MODEL 1. provides additional information about the performance of the CeSS algorithm.
|
|
|
- Godfrey Pope
- 6 years ago
- Views:
Transcription
1 MODEL 1 Sections 1 to 4 describe Model 1, a metabolic model of E. coli s Central Carbon Metabolism. Section 5 provides additional information about the performance of the CeSS algorithm. 1 Model structure: metabolites and reactions 1.1 Metabolite list Last column: 1 for external metabolites, 0 for internal. # Metabolite Name KEGG ID Ext. / Int. 1 Pyruvate C Thiamin diphosphat C (alpha-Hydroxyethyl)thiamine diphosphat C CO2 C ATP C H2O C AMP C Phosphoenolpyruvat C Orthophosphat C ADP C Acetyl-CoA C Formate C CoA C (S)-Malate C NAD+ C NADH C H+ C NADP+ C NADPH C Acetaldehyd C Acetyl phosphat C Acetate C Diphosphat C L-Glutamat C Oxoglutarat C NH3 C Oxalosuccinat C Oxaloacetat C Citrate C Succinate C Succinyl-Co C Acceptor C Fumarate C Reduced accepto C Glyoxylate C Isocitrate C Carboxy-1-hydroxypropyl-ThP C Phospho-D-glycerat C
2 39 (S)-Lactat C Ethanol C D-Glyceraldehyde 3-phosphat C Glycerone phosphat C D-Ribose 5-phosphat C Phospho-alpha-D-ribose 1-diphosphat C D-Ribulose 5-phosphat C Phospho-D-glyceroyl phosphat C beta-d-fructose 1,6-bisphosphat C cis-aconitat C Phospho-D-glycerat C Phospho-D-gluconat C D-Xylulose 5-phosphat C beta-d-glucose C beta-d-glucose 6-phosphat C Sedoheptulose 7-phosphat C Dihydrolipoamid C Lipoamide C D-Erythrose 4-phosphat C beta-d-fructose 6-phosphat C D-Glucono-1,5-lactone 6-phosphat C Dehydro-3-deoxy-6-phospho-D-gluconat C Enzyme N6-(dihydrolipoyl)lysin C (Dihydrolipoyllysine-residue acetyltransferase)-sacetyldihydrolipoyllysin C (Dihydrolipoyllysine-residue succinyltransferase)- C S-succinyldihydrolipoyllysin 64 Ferricytochrome b C Ferrocytochrome b C Enzyme N6-(lipoyl)lysin C sn-glycerol 3-phosphat C Glycerol C GlycerolEx 1 70 beta-d-glucoseex 1 71 GTP C GDP C O2 C Biomass Reaction list # Formula KEGG ID 1 Pyruvate + Thiamin diphosphate 2-(alpha-Hydroxyethyl)thiamine diphosphate + CO2 R Pyruvate + ATP + H2O AMP + Phosphoenolpyruvate + Orthophosphate R Pyruvate + ATP Phosphoenolpyruvate + ADP R Acetyl-CoA + Formate Pyruvate + CoA R (S)-Malate + NAD+ Pyruvate + CO2 + NADH + H+ R (S)-Malate + NADP+ Pyruvate + CO2 + H+ + NADPH R CoA + NAD+ + Acetaldehyde Acetyl-CoA + NADH + H+ R Orthophosphate + Acetyl-CoA CoA + Acetyl phosphate R
3 9 ATP + CoA + Acetate AMP + Acetyl-CoA + Diphosphate R H2O + NAD+ + L-Glutamate NADH + H+ + 2-Oxoglutarate + NH3 R Oxalosuccinate CO2 + 2-Oxoglutarate R ATP + Acetate ADP + Acetyl phosphate R ATP + Oxaloacetate CO2 + Phosphoenolpyruvate + ADP R (S)-Malate + NAD+ NADH + H+ + Oxaloacetate R CoA + Citrate H2O + Acetyl-CoA + Oxaloacetate R Citrate Acetate + Oxaloacetate R ATP + CoA + Succinate Orthophosphate + ADP + Succinyl-CoA R Succinate + Acceptor Fumarate + Reduced acceptor R CoA + (S)-Malate H2O + Acetyl-CoA + Glyoxylate R Isocitrate Succinate + Glyoxylate R Thiamin diphosphate + 2-Oxoglutarate CO2 + 3-Carboxy-1-hydroxypropyl-ThPP R Phospho-D-glycerate H2O + Phosphoenolpyruvate R NAD+ + (S)-Lactate Pyruvate + NADH + H+ R NAD+ + Ethanol NADH + H+ + Acetaldehyde R D-Glyceraldehyde 3-phosphate Glycerone phosphate R ATP + D-Ribose 5-phosphate AMP + 5-Phospho-alpha-D-ribose 1-diphosphate R D-Ribose 5-phosphate D-Ribulose 5-phosphate R Orthophosphate + NAD+ + D-Glyceraldehyde 3-phosphate NADH + H+ + 3-Phospho- R01061 D-glyceroyl phosphate 29 beta-d-fructose 1,6-bisphosphate D-Glyceraldehyde 3-phosphate + Glycerone phosphate R (S)-Malate H2O + Fumarate R Citrate Isocitrate R Citrate H2O + cis-aconitate R ATP + 3-Phospho-D-glycerate ADP + 3-Phospho-D-glyceroyl phosphate R Phospho-D-glycerate 3-Phospho-D-glycerate R NADP+ + 6-Phospho-D-gluconate CO2 + H+ + NADPH + D-Ribulose 5-phosphate R D-Ribulose 5-phosphate D-Xylulose 5-phosphate R ATP + beta-d-glucose ADP + beta-d-glucose 6-phosphate R D-Glyceraldehyde 3-phosphate + Sedoheptulose 7-phosphate D-Ribose 5-phosphate + R01641 D-Xylulose 5-phosphate 39 NAD+ + Dihydrolipoamide NADH + H+ + Lipoamide R D-Glyceraldehyde 3-phosphate + Sedoheptulose 7-phosphate D-Erythrose 4-phosphate R beta-d-fructose 6-phosphate 41 D-Glyceraldehyde 3-phosphate + beta-d-fructose 6-phosphate D-Xylulose 5-phosphate R D-Erythrose 4-phosphate 42 NADP+ + Isocitrate H+ + NADPH + Oxalosuccinate R Isocitrate H2O + cis-aconitate R H2O + D-Glucono-1,5-lactone 6-phosphate 6-Phospho-D-gluconate R Phospho-D-gluconate H2O + 2-Dehydro-3-deoxy-6-phospho-D-gluconate R Acetyl-CoA + Enzyme N6-(dihydrolipoyl)lysine CoA + [Dihydrolipoyllysine-residue R02569 acetyltransferase]s-acetyldihydrolipoyllysine 47 Succinyl-CoA + Enzyme N6-(dihydrolipoyl)lysine CoA + [Dihydrolipoyllysine-residue R02570 succinyltransferase]s-succinyldihydrolipoyllysine 48 NADP+ + beta-d-glucose 6-phosphate H+ + NADPH + D-Glucono-1,5-lactone 6- R02736 phosphate 49 Pyruvate + H2O + Ferricytochrome b1 CO2 + Acetate + Ferrocytochrome b1 R (alpha-Hydroxyethyl)thiamine diphosphate + Enzyme N6-(lipoyl)lysine Thiamin diphosphate + [Dihydrolipoyllysine-residue acetyltransferase]s-acetyldihydrolipoyllysine R
4 51 3-Carboxy-1-hydroxypropyl-ThPP + Enzyme N6-(lipoyl)lysine Thiamin diphosphate + R03316 [Dihydrolipoyllysine-residue succinyltransferase]s-succinyldihydrolipoyllysine 52 beta-d-glucose 6-phosphate beta-d-fructose 6-phosphate R ATP + beta-d-fructose 6-phosphate ADP + beta-d-fructose 1,6-bisphosphate R H2O + beta-d-fructose 1,6-bisphosphate Orthophosphate + beta-d-fructose 6- R04780 phosphate 55 2-Dehydro-3-deoxy-6-phospho-D-gluconate Pyruvate + D-Glyceraldehyde 3-phosphate R ATP + AMP 2ADP R H2O + sn-glycerol 3-phosphate Orthophosphate + Glycerol R NAD+ + sn-glycerol 3-phosphate NADH + H+ + Glycerone phosphate R GlycerolExt Glycerol 60 beta-d-glucoseext beta-d-glucose 61 NADPH NADP+ 62 ADP + GTP ATP + GDP ADP + Reduced acceptor O ATP + H2O + Acceptor ADP + NADH O ATP + H2O + NAD+ 65 ATP ADP Pyruvate CO ATP Phosphoenolpyruvate Oxoglutarate Oxaloacetate NAD Acetyl-CoA D-Glyceraldehyde-3- phosphate D-Ribose-5-phosphate beta-d-fructose-6-phosphate D- Erythrose-4-phosphate NADPH beta-d-glucose-6-phosphate O ADP NADH NADP CoA + Biomass 2 Simulated experiments The following tables describe the settings used to generate the simulated experimental data. Tables show the concentrations of enzymes and external metabolites. The initial concentrations of the internal metabolites are considered as model parameters and as such they are shown in subsection Enzyme concentrations Enzyme concentrations of the 66 reactions (rows 1 66) for 10 different simulations (columns A J). A B C D E F G H I J
5
6 2.2 External metabolites Initial concentrations of the 25 external metabolites (rows 1 25) for 10 different simulations (columns A J). A B C D E F G H I J Model parameters The following tables show the nominal values of the model parameters, p. The upper and lower bounds used in the optimizations are p L =0.1 p; p U =10 p. 3.1 Model parameters (1 490) Initial concentrations of the 49 internal metabolites (rows 1 49) for 10 different simulations (columns A J). A B C D E F G H I J
7 Model parameters ( ) Activation (k A ), Inhibition (k I ), and Michaelis-Menten (k M ) constants. The third column shows the corresponding reaction and metabolite number. Param. Type (Reaction, Param. number Metabolite) value 7
8 491 k A (3,7) k A (53,7) k A (53,10) k A (53,72) k I (54,5) k I (54,7) k I (53,8) k I (54,8) k I (54,9) k I (54,10) k I (15,16) k I (48,16) k I (15,25) k M (1,1) k M (2,1) k M (3,1) k M (4,1) k M (5,1) k M (6,1) k M (23,1) k M (49,1) k M (55,1) k M (66,1) k M (1,2) k M (21,2) k M (50,2) k M (51,2) k M (1,3) k M (50,3) k M (1,4) k M (5,4) k M (6,4) k M (11,4) k M (13,4) k M (21,4) k M (35,4) k M (49,4) k M (66,4) k M (2,5) k M (3,5) k M (9,5) k M (12,5) k M (13,5) k M (17,5) k M (26,5) k M (33,5) k M (37,5) k M (53,5) k M (56,5) k M (62,5)
