawde, Sangola, Solapur, Maharashtra, India. c Laboratoire Chimie Matériaux, Université Mohammed Premier, Oujda-60000, Morocco. ABSTRACT 4-propyl-2H-benzo[h]-chromen-2-one (FCS-304) is a derivative of coumarin which inhibits MAO-A. Selective MAO-A inhibitors are used in the treatment of neurological disorders such as depression. In this article, we have carried out molecular docking analysis of FCS-304 and other anti-depressent drugs to understand the structural features responsible for their potent antidepressant activity. The analysis reveals that the majority of interactions of FCS-304 are either hydrophobic or steric in nature whereas other anti-depressent drugs like Clorgyline, Iproniazide etc. have interacted MAO-A isoform through hydrophobic as well as polar interactions. In agreement with the previous results the presence of hydrophobic group at position number 4 is quite favorable. The present analysis indicates that presence of hydrogen donor at the end of chain present at the position 4 and hydrophobic group at position 8 could be advantageous. The analysis is useful for future drug designing. Keywords: MAO-A, FCS-304, Molecular docking, Drug Designing INTRODUCTION Coumarin (cromen-2-ona) is a major constituent of many natural as well as synthetic compounds. Not only coumarins are widely used in agricultural, paints, dye industries but have diverse pharmacological actions such as analgesic, anti-convulsant, neuro-protective, antidepressant etc [1-4]. Modern drug designing methods like QSAR, Molecular Docking etc. are widely used to understand the structural features responsible for pharmacological activities of 24
2 compounds [5]. The understanding can be utilized to increase the pharmacological properties of organic compounds. In continuation of our earlier work [6] herein we report the docking analysis of FCS-304 and comparison of its docking pose with marketed drugs as inhibitors of MAO [5, 6]. The objectives of this work are: (1) to understand structural characters responsible for MAO-A activity of FCS- 304 (2) to identify the types of interactions involved between FCS-304 with receptor site of Mao-A. (3) To compare docking pose of FCS-304 with marketed drugs (4) to analyze hydrophobic, electronic and steric characteristics of FCS-304. MATERIALS AND METHODS 2 Experimental Protocols: Computer-assisted simulated docking experiments were carried out in human MAO-A crystal structure (PDB ID: 2Z5X). Docking simulation study of FCS-304 using Argus Lab with the following protocol. (1) Enzyme structure was checked for missing atoms, bonds and contacts. Ramchandran plot was plotted to check the health of protein. (2) Hydrogen atoms were added to enzyme structure. Bound ligands were manually deleted from the enzyme. (3) The ligand molecules were constructed using ACD Chem Sketch 11.0 and optimized structure was used for docking. (4) The active site was generated and ligands were docked within the MAO-A active site. (5) The lowest energy conformation was selected and subjected to an energy minimization. RESULTS AND DISCUSSIONS 2.1 Modeling studies: Molecular modeling study and conformational alignment studies of the FCS-304 were performed in order to rationalize the obtained biological results (Fig. 1). The complete molecular docking studies were performed using human MAO-A crystal structure (PDB ID: 2z5x). Figure 1. (a) Energy minimized 3D conformation and (b) Molecular surface areas of FCS-304 red= mild polar, blue= H-bonding and white = hydrophobic region Before molecular docking the 3D structure of ligand was optimized. The pdb 2z5x was subjected to energy and residue optmisation. The health of protein was checked by plotting Ramchandran plot (Fig. 2). 25
3 (a) (b) Figure 2: (a) Docking pose of FCS-304 in the active site of MAO-A (b) Ramchandran Plot for pdb 2z5x after energy and residue optimization. Docking into the MAO-A active site revealed that the aliphatic chain at position 4 is responsible for hydrophobic interaction with Ile 180. In addition, the aromatic rings are also accountable for hydrophobic interaction with receptor. The polar part of molecule is more engaged in interaction with receptor. No H-bonding is present between FCS-304 and receptor (Fig.2). In order to get useful results that may be valuable in future drug designing we carried out receptor based electrostatic analysis also. (a) (b) Figure 3: (a) Structure of FCS-304 with numbering (b) Receptor based Electrostatics regions in the active site of MAO-A. Blue = donor, Red = acceptor and white = hydrophobic regions Figure 3 shows receptor based electrostatic regions around FCS-304. Interestingly, in agreement with the previous results [6], the presence of hydrophobic group at position number 4 is very favorable. A slight modification of chain at position 4 such as increasing the chain length with additional CH 2 -OH or simply OH may increase interaction with receptor resulting in tighter binding with MAO-A. The present analysis indicates that presence of hydrogen donor at the end of chain present at the position 4 and hydrophobic group at position 8 could be advantageous. 26
4 In order to get more productive and general results, we compared docking of FCS-304 with some marketed drugs like clorgyline, Iproniazide etc.; the docking poses show very interesting similarities and differences. Iproniazide Clorgyline Figure 4: a) Docking pose (b) Receptor based electrostatic contour map All MAO-A inhibitors have interact with residue like Tyr 407, Ile 180, Gln 215, Phe 208 etc. whereas clorogyline has additional interaction with Gly 67, Ala 68, Ile 207 that might be the reason behind making it to be very active even at very low concentration. Acknowledgement We are thankful to Dr. F. C. Raghuwanshi for his help and support throughout the work. REFERENCES [1] RD Jawarkar, VH Masand, KN Patil, DT Mahajan, MH Youssoufi, TB Hadda, SL Kumbhare, Der Pharm Chem, Accepted manuscript. [2]. SY Ariza, DC Rueda, J Rincón, EL Linares, MF Guerrero.VITAE, 2007,14 (2),
5 [3]. SY Kang, KY Lee, SH Sung, YC Kim. Journal of Natural Products, 2005,68(1), [4]. Y Chen, LD Kong, X Xia, HF Kung, L Zhang, Journal of Ethnopharmacology, 2005, 96 (3), [5]. VH Masand, KN Patil, DT Mahajan, RD Jawarkar, GM Nazerruddin, Der Pharma Chemica, 2010, 2(4), [6]. NE Vergel, JL López, F Orallo, D Viña, DM Buitrago, E Olmo, JA Mico, MF Guerrero, Life Sciences, 2010,86,
Design, docking study and ADME prediction of Chalcone derivatives as potent Tubulin inhibitors
 Journal of Computational Methods in Molecular Design, 2014, 4 (1):1-5 Scholars Research Library (http://scholarsresearchlibrary.com/archive.html) ISSN : 2231-3176 CODEN (USA): JCMMDA Design, docking study
Journal of Computational Methods in Molecular Design, 2014, 4 (1):1-5 Scholars Research Library (http://scholarsresearchlibrary.com/archive.html) ISSN : 2231-3176 CODEN (USA): JCMMDA Design, docking study
molecule on the screen in variety of representations and color schemes.
 4 5. DOCKING STUDIES: 5.. TOOLS AND MATERIALS USED 5... HEX Hex is an Interactive Molecular Graphics Program for calculating and displaying feasible docking modes of pairs of protein and DNA molecules.
4 5. DOCKING STUDIES: 5.. TOOLS AND MATERIALS USED 5... HEX Hex is an Interactive Molecular Graphics Program for calculating and displaying feasible docking modes of pairs of protein and DNA molecules.
