Universal Split Spinach Aptamer (USSA) for Nucleic Acid Analysis and DNA Computation
|
|
|
- Martina Phelps
- 6 years ago
- Views:
Transcription
1 Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2017 Supporting materials Universal Split Spinach Aptamer (USSA) for Nucleic Acid Analysis and DNA Computation Nanami Kikuchi 1 and Dmitry M. Kolpashchikov 1,2 1 Chemistry Department, University of Central Florida, Orlando, 32816, Florida, USA 2 National Center for Forensic Science and Burnett School of Biomedical Sciences, University of Central Florida, Orlando, 32816, Florida, USA Table of content 1) Materials and instruments 2) Table S1: oligonucleotides used in this study 3) Detailed Experimental Procedure 4) Figure S1. Design of USSA probe 5) Figure S2-S4. Suboptimal designs of USSA analyzing Mtb. F11/KZN analyte 6) Figure S5. Analysis of long analyte using USSA probe 7) References 1. Materials and instruments. All oligonucleotides were purchased from Integrated DNA Technologies, Inc. (Coralville, IA). DNAse/RNAse free water was purchased from Fisher Scientific and used for all assays including buffers, and for dissolution of oligonucleotides. Concentrations of oligonucleotide were determined based on UV light absorption at 260 nm. DFHBI was purchased from Lucerna, Inc. (New York, NY), KCl and MgCl 2 were purchased from Fisher Scientific. Trizma Hydrochloride (Tris-HCl), ph 7.40 was purchased from Sigma Aldrich. Two 2 Spinach buffer were prepared: 2 Spinach Buffer 1 (20 mm Tris-Cl, ph 7.4, 200 mm KCl, 10 mm MgCl 2 ); 2 Spinach Buffer 2 (20 mm Tris-HCl, ph 7.4, 200 mm KCl, 100 mm MgCl 2 ). All fluorescent spectra were taken using Fluorescence Spectrometer LS55 (PerkinElmer). Otherwise noted, excitation wavelength was set to 450 nm and emission was taken at 500 nm.
2 2. Table S1: oligonucleotides used in this study Name Sequence Purification SSA_m 5 -gtg ttg tgt /TEG/ UGG UGA AGG ACG GGU CCA GU SD SSA_f 5 -ACU GUU GAG UAG AGU GUG AGC UCC Gca gtg gcc cat acc catg c SD Adp_m aca aga tcg /TEG/ cca ctg aca caa cac SD Adp_f atg ggt atg ggg atct acg ggt t SD A m _DNA aac ccg tag atc cga tct tgt g SD A mm _DNA aac ccg tag atc cga act tgt g SD A m _RNA AAC CCG UAG AUC CGA UCU UGU G SD A mm _RNA AAC CCG UAG AUC CGA ACU UGU G SD Adp_m_NOR1 at ggg tat ggg c gtc ggc gct tgt SD Adp_f_NOR1 agcgccgactgttggc atacgtaagtgatcc gcca aca cca ctg aca caa cac SD I1 gcg gat cac tta cgt atg cca ac SD I2 ccc aca agc gcc gac SD Mtb. F11 ggg ttgaccc aca agc gcc gac tgt t ggc g ctg SD Mtb. KZN ggg ttgaccc aca agc gcc gac tgt c ggc g ctg SD f3 gcg cca aca aca caa cac SD f4 gcg cca aca a ctg aca caa cac SD f5 gcg cca aca ca ctg aca caa cac SD f6 gcg cca aca cca ctg aca caa cac SD f6peg gcg cca aca/teg/ cca ctg aca caa cac SD m3 tat ggg cca ctg gtc ggc gct tgt SD m4 t ggg tat ggg cc gtc ggc gct tgt SD m5 at ggg tat ggg c gtc ggc gct tgt SD m6 tat ggg tat ggg gtc ggc gct tgt SD m6peg tat ggg tat ggg /TEG/ gtc ggc gct tgt SD Amel Y Amel Y_mut (A G) c ccagtttaa gctctgatgg tt ggcctcaa gcctgtgttg ctccagcacc ctcctgcctg accattcgga t tgactcttt cctcct aaat a tggc tgtaa gtttattcat tcatgaacca ctgctcagga ag gttccatg aaagggcaaa aa c ccagtttaa gctctgatgg tt ggcctcaa gcctgtgttg ctccagcacc ctcctgcctg accattcgga t tgactcttt cctcct aaat g tggc tgtaa gtttattcat tcatgaacca SD SD
3 ctgctcagga ag gttccatg aaagggcaaa aa Adp_f_amelY at ggg tat ggg agg agg aaa gag tca atc cga atg gtc agg cag gag SD Adp_m_amelY gcca t attt/teg/cca ctg aca caa cac SD TEG-triethylene glycol linkers; SD, standard desalting; DNA sequences are in low case; SNS sites are bold underlined; analyte binding arms are underlined. 3. Detailed Experimental Procedure General Fluorescent assay for midna and mirna. For DNA analyte, the reaction mixtures contained DFHBI (1 μm) SSA_m (3.6 μm) and SSA_f (2 μm) strands, Adp_m (0.8 μm) and Adp_f (4.0 μm) and the indicated concentration of analyte in 60 μl of 1 Spinach 1 or Spinach 2 buffers. Control samples contained only DFHBI (1 μm) or DFHBI (1 μm) and SSA_m (3.6 μm) and SSA_f (2 μm) and Adp_m (0.8 μm) and Adp_f (4.0 μm) strands. For RNA analyte, DFHBI (0.5 μm) SSA_m (3.6 μm) and SSA_f (2.0 μm) and the indicated concentration of analyte in 60 μl of Spinach 2 buffers. Control samples contained only DFHBI (0.5 μm) or DFHBI (0.5 μm) SSA_m (3.6 μm) and SSA_f (2.0 μm) and Adp_m (0.8 μm) and Adp_f (4.0 μm) strands. All samples were incubated at room (22.5 o C) temperature. Fluorescent spectra were recorded at nm, with excitation at 450 nm after indicated incubation times. Data of three independent experiments were processed using Microsoft Excel. General Fluorescent assay for Mtb.F11 and Mtb.KZN. DFHBI (2 μm) SSA_m (3.6 μm) and SSA_f (2 μm) strands, Adp_m (0.8 μm) and Adp_f (4.0 μm) and the indicated concentration of analyte in 60 μl of 1 Spinach 1 or Spinach 2 buffers. Control samples contained only DFHBI (2 μm) or DFHBI (2 μm) and SSA_m (3.6 μm) and SSA_f (2 μm) strands. All samples were incubated at room (22.5 o C) temperature. Fluorescent spectra were recorded at nm, with excitation at 450 nm after indicated incubation times. Data of three independent experiments were processed using Microsoft Excel. Time dependence of USSA fluorescent response. DFHBI, SSA_m and SSA_f strands, Adp_m and Adp_f and 400 nm for DNA analyte, 100 nm for RNA analyte in 60 μl of 1 Spinach 2 buffers. Fluorescence measurements were taken after 0, 1, 2, 5, 10, 25 and 40 min. Control samples were ran parallel with the samples. Data of three independent experiments were processed using Microsoft Excel. Determining Limit of Detection. DFHBI, SSA_m and SSA_f strands, Adp_m and Adp_f matched DNA (1.4, 5.5, 14, 28, 138, 275 nm or 5.5 μm) or RNA (0.5, 2, 5, 10, 50, 100, 250 nm, 1 μm) analytes in 60 μl of 1 Spinach 2 buffers. matched or mismatched analyte was added for a total volume of 60 μl reaction mixture. Fluorescent spectra were measured after 30 min. Control samples were ran parallel with the samples. Data of three independent experiments were processed using Microsoft Excel. Selectivity Assay. For midna and mirna, DFHBI, SSA_m and SSA_f strands, Adp_m and Adp_f, and 100 nm matched or mismatched analytes in 60 μl of 1 Spinach 2 buffers. For
4 Mtb.F11 and Mtb.KZN, DFHBI, SSA_m and SSA_f strands, Adp_m and Adp_f, and the indicated concentration of analyte in 60 μl of 1 Spinach 2 buffers.. Control samples were ran parallel with the samples. Fluorescent spectra were measured after 30 min. Data of three independent experiments were processed using Microsoft Excel. NOT and NOR logic gates. DFHBI (0.5 µm), SSA_m and SSA_f strands (2.0 µm), Adp_f_NOR1 (1.0 µm) and Adp_m_NOR1 (0.3 µm), were added to 30 μl of 2 Spinach 2 buffer. Total volume was adjusted to 50 μl by H 2 O and 5 μl each of Input I1 and I2 (1.0 µm) were added for a total volume of 60 μl reaction mixture. Control samples were ran parallel with the samples. Fluorescent spectra were measured after 30 min. Data of three independent experiments were processed using Microsoft Excel. 4. Figure S1. Design of USSA probe Figure S1. Design of USSA with mirna analyte. Dotted black line is the triethylene glycol linkers. Ribunucleotides are shown in upper case; deoxyribonucleotide are in low case. SNP location on mirna underlined. We initially designed the adapter strands as indicated in Figure S2A but showed no selectivity towards fully matched analyte. Therefore strand Adp_m in Figure S1 and the main paper crossed over and was made complementary to SSA_f by 4 nucleotides which reduced the fluorescence signal of single mismatched analyte (Figure S2B). We also inserted a triethyleneglycol (TEG) linker connecting the SSA binding arm and the analyte binding arm which allowed great increase in the selectivity of the USSA (Figure S3C). Magnesium concentration in Spinach buffer was increased from 5 mm to 50 mm which allowed greater stability for hybridization which inducing higher signal for fully matched analyte (Figure S4). For any TEG containing strand combination, lower concentration of adapter-f was used which allowed lowering single mismatch analyte signal (Figure S4). We concluded that USSA in which strand_f is complementary to SSA_f by 6 nucleotides produced the greatest analyte dependent fluorescent increase (Figure S4).
