Supplementary Information
|
|
|
- Allen James
- 6 years ago
- Views:
Transcription
1 Journal : Nature Biotechnology Supplementary Information Targeted genome engineering in human cells with RNA-guided endonucleases Seung Woo Cho, Sojung Kim, Jong Min Kim, and Jin-Soo Kim* National Creative Research Initiatives Center for Genome Engineering and Department of Chemistry, Seoul National University, Gwanak-gu, Seoul , South Korea These authors contributed equally to this work. *Correspondence should be addressed to J.-S.K. (jskim01@snu.ac.kr). Table of Contents Supplementary Methods Supplementary Figure 1. Amino-acid sequence of the expressed version of Cas9 used in this study. Supplementary Figure 2. Reproducible genome editing in human cells with RGENs. Supplementary Figure 3. DNA sequences of mutant clones. Supplementary Figure 4. CCR5-specific RGEN did not induce off-target-associated chromosomal deletions. Supplementary Figure 5. Stable maintenance of RGEN-induced mutant cells. Supplementary Figure 6. RGEN-induced DSBs in cells detected by TP53BP1 staining. Supplementary Table 1. Oligonucleotides used in this study. 1
2 Supplementary Methods Construction of Cas9-encoding plasmids. The Cas9-coding sequence (4,104 bp), derived from Streptococcus pyogenes strain M1 GAS (NC_ ), was reconstituted using the human codon usage table and synthesized using oligonucleotides. First, 1-kbp DNA segments were assembled using overlapping ~35-mer oligonucleotides and Phusion polymerase (New England Biolabs) and cloned into T-vector (SolGent). A full-length Cas9 sequence was assembled using four 1-kbp DNA segments by overlap PCR. The Cas9- encoding DNA segment was subcloned into p3s, which was derived from pcdna3.1 (Invitrogen). In this vector, the Cas9 protein is expressed under the control of the CMV promoter and is fused to a peptide tag (NH 2 -GGSGPPKKKRKVYPYDVPDYA-COOH) containing the HA epitope and a nuclear localization signal (NLS) at the C-terminus (Supplementary Fig. 1). Expression and nuclear localization of the Cas9 protein in K562 cells were confirmed by western blotting using anti-ha antibody (Santa Cruz). In vitro DNA cleavage assay. The Cas9 cassette was subcloned into pet28-b(+) and transformed into BL21(DE3). The expression of Cas9 was induced using 0.5 mm IPTG for 4 h at 25. The Cas9 protein containing the His 6 -tag at the C terminus was purified using Ni- NTA agarose resin (Qiagen) and dialyzed against 20 mm HEPES (ph 7.5), 150 mm KCl, 1 mm DTT, and 10% glycerol. Purified Cas9 (50 nm) was incubated with super-coiled or predigested plasmid DNA (300 ng) and sgrna (50 nm) in a reaction volume of 20 μl in NEB buffer 3 for 1 h at 37. Digested DNA was analyzed by electrophoresis using 0.8% agarose gels. RNA preparation. RNA was in vitro transcribed through run-off reactions by T7 RNA polymerase using the MEGAshortscript T7 Kit (Ambion) according to the manufacturer s manual. Templates for RNA in vitro transcription were generated by annealing two complementary oligonuceotides (Supplementary Table 1). Transcribed RNA was resolved on a 8% denaturing urea-page gel. The gel slice containing RNA was cut out and transferred to elution buffer. RNA was recovered in nuclease-free water followed by phenol:chloroform extraction, chloroform extraction, and ethanol precipitation. Purified RNA was quantified by spectrometry. Cell culture. HEK 293T/17 (ATCC, CRL-11268) cells were maintained in Dulbecco s modified Eagle s medium (DMEM) supplemented with 100 units/ml penicillin, 100 μg/ml 2
3 streptomycin, 0.1 mm nonessential amino acids, and 10% fetal bovine serum (FBS). K562 (ATCC, CCL-243) cells were grown in RPMI-1640 with 10% FBS and the penicillin/streptomycin mix (100 U/ml and 100 μg/ml, respectively). Genome-editing assay. To introduce DSBs in mammalian cells using RGENs, 2x10 6 K562 cells were transfected with 20 μg of Cas9-encoding plasmid using the 4D-Nucleofector, SF Cell Line 4D-Nucleofector X Kit, Program FF-120 (Lonza) according to the manufacturer s protocol. After 24 h, 20 to 160 μg of in vitro transcribed sgrna was transfected into 1x10 6 K562 cells. Cells were collected at various time points after RNA transfection and genomic DNA was isolated. The region including the target site was PCR-amplified using the primers described in Supplementary Table 1. The amplicons were subjected to the T7E1 assay as described previously 6. For sequencing analysis, PCR products corresponding to genomic modifications were purified and cloned into the T-Blunt vector using the T-Blunt PCR Cloning Kit (SolGent). Cloned products were sequenced using the M13 primer. Reporter construct. The RFP-GFP reporter plasmids used in this study were constructed as described previously 5. Oligonucleotides corresponding to target sites (Supplementary Table 1) were synthesized (Macrogen) and annealed. The annealed oligonucleotides were ligated into a reporter vector digested with EcoRI and BamHI. Episomal reporter assay. HEK 293T cells were co-transfected with Cas9-encoding plasmid (0.8 μg) and the RFP-GFP reporter plasmid (0.2 μg) in a 24-well plate using Lipofectamine 2000 (Invitrogen). At 12h post transfection, sgrna (1 μg) prepared by in vitro transcription was transfected using Lipofectamine At 3 d post-transfection, transfected cells were subjected to flow cytometry and cells expressing both RFP and GFP were counted. 3
4 Supplementary Figures Supplementary Figure 1. Amino-acid sequence of the expressed version of Cas9 used in this study. The amino-acid sequence of the Cas9 protein encoded in our Cas9 expression plasmid is presented. The sequence of the nuclear localization signal and the HA epitope at the C-terminus is underlined. MDKKYSIGLDIGTNSVGWAVITDEYKVPSKKFKVLGNTDRHSIKKNLIGALLFDSGETAE 60 ATRLKRTARRRYTRRKNRICYLQEIFSNEMAKVDDSFFHRLEESFLVEEDKKHERHPIFG 180 NIVDEVAYHEKYPTIYHLRKKLVDSTDKADLRLIYLALAHMIKFRGHFLIEGDLNPDNSD 240 VDKLFIQLVQTYNQLFEENPINASGVDAKAILSARLSKSRRLENLIAQLPGEKKNGLFGN 300 LIALSLGLTPNFKSNFDLAEDAKLQLSKDTYDDDLDNLLAQIGDQYADLFLAAKNLSDAI 360 LLSDILRVNTEITKAPLSASMIKRYDEHHQDLTLLKALVRQQLPEKYKEIFFDQSKNGYA 420 GYIDGGASQEEFYKFIKPILEKMDGTEELLVKLNREDLLRKQRTFDNGSIPHQIHLGELH 480 AILRRQEDFYPFLKDNREKIEKILTFRIPYYVGPLARGNSRFAWMTRKSEETITPWNFEE 540 VVDKGASAQSFIERMTNFDKNLPNEKVLPKHSLLYEYFTVYNELTKVKYVTEGMRKPAFL 600 SGEQKKAIVDLLFKTNRKVTVKQLKEDYFKKIECFDSVEISGVEDRFNASLGTYHDLLKI 660 IKDKDFLDNEENEDILEDIVLTLTLFEDREMIEERLKTYAHLFDDKVMKQLKRRRYTGWG 720 RLSRKLINGIRDKQSGKTILDFLKSDGFANRNFMQLIHDDSLTFKEDIQKAQVSGQGDSL 780 HEHIANLAGSPAIKKGILQTVKVVDELVKVMGRHKPENIVIEMARENQTTQKGQKNSRER 840 MKRIEEGIKELGSQILKEHPVENTQLQNEKLYLYYLQNGRDMYVDQELDINRLSDYDVDH 900 IVPQSFLKDDSIDNKVLTRSDKNRGKSDNVPSEEVVKKMKNYWRQLLNAKLITQRKFDNL 960 TKAERGGLSELDKAGFIKRQLVETRQITKHVAQILDSRMNTKYDENDKLIREVKVITLKS 1020 KLVSDFRKDFQFYKVREINNYHHAHDAYLNAVVGTALIKKYPKLESEFVYGDYKVYDVRK 1080 MIAKSEQEIGKATAKYFFYSNIMNFFKTEITLANGEIRKRPLIETNGETGEIVWDKGRDF 1140 ATVRKVLSMPQVNIVKKTEVQTGGFSKESILPKRNSDKLIARKKDWDPKKYGGFDSPTVA 1200 YSVLVVAKVEKGKSKKLKSVKELLGITIMERSSFEKNPIDFLEAKGYKEVKKDLIIKLPK 1260 YSLFELENGRKRMLASAGELQKGNELALPSKYVNFLYLASHYEKLKGSPEDNEQKQLFVE 1320 QHKHYLDEIIEQISEFSKRVILADANLDKVLSAYNKHRDKPIREQAENIIHLFTLTNLGA 1380 PAAFKYFDTTIDRKRYTSTKEVLDATLIHQSITGLYETRIDLSQLGGDGGSGPPKKKRKV 1440 YPYDVPDYA* 4
