Introduction to Random Forests for Gene Expression Data. Utah State University Spring 2014 STAT 5570: Statistical Bioinformatics Notes 3.
|
|
|
- Ashlyn Bruce
- 5 years ago
- Views:
Transcription
1 Introduction to Random Forests for Gene Expression Data Utah State University Spring 2014 STAT 5570: Statistical Bioinformatics Notes 3.5 1
2 References Breiman, Machine Learning (2001) 45(1): Diaz-Uriarte and Alvarez de Andres, BMC Bioinformatics (2006) 7:3. Cutler, Cutler, and Stevens (2012) Random Forests. In Zhang and Ma, editors, Ensemble Machine Learning: Methods and Applications, pp
3 Gene Profiling / Selection Observe gene expression in different conditions healthy vs. diseased, e.g. Use simultaneous expression profiles of thousands of genes (what are the genes doing across arrays) Look at which genes are important in separating the two conditions; i.e., what determines the conditions signatures 3
4 Machine Learning Computational & statistical inference processes: observed data reusable algorithms for prediction Why machine? want minimal: human involvement Why learning? develop ability to predict Here, supervised learning: use knowledge of condition type 4
5 Machine Learning Methods Neural Network SVM (Support Vector Machine) RPART (Recursive PArtitioning and Regression Trees) CART (Classification and Regression Trees) Ensembling Learning (average of many trees) Boosting (Shapire et al., 1998) Bagging (Breiman, 1996) RandomForests (Breiman, 2001; Cutler & Stevens 2006; Cutler, Cutler, & Stevens 2008) 5
6 CART: Classification and Regression Trees Each individual (array) has data on many predictors (genes) and one response (disease state) Think of a tree, with splits based on levels of specific predictors Gene1 > 8.3 Diseased (18:21) Gene1 < 8.3 Healthy Diseased Choose predictors and (15:10) (3:11) split levels to maximize purity in new groups; the best split at each node Prediction made by: passing test cases down tree 6
7 CART generalized: Random Forests Rather than using all predictors and all individuals to make a single tree, make a forest of many (ntree) trees, each one based on a random selection of predictors and individuals Each tree is fit using a bootstrap sample of data (draw with replacement) and grown until each node is pure Each node is split using the best among a subset (of size mtry) of predictors randomly chosen at that node (default is sqrt. of # of predictors) (special case using all predictors: bagging) Prediction made by aggregating across the forest (majority vote or average) 7
8 How to measure goodness? Each tree fit on a training set (bootstrap sample), or the bag The left-over cases ( out-of-bag ) can be used as a test set for that tree (usually 1/3 of original data) The out-of-bag (OOB) error rate is the: % misclassification 8
9 What does RF give us? Kind of a black box but can look at variable importance For each tree, look at the OOB data: Permute values of predictor j among all OOB cases Pass OOB data down the tree, save the predictions For case i of OOB and predictor j, get: OOB error rate with variable j permuted OOB error rate before permutation Average across forest to get overall variable importance for each predictor j 9
10 Why variable importance? Recall: identifying conditions signatures Sort genes by some criterion Want smallest set of genes to achieve good diagnostic ability 10
11 Recall ALL Data RMA-preprocessed gene expression data Chiaretti et al., Blood (2004) 103(7) genes (hgu95av2 Affymetrix GeneChip) 128 samples (arrays) phenotypic data on all 128 patients, including: 95 B-cell cancer 33 T-cell cancer 11
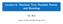
12 ALL subset: 4,400 genes, 30 arrays (15 B, 15 T) Look at top 25 most important genes from Random Forest. Color scale: purple (low) to orange (high) 12
13 ### First prepare objects for RF ### (similar to slide 13 of Notes 3.3) # load data library(affy); library(all); data(all) # (here, filter genes on raw scale, then return to log scale) # also, keep 30 arrays here JUST for computational # convenience (in-class) library(genefilter); e.mat <- 2^(exprs(ALL)[,c(81:110)]) ffun <- filterfun(povera(0.20,100)) t.fil <- genefilter(e.mat,ffun) small.eset <- log2(e.mat[t.fil,]) dim(small.eset) # 4400 genes, 30 arrays (15 B and 15 T) group <- c(rep('b',15),rep('t',15)) # group classification, in order 13
14 # One RF library(randomforest) set.seed(1234) print(date()) rf <- randomforest(x=t(small.eset),y=as.factor(group), ntree=10000) print(date()) # about 20 seconds # Make variable importance plot varimpplot(rf, n.var=25, main='all Subset Results') # Get names of most important genes imp.temp <- importance(rf) t <- order(imp.temp,decreasing=true) gn.imp <- rownames(imp.temp)[t] # these are in order most...least important 14
15 # Get expression values for 25 most important genes gn.25 <- gn.imp[1:25] # vector of top 25 genes, in order t <- is.element(rownames(small.eset),gn.25) sig.eset <- small.eset[t,] # matrix of expression values, # not necessarily in order of importance # Make a heatmap, with group differences obvious on plot # (similar to Notes 2.3) library(rcolorbrewer) hmcol <- colorramppalette(brewer.pal(9,"reds"))(256) colnames(sig.eset) <- group # This will label the heatmap columns csc <- rep(hmcol[50],30) csc[group=='t'] <- hmcol[200] # column side color will be dark for T and light for B heatmap(sig.eset,scale="row", col=hmcol,colsidecolors=csc) 15
16 Can focus on variable selection Iteratively fit many forests, each time discarding predictors with low importance from previous iterations Use bootstrap to assess standard error of error rates Choose the forest with the smallest number of genes whose error rate is within u standard errors of the minimum error rate of all forests (u = 0 or 1, typically) Reference: Diaz-Uriarte and Alvarez de Andres, BMC Bioinformatics (2006) 7:3. Online tool: 16
17 RF Variable Selection on ALL subset Subset: 30 arrays, 4400 genes (non-subset is the remaining 98 arrays) First forest: 10,000 trees; Subsequent forests: 2,000 trees At each iteration, drop 20% of variables (genes) until a 2-variable model is built; return the best of the series of models considered. 17
