Journal of Experimental Microbiology and Immunology (JEMI) Vol. 6:20-25 Copyright December 2004, M&I UBC
|
|
|
- Deborah McCoy
- 5 years ago
- Views:
Transcription
1 Preparing Plasmid Constructs to Investigate the Characteristics of Thiol Reductase and Flavin Reductase With Regard to Solubilizing Insoluble Proteinase Inhibitor 2 in Bacterial Protein Overexpression Systems NEHA SHAH Department of Microbiology and Immunology, UBC Some proteins produced in bacterial expression systems can be insoluble and aggregated making them difficult to study. One possible solution to solubilizing such insoluble proteins is to create a fusion protein containing the protein of interest coupled with a thioreductase protein. (Yasukawa et al. J. Biol. Chem. 270, (1995)) In this study we intended to assess whether thiol reductase or flavin reductase would both be equally effective at solubilizing the protein of interest; proteinase inhibitor 2. We attempted to create fusion protein constructs using the Novagen plasmid pet32a as a cloning vector since the pet32a contains thiol reductase. To compare reductase proteins, a flavin reductase gene was obtained for cloning. Due to shortage of time for this experiment, the constructs were not produced, however all of the fragments to be cloned and the control vector were produced. Additional time and resources would ensure that the constructs are produced and protein expression and analysis can be carried out to test the solubilizing properties of thiol and flavin reductases on proteinase inhibitor 2. An obstacle when studying protein chemistry using proteins produced by expression systems is the aggregation and inactivation that can result in insoluble proteins. Creating a recombinant fusion protein with thiol reductase (Trx) (6) improves the characterization of the protein of interest. The solubility of the insoluble proteins were compared in native form and in the presence of thiol reductase (6). Many tested proteins that formed had insoluble aggregates had significantly increased solubility when expressed with Trx. Two proteins in particular: Myb and p53 tumour suppressor had significantly increased solubility (6). A commercially available vector, produced by Novagen (1) also utilizes the trx sequence to enhance the production of active protein products when produced in a bacterial system. It is unclear whether the effect arises from the fusion protein properties or the reductase activity. To test our explanations we intended to couple a different reductase protein with an insoluble protein such as proteinase inhibitor 2 (PI2) to determing if there is an increase in the solubility of PI2 as an uncoupled protein. Another reductase protein, flavin reductase (Fre), is hypothesized to have similar solublizing properties as the Trx with the PI2 protein. Designing a construct with a fusion protein of PI2 and Trx and comparing the protein solubility to another construct with PI2 coupled to Fre will determine whether both are effected and determine which reductase is more effective at producing an active and soluble protein product. Due to cloning obstacles and time limitations, the constructs containing PI2 were not obtained, however more research will surely allow the solubility of proposed constructs to be studied. MATERIALS AND METHODS The constructs were based on the Novagen plasmid pet32a (1) which contains trx and is available as a common cloning vector. The cloning strategy involved creating three separate constructs with the PI2 gene to compare solubility in the presence of no reductase, trx, and fre. The trx was to be removed from the pet32a to create a control plasmid into which PI2 was inserted. The two experimental plasmids were to be produced by cloning in the PI2 gene into the pet32a with trx and also into a pet32a plasmid with the trx removed and replaced with the fre gene. This would allow us to accurately compare the PI2 gene as a fusion with either one of the two reductases, or none in the case of the control construct. The PI2 gene was obtained in a plasmid known as pe32. The PI2 gene was cloned out of the plasmid using PCR amplification and forward and reverse primers were designed with engineered restriction sites to allow cloning later into the pet32a plasmid (Fig. 1). The fre product was obtained by PCR amplification out of pes1 plasmid using forward and reverse primers (Fig. 2). The primers were designed to change one stop codon to a glycine as well as change an EcoR1 restriction site to an Nde1 site in order to allow the insertion of the fre into the vector once trx was removed from the pet32a using flanking Nde1 restriction sites. After producing the three constructs containing PI2 and either no reductase, trx, or fre, the plasmids were to be transformed into an E. coli strain such as BL21 to express the proteins and allow characterization of solubility of PI2 each of the different constructs. PI2 expressed as a fusion with the reductase proteins are expected to be of greater solubility than the PI2 protein alone. Preparation of Competent DH5α E. coli cells. A 5 ml culture of DH5α E. coli cells was grown in Luria Bertani (LB) (4) broth at 37 C in a shaking incubator overnight. This starter culture was added to 500 ml of LB broth in the morning and grown for several hours at 37 C until the culture was harvested after approximately 3 hours. The cells harvested and then washed with 25 ml of 100mM MgCl 2 twice followed by one wash with 25 ml of 100 mm CaCl 2. The culture was then resuspended in 20 ml of 100 mm CaCl 2 containing 10% glycerol. Aliquots of 200 µl were stored at - 80 C. 20
2 FIG. 1 Primer design to clone the PI2 gene from pe32 plasmid. Primers were designed using GenBank sequence ID for PI2 and the restriction sites of Nco1 and BamH1 were chosen due to the unique presence in the pet32a plasmid. The primers contain restriction enzyme sites to allow subsequent cloning into the pet32a with directional specificity by using enzymes cutting sticky unique ends. A PCR product of 597 bp is expected using these primers and template DNA. FIG. 2 Primer design to amplify flavin oxidoreductase (fre) gene out of the plasmid pes1. The yellow highlighted region indicates the primer-binding site of the start of the fre gene. The reverse primer contains mutations to replace the stop codon with a glycine to create a fusion protein with PI2, and introduce an Nde1 restriction enzyme site. The PCR product is expected to be 788 bp. 21
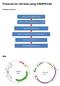
3 FIG. 3 Novagen pet32a vector (1). The Nco1 and BamH1 restriction sites are present in the multiple cloning site of the plasmid where the PI2 will be inserted. The vector contains the trxa sequence flanked by two Nde1 restriction sites. These Nde1 restriction sites will allow trx to be removed and fre inserted substituted to generate the constructs containing different reductase sequences with PI2 as a fusion protein. Amplification of the PI2 gene from the pe32 plasmid. As no plasmid map was available for the plasmid containing PI2, PCR amplification was done using GenBank sequence for PI2 as outlined in Fig. 1. Forward and reverse primers were designed to anneal to the PI2 sequence, and have overhangs containing unique restriction sites to allow for insertion into the vector later. The forward primer contained a restriction site for Nco1 and the reverse for BamH1. These restriction sites were so chosen because there was only one copy of each sequence present in the pet32a vector (Fig. 3). These sites allow the plasmid to be cut open within the multiple cloning sites so the PI2 fragment can be inserted in the correct orientation within the plasmid. Primers were ordered from the Nucleic Acid and Protein Sequencing facility at the University of British Columbia (NAPS) and rehydrated at a concentration of 20µM each. PCR was performed on uncut pe32 plasmid using Platinum Pfx DNA polymerase from Invitrogen Life Science (2). In all cases the amplification reactions were performed in total volumes of 30 ul containing 1 ul of extracted DNA, and final concentrations of: 1X PCR buffer, 1.5 mm MgCl 2, 0.2 mm each of four deoxynucleotide triphosphates, 0.5 um of each primer, and 1 unit Pfx polymerase. PCR conditions were based on standard PCR protocols. PCR cycles began with an initial denaturation at 94 C for 3 min followed by 40 cycles of denature at 94 C for 30 s, anneal at a gradient from 45 C to 65 C for 1 min, and extend at 72 C for 90 s followed by a final anneal at 72 C for 10 min. The gradient was used to attempt to obtain optimal PCR products from the reactions. Amplification of the fre gene from pes1 plasmid. Forward and reverse primers designed to flank the fre gene in the pes1 plasmid. Primer design and annealing sites are seen in Fig. 2. The forward primer anneals to start of the fre sequence and the reverse primer contains mutations to create a suitable sequence for cloning and fusion protein production. The reverse primer mutates a stop codon into a glycine to allow a fusion protein to be produced later with PI2. An Nde1 site was introduced in order to clone the fre fragment into the pet32a plasmid once the trx fragment was cut out using Nde1 sites. The fre gene will then insert where the trx was removed. Primers were ordered from NAPS and rehydrated to 20µM concentrations each. Each amplification reaction had final concentrations exactly as those in the PI2 reactions, except that pes1 plasmid was used as the template instead of pe32 plasmid. PCR began with an initial denaturation at 94 C for 3 min followed by 40 22