9 541 k M (63,5) k M (64,5) k M (65,5) k M (66,5) k M (2,6) k M (10,6) k M (15,6) k M (19,6) k M (22,6) k M (30,6) k M (32,6) k M (43,6) k M (44,6) k M (45,6) k M (49,6) k M (54,6) k M (57,6) k M (63,6) k M (64,6) k M (2,7) k M (9,7) k M (26,7) k M (56,7) k M (2,8) k M (3,8) k M (13,8) k M (22,8) k M (66,8) k M (2,9) k M (8,9) k M (17,9) k M (28,9) k M (54,9) k M (57,9) k M (3,10) k M (12,10) k M (13,10) k M (17,10) k M (33,10) k M (37,10) k M (53,10) k M (56,10) k M (62,10) k M (63,10) k M (64,10) k M (65,10) k M (66,10) k M (4,11) k M (7,11) k M (8,11) k M (9,11)
10 592 k M (15,11) k M (19,11) k M (46,11) k M (66,11) k M (4,12) k M (4,13) k M (7,13) k M (8,13) k M (9,13) k M (15,13) k M (17,13) k M (19,13) k M (46,13) k M (47,13) k M (66,13) k M (5,14) k M (6,14) k M (14,14) k M (19,14) k M (30,14) k M (5,15) k M (7,15) k M (10,15) k M (14,15) k M (23,15) k M (24,15) k M (28,15) k M (39,15) k M (58,15) k M (64,15) k M (66,15) k M (5,16) k M (7,16) k M (10,16) k M (14,16) k M (23,16) k M (24,16) k M (28,16) k M (39,16) k M (58,16) k M (64,16) k M (66,16) k M (5,17) k M (6,17) k M (7,17) k M (10,17) k M (14,17) k M (23,17) k M (24,17) k M (28,17) k M (35,17)
11 643 k M (39,17) k M (42,17) k M (48,17) k M (58,17) k M (6,18) k M (35,18) k M (42,18) k M (48,18) k M (61,18) k M (66,18) k M (6,19) k M (35,19) k M (42,19) k M (48,19) k M (61,19) k M (66,19) k M (7,20) k M (24,20) k M (8,21) k M (12,21) k M (9,22) k M (12,22) k M (16,22) k M (49,22) k M (9,23) k M (10,24) k M (10,25) k M (11,25) k M (21,25) k M (66,25) k M (10,26) k M (11,27) k M (42,27) k M (13,28) k M (14,28) k M (15,28) k M (16,28) k M (66,28) k M (15,29) k M (16,29) k M (31,29) k M (32,29) k M (17,30) k M (18,30) k M (20,30) k M (17,31) k M (47,31) k M (18,32) k M (63,32) k M (18,33) k M (30,33)
12 694 k M (18,34) k M (63,34) k M (19,35) k M (20,35) k M (20,36) k M (31,36) k M (42,36) k M (43,36) k M (21,37) k M (51,37) k M (22,38) k M (34,38) k M (23,39) k M (24,40) k M (25,41) k M (28,41) k M (29,41) k M (38,41) k M (40,41) k M (41,41) k M (55,41) k M (66,41) k M (25,42) k M (29,42) k M (58,42) k M (26,43) k M (27,43) k M (38,43) k M (66,43) k M (26,44) k M (27,45) k M (35,45) k M (36,45) k M (28,46) k M (33,46) k M (29,47) k M (53,47) k M (54,47) k M (32,48) k M (43,48) k M (33,49) k M (34,49) k M (35,50) k M (44,50) k M (45,50) k M (36,51) k M (38,51) k M (41,51) k M (37,52) k M (60,52) k M (37,53)
13 745 k M (48,53) k M (52,53) k M (66,53) k M (38,54) k M (40,54) k M (39,55) k M (39,56) k M (40,57) k M (41,57) k M (66,57) k M (40,58) k M (41,58) k M (52,58) k M (53,58) k M (54,58) k M (66,58) k M (44,59) k M (48,59) k M (45,60) k M (55,60) k M (46,61) k M (47,61) k M (46,62) k M (50,62) k M (47,63) k M (51,63) k M (49,64) k M (49,65) k M (50,66) k M (51,66) k M (57,67) k M (58,67) k M (57,68) k M (59,68) k M (59,69) k M (60,70) k M (62,71) k M (62,72) k M (63,73) k M (64,73) k M (66,73) k M (66,74) Model parameters ( ) Velocity (k v ) and equilibrium (k eq ) constants of the reactions. React. Param. k v Param. k eq # number number
14
15 Model outputs The 115 model outputs are: Outputs 1 49: concentrations of the 49 internal metabolites listed in table 1. Outputs : concentrations of the 66 reaction fluxes listed in table 2. 5 Influence of the tuning parameters in the performance of CeSS First we test what happens when the threads exchange information at different time intervals (τ). Figure 1 (top) shows the convergence curves, which plot the logarithm of the objective function value against the computation time. The performance of 10 individual, non cooperative threads is shown in black. It is compared to the performance of 10 cooperative threads which exchange information with different values of τ: 2.53 (approximately 12 hours in the hardware used, red curve), 2.70 (18 hours, green), 2.83 (24 hours, blue), 3.13 (48 hours, magenta), and 3.30 (72 hours, cyan). For this system, the performance of the cooperative algorithm is not significantly influenced by the choice of the time between information sharing: τ can range between 2.53 and 3.13 without affecting significantly the outcome of the optimization. Next we test the influence of the other cooperative optimization parameter, the number of threads η. We fix τ = 12 hours and compare the algorithm s performance for η = 5, 10, 20, and 30 threads. Results are shown in Fig. 1 (bottom); as expected, convergence improves when the number of threads increases. However, the improvement is not linear: the objective function value is very much reduced when changing from 5 to 10 threads, but it is only slightly reduced with 20 or 30 threads. 15
16 Objective Function (logarithmic) τ = 2.53 τ = 2.70 τ = 2.83 τ = 3.13 τ = 3.30 no coop Computation Time (hours) Objective function (logarithmic) threads 7 threads 10 threads 20 threads 30 threads Computation time (hours) Figure 1: Convergence curves, Model 1. TOP: different τ, η = 10. BOTTOM: different η, τ =
ORFs, unknown function GC (%) # of ORFs. Fen-75cm Fn (91) Nitrosopumilus maritimus
 Table S1 Comparison of genome properties for 10 distinct bins identified from two metagenomes. The mean coverage is calculated by dividing total read counts with read length. ORF, open reading frame. Nearest
Table S1 Comparison of genome properties for 10 distinct bins identified from two metagenomes. The mean coverage is calculated by dividing total read counts with read length. ORF, open reading frame. Nearest
Basic modeling approaches for biological systems -II. Mahesh Bule
 Basic modeling approaches for biological systems -II Mahesh Bule Modeling Metabolism Metabolism is general term for two kind of reactions Catabolic reactions (breakdown of complex compounds to get energy
Basic modeling approaches for biological systems -II Mahesh Bule Modeling Metabolism Metabolism is general term for two kind of reactions Catabolic reactions (breakdown of complex compounds to get energy
7.05 Spring 2004 April 23, Recitation #9
 7.05 Spring 004 April 3, 004 Recitation #9 ontact Information TA: Victor Sai Recitation: Friday, 3-4pm, -13 E-mail: sai@mit.edu ffice ours: Friday, 4-5pm, -13 Unit 4 Schedule Recitation/Exam Date Topic
7.05 Spring 004 April 3, 004 Recitation #9 ontact Information TA: Victor Sai Recitation: Friday, 3-4pm, -13 E-mail: sai@mit.edu ffice ours: Friday, 4-5pm, -13 Unit 4 Schedule Recitation/Exam Date Topic
Figure S1: Time course of the energy charge in trehalose-grown wild-type and adk1 mutant cells after sudden increase of glucose concentration to 110
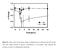 Figure S1: Time course of the energy charge in trehalose-grown wild-type and adk1 mutant cells after sudden increase of glucose concentration to 110 mmol/l. Data represent the average of at least two independent
Figure S1: Time course of the energy charge in trehalose-grown wild-type and adk1 mutant cells after sudden increase of glucose concentration to 110 mmol/l. Data represent the average of at least two independent
METABOLIC CAPABILITIES OF Actinobacillus succinogenes FOR SUCCINIC ACID PRODUCTION
 Brazilian Journal of Chemical Engineering ISSN 14-6632 Printed in Brazil www.abeq.org.br/bjche Vol. 31, No. 4, pp. 859-865, October - December, 214 dx.doi.org/1.159/14-6632.214314s2997 METABOLIC CAPABILITIES
Brazilian Journal of Chemical Engineering ISSN 14-6632 Printed in Brazil www.abeq.org.br/bjche Vol. 31, No. 4, pp. 859-865, October - December, 214 dx.doi.org/1.159/14-6632.214314s2997 METABOLIC CAPABILITIES
Metabolic Engineering for Fuels and Chemicals
 Metabolic Engineering for Fuels and Chemicals K.T. Shanmugam and Lonnie O. Ingram Dept. of Microbiology and Cell Science University of Florida Gainesville, Florida Florida Center for Renewable Chemicals
Metabolic Engineering for Fuels and Chemicals K.T. Shanmugam and Lonnie O. Ingram Dept. of Microbiology and Cell Science University of Florida Gainesville, Florida Florida Center for Renewable Chemicals