18 # Look at variable selection library(varselrf) set.seed(1234) print(date()) rfsel <- varselrf(t(small.eset),as.factor(group), ntree=10000, ntreeiterat=2000, vars.drop.frac=0.2) print(date()) # 40 seconds # rfsel$firstforest is the same as the slide 14 rf object rf.sig.gn <- rfsel$selected.vars # "38147_at" "38319_at" # set.seed(123) above gives genes: "2059_s_at" "38319_at" # Visualize these two genes exp.gn.1 <- small.eset[rownames(small.eset)==rf.sig.gn[1],] exp.gn.2 <- small.eset[rownames(small.eset)==rf.sig.gn[2],] use.pch <- c(rep(1,15),rep(16,15)) # Define plotting chars. use.col <- c(rep(1,15),rep(2,15)) # Define plotting colors plot(exp.gn.1,exp.gn.2,col=use.col,main='30 subset arrays', cex.main=1.5, cex.lab=1.5, xlab=rf.sig.gn[1], ylab=rf.sig.gn[2], pch=use.pch,cex=2) legend('bottomright', c('b-cell','t-cell'),pch=c(1,16),col=c(1,2),cex=1.5) 18
19 # Did this overfit these 30 arrays? # Look at JUST the other 98 # (first 80 are B-cell, last 18 are T-cell) eset.2 <- exprs(all)[,c(1:80,111:128)] group.2 <- c(rep(0,80),rep(1,18)) exp.gn.1 <- eset.2[rownames(eset.2)==rf.sig.gn[1],] exp.gn.2 <- eset.2[rownames(eset.2)==rf.sig.gn[2],] use.pch.2 <- c(rep(1,80),rep(16,18)) use.col.2 <- c(rep(1,80),rep(2,18)) plot(exp.gn.1,exp.gn.2,col=use.col.2, main='non-subset arrays', cex.main=1.5,cex=2, cex.lab=1.5, xlab=rf.sig.gn[1], ylab=rf.sig.gn[2], pch=use.pch.2) legend('bottomright', c('b-cell','t-cell'),pch=c(1,16),col=c(1,2),cex=1.5) 19
20 RF Variable Selection on ALL (full) Each condition has a profile / signature (across these genes) full ALL data: 12,625 genes, 128 arrays 20
21 # RF variable selection with full data set # set seed and define initial objects set.seed(123) eset <- exprs(all) # genes, 128 arrays cell <- c(rep(0,95),rep(1,33)) # first 95 are B-cell; last 33 are T-cell print(date()) rf.big <- varselrf(t(eset),as.factor(cell), ntree=10000, ntreeiterat=2000, vars.drop.frac=0.2) print(date()) # about 9 minutes rf.gn <- rf.big$selected.vars # "33039_at" "35016_at" "38319_at" 21
22 # make scatterplot matrix, with points colored by cell type t.rf <- is.element(rownames(eset),rf.gn) rf.eset <- t(eset[t.rf,]) # this rf.eset has rows for obs and columns for 3 genes use.pch <- c(rep(1,95),rep(16,33)) use.col <- cell+1 pairs(rf.eset,col=use.col,pch=use.pch,cex=1.5) # pairs function makes scatterplot matrix of rf.eset cols. # Now - make a profile plot (parallel coordinates plot) library(mass) parcoord(rf.eset, col=cell+1, lty=cell+1, lwd=3, var.label=true) legend(1.2,.15,c('b-cell','t-cell'),lty=c(1,2), lwd=3,col=c(1,2),bty='n') 22
23 Summary: RF for gene expression data Works well even with: many more variables than observations, many-valued categoricals, extensive missing values, badly unbalanced data Low error (comparable w/ boosting and SVM) Robustness (even with large noise in genes) Does not overfit Fast, and invariant to monotone transformations of the predictors Free! (Fortran code by Breiman and Cutler) Returns the importance of predictors (gene) Little need to tune parameters But, no: statistical inference 23
References. Introduction to Random Forests for Gene Expression Data. Machine Learning. Gene Profiling / Selection
 References Introduction to Random Forests for Gene Expression Data Utah State University Spring 2014 STAT 5570: Statistical Bioinformatics Notes 3.5 Breiman, Machine Learning (2001) 45(1): 5-32. Diaz-Uriarte
References Introduction to Random Forests for Gene Expression Data Utah State University Spring 2014 STAT 5570: Statistical Bioinformatics Notes 3.5 Breiman, Machine Learning (2001) 45(1): 5-32. Diaz-Uriarte
Today. Last time. Lecture 5: Discrimination (cont) Jane Fridlyand. Oct 13, 2005
 Biological question Experimental design Microarray experiment Failed Lecture : Discrimination (cont) Quality Measurement Image analysis Preprocessing Jane Fridlyand Pass Normalization Sample/Condition
Biological question Experimental design Microarray experiment Failed Lecture : Discrimination (cont) Quality Measurement Image analysis Preprocessing Jane Fridlyand Pass Normalization Sample/Condition
Copyright 2013, SAS Institute Inc. All rights reserved.
 IMPROVING PREDICTION OF CYBER ATTACKS USING ENSEMBLE MODELING June 17, 2014 82 nd MORSS Alexandria, VA Tom Donnelly, PhD Systems Engineer & Co-insurrectionist JMP Federal Government Team ABSTRACT Improving
IMPROVING PREDICTION OF CYBER ATTACKS USING ENSEMBLE MODELING June 17, 2014 82 nd MORSS Alexandria, VA Tom Donnelly, PhD Systems Engineer & Co-insurrectionist JMP Federal Government Team ABSTRACT Improving
CS6716 Pattern Recognition
 CS6716 Pattern Recognition Aaron Bobick School of Interactive Computing Administrivia Shray says the problem set is close to done Today chapter 15 of the Hastie book. Very few slides brought to you by
CS6716 Pattern Recognition Aaron Bobick School of Interactive Computing Administrivia Shray says the problem set is close to done Today chapter 15 of the Hastie book. Very few slides brought to you by
Random forest for gene selection and microarray data classification
 www.bioinformation.net Hypothesis Volume 7(3) Random forest for gene selection and microarray data classification Kohbalan Moorthy & Mohd Saberi Mohamad* Artificial Intelligence & Bioinformatics Research
www.bioinformation.net Hypothesis Volume 7(3) Random forest for gene selection and microarray data classification Kohbalan Moorthy & Mohd Saberi Mohamad* Artificial Intelligence & Bioinformatics Research
Lecture 6: Decision Tree, Random Forest, and Boosting
 Lecture 6: Decision Tree, Random Forest, and Boosting Tuo Zhao Schools of ISyE and CSE, Georgia Tech CS764/ISYE/CSE 674: Machine Learning/Computational Data Analysis Tree? Tuo Zhao Lecture 6: Decision
Lecture 6: Decision Tree, Random Forest, and Boosting Tuo Zhao Schools of ISyE and CSE, Georgia Tech CS764/ISYE/CSE 674: Machine Learning/Computational Data Analysis Tree? Tuo Zhao Lecture 6: Decision
Salford Predictive Modeler. Powerful machine learning software for developing predictive, descriptive, and analytical models.