4 cycles of denature at 94 C for 30 s, anneal at 54 C for 1 min, and extend at 72 C for 90 s followed by a final anneal at 72 C for 10 min. Cloning of fre gene into pet32a trx. pet32a trx vector was provided by Aileen Kartono. The PCR amplified fre fragment from pes1 plasmid was prepared for cloning by purification using Qiaquick PCR Purification Kit (5) to remove nucleotides and other reaction components. The fre fragment was digested using Nde1 restriction enzyme to generate a sticky ended fragment. The reaction was heat inactivated and then used to set up ligations into pet32a trx vector already linearized by the removal of trx. Several ligation reactions were set up as indicated in Table 1 (2). Many different conditions were tested to better the chances of obtaining a ligated product. Once allowed to ligate, each ligation reaction was transformed into chemically competent DH5α cells. degradation of some of the DNA products in the PCR reaction as well as overloading of the agarose gel lanes. The expected size of the gene product is 597bp and this size fragment is seen in lanes 8 through 12. These lanes correspond to the annealing gradient temperatures of 56.5 C, 59.3 C, 61.8 C, 63.8 C, and 65.0 C. These higher annealing temperatures are the preferred temperatures for amplifying the PI2 gene and it appears that lanes 10 and 11 have the brightest bands of the five positive lanes for the PI2 fragment. TABLE 1. Conditions for the ligation reactions of fre into the pet32a trx. Transformation of DH5α cells with ligation reaction. 2 µl of each ligation reaction in Table 1 was added to 100µl of chemically competent cells thawed on ice. The mixture was allowed to equilibrate on ice for 10 minutes and then incubated at 42 C for 90 seconds. After another 10 minutes on ice, 1 ml of SOC (4) broth was added. The cultures were incubated at 37 C in a shaking water bath for 3 hours. 200 µl of each of the cultures was then plated on an LB agar plate (4) containing 50µg/mL ampicillin to select for bacteria containing the plasmid of interest. The plates were incubated overnight at 37 C. Insertion of PI2 gene PCR fragment into pet32a and pet32a trx vector. PI2 PCR product purified from agarose gel after analyzing by gel using Qiagen Gel Extraction Kit (5). Used 600 ul of buffer QG for solubilizing agarose and elution in 2 x 20 ul buffer EB. Double digested PI2 purified fragment, pet32a vector, and pet32a Trx vector each with both restriction enzymes BamH1 and Nco1. After digestion, reactions were heat inactivated and ligations set up as in Table 2 and incubated at 4 C overnight (2). The reactions were then transformed using chemically competent DH5α cells and plated on LB agar plates (4) containing 50 µg/ml ampicillin. RESULTS Amplification of PI2 gene from pe32 plasmid. The agarose gel of the products of the gradient PCR to amplify PI2 showed many bands and background noise. There were many bands amplified as seen in Fig. 4 and there is some smearing seen in the lanes, especially in lane 2. This smearing could be the result of FIG. 4 Gel image of PI2 amplification from pe32 in gradient PCR. A 1% agarose gel image of the gradient PCR. Lane 1 contains 1ug of 1kB plus ladder (2). Lanes 2 to 12 contain PCR reaction samples ranging in gradient temperature from 45 C to 65 C. The 597 bp fragment from lanes 8 to 12 were excised from the gel for gel purification to isolate the correct fragment. Lanes 8 to 12 correspond to temperatures of 56.5 C, 59.3 C, 61.8 C, 63.8 C, and 65.0 C. Amplification of fre gene from pes1 plasmid. The amplification of the fre gene was successful and a PCR product of 788 bp. The agarose gel image of the PCR is not shown here but was identified as the correct length visually. The fre fragment was purified from the PCR reaction mixture using Qiagen PCR Purification Kit (5) before being used in any further cloning. Cloning of fre gene into pet32a trx vector. After incubation, no colonies were seen on any of the plates. The various ligation reaction conditions were all unsuccessful in producing a vector containing fre in place of the trx. A control of plasmid pes1 also containing the same antibiotic resistance was used as a transformation control, which resulted in the growth of numerous colonies. The growth of the control indicates that the competent cells were viable but the ligations were not successful. Insertion of PI2 PCR fragment into pet32a and pet32a trx vector. After incubation, no colonies were seen for either ligation reaction into the vectors. A control transformation was also prepared, which 23
5 TABLE 2. Conditions for the ligation reaction of pet32a trx and pet32a with PI2. again had numerous colonies on the agar. The ligation reactions were unsuccessful at producing a colony containing the PI2 gene. DISCUSSION Amplification of fre gene from pes1 plasmid. We obtained a fragment corresponding to the correct size of 788 bp which is the expected product of fre. The amplification of the product indicates that the primers and the PCR conditions were optimal for this gene and the yield was also high when analyzed on agarose gel. The fact that the PCR was successful indicates that the engineered mutations needed to clone the fre into the pet32a trx vector were also successful. The primers were designed to ensure that when fre was inserted into the vector, then the protein produced would be a complete fusion protein of the protein of the PI2 and fre. While analysis by agarose gel to assess the size of the fragment generated was sufficient identification for our purposes, a more reliable and accurate measure of the sequence of the product would be to sequence it to ensure that all the nucleotides are correct if the construct behaved improperly during protein expression experiments. With a correct sequence during the cloning stages, protein expression at later stages can be optimized when the nucleotide sequence codes for the correct proteins. Checking the sequence of the completed construct when it is obtained would assure that the fusion protein produced is indeed what is expected. Cloning of fre gene into pet32a Trx vector. We planned on cloning the Fre gene into the vector using the same restriction sites that remove the trx from the pet32a. These restriction sites for Nde1 were engineered onto fre for the purpose of cloning it into the pet32a trx vector. As the fre would insert directly in frame with the removal of the trx, a complete sequence was expected with no errors of frameshift or lack of protein production at a later stage. After many attempts of cloning the fre into pet32a trx there was no success at obtaining a construct. A variety of conditions were utilized as well as using fresh ligase enzyme and buffer, all to no success. This stage of cloning would need to be re-examined for possible problems inhibiting the ligation reactions. Such problems could be the Nde1 enzyme cutting the DNA but also creating an inhibition effect and blocking the ligation sites. Heat treatment of the DNA to be ligated was attempted prior to doing the ligations in order to release any protein that may be associating with the DNA, however this did not help to obtain a successful construct. The DNA fragments of fre and pet32a trx were heated to 95 C and very slowly cooled to 25 C. This step should have denatured any proteins present but still allowed the DNA to reanneal. As the Nde1 sites were already engineered and are a convenient cloning site, another way to obtain the same cut fragments for ligation would be to use another restriction enzyme that cuts at the same site as Nde1 but has different properties that may not inhibit subsequent ligations. Such an enzyme that cuts at the same site is an isoschizomer of Nde1 and an example of such a commercially available enzyme is BfuC1. This is a possible alternate method to obtaining the correct constructs. Insertion of PI2 PCR fragment into pet32a and pet32a Trx vector. As time was a factor in carrying out this experiment, only one attempt was made to clone the PI2 gene sequence into the two available vectors pet32a and pet32a trx. The digestion of the vectors and the PI2 sequence in preparation for cloning likely was successful at creating ends that would anneal given the appropriate conditions. The ligation may have been more successful at a higher temperature for a shorter period of time, for example 15 C for a few hours instead of the longer incubation that was attempted. It is predicted that the ligation would be successful on subsequent attempts because the restriction sites are sticky ends and since each end of 24
6 the gene has a different site, the fragment will insert in the correct direction to allow for protein expression later. The PI2 fragment primers were designed to keep the protein in frame with the vector to allow successful production of a fusion protein with either trx or fre. FURTHER RESEARCH The pet32a Trx vector construct containing fre must be obtained, and with further experimentation the construct can be produced successfully. The PI2 gene must be successfully inserted into all three vectors: pet32a, pet32a trx, and pet32a trx+fre. Any resulting colonies from these ligation reactions and transformation would need to be screened to check for the correct insert. If required, the plasmids could be sequenced to verify the identity prior to further experimentation. Once the positive constructs are available, the plasmid can be transformed into the strain of E. coli BL21 for protein expression analysis. Appropriate protein isolation techniques to isolate the fusion proteins would need to be designed and carried out to analyse the amount of soluble protein present compared to the amounts of aggregated insoluble proteins. ACKNOWLEDGEMENTS I would like to thank Dr. Bill Ramey and Jennifer Sibley for their expertise and knowledge to execute this project. I would also like to thank fellow classmates May Kazem and Aileen Kartono for their contributions to the project. The plasmids were kindly provided by Tai Man Louie. REFERENCES 1. EMD Biosciences, Novagen pet32a vector. 2. Invitrogen Corporation Pfx polymerase, Ligase protocols, 1Kb plus ladder specifications. 3. Louie, T.M., H. Yang, P. Karnchanaphanurach, X.S. Xie,, and L. Xun FAD is a preferred substrate and an inhibitor of Escherichia coli general NAD(P)H:flavin oxidoreductase. J Biol Chem. 277: Sambrook, J., E.F. Fritsch, and T. Maniatis Molecular Cloning: a laboratory manual, 3 rd ed. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, N.Y. 5. Qiagen Inc Qiaquick PCR Purification Kit, Qiaquick Gel Extraction Kit. 6. Yasukawa, T., C. Kanei-Ishii, T. Maekawa, J. Fujimoto, T. Yamamoto, and S. Ishii J Biol Chem. 270:
Journal of Experimental Microbiology and Immunology (JEMI) Vol. 6:26-34 Copyright December 2004, M&I UBC
 Cloning EDTA monooxygenase as a Model Protein to Characterize the Effects of Flavin oxidoreductase on Solubility of Proteins in Protein Overexpression Systems MIKAMEH KAZEM Department of Microbiology and
Cloning EDTA monooxygenase as a Model Protein to Characterize the Effects of Flavin oxidoreductase on Solubility of Proteins in Protein Overexpression Systems MIKAMEH KAZEM Department of Microbiology and
Attempts to Amplify EDTA Monooxygenase A for Cloning into the pet-32a Expression Vector
 Attempts to Amplify EDTA Monooxygenase A for Cloning into the pet-32a Expression Vector STEPHANIE YOUNG Department of Microbiology and Immunology, UBC One significant problem with bacterial overexpression
Attempts to Amplify EDTA Monooxygenase A for Cloning into the pet-32a Expression Vector STEPHANIE YOUNG Department of Microbiology and Immunology, UBC One significant problem with bacterial overexpression
Polymerase Chain Reaction
 Polymerase Chain Reaction Amplify your insert or verify its presence 3H Taq platinum PCR mix primers Ultrapure Water PCR tubes PCR machine A. Insert amplification For insert amplification, use the Taq
Polymerase Chain Reaction Amplify your insert or verify its presence 3H Taq platinum PCR mix primers Ultrapure Water PCR tubes PCR machine A. Insert amplification For insert amplification, use the Taq
Cold Fusion Cloning Kit. Cat. #s MC100A-1, MC101A-1. User Manual
 Fusion Cloning technology Cold Fusion Cloning Kit Store the master mixture and positive controls at -20 C Store the competent cells at -80 C. (ver. 120909) A limited-use label license covers this product.