5 5. Figure S2: Suboptimal designs of USSA analyzing Mtb. F11/KZN analyte Figure S2. Fluorescence complexes of USSA with the DNA analyte and responses of the two different combinations of stands m and f. Data for each combination is shown on the right side of each panel. Samples contained 3.6 µm SSA_f; 2.0 µm SSA_m; 4.0 µm Adp_f; 4.0µM Adp_m; 6.0 µm analyte; 20 µm DFHBI; in Spinach Buffer 2). Emission ( ex = 450 nm) was registered 500 nm after 30 min at 22.5 C. A, B) The bar graph shows the fluorescence intensity of samples containing DFHBI dye, no analyte (SSA+Adp), Am_DNA (SSA+Adp+matched analyte), and Amm_DNA (SSA+Adp+mismatched analyte).
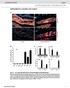
6 Figure S3: Suboptimal designs of USSA analyzing Mtb. F11/KZN analyte Figure S3. Fluorescence complexes of USSA with the DNA analyte and responses of the three different combinations of stands m and f. Data for each combination is shown on the right side of each panel. Samples contained 3.6 µm SSA_f; 2.0 µm SSA_m; 4.0 µm Adp_f; 4.0µM Adp_m; 6.0 µm analyte; 10 µm DFHBI; in 2 Spinach Buffer 1. Emission spectrum ( ex = 450 nm) were collected after 30 min at 22.5 C. A, B, C) The bar graph shows the fluorescence intensity of samples containing DFHBI dye, no analyte (SSA+Adp), Am_DNA (SSA+Adp+matched analyte), and Amm_DNA (SSA+Adp+mismatched analyte). C) Dashed lines represent triethylene glycol (TEG) linkers. Figure S4: Suboptimal designs of USSA analyzing Mtb. F11/KZN analyte Figure S4. Fluorescence complexes of USSA with the DNA analyte and responses of the two different combinations of stands m and f. Data for each combination is shown on the right side of each panel. Samples contained 3.6 µm SSA_f; 2.0 µm SSA_m; 0.4 µm Adp_f; 4.0µM Adp_m; 6.0 µm analyte; 5 µm DFHBI; in 2 Spinach Buffer 2. Emission spectrum ( ex = 450 nm) were collected after 30 min at 22.5 C. A, B, C) The bar graph shows the fluorescence intensity of samples containing DFHBI dye, no analyte (SSA+Adp), Am_DNA (SSA+Adp+matched analyte), and Amm_DNA (SSA+Adp+mismatched analyte). Deshed lines represents TEG linkers.
7 6) Figure S5. Analysis of long analyte using USSA probe Figure S5. Detection of a long DNA analyte by USSA_AMELY probe. A) Secondary structure of 152 nt AMELY analyte predicted by mfold. 1 The fragment bound by Adp_f_amelY and Adp_m_amelY are indicated by cyan and orange lines respectively. B) Fluorescent response of the USSA_AMELY probe in the presence of fully matched AMEL Y or single base mismatched Amel Y_mut (A G) analyte. No analyte is the probe response in the absence of analytes (negative control). The assay conditions: DFHBI (1 µm), SSA_m (2 µm), SSA_f (2 µm), Adp_f_amelY (2 µm), Adp_m_amelY (0.5 µm) in Spinach Buffer 2. C) The predicated secondary structure of USSA_AMELY probe with fully matched AMELY analyte. Single nucleotide substitution site is red underlined. For full sequence of Amel Y analyte see Table S1. D) Representative spectrums of USSA_AMELY probe in the presence of matched AmelY (green) or mismatched (Amel Y_mut (A G)) analytes. Blue line is USSA_AMELY probe is the absence of the analyte (negative control) In order to prove that SSA is suitable for the detection of long structure-forming analytes, we designed a probe for detection of AMELY (Figure S5A). DNA fragments of AMEL X and AMEL Y genes, which code for the amelogenin gene involved in the development of the enamel. 2 Different amelogenin sequences are located in X and Y chromosomes, which makes this gene to be a good marker for sex determination, in forensic and anthropological applications. 3 The analyte binding arm of the adaptor strand Adp_m_amelY was designed to unwind the secondary structure
8 of the analyte (Figure 3SA). The analyte binding arm of Adp_m_amelY was short to enable highly specific hybridization of signal nucleotide substitution. The signal-to-background ratio was ~20 under optimized concentrations of the USSA components (Figure S5B). Importantly, a single base substituted Adp_m_amelY produced fluorescence only slightly above the background (Figure S5B, last bar). 7) References 1) M. Zuker, Mfold web server for nucleic acid folding and hybridization prediction. Nucleic Acids Res., 2003, 31, ) B. A. Álvarez-Sandoval, L. R. Manzanilla, R. Montiel, Sex determination in highly fragmented human DNA by high-resolution melting (HRM) analysis. PLoS One, 2014, 9, e ) H. Nogami, H. Tsutsumi, T. Komuro, R. Mukoyama Rapid and simple sex determination method from dental pulp by loop-mediated isothermal amplification. Forensic Sci Int Genet. 2008, 2,
PGRP negatively regulates NOD-mediated cytokine production in rainbow trout liver cells
 Supplementary Information for: PGRP negatively regulates NOD-mediated cytokine production in rainbow trout liver cells Ju Hye Jang 1, Hyun Kim 2, Mi Jung Jang 2, Ju Hyun Cho 1,2,* 1 Research Institute
Supplementary Information for: PGRP negatively regulates NOD-mediated cytokine production in rainbow trout liver cells Ju Hye Jang 1, Hyun Kim 2, Mi Jung Jang 2, Ju Hyun Cho 1,2,* 1 Research Institute
Electronic Supplementary Information
 Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2017 Electronic Supplementary Information Dissecting binding of a β-barrel outer membrane
Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2017 Electronic Supplementary Information Dissecting binding of a β-barrel outer membrane
Supplement 1: Sequences of Capture Probes. Capture probes were /5AmMC6/CTG TAG GTG CGG GTG GAC GTA GTC
 Supplementary Appendixes Supplement 1: Sequences of Capture Probes. Capture probes were /5AmMC6/CTG TAG GTG CGG GTG GAC GTA GTC ACG TAG CTC CGG CTG GA-3 for vimentin, /5AmMC6/TCC CTC GCG CGT GGC TTC CGC
Supplementary Appendixes Supplement 1: Sequences of Capture Probes. Capture probes were /5AmMC6/CTG TAG GTG CGG GTG GAC GTA GTC ACG TAG CTC CGG CTG GA-3 for vimentin, /5AmMC6/TCC CTC GCG CGT GGC TTC CGC
SUPPLEMENTARY INFORMATION
 1. RNA/DNA sequences used in this study 2. Height and stiffness measurements on hybridized molecules 3. Stiffness maps at varying concentrations of target DNA 4. Stiffness measurements on RNA/DNA hybrids.
1. RNA/DNA sequences used in this study 2. Height and stiffness measurements on hybridized molecules 3. Stiffness maps at varying concentrations of target DNA 4. Stiffness measurements on RNA/DNA hybrids.
