The SVM2CRM data User s Guide
|
|
|
- Andrea Farmer
- 6 years ago
- Views:
Transcription
1 The SVM2CRM data User s Guide Guidantonio Malagoli Tagliazucchi, Silvio Bicciato Center for genome research University of Modena and Reggio Emilia, Modena October 31, 2017 October 31, 2017 Contents 1 Overview 1 2 CD4matrixInputSVMbin100window1000 object 2 3 trainpositive and trainnegative objects 2 4 External data 3 5 References 4 6 Session information 5 1 Overview The SVM2CRMdata package contains ChIP-seq data of methylation and acetylation maps from CD4+ T-cells([1]). The package contains also a list of p300 binding sites and two matrix contains the signals of the histone marks in correspondance of genomic regions with enhancers or background sequences. This package is useful to perform the tutorial analysis of SVM2CRM. In this tutorial we load a matrix with the signal of ChIP-seq data preprocess using cisrefindbed function of SVM2CRM package and two matrices containing the signal of the histone marks at level of putative enhancers and random genomic regions. 1
2 2 CD4matrixInputSVMbin100window1000 object This data.frame contains the signals of the histone modification from CD4+Tcells([1]) genome wide. In particular, this dataset contains 20 methylation and 17 acetylation ChIP-seq maps. We obtained this object using the function cisrefindbed from SVM2CRM package. For details about this function see the documentation in SVM2CRM package. Briefly,this function allowed the user to create one of the input require to perfomed the prediction of enhancers using SVM2CRM. In particular, this function reads external bed files of the histone modification (processed externally from R) and performed the smoothing of the signals genome-wide. When we processed this dataset we used cisrefindbed considering a bin.size of 100 bp, that is the values that we used to binarized the signals of the histone modification genome wide. Moreover the size of the window to perfomed the smoothing of the histone modification signals genome wide was set to 500, 1000, 2000 bp. This package contains the data.frame obtained using a window of 1000bp and considering only the chromosome 1. Before to perfomed the prediction of enhancers using your dataset and SVM2CRM we suggests to generate this data.frame using ad hoc script that contain only the code of cisrefindbed and that save the results of this analysis in a.rdata. This can help the user to reduce the computational time to perfomed the prediction genome-wide and avoid to create the input each time that the user want launch a new analysis (for e.g. using different parameters). > library(svm2crmdata) > setwd(system.file("data",package="svm2crmdata")) > load("cd4_matrixinputsvmbin100window1000.rda") > #The column contains the signal. The rows contains the genomic coordinate, while the > dim("cd4_matrixinputsvmbin100window1000") NULL 3 trainpositive and trainnegative objects The trainpositive and trainnegative objects were created using the function getsignal of SVM2CRM package. These data.frame contains the signals of the histone modification of 20 methylation and 17 acetylation ChIP-seq maps from ([1]) at p300 binding sites (train positive) and genomic random regions (train negative). In particular these data.frame contains the signals of the histone modifications in correspondence of p300 binding sites known 2
3 and genomic regions distant from TSSs of 1000bp. The creation of this matrices is mandatory because SVM2CRM require the presence of two datasets that contains the signals of the histone modification at level of putative enhancers (p300 binding sites) and in genomic regions different from distal cis-regulatory regions. Here we used getsignal from SVM2CRM to create respectivelly the trainpositive and trainnegative dataset. In particular, we download a list p300 binding sites and create a list of genomic-coordinates random regions using bedtools using a windows size of We used the same size of window (1000 bp) defined also to create the object described in the previously object (CD4matrixInputSVMbin100window1000).getSignal consider the signal of the histone marks around a particular windows (1000 bp) from the start of the genomic regions provided (e.g. the start of the p300 binding sites and background genomic regions). Also in this case we suggests to create this data.frame before to perfomed an entire analysis of SVM2CRM. > setwd(system.file("data",package="svm2crmdata")) > load("train_positive.rda") > load("train_negative.rda") 4 External data In SVM2CRMdata we provided ChIP-seq data from H2AK5ac, H2K23ac, H2AK9ac, H3K27ac, H3K4me1, H3K4me2, H3K4me3. This data contains the signals of the histone modification normalized using the procedure described in ([2]). The user can use these data to create the object described in the first section of this vignette. This object is one of the most important input required to perfomed the prediction of analysis with SVM2CRM. As described previously this object contains the genome wide signals of the histone modifications provided by the user. The user can create this input simple using cisrefindbed function from SVM2CRM. The parameters required are bin.size used to binnarized the signal of the histone modification (in this case 100bp), the size of the windows to consider the signals of the histone marks (each 1000 bp), and the function that the user want use to model the signals of the histone marks inside the windows (optional). The object that is generate using cisrefindbed several columns. A column with the chromosome, the start and end of the genomic windows, the signals of the histone marks inside the windows. The number of columns that contains the signals of the histone marks dependes on the window size. For details see section Introduction in the vignette of SVM2CRM 3
4 5 References References [1] Rosenfeld JA Wang Z, Zang C, Schones DE, Cuddapah S Barski A, Roh TY Cui K, Zhang MQ Peng W, and Zhao K. Combinatorial patterns of histone acetylations and methylations in the human genome. Nature Genetics, 7(40): , [2] Hiram A Firpi, Duygu Ucar, and Kai Tan. Discover regulatory dna elements using chromatin signatures and artificial neural network. Bioinformatics, 26(13): ,