Fusion Cloning technology Cold Fusion Cloning Kit Store the master mixture and positive controls at -20 C Store the competent cells at -80 C. (ver. 120909) A limited-use label license covers this product.
HE Swift Cloning Kit
 HE Swift Cloning Kit For high-efficient cloning of PCR products either blunt or sticky-end Kit Contents Contents VTT-BB05 phe Vector (35 ng/µl) 20 µl T4 DNA Ligase (3 U/µl) 20 µl 2 Reaction Buffer 100
HE Swift Cloning Kit For high-efficient cloning of PCR products either blunt or sticky-end Kit Contents Contents VTT-BB05 phe Vector (35 ng/µl) 20 µl T4 DNA Ligase (3 U/µl) 20 µl 2 Reaction Buffer 100
pgm-t Cloning Kit Cat. # : GVT202 Size : 20 Reactions Store at -20
 pgm-t Cloning Kit Cat. # : GVT202 Size : 20 Reactions Store at -20 1 Kit Contents Contents pgm-t Cloning Kit pgm-t Vector (50 ng/μl) 20 μl T4 DNA Ligase (3 U/μl) 20 μl 10X T4 DNA Ligation Buffer 30 μl
pgm-t Cloning Kit Cat. # : GVT202 Size : 20 Reactions Store at -20 1 Kit Contents Contents pgm-t Cloning Kit pgm-t Vector (50 ng/μl) 20 μl T4 DNA Ligase (3 U/μl) 20 μl 10X T4 DNA Ligation Buffer 30 μl
Construction of a Mutant pbr322 Using Site-Directed Mutagenesis to Investigate the Exclusion Effects of pbr322 During Co-transformation with puc19
 Construction of a Mutant pbr322 Using Site-Directed Mutagenesis to Investigate the Exclusion Effects of pbr322 During Co-transformation with puc19 IVANA KOMLJENOVIC Department of Microbiology and Immunology,
Construction of a Mutant pbr322 Using Site-Directed Mutagenesis to Investigate the Exclusion Effects of pbr322 During Co-transformation with puc19 IVANA KOMLJENOVIC Department of Microbiology and Immunology,
Journal of Experimental Microbiology and Immunology (JEMI) Vol. 16: Copyright April 2012, M&I UBC
 Production of a Recombinant Vector to Enable the Study of Thioredoxin Function as a Bound or Detached Solubilizer of Proteinase Inhibitor 2 in a Bacterial Protein Overexpression System Chris Duronio Department
Production of a Recombinant Vector to Enable the Study of Thioredoxin Function as a Bound or Detached Solubilizer of Proteinase Inhibitor 2 in a Bacterial Protein Overexpression System Chris Duronio Department
Cat # Box1 Box2. DH5a Competent E. coli cells CCK-20 (20 rxns) 40 µl 40 µl 50 µl x 20 tubes. Choo-Choo Cloning TM Enzyme Mix
 Molecular Cloning Laboratories User Manual Version 3.3 Product name: Choo-Choo Cloning Kits Cat #: CCK-10, CCK-20, CCK-096, CCK-384 Description: Choo-Choo Cloning is a highly efficient directional PCR
Molecular Cloning Laboratories User Manual Version 3.3 Product name: Choo-Choo Cloning Kits Cat #: CCK-10, CCK-20, CCK-096, CCK-384 Description: Choo-Choo Cloning is a highly efficient directional PCR
Gateway Cloning Protocol (Clough Lab Edition) This document is a modification of the Gateway cloning protocol developed by Manju in Chris Taylor's lab
 Gateway Cloning Protocol (Clough Lab Edition) This document is a modification of the Gateway cloning protocol developed by Manju in Chris Taylor's lab With the Gateway cloning system, a PCR fragment is
Gateway Cloning Protocol (Clough Lab Edition) This document is a modification of the Gateway cloning protocol developed by Manju in Chris Taylor's lab With the Gateway cloning system, a PCR fragment is
Data Sheet Quick PCR Cloning Kit
 Data Sheet Quick PCR Cloning Kit 6044 Cornerstone Ct. West, Ste. E DESCRIPTION: The Quick PCR Cloning Kit is a simple and highly efficient method to insert any gene or DNA fragment into a vector, without
Data Sheet Quick PCR Cloning Kit 6044 Cornerstone Ct. West, Ste. E DESCRIPTION: The Quick PCR Cloning Kit is a simple and highly efficient method to insert any gene or DNA fragment into a vector, without
Hetero-Stagger PCR Cloning Kit
 Product Name: Code No: Size: DynaExpress Hetero-Stagger PCR Cloning Kit DS150 20 reactions Kit Components: Box 1 (-20 ) phst-1 Vector, linearized Annealing Buffer Ligase Mixture phst Forward Sequence Primer
Product Name: Code No: Size: DynaExpress Hetero-Stagger PCR Cloning Kit DS150 20 reactions Kit Components: Box 1 (-20 ) phst-1 Vector, linearized Annealing Buffer Ligase Mixture phst Forward Sequence Primer
pgm-t Cloning Kit For direct cloning of PCR products generated by Taq DNA polymerases For research use only Cat. # : GVT202 Size : 20 Reactions
 pgm-t Cloning Kit For direct cloning of PCR products generated by Taq DNA polymerases Cat. # : GVT202 Size : 20 Reactions Store at -20 For research use only 1 pgm-t Cloning Kit Cat. No.: GVT202 Kit Contents
pgm-t Cloning Kit For direct cloning of PCR products generated by Taq DNA polymerases Cat. # : GVT202 Size : 20 Reactions Store at -20 For research use only 1 pgm-t Cloning Kit Cat. No.: GVT202 Kit Contents
Creating pentr vectors by BP reaction
 Creating pentr vectors by BP reaction Tanya Lepikhova and Rafael Martinez 15072011 Overview: 1. Design primers to add the attb sites to gene of interest 2. Perform PCR with a high fidelity DNA polymerase
Creating pentr vectors by BP reaction Tanya Lepikhova and Rafael Martinez 15072011 Overview: 1. Design primers to add the attb sites to gene of interest 2. Perform PCR with a high fidelity DNA polymerase
Determination of Exclusion Effect in Wild Type and Rop Deficient Mutated pbr322 Co-transformations
 Determination of Exclusion Effect in Wild Type and Rop Deficient Mutated pbr322 Co-transformations Suzana Sabaiduc and Andy Lo Microbiology and Immunology University of British Columbia pbr322 is an expression
Determination of Exclusion Effect in Wild Type and Rop Deficient Mutated pbr322 Co-transformations Suzana Sabaiduc and Andy Lo Microbiology and Immunology University of British Columbia pbr322 is an expression
YG1 Control. YG1 PstI XbaI. YG5 PstI XbaI Water Buffer DNA Enzyme1 Enzyme2
 9/9/03 Aim: Digestion and gel extraction of YG, YG3, YG5 and 8/C. Strain: E. coli DH5α Plasmid: Bba_J600, psbc3 4,, 3 6 7, 8 9 0, 3, 4 8/C 8/C SpeI PstI 3 3 x3 YG YG PstI XbaI.5 3.5 x YG3 YG3 PstI XbaI
9/9/03 Aim: Digestion and gel extraction of YG, YG3, YG5 and 8/C. Strain: E. coli DH5α Plasmid: Bba_J600, psbc3 4,, 3 6 7, 8 9 0, 3, 4 8/C 8/C SpeI PstI 3 3 x3 YG YG PstI XbaI.5 3.5 x YG3 YG3 PstI XbaI
Attempted Construction of a pet32a(+) Vector Containing EDTA Monooxygenase A
 Attempted Construction of a pet32a(+) Vector Containing EDTA Monooxygenase A Andrew Cameron, Stuart Gray, Anastasia Sribnaia and Tracy Withrow Department of Microbiology & Immunology, UBC Protein overexpression
Attempted Construction of a pet32a(+) Vector Containing EDTA Monooxygenase A Andrew Cameron, Stuart Gray, Anastasia Sribnaia and Tracy Withrow Department of Microbiology & Immunology, UBC Protein overexpression
peco TM -T7-nGST, Eco cloning Kit User Manual (Patent pending)
 peco TM -T7-nGST, Eco cloning Kit User Manual (Patent pending) Cloning PCR products for E Coli expression of N-term GST-tagged protein Cat# Contents Amounts Application IC-1004 peco-t7-ngst vector built-in
peco TM -T7-nGST, Eco cloning Kit User Manual (Patent pending) Cloning PCR products for E Coli expression of N-term GST-tagged protein Cat# Contents Amounts Application IC-1004 peco-t7-ngst vector built-in
GenBuilder TM Plus Cloning Kit User Manual
 GenBuilder TM Plus Cloning Kit User Manual Cat. No. L00744 Version 11242017 Ⅰ. Introduction... 2 I.1 Product Information... 2 I.2 Kit Contents and Storage... 2 I.3 GenBuilder Cloning Kit Workflow... 2
GenBuilder TM Plus Cloning Kit User Manual Cat. No. L00744 Version 11242017 Ⅰ. Introduction... 2 I.1 Product Information... 2 I.2 Kit Contents and Storage... 2 I.3 GenBuilder Cloning Kit Workflow... 2
GenBuilder TM Plus Cloning Kit User Manual
 GenBuilder TM Plus Cloning Kit User Manual Cat.no L00744 Version 11242017 Ⅰ. Introduction... 2 I.1 Product Information... 2 I.2 Kit Contents and Storage... 2 I.3 GenBuilder Cloning Kit Workflow... 2 Ⅱ.