 Powerful machine learning software for developing predictive, descriptive, and analytical models. The Company Minitab helps companies and institutions to spot trends, solve problems and discover valuable
Powerful machine learning software for developing predictive, descriptive, and analytical models. The Company Minitab helps companies and institutions to spot trends, solve problems and discover valuable
3 Ways to Improve Your Targeted Marketing with Analytics
 3 Ways to Improve Your Targeted Marketing with Analytics Introduction Targeted marketing is a simple concept, but a key element in a marketing strategy. The goal is to identify the potential customers
3 Ways to Improve Your Targeted Marketing with Analytics Introduction Targeted marketing is a simple concept, but a key element in a marketing strategy. The goal is to identify the potential customers
The KEGGgraph Package. Stat 6570 Michelle Carlsen April 21, 2014
 The KEGGgraph Package Stat 6570 Michelle Carlsen April 21, 2014 Kyoto Encyclopedia of Genes and Genomes An effort to map the whole of the genome Created a database of functions and utilities of the biological
The KEGGgraph Package Stat 6570 Michelle Carlsen April 21, 2014 Kyoto Encyclopedia of Genes and Genomes An effort to map the whole of the genome Created a database of functions and utilities of the biological
The parameter sensitivity of random forests
 Huang and Boutros BMC Bioinformatics (2016) 17:331 DOI 10.1186/s12859-016-1228-x METHODOLOGY ARTICLE Open Access The parameter sensitivity of random forests Barbara F.F. Huang 1 and Paul C. Boutros 1,2,3,4*
Huang and Boutros BMC Bioinformatics (2016) 17:331 DOI 10.1186/s12859-016-1228-x METHODOLOGY ARTICLE Open Access The parameter sensitivity of random forests Barbara F.F. Huang 1 and Paul C. Boutros 1,2,3,4*
Exploring Contributing Features of Pre-Graft Orthodontic Treatment of Cleft Lip and Palate Patients Using Random Forests
 Transactions on Science and Technology Vol. 5, No., 5 -, 28 Exploring Contributing Features of Pre-Graft Orthodontic Treatment of Cleft Lip and Palate Patients Using Random Forests Zaturrawiah Ali Omar
Transactions on Science and Technology Vol. 5, No., 5 -, 28 Exploring Contributing Features of Pre-Graft Orthodontic Treatment of Cleft Lip and Palate Patients Using Random Forests Zaturrawiah Ali Omar
Determining Method of Action in Drug Discovery Using Affymetrix Microarray Data
 Determining Method of Action in Drug Discovery Using Affymetrix Microarray Data Max Kuhn max.kuhn@pfizer.com Pfizer Global R&D Research Statistics Groton, CT Method of Action As the level of drug resistance
Determining Method of Action in Drug Discovery Using Affymetrix Microarray Data Max Kuhn max.kuhn@pfizer.com Pfizer Global R&D Research Statistics Groton, CT Method of Action As the level of drug resistance
Accurate Campaign Targeting Using Classification Algorithms
 Accurate Campaign Targeting Using Classification Algorithms Jieming Wei Sharon Zhang Introduction Many organizations prospect for loyal supporters and donors by sending direct mail appeals. This is an
Accurate Campaign Targeting Using Classification Algorithms Jieming Wei Sharon Zhang Introduction Many organizations prospect for loyal supporters and donors by sending direct mail appeals. This is an
Let the data speak: Machine learning methods for data editing and imputation
 Working Paper 31 UNITED NATIONS ECONOMIC COMMISSION FOR EUROPE CONFERENCE OF EUROPEAN STATISTICIANS Work Session on Statistical Data Editing (Budapest, Hungary, 14-16 September 2015) Topic (v): Emerging
Working Paper 31 UNITED NATIONS ECONOMIC COMMISSION FOR EUROPE CONFERENCE OF EUROPEAN STATISTICIANS Work Session on Statistical Data Editing (Budapest, Hungary, 14-16 September 2015) Topic (v): Emerging
Feature selection methods for SVM classification of microarray data
 Feature selection methods for SVM classification of microarray data Mike Love December 11, 2009 SVMs for microarray classification tasks Linear support vector machines have been used in microarray experiments
Feature selection methods for SVM classification of microarray data Mike Love December 11, 2009 SVMs for microarray classification tasks Linear support vector machines have been used in microarray experiments
Predicting elevated groundwater nitrate concentrations using a decision tree-based geospatial model in the Silver Springshed, FL
 Predicting elevated groundwater nitrate concentrations using a decision tree-based geospatial model in the Silver Springshed, FL Dean R. Dobberfuhl Andy Canion Lori McCloud Springs Research Questions Which
Predicting elevated groundwater nitrate concentrations using a decision tree-based geospatial model in the Silver Springshed, FL Dean R. Dobberfuhl Andy Canion Lori McCloud Springs Research Questions Which
Tree Depth in a Forest
 Tree Depth in a Forest Mark Segal Center for Bioinformatics & Molecular Biostatistics Division of Bioinformatics Department of Epidemiology and Biostatistics UCSF NUS / IMS Workshop on Classification and
Tree Depth in a Forest Mark Segal Center for Bioinformatics & Molecular Biostatistics Division of Bioinformatics Department of Epidemiology and Biostatistics UCSF NUS / IMS Workshop on Classification and
Linear model to forecast sales from past data of Rossmann drug Store
 Abstract Linear model to forecast sales from past data of Rossmann drug Store Group id: G3 Recent years, the explosive growth in data results in the need to develop new tools to process data into knowledge
Abstract Linear model to forecast sales from past data of Rossmann drug Store Group id: G3 Recent years, the explosive growth in data results in the need to develop new tools to process data into knowledge
In silico prediction of novel therapeutic targets using gene disease association data
 In silico prediction of novel therapeutic targets using gene disease association data, PhD, Associate GSK Fellow Scientific Leader, Computational Biology and Stats, Target Sciences GSK Big Data in Medicine
In silico prediction of novel therapeutic targets using gene disease association data, PhD, Associate GSK Fellow Scientific Leader, Computational Biology and Stats, Target Sciences GSK Big Data in Medicine
Advanced Statistical Methods: Beyond Linear Regression
 Advanced Statistical Methods: Beyond Linear Regression John R. Stevens Utah State University Notes 1. Case Study Data Sets Mathematics Educators Workshop 28 March 2009 1 http://www.stat.usu.edu/~jrstevens/pcmi
Advanced Statistical Methods: Beyond Linear Regression John R. Stevens Utah State University Notes 1. Case Study Data Sets Mathematics Educators Workshop 28 March 2009 1 http://www.stat.usu.edu/~jrstevens/pcmi
PREDICTION OF CONCRETE MIX COMPRESSIVE STRENGTH USING STATISTICAL LEARNING MODELS
 Journal of Engineering Science and Technology Vol. 13, No. 7 (2018) 1916-1925 School of Engineering, Taylor s University PREDICTION OF CONCRETE MIX COMPRESSIVE STRENGTH USING STATISTICAL LEARNING MODELS
Journal of Engineering Science and Technology Vol. 13, No. 7 (2018) 1916-1925 School of Engineering, Taylor s University PREDICTION OF CONCRETE MIX COMPRESSIVE STRENGTH USING STATISTICAL LEARNING MODELS
Random Forests. Parametrization and Dynamic Induction
 Random Forests Parametrization and Dynamic Induction Simon Bernard Document and Learning research team LITIS laboratory University of Rouen, France décembre 2014 Random Forest Classifiers Random Forests
Random Forests Parametrization and Dynamic Induction Simon Bernard Document and Learning research team LITIS laboratory University of Rouen, France décembre 2014 Random Forest Classifiers Random Forests
SPM 8.2. Salford Predictive Modeler
 SPM 8.2 Salford Predictive Modeler SPM 8.2 The SPM Salford Predictive Modeler software suite is a highly accurate and ultra-fast platform for developing predictive, descriptive, and analytical models from
SPM 8.2 Salford Predictive Modeler SPM 8.2 The SPM Salford Predictive Modeler software suite is a highly accurate and ultra-fast platform for developing predictive, descriptive, and analytical models from
BIOINF/BENG/BIMM/CHEM/CSE 184: Computational Molecular Biology. Lecture 2: Microarray analysis
 BIOINF/BENG/BIMM/CHEM/CSE 184: Computational Molecular Biology Lecture 2: Microarray analysis Genome wide measurement of gene transcription using DNA microarray Bruce Alberts, et al., Molecular Biology
BIOINF/BENG/BIMM/CHEM/CSE 184: Computational Molecular Biology Lecture 2: Microarray analysis Genome wide measurement of gene transcription using DNA microarray Bruce Alberts, et al., Molecular Biology
Evaluation of random forest regression for prediction of breeding value from genomewide SNPs
 c Indian Academy of Sciences RESEARCH ARTICLE Evaluation of random forest regression for prediction of breeding value from genomewide SNPs RUPAM KUMAR SARKAR 1,A.R.RAO 1, PRABINA KUMAR MEHER 1, T. NEPOLEAN
c Indian Academy of Sciences RESEARCH ARTICLE Evaluation of random forest regression for prediction of breeding value from genomewide SNPs RUPAM KUMAR SARKAR 1,A.R.RAO 1, PRABINA KUMAR MEHER 1, T. NEPOLEAN
Matrix Factorization-Based Data Fusion for Drug-Induced Liver Injury Prediction
 Matrix Factorization-Based Data Fusion for Drug-Induced Liver Injury Prediction Marinka Žitnik1 and Blaž Zupan 1,2 1 Faculty of Computer and Information Science, University of Ljubljana, Tržaška 25, 1000
Matrix Factorization-Based Data Fusion for Drug-Induced Liver Injury Prediction Marinka Žitnik1 and Blaž Zupan 1,2 1 Faculty of Computer and Information Science, University of Ljubljana, Tržaška 25, 1000
Introduction to gene expression microarray data analysis
 Introduction to gene expression microarray data analysis Outline Brief introduction: Technology and data. Statistical challenges in data analysis. Preprocessing data normalization and transformation. Useful
Introduction to gene expression microarray data analysis Outline Brief introduction: Technology and data. Statistical challenges in data analysis. Preprocessing data normalization and transformation. Useful
Prediction and Interpretation for Machine Learning Regression Methods
 Paper 1967-2018 Prediction and Interpretation for Machine Learning Regression Methods D. Richard Cutler, Utah State University ABSTRACT The last 30 years has seen extraordinary development of new tools
Paper 1967-2018 Prediction and Interpretation for Machine Learning Regression Methods D. Richard Cutler, Utah State University ABSTRACT The last 30 years has seen extraordinary development of new tools
Gene Set Enrichment Analysis! Robert Gentleman!