5 6 Session information The output in this vignette was produced under the following conditions: > sessioninfo() R Under development (unstable) ( r73567) Platform: x86_64-pc-linux-gnu (64-bit) Running under: Ubuntu LTS Matrix products: default BLAS: /home/biocbuild/bbs-3.7-bioc/r/lib/librblas.so LAPACK: /home/biocbuild/bbs-3.7-bioc/r/lib/librlapack.so locale: [1] LC_CTYPE=en_US.UTF-8 LC_NUMERIC=C [3] LC_TIME=en_US.UTF-8 LC_COLLATE=C [5] LC_MONETARY=en_US.UTF-8 LC_MESSAGES=en_US.UTF-8 [7] LC_PAPER=en_US.UTF-8 LC_NAME=C [9] LC_ADDRESS=C LC_TELEPHONE=C [11] LC_MEASUREMENT=en_US.UTF-8 LC_IDENTIFICATION=C attached base packages: [1] stats graphics grdevices utils datasets methods base other attached packages: [1] SVM2CRMdata_ loaded via a namespace (and not attached): [1] compiler_3.5.0 tools_
SVM2CRM: support vector machine for the cis-regulatory elements detections
 SVM2CRM: support vector machine for the cis-regulatory elements detections Guidantonio Malagoli Tagliazucchi, Silvio Bicciato Center for genome research University of Modena and Reggio Emilia, Modena October
SVM2CRM: support vector machine for the cis-regulatory elements detections Guidantonio Malagoli Tagliazucchi, Silvio Bicciato Center for genome research University of Modena and Reggio Emilia, Modena October
Analysis of genomic arrays using quantile smoothing
 Analysis of genomic arrays using quantile smoothing Jan Oosting, Paul Eilers & Renee Menezes Package quantsmooth. April 30, 2018 Contents 1 Usage 1 2 Session Information 7 Introduction Genomic arrays give
Analysis of genomic arrays using quantile smoothing Jan Oosting, Paul Eilers & Renee Menezes Package quantsmooth. April 30, 2018 Contents 1 Usage 1 2 Session Information 7 Introduction Genomic arrays give
IdeoViz Plot data along chromosome ideograms
 IdeoViz Plot data along chromosome ideograms Shraddha Pai (shraddha.pai@utoronto.ca), Jingliang Ren 27 January, 217 Plotting discrete or continuous dataseries in the context of chromosomal location has
IdeoViz Plot data along chromosome ideograms Shraddha Pai (shraddha.pai@utoronto.ca), Jingliang Ren 27 January, 217 Plotting discrete or continuous dataseries in the context of chromosomal location has
DBChIP: Detecting differential binding of transcription factors with ChIP-seq
 DBChIP: Detecting differential binding of transcription factors with ChIP-seq Kun Liang 1,2 and Sündüz Keleş 1,2 1 Department of Statistics, University of Wisconsin Madison, WI 53706. 2 Department of Biostatistics
DBChIP: Detecting differential binding of transcription factors with ChIP-seq Kun Liang 1,2 and Sündüz Keleş 1,2 1 Department of Statistics, University of Wisconsin Madison, WI 53706. 2 Department of Biostatistics
IdeoViz Plot data along chromosome ideograms
 IdeoViz Plot data along chromosome ideograms Shraddha Pai (shraddha.pai@utoronto.ca), Jingliang Ren 8 December, 2017 Plotting discrete or continuous dataseries in the context of chromosomal location has
IdeoViz Plot data along chromosome ideograms Shraddha Pai (shraddha.pai@utoronto.ca), Jingliang Ren 8 December, 2017 Plotting discrete or continuous dataseries in the context of chromosomal location has
Generating Grey Lists from Input Libraries
 Gord Brown Edited: 2014-12-29; Compiled: October 30, 2018 Contents 1 Introduction.............................. 1 2 Generating a Grey List........................ 1 3 Sample Data.............................
Gord Brown Edited: 2014-12-29; Compiled: October 30, 2018 Contents 1 Introduction.............................. 1 2 Generating a Grey List........................ 1 3 Sample Data.............................
An Introduction to exomepeak
 An Introduction to exomepeak Jia Meng, PhD Modified: 21 Dec, 2015. Compiled: October 30, 2018 1 Introduction The exomepeak R-package has been developed based on the MATLAB exome- Peak package, for the
An Introduction to exomepeak Jia Meng, PhD Modified: 21 Dec, 2015. Compiled: October 30, 2018 1 Introduction The exomepeak R-package has been developed based on the MATLAB exome- Peak package, for the
Using DeMAND, a package for Interrogating Drug Mechanism of Action using Network Dysregulation analysis
 Using DeMAND, a package for Interrogating Drug Mechanism of Action using Network Dysregulation analysis Jung Hoon Woo, Yishai Shimoni, Mukesh Bansal, and Andrea Califano Columbia University, New York,
Using DeMAND, a package for Interrogating Drug Mechanism of Action using Network Dysregulation analysis Jung Hoon Woo, Yishai Shimoni, Mukesh Bansal, and Andrea Califano Columbia University, New York,
DBChIP: Detecting differential binding of transcription factors with ChIP-seq
 DBChIP: Detecting differential binding of transcription factors with ChIP-seq Kun Liang 1,2 and Sündüz Keleş 1,2 1 Department of Statistics, University of Wisconsin Madison, WI 53706. 2 Department of Biostatistics
DBChIP: Detecting differential binding of transcription factors with ChIP-seq Kun Liang 1,2 and Sündüz Keleş 1,2 1 Department of Statistics, University of Wisconsin Madison, WI 53706. 2 Department of Biostatistics
User s guide to flowmap
 Chiaowen Joyce Hsiao 1, Yu Qian 2, Richard H. Scheuermann 2 [1em] 1 Center for Bioinformatics and Computational Biology (CBCB) and and Applied Mathematics and Scientific Computation, University of Maryland,
Chiaowen Joyce Hsiao 1, Yu Qian 2, Richard H. Scheuermann 2 [1em] 1 Center for Bioinformatics and Computational Biology (CBCB) and and Applied Mathematics and Scientific Computation, University of Maryland,
pint: probabilistic data integration for functional genomics
 pint: probabilistic data integration for functional genomics Olli-Pekka Huovilainen 1* and Leo Lahti 1,2 (1) Dpt. Information and Computer Science, Aalto University, Finland (2) Dpt. Veterinary Bioscience,
pint: probabilistic data integration for functional genomics Olli-Pekka Huovilainen 1* and Leo Lahti 1,2 (1) Dpt. Information and Computer Science, Aalto University, Finland (2) Dpt. Veterinary Bioscience,
Exploring protein interaction data using ppistats
 Exploring protein interaction data using ppistats Denise Scholtens and Tony Chiang Introduction ppistats is a package that provides tools for the description and analysis of protein interaction data represented
Exploring protein interaction data using ppistats Denise Scholtens and Tony Chiang Introduction ppistats is a package that provides tools for the description and analysis of protein interaction data represented
GEOsubmission. Alexandre Kuhn October 30, 2017
 GEOsubmission Alexandre Kuhn (kuhnam@mail.nih.gov) October 30, 2017 1 Summary The goal of GEOsubmission is to ease the submission of microarray datasets to the GEO repository. It generates a single file