Targeted metabolic profiling of the Tg197 mouse model reveals itaconic acid as a marker of Rheumatoid Arthritis.
 Targeted metabolic profiling of the Tg197 mouse model reveals itaconic acid as a marker of Rheumatoid Arthritis. Filippos Michopoulos 1,2,, Niki Karagianni 3, Nichola Whalley 1, Mike Firth 8, Christoforos
Targeted metabolic profiling of the Tg197 mouse model reveals itaconic acid as a marker of Rheumatoid Arthritis. Filippos Michopoulos 1,2,, Niki Karagianni 3, Nichola Whalley 1, Mike Firth 8, Christoforos
Supplementary Materials. The biochemical basis for thermoregulation in heat-producing flowers
 Supplementary Materials The biochemical basis for thermoregulation in heat-producing flowers Yui Umekawa, Roger S. Seymour 2, *, and Kikukatsu Ito United Graduate School of Agricultural Science, Iwate
Supplementary Materials The biochemical basis for thermoregulation in heat-producing flowers Yui Umekawa, Roger S. Seymour 2, *, and Kikukatsu Ito United Graduate School of Agricultural Science, Iwate
Supplemental Information. A Predictive Model for Selective Targeting. of the Warburg Effect through GAPDH. Inhibition with a Natural Product
 Cell Metabolism, Volume 26 Supplemental Information A Predictive Model for Selective Targeting of the Warburg Effect through GAPDH Inhibition with a Natural Product Maria V. Liberti, Ziwei Dai, Suzanne
Cell Metabolism, Volume 26 Supplemental Information A Predictive Model for Selective Targeting of the Warburg Effect through GAPDH Inhibition with a Natural Product Maria V. Liberti, Ziwei Dai, Suzanne
TBT4170 v2014. Kapittel 4 White Biotechnology: Cells as Synthetic Factories. Part 1
 TBT4170 v2014 Kapittel 4 White Biotechnology: Cells as Synthetic Factories Part 1 1 The single definition, OECD Biotechnology The application of science and technology to living organisms, as well as parts,
TBT4170 v2014 Kapittel 4 White Biotechnology: Cells as Synthetic Factories Part 1 1 The single definition, OECD Biotechnology The application of science and technology to living organisms, as well as parts,
Photosynthetic production of biofuels from CO 2 by cyanobacteria using Algenol s Direct to Ethanol process Strain development aspects
 Photosynthetic production of biofuels from CO 2 by cyanobacteria using Algenol s Direct to Ethanol process Strain development aspects Paul Roessler - Oct 1, 2014 Algal Biomass Summit Algenol Overview Advanced
Photosynthetic production of biofuels from CO 2 by cyanobacteria using Algenol s Direct to Ethanol process Strain development aspects Paul Roessler - Oct 1, 2014 Algal Biomass Summit Algenol Overview Advanced
All commands and actions are indicated as highlighted yellow steps CellNetAnalyzer and MatLab commands are in Courier Font
 Steady State Metabolic Modeling Homework In this lab we will use the application CellNetAnalyzer (CNA) to model metabolic pathways in E. coli and M. tuberculosis. This lab is a revised version from a lab
Steady State Metabolic Modeling Homework In this lab we will use the application CellNetAnalyzer (CNA) to model metabolic pathways in E. coli and M. tuberculosis. This lab is a revised version from a lab
Succinyl-proteome profiling of a high taxol containing hybrid Taxus. species (Taxus media) revealed involvement of succinylation in
 Succinyl-proteome profiling of a high taxol containing hybrid Taxus species (Taxus media) revealed involvement of succinylation in multiple metabolic pathways Chenjia Shen 1, Jie Xue 1, Tao Sun 1, Hong
Succinyl-proteome profiling of a high taxol containing hybrid Taxus species (Taxus media) revealed involvement of succinylation in multiple metabolic pathways Chenjia Shen 1, Jie Xue 1, Tao Sun 1, Hong
Metabolism and metabolic networks. Metabolism is the means by which cells acquire energy and building blocks for cellular material
 Metabolism and metabolic networks Metabolism is the means by which cells acquire energy and building blocks for cellular material Metabolism is organized into sequences of biochemical reactions, metabolic
Metabolism and metabolic networks Metabolism is the means by which cells acquire energy and building blocks for cellular material Metabolism is organized into sequences of biochemical reactions, metabolic
Pretreatment Fundamentals
 Pretreatment Fundamentals Bruce E. Dale, Richard T. Elander, Mark T. Holtzapple, Rajeev Kumar, Michael R. Ladisch, Yoon Y. Lee, Nate Mosier, Jack Saddler, Mohammed Moniruzzaman, Charles E. Wyman CAFI BIO
Pretreatment Fundamentals Bruce E. Dale, Richard T. Elander, Mark T. Holtzapple, Rajeev Kumar, Michael R. Ladisch, Yoon Y. Lee, Nate Mosier, Jack Saddler, Mohammed Moniruzzaman, Charles E. Wyman CAFI BIO
Biofuels: Renewable Transportation Fuels from Biomass
 National Renewable Energy Laboratory Biofuels: Renewable Transportation Fuels from Biomass Cynthia Riley Biotechnology Division for Fuels and Chemicals National Bioenergy Center Utility Federal Technology
National Renewable Energy Laboratory Biofuels: Renewable Transportation Fuels from Biomass Cynthia Riley Biotechnology Division for Fuels and Chemicals National Bioenergy Center Utility Federal Technology
Networks of Signal Transduction and Regulation in Cellular Systems
 Max Planck Institute for Dynamics of Complex Technical Systems Magdeburg Networks of Signal Transduction and Regulation in Cellular Systems E.D. Gilles 1 Magdeburg Capital of Saxony-Anhalt 2 3 MAX PLANCK
Max Planck Institute for Dynamics of Complex Technical Systems Magdeburg Networks of Signal Transduction and Regulation in Cellular Systems E.D. Gilles 1 Magdeburg Capital of Saxony-Anhalt 2 3 MAX PLANCK
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION Multilevel interaction of the DnaK/DnaJ(HSP70/HSP40) stress-responsive chaperone machine with the central metabolism Fréderic ANGLES, Marie-Pierre CASTANIE-CORNET, Nawel SLAMA,
SUPPLEMENTARY INFORMATION Multilevel interaction of the DnaK/DnaJ(HSP70/HSP40) stress-responsive chaperone machine with the central metabolism Fréderic ANGLES, Marie-Pierre CASTANIE-CORNET, Nawel SLAMA,
FROM DISCOVERY TO INSIGHT
 Agilent Pathway Architect FROM DISCOVERY TO INSIGHT METABOLOMICS PROTEOMICS PATHWAY ARCHITECT GENOMICS TRANSCRIPTOMICS AGILENT PATHWAY ARCHITECT SPEED DISCOVERY TO UNDERSTANDING Today s scientists face
Agilent Pathway Architect FROM DISCOVERY TO INSIGHT METABOLOMICS PROTEOMICS PATHWAY ARCHITECT GENOMICS TRANSCRIPTOMICS AGILENT PATHWAY ARCHITECT SPEED DISCOVERY TO UNDERSTANDING Today s scientists face
Metabolic Engineering
 saka Univ Metabolic Engineering Department of Bioinformatic Engineering, Graduate School of Information Science and Technology, saka University Dr. Hiroshi Shimizu Professor No. 2 Metabolic Engineering
saka Univ Metabolic Engineering Department of Bioinformatic Engineering, Graduate School of Information Science and Technology, saka University Dr. Hiroshi Shimizu Professor No. 2 Metabolic Engineering
Photosynthesis (in chloroplasts) Solar energy 6CO2 + 6H2O
 B-3.1 Summarize the overall process by which photosynthesis converts solar energy into chemical energy and interpret the chemical equation for the process. Photosynthesis (in chloroplasts) Solar energy
B-3.1 Summarize the overall process by which photosynthesis converts solar energy into chemical energy and interpret the chemical equation for the process. Photosynthesis (in chloroplasts) Solar energy
Transporter manipulation in food crops for increased yield
 ASPB 2018 Frank Skraly Transporter manipulation in food crops for increased yield July 17, 2018 Yield10 company overview Yield10 Bioscience (NasdaqCM:YTEN) is developing technologies to enhance global
ASPB 2018 Frank Skraly Transporter manipulation in food crops for increased yield July 17, 2018 Yield10 company overview Yield10 Bioscience (NasdaqCM:YTEN) is developing technologies to enhance global
GATOR1 regulates nitrogenic cataplerotic reactions of the mitochondrial TCA cycle. Jun Chen, Benjamin M. Sutter, Lei Shi, Benjamin P.