GenBuilder TM Plus Cloning Kit User Manual Cat.no L00744 Version 11242017 Ⅰ. Introduction... 2 I.1 Product Information... 2 I.2 Kit Contents and Storage... 2 I.3 GenBuilder Cloning Kit Workflow... 2 Ⅱ.
Construction of Recombinant Expression Vectors to Study the Effect of Thioredoxin on Heterologous Protein Solubility
 Construction of Recombinant Expression Vectors to Study the Effect of Thioredoxin on Heterologous Protein Solubility Liz Geum, Reed Huber, Nicole Leung, Matthew Lowe Department of Microbiology and Immunology,
Construction of Recombinant Expression Vectors to Study the Effect of Thioredoxin on Heterologous Protein Solubility Liz Geum, Reed Huber, Nicole Leung, Matthew Lowe Department of Microbiology and Immunology,
Ready_to_use Fast Seamless Cloning Kit. User Manual
 For general laboratory use. Not for use in diagnostic procedures. FOR IN VITRO USE ONLY. Ready_to_use Fast Seamless Cloning Kit User Manual 1 / 6 Tel: 021-58975266 Fax: 021-50800270 Email:tech@dogene.com
For general laboratory use. Not for use in diagnostic procedures. FOR IN VITRO USE ONLY. Ready_to_use Fast Seamless Cloning Kit User Manual 1 / 6 Tel: 021-58975266 Fax: 021-50800270 Email:tech@dogene.com
Lab Book igem Stockholm Lysostaphin. Week 9
 Lysostaphin Week 9 Summarized below are the experiments conducted this week in chronological order. Click on the experiment name to view it. To go back to this summary, click Summary in the footer. Summary
Lysostaphin Week 9 Summarized below are the experiments conducted this week in chronological order. Click on the experiment name to view it. To go back to this summary, click Summary in the footer. Summary
Antisense RNA Targeting the First Periplasmic Domain of YidC in Escherichia coli Appears to Induce Filamentation but Does Not Affect Cell Viability
 Antisense RNA Targeting the First Periplasmic Domain of YidC in Escherichia coli Appears to Induce Filamentation but Does Not Affect Cell Viability Riaaz Lalani, Nathaniel Susilo, Elisa Xiao, Andrea Xu
Antisense RNA Targeting the First Periplasmic Domain of YidC in Escherichia coli Appears to Induce Filamentation but Does Not Affect Cell Viability Riaaz Lalani, Nathaniel Susilo, Elisa Xiao, Andrea Xu
GenBuilder TM Cloning Kit User Manual
 GenBuilder TM Cloning Kit User Manual Cat.no L00701 Version 11242017 Ⅰ. Introduction... 2 I.1 Product Information... 2 I.2 Kit Contents and Storage... 2 I.3 GenBuilder Cloning Kit Workflow... 2 Ⅱ. DNA
GenBuilder TM Cloning Kit User Manual Cat.no L00701 Version 11242017 Ⅰ. Introduction... 2 I.1 Product Information... 2 I.2 Kit Contents and Storage... 2 I.3 GenBuilder Cloning Kit Workflow... 2 Ⅱ. DNA
Ligation Independent Cloning (LIC) Procedure
 Ligation Independent Cloning (LIC) Procedure Ligation Independent Cloning (LIC) LIC cloning allows insertion of DNA fragments without using restriction enzymes into specific vectors containing engineered
Ligation Independent Cloning (LIC) Procedure Ligation Independent Cloning (LIC) LIC cloning allows insertion of DNA fragments without using restriction enzymes into specific vectors containing engineered
Title: Understanding the impact of orientation on gene expression of lux operon in pkn800 transformation into Escherichia coli DH5α
 Seim - 1 Name: Darian Seim Title: Understanding the impact of orientation on gene expression of lux operon in pkn800 transformation into Escherichia coli DH5α Date: April 12 th, 2016 April 18 th, 2016
Seim - 1 Name: Darian Seim Title: Understanding the impact of orientation on gene expression of lux operon in pkn800 transformation into Escherichia coli DH5α Date: April 12 th, 2016 April 18 th, 2016
Subcloning 1. Overview 2. Method
 Subcloning 1. Overview: 1.1. Digest plasmids for vector + insert 1.2. CIP treat vector 1.3. Gel purify vector + insert 1.4. Ligate 1.5. Transform bacteria with ligation reactions 1.6. Qiagen or Eppendorf
Subcloning 1. Overview: 1.1. Digest plasmids for vector + insert 1.2. CIP treat vector 1.3. Gel purify vector + insert 1.4. Ligate 1.5. Transform bacteria with ligation reactions 1.6. Qiagen or Eppendorf
VDL101.3 CLONING TRANSGENE INTO pad5f35
 Purpose 1.1. The purpose of this protocol is to transfer a transgene from the pshuttlex plasmid to pad5/f35. 1.2. The starting material is 10 μg plasmid DNA. 1.3. This procedure is routinely performed
Purpose 1.1. The purpose of this protocol is to transfer a transgene from the pshuttlex plasmid to pad5/f35. 1.2. The starting material is 10 μg plasmid DNA. 1.3. This procedure is routinely performed
Lab Book igem Stockholm Lysostaphin. Week 8
 Lysostaphin Week 8 Summarized below are the experiments conducted this week in chronological order. Click on the experiment name to view it. To go back to this summary, click Summary in the footer. Summary
Lysostaphin Week 8 Summarized below are the experiments conducted this week in chronological order. Click on the experiment name to view it. To go back to this summary, click Summary in the footer. Summary
California Institute of Technology. Directed evolution. Dr. F.H. Arnold s lab
 Directed evolution. Dr. F.H. Arnold s lab May 4, 1999 Mutagenic PCR -[Mn] The amount of Mn used in the reaction should be titrated to produce the desired mutagenic rate. Libraries that have close to 30%
Directed evolution. Dr. F.H. Arnold s lab May 4, 1999 Mutagenic PCR -[Mn] The amount of Mn used in the reaction should be titrated to produce the desired mutagenic rate. Libraries that have close to 30%
In order to make our construct pah12 (Mlra) and pah05 (venus) biobrick compatitable PCR was conducted. Following PCR mixes were prepared
 BioBrick cloning Project: BioBricks Authors: Antti Koistinen Dates: 2016-09-15 to 2016-10-06 THURSDAY, 9/15 In order to make our construct pah12 (Mlra) and pah05 (venus) biobrick compatitable PCR was conducted.
BioBrick cloning Project: BioBricks Authors: Antti Koistinen Dates: 2016-09-15 to 2016-10-06 THURSDAY, 9/15 In order to make our construct pah12 (Mlra) and pah05 (venus) biobrick compatitable PCR was conducted.