5 Supplementary Figure 2. Reproducible genome editing in human cells with RGENs. (a) CCR5 locus. (b) C4BPB locus. The results of three independent experiments were shown. The average mutation frequencies were shown in Fig. 2. 5
6 Supplementary Figure 3. DNA sequences of mutant clones. RGEN-induced mutant clones were isolated by limiting dilution from populations of K562 cells transfected with the Cas9 plasmid and sgrna (100 μg and 40 μg used for CCR5 and C4BPB, respectively). The region of the target sequence complementary to the sgrna is shown in red. The PAM sequence recognized by Cas9 is shown in bold. The column on the right indicates the number of inserted or deleted bases. a. CCR5, 11% (= 5/46) mutated wt CAATCTATGACATCAATTATTATACATCGGAGCCCTGCCAAA clone 1 CAATCTATGACATCAATTATTATA------AGCCCTGCCAAA -6 clone 2 CAATCTATGACAT CGGAGCCCTGCCAAA -14 clone 3 CAATCTATGACAT CGGAGCCCTGCCAAA -14 clone 4 CAATCTATGACATCAATTATTATA------AGCCCTGCCAAA -6 clone 5 CAATCTATGACATCAATTATTATACAT//240bp//CGGAGCCCT +240 b. C4BPB, 21% (= 7/33) mutated wt TATGTGCAATGACCACTACATCCTCAAGGGCAGCAATCGGAG clone 1 TATGTGCAATGACCACTACATC AATCGGAG -12 clone 2 TATGTGCAATGA TCGGAG -24 clone 3 TATGTGCAATGACCACTACAT CAGCAATCGGAG -9 clone 4 TATGTGCAATGACCACTACATCC AGCAATCGGAG -8 clone 5a TATGTGCAATGACCACTACATCCT GAG -15 clone 5b TATGTGCAATGACCACTACATCCTTCAAGGGCAGCAATCGGAG +1 clone 6 TATGTGCAATGACCACTACATCCTCCTCAAGGGCAGCAATCGGAG +3 clone 7 TATGTGCAATGACCACTA----//78bp//---CAATCGGAG -15,+78 6
7 Supplementary Figure 4. CCR5 specific RGEN did not induce off-target-associated chromosomal deletions. The CCR5-specific RGEN and ZFN were expressed in human cells. PCR was used to detect the induction of the 15-kb chromosomal deletions in these cells. 7
8 Supplementary Figure 5. Stable maintenance of RGEN-induced mutant cells. Cells transected with RGENs were cultured for several days. Mutation frequencies were measured using the T7E1 assay at 3 d, 5 d, and 8 d post-transfection. 40 μg sgrna was used. 8
9 Supplementary Figure 6. RGEN-induced DSBs in cells detected by TP53BP1 staining. HeLa cells transfected with the C4BPB-specific RGEN (160 μg sgrna) were fixed on glass slides and incubated first with anti-tp53bp1 rabbit polyclonal antibodies (Bethyl Laboratories) and then with Alexa Fluor 488-conjugated secondary antibodies (Invitrogen- Molecular Probes). Cells were mounted in the presence of DAPI (Sigma) and examined under an immunofluorescence microscope (Zeiss). DAPI (blue) and TP53BP1 (green) images were merged. Etoposide (1 μm) was used as a positive control. The average number of foci per cell and the standard error of the mean are shown at the bottom of each picture. 9
10 Supplmentary Tables Supplementary Table 1. Oligonucleotides used in this study. Oligonucleotides used for the construction of the reporter plasmid Direction a Sequence (5' to 3') F AATTCATGACATCAATTATTATACATCGGAGGAG CCR5 R GATCCTCCTCCGATGTATAATAATTGATGTCATG Primers used in the T7E1 assay Gene Direction a Sequence (5' to 3') F1 CTCCATGGTGCTATAGAGCA CCR5 F2 GAGCCAAGCTCTCCATCTAGT R GCCCTGTCAAGAGTTGACAC C4BPB F1 R1 F2 R2 TATTTGGCTGGTTGAAAGGG AAAGTCATGAAATAAACACACCCA CTGCATTGATATGGTAGTACCATG GCTGTTCATTGCAATGGAATG Primers used for the amplification of off-target sites Gene Direction a Sequence (5' to 3') F1 GCTCCCACCTTAGTGCTCTG ADCY5 R1 GGTGGCAGGAACCTGTATGT F2 GTCATTGGCCAGAGATGTGGA R2 GTCCCATGACAGGCGTGTAT F GCCTGGCCAAGTTTCAGTTA KCNJ6 R1 TGGAGCCATTGGTTTGCATC R2 CCAGAACTAAGCCGTTTCTGAC F1 ATCACCGACAACCAGTTTCC CNTNAP2 F2 TGCAGTGCAGACTCTTTCCA R AAGGACACAGGGCAACTGAA F1 TGTGGAACGAGTGGTGACAG N/A Chr. 5 R1 GCTGGATTAGGAGGCAGGATTC F2 GTGCTGAGAACGCTTCATAGAG R2 GGACCAAACCACATTCTTCTCAC Primers used for the detection of chromosomal deletions Direction a Sequence (5' to 3') F CCACATCTCGTTCTCGGTTT Deletion R TCACAAGCCCACAGATATTT 10
11 In vitro transcription templates Direction a Sequence (5' to 3') b CCR5 C4BPB F R F R GAAATTAATACGACTCACTATAGGTGACATCAATTATTATACATGTTTT AGAGCTAGAAATAGCAAGTTAAAATAAGGCTAGTCCG CGGACTAGCCTTATTTTAACTTGCTATTTCTAGCTCTAAAACATGTATA ATAATTGATGTCACCTATAGTGAGTCGTATTAATTTC GAAATTAATACGACTCACTATAGGAATGACCACTACATCCTCAAGTTT TAGAGCTAGAAATAGCAAGTTAAAATAAGGCTAGTCCG CGGACTAGCCTTATTTTAACTTGCTATTTCTAGCTCTAAAACTTGAGGA TGTAGTGGTCATTCCTATAGTGAGTCGTATTAATTTC a F, forward; R, reverse b Sequences complementary to target DNA are shown in bold. The T7 promoter sequence is underlined. 11
XactEdit Cas9 Nuclease with NLS User Manual
 XactEdit Cas9 Nuclease with NLS User Manual An RNA-guided recombinant endonuclease for efficient targeted DNA cleavage Catalog Numbers CE1000-50K, CE1000-50, CE1000-250, CE1001-250, CE1001-1000 Table of
XactEdit Cas9 Nuclease with NLS User Manual An RNA-guided recombinant endonuclease for efficient targeted DNA cleavage Catalog Numbers CE1000-50K, CE1000-50, CE1000-250, CE1001-250, CE1001-1000 Table of
Transfection of CRISPR/Cas9 Nuclease NLS ribonucleoprotein (RNP) into adherent mammalian cells using Lipofectamine RNAiMAX
 Transfection of CRISPR/Cas9 Nuclease NLS ribonucleoprotein (RNP) into adherent mammalian cells using Lipofectamine RNAiMAX INTRODUCTION The CRISPR/Cas genome editing system consists of a single guide RNA
Transfection of CRISPR/Cas9 Nuclease NLS ribonucleoprotein (RNP) into adherent mammalian cells using Lipofectamine RNAiMAX INTRODUCTION The CRISPR/Cas genome editing system consists of a single guide RNA
Surrogate reporter-based enrichment of cells containing RNA-guided Cas9 nucleaseinduced
 Supplementary Data Surrogate reporter-based enrichment of cells containing RNA-guided Cas9 nucleaseinduced mutations Suresh Ramakrishna 1, Seung Woo Cho 2, Sojung Kim 2, Myungjae Song 1, Ramu Gopalappa
Supplementary Data Surrogate reporter-based enrichment of cells containing RNA-guided Cas9 nucleaseinduced mutations Suresh Ramakrishna 1, Seung Woo Cho 2, Sojung Kim 2, Myungjae Song 1, Ramu Gopalappa
CRISPR/Cas9 Gene Editing Tools
 CRISPR/Cas9 Gene Editing Tools - Guide-it Products for Successful CRISPR/Cas9 Gene Editing - Why choose Guide-it products? Optimized methods designed for speed and ease of use Complete kits that don t