 Supplementary Information GATOR1 regulates nitrogenic cataplerotic reactions of the mitochondrial TCA cycle Jun Chen, Benjamin M. Sutter, Lei Shi, Benjamin P. Tu* Department of Biochemistry University
Supplementary Information GATOR1 regulates nitrogenic cataplerotic reactions of the mitochondrial TCA cycle Jun Chen, Benjamin M. Sutter, Lei Shi, Benjamin P. Tu* Department of Biochemistry University
Mark Burk (PI), Nelson Barton, and John Trawick; Genomatica
 Development of an Integrated Biofuel and Chemical Refinery Final report for public Recipient Organization: Genomatica Supported in part by the US DOE award DE- EE0005002 to Genomatica Mark Burk (PI), Nelson
Development of an Integrated Biofuel and Chemical Refinery Final report for public Recipient Organization: Genomatica Supported in part by the US DOE award DE- EE0005002 to Genomatica Mark Burk (PI), Nelson
Model driven intracellular redox status modulation for increasing isobutanol production in Escherichia coli
 DOI 10.1186/s13068-015-0291-2 RESEARCH ARTICLE Open Access Model driven intracellular redox status modulation for increasing isobutanol production in Escherichia coli Jiao Liu 1,2, Haishan Qi 1,2, Cheng
DOI 10.1186/s13068-015-0291-2 RESEARCH ARTICLE Open Access Model driven intracellular redox status modulation for increasing isobutanol production in Escherichia coli Jiao Liu 1,2, Haishan Qi 1,2, Cheng
OMICS International Conferences
 About OMICS Group OMICS Group is an amalgamation of Open Access Publications and worldwide international science conferences and events. Established in the year 2007 with the sole aim of making the information
About OMICS Group OMICS Group is an amalgamation of Open Access Publications and worldwide international science conferences and events. Established in the year 2007 with the sole aim of making the information
Biomolecules: nucleotides
 Biomolecules: nucleotides Nucleotides and nucleic acids Nucleic acids are linear hetero-polymers adapted to maintain and transmit the genetic information Depending on their sugar moiety they can be classified
Biomolecules: nucleotides Nucleotides and nucleic acids Nucleic acids are linear hetero-polymers adapted to maintain and transmit the genetic information Depending on their sugar moiety they can be classified
Industrial microbiology
 Industrial microbiology pp. 166-173, 1032-1038, 1039-1045,1046-1050 Ed van Niel Ed.van_Niel@tmb.lth.se We are here Industrial microbiology biotechnology Why the increased interest Microbiological versus
Industrial microbiology pp. 166-173, 1032-1038, 1039-1045,1046-1050 Ed van Niel Ed.van_Niel@tmb.lth.se We are here Industrial microbiology biotechnology Why the increased interest Microbiological versus
Integrative Biology Accepted Manuscript
 Integrative Biology Accepted Manuscript This is an Accepted Manuscript, which has been through the Royal Society of Chemistry peer review process and has been accepted for publication. Accepted Manuscripts
Integrative Biology Accepted Manuscript This is an Accepted Manuscript, which has been through the Royal Society of Chemistry peer review process and has been accepted for publication. Accepted Manuscripts
BIOLOGY 311C - Brand Spring 2008
 BIOLOGY 311C - Brand Spring 2008 NAME (printed very legibly) Key UT-EID EXAMINATION 3 Before beginning, check to be sure that this exam contains 7 pages (including front and back) numbered consecutively,
BIOLOGY 311C - Brand Spring 2008 NAME (printed very legibly) Key UT-EID EXAMINATION 3 Before beginning, check to be sure that this exam contains 7 pages (including front and back) numbered consecutively,
Name period date AP BIO- 2 nd QTR 6 Week Test Review
 Name period date AP BIO- 2 nd QTR 6 Week Test Review Cellular Respiration 1. In the following reaction, C6H12O6 + 6 O2 6 CO2 + 6 H2O + Energy, identify which is reduced and which is oxidized: a. Glucose
Name period date AP BIO- 2 nd QTR 6 Week Test Review Cellular Respiration 1. In the following reaction, C6H12O6 + 6 O2 6 CO2 + 6 H2O + Energy, identify which is reduced and which is oxidized: a. Glucose
Abhishek Murarka, 1 James M. Clomburg 1 and Ramon Gonzalez 1,2 INTRODUCTION
 Microbiology (2010), 156, 1860 1872 DOI 10.1099/mic.0.036251-0 Metabolic flux analysis of wild-type Escherichia coli and mutants deficient in pyruvate-dissimilating enzymes during the fermentative metabolism
Microbiology (2010), 156, 1860 1872 DOI 10.1099/mic.0.036251-0 Metabolic flux analysis of wild-type Escherichia coli and mutants deficient in pyruvate-dissimilating enzymes during the fermentative metabolism
Pyruvate cycle increases aminoglycoside efficacy and provides respiratory energy in bacteria
 Pyruvate cycle increases aminoglycoside efficacy and provides respiratory energy in bacteria Yu-bin Su a,1, Bo Peng a,b,1,2, Hui Li a,c,1, Zhi-xue Cheng a,1, Tian-tuo Zhang d, Jia-xin Zhu d, Dan Li a,
Pyruvate cycle increases aminoglycoside efficacy and provides respiratory energy in bacteria Yu-bin Su a,1, Bo Peng a,b,1,2, Hui Li a,c,1, Zhi-xue Cheng a,1, Tian-tuo Zhang d, Jia-xin Zhu d, Dan Li a,
Correlations between the expression of genes encoding enzymes in the Krebs cycle and control of DNA replication in human cells
 Correlations between the expression of genes encoding enzymes in the Krebs cycle and control of DNA replication in human cells Aneta Wieczorek Duplication of genetic material is a key biological process
Correlations between the expression of genes encoding enzymes in the Krebs cycle and control of DNA replication in human cells Aneta Wieczorek Duplication of genetic material is a key biological process
number Done by Corrected by Doctor
 number 34 Done by Abdulrahman Alhanbali Corrected by Mohammad Mahmoud Tarabeih Doctor Diala Nucleotide metabolism 1 P age In this lecture we will talk about nucleotides; their structures, the synthesis
number 34 Done by Abdulrahman Alhanbali Corrected by Mohammad Mahmoud Tarabeih Doctor Diala Nucleotide metabolism 1 P age In this lecture we will talk about nucleotides; their structures, the synthesis
Metabolomics Pathway analysis. Anatoly Sorokin
 Metabolomics Pathway analysis Anatoly Sorokin Metabolomics Metabolomics is the "systema7c study of the unique chemical fingerprints that specific cellular processes leave behind", the study of their small-
Metabolomics Pathway analysis Anatoly Sorokin Metabolomics Metabolomics is the "systema7c study of the unique chemical fingerprints that specific cellular processes leave behind", the study of their small-
Microbiology Helmut Pospiech
 Microbiology 27.3.2018 Helmut Pospiech Microbial Metabolism Different ways to make a living Energy metabolism of Microorganisms Fermentation ADP +Pi Motility ATP Active transport (nutrient uptake) Any
Microbiology 27.3.2018 Helmut Pospiech Microbial Metabolism Different ways to make a living Energy metabolism of Microorganisms Fermentation ADP +Pi Motility ATP Active transport (nutrient uptake) Any
Chapter 18 NETWORKS: APPLICATIONS TO LYSINE FERMENTATIONS TABLE OF CONTENTS ...