 Gene Set Enrichment Analysis! Robert Gentleman! Outline! Description of the experimental setting! Defining gene sets! Description of the original GSEA algorithm! proposed by Mootha et al (2003)! Our approach
Gene Set Enrichment Analysis! Robert Gentleman! Outline! Description of the experimental setting! Defining gene sets! Description of the original GSEA algorithm! proposed by Mootha et al (2003)! Our approach
Predicting prokaryotic incubation times from genomic features Maeva Fincker - Final report
 Predicting prokaryotic incubation times from genomic features Maeva Fincker - mfincker@stanford.edu Final report Introduction We have barely scratched the surface when it comes to microbial diversity.
Predicting prokaryotic incubation times from genomic features Maeva Fincker - mfincker@stanford.edu Final report Introduction We have barely scratched the surface when it comes to microbial diversity.
Data Science in a pricing process
 Data Science in a pricing process Michaël Casalinuovo Consultant, ADDACTIS Software michael.casalinuovo@addactis.com Contents Nowadays, we live in a continuously changing market environment, Pricing has
Data Science in a pricing process Michaël Casalinuovo Consultant, ADDACTIS Software michael.casalinuovo@addactis.com Contents Nowadays, we live in a continuously changing market environment, Pricing has
Survival Outcome Prediction for Cancer Patients based on Gene Interaction Network Analysis and Expression Profile Classification
 Survival Outcome Prediction for Cancer Patients based on Gene Interaction Network Analysis and Expression Profile Classification Final Project Report Alexander Herrmann Advised by Dr. Andrew Gentles December
Survival Outcome Prediction for Cancer Patients based on Gene Interaction Network Analysis and Expression Profile Classification Final Project Report Alexander Herrmann Advised by Dr. Andrew Gentles December
Statistical analysis on microarray data: selection of gene
 Statistical analysis on microarray data: selection of gene prognosis signatures Kim-Anh Lê Cao 1 and Geoffrey J. McLachlan 2 1 ARC Centre of Excellence in Bioinformatics, Institute for Molecular Bioscience,
Statistical analysis on microarray data: selection of gene prognosis signatures Kim-Anh Lê Cao 1 and Geoffrey J. McLachlan 2 1 ARC Centre of Excellence in Bioinformatics, Institute for Molecular Bioscience,
Expression summarization
 Expression Quantification: Affy Affymetrix Genechip is an oligonucleotide array consisting of a several perfect match (PM) and their corresponding mismatch (MM) probes that interrogate for a single gene.
Expression Quantification: Affy Affymetrix Genechip is an oligonucleotide array consisting of a several perfect match (PM) and their corresponding mismatch (MM) probes that interrogate for a single gene.
Credit Card Marketing Classification Trees
 Credit Card Marketing Classification Trees From Building Better Models With JMP Pro, Chapter 6, SAS Press (2015). Grayson, Gardner and Stephens. Used with permission. For additional information, see community.jmp.com/docs/doc-7562.
Credit Card Marketing Classification Trees From Building Better Models With JMP Pro, Chapter 6, SAS Press (2015). Grayson, Gardner and Stephens. Used with permission. For additional information, see community.jmp.com/docs/doc-7562.
advanced analysis of gene expression microarray data aidong zhang World Scientific State University of New York at Buffalo, USA
 advanced analysis of gene expression microarray data aidong zhang State University of New York at Buffalo, USA World Scientific NEW JERSEY LONDON SINGAPORE BEIJING SHANGHAI HONG KONG TAIPEI CHENNAI Contents
advanced analysis of gene expression microarray data aidong zhang State University of New York at Buffalo, USA World Scientific NEW JERSEY LONDON SINGAPORE BEIJING SHANGHAI HONG KONG TAIPEI CHENNAI Contents
Gene Expression Data Analysis
 Gene Expression Data Analysis Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu BMIF 310, Fall 2009 Gene expression technologies (summary) Hybridization-based
Gene Expression Data Analysis Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu BMIF 310, Fall 2009 Gene expression technologies (summary) Hybridization-based
Explicating a biological basis for Chronic Fatigue Syndrome
 Explicating a biological basis for Chronic Fatigue Syndrome by Samar A. Abou-Gouda A thesis submitted to the School of Computing in conformity with the requirements for the degree of Master of Science
Explicating a biological basis for Chronic Fatigue Syndrome by Samar A. Abou-Gouda A thesis submitted to the School of Computing in conformity with the requirements for the degree of Master of Science
Using Decision Tree to predict repeat customers
 Using Decision Tree to predict repeat customers Jia En Nicholette Li Jing Rong Lim Abstract We focus on using feature engineering and decision trees to perform classification and feature selection on the
Using Decision Tree to predict repeat customers Jia En Nicholette Li Jing Rong Lim Abstract We focus on using feature engineering and decision trees to perform classification and feature selection on the
PREDICTION OF PIPE PERFORMANCE WITH MACHINE LEARNING USING R
 PREDICTION OF PIPE PERFORMANCE WITH MACHINE LEARNING USING R Name: XXXXXX Student Number: XXXXXXX 2016-11-29 1. Instruction As one of the most important infrastructures in cities, water mains buried underground
PREDICTION OF PIPE PERFORMANCE WITH MACHINE LEARNING USING R Name: XXXXXX Student Number: XXXXXXX 2016-11-29 1. Instruction As one of the most important infrastructures in cities, water mains buried underground
JMP Discovery Summit 2012
 JMP Discovery Summit 2012 New Methods for Developing Limited Data Sets for Predicting US Marine Corps Combat Losses Presented to JMP Discovery Summit 2012 SAS World Headquarters, Cary, NC Topic: Predictive
JMP Discovery Summit 2012 New Methods for Developing Limited Data Sets for Predicting US Marine Corps Combat Losses Presented to JMP Discovery Summit 2012 SAS World Headquarters, Cary, NC Topic: Predictive
Bioinformatics for Biologists
 Bioinformatics for Biologists Functional Genomics: Microarray Data Analysis Fran Lewitter, Ph.D. Head, Biocomputing Whitehead Institute Outline Introduction Working with microarray data Normalization Analysis
Bioinformatics for Biologists Functional Genomics: Microarray Data Analysis Fran Lewitter, Ph.D. Head, Biocomputing Whitehead Institute Outline Introduction Working with microarray data Normalization Analysis
Analysis of Microarray Data
 Analysis of Microarray Data Lecture 1: Experimental Design and Data Normalization George Bell, Ph.D. Senior Bioinformatics Scientist Bioinformatics and Research Computing Whitehead Institute Outline Introduction
Analysis of Microarray Data Lecture 1: Experimental Design and Data Normalization George Bell, Ph.D. Senior Bioinformatics Scientist Bioinformatics and Research Computing Whitehead Institute Outline Introduction
Progress Report: Predicting Which Recommended Content Users Click Stanley Jacob, Lingjie Kong
 Progress Report: Predicting Which Recommended Content Users Click Stanley Jacob, Lingjie Kong Machine learning models can be used to predict which recommended content users will click on a given website.