Molecular Docking Study of Some Novel Nitroimidazo[1,2-b]pyridazine
![Molecular Docking Study of Some Novel Nitroimidazo[1,2-b]pyridazine Molecular Docking Study of Some Novel Nitroimidazo[1,2-b]pyridazine](/thumbs/95/125396186.jpg) DOI:10.7598/cst2016.1227 Chemical Science Transactions ISSN:2278-3458 2016, 5(3), 700-710 RESEARCH ARTICLE Molecular Docking Study of Some Novel Nitroimidazo[1,2-b]pyridazine based Heterocyclics on Methicillin
DOI:10.7598/cst2016.1227 Chemical Science Transactions ISSN:2278-3458 2016, 5(3), 700-710 RESEARCH ARTICLE Molecular Docking Study of Some Novel Nitroimidazo[1,2-b]pyridazine based Heterocyclics on Methicillin
Multi-Criteria Drug Discovery: using CoMFA models to drive target specificity. Lei Wang, Ph.D.
 Multi-Criteria Drug Discovery: using CoMFA models to drive target specificity Lei Wang, Ph.D. The Problem Compounds exist that have some desirable properties to become a drug but The known compounds are
Multi-Criteria Drug Discovery: using CoMFA models to drive target specificity Lei Wang, Ph.D. The Problem Compounds exist that have some desirable properties to become a drug but The known compounds are
1. Introduction. 2. Materials and Method
 American Journal of Pharmacological Sciences, 2016, Vol. 4, No. 1, 1-6 Available online at http://pubs.sciepub.com/ajps/4/1/1 Science and Education Publishing DOI:10.12691/ajps-4-1-1 In-silico Designing
American Journal of Pharmacological Sciences, 2016, Vol. 4, No. 1, 1-6 Available online at http://pubs.sciepub.com/ajps/4/1/1 Science and Education Publishing DOI:10.12691/ajps-4-1-1 In-silico Designing
BIRKBECK COLLEGE (University of London)
 BIRKBECK COLLEGE (University of London) SCHOOL OF BIOLOGICAL SCIENCES M.Sc. EXAMINATION FOR INTERNAL STUDENTS ON: Postgraduate Certificate in Principles of Protein Structure MSc Structural Molecular Biology
BIRKBECK COLLEGE (University of London) SCHOOL OF BIOLOGICAL SCIENCES M.Sc. EXAMINATION FOR INTERNAL STUDENTS ON: Postgraduate Certificate in Principles of Protein Structure MSc Structural Molecular Biology
Ligand docking and binding site analysis with pymol and autodock/vina
 International Journal of Basic and Applied Sciences, 4 (2) (2015) 168-177 www.sciencepubco.com/index.php/ijbas Science Publishing Corporation doi: 10.14419/ijbas.v4i2.4123 Research Paper Ligand docking
International Journal of Basic and Applied Sciences, 4 (2) (2015) 168-177 www.sciencepubco.com/index.php/ijbas Science Publishing Corporation doi: 10.14419/ijbas.v4i2.4123 Research Paper Ligand docking
X-ray structures of fructosyl peptide oxidases revealing residues responsible for gating oxygen access in the oxidative half reaction
 X-ray structures of fructosyl peptide oxidases revealing residues responsible for gating oxygen access in the oxidative half reaction Tomohisa Shimasaki 1, Hiromi Yoshida 2, Shigehiro Kamitori 2 & Koji
X-ray structures of fructosyl peptide oxidases revealing residues responsible for gating oxygen access in the oxidative half reaction Tomohisa Shimasaki 1, Hiromi Yoshida 2, Shigehiro Kamitori 2 & Koji
Ligand docking. CS/CME/Biophys/BMI 279 Oct. 22 and 27, 2015 Ron Dror
 Ligand docking CS/CME/Biophys/BMI 279 Oct. 22 and 27, 2015 Ron Dror 1 Outline Goals of ligand docking Defining binding affinity (strength) Computing binding affinity: Simplifying the problem Ligand docking
Ligand docking CS/CME/Biophys/BMI 279 Oct. 22 and 27, 2015 Ron Dror 1 Outline Goals of ligand docking Defining binding affinity (strength) Computing binding affinity: Simplifying the problem Ligand docking
Structural bioinformatics
 Structural bioinformatics Why structures? The representation of the molecules in 3D is more informative New properties of the molecules are revealed, which can not be detected by sequences Eran Eyal Plant
Structural bioinformatics Why structures? The representation of the molecules in 3D is more informative New properties of the molecules are revealed, which can not be detected by sequences Eran Eyal Plant
COMPUTER AIDED DRUG DESIGN: AN OVERVIEW
 Available online on 19.09.2018 at http://jddtonline.info Journal of Drug Delivery and Therapeutics Open Access to Pharmaceutical and Medical Research 2011-18, publisher and licensee JDDT, This is an Open
Available online on 19.09.2018 at http://jddtonline.info Journal of Drug Delivery and Therapeutics Open Access to Pharmaceutical and Medical Research 2011-18, publisher and licensee JDDT, This is an Open
6-Foot Mini Toober Activity
 Big Idea The interaction between the substrate and enzyme is highly specific. Even a slight change in shape of either the substrate or the enzyme may alter the efficient and selective ability of the enzyme
Big Idea The interaction between the substrate and enzyme is highly specific. Even a slight change in shape of either the substrate or the enzyme may alter the efficient and selective ability of the enzyme
MOLECULAR DOCKING STUDIES OF DESIGNED BENZAMIDE DERIVATIVES AS HISTONE DEACETYLASE2 INHIBITORS
 Innovare Academic Sciences International Journal of Pharmacy and Pharmaceutical Sciences ISSN- 0975-1491 Vol 6, Issue 4, 2014 Original Article MOLECULAR DOCKING STUDIES OF DESIGNED BENZAMIDE DERIVATIVES
Innovare Academic Sciences International Journal of Pharmacy and Pharmaceutical Sciences ISSN- 0975-1491 Vol 6, Issue 4, 2014 Original Article MOLECULAR DOCKING STUDIES OF DESIGNED BENZAMIDE DERIVATIVES
Suppl. Figure 1: RCC1 sequence and sequence alignments. (a) Amino acid
 Supplementary Figures Suppl. Figure 1: RCC1 sequence and sequence alignments. (a) Amino acid sequence of Drosophila RCC1. Same colors are for Figure 1 with sequence of β-wedge that interacts with Ran in
Supplementary Figures Suppl. Figure 1: RCC1 sequence and sequence alignments. (a) Amino acid sequence of Drosophila RCC1. Same colors are for Figure 1 with sequence of β-wedge that interacts with Ran in
Dr. R. Sankar, BSE 631 (2018)
 Pauling, Corey and Branson Diffraction of DNA http://www.nature.com/scitable/topicpage/dna-is-a-structure-that-encodes-biological-6493050 In short, stereochemistry is important in determining which helices
Pauling, Corey and Branson Diffraction of DNA http://www.nature.com/scitable/topicpage/dna-is-a-structure-that-encodes-biological-6493050 In short, stereochemistry is important in determining which helices
Docking Studies on a Series of Polo-like Kinase 4 Inhibitors
 2018 International Conference on Medicine Sciences and Bioengineering (ICMSB 2018) ISBN: 978-1-60595-560-5 Docking Studies on a Series of Polo-like Kinase 4 Inhibitors Ru DING 1,a, Juan LIU 2,b, Rong TAN
2018 International Conference on Medicine Sciences and Bioengineering (ICMSB 2018) ISBN: 978-1-60595-560-5 Docking Studies on a Series of Polo-like Kinase 4 Inhibitors Ru DING 1,a, Juan LIU 2,b, Rong TAN
Each enzyme has a unique 3-D shape and recognizes and binds only the specific substrate of a reaction.