 Chapter 18 FLUX DETERMINATION IN CELLULAR BIOREACTION NETWORKS: APPLICATIONS TO LYSINE FERMENTATIONS Joseph J. VIlllino and Gregory Stephanopoulos TABLE OF CONTENTS I. Introduction.....'...2M II............................
Chapter 18 FLUX DETERMINATION IN CELLULAR BIOREACTION NETWORKS: APPLICATIONS TO LYSINE FERMENTATIONS Joseph J. VIlllino and Gregory Stephanopoulos TABLE OF CONTENTS I. Introduction.....'...2M II............................
Genome-scale metabolic network guided engineering of Streptomyces tsukubaensis for FK506 production improvement
 Huang et al. Microbial Cell Factories 213, 12:52 RESEARCH Genome-scale metabolic network guided engineering of Streptomyces tsukubaensis for FK56 production improvement Di Huang 1,3, Shanshan Li 1, Menglei
Huang et al. Microbial Cell Factories 213, 12:52 RESEARCH Genome-scale metabolic network guided engineering of Streptomyces tsukubaensis for FK56 production improvement Di Huang 1,3, Shanshan Li 1, Menglei
Investigating the effects of perturbations to pgi and eno gene expression on central carbon metabolism in Escherichia coli using
 Usui et al. Microbial Cell Factories 212, 11:87 RESEARCH Open Access Investigating the effects of perturbations to pgi and eno gene expression on central carbon metabolism in Escherichia coli using 13
Usui et al. Microbial Cell Factories 212, 11:87 RESEARCH Open Access Investigating the effects of perturbations to pgi and eno gene expression on central carbon metabolism in Escherichia coli using 13
*A table of standard reduction potentials may be found on the last page of the exam.
 Name: Page 1 Ecology 1.018J/7.30 Quiz 1 October 2, 2008 Please put your name on every page! Space is provided for your answers; if you need more room, use the back of the *same* page. This exam is worth
Name: Page 1 Ecology 1.018J/7.30 Quiz 1 October 2, 2008 Please put your name on every page! Space is provided for your answers; if you need more room, use the back of the *same* page. This exam is worth
The Carboxylate Platform
 The Carboxylate Platform Nigel Horan Lecture Outline The industry why it should innovate What is the carboxylate platform? Potential benefits Retrofitting and new build Conclusions AD Industry Now over
The Carboxylate Platform Nigel Horan Lecture Outline The industry why it should innovate What is the carboxylate platform? Potential benefits Retrofitting and new build Conclusions AD Industry Now over
number Done by Corrected by Doctor Diala
 number 34 Done by Abdulrahman Alhanbali Corrected by Mohammad Mahmoud Tarabeih Doctor Diala 1 P a g e Nucleotide metabolism In this lecture we will talk about nucleotides; their structures, the synthesis
number 34 Done by Abdulrahman Alhanbali Corrected by Mohammad Mahmoud Tarabeih Doctor Diala 1 P a g e Nucleotide metabolism In this lecture we will talk about nucleotides; their structures, the synthesis
Increased 3-hydroxypropionic acid production from glycerol, by modification of central metabolism in Escherichia coli
 Tokuyama et al. Microbial Cell Factories 214, 13:64 RESEARCH Open Access Increased 3-hydroxypropionic acid production from glycerol, by modification of central metabolism in Escherichia coli Kento Tokuyama
Tokuyama et al. Microbial Cell Factories 214, 13:64 RESEARCH Open Access Increased 3-hydroxypropionic acid production from glycerol, by modification of central metabolism in Escherichia coli Kento Tokuyama
CM4125 Bioprocess Engineering Lab: Week 4: Introduction to Metabolic Engineering and Metabolic Flux Analysis
 CM4125 Bioprocess Engineering Lab: Week 4: Introduction to Metabolic Engineering and Metabolic Flux Analysis Instructors: : David R. Shonnard 1, Susan T. Bagley 2 Laboratory Teaching Assistant: : Abraham
CM4125 Bioprocess Engineering Lab: Week 4: Introduction to Metabolic Engineering and Metabolic Flux Analysis Instructors: : David R. Shonnard 1, Susan T. Bagley 2 Laboratory Teaching Assistant: : Abraham
Metabolic Engineering of New Routes to Biofuels
 Metabolic Engineering of New Routes to Biofuels Steve Wilkinson Manchester Interdisciplinary Biocentre, Manchester University UK. Overview Metabolism, enzymes, genes Bioethanol production in yeast Metabolic
Metabolic Engineering of New Routes to Biofuels Steve Wilkinson Manchester Interdisciplinary Biocentre, Manchester University UK. Overview Metabolism, enzymes, genes Bioethanol production in yeast Metabolic
METABOLIC MODELLING FOR THE CONTROL OF INTRACELLULAR PROCESSES IN PLANT CELLS CULTURES
 2.5 ADP + 2.5 Pi + NADH + O 2 2.5 ATP + NAD - 2 METABOLIC MODELLING FOR THE CONTROL OF INTRACELLULAR PROCESSES IN PLANT CELLS CULTURES Mathieu Cloutier Michel Perrier Mario Jolicoeur Montpellier, Mercredi
2.5 ADP + 2.5 Pi + NADH + O 2 2.5 ATP + NAD - 2 METABOLIC MODELLING FOR THE CONTROL OF INTRACELLULAR PROCESSES IN PLANT CELLS CULTURES Mathieu Cloutier Michel Perrier Mario Jolicoeur Montpellier, Mercredi
Application of metabolic flux and transcriptome. analyses to understanding the physiology of. engineered Geobacillus thermoglucosidasius
 Application of metabolic flux and transcriptome analyses to understanding the physiology of engineered Geobacillus thermoglucosidasius Charlotte Emily Ward Imperial College London, Department of Life Sciences
Application of metabolic flux and transcriptome analyses to understanding the physiology of engineered Geobacillus thermoglucosidasius Charlotte Emily Ward Imperial College London, Department of Life Sciences
Extreme Pathway Analysis of Human Red Blood Cell Metabolism
 808 Biophysical Journal Volume 83 August 2002 808 818 Extreme Pathway Analysis of Human Red Blood Cell Metabolism Sharon J. Wiback and Bernhard O. Palsson Department of Bioengineering, University of California,
808 Biophysical Journal Volume 83 August 2002 808 818 Extreme Pathway Analysis of Human Red Blood Cell Metabolism Sharon J. Wiback and Bernhard O. Palsson Department of Bioengineering, University of California,
Minimal Reaction Sets for Escherichia coli Metabolism under Different Growth Requirements and Uptake Environments
 Biotechnol. Prog. 2001, 17, 791 797 791 ARTICLES inimal Reaction Sets for Escherichia coli etabolism under Different Growth Requirements and Uptake Environments Anthony P. Burgard, Shankar Vaidyaraman,
Biotechnol. Prog. 2001, 17, 791 797 791 ARTICLES inimal Reaction Sets for Escherichia coli etabolism under Different Growth Requirements and Uptake Environments Anthony P. Burgard, Shankar Vaidyaraman,
Introduction to Enzymes
 Introduction to Enzymes Enzyme Engineering What is enzymes? Life depends on well-orchestrated series of chemical reactions : E. coli has 4288 proteins, 2656 of which are characterized, and 64% (1701) of
Introduction to Enzymes Enzyme Engineering What is enzymes? Life depends on well-orchestrated series of chemical reactions : E. coli has 4288 proteins, 2656 of which are characterized, and 64% (1701) of
(12) Patent Application Publication (10) Pub. No.: US 2002/ A1
 (19) United States US 2002O168654A1 (12) Patent Application Publication (10) Pub. No. US 2002/0168654A1 Maranas et al. (43) Pub. Date Nov. 14, 2002 (54) METHOD AND SYSTEM FOR MODELING CELLULAR METABOLISM
(19) United States US 2002O168654A1 (12) Patent Application Publication (10) Pub. No. US 2002/0168654A1 Maranas et al. (43) Pub. Date Nov. 14, 2002 (54) METHOD AND SYSTEM FOR MODELING CELLULAR METABOLISM
Computational Methods in Bioinformatics
 Computational Methods in Bioinformatics Ying Xu 2017/12/6 1 What We Intend to Teach A general introduction to the field of bioinformatics what problems people have been and are currently working on how