Progress Report: Predicting Which Recommended Content Users Click Stanley Jacob, Lingjie Kong Machine learning models can be used to predict which recommended content users will click on a given website.
A computational pipeline for the development of multi-marker bio-signature panels and ensemble classifiers. Günther et al.
 A computational pipeline for the development of multi-marker bio-signature panels and ensemble s Günther et al. Günther et al. BMC Bioinformatics 2012, 13:326 Günther et al. BMC Bioinformatics 2012, 13:326
A computational pipeline for the development of multi-marker bio-signature panels and ensemble s Günther et al. Günther et al. BMC Bioinformatics 2012, 13:326 Günther et al. BMC Bioinformatics 2012, 13:326
A random forests approach to prioritize Highway Safety Manual (HSM) variables for data collection
 JOURNAL OF ADVANCED TRANSPORTATION J. Adv. Transp. (2016); 50:522 540 Published online 29 December 2015 in Wiley Online Library (wileyonlinelibrary.com)..1358 A random forests approach to prioritize Highway
JOURNAL OF ADVANCED TRANSPORTATION J. Adv. Transp. (2016); 50:522 540 Published online 29 December 2015 in Wiley Online Library (wileyonlinelibrary.com)..1358 A random forests approach to prioritize Highway
From Fraud Analytics Using Descriptive, Predictive, and Social Network Techniques. Full book available for purchase here.
 From Fraud Analytics Using Descriptive, Predictive, and Social Network Techniques. Full book available for purchase here. Contents List of Figures xv Foreword xxiii Preface xxv Acknowledgments xxix Chapter
From Fraud Analytics Using Descriptive, Predictive, and Social Network Techniques. Full book available for purchase here. Contents List of Figures xv Foreword xxiii Preface xxv Acknowledgments xxix Chapter
Identification of biological themes in microarray data from a mouse heart development time series using GeneSifter
 Identification of biological themes in microarray data from a mouse heart development time series using GeneSifter VizX Labs, LLC Seattle, WA 98119 Abstract Oligonucleotide microarrays were used to study
Identification of biological themes in microarray data from a mouse heart development time series using GeneSifter VizX Labs, LLC Seattle, WA 98119 Abstract Oligonucleotide microarrays were used to study
Forecasting GDP Growth, or How Can Random Forests Improve Predictions in Economics?
 Bachelor Thesis Statistics Forecasting GDP Growth, or How Can Random Forests Improve Predictions in Economics? Nils Adriansson & Ingrid Mattsson February, 2015 Supervisor: Måns Thulin UPPSALA UNIVERSITY
Bachelor Thesis Statistics Forecasting GDP Growth, or How Can Random Forests Improve Predictions in Economics? Nils Adriansson & Ingrid Mattsson February, 2015 Supervisor: Måns Thulin UPPSALA UNIVERSITY
Indicator Development for Forest Governance. IFRI Presentation
 Indicator Development for Forest Governance IFRI Presentation Plan for workshop: 4 parts Creating useful and reliable indicators IFRI research and data Results of analysis Monitoring to improve interventions
Indicator Development for Forest Governance IFRI Presentation Plan for workshop: 4 parts Creating useful and reliable indicators IFRI research and data Results of analysis Monitoring to improve interventions
Illuminating Genetic Networks with Random Forest
 ? Illuminating Genetic Networks with Random Forest ANDREAS BEYER University of Cologne Outline Random Forest Applications QTL mapping Epistasis (analyzing model structure) 2 Random Forest HOW DOES IT WORK?
? Illuminating Genetic Networks with Random Forest ANDREAS BEYER University of Cologne Outline Random Forest Applications QTL mapping Epistasis (analyzing model structure) 2 Random Forest HOW DOES IT WORK?