GEOsubmission Alexandre Kuhn (kuhnam@mail.nih.gov) October 30, 2017 1 Summary The goal of GEOsubmission is to ease the submission of microarray datasets to the GEO repository. It generates a single file
Introduction to the Data Analysis of the Roche xcelligence System with RTCA Package
 Introduction to the Data Analysis of the Roche xcelligence System with RTCA Package Jitao David Zhang October 30, 2017 1 Introduction to the xcelligence System The xcelligence System, also known as RT-CES
Introduction to the Data Analysis of the Roche xcelligence System with RTCA Package Jitao David Zhang October 30, 2017 1 Introduction to the xcelligence System The xcelligence System, also known as RT-CES
ichip: A Package for Analyzing Multi-platform ChIP-chip data with Various Sample Sizes
 ichip: A Package for Analyzing Multi-platform ChIP-chip data with Various Sample Sizes Qianxing Mo October 30, 2018 Contents Department of Biostatistics & Bioinformatics H. Lee Moffitt Cancer Center &
ichip: A Package for Analyzing Multi-platform ChIP-chip data with Various Sample Sizes Qianxing Mo October 30, 2018 Contents Department of Biostatistics & Bioinformatics H. Lee Moffitt Cancer Center &
immunoclust - Automated Pipeline for Population Detection in Flow Cytometry
 - Automated Pipeline for Population Detection in Flow Cytometry Till Sörensen 1 1 tillantoni.soerensen@charite.de June 1, 18 Contents 1 Licensing............................... Overview................................
- Automated Pipeline for Population Detection in Flow Cytometry Till Sörensen 1 1 tillantoni.soerensen@charite.de June 1, 18 Contents 1 Licensing............................... Overview................................
immunoclust - Automated Pipeline for Population Detection in Flow Cytometry
 immunoclust - Automated Pipeline for Population Detection in Flow Cytometry Till Sörensen * April 24, 2017 Contents 1 Licensing 2 2 Overview 2 3 Getting started 2 4 Example Illustrating the immunoclust
immunoclust - Automated Pipeline for Population Detection in Flow Cytometry Till Sörensen * April 24, 2017 Contents 1 Licensing 2 2 Overview 2 3 Getting started 2 4 Example Illustrating the immunoclust
An Introduction Mixture Model Normalization with the metabomxtr Package
 An Introduction Mixture Model Normalization with the metabomxtr Package Michael Nodzenski, Anna C. Reisetter, Denise M. Scholtens April 0, 0 Introduction Controlling technical variability in metabolite
An Introduction Mixture Model Normalization with the metabomxtr Package Michael Nodzenski, Anna C. Reisetter, Denise M. Scholtens April 0, 0 Introduction Controlling technical variability in metabolite
Using the attract Package To Find the Gene Expression Modules That Represent The Drivers of Kauffman s Attractor Landscape
 Using the attract Package To Find the Gene Expression Modules That Represent The Drivers of Kauffman s Attractor Landscape Jessica Mar January 5, 2019 1 Introduction A mammalian organism is made up of
Using the attract Package To Find the Gene Expression Modules That Represent The Drivers of Kauffman s Attractor Landscape Jessica Mar January 5, 2019 1 Introduction A mammalian organism is made up of
msgbsr: an R package to analyse methylation sensitive genotyping by sequencing (MS-GBS) data
 msgbsr: an R package to analyse methylation sensitive genotyping by sequencing (MS-GBS) data Benjamin Mayne October 30, 2018 Contents 1 Introduction 2 2 Reading data into R 2 3 Confirmation of correct
msgbsr: an R package to analyse methylation sensitive genotyping by sequencing (MS-GBS) data Benjamin Mayne October 30, 2018 Contents 1 Introduction 2 2 Reading data into R 2 3 Confirmation of correct
immunoclust - Automated Pipeline for Population Detection in Flow Cytometry
 immunoclust - Automated Pipeline for Population Detection in Flow Cytometry Till Sörensen April 16, 2015 Contents 1 Licensing 2 2 Overview 2 3 Getting started 2 4 Example Illustrating the immunoclust Pipeline
immunoclust - Automated Pipeline for Population Detection in Flow Cytometry Till Sörensen April 16, 2015 Contents 1 Licensing 2 2 Overview 2 3 Getting started 2 4 Example Illustrating the immunoclust Pipeline
daglogo guide Jianhong Ou, Lihua Julie Zhu April 16, Introduction 1 4 using daglogo to analysis Catobolite Activator Protein 9 5 References 11
 daglogo guide Jianhong Ou, Lihua Julie Zhu April 16, 15 Contents 1 Introduction 1 2 Prepare environment 2 3 Examples of using daglogo 2 3.1 Step 1, fetch sequences.................................. 2 3.2
daglogo guide Jianhong Ou, Lihua Julie Zhu April 16, 15 Contents 1 Introduction 1 2 Prepare environment 2 3 Examples of using daglogo 2 3.1 Step 1, fetch sequences.................................. 2 3.2
The CopyNumber450k User s Guide: Calling CNVs from Illumina 450k Methylation Arrays
 The CopyNumber450k User s Guide: Calling CNVs from Illumina 450k Methylation Arrays Simon Papillon-Cavanagh 1,3, Jean-Philippe Fortin 2, and Nicolas De Jay 3 1 McGill University and Genome Quebec Innovation
The CopyNumber450k User s Guide: Calling CNVs from Illumina 450k Methylation Arrays Simon Papillon-Cavanagh 1,3, Jean-Philippe Fortin 2, and Nicolas De Jay 3 1 McGill University and Genome Quebec Innovation
mirnatap example use
 mirnatap example use Maciej Pajak, Ian Simpson April 30, 2018 Contents 1 Introduction 2 2 Installation 2 3 Workflow 3 4 Session Information 5 References 6 1 1 Introduction mirnatap package is designed
mirnatap example use Maciej Pajak, Ian Simpson April 30, 2018 Contents 1 Introduction 2 2 Installation 2 3 Workflow 3 4 Session Information 5 References 6 1 1 Introduction mirnatap package is designed
prot2d : Statistical Tools for volume data from 2D Gel Electrophoresis
 prot2d : Statistical Tools for volume data from 2D Gel Electrophoresis Sébastien Artigaud * April 30, 2018 Contents 1 Introduction 1 2 Input data for prot2d 2 3 Getting started 2 4 prot2d workflow 2 4.1
prot2d : Statistical Tools for volume data from 2D Gel Electrophoresis Sébastien Artigaud * April 30, 2018 Contents 1 Introduction 1 2 Input data for prot2d 2 3 Getting started 2 4 prot2d workflow 2 4.1
MGFM: Marker Gene Finder in Microarray gene expression data
 MGFM: Marker Gene Finder in Microarray gene expression data Khadija El Amrani * June 13, 2018 Contents 1 Introduction 1 2 Requirements 2 3 Contents of the package 2 3.1 getmarkergenes.....................................