Supplementary. Table 1: Oligonucleotides and Plasmids. complementary to positions from 77 of the SRα '- GCT CTA GAG AAC TTG AAG TAC AGA CTG C
 Supplementary Table 1: Oligonucleotides and Plasmids 913954 5'- GCT CTA GAG AAC TTG AAG TAC AGA CTG C 913955 5'- CCC AAG CTT ACA GTG TGG CCA TTC TGC TG 223396 5'- CGA CGC GTA CAG TGT GGC CAT TCT GCT G
Supplementary Table 1: Oligonucleotides and Plasmids 913954 5'- GCT CTA GAG AAC TTG AAG TAC AGA CTG C 913955 5'- CCC AAG CTT ACA GTG TGG CCA TTC TGC TG 223396 5'- CGA CGC GTA CAG TGT GGC CAT TCT GCT G
Lecture 10, 20/2/2002: The process of solution development - The CODEHOP strategy for automatic design of consensus-degenerate primers for PCR
 Lecture 10, 20/2/2002: The process of solution development - The CODEHOP strategy for automatic design of consensus-degenerate primers for PCR 1 The problem We wish to clone a yet unknown gene from a known
Lecture 10, 20/2/2002: The process of solution development - The CODEHOP strategy for automatic design of consensus-degenerate primers for PCR 1 The problem We wish to clone a yet unknown gene from a known
Supplemental Data Supplemental Figure 1.
 Supplemental Data Supplemental Figure 1. Silique arrangement in the wild-type, jhs, and complemented lines. Wild-type (WT) (A), the jhs1 mutant (B,C), and the jhs1 mutant complemented with JHS1 (Com) (D)
Supplemental Data Supplemental Figure 1. Silique arrangement in the wild-type, jhs, and complemented lines. Wild-type (WT) (A), the jhs1 mutant (B,C), and the jhs1 mutant complemented with JHS1 (Com) (D)
Y-chromosomal haplogroup typing Using SBE reaction
 Schematic of multiplex PCR followed by SBE reaction Multiplex PCR Exo SAP purification SBE reaction 5 A 3 ddatp ddgtp 3 T 5 A G 3 T 5 3 5 G C 5 3 3 C 5 ddttp ddctp 5 T 3 T C 3 A 5 3 A 5 5 C 3 3 G 5 3 G
Schematic of multiplex PCR followed by SBE reaction Multiplex PCR Exo SAP purification SBE reaction 5 A 3 ddatp ddgtp 3 T 5 A G 3 T 5 3 5 G C 5 3 3 C 5 ddttp ddctp 5 T 3 T C 3 A 5 3 A 5 5 C 3 3 G 5 3 G
Hes6. PPARα. PPARγ HNF4 CD36
 SUPPLEMENTARY INFORMATION Supplementary Table Positions and Sequences of ChIP primers -63 AGGTCACTGCCA -79 AGGTCTGCTGTG Hes6-0067 GGGCAaAGTTCA ACOT -395 GGGGCAgAGTTCA PPARα -309 GGCTCAaAGTTCAaGTTCA CPTa
SUPPLEMENTARY INFORMATION Supplementary Table Positions and Sequences of ChIP primers -63 AGGTCACTGCCA -79 AGGTCTGCTGTG Hes6-0067 GGGCAaAGTTCA ACOT -395 GGGGCAgAGTTCA PPARα -309 GGCTCAaAGTTCAaGTTCA CPTa
Supporting Information
 Supporting Information Table S1. Oligonucleotide sequences used in this work Oligo DNA A B C D CpG-A CpG-B CpG-C CpG-D Sequence 5 ACA TTC CTA AGT CTG AAA CAT TAC AGC TTG CTA CAC GAG AAG AGC CGC CAT AGT
Supporting Information Table S1. Oligonucleotide sequences used in this work Oligo DNA A B C D CpG-A CpG-B CpG-C CpG-D Sequence 5 ACA TTC CTA AGT CTG AAA CAT TAC AGC TTG CTA CAC GAG AAG AGC CGC CAT AGT
RPA-AB RPA-C Supplemental Figure S1: SDS-PAGE stained with Coomassie Blue after protein purification.
 RPA-AB RPA-C (a) (b) (c) (d) (e) (f) Supplemental Figure S: SDS-PAGE stained with Coomassie Blue after protein purification. (a) RPA; (b) RPA-AB; (c) RPA-CDE; (d) RPA-CDE core; (e) RPA-DE; and (f) RPA-C
RPA-AB RPA-C (a) (b) (c) (d) (e) (f) Supplemental Figure S: SDS-PAGE stained with Coomassie Blue after protein purification. (a) RPA; (b) RPA-AB; (c) RPA-CDE; (d) RPA-CDE core; (e) RPA-DE; and (f) RPA-C
Supporting information for Biochemistry, 1995, 34(34), , DOI: /bi00034a013
 Supporting information for Biochemistry, 1995, 34(34), 10807 10815, DOI: 10.1021/bi00034a013 LESNIK 10807-1081 Terms & Conditions Electronic Supporting Information files are available without a subscription
Supporting information for Biochemistry, 1995, 34(34), 10807 10815, DOI: 10.1021/bi00034a013 LESNIK 10807-1081 Terms & Conditions Electronic Supporting Information files are available without a subscription
Figure S1. Characterization of the irx9l-1 mutant. (A) Diagram of the Arabidopsis IRX9L gene drawn based on information from TAIR (the Arabidopsis
 1 2 3 4 5 6 7 8 9 10 11 12 Figure S1. Characterization of the irx9l-1 mutant. (A) Diagram of the Arabidopsis IRX9L gene drawn based on information from TAIR (the Arabidopsis Information Research). Exons
1 2 3 4 5 6 7 8 9 10 11 12 Figure S1. Characterization of the irx9l-1 mutant. (A) Diagram of the Arabidopsis IRX9L gene drawn based on information from TAIR (the Arabidopsis Information Research). Exons
Multi-Amplified Sensing of MicroRNA by a Small DNA Fragment-Driven Enzymatic Cascade Reaction
 Supporting Information for Multi-Amplified Sensing of MicroRNA by a Small DNA Fragment-Driven Enzymatic Cascade Reaction Eunjung Kim, Philip D. Howes, Spencer W. Crowder, and Molly M. Stevens* Department
Supporting Information for Multi-Amplified Sensing of MicroRNA by a Small DNA Fragment-Driven Enzymatic Cascade Reaction Eunjung Kim, Philip D. Howes, Spencer W. Crowder, and Molly M. Stevens* Department
DNA sentences. How are proteins coded for by DNA? Materials. Teacher instructions. Student instructions. Reflection
 DNA sentences How are proteins coded for by DNA? Deoxyribonucleic acid (DNA) is the molecule of life. DNA is one of the most recognizable nucleic acids, a double-stranded helix. The process by which DNA
DNA sentences How are proteins coded for by DNA? Deoxyribonucleic acid (DNA) is the molecule of life. DNA is one of the most recognizable nucleic acids, a double-stranded helix. The process by which DNA
Supplemental Data. mir156-regulated SPL Transcription. Factors Define an Endogenous Flowering. Pathway in Arabidopsis thaliana
 Cell, Volume 138 Supplemental Data mir156-regulated SPL Transcription Factors Define an Endogenous Flowering Pathway in Arabidopsis thaliana Jia-Wei Wang, Benjamin Czech, and Detlef Weigel Table S1. Interaction
Cell, Volume 138 Supplemental Data mir156-regulated SPL Transcription Factors Define an Endogenous Flowering Pathway in Arabidopsis thaliana Jia-Wei Wang, Benjamin Czech, and Detlef Weigel Table S1. Interaction
Supplemental Information. Human Senataxin Resolves RNA/DNA Hybrids. Formed at Transcriptional Pause Sites. to Promote Xrn2-Dependent Termination
 Supplemental Information Molecular Cell, Volume 42 Human Senataxin Resolves RNA/DNA Hybrids Formed at Transcriptional Pause Sites to Promote Xrn2-Dependent Termination Konstantina Skourti-Stathaki, Nicholas
Supplemental Information Molecular Cell, Volume 42 Human Senataxin Resolves RNA/DNA Hybrids Formed at Transcriptional Pause Sites to Promote Xrn2-Dependent Termination Konstantina Skourti-Stathaki, Nicholas
Gene synthesis by circular assembly amplification
 Gene synthesis by circular assembly amplification Duhee Bang & George M Church Supplementary figures and text: Supplementary Figure 1. Dpo4 gene (1.05kb) construction by various methods. Supplementary
Gene synthesis by circular assembly amplification Duhee Bang & George M Church Supplementary figures and text: Supplementary Figure 1. Dpo4 gene (1.05kb) construction by various methods. Supplementary
Supplementary Figure 1A A404 Cells +/- Retinoic Acid
 Supplementary Figure 1A A44 Cells +/- Retinoic Acid 1 1 H3 Lys4 di-methylation SM-actin VEC cfos (-) RA (+) RA 14 1 1 8 6 4 H3 Lys79 di-methylation SM-actin VEC cfos (-) RA (+) RA Supplementary Figure
Supplementary Figure 1A A44 Cells +/- Retinoic Acid 1 1 H3 Lys4 di-methylation SM-actin VEC cfos (-) RA (+) RA 14 1 1 8 6 4 H3 Lys79 di-methylation SM-actin VEC cfos (-) RA (+) RA Supplementary Figure
Electronic Supplementary Information
 Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2014 Electronic Supplementary Information Multiplexed Detection of Lung Cancer Cells at the Single-Molecule
Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2014 Electronic Supplementary Information Multiplexed Detection of Lung Cancer Cells at the Single-Molecule
Arabidopsis actin depolymerizing factor AtADF4 mediates defense signal transduction triggered by the Pseudomonas syringae effector AvrPphB
 Arabidopsis actin depolymerizing factor mediates defense signal transduction triggered by the Pseudomonas syringae effector AvrPphB Files in this Data Supplement: Supplemental Table S1 Supplemental Table