Computational Methods in Bioinformatics Ying Xu 2017/12/6 1 What We Intend to Teach A general introduction to the field of bioinformatics what problems people have been and are currently working on how
Enzymes That Bind Nucleotides
 Enzymes That Bind Nucleotides Nucleotides play a central role in cellular metabolism, in which they are the major currency of energy exchange. They channel the energy released during the catabolism of
Enzymes That Bind Nucleotides Nucleotides play a central role in cellular metabolism, in which they are the major currency of energy exchange. They channel the energy released during the catabolism of
Microbiology Exam 2, 2/27/07 Name
 Microbiology Exam 2, 2/27/07 Name 1. (2 pts) Catabolism refers to D, whereas anabolism refers to A. (be sure to fill in the blanks!) a. synthesis of cell structures b. transfer of electrons c. uptake of
Microbiology Exam 2, 2/27/07 Name 1. (2 pts) Catabolism refers to D, whereas anabolism refers to A. (be sure to fill in the blanks!) a. synthesis of cell structures b. transfer of electrons c. uptake of
BIOLOGICAL PHOSPHOROUS REMOVAL AN OPERATOR S GUIDE
 BIOLOGICAL PHOSPHOROUS REMOVAL AN OPERATOR S GUIDE ABSTRACT If you have ever faced a rising effluent phosphorous concentration and you are relying on biological phosphorous removal, the information offered
BIOLOGICAL PHOSPHOROUS REMOVAL AN OPERATOR S GUIDE ABSTRACT If you have ever faced a rising effluent phosphorous concentration and you are relying on biological phosphorous removal, the information offered
Computational and Experimental Analysis of Redundancy in the Central Metabolism of Geobacter sulfurreducens
 Computational and Experimental Analysis of Redundancy in the Central Metabolism of Geobacter sulfurreducens Daniel Segura 1,2, Radhakrishnan Mahadevan 3,4*, Katy Juárez 1,5, Derek R. Lovley 1 1 Department
Computational and Experimental Analysis of Redundancy in the Central Metabolism of Geobacter sulfurreducens Daniel Segura 1,2, Radhakrishnan Mahadevan 3,4*, Katy Juárez 1,5, Derek R. Lovley 1 1 Department
Computational and Experimental Analysis of Redundancy in the Central Metabolism of Geobacter Sulfurreducens
 University of Massachusetts Amherst From the SelectedWorks of Derek Lovley 2008 Computational and Experimental Analysis of Redundancy in the Central Metabolism of Geobacter Sulfurreducens Derek Lovley,
University of Massachusetts Amherst From the SelectedWorks of Derek Lovley 2008 Computational and Experimental Analysis of Redundancy in the Central Metabolism of Geobacter Sulfurreducens Derek Lovley,
Take-Home Quiz II. Summer 2005 Semester
 General Instructions and Information: Obtain an answer sheet from the instructor and legibly write your name in the appropriate space. After placing your name, you must enter your Patron ID Number (NOT
General Instructions and Information: Obtain an answer sheet from the instructor and legibly write your name in the appropriate space. After placing your name, you must enter your Patron ID Number (NOT
(Received for publication, February 27, 1978, and in revised form, January 30, 1980) or galactose, glycolytic activity decreases dramatically, by as
 THE JOURNAL OF BOLO(i1CAL CHEMSTRY Vd. 255. No. 12. ssue of dune 25. pp 5616-5626, 1980 Prmted m SA. The Pentose Cycle CONTROL AND ESSENTAL FUNCTON N HeLa CELL NUCLEC ACD SYNTHESS* (Received for publication,
THE JOURNAL OF BOLO(i1CAL CHEMSTRY Vd. 255. No. 12. ssue of dune 25. pp 5616-5626, 1980 Prmted m SA. The Pentose Cycle CONTROL AND ESSENTAL FUNCTON N HeLa CELL NUCLEC ACD SYNTHESS* (Received for publication,
BIOCHEMICAL AND BIOPHYSICAL RESEARCH COMMUNICATIONS
 ISOLATION OF MUTANTS OF ESCHERICHIA COLI LACKING NAD- AND NADP-LINKED M~ic ENZYME ACTIVITIES Eric J. Hansen and Elliot Juni Department of Microbiology The University of Michigan Ann Arbor, Michigan 48104
ISOLATION OF MUTANTS OF ESCHERICHIA COLI LACKING NAD- AND NADP-LINKED M~ic ENZYME ACTIVITIES Eric J. Hansen and Elliot Juni Department of Microbiology The University of Michigan Ann Arbor, Michigan 48104
1) Introduction. 2. Basic Concepts. A Brief Guide to Enzyme Nomenclature and Classification Keith Tipton and Andrew McDonald
 A Brief Guide to Enzyme Nomenclature and Classification Keith Tipton and Andrew McDonald 1) Introduction NC-IUBMB Enzyme List, or, to give it its full title, Recommendations of the Nomenclature Committee
A Brief Guide to Enzyme Nomenclature and Classification Keith Tipton and Andrew McDonald 1) Introduction NC-IUBMB Enzyme List, or, to give it its full title, Recommendations of the Nomenclature Committee
Perturbation-based Control of Industrial Fed-batch Bioprocesses
 Perturbation-based Control of Industrial Fed-batch Bioprocesses Johnsson, Ola 215 Document Version: Publisher's PDF, also known as Version of record Link to publication Citation for published version (APA):
Perturbation-based Control of Industrial Fed-batch Bioprocesses Johnsson, Ola 215 Document Version: Publisher's PDF, also known as Version of record Link to publication Citation for published version (APA):
Chapter 12 Respiration
 2.2 Cell Metabolism Learning Objectives Chapter 12 Respiration 2.2.5 Respiration 1. Define, give the role and balanced equation for "aerobic respiration". 2. Explain the stages and molecules involved in
2.2 Cell Metabolism Learning Objectives Chapter 12 Respiration 2.2.5 Respiration 1. Define, give the role and balanced equation for "aerobic respiration". 2. Explain the stages and molecules involved in
Exometabolome analysis reveals hypoxia at the up-scaling of a Saccharomyces cerevisiae high-cell density fed-batch biopharmaceutical process
 Fu et al. Microbial Cell Factories 4, 3:3 http://www.microbialcellfactories.com/content/3//3 RESEARCH Open Access Exometabolome analysis reveals hypoxia at the up-scaling of a Saccharomyces cerevisiae
Fu et al. Microbial Cell Factories 4, 3:3 http://www.microbialcellfactories.com/content/3//3 RESEARCH Open Access Exometabolome analysis reveals hypoxia at the up-scaling of a Saccharomyces cerevisiae
TU Dortmund University Department of Biochemical and Chemical Engineering Process Dynamics and Operations Group Prof. Dr.-Ing.
 TU Dortmund University Department of Biochemical and Chemical Engineering Process Dynamics and Operations Group Prof. Dr.-Ing. Sebastian Engell Bachelor Thesis Modelling the Ethanol production by Saccharomyces
TU Dortmund University Department of Biochemical and Chemical Engineering Process Dynamics and Operations Group Prof. Dr.-Ing. Sebastian Engell Bachelor Thesis Modelling the Ethanol production by Saccharomyces
Preparation and Application of Proteomic Antibody Bank against Whole Water-Soluble Proteins from Rapid-Growing Bamboo Shoots
 Preparation and Application of Proteomic Antibody Bank against Whole Water-Soluble Proteins from Rapid-Growing Bamboo Shoots Yu-Jen Wu, Dai-Jer Wu, Jiann-Shing Wu and Rong-Huay Juang * * Major affiliation:
Preparation and Application of Proteomic Antibody Bank against Whole Water-Soluble Proteins from Rapid-Growing Bamboo Shoots Yu-Jen Wu, Dai-Jer Wu, Jiann-Shing Wu and Rong-Huay Juang * * Major affiliation:
green B 1 ) into a single unit to model the substrate in this reaction. enzyme
 Teacher Key Objectives You will use the model pieces in the kit to: Simulate enzymatic actions. Explain enzymatic specificity. Investigate two types of enzyme inhibitors used in regulating enzymatic activity.
Teacher Key Objectives You will use the model pieces in the kit to: Simulate enzymatic actions. Explain enzymatic specificity. Investigate two types of enzyme inhibitors used in regulating enzymatic activity.