MGFM: Marker Gene Finder in Microarray gene expression data Khadija El Amrani * June 13, 2018 Contents 1 Introduction 1 2 Requirements 2 3 Contents of the package 2 3.1 getmarkergenes.....................................
1 curatedbladderdata: Clinically Annotated Data for the Bladder Cancer Transcriptome... 1
 Markus Riester November 1, 2018 Contents 1 : Clinically Annotated Data for the Bladder Cancer Transcriptome........................ 1 2 Load data sets............................ 1 3 Load datasets based
Markus Riester November 1, 2018 Contents 1 : Clinically Annotated Data for the Bladder Cancer Transcriptome........................ 1 2 Load data sets............................ 1 3 Load datasets based
methyanalysis: an R package for DNA methylation data analysis and visualization
 methyanalysis: an R package for DNA methylation data analysis and visualization Pan Du and Richard Bourgon April 12, 2014 Department of Bioinformatics and Computational Biology Genentech Inc., South San
methyanalysis: an R package for DNA methylation data analysis and visualization Pan Du and Richard Bourgon April 12, 2014 Department of Bioinformatics and Computational Biology Genentech Inc., South San
traser: TRait-Associated SNP EnRichment analyses
 traser: TRait-Associated SNP EnRichment analyses Li Chen, Zhaohui S.Qin [1em]Department of Biostatistics and Bioinformatics Emory University Atlanta, GA 303022 [1em] li.chen@emory.edu,zhaohui.qin@emory.edu
traser: TRait-Associated SNP EnRichment analyses Li Chen, Zhaohui S.Qin [1em]Department of Biostatistics and Bioinformatics Emory University Atlanta, GA 303022 [1em] li.chen@emory.edu,zhaohui.qin@emory.edu
The REDseq user s guide
 The REDseq user s guide Lihua Julie Zhu * April 30, 2018 Contents 1 Introduction 1 2 Examples of using REDseq 2 2.1 Task 1: Build a RE map for a genome..................... 2 2.2 Task 2: Assign mapped
The REDseq user s guide Lihua Julie Zhu * April 30, 2018 Contents 1 Introduction 1 2 Examples of using REDseq 2 2.1 Task 1: Build a RE map for a genome..................... 2 2.2 Task 2: Assign mapped
The REDseq user s guide
 The REDseq user s guide Lihua Julie Zhu * June 13, 2018 Contents 1 Introduction 1 2 Examples of using REDseq 2 2.1 Task 1: Build a RE map for a genome..................... 2 2.2 Task 2: Assign mapped sequence
The REDseq user s guide Lihua Julie Zhu * June 13, 2018 Contents 1 Introduction 1 2 Examples of using REDseq 2 2.1 Task 1: Build a RE map for a genome..................... 2 2.2 Task 2: Assign mapped sequence
Finding Chimeric Sequences
 Finding Chimeric Sequences Erik S. Wright October 30, 2018 Contents 1 Introduction 1 2 Chimera Finding Problems 1 3 Getting Started 2 4 Obtaining/Building A Reference Database 2 5 Steps to Find Chimeras
Finding Chimeric Sequences Erik S. Wright October 30, 2018 Contents 1 Introduction 1 2 Chimera Finding Problems 1 3 Getting Started 2 4 Obtaining/Building A Reference Database 2 5 Steps to Find Chimeras
GeneExpressionSignature: Computing pairwise distances between different biological states
 GeneExpressionSignature: Computing pairwise distances between different biological states Yang Cao, Lu Han, Fei Li, Xiaochen Bo October 30, 2017 Contents 1 Introduction 1 2 Getting Started 2 2.1 Data Ranking.............................
GeneExpressionSignature: Computing pairwise distances between different biological states Yang Cao, Lu Han, Fei Li, Xiaochen Bo October 30, 2017 Contents 1 Introduction 1 2 Getting Started 2 2.1 Data Ranking.............................
Design Microarray Probes
 Design Microarray Probes Erik S. Wright October 30, 2017 Contents 1 Introduction 1 2 Getting Started 1 2.1 Startup................................................ 1 2.2 Creating a Sequence Database....................................
Design Microarray Probes Erik S. Wright October 30, 2017 Contents 1 Introduction 1 2 Getting Started 1 2.1 Startup................................................ 1 2.2 Creating a Sequence Database....................................
mirnatap example use
 mirnatap example use Maciej Pajak, Ian Simpson September 9, 2015 Contents 1 Introduction 2 2 Installation 2 3 Workflow 2 4 Session Information 5 References 6 1 1 Introduction mirnatap package is designed
mirnatap example use Maciej Pajak, Ian Simpson September 9, 2015 Contents 1 Introduction 2 2 Installation 2 3 Workflow 2 4 Session Information 5 References 6 1 1 Introduction mirnatap package is designed
cogena, a workflow for co-expressed gene-set enrichment analysis Zhilong Jia, Michael R. Barnes
 cogena, a workflow for co-expressed gene-set enrichment analysis Zhilong Jia, Michael R. Barnes 2018-04-30 Contents 1 Abstract 1 2 A quick start 2 3 Data Input 2 3.1 Input data required..........................................
cogena, a workflow for co-expressed gene-set enrichment analysis Zhilong Jia, Michael R. Barnes 2018-04-30 Contents 1 Abstract 1 2 A quick start 2 3 Data Input 2 3.1 Input data required..........................................