CRISPR/Cas9 Gene Editing Tools - Guide-it Products for Successful CRISPR/Cas9 Gene Editing - Why choose Guide-it products? Optimized methods designed for speed and ease of use Complete kits that don t
CRISPR/Cas9 Gene Editing Tools
 CRISPR/Cas9 Gene Editing Tools - Separations Simply Spectacular INDELS Identify indels Determine if one or both copies of your gene have indels The Guide-it Genotype Confirmation Kit: Simple detection
CRISPR/Cas9 Gene Editing Tools - Separations Simply Spectacular INDELS Identify indels Determine if one or both copies of your gene have indels The Guide-it Genotype Confirmation Kit: Simple detection
Simple protocol for gene editing using GenCrisprTM Cas9 nuclease
 Simple protocol for gene editing using GenCrisprTM Cas9 nuclease Contents Protocol Step 1: Choose the target DNA sequence Step 2: Design sgrna Step 3: Preparation for sgrna 3.1 In vitro transcription of
Simple protocol for gene editing using GenCrisprTM Cas9 nuclease Contents Protocol Step 1: Choose the target DNA sequence Step 2: Design sgrna Step 3: Preparation for sgrna 3.1 In vitro transcription of
Supplementary data. sienigma. F-Enigma F-EnigmaSM. a-p53
 Supplementary data Supplemental Figure 1 A sienigma #2 sienigma sicontrol a-enigma - + ++ - - - - - - + ++ - - - - - - ++ B sienigma F-Enigma F-EnigmaSM a-flag HLK3 cells - - - + ++ + ++ - + - + + - -
Supplementary data Supplemental Figure 1 A sienigma #2 sienigma sicontrol a-enigma - + ++ - - - - - - + ++ - - - - - - ++ B sienigma F-Enigma F-EnigmaSM a-flag HLK3 cells - - - + ++ + ++ - + - + + - -
Protocols for cell lines using CRISPR/CAS
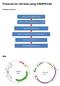 Protocols for cell lines using CRISPR/CAS Procedure overview Map Preparation of CRISPR/CAS plasmids Expression vectors for guide RNA (U6-gRNA) and Cas9 gene (CMV-p-Cas9) are ampicillin-resist ant and stable
Protocols for cell lines using CRISPR/CAS Procedure overview Map Preparation of CRISPR/CAS plasmids Expression vectors for guide RNA (U6-gRNA) and Cas9 gene (CMV-p-Cas9) are ampicillin-resist ant and stable
Confocal immunofluorescence microscopy
 Confocal immunofluorescence microscopy HL-6 and cells were cultured and cytospun onto glass slides. The cells were double immunofluorescence stained for Mt NPM1 and fibrillarin (nucleolar marker). Briefly,
Confocal immunofluorescence microscopy HL-6 and cells were cultured and cytospun onto glass slides. The cells were double immunofluorescence stained for Mt NPM1 and fibrillarin (nucleolar marker). Briefly,
Guide-it sgrna In Vitro Transcription and Screening Systems User Manual
 Clontech Laboratories, Inc. Guide-it sgrna In Vitro Transcription and Screening Systems User Manual Cat. Nos. 631438, 631439 & 631440 (042114) Clontech Laboratories, Inc. A Takara Bio Company 1290 Terra
Clontech Laboratories, Inc. Guide-it sgrna In Vitro Transcription and Screening Systems User Manual Cat. Nos. 631438, 631439 & 631440 (042114) Clontech Laboratories, Inc. A Takara Bio Company 1290 Terra
Contents... vii. List of Figures... xii. List of Tables... xiv. Abbreviatons... xv. Summary... xvii. 1. Introduction In vitro evolution...
 vii Contents Contents... vii List of Figures... xii List of Tables... xiv Abbreviatons... xv Summary... xvii 1. Introduction...1 1.1 In vitro evolution... 1 1.2 Phage Display Technology... 3 1.3 Cell surface
vii Contents Contents... vii List of Figures... xii List of Tables... xiv Abbreviatons... xv Summary... xvii 1. Introduction...1 1.1 In vitro evolution... 1 1.2 Phage Display Technology... 3 1.3 Cell surface
Supplementary Information
 1 2 3 4 5 6 7 8 9 10 11 12 Supplementary Information Supplementary Figure 1: Failure of NgAgo to cleave episomal or chromosomal targets in human cells. Supplementary Figure 2: Each component of NgAgo system
1 2 3 4 5 6 7 8 9 10 11 12 Supplementary Information Supplementary Figure 1: Failure of NgAgo to cleave episomal or chromosomal targets in human cells. Supplementary Figure 2: Each component of NgAgo system
Ligation Independent Cloning (LIC) Procedure
 Ligation Independent Cloning (LIC) Procedure Ligation Independent Cloning (LIC) LIC cloning allows insertion of DNA fragments without using restriction enzymes into specific vectors containing engineered
Ligation Independent Cloning (LIC) Procedure Ligation Independent Cloning (LIC) LIC cloning allows insertion of DNA fragments without using restriction enzymes into specific vectors containing engineered
Supplementary Information: Materials and Methods. GST and GST-p53 were purified according to standard protocol after
 Supplementary Information: Materials and Methods Recombinant protein expression and in vitro kinase assay. GST and GST-p53 were purified according to standard protocol after induction with.5mm IPTG for
Supplementary Information: Materials and Methods Recombinant protein expression and in vitro kinase assay. GST and GST-p53 were purified according to standard protocol after induction with.5mm IPTG for
Supplementary Information
 Supplementary Information Vidigal and Ventura a wt locus 5 region 3 region CCTCTGCCACTGCGAGGGCGTCCAATGGTGCTTG(...)AACAGGTGGAATATCCCTACTCTA predicted deletion clone 1 clone 2 clone 3 CCTCTGCCACTGCGAGGGCGTC-AGGTGGAATATCCCTACTCTA
Supplementary Information Vidigal and Ventura a wt locus 5 region 3 region CCTCTGCCACTGCGAGGGCGTCCAATGGTGCTTG(...)AACAGGTGGAATATCCCTACTCTA predicted deletion clone 1 clone 2 clone 3 CCTCTGCCACTGCGAGGGCGTC-AGGTGGAATATCCCTACTCTA
Supplementary Information
 Supplementary Information Self-assembled Messenger RNA Nanoparticles (mrna-nps) for Efficient Gene Expression Hyejin Kim 1, Yongkuk Park 1 and Jong Bum Lee 1, * 1 Department of Chemical Engineering, University
Supplementary Information Self-assembled Messenger RNA Nanoparticles (mrna-nps) for Efficient Gene Expression Hyejin Kim 1, Yongkuk Park 1 and Jong Bum Lee 1, * 1 Department of Chemical Engineering, University
Design. Construction. Characterization
 Design Construction Characterization DNA mrna (messenger) A C C transcription translation C A C protein His A T G C T A C G Plasmids replicon copy number incompatibility selection marker origin of replication
Design Construction Characterization DNA mrna (messenger) A C C transcription translation C A C protein His A T G C T A C G Plasmids replicon copy number incompatibility selection marker origin of replication
Genomic DNA was extracted from 3 to 5 ml of blood collected in EDTA blood collection tubes
 Supplementary information Methods DNA and RNA extraction Genomic DNA was extracted from to ml of blood collected in EDTA blood collection tubes using the Gentra Puregene Blood kit (Qiagen, California,
Supplementary information Methods DNA and RNA extraction Genomic DNA was extracted from to ml of blood collected in EDTA blood collection tubes using the Gentra Puregene Blood kit (Qiagen, California,