Arabidopsis actin depolymerizing factor mediates defense signal transduction triggered by the Pseudomonas syringae effector AvrPphB Files in this Data Supplement: Supplemental Table S1 Supplemental Table
Codon Bias with PRISM. 2IM24/25, Fall 2007
 Codon Bias with PRISM 2IM24/25, Fall 2007 from RNA to protein mrna vs. trna aminoacid trna anticodon mrna codon codon-anticodon matching Watson-Crick base pairing A U and C G binding first two nucleotide
Codon Bias with PRISM 2IM24/25, Fall 2007 from RNA to protein mrna vs. trna aminoacid trna anticodon mrna codon codon-anticodon matching Watson-Crick base pairing A U and C G binding first two nucleotide
Supplementary Information. Construction of Lasso Peptide Fusion Proteins
 Supplementary Information Construction of Lasso Peptide Fusion Proteins Chuhan Zong 1, Mikhail O. Maksimov 2, A. James Link 2,3 * Departments of 1 Chemistry, 2 Chemical and Biological Engineering, and
Supplementary Information Construction of Lasso Peptide Fusion Proteins Chuhan Zong 1, Mikhail O. Maksimov 2, A. James Link 2,3 * Departments of 1 Chemistry, 2 Chemical and Biological Engineering, and
Supporting Information
 Supporting Information Transfection of DNA Cages into Mammalian Cells Email: a.turberfield@physics.ox.ac.uk Table of Contents Supporting Figure 1 DNA tetrahedra used in transfection experiments 2 Supporting
Supporting Information Transfection of DNA Cages into Mammalian Cells Email: a.turberfield@physics.ox.ac.uk Table of Contents Supporting Figure 1 DNA tetrahedra used in transfection experiments 2 Supporting
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/10/494/eaan6284/dc1 Supplementary Materials for Activation of master virulence regulator PhoP in acidic ph requires the Salmonella-specific protein UgtL Jeongjoon
www.sciencesignaling.org/cgi/content/full/10/494/eaan6284/dc1 Supplementary Materials for Activation of master virulence regulator PhoP in acidic ph requires the Salmonella-specific protein UgtL Jeongjoon
Supporting Online Information
 Supporting Online Information Isolation of Human Genomic DNA Sequences with Expanded Nucleobase Selectivity Preeti Rathi, Sara Maurer, Grzegorz Kubik and Daniel Summerer* Department of Chemistry and Chemical
Supporting Online Information Isolation of Human Genomic DNA Sequences with Expanded Nucleobase Selectivity Preeti Rathi, Sara Maurer, Grzegorz Kubik and Daniel Summerer* Department of Chemistry and Chemical
A high efficient electrochemiluminescence resonance energy. transfer system in one nanostructure: its application for
 Supporting Information for A high efficient electrochemiluminescence resonance energy transfer system in one nanostructure: its application for ultrasensitive detection of microrna in cancer cells Zhaoyang
Supporting Information for A high efficient electrochemiluminescence resonance energy transfer system in one nanostructure: its application for ultrasensitive detection of microrna in cancer cells Zhaoyang
Phosphate and R2D2 Restrict the Substrate Specificity of Dicer-2, an ATP- Driven Ribonuclease
 Supplemental Information Molecular Cell, Volume 42 hosphate and R2D2 Restrict the Substrate Specificity of Dicer-2, an AT- Driven Ribonuclease Elif Sarinay Cenik, Ryuya Fukunaga, Gang Lu, Robert Dutcher,
Supplemental Information Molecular Cell, Volume 42 hosphate and R2D2 Restrict the Substrate Specificity of Dicer-2, an AT- Driven Ribonuclease Elif Sarinay Cenik, Ryuya Fukunaga, Gang Lu, Robert Dutcher,
Amplified Analysis of DNA by the Autonomous Assembly of Polymers Consisting of DNAzyme Wires
 Supporting information for the paper: Amplified Analysis of DNA by the Autonomous Assembly of Polymers Consisting of DNAzyme Wires Fuan Wang, Johann Elbaz, Ron Orbach, Nimrod Magen and Itamar Willner*
Supporting information for the paper: Amplified Analysis of DNA by the Autonomous Assembly of Polymers Consisting of DNAzyme Wires Fuan Wang, Johann Elbaz, Ron Orbach, Nimrod Magen and Itamar Willner*
Materials Protein synthesis kit. This kit consists of 24 amino acids, 24 transfer RNAs, four messenger RNAs and one ribosome (see below).
 Protein Synthesis Instructions The purpose of today s lab is to: Understand how a cell manufactures proteins from amino acids, using information stored in the genetic code. Assemble models of four very
Protein Synthesis Instructions The purpose of today s lab is to: Understand how a cell manufactures proteins from amino acids, using information stored in the genetic code. Assemble models of four very
Cat. # Product Size DS130 DynaExpress TA PCR Cloning Kit (ptakn-2) 20 reactions Box 1 (-20 ) ptakn-2 Vector, linearized 20 µl (50 ng/µl) 1
 Product Name: Kit Component TA PCR Cloning Kit (ptakn-2) Cat. # Product Size DS130 TA PCR Cloning Kit (ptakn-2) 20 reactions Box 1 (-20 ) ptakn-2 Vector, linearized 20 µl (50 ng/µl) 1 2 Ligation Buffer
Product Name: Kit Component TA PCR Cloning Kit (ptakn-2) Cat. # Product Size DS130 TA PCR Cloning Kit (ptakn-2) 20 reactions Box 1 (-20 ) ptakn-2 Vector, linearized 20 µl (50 ng/µl) 1 2 Ligation Buffer
Molecular Level of Genetics
 Molecular Level of Genetics Most of the molecules found in humans and other living organisms fall into one of four categories: 1. carbohydrates (sugars and starches) 2. lipids (fats, oils, and waxes) 3.
Molecular Level of Genetics Most of the molecules found in humans and other living organisms fall into one of four categories: 1. carbohydrates (sugars and starches) 2. lipids (fats, oils, and waxes) 3.
PROTEIN SYNTHESIS Study Guide
 PART A. Read the following: PROTEIN SYNTHESIS Study Guide Protein synthesis is the process used by the body to make proteins. The first step of protein synthesis is called Transcription. It occurs in the
PART A. Read the following: PROTEIN SYNTHESIS Study Guide Protein synthesis is the process used by the body to make proteins. The first step of protein synthesis is called Transcription. It occurs in the
Supplementary Figure 1. Ratiometric fluorescence visualization of DNA cleavage by
 Supplementary Figure 1. Ratiometric fluorescence visualization of DNA cleavage by Cas9:gRNA. (a) A labeled by Cy3 and Cy5 with an inter-probe distance of > 30 bp was tethered to a PEG-coated surface via
Supplementary Figure 1. Ratiometric fluorescence visualization of DNA cleavage by Cas9:gRNA. (a) A labeled by Cy3 and Cy5 with an inter-probe distance of > 30 bp was tethered to a PEG-coated surface via
Electronic Supplementary Information
 Electronic Supplementary Information Ultrasensitive and Selective DNA Detection Based on Nicking Endonuclease Assisted Signal Amplification and Its Application in Cancer Cells Detection Sai Bi, Jilei Zhang,
Electronic Supplementary Information Ultrasensitive and Selective DNA Detection Based on Nicking Endonuclease Assisted Signal Amplification and Its Application in Cancer Cells Detection Sai Bi, Jilei Zhang,
Supplemental Data. Bennett et al. (2010). Plant Cell /tpc
 BRN1 ---------MSSSNGGVPPGFRFHPTDEELLHYYLKKKISYEKFEMEVIKEVDLNKIEPWDLQDRCKIGSTPQNEWYFFSHKDRKYPTGS 81 BRN2 --------MGSSSNGGVPPGFRFHPTDEELLHYYLKKKISYQKFEMEVIREVDLNKLEPWDLQERCKIGSTPQNEWYFFSHKDRKYPTGS 82 SMB
BRN1 ---------MSSSNGGVPPGFRFHPTDEELLHYYLKKKISYEKFEMEVIKEVDLNKIEPWDLQDRCKIGSTPQNEWYFFSHKDRKYPTGS 81 BRN2 --------MGSSSNGGVPPGFRFHPTDEELLHYYLKKKISYQKFEMEVIREVDLNKLEPWDLQERCKIGSTPQNEWYFFSHKDRKYPTGS 82 SMB
Disease and selection in the human genome 3
 Disease and selection in the human genome 3 Ka/Ks revisited Please sit in row K or forward RBFD: human populations, adaptation and immunity Neandertal Museum, Mettman Germany Sequence genome Measure expression
Disease and selection in the human genome 3 Ka/Ks revisited Please sit in row K or forward RBFD: human populations, adaptation and immunity Neandertal Museum, Mettman Germany Sequence genome Measure expression
ΔPDD1 x ΔPDD1. ΔPDD1 x wild type. 70 kd Pdd1. Pdd3
 Supplemental Fig. S1 ΔPDD1 x wild type ΔPDD1 x ΔPDD1 70 kd Pdd1 50 kd 37 kd Pdd3 Supplemental Fig. S1. ΔPDD1 strains express no detectable Pdd1 protein. Western blot analysis of whole-protein extracts
Supplemental Fig. S1 ΔPDD1 x wild type ΔPDD1 x ΔPDD1 70 kd Pdd1 50 kd 37 kd Pdd3 Supplemental Fig. S1. ΔPDD1 strains express no detectable Pdd1 protein. Western blot analysis of whole-protein extracts
Add 5µl of 3N NaOH to DNA sample (final concentration 0.3N NaOH).