The ChIP-Seq project. Giovanna Ambrosini, Philipp Bucher. April 19, 2010 Lausanne. EPFL-SV Bucher Group
 The ChIP-Seq project Giovanna Ambrosini, Philipp Bucher EPFL-SV Bucher Group April 19, 2010 Lausanne Overview Focus on technical aspects Description of applications (C programs) Where to find binaries,
The ChIP-Seq project Giovanna Ambrosini, Philipp Bucher EPFL-SV Bucher Group April 19, 2010 Lausanne Overview Focus on technical aspects Description of applications (C programs) Where to find binaries,
Human Fibroblast IMR90 Hi-C Data (Dixon et al.)
 Human Fibroblast IMR90 Hi-C Data (Dixon et al.) Nicolas Servant October 31, 2017 1 Introduction The Hi-C technic was first introduced by Lieberman-Aiden et al. [2009]. In the continuity with 3C, 4C and
Human Fibroblast IMR90 Hi-C Data (Dixon et al.) Nicolas Servant October 31, 2017 1 Introduction The Hi-C technic was first introduced by Lieberman-Aiden et al. [2009]. In the continuity with 3C, 4C and
prot2d : Statistical Tools for volume data from 2D Gel Electrophoresis
 prot2d : Statistical Tools for volume data from 2D Gel Electrophoresis Sébastien Artigaud * October 16, 2018 Contents 1 Introduction 1 2 Input data for prot2d 2 3 Getting started 2 4 prot2d workflow 2
prot2d : Statistical Tools for volume data from 2D Gel Electrophoresis Sébastien Artigaud * October 16, 2018 Contents 1 Introduction 1 2 Input data for prot2d 2 3 Getting started 2 4 prot2d workflow 2
FlipFlop: Fast Lasso-based Isoform Prediction as a Flow Problem
 FlipFlop: Fast Lasso-based Isoform Prediction as a Flow Problem Elsa Bernard Laurent Jacob Julien Mairal Jean-Philippe Vert October 30, 2017 Abstract FlipFlop implements a fast method for de novo transcript
FlipFlop: Fast Lasso-based Isoform Prediction as a Flow Problem Elsa Bernard Laurent Jacob Julien Mairal Jean-Philippe Vert October 30, 2017 Abstract FlipFlop implements a fast method for de novo transcript
Bioconductor mapkl Package
 Argiris Sakellariou 1,2 [1cm] 1 Biomedical Informatics Unit, Biomedical Research Foundation of the Academy of Athens, Athens, Greece [0cm] 2 Department of Informatics and Telecommunications, National and
Argiris Sakellariou 1,2 [1cm] 1 Biomedical Informatics Unit, Biomedical Research Foundation of the Academy of Athens, Athens, Greece [0cm] 2 Department of Informatics and Telecommunications, National and
Measuring Protein-DNA interactions
 Measuring Protein-DNA interactions How is Biological Complexity Achieved? Mediated by Transcription Factors (TFs) 2 Transcription Factors are genetic switches 3 Regulation of Gene Expression by Transcription
Measuring Protein-DNA interactions How is Biological Complexity Achieved? Mediated by Transcription Factors (TFs) 2 Transcription Factors are genetic switches 3 Regulation of Gene Expression by Transcription
flowvs: Variance stabilization in flow cytometry (and microarrays)
 flowvs: Variance stabilization in flow cytometry (and microarrays) Ariful Azad, Bartek Rajwa, Alex Pothen January 4, 2019 Email: azad@lbl.gov Contents 1 Licensing 2 2 Variance stabilization in flow cytometry
flowvs: Variance stabilization in flow cytometry (and microarrays) Ariful Azad, Bartek Rajwa, Alex Pothen January 4, 2019 Email: azad@lbl.gov Contents 1 Licensing 2 2 Variance stabilization in flow cytometry
flowvs: Variance stabilization in flow cytometry (and microarrays)
 flowvs: Variance stabilization in flow cytometry (and microarrays) Ariful Azad, Bartek Rajwa, Alex Pothen January 5, 2019 Email: azad@lbl.gov Contents 1 Licensing 2 2 Variance stabilization in flow cytometry
flowvs: Variance stabilization in flow cytometry (and microarrays) Ariful Azad, Bartek Rajwa, Alex Pothen January 5, 2019 Email: azad@lbl.gov Contents 1 Licensing 2 2 Variance stabilization in flow cytometry
A Brief History. Bootstrapping. Bagging. Boosting (Schapire 1989) Adaboost (Schapire 1995)
 A Brief History Bootstrapping Bagging Boosting (Schapire 1989) Adaboost (Schapire 1995) What s So Good About Adaboost Improves classification accuracy Can be used with many different classifiers Commonly
A Brief History Bootstrapping Bagging Boosting (Schapire 1989) Adaboost (Schapire 1995) What s So Good About Adaboost Improves classification accuracy Can be used with many different classifiers Commonly
Oscope: a statistical pipeline for identifying oscillatory genes in unsynchronized single cell RNA-seq experiments
 Oscope: a statistical pipeline for identifying oscillatory genes in unsynchronized single cell RNA-seq experiments Ning Leng and Christina Kendziorski June 13, 2018 Contents 1 Introduction..............................
Oscope: a statistical pipeline for identifying oscillatory genes in unsynchronized single cell RNA-seq experiments Ning Leng and Christina Kendziorski June 13, 2018 Contents 1 Introduction..............................
Oscope: a statistical pipeline for identifying oscillatory genes in unsynchronized single cell RNA-seq experiments
 Oscope: a statistical pipeline for identifying oscillatory genes in unsynchronized single cell RNA-seq experiments Ning Leng and Christina Kendziorski October 30, 2018 Contents 1 Introduction..............................