 1 Enzyme = protein molecule that serves as a biological catalyst. allow life to go on. speed up and regulate metabolic reactions. Catalyst= a chemical that speeds up the rate of a reaction without itself
1 Enzyme = protein molecule that serves as a biological catalyst. allow life to go on. speed up and regulate metabolic reactions. Catalyst= a chemical that speeds up the rate of a reaction without itself
In silico drug designing studies on dengue virus envelope protein. Received: / Revised Accepted: / Published:
 World Journal of Pharmaceutical Sciences ISSN (Print): 2321-3310; ISSN (Online): 2321-3086 Available online at: http://www.wjpsonline.org/ Original Article In silico drug designing studies on dengue virus
World Journal of Pharmaceutical Sciences ISSN (Print): 2321-3310; ISSN (Online): 2321-3086 Available online at: http://www.wjpsonline.org/ Original Article In silico drug designing studies on dengue virus
In silico screening of 3,4 dihydropyrimidones as focal adhesion tyrosine kinase inhibitors
 Research Article ISS: 0974-6943 Gopinathan arasimhan et al. / Journal of Pharmacy Research 2015,9(1), Available online through www.jprsolutions.info In silico screening of 3,4 dihydropyrimidones as focal
Research Article ISS: 0974-6943 Gopinathan arasimhan et al. / Journal of Pharmacy Research 2015,9(1), Available online through www.jprsolutions.info In silico screening of 3,4 dihydropyrimidones as focal
Introduction to docking using VSDMIP 1.5
 Introduction to docking using VSDMIP 1.5 Alvaro Cortés Cabrera, Almudena Perona and Antonio Morreale Centro de Biología Molecular Severo Ochoa. UAM CSIC C/Nicolás Cabrera, 1. Campus UAM. 28049, Madrid.
Introduction to docking using VSDMIP 1.5 Alvaro Cortés Cabrera, Almudena Perona and Antonio Morreale Centro de Biología Molecular Severo Ochoa. UAM CSIC C/Nicolás Cabrera, 1. Campus UAM. 28049, Madrid.
Introduction to molecular docking
 Introduction to molecular docking Alvaro Cortes Cabrera, Javier Klett, Antonio Morreale Centro de Biología Molecular Severo Ochoa. UAM CSIC C/Nicolas Cabrera, 1. Campos Cantoblanco UAM. 28049, Madrid.
Introduction to molecular docking Alvaro Cortes Cabrera, Javier Klett, Antonio Morreale Centro de Biología Molecular Severo Ochoa. UAM CSIC C/Nicolas Cabrera, 1. Campos Cantoblanco UAM. 28049, Madrid.
VIRTUAL SCREENING STUDIES OF TWO CLOSELY RELATED WITHANOLIDES TO CONTROL CELL PROLIFERATION AND INDUCTION OF CELL SENESCENCE
 Rasayan J. Chem., 11(1), 339-344(2018) http://dx.doi.org/10.7324/rjc.2018.1111848 Vol. 11 No. 1 339-344 January - March 2018 ISSN: 0974-1496 e-issn: 0976-0083 CODEN: RJCABP http://www.rasayanjournal.com
Rasayan J. Chem., 11(1), 339-344(2018) http://dx.doi.org/10.7324/rjc.2018.1111848 Vol. 11 No. 1 339-344 January - March 2018 ISSN: 0974-1496 e-issn: 0976-0083 CODEN: RJCABP http://www.rasayanjournal.com
Pharmacophore modeling, virtual computational screening and biological evaluation studies
 Pharmacophore modeling, virtual computational screening and biological evaluation studies Drug discovery process plays an important role in identifying new investigational drug-likes and developing new
Pharmacophore modeling, virtual computational screening and biological evaluation studies Drug discovery process plays an important role in identifying new investigational drug-likes and developing new
Computer Simulation of Biosynthetic Modifications to Improve Binding Activity. by Kara Luo MIT PRIMES 2015 Mentored by Gil Alterovitz
 Computer Simulation of Biosynthetic Modifications to Improve Binding Activity by Kara Luo MIT PRIMES 2015 Mentored by Gil Alterovitz Super Bug - Enterococcus Faecium Potentially lethal, worldwide infection
Computer Simulation of Biosynthetic Modifications to Improve Binding Activity by Kara Luo MIT PRIMES 2015 Mentored by Gil Alterovitz Super Bug - Enterococcus Faecium Potentially lethal, worldwide infection
Electronic Supplementary Information
 Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 2018 Electronic Supplementary Information Unraveling the effects of amino acid substitutions
Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 2018 Electronic Supplementary Information Unraveling the effects of amino acid substitutions
Synthetic Libraries of linear Oligomers
 ynthetic Libraries of linear ligomers tephan eiffert Peptides: highly valuable lead structures for drug discovery Peptidomimetics * high ability to bind selectively at a large number of biological receptors
ynthetic Libraries of linear ligomers tephan eiffert Peptides: highly valuable lead structures for drug discovery Peptidomimetics * high ability to bind selectively at a large number of biological receptors
Solving Structure Based Design Problems using Discovery Studio 1.7 Building a Flexible Docking Protocol
 Solving Structure Based Design Problems using Discovery Studio 1.7 Building a Flexible Docking Protocol C. M. (Venkat) Venkatachalam Fellow, Life Sciences Dipesh Risal Marketing, Life Sciences Overview
Solving Structure Based Design Problems using Discovery Studio 1.7 Building a Flexible Docking Protocol C. M. (Venkat) Venkatachalam Fellow, Life Sciences Dipesh Risal Marketing, Life Sciences Overview
Structure and Function of the First Full-Length Murein Peptide Ligase (Mpl) Cell Wall Recycling Protein
 Paper Presentation PLoS ONE 2011 Structure and Function of the First Full-Length Murein Peptide Ligase (Mpl) Cell Wall Recycling Protein Debanu Das, Mireille Herve, Julie Feuerhelm, etc. and Dominique
Paper Presentation PLoS ONE 2011 Structure and Function of the First Full-Length Murein Peptide Ligase (Mpl) Cell Wall Recycling Protein Debanu Das, Mireille Herve, Julie Feuerhelm, etc. and Dominique
Drug DNA interaction. Modeling DNA ligand interaction of intercalating ligands
 Drug DNA interaction DNA as carrier of genetic information is a major target for drug interaction because of the ability to interfere with transcription (gene expression and protein synthesis) and DNA
Drug DNA interaction DNA as carrier of genetic information is a major target for drug interaction because of the ability to interfere with transcription (gene expression and protein synthesis) and DNA
NANYANG TECHNOLOGICAL UNIVERSITY SEMESTER I EXAMINATION CBC922 Medicinal Chemistry. NOVEMBER TIME ALLOWED: 120 min
 AYAG TECLGICAL UIVERSITY SEMESTER I EXAMIATI 2006-2007 CBC922 Medicinal Chemistry VEMBER 2006 - TIME ALLWED 120 min ISTRUCTIS T CADIDATES 1. This examination paper contains TW (2) parts and comprises SIX
AYAG TECLGICAL UIVERSITY SEMESTER I EXAMIATI 2006-2007 CBC922 Medicinal Chemistry VEMBER 2006 - TIME ALLWED 120 min ISTRUCTIS T CADIDATES 1. This examination paper contains TW (2) parts and comprises SIX
Recent Progress of Interprotein s Research Activities. Interprotein Corporation
 Recent Progress of Interprotein s Research Activities - INTENDD and AI-guided INTENDD - Interprotein Corporation 1 Interprotein Corporation Location: Osaka, Japan Year Established: 2001 CEO & President:
Recent Progress of Interprotein s Research Activities - INTENDD and AI-guided INTENDD - Interprotein Corporation 1 Interprotein Corporation Location: Osaka, Japan Year Established: 2001 CEO & President:
From mechanism to medicne
 From mechanism to medicne a look at proteins and drug design Chem 342 δ δ δ+ M 2009 δ+ δ+ δ M Drug Design - an Iterative Approach @ DSU Structural Analysis of Receptor Structural Analysis of Ligand-Receptor
From mechanism to medicne a look at proteins and drug design Chem 342 δ δ δ+ M 2009 δ+ δ+ δ M Drug Design - an Iterative Approach @ DSU Structural Analysis of Receptor Structural Analysis of Ligand-Receptor
Amino Acids and Proteins
 Various Functions of Proteins SB203 Amino Acids and Proteins Jirundon Yuvaniyama, Ph.D. Department of Biochemistry Faculty of Science Mahidol University Enzymes Transport proteins utrient and storage proteins