 Bisulfite Treatment of DNA Dilute DNA sample to 2µg DNA in 50µl ddh 2 O. Add 5µl of 3N NaOH to DNA sample (final concentration 0.3N NaOH). Incubate in a 37ºC water bath for 30 minutes. To 55µl samples
Bisulfite Treatment of DNA Dilute DNA sample to 2µg DNA in 50µl ddh 2 O. Add 5µl of 3N NaOH to DNA sample (final concentration 0.3N NaOH). Incubate in a 37ºC water bath for 30 minutes. To 55µl samples
PCR analysis was performed to show the presence and the integrity of the var1csa and var-
 Supplementary information: Methods: Table S1: Primer Name Nucleotide sequence (5-3 ) DBL3-F tcc ccg cgg agt gaa aca tca tgt gac tg DBL3-R gac tag ttt ctt tca ata aat cac tcg c DBL5-F cgc cct agg tgc ttc
Supplementary information: Methods: Table S1: Primer Name Nucleotide sequence (5-3 ) DBL3-F tcc ccg cgg agt gaa aca tca tgt gac tg DBL3-R gac tag ttt ctt tca ata aat cac tcg c DBL5-F cgc cct agg tgc ttc
Supplementary Figures
 Supplementary Figures Supplementary Fig. 1 Characterization of GSCs. a. Immunostaining of primary GSC spheres from GSC lines. Nestin (neural progenitor marker, red), TLX (green). Merged images of nestin,
Supplementary Figures Supplementary Fig. 1 Characterization of GSCs. a. Immunostaining of primary GSC spheres from GSC lines. Nestin (neural progenitor marker, red), TLX (green). Merged images of nestin,
Dierks Supplementary Fig. S1
 Dierks Supplementary Fig. S1 ITK SYK PH TH K42R wt K42R (kinase deficient) R29C E42K Y323F R29C E42K Y323F (reduced phospholipid binding) (enhanced phospholipid binding) (reduced Cbl binding) E42K Y323F
Dierks Supplementary Fig. S1 ITK SYK PH TH K42R wt K42R (kinase deficient) R29C E42K Y323F R29C E42K Y323F (reduced phospholipid binding) (enhanced phospholipid binding) (reduced Cbl binding) E42K Y323F
SUPPORTING INFORMATION
 SUPPORTING INFORMATION Investigation of the Biosynthesis of the Lasso Peptide Chaxapeptin Using an E. coli-based Production System Helena Martin-Gómez, Uwe Linne, Fernando Albericio, Judit Tulla-Puche,*
SUPPORTING INFORMATION Investigation of the Biosynthesis of the Lasso Peptide Chaxapeptin Using an E. coli-based Production System Helena Martin-Gómez, Uwe Linne, Fernando Albericio, Judit Tulla-Puche,*
DNA Computing Circuits Using Libraries of. DNAzyme Subunits
 SUPPLEMENTARY INFORMATION Supplementary Information for the paper: DNA Computing Circuits Using Libraries of DNAzyme Subunits Johann Elbaz a, Oleg lioubashevski a, Fuan Wang a, Françoise Remacle b, Raphael
SUPPLEMENTARY INFORMATION Supplementary Information for the paper: DNA Computing Circuits Using Libraries of DNAzyme Subunits Johann Elbaz a, Oleg lioubashevski a, Fuan Wang a, Françoise Remacle b, Raphael
II 0.95 DM2 (RPP1) DM3 (At3g61540) b
 Table S2. F 2 Segregation Ratios at 16 C, Related to Figure 2 Cross n c Phenotype Model e 2 Locus A Locus B Normal F 1 -like Enhanced d Uk-1/Uk-3 149 64 36 49 DM2 (RPP1) DM1 (SSI4) a Bla-1/Hh-0 F 3 111
Table S2. F 2 Segregation Ratios at 16 C, Related to Figure 2 Cross n c Phenotype Model e 2 Locus A Locus B Normal F 1 -like Enhanced d Uk-1/Uk-3 149 64 36 49 DM2 (RPP1) DM1 (SSI4) a Bla-1/Hh-0 F 3 111
SAY IT WITH DNA: Protein Synthesis Activity by Larry Flammer
 TEACHER S GUIDE SAY IT WITH DNA: Protein Synthesis Activity by Larry Flammer SYNOPSIS This activity uses the metaphor of decoding a secret message for the Protein Synthesis process. Students teach themselves
TEACHER S GUIDE SAY IT WITH DNA: Protein Synthesis Activity by Larry Flammer SYNOPSIS This activity uses the metaphor of decoding a secret message for the Protein Synthesis process. Students teach themselves
Supporting Information. Copyright Wiley-VCH Verlag GmbH & Co. KGaA, Weinheim, 2006
 Supporting Information Copyright Wiley-VCH Verlag GmbH & Co. KGaA, 69451 Weinheim, 2006 Copyright Wiley-VCH Verlag GmbH & Co. KGaA, 69451 Weinheim, 2006 Supporting Information for Expanding the Genetic
Supporting Information Copyright Wiley-VCH Verlag GmbH & Co. KGaA, 69451 Weinheim, 2006 Copyright Wiley-VCH Verlag GmbH & Co. KGaA, 69451 Weinheim, 2006 Supporting Information for Expanding the Genetic
Multiplexing Genome-scale Engineering
 Multiplexing Genome-scale Engineering Harris Wang, Ph.D. Department of Systems Biology Department of Pathology & Cell Biology http://wanglab.c2b2.columbia.edu Rise of Genomics An Expanding Toolbox Esvelt
Multiplexing Genome-scale Engineering Harris Wang, Ph.D. Department of Systems Biology Department of Pathology & Cell Biology http://wanglab.c2b2.columbia.edu Rise of Genomics An Expanding Toolbox Esvelt
ORFs and genes. Please sit in row K or forward
 ORFs and genes Please sit in row K or forward https://www.flickr.com/photos/teseum/3231682806/in/photostream/ Question: why do some strains of Vibrio cause cholera and others don t? Methods Mechanisms
ORFs and genes Please sit in row K or forward https://www.flickr.com/photos/teseum/3231682806/in/photostream/ Question: why do some strains of Vibrio cause cholera and others don t? Methods Mechanisms
evaluated with UAS CLB eliciting UAS CIT -N Libraries increase in the
 Supplementary Figures Supplementary Figure 1: Promoter scaffold library assemblies. Many ensembless of libraries were evaluated in this work. As a legend, the box outline color in top half of the figure
Supplementary Figures Supplementary Figure 1: Promoter scaffold library assemblies. Many ensembless of libraries were evaluated in this work. As a legend, the box outline color in top half of the figure
Supplementary Information
 Supplementary Information A general solution for opening double-stranded DNA for isothermal amplification Gangyi Chen, Juan Dong, Yi Yuan, Na Li, Xin Huang, Xin Cui* and Zhuo Tang* Supplementary Materials
Supplementary Information A general solution for opening double-stranded DNA for isothermal amplification Gangyi Chen, Juan Dong, Yi Yuan, Na Li, Xin Huang, Xin Cui* and Zhuo Tang* Supplementary Materials
NAME:... MODEL ANSWER... STUDENT NUMBER:... Maximum marks: 50. Internal Examiner: Hugh Murrell, Computer Science, UKZN
 COMP710, Bioinformatics with Julia, Test One, Thursday the 20 th of April, 2017, 09h30-11h30 1 NAME:...... MODEL ANSWER... STUDENT NUMBER:...... Maximum marks: 50 Internal Examiner: Hugh Murrell, Computer
COMP710, Bioinformatics with Julia, Test One, Thursday the 20 th of April, 2017, 09h30-11h30 1 NAME:...... MODEL ANSWER... STUDENT NUMBER:...... Maximum marks: 50 Internal Examiner: Hugh Murrell, Computer
SUPPORTING INFORMATION FILE
 Intrinsic and extrinsic connections of Tet3 dioxygenase with CXXC zinc finger modules Nan Liu, Mengxi Wang, Wen Deng, Christine S. Schmidt, Weihua Qin, Heinrich Leonhardt and Fabio Spada Department of
Intrinsic and extrinsic connections of Tet3 dioxygenase with CXXC zinc finger modules Nan Liu, Mengxi Wang, Wen Deng, Christine S. Schmidt, Weihua Qin, Heinrich Leonhardt and Fabio Spada Department of
Degenerate Code. Translation. trna. The Code is Degenerate trna / Proofreading Ribosomes Translation Mechanism
 Translation The Code is Degenerate trna / Proofreading Ribosomes Translation Mechanism Degenerate Code There are 64 possible codon triplets There are 20 naturally-encoding amino acids Several codons specify
Translation The Code is Degenerate trna / Proofreading Ribosomes Translation Mechanism Degenerate Code There are 64 possible codon triplets There are 20 naturally-encoding amino acids Several codons specify