Oscope: a statistical pipeline for identifying oscillatory genes in unsynchronized single cell RNA-seq experiments Ning Leng and Christina Kendziorski October 30, 2018 Contents 1 Introduction..............................
methyanalysis: an R package for DNA methylation data analysis and visualization
 methyanalysis: an R package for DNA methylation data analysis and visualization Pan Du * and Richard Bourgon June 13, 2018 Department of Bioinformatics and Computational Biology Genentech Inc., South San
methyanalysis: an R package for DNA methylation data analysis and visualization Pan Du * and Richard Bourgon June 13, 2018 Department of Bioinformatics and Computational Biology Genentech Inc., South San
seqpattern: an R package for visualisation of oligonucleotide sequence patterns and motif occurrences
 : an R package for visualisation of oligonucleotide sequence patterns and motif occurrences Vanja Haberle 1 1 vanja.haberle@gm May 8, 2015 Contents 1 Introduction.................................... 2
: an R package for visualisation of oligonucleotide sequence patterns and motif occurrences Vanja Haberle 1 1 vanja.haberle@gm May 8, 2015 Contents 1 Introduction.................................... 2
ChIP-seq data analysis with Chipster. Eija Korpelainen CSC IT Center for Science, Finland
 ChIP-seq data analysis with Chipster Eija Korpelainen CSC IT Center for Science, Finland chipster@csc.fi What will I learn? Short introduction to ChIP-seq Analyzing ChIP-seq data Central concepts Analysis
ChIP-seq data analysis with Chipster Eija Korpelainen CSC IT Center for Science, Finland chipster@csc.fi What will I learn? Short introduction to ChIP-seq Analyzing ChIP-seq data Central concepts Analysis
methyanalysis: an R package for DNA methylation data analysis and visualization
 methyanalysis: an R package for DNA methylation data analysis and visualization Pan Du * and Richard Bourgon October 30, 2017 Department of Bioinformatics and Computational Biology Genentech Inc., South
methyanalysis: an R package for DNA methylation data analysis and visualization Pan Du * and Richard Bourgon October 30, 2017 Department of Bioinformatics and Computational Biology Genentech Inc., South
seqpattern: an R package for visualisation of oligonucleotide sequence patterns and motif occurrences
 : an R package for visualisation of oligonucleotide sequence patterns and motif occurrences Vanja Haberle 1 1 vanja.haberle@gm May 8, 2015 Contents 1 Introduction.................................... 2
: an R package for visualisation of oligonucleotide sequence patterns and motif occurrences Vanja Haberle 1 1 vanja.haberle@gm May 8, 2015 Contents 1 Introduction.................................... 2
Transcription start site classification
 Transcription start site classification Max Libbrecht, Matt Fisher, Roy Frostig, Hrysoula Papadakis, Anshul Kundaje, Serafim Batzoglou December 11, 2009 Abstract Understanding the mechanisms of gene expression
Transcription start site classification Max Libbrecht, Matt Fisher, Roy Frostig, Hrysoula Papadakis, Anshul Kundaje, Serafim Batzoglou December 11, 2009 Abstract Understanding the mechanisms of gene expression
Design Primers That Yield Group-Specific Signatures
 Design Primers That Yield Group-Specific Signatures Erik S. Wright October 30, 2017 Contents 1 Introduction 1 2 Getting Started 2 2.1 Startup................................................ 2 2.2 Creating
Design Primers That Yield Group-Specific Signatures Erik S. Wright October 30, 2017 Contents 1 Introduction 1 2 Getting Started 2 2.1 Startup................................................ 2 2.2 Creating
Bayesian Inference of Regulatory influence on Expression (birte)
 Bayesian Inference of Regulatory influence on Expression (birte) Holger Fröhlich October 30, 2017 1 Introduction Expression levels of mrna is regulated by different processes, comprising inhibition or
Bayesian Inference of Regulatory influence on Expression (birte) Holger Fröhlich October 30, 2017 1 Introduction Expression levels of mrna is regulated by different processes, comprising inhibition or
I/O Suite, VCF (1000 Genome) and HapMap
 I/O Suite, VCF (1000 Genome) and HapMap Hin-Tak Leung April 13, 2013 Contents 1 Introduction 1 1.1 Ethnic Composition of 1000G vs HapMap........................ 2 2 1000 Genome vs HapMap YRI (Africans)
I/O Suite, VCF (1000 Genome) and HapMap Hin-Tak Leung April 13, 2013 Contents 1 Introduction 1 1.1 Ethnic Composition of 1000G vs HapMap........................ 2 2 1000 Genome vs HapMap YRI (Africans)
Opening up the blackbox: An interpretable deep neural network-based classifier for cell-type specific enhancer predictions
 Opening up the blackbox: An interpretable deep neural network-based classifier for cell-type specific enhancer predictions Seong Gon Kim*, Nawanol Theera-Ampornpunt*, Chih-Hao Fang, Mrudul Harwani, Ananth
Opening up the blackbox: An interpretable deep neural network-based classifier for cell-type specific enhancer predictions Seong Gon Kim*, Nawanol Theera-Ampornpunt*, Chih-Hao Fang, Mrudul Harwani, Ananth
Go to Bottom Left click WashU Epigenome Browser. Click
 Now you are going to look at the Human Epigenome Browswer. It has a more sophisticated but weirder interface than the UCSC Genome Browser. All the data that you will view as tracks is in reality just files
Now you are going to look at the Human Epigenome Browswer. It has a more sophisticated but weirder interface than the UCSC Genome Browser. All the data that you will view as tracks is in reality just files
What we ll do today. Types of stem cells. Do engineered ips and ES cells have. What genes are special in stem cells?
 Do engineered ips and ES cells have similar molecular signatures? What we ll do today Research questions in stem cell biology Comparing expression and epigenetics in stem cells asuring gene expression
Do engineered ips and ES cells have similar molecular signatures? What we ll do today Research questions in stem cell biology Comparing expression and epigenetics in stem cells asuring gene expression
Do engineered ips and ES cells have similar molecular signatures?