Conversion of plasmids into Gateway compatible cloning
 Conversion of plasmids into Gateway compatible cloning Rafael Martinez 14072011 Overview: 1. Select the right Gateway cassette (A, B or C). 2. Design primers to amplify the right Gateway cassette from
Conversion of plasmids into Gateway compatible cloning Rafael Martinez 14072011 Overview: 1. Select the right Gateway cassette (A, B or C). 2. Design primers to amplify the right Gateway cassette from
Gene Jockeying: Tricks of the Trade so that your Ligations work Every Time! Bevin Engelward
 Gene Jockeying: Tricks of the Trade so that your Ligations work Every Time! Bevin Engelward We will first review a simple example of a ligation. Know Your Vectors! -Find out as much as you can about your
Gene Jockeying: Tricks of the Trade so that your Ligations work Every Time! Bevin Engelward We will first review a simple example of a ligation. Know Your Vectors! -Find out as much as you can about your
Figure 1. Map of cloning vector pgem T-Easy (bacterial plasmid DNA)
 Texas A&M University-Corpus Christi CHEM4402 Biochemistry II Laboratory Laboratory 6: Ligation & Bacterial Transformation (Bring your text and laptop to class if you wish to work on your assignment during
Texas A&M University-Corpus Christi CHEM4402 Biochemistry II Laboratory Laboratory 6: Ligation & Bacterial Transformation (Bring your text and laptop to class if you wish to work on your assignment during
Fast and efficient site-directed mutagenesis with Platinum SuperFi DNA Polymerase
 APPLICATION NOTE Platinum Superi Polymerase ast and efficient site-directed mutagenesis with Platinum Superi Polymerase Introduction Site-directed mutagenesis is one of the most essential techniques to
APPLICATION NOTE Platinum Superi Polymerase ast and efficient site-directed mutagenesis with Platinum Superi Polymerase Introduction Site-directed mutagenesis is one of the most essential techniques to
Diagnosis Sanger. Interpreting and Troubleshooting Chromatograms. Volume 1: Help! No Data! GENEWIZ Technical Support
 Diagnosis Sanger Interpreting and Troubleshooting Chromatograms GENEWIZ Technical Support DNAseq@genewiz.com Troubleshooting This troubleshooting guide is based on common issues seen from samples within
Diagnosis Sanger Interpreting and Troubleshooting Chromatograms GENEWIZ Technical Support DNAseq@genewiz.com Troubleshooting This troubleshooting guide is based on common issues seen from samples within
Designing and creating your gene knockout Background The rada gene was identified as a gene, that when mutated, caused cells to become hypersensitive
 Designing and creating your gene knockout Background The rada gene was identified as a gene, that when mutated, caused cells to become hypersensitive to ionizing radiation. However, why these mutants are
Designing and creating your gene knockout Background The rada gene was identified as a gene, that when mutated, caused cells to become hypersensitive to ionizing radiation. However, why these mutants are
Enhanced Arginase production: rocf
 Enhanced Arginase production: rocf Purpose and Justification: Bacillus subtilis produces urease, which catalyses the hydrolysis of urea into ammonium and carbonate. Since the cell wall of the bacteria
Enhanced Arginase production: rocf Purpose and Justification: Bacillus subtilis produces urease, which catalyses the hydrolysis of urea into ammonium and carbonate. Since the cell wall of the bacteria
Development of Positive Control for Hepatitis B Virus
 Human Journals Research Article December 2015 Vol.:2, Issue:2 All rights are reserved by Saurabh Bandhavkar et al. Development of Positive Control for Hepatitis B Virus Keywords: Hepatitis B virus, pbluescript,
Human Journals Research Article December 2015 Vol.:2, Issue:2 All rights are reserved by Saurabh Bandhavkar et al. Development of Positive Control for Hepatitis B Virus Keywords: Hepatitis B virus, pbluescript,
Lecture Four. Molecular Approaches I: Nucleic Acids
 Lecture Four. Molecular Approaches I: Nucleic Acids I. Recombinant DNA and Gene Cloning Recombinant DNA is DNA that has been created artificially. DNA from two or more sources is incorporated into a single
Lecture Four. Molecular Approaches I: Nucleic Acids I. Recombinant DNA and Gene Cloning Recombinant DNA is DNA that has been created artificially. DNA from two or more sources is incorporated into a single
10X ligation buffer ligase 1 vector DNA insert DNA H 2 O. 10 µl Total Volume. 10X ligation buffer ligase 1 vector DNA insert DNA
 Biol/Chem 475 S07 Study problems for quiz 1 See also questions posed in lab handouts including ligase handout Answers to questions 1&2 included at the end of this document. 1. You plan to clone a 1.0 kb
Biol/Chem 475 S07 Study problems for quiz 1 See also questions posed in lab handouts including ligase handout Answers to questions 1&2 included at the end of this document. 1. You plan to clone a 1.0 kb
Perform HiFi PCR Purify PCR-mixture using PCR-clean up kit and determine concentration
 IGEM EXPERIMENT 2 CONSTRUCTION OF PSB#X#-T7CCDB Strategy Cloning Flow Chart (using restriction/ligation): 1. In silico design of cloning Design a cloning strategy (in case of insert/backbone): o Using
IGEM EXPERIMENT 2 CONSTRUCTION OF PSB#X#-T7CCDB Strategy Cloning Flow Chart (using restriction/ligation): 1. In silico design of cloning Design a cloning strategy (in case of insert/backbone): o Using
NZYGene Synthesis kit
 Kit components Component Concentration Amount NZYGene Synthesis kit Catalogue number: MB33901, 10 reactions GS DNA Polymerase 1U/ μl 30 μl Reaction Buffer for GS DNA Polymerase 10 150 μl dntp mix 2 mm
Kit components Component Concentration Amount NZYGene Synthesis kit Catalogue number: MB33901, 10 reactions GS DNA Polymerase 1U/ μl 30 μl Reaction Buffer for GS DNA Polymerase 10 150 μl dntp mix 2 mm
Cytochrome c heme lyase can mature a fusion peptide composed of the amino-terminal residues of horse cytochrome c
 Cytochrome c heme lyase can mature a fusion peptide composed of the amino-terminal residues of horse cytochrome c Wesley B. Asher a and Kara L. Bren a * a Department of Chemistry, University of Rochester,
Cytochrome c heme lyase can mature a fusion peptide composed of the amino-terminal residues of horse cytochrome c Wesley B. Asher a and Kara L. Bren a * a Department of Chemistry, University of Rochester,
Attempt to Construct a Rop + puc19 from pbr322 Mutant lacking the Tetracycline Resistance Gene
 Attempt to Construct a Rop + puc19 from pbr322 Mutant lacking the Tetracycline Resistance Gene DELIA WANG Department of Microbiology and Immunology, UBC The two commonly used plasmid vectors, pbr322 and
Attempt to Construct a Rop + puc19 from pbr322 Mutant lacking the Tetracycline Resistance Gene DELIA WANG Department of Microbiology and Immunology, UBC The two commonly used plasmid vectors, pbr322 and
Guide-it Indel Identification Kit User Manual
 Clontech Laboratories, Inc. Guide-it Indel Identification Kit User Manual Cat. No. 631444 (120114) Clontech Laboratories, Inc. A Takara Bio Company 1290 Terra Bella Avenue, Mountain View, CA 94043, USA
Clontech Laboratories, Inc. Guide-it Indel Identification Kit User Manual Cat. No. 631444 (120114) Clontech Laboratories, Inc. A Takara Bio Company 1290 Terra Bella Avenue, Mountain View, CA 94043, USA
MUTAGENESIS OF THE PROMOTER OF THE OIL PALM HOMOGENTISATE GERANYLGERANYL TRANSFERASE GENE (HGGT) BY PCR-DRIVEN OVERLAP EXTENSION
 ICBAA2017-30 MUTAGENESIS OF THE PROMOTER OF THE OIL PALM HOMOGENTISATE GERANYLGERANYL TRANSFERASE GENE (HGGT) BY PCR-DRIVEN OVERLAP EXTENSION Mohd Shahrul Nizwanshah Karim and Siti Nor Akmar Abdullah Laboratory
ICBAA2017-30 MUTAGENESIS OF THE PROMOTER OF THE OIL PALM HOMOGENTISATE GERANYLGERANYL TRANSFERASE GENE (HGGT) BY PCR-DRIVEN OVERLAP EXTENSION Mohd Shahrul Nizwanshah Karim and Siti Nor Akmar Abdullah Laboratory
Rapid amplification of cdna ends (RACE)
 Rapid amplification of cdna ends (RACE) Rapid amplification of cdna ends (RACE) is a technique used in molecular biology to obtain the full length sequence of an RNA transcript found within a cell. RACE
Rapid amplification of cdna ends (RACE) Rapid amplification of cdna ends (RACE) is a technique used in molecular biology to obtain the full length sequence of an RNA transcript found within a cell. RACE
Construction of an enlarged puc19 vector with a rop gene designed to study plasmid maintenance in Escherichia coli
 Construction of an enlarged puc19 vector with a rop gene designed to study plasmid maintenance in Escherichia coli Benson Chang, Arnab Ray, Thomas Tsuei, Rachel Wan Department of Microbiology and Immunology,
Construction of an enlarged puc19 vector with a rop gene designed to study plasmid maintenance in Escherichia coli Benson Chang, Arnab Ray, Thomas Tsuei, Rachel Wan Department of Microbiology and Immunology,
Transformation (The method of CaCl 2 )
 PROTOCOLS E. coli Transformation (The method of CaCl 2 ) Ligation PRODUCTION competent cells of E. COLI. (Rubidium cells). Gel DNA Recovery Kit Electroporation Digestiones Plasmid purification (The method
PROTOCOLS E. coli Transformation (The method of CaCl 2 ) Ligation PRODUCTION competent cells of E. COLI. (Rubidium cells). Gel DNA Recovery Kit Electroporation Digestiones Plasmid purification (The method
TECHNICAL BULLETIN. In Vitro Bacterial Split Fluorescent Protein Fold n Glow Solubility Assay Kits
 In Vitro Bacterial Split Fluorescent Protein Fold n Glow Solubility Assay Kits Catalog Numbers APPA001 In Vitro Bacterial Split GFP "Fold 'n' Glow" Solubility Assay Kit (Green) APPA008 In Vitro Bacterial
In Vitro Bacterial Split Fluorescent Protein Fold n Glow Solubility Assay Kits Catalog Numbers APPA001 In Vitro Bacterial Split GFP "Fold 'n' Glow" Solubility Assay Kit (Green) APPA008 In Vitro Bacterial
Recombinants and Transformation
 Jesse Ruben Partner Roman Verner BMB 442 Recombinants and Transformation Introduction The goal of this experiment was to take two antibiotic resistance genes for ampicillin and kanamycin from plasmids
Jesse Ruben Partner Roman Verner BMB 442 Recombinants and Transformation Introduction The goal of this experiment was to take two antibiotic resistance genes for ampicillin and kanamycin from plasmids
Supplementary Information
 Single day construction of multi-gene circuits with 3G assembly Andrew D. Halleran 1, Anandh Swaminathan 2, and Richard M. Murray 1, 2 1. Bioengineering, California Institute of Technology, Pasadena, CA.