hcd1tg/hj1tg/ ApoE-/- hcd1tg/hj1tg/ ApoE+/+
 ApoE+/+ ApoE-/- ApoE-/- H&E (1x) Supplementary Figure 1. No obvious pathology is observed in the colon of diseased ApoE-/me. Colon samples were fixed in 1% formalin and laid out in Swiss rolls for paraffin
ApoE+/+ ApoE-/- ApoE-/- H&E (1x) Supplementary Figure 1. No obvious pathology is observed in the colon of diseased ApoE-/me. Colon samples were fixed in 1% formalin and laid out in Swiss rolls for paraffin
strain devoid of the aox1 gene [1]. Thus, the identification of AOX1 in the intracellular
![strain devoid of the aox1 gene [1]. Thus, the identification of AOX1 in the intracellular strain devoid of the aox1 gene [1]. Thus, the identification of AOX1 in the intracellular](/thumbs/72/66361212.jpg) Additional file 2 Identification of AOX1 in P. pastoris GS115 with a Mut s phenotype Results and Discussion The HBsAg producing strain was originally identified as a Mut s (methanol utilization slow) strain
Additional file 2 Identification of AOX1 in P. pastoris GS115 with a Mut s phenotype Results and Discussion The HBsAg producing strain was originally identified as a Mut s (methanol utilization slow) strain
Supplemental material
 Supplemental material Diversity of O-antigen repeat-unit structures can account for the substantial sequence variation of Wzx translocases Yaoqin Hong and Peter R. Reeves School of Molecular Bioscience,
Supplemental material Diversity of O-antigen repeat-unit structures can account for the substantial sequence variation of Wzx translocases Yaoqin Hong and Peter R. Reeves School of Molecular Bioscience,
Search for and Analysis of Single Nucleotide Polymorphisms (SNPs) in Rice (Oryza sativa, Oryza rufipogon) and Establishment of SNP Markers
 DNA Research 9, 163 171 (2002) Search for and Analysis of Single Nucleotide Polymorphisms (SNPs) in Rice (Oryza sativa, Oryza rufipogon) and Establishment of SNP Markers Shinobu Nasu, Junko Suzuki, Rieko
DNA Research 9, 163 171 (2002) Search for and Analysis of Single Nucleotide Polymorphisms (SNPs) in Rice (Oryza sativa, Oryza rufipogon) and Establishment of SNP Markers Shinobu Nasu, Junko Suzuki, Rieko
Human Gene,cs 06: Gene Expression. Diversity of cell types. How do cells become different? 9/19/11. neuron
 Human Gene,cs 06: Gene Expression 20110920 Diversity of cell types neuron How do cells become different? A. Each type of cell has different DNA in its nucleus B. Each cell has different genes C. Each type
Human Gene,cs 06: Gene Expression 20110920 Diversity of cell types neuron How do cells become different? A. Each type of cell has different DNA in its nucleus B. Each cell has different genes C. Each type
A green method of staining DNA in polyacrylamide gel electrophoresis based on fluorescent copper nanoclusters synthesized in situ
 Electronic Supplementary Material A green method of staining DNA in polyacrylamide gel electrophoresis based on fluorescent copper nanoclusters synthesized in situ Xiaoli Zhu 1, Hai Shi 1, Yalan Shen 1,
Electronic Supplementary Material A green method of staining DNA in polyacrylamide gel electrophoresis based on fluorescent copper nanoclusters synthesized in situ Xiaoli Zhu 1, Hai Shi 1, Yalan Shen 1,
Project 07/111 Final Report October 31, Project Title: Cloning and expression of porcine complement C3d for enhanced vaccines
 Project 07/111 Final Report October 31, 2007. Project Title: Cloning and expression of porcine complement C3d for enhanced vaccines Project Leader: Dr Douglas C. Hodgins (519-824-4120 Ex 54758, fax 519-824-5930)
Project 07/111 Final Report October 31, 2007. Project Title: Cloning and expression of porcine complement C3d for enhanced vaccines Project Leader: Dr Douglas C. Hodgins (519-824-4120 Ex 54758, fax 519-824-5930)
Supplementary Information
 Supplementary Information Minimal mechanistic model of sirna-dependent target RNA slicing by recombinant human Argonaute2 protein Andrea Deerberg*, Sarah Willkomm* and Tobias Restle Institute of Molecular
Supplementary Information Minimal mechanistic model of sirna-dependent target RNA slicing by recombinant human Argonaute2 protein Andrea Deerberg*, Sarah Willkomm* and Tobias Restle Institute of Molecular
Table S1. Bacterial strains (Related to Results and Experimental Procedures)
 Table S1. Bacterial strains (Related to Results and Experimental Procedures) Strain number Relevant genotype Source or reference 1045 AB1157 Graham Walker (Donnelly and Walker, 1989) 2458 3084 (MG1655)
Table S1. Bacterial strains (Related to Results and Experimental Procedures) Strain number Relevant genotype Source or reference 1045 AB1157 Graham Walker (Donnelly and Walker, 1989) 2458 3084 (MG1655)
ANCIENT BACTERIA? 250 million years later, scientists revive life forms
 ANCIENT BACTERIA? 250 million years later, scientists revive life forms Thursday, October 19, 2000 U.S. researchers say they have revived bacteria that have been dormant for more then 250 million years,
ANCIENT BACTERIA? 250 million years later, scientists revive life forms Thursday, October 19, 2000 U.S. researchers say they have revived bacteria that have been dormant for more then 250 million years,
Supporting Information. Trifluoroacetophenone-Linked Nucleotides and DNA for Studying of DNA-protein Interactions by 19 F NMR Spectroscopy
 Supporting Information Trifluoroacetophenone-Linked Nucleotides and DNA for Studying of DNA-protein Interactions by 19 F NMR Spectroscopy Agata Olszewska, Radek Pohl and Michal Hocek # * Institute of Organic
Supporting Information Trifluoroacetophenone-Linked Nucleotides and DNA for Studying of DNA-protein Interactions by 19 F NMR Spectroscopy Agata Olszewska, Radek Pohl and Michal Hocek # * Institute of Organic
Chapter 3: Information Storage and Transfer in Life
 Chapter 3: Information Storage and Transfer in Life The trapped scientist examples are great for conceptual purposes, but they do not accurately model how information in life changes because they do not
Chapter 3: Information Storage and Transfer in Life The trapped scientist examples are great for conceptual purposes, but they do not accurately model how information in life changes because they do not
Anti-Pim-1 (Cat#3247), anti-met (Cat#3127), anti-ron (Cat#2654), Anti-EGFR
 Supplementary Methods Antibodies Anti-Pim-1 (Cat#3247), anti-met (Cat#3127), anti-ron (Cat#2654), Anti-EGFR (Cat#2646), anti-igf1r (Cat#3018), anti-insr (Cat#3020), anti-akt (pan, Cat#4691), anti-phospho-akt
Supplementary Methods Antibodies Anti-Pim-1 (Cat#3247), anti-met (Cat#3127), anti-ron (Cat#2654), Anti-EGFR (Cat#2646), anti-igf1r (Cat#3018), anti-insr (Cat#3020), anti-akt (pan, Cat#4691), anti-phospho-akt
Legends for supplementary figures 1-3
 High throughput resistance profiling of Plasmodium falciparum infections based on custom dual indexing and Illumina next generation sequencing-technology Sidsel Nag 1,2 *, Marlene D. Dalgaard 3, Poul-Erik
High throughput resistance profiling of Plasmodium falciparum infections based on custom dual indexing and Illumina next generation sequencing-technology Sidsel Nag 1,2 *, Marlene D. Dalgaard 3, Poul-Erik
G+C content. 1 Introduction. 2 Chromosomes Topology & Counts. 3 Genome size. 4 Replichores and gene orientation. 5 Chirochores.