 Do engineered ips and ES cells have similar molecular signatures? Comparing expression and epigenetics in stem cells George Bell, Ph.D. Bioinformatics and Research Computing 2012 Spring Lecture Series
Do engineered ips and ES cells have similar molecular signatures? Comparing expression and epigenetics in stem cells George Bell, Ph.D. Bioinformatics and Research Computing 2012 Spring Lecture Series
A Supervised Learning Approach to the Prediction of Hi-C Data
 A Supervised Learning Approach to the Prediction of Hi-C Data Tyler Derr YUE Lab The Department of Biochemistry & Molecular Biology Institute for Personalized Medicine Pennsylvania State University College
A Supervised Learning Approach to the Prediction of Hi-C Data Tyler Derr YUE Lab The Department of Biochemistry & Molecular Biology Institute for Personalized Medicine Pennsylvania State University College
The ENCODE Encyclopedia. & Variant Annotation Using RegulomeDB and HaploReg
 The ENCODE Encyclopedia & Variant Annotation Using RegulomeDB and HaploReg Jill E. Moore Weng Lab University of Massachusetts Medical School October 10, 2015 Where s the Encyclopedia? ENCODE: Encyclopedia
The ENCODE Encyclopedia & Variant Annotation Using RegulomeDB and HaploReg Jill E. Moore Weng Lab University of Massachusetts Medical School October 10, 2015 Where s the Encyclopedia? ENCODE: Encyclopedia
The opossom Package. Henry Löffler-Wirth, Martin Kalcher. October 30, 2017
 The opossom Package Henry Löffler-Wirth, Martin Kalcher October 30, 2017 High-throughput technologies such as whole genome transcriptional profiling revolutionized molecular biology and provide an incredible
The opossom Package Henry Löffler-Wirth, Martin Kalcher October 30, 2017 High-throughput technologies such as whole genome transcriptional profiling revolutionized molecular biology and provide an incredible
Supplemental Figure 1.
 Supplemental Data. Charron et al. Dynamic landscapes of four histone modifications during de-etiolation in Arabidopsis. Plant Cell (2009). 10.1105/tpc.109.066845 Supplemental Figure 1. Immunodetection
Supplemental Data. Charron et al. Dynamic landscapes of four histone modifications during de-etiolation in Arabidopsis. Plant Cell (2009). 10.1105/tpc.109.066845 Supplemental Figure 1. Immunodetection
goseq: Gene Ontology testing for RNA-seq datasets
 goseq: Gene Ontology testing for RNA-seq datasets Matthew D. Young Nadia Davidson Matthew J. Wakefield Gordon K. Smyth Alicia Oshlack 8 September 2017 1 Introduction This document gives an introduction
goseq: Gene Ontology testing for RNA-seq datasets Matthew D. Young Nadia Davidson Matthew J. Wakefield Gordon K. Smyth Alicia Oshlack 8 September 2017 1 Introduction This document gives an introduction
goseq: Gene Ontology testing for RNA-seq datasets
 goseq: Gene Ontology testing for RNA-seq datasets Matthew D. Young Nadia Davidson Matthew J. Wakefield Gordon K. Smyth Alicia Oshlack 8 September 2017 1 Introduction This document gives an introduction
goseq: Gene Ontology testing for RNA-seq datasets Matthew D. Young Nadia Davidson Matthew J. Wakefield Gordon K. Smyth Alicia Oshlack 8 September 2017 1 Introduction This document gives an introduction
Bioinformatics of Transcriptional Regulation
 Bioinformatics of Transcriptional Regulation Carl Herrmann IPMB & DKFZ c.herrmann@dkfz.de Wechselwirkung von Maßnahmen und Auswirkungen Einflussmöglichkeiten in einem Dialog From genes to active compounds
Bioinformatics of Transcriptional Regulation Carl Herrmann IPMB & DKFZ c.herrmann@dkfz.de Wechselwirkung von Maßnahmen und Auswirkungen Einflussmöglichkeiten in einem Dialog From genes to active compounds
RIPSeeker: a statistical package for identifying protein-associated transcripts from RIP-seq experiments
 RIPSeeker: a statistical package for identifying protein-associated transcripts from RIP-seq experiments Yue Li yueli@cs.toronto.edu April 30, 2018 1 Introduction Ribonucleoprotein (RNP) immunoprecipitation
RIPSeeker: a statistical package for identifying protein-associated transcripts from RIP-seq experiments Yue Li yueli@cs.toronto.edu April 30, 2018 1 Introduction Ribonucleoprotein (RNP) immunoprecipitation
Decoding Chromatin States with Epigenome Data Advanced Topics in Computa8onal Genomics
 Decoding Chromatin States with Epigenome Data 02-715 Advanced Topics in Computa8onal Genomics HMMs for Decoding Chromatin States Epigene8c modifica8ons of the genome have been associated with Establishing
Decoding Chromatin States with Epigenome Data 02-715 Advanced Topics in Computa8onal Genomics HMMs for Decoding Chromatin States Epigene8c modifica8ons of the genome have been associated with Establishing
Gaussian derivative wavelets identify dynamic changes in histone modification
 Gaussian derivative wavelets identify dynamic changes in histone modification Nha Nguyen Department of Genetics, Institute for Diabetes, Obesity and Metabolism, School of Medicine, University of Pennsylvania,
Gaussian derivative wavelets identify dynamic changes in histone modification Nha Nguyen Department of Genetics, Institute for Diabetes, Obesity and Metabolism, School of Medicine, University of Pennsylvania,
Bayesian Inference of Regulatory influence on Expression (birte)
 Bayesian Inference of Regulatory influence on Expression (birte) Holger Fröhlich October 30, 2017 1 Introduction Expression levels of mrna is regulated by different processes, comprising inhibition or
Bayesian Inference of Regulatory influence on Expression (birte) Holger Fröhlich October 30, 2017 1 Introduction Expression levels of mrna is regulated by different processes, comprising inhibition or
Introduction to iplant Collaborative Jinyu Yang Bioinformatics and Mathematical Biosciences Lab
 Introduction to iplant Collaborative Jinyu Yang Bioinformatics and Mathematical Biosciences Lab May/27 th /2016 About iplant What is iplant Collaborative? 1. A virtual organization 2. Created by a cooperative