RNA oligonucleotides and 2 -O-methylated oligonucleotides were synthesized by. 5 AGACACAAACACCAUUGUCACACUCCACAGC; Rand-2 OMe,
 Materials and methods Oligonucleotides and DNA constructs RNA oligonucleotides and 2 -O-methylated oligonucleotides were synthesized by Dharmacon Inc. (Lafayette, CO). The sequences were: 122-2 OMe, 5
Materials and methods Oligonucleotides and DNA constructs RNA oligonucleotides and 2 -O-methylated oligonucleotides were synthesized by Dharmacon Inc. (Lafayette, CO). The sequences were: 122-2 OMe, 5
Guide-it sgrna In Vitro Transcription and Screening Systems User Manual
 Guide-it sgrna In Vitro Transcription and Screening Systems User Manual Cat. Nos. 632638, 632639, 632635, 632636, 632637 (040618) 1290 Terra Bella Avenue, Mountain View, CA 94043, USA U.S. Technical Support:
Guide-it sgrna In Vitro Transcription and Screening Systems User Manual Cat. Nos. 632638, 632639, 632635, 632636, 632637 (040618) 1290 Terra Bella Avenue, Mountain View, CA 94043, USA U.S. Technical Support:
Nature Medicine doi: /nm.2548
 Supplementary Table 1: Genotypes of offspring and embryos from matings of Pmm2 WT/F118L mice with Pmm2 WT/R137H mice total events Pmm2 WT/WT Pmm2 WT/R137H Pmm2 WT/F118L Pmm2 R137H/F118L offspring 117 (100%)
Supplementary Table 1: Genotypes of offspring and embryos from matings of Pmm2 WT/F118L mice with Pmm2 WT/R137H mice total events Pmm2 WT/WT Pmm2 WT/R137H Pmm2 WT/F118L Pmm2 R137H/F118L offspring 117 (100%)
Conversion of plasmids into Gateway compatible cloning
 Conversion of plasmids into Gateway compatible cloning Rafael Martinez 14072011 Overview: 1. Select the right Gateway cassette (A, B or C). 2. Design primers to amplify the right Gateway cassette from
Conversion of plasmids into Gateway compatible cloning Rafael Martinez 14072011 Overview: 1. Select the right Gateway cassette (A, B or C). 2. Design primers to amplify the right Gateway cassette from
Protocol for Genome Editing via the RNA-guided Cas9 Nuclease in. Zebrafish Embryos 1
 Protocol for Genome Editing via the RNA-guided Cas9 Nuclease in Zebrafish Embryos 1 1. In vitro synthesis of capped Cas9 mrna The full length of humanized Cas9 cdnas with double NLS were cloned into pxt7
Protocol for Genome Editing via the RNA-guided Cas9 Nuclease in Zebrafish Embryos 1 1. In vitro synthesis of capped Cas9 mrna The full length of humanized Cas9 cdnas with double NLS were cloned into pxt7
CHAPTER 9 DNA Technologies
 CHAPTER 9 DNA Technologies Recombinant DNA Artificially created DNA that combines sequences that do not occur together in the nature Basis of much of the modern molecular biology Molecular cloning of genes
CHAPTER 9 DNA Technologies Recombinant DNA Artificially created DNA that combines sequences that do not occur together in the nature Basis of much of the modern molecular biology Molecular cloning of genes
SUPPLEMENTARY INFORMATION
 R2A MASNSEKNPLL-SDEKPKSTEENKSS-KPESASGSSTSSAMP---GLNFNAFDFSNMASIL 56 R2B MASSSEKTPLIPSDEKNDTKEESKSTTKPESGSGAPPSPS-PTDPGLDFNAFDFSGMAGIL 60 R2A NDPSIREMAEQIAKDPAFNQLAEQLQRSIPNAGQEGGFPNFDPQQYVNTMQQVMHNPEFK
R2A MASNSEKNPLL-SDEKPKSTEENKSS-KPESASGSSTSSAMP---GLNFNAFDFSNMASIL 56 R2B MASSSEKTPLIPSDEKNDTKEESKSTTKPESGSGAPPSPS-PTDPGLDFNAFDFSGMAGIL 60 R2A NDPSIREMAEQIAKDPAFNQLAEQLQRSIPNAGQEGGFPNFDPQQYVNTMQQVMHNPEFK
Supplemental Information
 Supplemental Information ATP-dependent unwinding of U4/U6 snrnas by the Brr2 helicase requires the C-terminus of Prp8 Corina Maeder 1,3, Alan K. Kutach 1,2,3, and Christine Guthrie 1 1 Department of Biochemistry
Supplemental Information ATP-dependent unwinding of U4/U6 snrnas by the Brr2 helicase requires the C-terminus of Prp8 Corina Maeder 1,3, Alan K. Kutach 1,2,3, and Christine Guthrie 1 1 Department of Biochemistry
Torsional Constraints of DNA Substrates Impact Cas9 Cleavage
 Supporting Information Torsional Constraints of DNA Substrates Impact Cas9 Cleavage Michael H. Räz, Kumi Hidaka, Shana J. Sturla, Hiroshi Sugiyama, *,, and Masayuki Endo *, Institute for Integrated Cell-Material
Supporting Information Torsional Constraints of DNA Substrates Impact Cas9 Cleavage Michael H. Räz, Kumi Hidaka, Shana J. Sturla, Hiroshi Sugiyama, *,, and Masayuki Endo *, Institute for Integrated Cell-Material
RNA was isolated using NucleoSpin RNA II (Macherey-Nagel, Bethlehem, PA) according to the
 Supplementary Methods RT-PCR and real-time PCR analysis RNA was isolated using NucleoSpin RNA II (Macherey-Nagel, Bethlehem, PA) according to the manufacturer s protocol and quantified by measuring the
Supplementary Methods RT-PCR and real-time PCR analysis RNA was isolated using NucleoSpin RNA II (Macherey-Nagel, Bethlehem, PA) according to the manufacturer s protocol and quantified by measuring the
Supplemental Experimental Procedures
 Supplemental Experimental Procedures Generation of BLM protein segments and mutants. For immunoprecipitation experiments with topoisomerase IIα, BLM N-terminal segments were generated by PCR amplification
Supplemental Experimental Procedures Generation of BLM protein segments and mutants. For immunoprecipitation experiments with topoisomerase IIα, BLM N-terminal segments were generated by PCR amplification
TrueORF TM cdna Clones and PrecisionShuttle TM Vector System
 TrueORF TM cdna Clones and PrecisionShuttle TM Vector System Application Guide Table of Contents Package Contents and Storage Conditions... 2 Related, Optional Reagents... 2 Related Products... 2 Available
TrueORF TM cdna Clones and PrecisionShuttle TM Vector System Application Guide Table of Contents Package Contents and Storage Conditions... 2 Related, Optional Reagents... 2 Related Products... 2 Available
CHAPTER 4 Cloning, expression, purification and preparation of site-directed mutants of NDUFS3 and NDUFS7
 CHAPTER 4 Cloning, expression, purification and preparation of site-directed mutants of NDUFS3 and NDUFS7 subunits of human mitochondrial Complex-I Q module N DUFS2, 3, 7 and 8 form the core subunits of
CHAPTER 4 Cloning, expression, purification and preparation of site-directed mutants of NDUFS3 and NDUFS7 subunits of human mitochondrial Complex-I Q module N DUFS2, 3, 7 and 8 form the core subunits of
pdsipher and pdsipher -GFP shrna Vector User s Guide
 pdsipher and pdsipher -GFP shrna Vector User s Guide NOTE: PLEASE READ THE ENTIRE PROTOCOL CAREFULLY BEFORE USE Page 1. Introduction... 1 2. Vector Overview... 1 3. Vector Maps 2 4. Materials Provided...
pdsipher and pdsipher -GFP shrna Vector User s Guide NOTE: PLEASE READ THE ENTIRE PROTOCOL CAREFULLY BEFORE USE Page 1. Introduction... 1 2. Vector Overview... 1 3. Vector Maps 2 4. Materials Provided...