 1 Introduction 2 Chromosomes Topology & Counts 3 Genome size 4 Replichores and gene orientation 5 Chirochores 6 7 Codon usage 121 marc.bailly-bechet@univ-lyon1.fr Bacterial genome structures Introduction
1 Introduction 2 Chromosomes Topology & Counts 3 Genome size 4 Replichores and gene orientation 5 Chirochores 6 7 Codon usage 121 marc.bailly-bechet@univ-lyon1.fr Bacterial genome structures Introduction
Overexpression Normal expression Overexpression Normal expression. 26 (21.1%) N (%) P-value a N (%)
 SUPPLEMENTARY TABLES Table S1. Alteration of ZNF322A protein expression levels in relation to clinicopathological parameters in 123 Asian and 74 Caucasian lung cancer patients. Asian patients Caucasian
SUPPLEMENTARY TABLES Table S1. Alteration of ZNF322A protein expression levels in relation to clinicopathological parameters in 123 Asian and 74 Caucasian lung cancer patients. Asian patients Caucasian
Electronic Supplementary Information Sensitive detection of polynucleotide kinase using rolling circle amplification-induced chemiluminescence
 Electronic Supplementary Material (ESI) for Chemical Communications. This journal is The Royal Society of Chemistry 2014 Electronic Supplementary Information Sensitive detection of polynucleotide kinase
Electronic Supplementary Material (ESI) for Chemical Communications. This journal is The Royal Society of Chemistry 2014 Electronic Supplementary Information Sensitive detection of polynucleotide kinase
Just one nucleotide! Exploring the effects of random single nucleotide mutations
 Dr. Beatriz Gonzalez In-Class Worksheet Name: Learning Objectives: Just one nucleotide! Exploring the effects of random single nucleotide mutations Given a coding DNA sequence, determine the mrna Based
Dr. Beatriz Gonzalez In-Class Worksheet Name: Learning Objectives: Just one nucleotide! Exploring the effects of random single nucleotide mutations Given a coding DNA sequence, determine the mrna Based
UNIT I RNA AND TYPES R.KAVITHA,M.PHARM LECTURER DEPARTMENT OF PHARMACEUTICS SRM COLLEGE OF PHARMACY KATTANKULATUR
 UNIT I RNA AND TYPES R.KAVITHA,M.PHARM LECTURER DEPARTMENT OF PHARMACEUTICS SRM COLLEGE OF PHARMACY KATTANKULATUR RNA, as previously mentioned, is an acronym for ribonucleic acid. There are many forms
UNIT I RNA AND TYPES R.KAVITHA,M.PHARM LECTURER DEPARTMENT OF PHARMACEUTICS SRM COLLEGE OF PHARMACY KATTANKULATUR RNA, as previously mentioned, is an acronym for ribonucleic acid. There are many forms
Biomolecules: lecture 6
 Biomolecules: lecture 6 - to learn the basics on how DNA serves to make RNA = transcription - to learn how the genetic code instructs protein synthesis - to learn the basics on how proteins are synthesized
Biomolecules: lecture 6 - to learn the basics on how DNA serves to make RNA = transcription - to learn how the genetic code instructs protein synthesis - to learn the basics on how proteins are synthesized
Lecture 19A. DNA computing
 Lecture 19A. DNA computing What exactly is DNA (deoxyribonucleic acid)? DNA is the material that contains codes for the many physical characteristics of every living creature. Your cells use different
Lecture 19A. DNA computing What exactly is DNA (deoxyribonucleic acid)? DNA is the material that contains codes for the many physical characteristics of every living creature. Your cells use different
A netlike rolling circle nucleic acid amplification technique
 Electronic Supplementary Material (ESI) for Analyst. This journal is The Royal Society of Chemistry 2014 Supplementary Information A netlike rolling circle nucleic acid amplification technique Xiaoli Zhu,
Electronic Supplementary Material (ESI) for Analyst. This journal is The Royal Society of Chemistry 2014 Supplementary Information A netlike rolling circle nucleic acid amplification technique Xiaoli Zhu,
CONVERGENT EVOLUTION. Def n acquisition of some biological trait but different lineages
 CONVERGENT EVOLUTION Def n acquisition of some biological trait but different lineages Living Rock cactus Baseball plant THE QUESTION From common ancestor or independent acquisition? By Lineage By Convergence
CONVERGENT EVOLUTION Def n acquisition of some biological trait but different lineages Living Rock cactus Baseball plant THE QUESTION From common ancestor or independent acquisition? By Lineage By Convergence
Supplementary information
 Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2017 Supplementary information Structural insights into the EthR-DNA interaction from native mass spectrometry
Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2017 Supplementary information Structural insights into the EthR-DNA interaction from native mass spectrometry
Mismatch discrimination of lipidated DNA and LNA-probes (LiNAs) in hybridization-controlled liposome assembly
 Electronic Supplementary Material (ESI) for Organic & Biomolecular Chemistry. This journal is The Royal Society of Chemistry 2016 SUPPORTING INFORMATION FOR Mismatch discrimination of lipidated DNA and
Electronic Supplementary Material (ESI) for Organic & Biomolecular Chemistry. This journal is The Royal Society of Chemistry 2016 SUPPORTING INFORMATION FOR Mismatch discrimination of lipidated DNA and
Genomics and Gene Recognition Genes and Blue Genes
 Genomics and Gene Recognition Genes and Blue Genes November 1, 2004 Prokaryotic Gene Structure prokaryotes are simplest free-living organisms studying prokaryotes can give us a sense what is the minimum
Genomics and Gene Recognition Genes and Blue Genes November 1, 2004 Prokaryotic Gene Structure prokaryotes are simplest free-living organisms studying prokaryotes can give us a sense what is the minimum
Protein Synthesis. Application Based Questions
 Protein Synthesis Application Based Questions MRNA Triplet Codons Note: Logic behind the single letter abbreviations can be found at: http://www.biology.arizona.edu/biochemistry/problem_sets/aa/dayhoff.html
Protein Synthesis Application Based Questions MRNA Triplet Codons Note: Logic behind the single letter abbreviations can be found at: http://www.biology.arizona.edu/biochemistry/problem_sets/aa/dayhoff.html
INTRODUCTION TO THE MOLECULAR GENETICS OF THE COLOR MUTATIONS IN ROCK POCKET MICE
 The Making of the The Fittest: Making of the Fittest Natural Selection Natural and Adaptation Selection and Adaptation Educator Materials TEACHER MATERIALS INTRODUCTION TO THE MOLECULAR GENETICS OF THE
The Making of the The Fittest: Making of the Fittest Natural Selection Natural and Adaptation Selection and Adaptation Educator Materials TEACHER MATERIALS INTRODUCTION TO THE MOLECULAR GENETICS OF THE
(a) Which enzyme(s) make 5' - 3' phosphodiester bonds? (c) Which enzyme(s) make single-strand breaks in DNA backbones?