Outline. Analysis of Microarray Data. Most important design question. General experimental issues
 Outline Analysis of Microarray Data Lecture 1: Experimental Design and Data Normalization Introduction to microarrays Experimental design Data normalization Other data transformation Exercises George Bell,
Outline Analysis of Microarray Data Lecture 1: Experimental Design and Data Normalization Introduction to microarrays Experimental design Data normalization Other data transformation Exercises George Bell,
Bioinformatics. Microarrays: designing chips, clustering methods. Fran Lewitter, Ph.D. Head, Biocomputing Whitehead Institute
 Bioinformatics Microarrays: designing chips, clustering methods Fran Lewitter, Ph.D. Head, Biocomputing Whitehead Institute Course Syllabus Jan 7 Jan 14 Jan 21 Jan 28 Feb 4 Feb 11 Feb 18 Feb 25 Sequence
Bioinformatics Microarrays: designing chips, clustering methods Fran Lewitter, Ph.D. Head, Biocomputing Whitehead Institute Course Syllabus Jan 7 Jan 14 Jan 21 Jan 28 Feb 4 Feb 11 Feb 18 Feb 25 Sequence
An Implementation of genetic algorithm based feature selection approach over medical datasets
 An Implementation of genetic algorithm based feature selection approach over medical s Dr. A. Shaik Abdul Khadir #1, K. Mohamed Amanullah #2 #1 Research Department of Computer Science, KhadirMohideen College,
An Implementation of genetic algorithm based feature selection approach over medical s Dr. A. Shaik Abdul Khadir #1, K. Mohamed Amanullah #2 #1 Research Department of Computer Science, KhadirMohideen College,
AFFYMETRIX c Technology and Preprocessing Methods
 Analysis of Genomic and Proteomic Data AFFYMETRIX c Technology and Preprocessing Methods bhaibeka@ulb.ac.be Université Libre de Bruxelles Institut Jules Bordet Table of Contents AFFYMETRIX c Technology
Analysis of Genomic and Proteomic Data AFFYMETRIX c Technology and Preprocessing Methods bhaibeka@ulb.ac.be Université Libre de Bruxelles Institut Jules Bordet Table of Contents AFFYMETRIX c Technology
Supervised Learning Using Artificial Prediction Markets
 Supervised Learning Using Artificial Prediction Markets Adrian Barbu Department of Statistics Florida State University Joint work with Nathan Lay, FSU Dept. of Scientific Computing 1 Main Contributions
Supervised Learning Using Artificial Prediction Markets Adrian Barbu Department of Statistics Florida State University Joint work with Nathan Lay, FSU Dept. of Scientific Computing 1 Main Contributions
Random Forests. Parametrization, Tree Selection and Dynamic Induction
 Random Forests Parametrization, Tree Selection and Dynamic Induction Simon Bernard Document and Learning research team LITIS lab. University of Rouen, France décembre 2014 Random Forest Classifiers Random
Random Forests Parametrization, Tree Selection and Dynamic Induction Simon Bernard Document and Learning research team LITIS lab. University of Rouen, France décembre 2014 Random Forest Classifiers Random
Analysis of Microarray Data
 Analysis of Microarray Data Lecture 1: Experimental Design and Data Normalization George Bell, Ph.D. Senior Bioinformatics Scientist Bioinformatics and Research Computing Whitehead Institute Outline Introduction
Analysis of Microarray Data Lecture 1: Experimental Design and Data Normalization George Bell, Ph.D. Senior Bioinformatics Scientist Bioinformatics and Research Computing Whitehead Institute Outline Introduction
Knowledge-Guided Analysis with KnowEnG Lab
 Han Sinha Song Weinshilboum Knowledge-Guided Analysis with KnowEnG Lab KnowEnG Center Powerpoint by Charles Blatti Knowledge-Guided Analysis KnowEnG Center 2017 1 Exercise In this exercise we will be doing
Han Sinha Song Weinshilboum Knowledge-Guided Analysis with KnowEnG Lab KnowEnG Center Powerpoint by Charles Blatti Knowledge-Guided Analysis KnowEnG Center 2017 1 Exercise In this exercise we will be doing
Microarray Data Analysis. Normalization
 Microarray Data Analysis Normalization Outline General issues Normalization for two colour microarrays Normalization and other stuff for one color microarrays 2 Preprocessing: normalization The word normalization
Microarray Data Analysis Normalization Outline General issues Normalization for two colour microarrays Normalization and other stuff for one color microarrays 2 Preprocessing: normalization The word normalization
Sales prediction in online banking
 Sales prediction in online banking Mathias Iden Erlend S. Stenberg Master of Science in Computer Science Submission date: June 2018 Supervisor: Jon Atle Gulla, IDI Norwegian University of Science and Technology
Sales prediction in online banking Mathias Iden Erlend S. Stenberg Master of Science in Computer Science Submission date: June 2018 Supervisor: Jon Atle Gulla, IDI Norwegian University of Science and Technology
Preprocessing Methods for Two-Color Microarray Data
 Preprocessing Methods for Two-Color Microarray Data 1/15/2011 Copyright 2011 Dan Nettleton Preprocessing Steps Background correction Transformation Normalization Summarization 1 2 What is background correction?
Preprocessing Methods for Two-Color Microarray Data 1/15/2011 Copyright 2011 Dan Nettleton Preprocessing Steps Background correction Transformation Normalization Summarization 1 2 What is background correction?
Copyr i g ht 2012, SAS Ins titut e Inc. All rights res er ve d. ENTERPRISE MINER: ANALYTICAL MODEL DEVELOPMENT
 ENTERPRISE MINER: ANALYTICAL MODEL DEVELOPMENT ANALYTICAL MODEL DEVELOPMENT AGENDA Enterprise Miner: Analytical Model Development The session looks at: - Supervised and Unsupervised Modelling - Classification
ENTERPRISE MINER: ANALYTICAL MODEL DEVELOPMENT ANALYTICAL MODEL DEVELOPMENT AGENDA Enterprise Miner: Analytical Model Development The session looks at: - Supervised and Unsupervised Modelling - Classification
CS4220: Knowledge Discovery Methods for Bioinformatics Unit 4: Batch Effects. Wong Limsoon
 : Knowledge Discovery Methods for Bioinformatics Unit 4: Batch Effects Wong Limsoon 2 Plan Batch effects Visualization Normalization PC1 removal Batch effect-resistant feature selection Batch effect-resistant
: Knowledge Discovery Methods for Bioinformatics Unit 4: Batch Effects Wong Limsoon 2 Plan Batch effects Visualization Normalization PC1 removal Batch effect-resistant feature selection Batch effect-resistant
Euro area GDP forecasting using large survey datasets
 Euro area GDP forecasting using large survey datasets A random forest approach Olivier Biau, Angela D'Elia Directorate General for Financial and Economic Affairs - European Commission * Abstract Recent
Euro area GDP forecasting using large survey datasets A random forest approach Olivier Biau, Angela D'Elia Directorate General for Financial and Economic Affairs - European Commission * Abstract Recent
An Innovative System of Disaggregate Models for Trip Generation and Trip Attributes Using Random Forest Concept
 An Innovative System of Disaggregate Models for Trip Generation and Trip Attributes Using Random Forest Concept Milad Ghasri, Taha Hossein Rashidi Abstract This research attempts to address the gap between
An Innovative System of Disaggregate Models for Trip Generation and Trip Attributes Using Random Forest Concept Milad Ghasri, Taha Hossein Rashidi Abstract This research attempts to address the gap between
Exam 1 from a Past Semester
 Exam from a Past Semester. Provide a brief answer to each of the following questions. a) What do perfect match and mismatch mean in the context of Affymetrix GeneChip technology? Be as specific as possible
Exam from a Past Semester. Provide a brief answer to each of the following questions. a) What do perfect match and mismatch mean in the context of Affymetrix GeneChip technology? Be as specific as possible
Towards personalized treatment of cancer
 Mathematical Statistics Stockholm University Towards personalized treatment of cancer Tomas Jacobsson Examensarbete 2012:3 Postal address: Mathematical Statistics Dept. of Mathematics Stockholm University
Mathematical Statistics Stockholm University Towards personalized treatment of cancer Tomas Jacobsson Examensarbete 2012:3 Postal address: Mathematical Statistics Dept. of Mathematics Stockholm University
Data Mining for Biological Data Analysis
 Data Mining for Biological Data Analysis Data Mining and Text Mining (UIC 583 @ Politecnico di Milano) References Data Mining Course by Gregory-Platesky Shapiro available at www.kdnuggets.com Jiawei Han
Data Mining for Biological Data Analysis Data Mining and Text Mining (UIC 583 @ Politecnico di Milano) References Data Mining Course by Gregory-Platesky Shapiro available at www.kdnuggets.com Jiawei Han
Normalization. Getting the numbers comparable. DNA Microarray Bioinformatics - #27612
 Normalization Getting the numbers comparable The DNA Array Analysis Pipeline Question Experimental Design Array design Probe design Sample Preparation Hybridization Buy Chip/Array Image analysis Expression
Normalization Getting the numbers comparable The DNA Array Analysis Pipeline Question Experimental Design Array design Probe design Sample Preparation Hybridization Buy Chip/Array Image analysis Expression
Lab 1: A review of linear models
 Lab 1: A review of linear models The purpose of this lab is to help you review basic statistical methods in linear models and understanding the implementation of these methods in R. In general, we need
Lab 1: A review of linear models The purpose of this lab is to help you review basic statistical methods in linear models and understanding the implementation of these methods in R. In general, we need
Unravelling Airbnb Predicting Price for New Listing
 Unravelling Airbnb Predicting Price for New Listing Paridhi Choudhary H John Heinz III College Carnegie Mellon University Pittsburgh, PA 15213 paridhic@andrew.cmu.edu Aniket Jain H John Heinz III College
Unravelling Airbnb Predicting Price for New Listing Paridhi Choudhary H John Heinz III College Carnegie Mellon University Pittsburgh, PA 15213 paridhic@andrew.cmu.edu Aniket Jain H John Heinz III College
Machine Learning Models for Sales Time Series Forecasting
 Article Machine Learning Models for Sales Time Series Forecasting Bohdan M. Pavlyshenko SoftServe, Inc., Ivan Franko National University of Lviv * Correspondence: bpavl@softserveinc.com, b.pavlyshenko@gmail.com
Article Machine Learning Models for Sales Time Series Forecasting Bohdan M. Pavlyshenko SoftServe, Inc., Ivan Franko National University of Lviv * Correspondence: bpavl@softserveinc.com, b.pavlyshenko@gmail.com
Preface to the third edition Preface to the first edition Acknowledgments
 Contents Foreword Preface to the third edition Preface to the first edition Acknowledgments Part I PRELIMINARIES XXI XXIII XXVII XXIX CHAPTER 1 Introduction 3 1.1 What Is Business Analytics?................