Programmable Sequence-Specific Transcriptional Regulation of Mammalian Genome Using Designer TAL Effectors
 Supplementary Information Programmable Sequence-Specific Transcriptional Regulation of Mammalian Genome Using Designer TAL Effectors Feng Zhang 1,2,3,5,7 *,±, Le Cong 2,3,4 *, Simona Lodato 5,6, Sriram
Supplementary Information Programmable Sequence-Specific Transcriptional Regulation of Mammalian Genome Using Designer TAL Effectors Feng Zhang 1,2,3,5,7 *,±, Le Cong 2,3,4 *, Simona Lodato 5,6, Sriram
(Supplementary Methods online)
 (Supplementary Methods online) Production and purification of either LC-antisense or control molecules Recombinant phagemids and the phagemid vector were transformed into XL-1 Blue competent bacterial
(Supplementary Methods online) Production and purification of either LC-antisense or control molecules Recombinant phagemids and the phagemid vector were transformed into XL-1 Blue competent bacterial
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature12119 SUPPLEMENTARY FIGURES AND LEGENDS pre-let-7a- 1 +14U pre-let-7a- 1 Ddx3x Dhx30 Dis3l2 Elavl1 Ggt5 Hnrnph 2 Osbpl5 Puf60 Rnpc3 Rpl7 Sf3b3 Sf3b4 Tia1 Triobp U2af1 U2af2 1 6 2 4 3
doi:10.1038/nature12119 SUPPLEMENTARY FIGURES AND LEGENDS pre-let-7a- 1 +14U pre-let-7a- 1 Ddx3x Dhx30 Dis3l2 Elavl1 Ggt5 Hnrnph 2 Osbpl5 Puf60 Rnpc3 Rpl7 Sf3b3 Sf3b4 Tia1 Triobp U2af1 U2af2 1 6 2 4 3
Schematic representation of the endogenous PALB2 locus and gene-disruption constructs
 Supplementary Figures Supplementary Figure 1. Generation of PALB2 -/- and BRCA2 -/- /PALB2 -/- DT40 cells. (A) Schematic representation of the endogenous PALB2 locus and gene-disruption constructs carrying
Supplementary Figures Supplementary Figure 1. Generation of PALB2 -/- and BRCA2 -/- /PALB2 -/- DT40 cells. (A) Schematic representation of the endogenous PALB2 locus and gene-disruption constructs carrying
The Expression of Recombinant Sheep Prion Protein (RecShPrPC) and its Detection Using Western Blot and Immuno-PCR
 The Expression of Recombinant Sheep Prion Protein (RecShPrPC) and its Detection Using Western Blot and Immuno-PCR S. Thomas, C. S. Fernando, J. Roach, U. DeSilva and C. A. Mireles DeWitt The objective
The Expression of Recombinant Sheep Prion Protein (RecShPrPC) and its Detection Using Western Blot and Immuno-PCR S. Thomas, C. S. Fernando, J. Roach, U. DeSilva and C. A. Mireles DeWitt The objective
Targeted Gene Mutation in Rice Using a CRISPR-Cas9 System Kabin Xie 1, Bastian Minkenberg 2 and Yinong Yang 2*
 Targeted Gene Mutation in Rice Using a CRISPR-Cas9 System Kabin Xie 1, Bastian Minkenberg 2 and Yinong Yang 2* 1 Department of Plant Pathology and Environmental Microbiology, Pennsylvania State University,
Targeted Gene Mutation in Rice Using a CRISPR-Cas9 System Kabin Xie 1, Bastian Minkenberg 2 and Yinong Yang 2* 1 Department of Plant Pathology and Environmental Microbiology, Pennsylvania State University,
Molecular Genetics Techniques. BIT 220 Chapter 20
 Molecular Genetics Techniques BIT 220 Chapter 20 What is Cloning? Recombinant DNA technologies 1. Producing Recombinant DNA molecule Incorporate gene of interest into plasmid (cloning vector) 2. Recombinant
Molecular Genetics Techniques BIT 220 Chapter 20 What is Cloning? Recombinant DNA technologies 1. Producing Recombinant DNA molecule Incorporate gene of interest into plasmid (cloning vector) 2. Recombinant
Supplementary Materials
 Supplementary Materials Supplementary Methods Supplementary Discussion Supplementary Figure 1 Calculated frequencies of embryo cells bearing bi-allelic alterations. Targeted indel mutations induced by
Supplementary Materials Supplementary Methods Supplementary Discussion Supplementary Figure 1 Calculated frequencies of embryo cells bearing bi-allelic alterations. Targeted indel mutations induced by
Supporting Online Material for
 www.sciencemag.org/cgi/content/full/1137999/dc1 Supporting Online Material for Disrupting the Pairing Between let-7 and Enhances Oncogenic Transformation Christine Mayr, Michael T. Hemann, David P. Bartel*
www.sciencemag.org/cgi/content/full/1137999/dc1 Supporting Online Material for Disrupting the Pairing Between let-7 and Enhances Oncogenic Transformation Christine Mayr, Michael T. Hemann, David P. Bartel*
CRISPR/Cas9 Genome Editing: Transfection Methods
 CRISPR/ Genome Editing: Transfection Methods For over 20 years Mirus Bio has developed and manufactured high performance transfection products and technologies. That expertise is now being applied to the
CRISPR/ Genome Editing: Transfection Methods For over 20 years Mirus Bio has developed and manufactured high performance transfection products and technologies. That expertise is now being applied to the
Guide-it Indel Identification Kit User Manual
 Clontech Laboratories, Inc. Guide-it Indel Identification Kit User Manual Cat. No. 631444 (120114) Clontech Laboratories, Inc. A Takara Bio Company 1290 Terra Bella Avenue, Mountain View, CA 94043, USA
Clontech Laboratories, Inc. Guide-it Indel Identification Kit User Manual Cat. No. 631444 (120114) Clontech Laboratories, Inc. A Takara Bio Company 1290 Terra Bella Avenue, Mountain View, CA 94043, USA
CRISPR Ribonucleoprotein (RNP) User Manual
 CRISPR Ribonucleoprotein (RNP) User Manual Description This user manual describes how to use GenScript s CRISPR Ribonucleoprotein (RNP) products for targeted genome editing. The RNP system is comprised
CRISPR Ribonucleoprotein (RNP) User Manual Description This user manual describes how to use GenScript s CRISPR Ribonucleoprotein (RNP) products for targeted genome editing. The RNP system is comprised
Supplementary Information: Materials and Methods. Immunoblot and immunoprecipitation. Cells were washed in phosphate buffered
 Supplementary Information: Materials and Methods Immunoblot and immunoprecipitation. Cells were washed in phosphate buffered saline (PBS) and lysed in TNN lysis buffer (50mM Tris at ph 8.0, 120mM NaCl
Supplementary Information: Materials and Methods Immunoblot and immunoprecipitation. Cells were washed in phosphate buffered saline (PBS) and lysed in TNN lysis buffer (50mM Tris at ph 8.0, 120mM NaCl
Chapter 6 - Molecular Genetic Techniques
 Chapter 6 - Molecular Genetic Techniques Two objects of molecular & genetic technologies For analysis For generation Molecular genetic technologies! For analysis DNA gel electrophoresis Southern blotting
Chapter 6 - Molecular Genetic Techniques Two objects of molecular & genetic technologies For analysis For generation Molecular genetic technologies! For analysis DNA gel electrophoresis Southern blotting
Direct Cytosolic Delivery of CRISPR/Cas9- Ribonucleoprotein for Efficient Gene Editing
 Supplementary information Direct Cytosolic Delivery of CRISPR/Cas9- Ribonucleoprotein for Efficient Gene Editing Rubul Mout, Moumita Ray, Gulen Yesilbag Tonga, Yi-Wei Lee, Tristan Tay, Kanae Sasaki, Vincent
Supplementary information Direct Cytosolic Delivery of CRISPR/Cas9- Ribonucleoprotein for Efficient Gene Editing Rubul Mout, Moumita Ray, Gulen Yesilbag Tonga, Yi-Wei Lee, Tristan Tay, Kanae Sasaki, Vincent
Supporting Online Material for
 www.sciencemag.org/cgi/content/full/1154040/dc1 Supporting Online Material for Selective Blockade of MicroRNA Processing by Lin-28 Srinivas R. Viswanathan, George Q. Daley,* Richard I. Gregory* *To whom
www.sciencemag.org/cgi/content/full/1154040/dc1 Supporting Online Material for Selective Blockade of MicroRNA Processing by Lin-28 Srinivas R. Viswanathan, George Q. Daley,* Richard I. Gregory* *To whom
Supplemental Materials and Methods
 Supplemental Materials and Methods In situ hybridization In situ hybridization analysis of HFE2 and genin mrna in rat liver tissues was performed as previously described (1). Briefly, the digoxigenin-labeled
Supplemental Materials and Methods In situ hybridization In situ hybridization analysis of HFE2 and genin mrna in rat liver tissues was performed as previously described (1). Briefly, the digoxigenin-labeled
PCR Cloning II fusion domains for purification His 6 GST chitin binding protein MBP
 PCR cloning : why secondary amplification? May 26, 2009 secondary amplification amplification of the coding sequence alone or of defined fragments ligation into expression vector insertion of fusion domains
PCR cloning : why secondary amplification? May 26, 2009 secondary amplification amplification of the coding sequence alone or of defined fragments ligation into expression vector insertion of fusion domains
CRISPR-based gene disruption in murine HSPCs. Ayumi Kitano and Daisuke Nakada
 CRISPR-based gene disruption in murine HSPCs Ayumi Kitano and Daisuke Nakada Nakada Lab Protocol INTRODUCTION We recently described fast, efficient, and cost-effective methods to directly modify the genomes
CRISPR-based gene disruption in murine HSPCs Ayumi Kitano and Daisuke Nakada Nakada Lab Protocol INTRODUCTION We recently described fast, efficient, and cost-effective methods to directly modify the genomes
A novel mechanism of post-translational modulation of HMGA functions by the histone chaperone nucleophosmin.