Single day construction of multi-gene circuits with 3G assembly Andrew D. Halleran 1, Anandh Swaminathan 2, and Richard M. Murray 1, 2 1. Bioengineering, California Institute of Technology, Pasadena, CA.
The GeneEditor TM in vitro Mutagenesis System: Site- Directed Mutagenesis Using Altered Beta-Lactamase Specificity
 Promega Notes Magazine Number 62, 1997, p. 02 The GeneEditor TM in vitro Mutagenesis System: Site- Directed Mutagenesis Using Altered Beta-Lactamase Specificity By Christine Andrews and Scott Lesley Promega
Promega Notes Magazine Number 62, 1997, p. 02 The GeneEditor TM in vitro Mutagenesis System: Site- Directed Mutagenesis Using Altered Beta-Lactamase Specificity By Christine Andrews and Scott Lesley Promega
Molecular cloning: 6 RBS+ arac/ laci
 Molecular cloning: 6 RBS+ arac/ laci Resource: 6 RBS: from the parts: B0030, B0032, B0034, J61100, J61107, J61127; renamed as RBS1 RBS2 RBS6; arac: C0080 laci: C0012 July 6 th Plasmid mini prep: 6 RBS;
Molecular cloning: 6 RBS+ arac/ laci Resource: 6 RBS: from the parts: B0030, B0032, B0034, J61100, J61107, J61127; renamed as RBS1 RBS2 RBS6; arac: C0080 laci: C0012 July 6 th Plasmid mini prep: 6 RBS;
FROM EXPERIMENTS IN BACTERIAL GENETICS AND GENE TECHNIQUE
 Uppsala 2001-04-01 REPORT FROM EXPERIMENTS IN BACTERIAL GENETICS AND GENE TECHNIQUE Laboratory assistants: Maria Jönsson Amera Gibreel Students: Contents ASSIGNMENT:... 3 INTRODUCTION:... 3 MATERIAL AND
Uppsala 2001-04-01 REPORT FROM EXPERIMENTS IN BACTERIAL GENETICS AND GENE TECHNIQUE Laboratory assistants: Maria Jönsson Amera Gibreel Students: Contents ASSIGNMENT:... 3 INTRODUCTION:... 3 MATERIAL AND
KOD -Plus- Mutagenesis Kit
 Instruction manual KOD -Plus- Mutagenesis Kit 0811 F0936K KOD -Plus- Mutagenesis Kit SMK-101 20 reactions Store at -20 C Contents [1] Introduction [2] Flow chart [3] Components [4] Notes [5] Protocol 1.
Instruction manual KOD -Plus- Mutagenesis Kit 0811 F0936K KOD -Plus- Mutagenesis Kit SMK-101 20 reactions Store at -20 C Contents [1] Introduction [2] Flow chart [3] Components [4] Notes [5] Protocol 1.
Journal of Experimental Microbiology and Immunology (JEMI) Vol. 20: Copyright April 2016, M&I UBC
 The Major Periplasmic Domain of YidC May Be Required for Polar Localization of a Green Fluorescence Protein Tagged YidC Variant Protein in Escherichia coli Peter Xu, Kevin He, Steven Yan Department of
The Major Periplasmic Domain of YidC May Be Required for Polar Localization of a Green Fluorescence Protein Tagged YidC Variant Protein in Escherichia coli Peter Xu, Kevin He, Steven Yan Department of
Puro. Knockout Detection (KOD) Kit
 Puro Knockout Detection (KOD) Kit Cat. No. CC-03 18 Oct. 2016 Contents I. Kit Contents and Storage II. Product Overview III. Methods Experimental Outline Genomic DNA Preparation Obtain Hybrid DNA Digest
Puro Knockout Detection (KOD) Kit Cat. No. CC-03 18 Oct. 2016 Contents I. Kit Contents and Storage II. Product Overview III. Methods Experimental Outline Genomic DNA Preparation Obtain Hybrid DNA Digest
Generation of Non-typeable Haemophilus influenzae Directed Gene Deletion Mutants Jeroen D. Langereis *
 Generation of Non-typeable Haemophilus influenzae Directed Gene Deletion Mutants Jeroen D. Langereis * Laboratory of Pediatric Infectious Diseases, Department of Pediatrics and Laboratory of Medical Immunology,
Generation of Non-typeable Haemophilus influenzae Directed Gene Deletion Mutants Jeroen D. Langereis * Laboratory of Pediatric Infectious Diseases, Department of Pediatrics and Laboratory of Medical Immunology,
Vector Linearization. igem TU/e 2016 Biomedical Engineering
 igem TU/e 2016 Biomedical Engineering Eindhoven University of Technology Room: Ceres 0.04 Den Dolech 2, 5612 AZ Eindhoven The Netherlands Tel. no. +31 50 247 55 59 2016.igem.org/Team:TU_Eindhoven Vector
igem TU/e 2016 Biomedical Engineering Eindhoven University of Technology Room: Ceres 0.04 Den Dolech 2, 5612 AZ Eindhoven The Netherlands Tel. no. +31 50 247 55 59 2016.igem.org/Team:TU_Eindhoven Vector
igem LMU- Munich Munich Cloning procedure
 igem LM- Munich 2014 - igem@bio.lmu.de - http://2014.igem.org/team:lm- Munich Cloning procedure Overview of a cloning procedure: 1. Digestion of a PCR product ( insert) and a vector 2. Dephosphorylation
igem LM- Munich 2014 - igem@bio.lmu.de - http://2014.igem.org/team:lm- Munich Cloning procedure Overview of a cloning procedure: 1. Digestion of a PCR product ( insert) and a vector 2. Dephosphorylation
PCR Cloning Protocol
 Modifications to EXPERIMENTS 21 and 24: PCR and Molecular Cloning This experiment was designed by Dylan Dodd, based on research completed in Dr. Isaac Cann s lab*, with modifications and editing of content
Modifications to EXPERIMENTS 21 and 24: PCR and Molecular Cloning This experiment was designed by Dylan Dodd, based on research completed in Dr. Isaac Cann s lab*, with modifications and editing of content
Amgen Laboratory Series. Tabs C and E
 Amgen Laboratory Series Tabs C and E Chapter 2A Goals Describe the characteristics of plasmids Explain how plasmids are used in cloning a gene Describe the function of restriction enzymes Explain how to
Amgen Laboratory Series Tabs C and E Chapter 2A Goals Describe the characteristics of plasmids Explain how plasmids are used in cloning a gene Describe the function of restriction enzymes Explain how to
Product Information (1) General Information
 Table of Contents Product Information 2 (1) General Information 2 (2) Features 3 (3) Kit Contents 3 (4) Storage and Expiration 3 Standard Protocol for Preparation of Competent Cells 4 Instant Protocol
Table of Contents Product Information 2 (1) General Information 2 (2) Features 3 (3) Kit Contents 3 (4) Storage and Expiration 3 Standard Protocol for Preparation of Competent Cells 4 Instant Protocol
Supporting Information