Various Functions of Proteins SB203 Amino Acids and Proteins Jirundon Yuvaniyama, Ph.D. Department of Biochemistry Faculty of Science Mahidol University Enzymes Transport proteins utrient and storage proteins
Fundamentals of Protein Structure
 Outline Fundamentals of Protein Structure Yu (Julie) Chen and Thomas Funkhouser Princeton University CS597A, Fall 2005 Protein structure Primary Secondary Tertiary Quaternary Forces and factors Levels
Outline Fundamentals of Protein Structure Yu (Julie) Chen and Thomas Funkhouser Princeton University CS597A, Fall 2005 Protein structure Primary Secondary Tertiary Quaternary Forces and factors Levels
If you wish to have extra practice with swiss pdb viewer or to familiarize yourself with how to use the program here is a tutorial:
 Name (s): Swiss PDB viewer assignment chapter 4. If you wish to have extra practice with swiss pdb viewer or to familiarize yourself with how to use the program here is a tutorial: http://spdbv.vital-it.ch/themolecularlevel/spvtut/index.html
Name (s): Swiss PDB viewer assignment chapter 4. If you wish to have extra practice with swiss pdb viewer or to familiarize yourself with how to use the program here is a tutorial: http://spdbv.vital-it.ch/themolecularlevel/spvtut/index.html
INTERNATIONAL JOURNAL OF RESEARCH IN PHARMACY AND CHEMISTRY
 ITERATIAL JURAL F RESEARCH I PHARMACY AD CHEMISTRY Available online at www.ijrpc.com Research Article PHARMACPHRE BASED 3D QSAR AALYSIS F A EW CLASS F HIGH- AFFIITY LIGADS FR ECDYSE RECEPTRS FR DEVELPIG
ITERATIAL JURAL F RESEARCH I PHARMACY AD CHEMISTRY Available online at www.ijrpc.com Research Article PHARMACPHRE BASED 3D QSAR AALYSIS F A EW CLASS F HIGH- AFFIITY LIGADS FR ECDYSE RECEPTRS FR DEVELPIG
Alexandre R. Gingras, Umakhanth Venkatraman Girija, Anthony H. Keeble, Roshni Panchal, Daniel A. Mitchell, Peter C.E. Moody, and Russell Wallis
 Structure, 19 Supplemental Information Structural Basis of Mannan-Binding Lectin Recognition by Its Associated Serine Protease MASP-1: Implications for Complement Activation Alexandre R. Gingras, Umakhanth
Structure, 19 Supplemental Information Structural Basis of Mannan-Binding Lectin Recognition by Its Associated Serine Protease MASP-1: Implications for Complement Activation Alexandre R. Gingras, Umakhanth
What s New in Discovery Studio 2.5.5
 What s New in Discovery Studio 2.5.5 Discovery Studio takes modeling and simulations to the next level. It brings together the power of validated science on a customizable platform for drug discovery research.
What s New in Discovery Studio 2.5.5 Discovery Studio takes modeling and simulations to the next level. It brings together the power of validated science on a customizable platform for drug discovery research.
Molekulare Mechanismen der Signaltransduktion
 Molekulare Mechanismen der Signaltransduktion 12 Mechanism of auxin perception slides: http://tinyurl.com/modul-mms doi: 10.1038/nature05731 previous model http://www.plantcell.org/content/17/9/2425/f1.large.jpg
Molekulare Mechanismen der Signaltransduktion 12 Mechanism of auxin perception slides: http://tinyurl.com/modul-mms doi: 10.1038/nature05731 previous model http://www.plantcell.org/content/17/9/2425/f1.large.jpg
Specificity: Induced Fit
 Specificity: Induced Fit Conformational changes may occur upon ligand binding (Daniel Koshland in 1958) This adaptation is called the induced fit Induced fit allows for tighter binding of the ligand Induced
Specificity: Induced Fit Conformational changes may occur upon ligand binding (Daniel Koshland in 1958) This adaptation is called the induced fit Induced fit allows for tighter binding of the ligand Induced
Research Article INTRODUCTION. R. Priyadharsini 1 *, T. Kalaivani 1, M. Keerthana 1, U. Logeshwari 1, S. Nivedhitha 1, S. Bhuvaneswari 2 ABSTRACT
 Research Article In silico identification of newer potential spleen tyrosine kinase and protein arginine deiminase 4 inhibitors as potent antirheumatoid arthritic agents R. Priyadharsini 1 *, T. Kalaivani
Research Article In silico identification of newer potential spleen tyrosine kinase and protein arginine deiminase 4 inhibitors as potent antirheumatoid arthritic agents R. Priyadharsini 1 *, T. Kalaivani
DNA/Protein Binding, Molecular Docking and in Vitro Anti-cancer Activity of some Thioether-Dipyrrinato Complexes
 DNA/Protein Binding, Molecular Docking and in Vitro Anti-cancer Activity of some Thioether-Dipyrrinato Complexes Rakesh Kumar Gupta, Gunjan Sharma, ξ Rampal Pandey, Amit Kumar, Biplob Koch, ξ Pei- Zhou
DNA/Protein Binding, Molecular Docking and in Vitro Anti-cancer Activity of some Thioether-Dipyrrinato Complexes Rakesh Kumar Gupta, Gunjan Sharma, ξ Rampal Pandey, Amit Kumar, Biplob Koch, ξ Pei- Zhou
Docking studies on short and ultra-short peptides against beta-secretase (BACE1) enzyme
 Research Article Docking studies on short and ultra-short peptides against beta-secretase (BACE1) enzyme Pandurangan Perumal 1, Liew Chun Lap 1, Ling Yu Luong 1, Tan Ling Hoi 1, Pavadai Parasuraman 2 ABSTRACT
Research Article Docking studies on short and ultra-short peptides against beta-secretase (BACE1) enzyme Pandurangan Perumal 1, Liew Chun Lap 1, Ling Yu Luong 1, Tan Ling Hoi 1, Pavadai Parasuraman 2 ABSTRACT
Docking Studies of the Binding Mode of Dictyostatin and Its Analogues to the Taxoid Binding Site on Tubulin
 Docking Studies of the Binding Mode of Dictyostatin and Its Analogues to the Taxoid Binding Site on Tubulin Christopher B. Hackmeyer Spring Hill College Billy W. Day, Ph.D. Department of Pharmaceutical
Docking Studies of the Binding Mode of Dictyostatin and Its Analogues to the Taxoid Binding Site on Tubulin Christopher B. Hackmeyer Spring Hill College Billy W. Day, Ph.D. Department of Pharmaceutical
LIST OF ACRONYMS & ABBREVIATIONS
 LIST OF ACRONYMS & ABBREVIATIONS ALA ARG ASN ATD CRD CYS GLN GLU GLY GPCR HIS hstr ILE LEU LYS MET mglur1 NHDC PDB PHE PRO SER T1R2 T1R3 TMD TRP TYR THR 7-TM VFTM ZnSO 4 Alanine, A Arginine, R Asparagines,
LIST OF ACRONYMS & ABBREVIATIONS ALA ARG ASN ATD CRD CYS GLN GLU GLY GPCR HIS hstr ILE LEU LYS MET mglur1 NHDC PDB PHE PRO SER T1R2 T1R3 TMD TRP TYR THR 7-TM VFTM ZnSO 4 Alanine, A Arginine, R Asparagines,
Antibody-Antigen recognition. Structural Biology Weekend Seminar Annegret Kramer
 Antibody-Antigen recognition Structural Biology Weekend Seminar 10.07.2005 Annegret Kramer Contents Function and structure of antibodies Features of antibody-antigen interfaces Examples of antibody-antigen
Antibody-Antigen recognition Structural Biology Weekend Seminar 10.07.2005 Annegret Kramer Contents Function and structure of antibodies Features of antibody-antigen interfaces Examples of antibody-antigen
CHAPTER Discovery of novel HSP90 inhibitors. One promising therapeutic strategy for treating cancer is to
 106 CHAPTER 4 4.1 Introduction 4. Discovery of novel HSP90 inhibitors ne promising therapeutic strategy for treating cancer is to specifically target signal transduction pathways that have a key role in
106 CHAPTER 4 4.1 Introduction 4. Discovery of novel HSP90 inhibitors ne promising therapeutic strategy for treating cancer is to specifically target signal transduction pathways that have a key role in
Supporting Information Contents
 Supporting Information Choy Theng Loh, Kiyoshi Ozawa, Kellie L. Tuck, Nicholas Barlow, Thomas Huber, Gottfried Otting, and Bim Graham Lanthanide tags for site-specific ligation to an unnatural amino acid
Supporting Information Choy Theng Loh, Kiyoshi Ozawa, Kellie L. Tuck, Nicholas Barlow, Thomas Huber, Gottfried Otting, and Bim Graham Lanthanide tags for site-specific ligation to an unnatural amino acid
Advanced in silico Drug Design KFC/ADD Tutorial: Molecular Docking Intro. Karel Berka, Ph.D. Jindřich Fanfrlík, Ph.D. Martin Lepšík, Ph.D.