CEE 370 Environmental Engineering Principles. Environmental Microbiology
 Updated: 1 October 015 Print version CEE 370 Environmental Engineering Principles Lecture #13 Environmental Biology II Metabolism Reading: Davis & Masten, Chapter 3 David Reckhow CEE 370 L#13 1 Environmental
Updated: 1 October 015 Print version CEE 370 Environmental Engineering Principles Lecture #13 Environmental Biology II Metabolism Reading: Davis & Masten, Chapter 3 David Reckhow CEE 370 L#13 1 Environmental
Mutual Regulation of CRP and N(epsilon)-Lysine Acetylation in Escherichia Coli
 Loyola University Chicago Loyola ecommons Dissertations Theses and Dissertations 2017 Mutual Regulation of CRP and N(epsilon)-Lysine Acetylation in Escherichia Coli Robert James Davis Loyola University
Loyola University Chicago Loyola ecommons Dissertations Theses and Dissertations 2017 Mutual Regulation of CRP and N(epsilon)-Lysine Acetylation in Escherichia Coli Robert James Davis Loyola University
METABOLIC ENGINEERING OF BACILLUS FOR ENHANCED PRODUCT AND CELLULAR YIELDS. Zhiwei Pan
 METABOLIC ENGINEERING OF BACILLUS FOR ENHANCED PRODUCT AND CELLULAR YIELDS by Zhiwei Pan B.E. in Fermentation Engineering, Wuxi University of Light Industry, 1995 M.S. in Biochemical Engineering, East
METABOLIC ENGINEERING OF BACILLUS FOR ENHANCED PRODUCT AND CELLULAR YIELDS by Zhiwei Pan B.E. in Fermentation Engineering, Wuxi University of Light Industry, 1995 M.S. in Biochemical Engineering, East
Constraint-Based Workshops
 Constraint-Based Workshops 10. Flux Coupling (again) & Metabolic Engineering February 28 th, 2008 Constraint-Based Methods Optimal Solutions 1. FBA 2. Flux Variability Flux Dependencies 1. Robustness 2.
Constraint-Based Workshops 10. Flux Coupling (again) & Metabolic Engineering February 28 th, 2008 Constraint-Based Methods Optimal Solutions 1. FBA 2. Flux Variability Flux Dependencies 1. Robustness 2.
CEE 370 Environmental Engineering Principles
 Updated: 12 October 2015 Print version CEE 370 Environmental Engineering Principles Lecture #13 Environmental Biology II Metabolism Reading: Davis & Masten, Chapter 3 David Reckhow CEE 370 L#13 1 Environmental
Updated: 12 October 2015 Print version CEE 370 Environmental Engineering Principles Lecture #13 Environmental Biology II Metabolism Reading: Davis & Masten, Chapter 3 David Reckhow CEE 370 L#13 1 Environmental
Enzymes. 13. Explain the active site theory to examine enzyme function
 Name: 2.2 Cell Metabolism Objectives At the end of this sub section students should be able to: 2.2.1 Metabolism 1. Define the term: metabolism. 2.2.2 Sources of energy 2. State that solar energy is source
Name: 2.2 Cell Metabolism Objectives At the end of this sub section students should be able to: 2.2.1 Metabolism 1. Define the term: metabolism. 2.2.2 Sources of energy 2. State that solar energy is source
SUPPLEMENTAL MATERIALS AND METHODS
 SUPPLEMENTAL MATERIALS AND METHODS Cell Lines and Reagents Human CRC cell lines HCT116, HKe-3, HKh-2, DLD-1, and DKO-4 were generated as described previously (11). For the normoxic culture, cells were
SUPPLEMENTAL MATERIALS AND METHODS Cell Lines and Reagents Human CRC cell lines HCT116, HKe-3, HKh-2, DLD-1, and DKO-4 were generated as described previously (11). For the normoxic culture, cells were
Adenine and adenosine salvage pathways in erythrocytes and the role of S-adenosylhomocysteine hydrolase
 Adenine and adenosine salvage pathways in erythrocytes and the role of S-adenosylhomocysteine hydrolase A theoretical study using elementary flux modes Stefan Schuster and Dimitar Kenanov Department of
Adenine and adenosine salvage pathways in erythrocytes and the role of S-adenosylhomocysteine hydrolase A theoretical study using elementary flux modes Stefan Schuster and Dimitar Kenanov Department of
Scaling up Industrial Biotechnology
 Scaling up Industrial Biotechnology 7 th Life Science Symposium Bioenergy May 10 th 2016 Senior Engineering Fellow Outline Scaling up Industrial Biotechnology A fermentation guy Genomatica What is scale-up?
Scaling up Industrial Biotechnology 7 th Life Science Symposium Bioenergy May 10 th 2016 Senior Engineering Fellow Outline Scaling up Industrial Biotechnology A fermentation guy Genomatica What is scale-up?
Life's Chemical and Biological Processes
 Life's Chemical and Biological Processes AP Biology 1st June 2006 Contents 1 Cellular Respiration 2 1.1 Introduction............................ 2 1.2 Glycolysis............................. 3 1.3 Aerobic
Life's Chemical and Biological Processes AP Biology 1st June 2006 Contents 1 Cellular Respiration 2 1.1 Introduction............................ 2 1.2 Glycolysis............................. 3 1.3 Aerobic
Intracellular Carbon Fluxes in Riboflavin-Producing Bacillus subtilis during Growth on Two-Carbon Substrate Mixtures
 APPLIED AND ENVIRONMENTAL MICROBIOLOGY, Apr. 2002, p. 1760 1771 Vol. 68, No. 4 0099-2240/02/$04.00 0 DOI: 10.1128/AEM.68.4.1760 1771.2002 Copyright 2002, American Society for Microbiology. All Rights Reserved.
APPLIED AND ENVIRONMENTAL MICROBIOLOGY, Apr. 2002, p. 1760 1771 Vol. 68, No. 4 0099-2240/02/$04.00 0 DOI: 10.1128/AEM.68.4.1760 1771.2002 Copyright 2002, American Society for Microbiology. All Rights Reserved.
PROLONGING A HIGH RATE OF SUCCINATE PRODUCTION IN DUAL-PHASE PRABU DIRAVIDAN KIZHSEVUR VIJAYAN. (Under the direction of Mark A.
 PROLONGING A HIGH RATE OF SUCCINATE PRODUCTION IN DUAL-PHASE FERMENTATION USING ESCHERICHIA COLI AFP111 By PRABU DIRAVIDAN KIZHSEVUR VIJAYAN (Under the direction of Mark A. Eiteman) ABSTRACT The commodity
PROLONGING A HIGH RATE OF SUCCINATE PRODUCTION IN DUAL-PHASE FERMENTATION USING ESCHERICHIA COLI AFP111 By PRABU DIRAVIDAN KIZHSEVUR VIJAYAN (Under the direction of Mark A. Eiteman) ABSTRACT The commodity
Metabolism BIOL 3702: Chapter 10
 Metabolism BIOL 3702: Chapter 10 Introduction to Metabolism u Metabolism is the sum total of all the chemical reactions occurring in a cell u Two major parts of metabolism: v Catabolism Ø Large, more complex
Metabolism BIOL 3702: Chapter 10 Introduction to Metabolism u Metabolism is the sum total of all the chemical reactions occurring in a cell u Two major parts of metabolism: v Catabolism Ø Large, more complex
Alessandro Ciranna 1*, Sudhanshu S Pawar 2, Ville Santala 1, Matti Karp 1 and Ed WJ van Niel 2
 Ciranna et al. Microbial Cell Factories 2014, 13:48 RESEARCH Open Access Assessment of metabolic flux distribution in the thermophilic hydrogen producer Caloramator celer as affected by external ph and
Ciranna et al. Microbial Cell Factories 2014, 13:48 RESEARCH Open Access Assessment of metabolic flux distribution in the thermophilic hydrogen producer Caloramator celer as affected by external ph and
Analysis of gluconeogenic and anaplerotic enzymes in Campylobacter jejuni: an essential role for phosphoenolpyruvate carboxykinase
 Microbiology (2002), 148, 685 694 Printed in Great Britain Analysis of gluconeogenic and anaplerotic enzymes in Campylobacter jejuni: an essential role for phosphoenolpyruvate carboxykinase Jyoti Velayudhan
Microbiology (2002), 148, 685 694 Printed in Great Britain Analysis of gluconeogenic and anaplerotic enzymes in Campylobacter jejuni: an essential role for phosphoenolpyruvate carboxykinase Jyoti Velayudhan
Scientific Background Synechococcus pigment types
 Scientific Background Synechococcus pigment types Oligotrophic Mesotrophic Coastal 0 m Nutrients 50 m 100 m 150 m Scientific Background Synechococcus Genomes BL107 CA RS9917 Clade I Clade II Clade III
Scientific Background Synechococcus pigment types Oligotrophic Mesotrophic Coastal 0 m Nutrients 50 m 100 m 150 m Scientific Background Synechococcus Genomes BL107 CA RS9917 Clade I Clade II Clade III