 A novel mechanism of post-translational modulation of HMGA functions by the histone chaperone nucleophosmin. Laura Arnoldo 1,#, Riccardo Sgarra 1,#, Eusebio Chiefari 2, Stefania Iiritano 2, Biagio Arcidiacono
A novel mechanism of post-translational modulation of HMGA functions by the histone chaperone nucleophosmin. Laura Arnoldo 1,#, Riccardo Sgarra 1,#, Eusebio Chiefari 2, Stefania Iiritano 2, Biagio Arcidiacono
Application of Molecular Biology tools for cloning of a foreign gene
 IFM/Kemi Linköpings Universitet September 2013/LGM Labmanual Project course Application of Molecular Biology tools for cloning of a foreign gene Table of contents Introduction... 3 Amplification of a gene
IFM/Kemi Linköpings Universitet September 2013/LGM Labmanual Project course Application of Molecular Biology tools for cloning of a foreign gene Table of contents Introduction... 3 Amplification of a gene
Molecular Cell Biology - Problem Drill 11: Recombinant DNA
 Molecular Cell Biology - Problem Drill 11: Recombinant DNA Question No. 1 of 10 1. Which of the following statements about the sources of DNA used for molecular cloning is correct? Question #1 (A) cdna
Molecular Cell Biology - Problem Drill 11: Recombinant DNA Question No. 1 of 10 1. Which of the following statements about the sources of DNA used for molecular cloning is correct? Question #1 (A) cdna
Bioimaging of microrna-294 expression-dependent color change. in cells by a dual fluorophore-based molecular beacon
 Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2014 Electronic Supplementary Information for Chemical Communications Bioimaging of microrna-294 expression-dependent
Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2014 Electronic Supplementary Information for Chemical Communications Bioimaging of microrna-294 expression-dependent
AmpliScribe T7-Flash Biotin-RNA Transcription Kit
 AmpliScribe T7-Flash Biotin-RNA Transcription Kit Cat. No. ASB71110 Available exclusively thru Lucigen. lucigen.com/epibio www.lucigen.com MA276E AmpliScribe T7-Flash Biotin-RNA Transcription Kit 12/2016
AmpliScribe T7-Flash Biotin-RNA Transcription Kit Cat. No. ASB71110 Available exclusively thru Lucigen. lucigen.com/epibio www.lucigen.com MA276E AmpliScribe T7-Flash Biotin-RNA Transcription Kit 12/2016
Recitation CHAPTER 9 DNA Technologies
 Recitation CHAPTER 9 DNA Technologies DNA Cloning: General Scheme A cloning vector and eukaryotic chromosomes are separately cleaved with the same restriction endonuclease. (A single chromosome is shown
Recitation CHAPTER 9 DNA Technologies DNA Cloning: General Scheme A cloning vector and eukaryotic chromosomes are separately cleaved with the same restriction endonuclease. (A single chromosome is shown
Site-directed mutagenesis of proteins
 IFM/Kemi Linköpings Universitet August 2013/LGM Labmanual Site-directed mutagenesis of proteins Figur 1: Flow-chart of the site-directed mutagenesis lab exercise 2 Site-specific mutagenesis Introduction
IFM/Kemi Linköpings Universitet August 2013/LGM Labmanual Site-directed mutagenesis of proteins Figur 1: Flow-chart of the site-directed mutagenesis lab exercise 2 Site-specific mutagenesis Introduction
Efficient Method for Isolation of High Quality Concentrated Cellular RNA with Extremely Low Levels of Genomic DNA Contamination Application
 Efficient Method for Isolation of High Quality Concentrated Cellular RNA with Extremely Low Levels of Genomic DNA Contamination Application Gene Expression Authors Ilgar Abbaszade, Claudia Robbins, John
Efficient Method for Isolation of High Quality Concentrated Cellular RNA with Extremely Low Levels of Genomic DNA Contamination Application Gene Expression Authors Ilgar Abbaszade, Claudia Robbins, John
CELLTECHGEN For Research Only. Construction of sgrna expression vector for Lenti-virus system (Example: Lenti-U6 sgrna-ef1 -Puro vector)
 Construction of sgrna expression vector for Lenti-virus system (Example: Lenti-U6 sgrna-ef1 -Puro vector) Catalog number: CTG-CAS9-11 Introduction The vector Lenti-U6 sgrna-ef1 -Puro is designed for expression
Construction of sgrna expression vector for Lenti-virus system (Example: Lenti-U6 sgrna-ef1 -Puro vector) Catalog number: CTG-CAS9-11 Introduction The vector Lenti-U6 sgrna-ef1 -Puro is designed for expression
Fast and efficient site-directed mutagenesis with Platinum SuperFi DNA Polymerase
 APPLICATION NOTE Platinum Superi Polymerase ast and efficient site-directed mutagenesis with Platinum Superi Polymerase Introduction Site-directed mutagenesis is one of the most essential techniques to
APPLICATION NOTE Platinum Superi Polymerase ast and efficient site-directed mutagenesis with Platinum Superi Polymerase Introduction Site-directed mutagenesis is one of the most essential techniques to
FROM EXPERIMENTS IN BACTERIAL GENETICS AND GENE TECHNIQUE
 Uppsala 2001-04-01 REPORT FROM EXPERIMENTS IN BACTERIAL GENETICS AND GENE TECHNIQUE Laboratory assistants: Maria Jönsson Amera Gibreel Students: Contents ASSIGNMENT:... 3 INTRODUCTION:... 3 MATERIAL AND
Uppsala 2001-04-01 REPORT FROM EXPERIMENTS IN BACTERIAL GENETICS AND GENE TECHNIQUE Laboratory assistants: Maria Jönsson Amera Gibreel Students: Contents ASSIGNMENT:... 3 INTRODUCTION:... 3 MATERIAL AND
SUPPLEMENTARY INFORMATION
 The Supplementary Information (SI) Methods Cell culture and transfections H1299, U2OS, 293, HeLa cells were maintained in DMEM medium supplemented with 10% fetal bovine serum. H1299 and 293 cells were
The Supplementary Information (SI) Methods Cell culture and transfections H1299, U2OS, 293, HeLa cells were maintained in DMEM medium supplemented with 10% fetal bovine serum. H1299 and 293 cells were
Nature Methods: doi: /nmeth Supplementary Figure 1. Comparison of REC2 and BCL-xL domains and modeling domain replacement.
 Supplementary Figure 1 Comparison of REC2 and BCL-xL domains and modeling domain replacement. (a) The REC2 domain of Cas9 and (b) BCL-xL display similar globular structures and volumes. (c, d) Modeling
Supplementary Figure 1 Comparison of REC2 and BCL-xL domains and modeling domain replacement. (a) The REC2 domain of Cas9 and (b) BCL-xL display similar globular structures and volumes. (c, d) Modeling
Supplementary Methods
 Supplementary Methods Reverse transcribed Quantitative PCR. Total RNA was isolated from bone marrow derived macrophages using RNeasy Mini Kit (Qiagen), DNase-treated (Promega RQ1), and reverse transcribed
Supplementary Methods Reverse transcribed Quantitative PCR. Total RNA was isolated from bone marrow derived macrophages using RNeasy Mini Kit (Qiagen), DNase-treated (Promega RQ1), and reverse transcribed
Supplementary
 Supplementary information Supplementary Material and Methods Plasmid construction The transposable element vectors for inducible expression of RFP-FUS wt and EGFP-FUS R521C and EGFP-FUS P525L were derived
Supplementary information Supplementary Material and Methods Plasmid construction The transposable element vectors for inducible expression of RFP-FUS wt and EGFP-FUS R521C and EGFP-FUS P525L were derived
Oligonucleotides containing either a single cyclo-da lesion or a single CPD in an
 Materials and Methods Preparation of Constructs Oligonucleotides containing either a single cyclo-da lesion or a single CPD in an unique MfeI site were prepared as described previously (Brooks et al.,
Materials and Methods Preparation of Constructs Oligonucleotides containing either a single cyclo-da lesion or a single CPD in an unique MfeI site were prepared as described previously (Brooks et al.,
SUPPORTING INFORMATION. A cleavage-responsive stem-loop hairpin for assaying guide RNA activity
 SUPPORTING INFORMATION A cleavage-responsive stem-loop hairpin for assaying guide RNA activity Tara R. deboer 1, Noreen Wauford 1, Jing-Yi Chung, Miguel Salvador Torres Perez, and Niren Murthy* University
SUPPORTING INFORMATION A cleavage-responsive stem-loop hairpin for assaying guide RNA activity Tara R. deboer 1, Noreen Wauford 1, Jing-Yi Chung, Miguel Salvador Torres Perez, and Niren Murthy* University
SUPPLEMENTARY INFORMATION
 (Supplementary Methods and Materials) GST pull-down assay GST-fusion proteins Fe65 365-533, and Fe65 538-700 were expressed in BL21 bacterial cells and purified with glutathione-agarose beads (Sigma).