 EXAMPLE QUESTIONS AND ANSWERS 1. Topoisomerase does which one of the following? (a) Makes new DNA strands. (b) Unties knots in DNA molecules. (c) Joins the ends of double-stranded DNA molecules. (d) Is
EXAMPLE QUESTIONS AND ANSWERS 1. Topoisomerase does which one of the following? (a) Makes new DNA strands. (b) Unties knots in DNA molecules. (c) Joins the ends of double-stranded DNA molecules. (d) Is
Quantitative reverse-transcription PCR. Transcript levels of flgs, flgr, flia and flha were
 1 Supplemental methods 2 3 4 5 6 7 8 9 1 11 12 13 14 15 16 17 18 19 21 22 23 Quantitative reverse-transcription PCR. Transcript levels of flgs, flgr, flia and flha were monitored by quantitative reverse-transcription
1 Supplemental methods 2 3 4 5 6 7 8 9 1 11 12 13 14 15 16 17 18 19 21 22 23 Quantitative reverse-transcription PCR. Transcript levels of flgs, flgr, flia and flha were monitored by quantitative reverse-transcription
SUPPLEMENTARY INFORMATION
 doi: 10.1038/nature07182 SUPPLEMENTAL FIGURES AND TABLES Fig. S1. myf5-expressing cells give rise to brown fat depots and skeletal muscle (a) Perirenal BAT from control (cre negative) and myf5-cre:r26r3-yfp
doi: 10.1038/nature07182 SUPPLEMENTAL FIGURES AND TABLES Fig. S1. myf5-expressing cells give rise to brown fat depots and skeletal muscle (a) Perirenal BAT from control (cre negative) and myf5-cre:r26r3-yfp
Fishy Amino Acid Codon. UUU Phe UCU Ser UAU Tyr UGU Cys. UUC Phe UCC Ser UAC Tyr UGC Cys. UUA Leu UCA Ser UAA Stop UGA Stop
 Fishy Code Slips Fish 1 GGTTATAGAGGTACTACC Fish 2 GGCTTCAGAGGTACTACC Fish 3 CATAGCAGAGGTACTACC Fish 4 GGTTATTCTGTCTTATTG Fish 5 GGCTTCTCTGTCTTATTG Fish 6 CATAGCGCTGCAACTACC Fishy Amino Acid Codon UUU Phe
Fishy Code Slips Fish 1 GGTTATAGAGGTACTACC Fish 2 GGCTTCAGAGGTACTACC Fish 3 CATAGCAGAGGTACTACC Fish 4 GGTTATTCTGTCTTATTG Fish 5 GGCTTCTCTGTCTTATTG Fish 6 CATAGCGCTGCAACTACC Fishy Amino Acid Codon UUU Phe
Protein Synthesis: Transcription and Translation
 Review Protein Synthesis: Transcription and Translation Central Dogma of Molecular Biology Protein synthesis requires two steps: transcription and translation. DNA contains codes Three bases in DNA code
Review Protein Synthesis: Transcription and Translation Central Dogma of Molecular Biology Protein synthesis requires two steps: transcription and translation. DNA contains codes Three bases in DNA code
Supporting Information
 Supporting Information An Enzyme-Powered Three Dimensional DNA Nanomachine for DNA Walking, Payload Release, and Biosensing Xiaolong Yang, Yanan Tang, Sean D. Mason, Junbo Chen, Feng Li * Department of
Supporting Information An Enzyme-Powered Three Dimensional DNA Nanomachine for DNA Walking, Payload Release, and Biosensing Xiaolong Yang, Yanan Tang, Sean D. Mason, Junbo Chen, Feng Li * Department of
7.016 Problem Set 3. 1 st Pedigree
 7.016 Problem Set 3 Question 1 The following human pedigree shows the inheritance pattern of a specific disease within a family. Assume that the individuals marrying into the family for all generations
7.016 Problem Set 3 Question 1 The following human pedigree shows the inheritance pattern of a specific disease within a family. Assume that the individuals marrying into the family for all generations
1. DNA, RNA structure. 2. DNA replication. 3. Transcription, translation
 1. DNA, RNA structure 2. DNA replication 3. Transcription, translation DNA and RNA are polymers of nucleotides DNA is a nucleic acid, made of long chains of nucleotides Nucleotide Phosphate group Nitrogenous
1. DNA, RNA structure 2. DNA replication 3. Transcription, translation DNA and RNA are polymers of nucleotides DNA is a nucleic acid, made of long chains of nucleotides Nucleotide Phosphate group Nitrogenous
Table S1. Sequences of mutagenesis primers used to create altered rdpa- and sdpa genes
 Supplementary Table and Figures for Structural Basis for the Enantiospecificities of R- and S-Specific Phenoxypropionate/α-Ketoglutarate Dioxygenases by Tina A. Müller, Maria I. Zavodszky, Michael Feig,
Supplementary Table and Figures for Structural Basis for the Enantiospecificities of R- and S-Specific Phenoxypropionate/α-Ketoglutarate Dioxygenases by Tina A. Müller, Maria I. Zavodszky, Michael Feig,
Bioinformatics CSM17 Week 6: DNA, RNA and Proteins
 Bioinformatics CSM17 Week 6: DNA, RNA and Proteins Transcription (reading the DNA template) Translation (RNA -> protein) Protein Structure Transcription - reading the data enzyme - transcriptase gene opens
Bioinformatics CSM17 Week 6: DNA, RNA and Proteins Transcription (reading the DNA template) Translation (RNA -> protein) Protein Structure Transcription - reading the data enzyme - transcriptase gene opens
Deoxyribonucleic Acid DNA. Structure of DNA. Structure of DNA. Nucleotide. Nucleotides 5/13/2013
 Deoxyribonucleic Acid DNA The Secret of Life DNA is the molecule responsible for controlling the activities of the cell It is the hereditary molecule DNA directs the production of protein In 1953, Watson
Deoxyribonucleic Acid DNA The Secret of Life DNA is the molecule responsible for controlling the activities of the cell It is the hereditary molecule DNA directs the production of protein In 1953, Watson
Level 2 Biology, 2017
 91159 911590 2SUPERVISOR S Level 2 Biology, 2017 91159 Demonstrate understanding of gene expression 2.00 p.m. Wednesday 22 November 2017 Credits: Four Achievement Achievement with Merit Achievement with
91159 911590 2SUPERVISOR S Level 2 Biology, 2017 91159 Demonstrate understanding of gene expression 2.00 p.m. Wednesday 22 November 2017 Credits: Four Achievement Achievement with Merit Achievement with
An engineered tryptophan zipper-type peptide as a molecular recognition scaffold
 SUPPLEMENTARY MATERIAL An engineered tryptophan zipper-type peptide as a molecular recognition scaffold Zihao Cheng and Robert E. Campbell* Supplementary Methods Library construction for FRET-based screening
SUPPLEMENTARY MATERIAL An engineered tryptophan zipper-type peptide as a molecular recognition scaffold Zihao Cheng and Robert E. Campbell* Supplementary Methods Library construction for FRET-based screening
The B1 Protein Guides the Biosynthesis of a Lasso Peptide
 The B1 Protein Guides the Biosynthesis of a Lasso Peptide Shaozhou Zhu 1,2, Christopher D. Fage 1, Julian D. Hegemann 1, Andreas Mielcarek 1, Dushan Yan 1, Uwe Linne 1 & Mohamed A. Marahiel*,1 1 Department
The B1 Protein Guides the Biosynthesis of a Lasso Peptide Shaozhou Zhu 1,2, Christopher D. Fage 1, Julian D. Hegemann 1, Andreas Mielcarek 1, Dushan Yan 1, Uwe Linne 1 & Mohamed A. Marahiel*,1 1 Department
Supplemental Table 1. Primers used for PCR.
 Supplemental Table 1. Primers used for PCR. Gene Type Primer Sequence Genotyping and semi-quantitative RT-PCR F 5 -TTG CCC GAT CAC CAT CTG TA-3 rwa1-1 R 5 -TGT AGC GAT CAA GGC CTG ATC TAA-3 LB 5 -TAG CAT
Supplemental Table 1. Primers used for PCR. Gene Type Primer Sequence Genotyping and semi-quantitative RT-PCR F 5 -TTG CCC GAT CAC CAT CTG TA-3 rwa1-1 R 5 -TGT AGC GAT CAA GGC CTG ATC TAA-3 LB 5 -TAG CAT
Lecture 11: Gene Prediction
 Lecture 11: Gene Prediction Study Chapter 6.11-6.14 1 Gene: A sequence of nucleotides coding for protein Gene Prediction Problem: Determine the beginning and end positions of genes in a genome Where are
Lecture 11: Gene Prediction Study Chapter 6.11-6.14 1 Gene: A sequence of nucleotides coding for protein Gene Prediction Problem: Determine the beginning and end positions of genes in a genome Where are
Rapid detection of a Dengue virus RNA sequence with single molecule sensitivity using tandem toehold-mediated displacement reactions
 Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2018 Electronic Supplementary Information (ESI) for Rapid detection of a Dengue virus RNA sequence with
Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2018 Electronic Supplementary Information (ESI) for Rapid detection of a Dengue virus RNA sequence with
Supporting Information
 Supporting Information Wiley-VCH 2007 69451 Weinheim, Germany Site-Specific Control of Distances between Gold Nanoparticles using Phosphorothioate Anchors on DNA and a Short Bifunctional Molecular Fastener
Supporting Information Wiley-VCH 2007 69451 Weinheim, Germany Site-Specific Control of Distances between Gold Nanoparticles using Phosphorothioate Anchors on DNA and a Short Bifunctional Molecular Fastener