 Advanced in silico Drug Design KFC/ADD Tutorial: Molecular Docking Intro Karel Berka, Ph.D. Jindřich Fanfrlík, Ph.D. Martin Lepšík, Ph.D. UP Olomouc, 1.-3.2. 2016 Tutorial preparation Programs installation:
Advanced in silico Drug Design KFC/ADD Tutorial: Molecular Docking Intro Karel Berka, Ph.D. Jindřich Fanfrlík, Ph.D. Martin Lepšík, Ph.D. UP Olomouc, 1.-3.2. 2016 Tutorial preparation Programs installation:
1) The penicillin family of antibiotics, discovered by Alexander Fleming in 1928, has the following general structure: O O
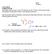 ame: TF ame: LS1a Fall 06 Problem Set #3 Due Friday 10/13 at noon in your TF s drop box on the 2 nd floor of the Science Center All questions including the (*extra*) ones should be turned in 1) The penicillin
ame: TF ame: LS1a Fall 06 Problem Set #3 Due Friday 10/13 at noon in your TF s drop box on the 2 nd floor of the Science Center All questions including the (*extra*) ones should be turned in 1) The penicillin
Solutions to 7.02 Quiz II 10/27/05
 Solutions to 7.02 Quiz II 10/27/05 Class Average = 83 Standard Deviation = 9 Range Grade % 87-100 A 43 74-86 B 39 55-73 C 17 > 54 D 1 Question 1 (56 points) While studying deep sea bacteria, you discover
Solutions to 7.02 Quiz II 10/27/05 Class Average = 83 Standard Deviation = 9 Range Grade % 87-100 A 43 74-86 B 39 55-73 C 17 > 54 D 1 Question 1 (56 points) While studying deep sea bacteria, you discover
Programme Good morning and summary of last week Levels of Protein Structure - I Levels of Protein Structure - II
 Programme 8.00-8.10 Good morning and summary of last week 8.10-8.30 Levels of Protein Structure - I 8.30-9.00 Levels of Protein Structure - II 9.00-9.15 Break 9.15-11.15 Exercise: Building a protein model
Programme 8.00-8.10 Good morning and summary of last week 8.10-8.30 Levels of Protein Structure - I 8.30-9.00 Levels of Protein Structure - II 9.00-9.15 Break 9.15-11.15 Exercise: Building a protein model
Structure and Possible Mechanism of the CcbJ Methyltransferase from Streptomyces caelestis
 Supplemental material to accompany Structure and Possible Mechanism of the CcbJ Methyltransferase from Streptomyces caelestis Jacob Bauer, a Gabriela Ondrovičová, a Lucie Najmanová, b Vladimír Pevala,
Supplemental material to accompany Structure and Possible Mechanism of the CcbJ Methyltransferase from Streptomyces caelestis Jacob Bauer, a Gabriela Ondrovičová, a Lucie Najmanová, b Vladimír Pevala,
MOLECULAR DOCKING STUDIES FOR IDENTIFICATION OF POTENT PHOSPHOINOSITIDE 3-KINASES (PI3KS) INHIBITORS
 CHAPTER 7 MLECULAR DCKIG STUDIES FR IDETIFICATI F PTET PHSPHISITIDE 3-KIASES (PI3KS) IHIBITRS 7.1 Introduction Phosphoinositide 3 kinases (PI3Ks) constitute a class of enzymes that catalyse phosphorylation
CHAPTER 7 MLECULAR DCKIG STUDIES FR IDETIFICATI F PTET PHSPHISITIDE 3-KIASES (PI3KS) IHIBITRS 7.1 Introduction Phosphoinositide 3 kinases (PI3Ks) constitute a class of enzymes that catalyse phosphorylation
BC 367, Exam 2 November 13, Part I. Multiple Choice (3 pts each)- Please circle the single best answer.
 Name BC 367, Exam 2 November 13, 2008 Part I. Multiple Choice (3 pts each)- Please circle the single best answer. 1. The enzyme pyruvate dehydrogenase catalyzes the following reaction. What kind of enzyme
Name BC 367, Exam 2 November 13, 2008 Part I. Multiple Choice (3 pts each)- Please circle the single best answer. 1. The enzyme pyruvate dehydrogenase catalyzes the following reaction. What kind of enzyme
Molecular docking studies of indole derivatives containing cyanide group as hepatitis C Ns5b polymerase inhibitor
 2015; 4(9): 43-48 ISS: 2277-7695 TPI 2015; 4(9): 43-48 2015 TPI www.thepharmajournal.com Received: 22-09-2015 Accepted: 26-10-2015 Thayaillany Rajandran a) Faculty of Pharmacy, Asia Metropolitan University,
2015; 4(9): 43-48 ISS: 2277-7695 TPI 2015; 4(9): 43-48 2015 TPI www.thepharmajournal.com Received: 22-09-2015 Accepted: 26-10-2015 Thayaillany Rajandran a) Faculty of Pharmacy, Asia Metropolitan University,
Can inhibitors of HIV integrase work on HCV polymerase?