(12) Patent Application Publication (10) Pub. No.: US 2015/ A1
 US 20150203877A1 (19) United States (12) Patent Application Publication (10) Pub. No.: US 2015/0203877 A1 Rush et al. (43) Pub. Date: Jul. 23, 2015 (54) YEAST CELLS HAVING REDUCTIVE TCA PATHWAY FROM PYRUVATE
US 20150203877A1 (19) United States (12) Patent Application Publication (10) Pub. No.: US 2015/0203877 A1 Rush et al. (43) Pub. Date: Jul. 23, 2015 (54) YEAST CELLS HAVING REDUCTIVE TCA PATHWAY FROM PYRUVATE
Growth rate dependency of de novo resveratrol production in chemostat cultures of an engineered Saccharomyces cerevisiae strain
 DOI 10.1186/s12934-015-0321-6 RESEARCH Open Access Growth rate dependency of de novo resveratrol production in chemostat cultures of an engineered Saccharomyces cerevisiae strain Tim Vos, Pilar de la Torre
DOI 10.1186/s12934-015-0321-6 RESEARCH Open Access Growth rate dependency of de novo resveratrol production in chemostat cultures of an engineered Saccharomyces cerevisiae strain Tim Vos, Pilar de la Torre
Active Role of Oxygen and NADH Oxidase in Growth and Energy Metabolism of Leuconostoc
 Journal of General Microbiology (1 986), 132, 1789-1 796. Printed in Great Britain 1789 Active Role of Oxygen and NADH Oxidase in Growth and Energy Metabolism of Leuconostoc By CON A. LUCEY A SEAMUS COON*
Journal of General Microbiology (1 986), 132, 1789-1 796. Printed in Great Britain 1789 Active Role of Oxygen and NADH Oxidase in Growth and Energy Metabolism of Leuconostoc By CON A. LUCEY A SEAMUS COON*
Engineering E. coli for xylitol production during growth on xylose
 Engineering E. coli for xylitol production during growth on xylose Olubolaji Akinterinwa & Patrick C. Cirino IBE 2008 Annual Meeting, March 7th 008-17 Advances in Engineering Microbial Metabolism Xylitol
Engineering E. coli for xylitol production during growth on xylose Olubolaji Akinterinwa & Patrick C. Cirino IBE 2008 Annual Meeting, March 7th 008-17 Advances in Engineering Microbial Metabolism Xylitol
Please sign below if you wish to have your grades posted by the last five digits of your SSN
 BIO 226R EXAM II (Sample) PRINT YOUR NAME SSN Please sign below if you wish to have your grades posted by the last five digits of your SSN Signature BIO 226R Exam II has 6 pages, and 27 questions. There
BIO 226R EXAM II (Sample) PRINT YOUR NAME SSN Please sign below if you wish to have your grades posted by the last five digits of your SSN Signature BIO 226R Exam II has 6 pages, and 27 questions. There
College of pharmacy Third stage Dr.Rafeef Amer
 College of pharmacy Third stage Dr.Rafeef Amer Hhhhhhhhhhhhhhhhhhhhhhhhhhhhhh hhhhhhhhhhhhhhhhvyyyyyyyt``` Enzyme Enzymes are very efficient catalysts for biochemical reactions. present in the cell in
College of pharmacy Third stage Dr.Rafeef Amer Hhhhhhhhhhhhhhhhhhhhhhhhhhhhhh hhhhhhhhhhhhhhhhvyyyyyyyt``` Enzyme Enzymes are very efficient catalysts for biochemical reactions. present in the cell in
ENGINEERING THE METABOLISM OF Escherichia coli FOR THE SYNTHESIS OF OXIDIZED PRODUCTS: ACETATE AND PYRUVATE PRODUCTION
 ENGINEERING THE METABOLISM OF Escherichia coli FOR THE SYNTHESIS OF OXIDIZED PRODUCTS: ACETATE AND PYRUVATE PRODUCTION By THOMAS B. CAUSEY A DISSERTATION PRESENTED TO THE GRADUATE SCHOOL OF THE UNIVERSITY
ENGINEERING THE METABOLISM OF Escherichia coli FOR THE SYNTHESIS OF OXIDIZED PRODUCTS: ACETATE AND PYRUVATE PRODUCTION By THOMAS B. CAUSEY A DISSERTATION PRESENTED TO THE GRADUATE SCHOOL OF THE UNIVERSITY
Metabolomics uncovers the regulatory pathway of acyl-homoserine. lactones-based quorum sensing in anammox consortia
 Supporting information Metabolomics uncovers the regulatory pathway of acyl-homoserine lactones-based quorum sensing in anammox consortia Xi Tang 1,2, Yongzhao Guo 1,3, Shanshan Wu 1,2, Liming Chen 1,2,
Supporting information Metabolomics uncovers the regulatory pathway of acyl-homoserine lactones-based quorum sensing in anammox consortia Xi Tang 1,2, Yongzhao Guo 1,3, Shanshan Wu 1,2, Liming Chen 1,2,
Qualitative Cellular Dynamics Analysis with Gene Regulatory Information and Metabolic Pathways
 The First International Symposium on Optimization and Systems Biology (OSB 07) Beijing, China, August 8 10, 2007 Copyright 2007 ORSC & APORC pp. 143 150 Qualitative Cellular Dynamics Analysis with Gene
The First International Symposium on Optimization and Systems Biology (OSB 07) Beijing, China, August 8 10, 2007 Copyright 2007 ORSC & APORC pp. 143 150 Qualitative Cellular Dynamics Analysis with Gene
Center for Engineering in Medicine Newsletter
 Center for Engineering in Medicine Newsletter October 200 Volume 2, Issue 2 Inside this Issue Profile & Minisymposium 2-4 Metabolic Engineering 4 Personnel Changes 5 Seminar Series Outdoor Club From the
Center for Engineering in Medicine Newsletter October 200 Volume 2, Issue 2 Inside this Issue Profile & Minisymposium 2-4 Metabolic Engineering 4 Personnel Changes 5 Seminar Series Outdoor Club From the
AGILENT SOLUTIONS FOR METABOLOMICS
 YOUR Path To Succcess AGILENT SOLUTIONS FOR METABOLOMICS Understanding METABOLOMICS Agilent is the leading global solution vendor for measuring metabolism, offering researchers a broad array of cutting-edge
YOUR Path To Succcess AGILENT SOLUTIONS FOR METABOLOMICS Understanding METABOLOMICS Agilent is the leading global solution vendor for measuring metabolism, offering researchers a broad array of cutting-edge
while Bacilli is the class to which the order Lactobacillales belongs to.
 Interactive Questions Question 1: Lactic acid bacteria belong to which order? Lactobacillaceae Lactobacillales Lactobacillales is the correct order. Lactobacillaceae is one family within this order, while
Interactive Questions Question 1: Lactic acid bacteria belong to which order? Lactobacillaceae Lactobacillales Lactobacillales is the correct order. Lactobacillaceae is one family within this order, while
Enzymes, ATP and Bioenergetics
 Harriet Wilson, Lecture Notes Bio. Sci. 4 - Microbiology Sierra College Enzymes, ATP and Bioenergetics Bioenergetics Bioenergetics can be defined as energy transfer mechanisms occurring within living organisms.
Harriet Wilson, Lecture Notes Bio. Sci. 4 - Microbiology Sierra College Enzymes, ATP and Bioenergetics Bioenergetics Bioenergetics can be defined as energy transfer mechanisms occurring within living organisms.
Optimization of enzyme parameters for fermentative production of biorenewable fuels and chemicals
 Chemical and Biological Engineering Publications Chemical and Biological Engineering 10-2012 Optimization of enzyme parameters for fermentative production of biorenewable fuels and chemicals Laura R. Jarboe
Chemical and Biological Engineering Publications Chemical and Biological Engineering 10-2012 Optimization of enzyme parameters for fermentative production of biorenewable fuels and chemicals Laura R. Jarboe
Nucleotide Metabolism Biochemistry by Lippincott pp
 Nucleotide Metabolism Biochemistry by Lippincott pp 291-306 Metabolism: CONCEPT Ø Metabolism is the totality of an organism s chemical reactions. Ø A metabolic pathway begins with a specific molecule and
Nucleotide Metabolism Biochemistry by Lippincott pp 291-306 Metabolism: CONCEPT Ø Metabolism is the totality of an organism s chemical reactions. Ø A metabolic pathway begins with a specific molecule and
Enzymes III. Dr. Kevin Ahern
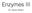 Enzymes III Dr. Kevin Ahern Enzyme Inhibition Enzyme Inhibition Competitive Inhibitor Resembles Natural Substrate and Competes with it for Binding to the Active Site Enzyme Inhibition Normal Substrate
Enzymes III Dr. Kevin Ahern Enzyme Inhibition Enzyme Inhibition Competitive Inhibitor Resembles Natural Substrate and Competes with it for Binding to the Active Site Enzyme Inhibition Normal Substrate