(Supplementary Methods and Materials) GST pull-down assay GST-fusion proteins Fe65 365-533, and Fe65 538-700 were expressed in BL21 bacterial cells and purified with glutathione-agarose beads (Sigma).
Selected Techniques Part I
 1 Selected Techniques Part I Gel Electrophoresis Can be both qualitative and quantitative Qualitative About what size is the fragment? How many fragments are present? Is there in insert or not? Quantitative
1 Selected Techniques Part I Gel Electrophoresis Can be both qualitative and quantitative Qualitative About what size is the fragment? How many fragments are present? Is there in insert or not? Quantitative
SUPPLEMENTAL MATERIALS. Chromatin immunoprecipitation assays. Single-cell suspensions of pituitary
 SUPPLEMENTAL MATERIALS METHODS Chromatin immunoprecipitation assays. Single-cell suspensions of pituitary and liver cells were prepared from the indicated transgenic mice, as described (Ho et al, 2002).
SUPPLEMENTAL MATERIALS METHODS Chromatin immunoprecipitation assays. Single-cell suspensions of pituitary and liver cells were prepared from the indicated transgenic mice, as described (Ho et al, 2002).
Supporting Information
 Supporting Information Wiley-VCH 2006 69451 Weinheim, Germany RNA ligands that distinguish metabolite-induced conformations in the TPP riboswitch Günter Mayer, Marie-Sophie L. Raddatz, Julia D. Grunwald,
Supporting Information Wiley-VCH 2006 69451 Weinheim, Germany RNA ligands that distinguish metabolite-induced conformations in the TPP riboswitch Günter Mayer, Marie-Sophie L. Raddatz, Julia D. Grunwald,
Alt-R CRISPR-Cpf1 System:
 user guide Alt-R CRISPR-Cpf1 System: Delivery of ribonucleoprotein complexes in HEK-293 cells using the Amaxa Nucleofector System See what more we can do for you at www.idtdna.com. For Research Use Only
user guide Alt-R CRISPR-Cpf1 System: Delivery of ribonucleoprotein complexes in HEK-293 cells using the Amaxa Nucleofector System See what more we can do for you at www.idtdna.com. For Research Use Only
GenBuilder TM Plus Cloning Kit User Manual
 GenBuilder TM Plus Cloning Kit User Manual Cat. No. L00744 Version 11242017 Ⅰ. Introduction... 2 I.1 Product Information... 2 I.2 Kit Contents and Storage... 2 I.3 GenBuilder Cloning Kit Workflow... 2
GenBuilder TM Plus Cloning Kit User Manual Cat. No. L00744 Version 11242017 Ⅰ. Introduction... 2 I.1 Product Information... 2 I.2 Kit Contents and Storage... 2 I.3 GenBuilder Cloning Kit Workflow... 2
Supplementary information
 Supplementary information The E3 ligase RNF8 regulates KU80 removal and NHEJ repair Lin Feng 1, Junjie Chen 1 1 Department of Experimental Radiation Oncology, The University of Texas M. D. Anderson Cancer
Supplementary information The E3 ligase RNF8 regulates KU80 removal and NHEJ repair Lin Feng 1, Junjie Chen 1 1 Department of Experimental Radiation Oncology, The University of Texas M. D. Anderson Cancer
Fisher (Fairlawn, NJ) and Sigma-Aldrich (St. Louis, MO) and were used without further. (Promega) and DpnI (New England Biolabs, Beverly, MA).
 175 Appendix III Chapter 4 Methods General. Unless otherwise noted, reagents were purchased from the commercial suppliers Fisher (Fairlawn, NJ) and Sigma-Aldrich (St. Louis, MO) and were used without further
175 Appendix III Chapter 4 Methods General. Unless otherwise noted, reagents were purchased from the commercial suppliers Fisher (Fairlawn, NJ) and Sigma-Aldrich (St. Louis, MO) and were used without further
GenBuilder TM Plus Cloning Kit User Manual
 GenBuilder TM Plus Cloning Kit User Manual Cat.no L00744 Version 11242017 Ⅰ. Introduction... 2 I.1 Product Information... 2 I.2 Kit Contents and Storage... 2 I.3 GenBuilder Cloning Kit Workflow... 2 Ⅱ.
GenBuilder TM Plus Cloning Kit User Manual Cat.no L00744 Version 11242017 Ⅰ. Introduction... 2 I.1 Product Information... 2 I.2 Kit Contents and Storage... 2 I.3 GenBuilder Cloning Kit Workflow... 2 Ⅱ.
Amplified segment of DNA can be purified from bacteria in sufficient quantity and quality for :
 Transformation Insertion of DNA of interest Amplification Amplified segment of DNA can be purified from bacteria in sufficient quantity and quality for : DNA Sequence. Understand relatedness of genes and
Transformation Insertion of DNA of interest Amplification Amplified segment of DNA can be purified from bacteria in sufficient quantity and quality for : DNA Sequence. Understand relatedness of genes and
The retroviral vectors encoding WT human SIRT1 or a mutant of SIRT in which a
 Supporting online material Constructs The retroviral vectors encoding WT human SIRT1 or a mutant of SIRT in which a critical histidine in the deacetylase domain of SIRT1 has been replaced by a tyrosine
Supporting online material Constructs The retroviral vectors encoding WT human SIRT1 or a mutant of SIRT in which a critical histidine in the deacetylase domain of SIRT1 has been replaced by a tyrosine
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature25164 Supplementary Methods Supplementary Table 1. Two-way ANOVA analysis of ABR and DPOAE thresholds. Supplementary Table 2. The number of injected and uninjected ears tested for ABRs
doi:10.1038/nature25164 Supplementary Methods Supplementary Table 1. Two-way ANOVA analysis of ABR and DPOAE thresholds. Supplementary Table 2. The number of injected and uninjected ears tested for ABRs
8Br-cAMP was purchased from Sigma (St. Louis, MO). Silencer Negative Control sirna #1 and
 1 Supplemental information 2 3 Materials and Methods 4 Reagents and animals 5 8Br-cAMP was purchased from Sigma (St. Louis, MO). Silencer Negative Control sirna #1 and 6 Silencer Select Pre-designed sirna
1 Supplemental information 2 3 Materials and Methods 4 Reagents and animals 5 8Br-cAMP was purchased from Sigma (St. Louis, MO). Silencer Negative Control sirna #1 and 6 Silencer Select Pre-designed sirna
Supplementary Figure Legends
 Supplementary Figure Legends Figure S1 gene targeting strategy for disruption of chicken gene, related to Figure 1 (f)-(i). (a) The locus and the targeting constructs showing HpaI restriction sites. The
Supplementary Figure Legends Figure S1 gene targeting strategy for disruption of chicken gene, related to Figure 1 (f)-(i). (a) The locus and the targeting constructs showing HpaI restriction sites. The
Deep sequencing reveals global patterns of mrna recruitment
 Supplementary information for: Deep sequencing reveals global patterns of mrna recruitment during translation initiation Rong Gao 1#*, Kai Yu 1#, Ju-Kui Nie 1,Teng-Fei Lian 1, Jian-Shi Jin 1, Anders Liljas
Supplementary information for: Deep sequencing reveals global patterns of mrna recruitment during translation initiation Rong Gao 1#*, Kai Yu 1#, Ju-Kui Nie 1,Teng-Fei Lian 1, Jian-Shi Jin 1, Anders Liljas
EnGen Mutation Detection Kit
 DNA MODIFYING ENZYMES EnGen Mutation Detection Kit Instruction Manual NEB #E3321S 25 reactions Version 2.0 6/18 be INSPIRED drive DISCOVERY stay GENUINE This product is intended for research purposes only.
DNA MODIFYING ENZYMES EnGen Mutation Detection Kit Instruction Manual NEB #E3321S 25 reactions Version 2.0 6/18 be INSPIRED drive DISCOVERY stay GENUINE This product is intended for research purposes only.