 [245g] Can inhibitors of HIV integrase work on HCV polymerase? Marino Artico 1, Roberta Costi 1, Roberto Di Santo 1, Maurizio Fermeglia 2, Marco Ferrone 2, Stefano Manfredini 3, Sabrina Pricl 2 1 Department
[245g] Can inhibitors of HIV integrase work on HCV polymerase? Marino Artico 1, Roberta Costi 1, Roberto Di Santo 1, Maurizio Fermeglia 2, Marco Ferrone 2, Stefano Manfredini 3, Sabrina Pricl 2 1 Department
Targeting of the disease related proteome by small molecules
 Targeting of the disease related proteome by small molecules Workshop on chemical information 2018 EPFL Modest v. Korff utline Introduction Data sources Diseases Proteins Compounds Data relations Diseases
Targeting of the disease related proteome by small molecules Workshop on chemical information 2018 EPFL Modest v. Korff utline Introduction Data sources Diseases Proteins Compounds Data relations Diseases
CS597A Structural Bioinformatics
 Outline CS597A Thomas Funkhouser Princeton University Fall 27 Overview of structural bioinformatics Challenges Applications Overview of course Lectures Coursework Projects Bioinformatics Definition: The
Outline CS597A Thomas Funkhouser Princeton University Fall 27 Overview of structural bioinformatics Challenges Applications Overview of course Lectures Coursework Projects Bioinformatics Definition: The
ECS 129: Structural Bioinformatics March 15, 2016
 Notes: ES 129: Structural Bioinformatics March 15, 2016 1) The final exam is open book, open notes. 2) The final is divided into 2 parts, and graded over 100 points (with 8 points extra credit) 3) You
Notes: ES 129: Structural Bioinformatics March 15, 2016 1) The final exam is open book, open notes. 2) The final is divided into 2 parts, and graded over 100 points (with 8 points extra credit) 3) You
Solutions to Problem Set 1
 MIT Department of Biology 7.014 Introductory Biology, Spring 004 Question 1 Solutions to 7.014 Problem Set 1 a) Describe the conditions of the atmosphere on prebiotic earth and how these conditions differ
MIT Department of Biology 7.014 Introductory Biology, Spring 004 Question 1 Solutions to 7.014 Problem Set 1 a) Describe the conditions of the atmosphere on prebiotic earth and how these conditions differ
Introduction to Drug Design and Discovery
 Introduction to Drug Design and Discovery Course: Drug Design Course code: 0510412 Dr. Balakumar Chandrasekaran Dr. Bilal Al-Jaidi Assistant Professors, Pharmaceutical Medicinal Chemistry, Faculty of Pharmacy,
Introduction to Drug Design and Discovery Course: Drug Design Course code: 0510412 Dr. Balakumar Chandrasekaran Dr. Bilal Al-Jaidi Assistant Professors, Pharmaceutical Medicinal Chemistry, Faculty of Pharmacy,
AN ALTERNATIVE TO ENDOSULFAN: STRUCTURE BASED PHARMACOPHORE MODELING, GLIDE AND MM-GBSA ANALYSIS OF ECDYSONE RECEPTOR AND MOLECULAR DYNAMICS STUDIES
 International Journal of Bio-Technology and Research (IJBTR) ISSN(P): 2249-6858; ISSN(E): 2249-796X Vol. 5, Issue 1, Feb 2015, 1-10 TJPRC Pvt. Ltd. AN ALTERNATIVE TO ENDOSULFAN: STRUCTURE BASED PHARMACOPHORE
International Journal of Bio-Technology and Research (IJBTR) ISSN(P): 2249-6858; ISSN(E): 2249-796X Vol. 5, Issue 1, Feb 2015, 1-10 TJPRC Pvt. Ltd. AN ALTERNATIVE TO ENDOSULFAN: STRUCTURE BASED PHARMACOPHORE
Conformational properties of enzymes
 Conformational properties of enzymes; Physics of enzyme substrate interactions; Electronic conformational interactions, cooperative properties of enzymes Mitesh Shrestha Conformational properties of enzymes
Conformational properties of enzymes; Physics of enzyme substrate interactions; Electronic conformational interactions, cooperative properties of enzymes Mitesh Shrestha Conformational properties of enzymes
BMB/Bi/Ch 170 Fall 2017 Problem Set 1: Proteins I
 BMB/Bi/Ch 170 Fall 2017 Problem Set 1: Proteins I Please use ray-tracing feature for all the images you are submitting. Use either the Ray button on the right side of the command window in PyMOL or variations
BMB/Bi/Ch 170 Fall 2017 Problem Set 1: Proteins I Please use ray-tracing feature for all the images you are submitting. Use either the Ray button on the right side of the command window in PyMOL or variations
RNA as drug target: docking studies. Florent Barbault ITODYS CNRS (UMR7086) Univ. Paris7 Group of Molec. Modeling & Chem. Info.
 RNA as drug target: docking studies Florent Barbault ITODYS CNRS (UMR7086) Univ. Paris7 Group of Molec. Modeling & Chem. Info. Overview Why targeting RNA Drug design strategy Parametrisation of the scoring
RNA as drug target: docking studies Florent Barbault ITODYS CNRS (UMR7086) Univ. Paris7 Group of Molec. Modeling & Chem. Info. Overview Why targeting RNA Drug design strategy Parametrisation of the scoring
Computational methods in drug discovery
 Computational methods in drug discovery COURSE HIGHLIGHTS v Structure-based drug discovery Ø Binding site identification Ø Molecular docking v Biologics Ø Molecular modelling Ø Protein-Protein docking
Computational methods in drug discovery COURSE HIGHLIGHTS v Structure-based drug discovery Ø Binding site identification Ø Molecular docking v Biologics Ø Molecular modelling Ø Protein-Protein docking
RMSD-BASED CLUSTERING AND QR FACTORIZATION. Rob Swift, UCI Tuesday, August 2, 2011
 RMSD-BASED CLUSTERING AND QR FACTORIZATION Rob Swift, UCI Tuesday, August 2, 2011 Outline Biomolecular flexibility, a few examples CADD pipeline overview RMSD clustering QR factorization Our evolving ideas
RMSD-BASED CLUSTERING AND QR FACTORIZATION Rob Swift, UCI Tuesday, August 2, 2011 Outline Biomolecular flexibility, a few examples CADD pipeline overview RMSD clustering QR factorization Our evolving ideas
Supplementary Information for. Structure of human tyrosylprotein sulfotransferase-2 reveals the mechanism of protein tyrosine sulfation reaction
 Supplementary Information for Structure of human tyrosylprotein sulfotransferase-2 reveals the mechanism of protein tyrosine sulfation reaction Takamasa Teramoto, Yukari Fujikawa, Yoshirou Kawaguchi, Katsuhisa
Supplementary Information for Structure of human tyrosylprotein sulfotransferase-2 reveals the mechanism of protein tyrosine sulfation reaction Takamasa Teramoto, Yukari Fujikawa, Yoshirou Kawaguchi, Katsuhisa
Subtract v1.0: Measuring the volume of protein binding site cavities. Panos Kakoulidis, Ioannis Galdadas and Zoe Cournia
 Subtract v1.0: Measuring the volume of protein binding site cavities Panos Kakoulidis, Ioannis Galdadas and Zoe Cournia Biomedical Research Foundation, 4 Soranou Ephessiou, 11527, Athens, Greece Researchers
Subtract v1.0: Measuring the volume of protein binding site cavities Panos Kakoulidis, Ioannis Galdadas and Zoe Cournia Biomedical Research Foundation, 4 Soranou Ephessiou, 11527, Athens, Greece Researchers
Interaction of 4-Nitroaniline with Serum Albumin
 Applied Mechanics and Materials Online: 214-2-6 ISSN: 1662-7482, Vols. 522-524, pp 337-34 doi:1.428/www.scientific.net/amm.522-524.337 214 Trans Tech Publications, Switzerland Interaction of 4-Nitroaniline