 Supporting Information Development of a 2,4-Dinitrotoluene-Responsive Synthetic Riboswitch in E. coli cells Molly E. Davidson, Svetlana V. Harbaugh, Yaroslav G. Chushak, Morley O. Stone, Nancy Kelley-
Supporting Information Development of a 2,4-Dinitrotoluene-Responsive Synthetic Riboswitch in E. coli cells Molly E. Davidson, Svetlana V. Harbaugh, Yaroslav G. Chushak, Morley O. Stone, Nancy Kelley-
PCR Cloning II fusion domains for purification His 6 GST chitin binding protein MBP
 PCR cloning : why secondary amplification? May 26, 2009 secondary amplification amplification of the coding sequence alone or of defined fragments ligation into expression vector insertion of fusion domains
PCR cloning : why secondary amplification? May 26, 2009 secondary amplification amplification of the coding sequence alone or of defined fragments ligation into expression vector insertion of fusion domains
Molecular Techniques Third-year Biology
 PLANNING Genetics Lab practices Molecular Techniques. Genetics Lab practices protocol. 2015-16 PCR-DIRECTED MUTAGENESIS, MOLECULAR CLONING AND RESTRICTION ANALYSIS Sessions 1 & 2 (2x3 hours): PCR-directed
PLANNING Genetics Lab practices Molecular Techniques. Genetics Lab practices protocol. 2015-16 PCR-DIRECTED MUTAGENESIS, MOLECULAR CLONING AND RESTRICTION ANALYSIS Sessions 1 & 2 (2x3 hours): PCR-directed
Application of Molecular Biology tools for cloning of a foreign gene
 IFM/Kemi Linköpings Universitet September 2013/LGM Labmanual Project course Application of Molecular Biology tools for cloning of a foreign gene Table of contents Introduction... 3 Amplification of a gene
IFM/Kemi Linköpings Universitet September 2013/LGM Labmanual Project course Application of Molecular Biology tools for cloning of a foreign gene Table of contents Introduction... 3 Amplification of a gene
Genetic Compensation of E. coli C156 with pbad-ompa for the Investigation of Conjugation Efficiency
 Genetic Compensation of E. coli C156 with pbad-ompa for the Investigation of Conjugation Efficiency Anthony Hsieh and Esten Williams Department of Microbiology & Immunology, University of British Columbia
Genetic Compensation of E. coli C156 with pbad-ompa for the Investigation of Conjugation Efficiency Anthony Hsieh and Esten Williams Department of Microbiology & Immunology, University of British Columbia
Understanding the Cellular Mechanism of the Excess Microsporocytes I (EMSI) Gene. Andrew ElBardissi, The Pennsylvania State University
 Understanding the Cellular Mechanism of the Excess Microsporocytes I (EMSI) Gene Andrew ElBardissi, The Pennsylvania State University Abstract: Hong Ma, The Pennsylvania State University The Excess Microsporocytes
Understanding the Cellular Mechanism of the Excess Microsporocytes I (EMSI) Gene Andrew ElBardissi, The Pennsylvania State University Abstract: Hong Ma, The Pennsylvania State University The Excess Microsporocytes
Protocols for cell lines using CRISPR/CAS
 Protocols for cell lines using CRISPR/CAS Procedure overview Map Preparation of CRISPR/CAS plasmids Expression vectors for guide RNA (U6-gRNA) and Cas9 gene (CMV-p-Cas9) are ampicillin-resist ant and stable
Protocols for cell lines using CRISPR/CAS Procedure overview Map Preparation of CRISPR/CAS plasmids Expression vectors for guide RNA (U6-gRNA) and Cas9 gene (CMV-p-Cas9) are ampicillin-resist ant and stable
Purification and Sequencing of DNA Guides from Prokaryotic Argonaute Daan C. Swarts *, Edze R. Westra, Stan J. J. Brouns and John van der Oost
 Purification and Sequencing of DNA Guides from Prokaryotic Argonaute Daan C. Swarts *, Edze R. Westra, Stan J. J. Brouns and John van der Oost Department of Agrotechnology and Food Sciences, Wageningen
Purification and Sequencing of DNA Guides from Prokaryotic Argonaute Daan C. Swarts *, Edze R. Westra, Stan J. J. Brouns and John van der Oost Department of Agrotechnology and Food Sciences, Wageningen
ITS Sequencing in Millepora. 10/09 Subcloning DNA Fragments into pbluescript Preparation of pbluescript Vector
 Page 1 of 5 10/09 Subcloning DNA Fragments into pbluescript Preparation of pbluescript Vector 1. Digest 1 µg of pbluescript with Eco RI 2. Following digestion, add 0.1 volumes of 3M sodium acetate (ph
Page 1 of 5 10/09 Subcloning DNA Fragments into pbluescript Preparation of pbluescript Vector 1. Digest 1 µg of pbluescript with Eco RI 2. Following digestion, add 0.1 volumes of 3M sodium acetate (ph
igem 2016 Microbiology BMB SDU
 igem 2016 Microbiology BMB SDU Project type: Bacteriocin Project title: IMPACT (Intein Mediated Purification with an Affinity Chitin-binding Tag) Sub project: Purify hybrid-bacteriocin Laterosporulin-Thuricin
igem 2016 Microbiology BMB SDU Project type: Bacteriocin Project title: IMPACT (Intein Mediated Purification with an Affinity Chitin-binding Tag) Sub project: Purify hybrid-bacteriocin Laterosporulin-Thuricin
Construction of pcxz14w, a Novel puc19-derived Plasmid Encoding the rop Gene
 Construction of pcxz14w, a Novel puc19-derived Plasmid Encoding the rop Gene Shary Chen, Ziyan Xu, Wenchen Zhao Department of Microbiology and Immunology, University of British Columbia When the ColE1-derived
Construction of pcxz14w, a Novel puc19-derived Plasmid Encoding the rop Gene Shary Chen, Ziyan Xu, Wenchen Zhao Department of Microbiology and Immunology, University of British Columbia When the ColE1-derived
CLONING INVERTED REPEATS IN HIGH THROUGHPUT
 CLONING INVERTED REPEATS IN HIGH THROUGHPUT This protocol is calculated for cloning one 96 well plate 1. design primers: as standard we include a EcoRI restriction site on the 5 primer and XbaI on the
CLONING INVERTED REPEATS IN HIGH THROUGHPUT This protocol is calculated for cloning one 96 well plate 1. design primers: as standard we include a EcoRI restriction site on the 5 primer and XbaI on the
Protocols. We used a buffer mix with polymerase (either Mango Mix (forhandler) or 2x High Fidelity) and the appropriate primers and template DNA.