Supporting Information
 Supporting Information Deng et al. 10.1073/pnas.1515692112 SI Materials and Methods FPLC. All fusion proteins were expressed and purified through a three-step FPLC purification protocol, as described (20),
Supporting Information Deng et al. 10.1073/pnas.1515692112 SI Materials and Methods FPLC. All fusion proteins were expressed and purified through a three-step FPLC purification protocol, as described (20),
For designing necessary primers and oligos for grna, online CRISPR designing tools can be used (e.g.
 DESIGN OF grnas For designing necessary primers and oligos for grna, online CRISPR designing tools can be used (e.g. http://crispr.mit.edu/, https://benchling.com). Assembly of grna transcriptional cassettes
DESIGN OF grnas For designing necessary primers and oligos for grna, online CRISPR designing tools can be used (e.g. http://crispr.mit.edu/, https://benchling.com). Assembly of grna transcriptional cassettes
3 Designing Primers for Site-Directed Mutagenesis
 3 Designing Primers for Site-Directed Mutagenesis 3.1 Learning Objectives During the next two labs you will learn the basics of site-directed mutagenesis: you will design primers for the mutants you designed
3 Designing Primers for Site-Directed Mutagenesis 3.1 Learning Objectives During the next two labs you will learn the basics of site-directed mutagenesis: you will design primers for the mutants you designed
Lecture Four. Molecular Approaches I: Nucleic Acids
 Lecture Four. Molecular Approaches I: Nucleic Acids I. Recombinant DNA and Gene Cloning Recombinant DNA is DNA that has been created artificially. DNA from two or more sources is incorporated into a single
Lecture Four. Molecular Approaches I: Nucleic Acids I. Recombinant DNA and Gene Cloning Recombinant DNA is DNA that has been created artificially. DNA from two or more sources is incorporated into a single
N173A. residue number
 a 0.3 Δω 1 ( 1 H Ν ) [ppm] 0-0.3 N173A -0.6 Y177A b Δω 3 ( 13 C α ) [ppm] 2 0-2 c Δω 3 ( 13 C β ) [ppm] 2 0-2 -4 Supplementary Figure 1 140 150 160 170 180 190 200 residue number Comparison of chemical
a 0.3 Δω 1 ( 1 H Ν ) [ppm] 0-0.3 N173A -0.6 Y177A b Δω 3 ( 13 C α ) [ppm] 2 0-2 c Δω 3 ( 13 C β ) [ppm] 2 0-2 -4 Supplementary Figure 1 140 150 160 170 180 190 200 residue number Comparison of chemical
PDIP46 (DNA polymerase δ interacting protein 46) is an activating factor for human DNA polymerase δ
 PDIP46 (DNA polymerase δ interacting protein 46) is an activating factor for human DNA polymerase δ Supplementary Material Figure S1. PDIP46 is associated with Pol isolated by immunoaffinity chromatography.
PDIP46 (DNA polymerase δ interacting protein 46) is an activating factor for human DNA polymerase δ Supplementary Material Figure S1. PDIP46 is associated with Pol isolated by immunoaffinity chromatography.
Supplementary Figure S1 Purification of deubiquitinases HEK293 cells were transfected with the indicated DUB-expressing plasmids.
 Supplementary Figure S1 Purification of deubiquitinases HEK293 cells were transfected with the indicated DUB-expressing plasmids. The cells were harvested 72 h after transfection. FLAG-tagged deubiquitinases
Supplementary Figure S1 Purification of deubiquitinases HEK293 cells were transfected with the indicated DUB-expressing plasmids. The cells were harvested 72 h after transfection. FLAG-tagged deubiquitinases
Ultrasensitive detection of antibodies using a new Tus-Ter-lock immunopcr system
 ELECTRONIC SUPPLEMENTARY INFORMATION Ultrasensitive detection of antibodies using a new Tus-Ter-lock immunopcr system Isabelle Morin, Nicholas E. Dixon and Patrick M. Schaeffer Materials and methods Cloning,
ELECTRONIC SUPPLEMENTARY INFORMATION Ultrasensitive detection of antibodies using a new Tus-Ter-lock immunopcr system Isabelle Morin, Nicholas E. Dixon and Patrick M. Schaeffer Materials and methods Cloning,
Combining Techniques to Answer Molecular Questions
 Combining Techniques to Answer Molecular Questions UNIT FM02 How to cite this article: Curr. Protoc. Essential Lab. Tech. 9:FM02.1-FM02.5. doi: 10.1002/9780470089941.etfm02s9 INTRODUCTION This manual is
Combining Techniques to Answer Molecular Questions UNIT FM02 How to cite this article: Curr. Protoc. Essential Lab. Tech. 9:FM02.1-FM02.5. doi: 10.1002/9780470089941.etfm02s9 INTRODUCTION This manual is
Four different active promoter genes were chosen, ATXN7L2, PSRC1, CELSR2 and
 SUPPLEMENTARY MATERIALS AND METHODS Chromatin Immunoprecipitation for qpcr analysis Four different active promoter genes were chosen, ATXN7L2, PSRC1, CELSR2 and IL24, all located on chromosome 1. Primer
SUPPLEMENTARY MATERIALS AND METHODS Chromatin Immunoprecipitation for qpcr analysis Four different active promoter genes were chosen, ATXN7L2, PSRC1, CELSR2 and IL24, all located on chromosome 1. Primer
Supporting Information
 Supporting Information Development of a 2,4-Dinitrotoluene-Responsive Synthetic Riboswitch in E. coli cells Molly E. Davidson, Svetlana V. Harbaugh, Yaroslav G. Chushak, Morley O. Stone, Nancy Kelley-
Supporting Information Development of a 2,4-Dinitrotoluene-Responsive Synthetic Riboswitch in E. coli cells Molly E. Davidson, Svetlana V. Harbaugh, Yaroslav G. Chushak, Morley O. Stone, Nancy Kelley-
Data Sheet Quick PCR Cloning Kit
 Data Sheet Quick PCR Cloning Kit 6044 Cornerstone Ct. West, Ste. E DESCRIPTION: The Quick PCR Cloning Kit is a simple and highly efficient method to insert any gene or DNA fragment into a vector, without
Data Sheet Quick PCR Cloning Kit 6044 Cornerstone Ct. West, Ste. E DESCRIPTION: The Quick PCR Cloning Kit is a simple and highly efficient method to insert any gene or DNA fragment into a vector, without
CELLTECHGEN For Research Only. Construction of all-in-one vector for Lenti-virus system (Example: Lenti-EF1 -Cas9-EGFP-U6 sgrna vector)
 Construction of all-in-one vector for Lenti-virus system (Example: Lenti-EF1 -Cas9-EGFP-U6 sgrna vector) Catalog number: CTG-CAS9-18 Introduction The vector Lenti-EF1 -Cas9-EGFP-U6 sgrna is designed for
Construction of all-in-one vector for Lenti-virus system (Example: Lenti-EF1 -Cas9-EGFP-U6 sgrna vector) Catalog number: CTG-CAS9-18 Introduction The vector Lenti-EF1 -Cas9-EGFP-U6 sgrna is designed for
Supplemental Data. LMO4 Controls the Balance between Excitatory. and Inhibitory Spinal V2 Interneurons
 Neuron, Volume 61 Supplemental Data LMO4 Controls the Balance between Excitatory and Inhibitory Spinal V2 Interneurons Kaumudi Joshi, Seunghee Lee, Bora Lee, Jae W. Lee, and Soo-Kyung Lee Supplemental
Neuron, Volume 61 Supplemental Data LMO4 Controls the Balance between Excitatory and Inhibitory Spinal V2 Interneurons Kaumudi Joshi, Seunghee Lee, Bora Lee, Jae W. Lee, and Soo-Kyung Lee Supplemental
NZYGene Synthesis kit
 Kit components Component Concentration Amount NZYGene Synthesis kit Catalogue number: MB33901, 10 reactions GS DNA Polymerase 1U/ μl 30 μl Reaction Buffer for GS DNA Polymerase 10 150 μl dntp mix 2 mm
Kit components Component Concentration Amount NZYGene Synthesis kit Catalogue number: MB33901, 10 reactions GS DNA Polymerase 1U/ μl 30 μl Reaction Buffer for GS DNA Polymerase 10 150 μl dntp mix 2 mm
Document S1. Supplemental Experimental Procedures and Three Figures (see next page)
 Supplemental Data Document S1. Supplemental Experimental Procedures and Three Figures (see next page) Table S1. List of Candidate Genes Identified from the Screen. Candidate genes, corresponding dsrnas
Supplemental Data Document S1. Supplemental Experimental Procedures and Three Figures (see next page) Table S1. List of Candidate Genes Identified from the Screen. Candidate genes, corresponding dsrnas