Applied Mechanics and Materials Online: 214-2-6 ISSN: 1662-7482, Vols. 522-524, pp 337-34 doi:1.428/www.scientific.net/amm.522-524.337 214 Trans Tech Publications, Switzerland Interaction of 4-Nitroaniline
 Problem: The GC base pairs are more stable than AT base pairs. Why? 5. Triple-stranded DNA was first observed in 1957. Scientists later discovered that the formation of triplestranded DNA involves a type
Problem: The GC base pairs are more stable than AT base pairs. Why? 5. Triple-stranded DNA was first observed in 1957. Scientists later discovered that the formation of triplestranded DNA involves a type
Ligand immobilization using thiol-disulphide exchange
 A P P L I C A T I T E 9 Ligand immobilization using thiol-disulphide exchange Abstract This Application ote describes an immobilization procedure based on thioldisulphide exchange, providing a valuable
A P P L I C A T I T E 9 Ligand immobilization using thiol-disulphide exchange Abstract This Application ote describes an immobilization procedure based on thioldisulphide exchange, providing a valuable
Drug Design 2. Overview. Drug Discovery Pipeline. Drug Design. Oliver Kohlbacher Winter 2009/ Success Stories
 Drug Design 2 Oliver Kohlbacher Winter 2009/2010 13. Success Stories Abt. Simulation biologischer Systeme WSI/ZBIT, Eberhard Karls Universität Tübingen Overview Drug Discovery Pipeline reloaded Successful
Drug Design 2 Oliver Kohlbacher Winter 2009/2010 13. Success Stories Abt. Simulation biologischer Systeme WSI/ZBIT, Eberhard Karls Universität Tübingen Overview Drug Discovery Pipeline reloaded Successful
Bioinformation Volume 5 open access
 Molecular docking analysis of new generation cephalosporins interactions with recently known SHV-variants Asad Ullah Khan 1 *, Mohd Hassan Baig 1, Gulshan Wadhwa 2 1 Interdisciplinary Biotechnology Unit,
Molecular docking analysis of new generation cephalosporins interactions with recently known SHV-variants Asad Ullah Khan 1 *, Mohd Hassan Baig 1, Gulshan Wadhwa 2 1 Interdisciplinary Biotechnology Unit,
Basic concepts of molecular biology
 Basic concepts of molecular biology Gabriella Trucco Email: gabriella.trucco@unimi.it Life The main actors in the chemistry of life are molecules called proteins nucleic acids Proteins: many different
Basic concepts of molecular biology Gabriella Trucco Email: gabriella.trucco@unimi.it Life The main actors in the chemistry of life are molecules called proteins nucleic acids Proteins: many different
The Disordered Structural Ensembles of Vasopressin and Oxytocin and Their Mutants
 The Disordered Structural Ensembles of Vasopressin and Oxytocin and Their Mutants Eugene Yedvabny, Paul S. Nerenberg 4, Clare So and Teresa Head-Gordon,,3* Department of Chemistry, Department of Bioengineering,
The Disordered Structural Ensembles of Vasopressin and Oxytocin and Their Mutants Eugene Yedvabny, Paul S. Nerenberg 4, Clare So and Teresa Head-Gordon,,3* Department of Chemistry, Department of Bioengineering,
Fig. 1. A schematic illustration of pipeline from gene to drug : integration of virtual and real experiments.
 1 Integration of Structure-Based Drug Design and SPR Biosensor Technology in Discovery of New Lead Compounds Alexis S. Ivanov Institute of Biomedical Chemistry RAMS, 10, Pogodinskaya str., Moscow, 119121,
1 Integration of Structure-Based Drug Design and SPR Biosensor Technology in Discovery of New Lead Compounds Alexis S. Ivanov Institute of Biomedical Chemistry RAMS, 10, Pogodinskaya str., Moscow, 119121,
Drug Discovery Targeting the GHKL Superfamily - HSP90, DNA Gyrase, Histidine Kinases
 Drug Discovery Targeting the GHKL Superfamily - HSP90, DNA Gyrase, Histidine Kinases (Jason) Xinshan Kang, Boris Klebansky, Richard Fine BioPredict, Inc., 660 KinderKamack Rd., radell, NJ 07649 232 nd
Drug Discovery Targeting the GHKL Superfamily - HSP90, DNA Gyrase, Histidine Kinases (Jason) Xinshan Kang, Boris Klebansky, Richard Fine BioPredict, Inc., 660 KinderKamack Rd., radell, NJ 07649 232 nd
Research Article. Development of rifampicin derivatives sensitive to the rpob mutated Mycobacterium tuberculosis: An insilico approach
 Available online www.jocpr.com Journal of Chemical and Pharmaceutical Research, 2015, 7(5):153-159 Research Article ISSN : 0975-7384 CODEN(USA) : JCPRC5 Development of rifampicin derivatives sensitive
Available online www.jocpr.com Journal of Chemical and Pharmaceutical Research, 2015, 7(5):153-159 Research Article ISSN : 0975-7384 CODEN(USA) : JCPRC5 Development of rifampicin derivatives sensitive
Docking. Why? Docking : finding the binding orientation of two molecules with known structures
 Docking : finding the binding orientation of two molecules with known structures Docking According to the molecules involved: Protein-Ligand docking Protein-Protein docking Specific docking algorithms
Docking : finding the binding orientation of two molecules with known structures Docking According to the molecules involved: Protein-Ligand docking Protein-Protein docking Specific docking algorithms
Ch Biophysical Chemistry
 Ch 247.53. Biophysical Chemistry Nina Rosario L. Rojas 2012-2013 sem 1 Review of Protein Structure Why structure? Primary, secondary, tertiary structure Disulfide bonds scheme 2 STRUCTURE- REGULAR STRUCTURE
Ch 247.53. Biophysical Chemistry Nina Rosario L. Rojas 2012-2013 sem 1 Review of Protein Structure Why structure? Primary, secondary, tertiary structure Disulfide bonds scheme 2 STRUCTURE- REGULAR STRUCTURE
Compound Re-Profiling. Dr Robert Scoffin CEO, Cresset
 Compound Re-Profiling Dr Robert Scoffin CEO, Cresset What is it? F F F N N H 2 N S O O Br > Compound Re-Profiling or Re-Purposing is the process of finding a new clinical use for an existing treatment
Compound Re-Profiling Dr Robert Scoffin CEO, Cresset What is it? F F F N N H 2 N S O O Br > Compound Re-Profiling or Re-Purposing is the process of finding a new clinical use for an existing treatment
Abbreviations. Inhibitor synthesis
 Supplementary Material I for: Bailey et al. manuscript An analysis of subdomain orientation, conformational change and disorder in relation to crystal packing of aspartic proteinases. Abbreviations Boc
Supplementary Material I for: Bailey et al. manuscript An analysis of subdomain orientation, conformational change and disorder in relation to crystal packing of aspartic proteinases. Abbreviations Boc
COMP364. Introduction to RNA secondary structure prediction. Jérôme Waldispühl School of Computer Science, McGill
 COMP364 Introduction to RNA secondary structure prediction Jérôme Waldispühl School of Computer Science, McGill RNA world In prebiotic world, RNA thought to have filled two distinct roles: 1. an information
COMP364 Introduction to RNA secondary structure prediction Jérôme Waldispühl School of Computer Science, McGill RNA world In prebiotic world, RNA thought to have filled two distinct roles: 1. an information
Prediction of Protein-Protein Binding Sites and Epitope Mapping. Copyright 2017 Chemical Computing Group ULC All Rights Reserved.