 Protocols PCRs Colony PCRs We used a buffer mix with polymerase (either Mango Mix (forhandler) or 2x High Fidelity) and the appropriate primers and template DNA. For a 10 µl reaction 2x Mango Mix/2x High
Protocols PCRs Colony PCRs We used a buffer mix with polymerase (either Mango Mix (forhandler) or 2x High Fidelity) and the appropriate primers and template DNA. For a 10 µl reaction 2x Mango Mix/2x High
DNA miniprep by Alkaline Lysis (activity)
 DNA miniprep by Alkaline Lysis (activity) Contents 1 Alkaline Lysis 2 Exercise 1: Plasmid DNA Mini-Prep by Alkaline Lysis 3 Identification of Plasmid DNA 4 Exercise 2: Restriction Digestion Identification
DNA miniprep by Alkaline Lysis (activity) Contents 1 Alkaline Lysis 2 Exercise 1: Plasmid DNA Mini-Prep by Alkaline Lysis 3 Identification of Plasmid DNA 4 Exercise 2: Restriction Digestion Identification
CHAPTER 5 PTP-1B CLONING AND RECOMBINANT PROTEIN EXPRESSION. Recombinant DNA technology has revolutionized molecular biology and
 204 CHAPTER 5 PTP-1B CLONING AND RECOMBINANT PROTEIN EXPRESSION SUMMARY Recombinant DNA technology has revolutionized molecular biology and genetics. Today, virtually any segment of DNA, the genetic material
204 CHAPTER 5 PTP-1B CLONING AND RECOMBINANT PROTEIN EXPRESSION SUMMARY Recombinant DNA technology has revolutionized molecular biology and genetics. Today, virtually any segment of DNA, the genetic material
Generation of gene knockout vectors for Dictyostelium discoideum
 Generation of gene knockout vectors for Dictyostelium discoideum Instruction manual Last date of revision April 2012 2015 Version PR29-0001 PR29-0003 www.stargate.com This manual can be downloaded under
Generation of gene knockout vectors for Dictyostelium discoideum Instruction manual Last date of revision April 2012 2015 Version PR29-0001 PR29-0003 www.stargate.com This manual can be downloaded under
Farnham Lab Protocol for CRISPR/Cas 9- mediated enhancer deletion using a puromycin selection vector Created by Yu (Phoebe) Guo
 Farnham Lab Protocol for CRISPR/Cas 9- mediated enhancer deletion using a puromycin selection vector Created by Yu (Phoebe) Guo 20151024 Part A. Design grna oligos 1. Design grnas Go to Optimized CRISPR
Farnham Lab Protocol for CRISPR/Cas 9- mediated enhancer deletion using a puromycin selection vector Created by Yu (Phoebe) Guo 20151024 Part A. Design grna oligos 1. Design grnas Go to Optimized CRISPR
Recombinant DNA recombinant DNA DNA cloning gene cloning
 DNA Technology Recombinant DNA In recombinant DNA, DNA from two different sources, often two species, are combined into the same DNA molecule. DNA cloning permits production of multiple copies of a specific
DNA Technology Recombinant DNA In recombinant DNA, DNA from two different sources, often two species, are combined into the same DNA molecule. DNA cloning permits production of multiple copies of a specific
Presto Mini Plasmid Kit
 Instruction Manual Ver. 03.06.17 For Research Use Only Presto Mini Plasmid Kit PDH004 (4 Preparation Sample Kit) PDH100 (100 Preparation Kit) PDH300 (300 Preparation Kit) Advantages Sample: 1-7 ml of cultured
Instruction Manual Ver. 03.06.17 For Research Use Only Presto Mini Plasmid Kit PDH004 (4 Preparation Sample Kit) PDH100 (100 Preparation Kit) PDH300 (300 Preparation Kit) Advantages Sample: 1-7 ml of cultured
Polymerase Chain Reaction (PCR)
 Laboratory for Environmental Pathogens Research Department of Environmental Sciences University of Toledo Polymerase Chain Reaction (PCR) Background information The polymerase chain reaction (PCR) is an
Laboratory for Environmental Pathogens Research Department of Environmental Sciences University of Toledo Polymerase Chain Reaction (PCR) Background information The polymerase chain reaction (PCR) is an
CSS451 Spring 2010 Polymerase Chain Reaction Laboratory
 CSS451 Spring 2010 Polymerase Chain Reaction Laboratory The purpose of the polymerase chain reaction (PCR) is to amplify specific segments of DNA. If one knows the DNA sequence of regions of DNA that flank
CSS451 Spring 2010 Polymerase Chain Reaction Laboratory The purpose of the polymerase chain reaction (PCR) is to amplify specific segments of DNA. If one knows the DNA sequence of regions of DNA that flank
mrnaexpress mrna Synthesis Kit Cat. #MR-KIT-1
 mrnaexpress mrna Synthesis Kit Cat. #MR-KIT-1 User Manual Check Individual Components for Storage conditions ver. 2-070918 A limited-use label license covers this product. By use of this product, you accept
mrnaexpress mrna Synthesis Kit Cat. #MR-KIT-1 User Manual Check Individual Components for Storage conditions ver. 2-070918 A limited-use label license covers this product. By use of this product, you accept
Vector Linearization. igem TU/e 2015 Biomedical Engineering
 igem TU/e 2015 Biomedical Engineering Eindhoven University of Technology Room: Ceres 0.04 Den Dolech 2, 5612 AZ Eindhoven The Netherlands Tel. no. +31 50 247 55 59 2015.igem.org/Team:TU_Eindhoven Vector
igem TU/e 2015 Biomedical Engineering Eindhoven University of Technology Room: Ceres 0.04 Den Dolech 2, 5612 AZ Eindhoven The Netherlands Tel. no. +31 50 247 55 59 2015.igem.org/Team:TU_Eindhoven Vector
pcmv6-neo Vector Application Guide Table of Contents Package Contents and Storage Conditions Each kit comes with the following components:
 pcmv6-neo Vector Application Guide Table of Contents Package Contents and Storage Conditions... 1 Product Description... 1 Introduction... 1 Production and Quality Assurance... 2 Methods... 2 Other required
pcmv6-neo Vector Application Guide Table of Contents Package Contents and Storage Conditions... 1 Product Description... 1 Introduction... 1 Production and Quality Assurance... 2 Methods... 2 Other required
Recombineering Manual
 Recombineering Manual Anthony Popkie The Phiel Laboratory The Research Institute at Nationwide Childrenʼs Hospital 1 BAC Transformation BACs may be transformed into either DY380, EL250 or EL350 cells.
Recombineering Manual Anthony Popkie The Phiel Laboratory The Research Institute at Nationwide Childrenʼs Hospital 1 BAC Transformation BACs may be transformed into either DY380, EL250 or EL350 cells.
Modifications to EXPERIMENTS 21 and 24: PCR and Molecular Cloning
 Modifications to EXPERIMENTS 21 and 24: PCR and Molecular Cloning This experiment was designed by Dylan Dodd, based on research completed in Dr. Isaac Cann s lab*, with modifications and editing of content
Modifications to EXPERIMENTS 21 and 24: PCR and Molecular Cloning This experiment was designed by Dylan Dodd, based on research completed in Dr. Isaac Cann s lab*, with modifications and editing of content
BACTERIAL PRODUCTION EXPRESSION METHOD OVERVIEW: PEF # GENE NAME EXPRESSION VECTOR MOLECULAR WEIGHT kda (full-length) 34.
 BACTERIAL PRODUCTION PEF # GENE NAME EXPRESSION VECTOR MOLECULAR WEIGHT 2015-XXXX XXXX pet-32a 50.9 kda (full-length) 34.0 kda (cleaved) EXPRESSION METHOD OVERVIEW: Plasmid DNA was transformed into BL21
BACTERIAL PRODUCTION PEF # GENE NAME EXPRESSION VECTOR MOLECULAR WEIGHT 2015-XXXX XXXX pet-32a 50.9 kda (full-length) 34.0 kda (cleaved) EXPRESSION METHOD OVERVIEW: Plasmid DNA was transformed into BL21
BIO/CHEM 475 Molecular Biology Laboratory Spring 2007 Biol/Chem 475 Part 2 of Cloning Lab
 BIO/CHEM 475 Molecular Biology Laboratory Spring 2007 Biol/Chem 475 Part 2 of Cloning Lab Week 5 Analysis of pgem recombinant clones using PCR Clonecheck to determine size of insert; select clones for
BIO/CHEM 475 Molecular Biology Laboratory Spring 2007 Biol/Chem 475 Part 2 of Cloning Lab Week 5 Analysis of pgem recombinant clones using PCR Clonecheck to determine size of insert; select clones for
TIANgel Mini DNA Purification Kit
 TIANgel Mini DNA Purification Kit For DNA purification from agarose and polyacrylamide gels www.tiangen.com/en DP130419 TIANgel Mini DNA Purification Kit Kit Contents (Spin column) Cat. no. DP208 Contents
TIANgel Mini DNA Purification Kit For DNA purification from agarose and polyacrylamide gels www.tiangen.com/en DP130419 TIANgel Mini DNA Purification Kit Kit Contents (Spin column) Cat. no. DP208 Contents
Rotation Report Sample Version 3. Due Date: August 9, Analysis of the Guanine Nucleotide Exchange Activity of the S. cerevisiae Ats1 Protein
 Rotation Report Sample Version 3 Due Date: August 9, 1998 Analysis of the Guanine Nucleotide Exchange Activity of the S. cerevisiae Ats1 Protein Anita H. Corbett Advisor: Amy Jones Rotation 1 Abstract:
Rotation Report Sample Version 3 Due Date: August 9, 1998 Analysis of the Guanine Nucleotide Exchange Activity of the S. cerevisiae Ats1 Protein Anita H. Corbett Advisor: Amy Jones Rotation 1 Abstract:
Overview: The DNA Toolbox
 Overview: The DNA Toolbox Sequencing of the genomes of more than 7,000 species was under way in 2010 DNA sequencing has depended on advances in technology, starting with making recombinant DNA In recombinant
Overview: The DNA Toolbox Sequencing of the genomes of more than 7,000 species was under way in 2010 DNA sequencing has depended on advances in technology, starting with making recombinant DNA In recombinant
Amplified restriction fragments for genomic enrichment. 1 Reagents and Equipment. Tom Parchman Zach Gompert
 1 Amplified restriction fragments for genomic enrichment (version 2.3 August 2011) Tom Parchman (tparchma@uwyo.edu) Zach Gompert (zgompert@uwyo.edu) Alex Buerkle (buerkle@uwyo.edu) University of Wyoming
1 Amplified restriction fragments for genomic enrichment (version 2.3 August 2011) Tom Parchman (tparchma@uwyo.edu) Zach Gompert (zgompert@uwyo.edu) Alex Buerkle (buerkle@uwyo.edu) University of Wyoming
Justin Veazey. Experiment 3; Analysis of digestion products of puc19, GFPuv, and pgem-t easy
 Veazey 1 Justin Veazey 7A Experiment 3; Analysis of digestion products of puc19, GFPuv, and pgem-t easy Construction of recombinants GFPuv-pGEM-T easy and GFPuv-pUC19 Transformation and analysis of recombinant
Veazey 1 Justin Veazey 7A Experiment 3; Analysis of digestion products of puc19, GFPuv, and pgem-t easy Construction of recombinants GFPuv-pGEM-T easy and GFPuv-pUC19 Transformation and analysis of recombinant