Contents Foreword Preface to the third edition Preface to the first edition Acknowledgments Part I PRELIMINARIES XXI XXIII XXVII XXIX CHAPTER 1 Introduction 3 1.1 What Is Business Analytics?................
Data Mining Applications with R
 Data Mining Applications with R Yanchang Zhao Senior Data Miner, RDataMining.com, Australia Associate Professor, Yonghua Cen Nanjing University of Science and Technology, China AMSTERDAM BOSTON HEIDELBERG
Data Mining Applications with R Yanchang Zhao Senior Data Miner, RDataMining.com, Australia Associate Professor, Yonghua Cen Nanjing University of Science and Technology, China AMSTERDAM BOSTON HEIDELBERG
Forecasting Seasonal Footwear Demand Using Machine Learning. By Majd Kharfan & Vicky Chan, SCM 2018 Advisor: Tugba Efendigil
 Forecasting Seasonal Footwear Demand Using Machine Learning By Majd Kharfan & Vicky Chan, SCM 2018 Advisor: Tugba Efendigil 1 Agenda Ø Ø Ø Ø Ø Ø Ø The State Of Fashion Industry Research Objectives AI In
Forecasting Seasonal Footwear Demand Using Machine Learning By Majd Kharfan & Vicky Chan, SCM 2018 Advisor: Tugba Efendigil 1 Agenda Ø Ø Ø Ø Ø Ø Ø The State Of Fashion Industry Research Objectives AI In
Machine Learning Techniques For Particle Identification
 Machine Learning Techniques For Particle Identification 06.March.2018 I Waleed Esmail, Tobias Stockmanns, Michael Kunkel, James Ritman Institut für Kernphysik (IKP), Forschungszentrum Jülich Outlines:
Machine Learning Techniques For Particle Identification 06.March.2018 I Waleed Esmail, Tobias Stockmanns, Michael Kunkel, James Ritman Institut für Kernphysik (IKP), Forschungszentrum Jülich Outlines:
Mixture modeling for genome-wide localization of transcription factors
 Mixture modeling for genome-wide localization of transcription factors Sündüz Keleş 1,2 and Heejung Shim 1 1 Department of Statistics 2 Department of Biostatistics & Medical Informatics University of Wisconsin,
Mixture modeling for genome-wide localization of transcription factors Sündüz Keleş 1,2 and Heejung Shim 1 1 Department of Statistics 2 Department of Biostatistics & Medical Informatics University of Wisconsin,
Ensemble Modeling. Toronto Data Mining Forum November 2017 Helen Ngo
 Ensemble Modeling Toronto Data Mining Forum November 2017 Helen Ngo Agenda Introductions Why Ensemble Models? Simple & Complex ensembles Thoughts: Post-real-life Experimentation Downsides of Ensembles
Ensemble Modeling Toronto Data Mining Forum November 2017 Helen Ngo Agenda Introductions Why Ensemble Models? Simple & Complex ensembles Thoughts: Post-real-life Experimentation Downsides of Ensembles
Prospects for Economics in the Machine Learning and Big Data Era
 Prospects for Economics in the Machine Learning and Big Data Era SEPT 29, 2018 WILLIAM CHOW Introduction 2 Is Big Data and the associated treatment of it a fad? No. Because the economics profession has
Prospects for Economics in the Machine Learning and Big Data Era SEPT 29, 2018 WILLIAM CHOW Introduction 2 Is Big Data and the associated treatment of it a fad? No. Because the economics profession has
Applications of Machine Learning to Predict Yelp Ratings
 Applications of Machine Learning to Predict Yelp Ratings Kyle Carbon Aeronautics and Astronautics kcarbon@stanford.edu Kacyn Fujii Electrical Engineering khfujii@stanford.edu Prasanth Veerina Computer
Applications of Machine Learning to Predict Yelp Ratings Kyle Carbon Aeronautics and Astronautics kcarbon@stanford.edu Kacyn Fujii Electrical Engineering khfujii@stanford.edu Prasanth Veerina Computer
Supplementary Material. Model-specification uncertainty in future area burned by wildfires in Canada
 1 10.1071/WF17123_AC CSIRO 2018 Supplementary Material: International Journal of Wildland Fire, 27(3), 164 175. Supplementary Material Model-specification uncertainty in future area burned by wildfires
1 10.1071/WF17123_AC CSIRO 2018 Supplementary Material: International Journal of Wildland Fire, 27(3), 164 175. Supplementary Material Model-specification uncertainty in future area burned by wildfires
Decision Trees And Random Forests A Visual Introduction For Beginners
 Decision Trees And Random Forests A Visual Introduction For Beginners We have made it easy for you to find a PDF Ebooks without any digging. And by having access to our ebooks online or by storing it on
Decision Trees And Random Forests A Visual Introduction For Beginners We have made it easy for you to find a PDF Ebooks without any digging. And by having access to our ebooks online or by storing it on
Predicting the profitability level of companies regarding the five comparability factors
 VU University Amsterdam MSc. Business Analytics Research Paper Predicting the profitability level of companies regarding the five comparability factors March 31, 2017 Manon Wintgens (2558262) Manon Wintgens
VU University Amsterdam MSc. Business Analytics Research Paper Predicting the profitability level of companies regarding the five comparability factors March 31, 2017 Manon Wintgens (2558262) Manon Wintgens
Dallas J. Elgin, Ph.D. IMPAQ International Randi Walters, Ph.D. Casey Family Programs APPAM Fall Research Conference
 Utilizing Predictive Modeling to Improve Policy through Improved Targeting of Agency Resources: A Case Study on Placement Instability among Foster Children Dallas J. Elgin, Ph.D. IMPAQ International Randi
Utilizing Predictive Modeling to Improve Policy through Improved Targeting of Agency Resources: A Case Study on Placement Instability among Foster Children Dallas J. Elgin, Ph.D. IMPAQ International Randi
A STUDY ON STATISTICAL BASED FEATURE SELECTION METHODS FOR CLASSIFICATION OF GENE MICROARRAY DATASET
 A STUDY ON STATISTICAL BASED FEATURE SELECTION METHODS FOR CLASSIFICATION OF GENE MICROARRAY DATASET 1 J.JEYACHIDRA, M.PUNITHAVALLI, 1 Research Scholar, Department of Computer Science and Applications,
A STUDY ON STATISTICAL BASED FEATURE SELECTION METHODS FOR CLASSIFICATION OF GENE MICROARRAY DATASET 1 J.JEYACHIDRA, M.PUNITHAVALLI, 1 Research Scholar, Department of Computer Science and Applications,
Predictive Analytics
 Predictive Analytics Mani Janakiram, PhD Director, Supply Chain Intelligence & Analytics, Intel Corp. Adjunct Professor of Supply Chain, ASU October 2017 "Prediction is very difficult, especially if it's