 Prediction of Protein-Protein Binding Sites and Epitope Mapping Epitope Mapping Antibodies interact with antigens at epitopes Epitope is collection residues on antigen Continuous (sequence) or non-continuous
Prediction of Protein-Protein Binding Sites and Epitope Mapping Epitope Mapping Antibodies interact with antigens at epitopes Epitope is collection residues on antigen Continuous (sequence) or non-continuous
Protocol S1: Supporting Information
 Protocol S1: Supporting Information Basis for the specificity of the kinase domain of Abl for peptide substrates The crystal structures reported in this work were obtained using two different ATP analog-peptide
Protocol S1: Supporting Information Basis for the specificity of the kinase domain of Abl for peptide substrates The crystal structures reported in this work were obtained using two different ATP analog-peptide
Four levels of protein Structure
 Proteins (polypeptides) Four levels of protein Structure Primary Structure (1 structure): Secondary Structure (2 structure): Tertiary Structure (3 structure): Quaternary Structure (4 structure): Proteins
Proteins (polypeptides) Four levels of protein Structure Primary Structure (1 structure): Secondary Structure (2 structure): Tertiary Structure (3 structure): Quaternary Structure (4 structure): Proteins
Fluorescense. Aromatic molecules often fluoresce (but DNA bases don't) Excitation ~ absorbance (Differences due to: ν h ' Fluorescence
 Fluorescense Two types of spectra: Excitation: Detection λ fixed and scan excitation λ. Emission: Excitation λ fixed (usually at A max ) & scan detection λ. What goes up must come down Radiationless decay
Fluorescense Two types of spectra: Excitation: Detection λ fixed and scan excitation λ. Emission: Excitation λ fixed (usually at A max ) & scan detection λ. What goes up must come down Radiationless decay
2018 Protein Modeling Exam Key
 2018 Protein Modeling Exam Key Multiple Choice: 1. Which of the following amino acids has a negative charge at ph 7? a. Gln b. Glu c. Ser d. Cys 2. Which of the following is an example of secondary structure?
2018 Protein Modeling Exam Key Multiple Choice: 1. Which of the following amino acids has a negative charge at ph 7? a. Gln b. Glu c. Ser d. Cys 2. Which of the following is an example of secondary structure?
CURRICULUM VITAE. Lecturer (Pharmaceutical Chemistry Department, Faculty of Pharmacy, Al- Azhar University, Cairo, Egypt)
 CURRICULUM VITAE Farag F. S. Selim, PhD CONTACT DETAILS Name: Farag Farouk Sherbiny Selim Nationality: Egyptian Position: Lecturer (Pharmaceutical Chemistry Department, Faculty of Pharmacy, Al- Azhar University,
CURRICULUM VITAE Farag F. S. Selim, PhD CONTACT DETAILS Name: Farag Farouk Sherbiny Selim Nationality: Egyptian Position: Lecturer (Pharmaceutical Chemistry Department, Faculty of Pharmacy, Al- Azhar University,
systemsdock Operation Manual
 systemsdock Operation Manual Version 2.0 2016 April systemsdock is being developed by Okinawa Institute of Science and Technology http://www.oist.jp/ Integrated Open Systems Unit http://openbiology.unit.oist.jp/_new/
systemsdock Operation Manual Version 2.0 2016 April systemsdock is being developed by Okinawa Institute of Science and Technology http://www.oist.jp/ Integrated Open Systems Unit http://openbiology.unit.oist.jp/_new/
Protein Folding Problem I400: Introduction to Bioinformatics
 Protein Folding Problem I400: Introduction to Bioinformatics November 29, 2004 Protein biomolecule, macromolecule more than 50% of the dry weight of cells is proteins polymer of amino acids connected into
Protein Folding Problem I400: Introduction to Bioinformatics November 29, 2004 Protein biomolecule, macromolecule more than 50% of the dry weight of cells is proteins polymer of amino acids connected into
Supporting Information. A general chemiluminescence strategy. for measuring aptamer-target binding and target concentration
 Supporting Information A general chemiluminescence strategy for measuring aptamer-target binding and target concentration Shiyuan Li, Duyu Chen, Qingtong Zhou, Wei Wang, Lingfeng Gao, Jie Jiang, Haojun
Supporting Information A general chemiluminescence strategy for measuring aptamer-target binding and target concentration Shiyuan Li, Duyu Chen, Qingtong Zhou, Wei Wang, Lingfeng Gao, Jie Jiang, Haojun
Pharmaceutical Informatics and the Pathway to Personalized Medicines
 Pharmaceutical Informatics and the Pathway to Personalized Medicines Sangtae Kim and Venkat Venkatasubramanian Purdue University ICCS2007 DDDAS Workshop Beijing May 26-28, 2007 Outline The Grand Challenge
Pharmaceutical Informatics and the Pathway to Personalized Medicines Sangtae Kim and Venkat Venkatasubramanian Purdue University ICCS2007 DDDAS Workshop Beijing May 26-28, 2007 Outline The Grand Challenge
Protein-Ligand Analysis with SID
 Protein-Ligand Analysis with SID A new post-simulation analysis tool Dmitry Lupyan, PhD Understanding molecular processes with MD Drug discovery and design Protein-protein interactions + Protein-DNA interactions
Protein-Ligand Analysis with SID A new post-simulation analysis tool Dmitry Lupyan, PhD Understanding molecular processes with MD Drug discovery and design Protein-protein interactions + Protein-DNA interactions
SUPPLEMENTARY INFORMATION Figures. Supplementary Figure 1 a. Page 1 of 30. Nature Chemical Biology: doi: /nchembio.2528
 SUPPLEMENTARY INFORMATION Figures Supplementary Figure 1 a b c Page 1 of 0 11 Supplementary Figure 1: Biochemical characterisation and binding validation of the reversible USP inhibitor 1. a, Biochemical
SUPPLEMENTARY INFORMATION Figures Supplementary Figure 1 a b c Page 1 of 0 11 Supplementary Figure 1: Biochemical characterisation and binding validation of the reversible USP inhibitor 1. a, Biochemical
static MM_Index snap(mm_index corect, MM_Index ligct, int imatch0, int *moleatoms, i
 BIOLUMINATE static MM_Index snap(mm_index corect, MM_Index ligct, int imatch0, int *moleatoms, int *refcoreatoms){int ncoreat = :vector mappings; PhpCoreMapping mapping; for COMMON(glidelig).
BIOLUMINATE static MM_Index snap(mm_index corect, MM_Index ligct, int imatch0, int *moleatoms, int *refcoreatoms){int ncoreat = :vector mappings; PhpCoreMapping mapping; for COMMON(glidelig).
Insilico Investigation of Potential Inhibitors Targetting Protein Associated with Vitiligo Disorder
 Volume: 3, Issue: 8, 4-8 Aug 2015 www.biosciencejournals.com ISSN: 2321-9122 Impact Factor: 3.742 Assistant Professor, Department of Biotechnology, Selvam College of Technology, Namakkal-637001. Insilico
Volume: 3, Issue: 8, 4-8 Aug 2015 www.biosciencejournals.com ISSN: 2321-9122 Impact Factor: 3.742 Assistant Professor, Department of Biotechnology, Selvam College of Technology, Namakkal-637001. Insilico
4/10/2011. Rosetta software package. Rosetta.. Conformational sampling and scoring of models in Rosetta.
 Rosetta.. Ph.D. Thomas M. Frimurer Novo Nordisk Foundation Center for Potein Reseach Center for Basic Metabilic Research Breif introduction to Rosetta Rosetta docking example Rosetta software package Breif
Rosetta.. Ph.D. Thomas M. Frimurer Novo Nordisk Foundation Center for Potein Reseach Center for Basic Metabilic Research Breif introduction to Rosetta Rosetta docking example Rosetta software package Breif