Predictive Analytics Mani Janakiram, PhD Director, Supply Chain Intelligence & Analytics, Intel Corp. Adjunct Professor of Supply Chain, ASU October 2017 "Prediction is very difficult, especially if it's
FINAL PROJECT REPORT IME672. Group Number 6
 FINAL PROJECT REPORT IME672 Group Number 6 Ayushya Agarwal 14168 Rishabh Vaish 14553 Rohit Bansal 14564 Abhinav Sharma 14015 Dil Bag Singh 14222 Introduction Cell2Cell, The Churn Game. The cellular telephone
FINAL PROJECT REPORT IME672 Group Number 6 Ayushya Agarwal 14168 Rishabh Vaish 14553 Rohit Bansal 14564 Abhinav Sharma 14015 Dil Bag Singh 14222 Introduction Cell2Cell, The Churn Game. The cellular telephone
Rank hotels on Expedia.com to maximize purchases
 Rank hotels on Expedia.com to maximize purchases Nishith Khantal, Valentina Kroshilina, Deepak Maini December 14, 2013 1 Introduction For an online travel agency (OTA), matching users to hotel inventory
Rank hotels on Expedia.com to maximize purchases Nishith Khantal, Valentina Kroshilina, Deepak Maini December 14, 2013 1 Introduction For an online travel agency (OTA), matching users to hotel inventory
Learning how people respond to changes in energy prices
 Learning how people respond to changes in energy prices Ben Lim bclimatestanford.edu Jonathan Roth jmrothstanford.edu Andrew Sonta asontastanford.edu Abstract We investigate the problem of predicting day-ahead
Learning how people respond to changes in energy prices Ben Lim bclimatestanford.edu Jonathan Roth jmrothstanford.edu Andrew Sonta asontastanford.edu Abstract We investigate the problem of predicting day-ahead
{saharonr, lastgift>35
 KDD-Cup 99 : Knowledge Discovery In a Charitable Organization s Donor Database Saharon Rosset and Aron Inger Amdocs (Israel) Ltd. 8 Hapnina St. Raanana, Israel, 43000 {saharonr, aroni}@amdocs.com 1. INTRODUCTION
KDD-Cup 99 : Knowledge Discovery In a Charitable Organization s Donor Database Saharon Rosset and Aron Inger Amdocs (Israel) Ltd. 8 Hapnina St. Raanana, Israel, 43000 {saharonr, aroni}@amdocs.com 1. INTRODUCTION
Gene Set Enrichment Analysis
 Gene Set Enrichment Analysis Martin Morgan Fred Hutchinson Cancer Research Center Seattle, WA, USA 28 April 2009 Motivation Many analyses: Exploratory, even in designed experiments: which of 1000 s of
Gene Set Enrichment Analysis Martin Morgan Fred Hutchinson Cancer Research Center Seattle, WA, USA 28 April 2009 Motivation Many analyses: Exploratory, even in designed experiments: which of 1000 s of
Identification of SNP Interactions Using Logic Regression
 Identification of SNP Interactions Using Logic Regression Holger Schwender and Katja Ickstadt Collaborative Research Center 475 Department of Statistics University of Dortmund holger.schwender@udo.edu
Identification of SNP Interactions Using Logic Regression Holger Schwender and Katja Ickstadt Collaborative Research Center 475 Department of Statistics University of Dortmund holger.schwender@udo.edu
Bioinformatics : Gene Expression Data Analysis
 05.12.03 Bioinformatics : Gene Expression Data Analysis Aidong Zhang Professor Computer Science and Engineering What is Bioinformatics Broad Definition The study of how information technologies are used
05.12.03 Bioinformatics : Gene Expression Data Analysis Aidong Zhang Professor Computer Science and Engineering What is Bioinformatics Broad Definition The study of how information technologies are used
Approaches for Identifying Consumer Preferences for the Design of Technology Products: A Case Study of Residential Solar Panels
 Approaches for Identifying Consumer Preferences for the Design of Technology Products: A Case Study of Residential Solar Panels Heidi Q. Chen Graduate Student Mechanical Engineering Massachusetts Institute
Approaches for Identifying Consumer Preferences for the Design of Technology Products: A Case Study of Residential Solar Panels Heidi Q. Chen Graduate Student Mechanical Engineering Massachusetts Institute
2015 The MathWorks, Inc. 1
 2015 The MathWorks, Inc. 1 MATLAB 을이용한머신러닝 ( 기본 ) Senior Application Engineer 엄준상과장 2015 The MathWorks, Inc. 2 Machine Learning is Everywhere Solution is too complex for hand written rules or equations
2015 The MathWorks, Inc. 1 MATLAB 을이용한머신러닝 ( 기본 ) Senior Application Engineer 엄준상과장 2015 The MathWorks, Inc. 2 Machine Learning is Everywhere Solution is too complex for hand written rules or equations
Detecting gene gene interactions that underlie human diseases
 Genome-wide association studies Detecting gene gene interactions that underlie human diseases Heather J. Cordell Abstract Following the identification of several disease-associated polymorphisms by genome-wide
Genome-wide association studies Detecting gene gene interactions that underlie human diseases Heather J. Cordell Abstract Following the identification of several disease-associated polymorphisms by genome-wide
Detecting gene gene interactions that underlie human diseases
 REVIEWS Genome-wide association studies Detecting gene gene interactions that underlie human diseases Heather J. Cordell Abstract Following the identification of several disease-associated polymorphisms
REVIEWS Genome-wide association studies Detecting gene gene interactions that underlie human diseases Heather J. Cordell Abstract Following the identification of several disease-associated polymorphisms
science and applications
 Introduction to data science and applications 1 What s possible with data and analytics? 2 Facebook can predict break-ups? http://www.huffingtonpost.com/2014/02/14/facebook-relationship-study_n_4784291.html
Introduction to data science and applications 1 What s possible with data and analytics? 2 Facebook can predict break-ups? http://www.huffingtonpost.com/2014/02/14/facebook-relationship-study_n_4784291.html
USING R IN SAS ENTERPRISE MINER EDMONTON USER GROUP
 USING R IN SAS ENTERPRISE MINER EDMONTON USER GROUP INTRODUCTION PAT VALENTE, MA Solution Specialist, Data Sciences at SAS. Training in Economics and Statistics. 20 years experience in business areas including
USING R IN SAS ENTERPRISE MINER EDMONTON USER GROUP INTRODUCTION PAT VALENTE, MA Solution Specialist, Data Sciences at SAS. Training in Economics and Statistics. 20 years experience in business areas including