Introduction to iplant Collaborative Jinyu Yang Bioinformatics and Mathematical Biosciences Lab May/27 th /2016 About iplant What is iplant Collaborative? 1. A virtual organization 2. Created by a cooperative
ChromaSig: A Probabilistic Approach to Finding Common Chromatin Signatures in the Human Genome
 : A Probabilistic Approach to Finding Common Chromatin Signatures in the Human Genome Gary Hon 1,2, Bing Ren 1,2,3 *, Wei Wang 1,4 * 1 Bioinformatics Program, University of California San Diego, La Jolla,
: A Probabilistic Approach to Finding Common Chromatin Signatures in the Human Genome Gary Hon 1,2, Bing Ren 1,2,3 *, Wei Wang 1,4 * 1 Bioinformatics Program, University of California San Diego, La Jolla,
Gene expression analysis. Biosciences 741: Genomics Fall, 2013 Week 5. Gene expression analysis
 Gene expression analysis Biosciences 741: Genomics Fall, 2013 Week 5 Gene expression analysis From EST clusters to spotted cdna microarrays Long vs. short oligonucleotide microarrays vs. RT-PCR Methods
Gene expression analysis Biosciences 741: Genomics Fall, 2013 Week 5 Gene expression analysis From EST clusters to spotted cdna microarrays Long vs. short oligonucleotide microarrays vs. RT-PCR Methods
Deep learning frameworks for regulatory genomics and epigenomics
 Deep learning frameworks for regulatory genomics and epigenomics Chuan Sheng Foo Avanti Shrikumar Nicholas Sinnott- Armstrong ANSHUL KUNDAJE Genetics, Computer science Stanford University Johnny Israeli
Deep learning frameworks for regulatory genomics and epigenomics Chuan Sheng Foo Avanti Shrikumar Nicholas Sinnott- Armstrong ANSHUL KUNDAJE Genetics, Computer science Stanford University Johnny Israeli
Genome 541! Unit 4, lecture 3! Genomics assays
 Genome 541! Unit 4, lecture 3! Genomics assays I d like a bit more background on the assays and bioterminology.!! The phantom peak concept was confusing.! I didn t quite understand what the phantom peak
Genome 541! Unit 4, lecture 3! Genomics assays I d like a bit more background on the assays and bioterminology.!! The phantom peak concept was confusing.! I didn t quite understand what the phantom peak
CS273B: Deep learning for Genomics and Biomedicine
 CS273B: Deep learning for Genomics and Biomedicine Lecture 2: Convolutional neural networks and applications to functional genomics 09/28/2016 Anshul Kundaje, James Zou, Serafim Batzoglou Outline Anatomy
CS273B: Deep learning for Genomics and Biomedicine Lecture 2: Convolutional neural networks and applications to functional genomics 09/28/2016 Anshul Kundaje, James Zou, Serafim Batzoglou Outline Anatomy
TargetScore: Infer microrna targets using microrna-overexpression data and sequence information (Li et al. [2013])
![TargetScore: Infer microrna targets using microrna-overexpression data and sequence information (Li et al. [2013]) TargetScore: Infer microrna targets using microrna-overexpression data and sequence information (Li et al. [2013])](/thumbs/81/84144159.jpg) TargetScore: Infer microrna targets using microrna-overexpression data and sequence information (Li et al. [2013]) Yue Li yueli@cs.toronto.edu April 30, 2018 1 Introduction MicroRNAs (mirnas) are known
TargetScore: Infer microrna targets using microrna-overexpression data and sequence information (Li et al. [2013]) Yue Li yueli@cs.toronto.edu April 30, 2018 1 Introduction MicroRNAs (mirnas) are known
APPLICATION NOTE. Abstract. Introduction
 From minuscule amounts to magnificent results: reliable ChIP-seq data from 1, cells with the True MicroChIP and the MicroPlex Library Preparation kits Abstract Diagenode has developed groundbreaking solutions
From minuscule amounts to magnificent results: reliable ChIP-seq data from 1, cells with the True MicroChIP and the MicroPlex Library Preparation kits Abstract Diagenode has developed groundbreaking solutions
goseq: Gene Ontology testing for RNA-seq datasets
 goseq: Gene Ontology testing for RNA-seq datasets Matthew D. Young myoung@wehi.edu.au Matthew J. Wakefield Alicia Oshlack 3 March 2011 Gordon K. Smyth 1 Introduction This document gives an introduction
goseq: Gene Ontology testing for RNA-seq datasets Matthew D. Young myoung@wehi.edu.au Matthew J. Wakefield Alicia Oshlack 3 March 2011 Gordon K. Smyth 1 Introduction This document gives an introduction
Bio5488 Practice Midterm (2018) 1. Next-gen sequencing
 1. Next-gen sequencing 1. You have found a new strain of yeast that makes fantastic wine. You d like to sequence this strain to ascertain the differences from S. cerevisiae. To accurately call a base pair,
1. Next-gen sequencing 1. You have found a new strain of yeast that makes fantastic wine. You d like to sequence this strain to ascertain the differences from S. cerevisiae. To accurately call a base pair,
Package DBChIP. May 12, 2018
 Type Package Package May 12, 2018 Title Differential Binding of Transcription Factor with ChIP-seq Version 1.24.0 Date 2012-06-21 Author Kun Liang Maintainer Kun Liang Depends R
Type Package Package May 12, 2018 Title Differential Binding of Transcription Factor with ChIP-seq Version 1.24.0 Date 2012-06-21 Author Kun Liang Maintainer Kun Liang Depends R
HebbPlot: an intelligent tool for learning and visualizing chromatin mark signatures
 Girgis et al. BMC Bioinformatics (2018) 19:310 https://doi.org/10.1186/s12859-018-2312-1 SOFTWARE Open Access HebbPlot: an intelligent tool for learning and visualizing chromatin mark signatures Hani Z.
Girgis et al. BMC Bioinformatics (2018) 19:310 https://doi.org/10.1186/s12859-018-2312-1 SOFTWARE Open Access HebbPlot: an intelligent tool for learning and visualizing chromatin mark signatures Hani Z.
Analyzing ChIP-seq data. R. Gentleman, D. Sarkar, S. Tapscott, Y. Cao, Z. Yao, M. Lawrence, P. Aboyoun, M. Morgan, L. Ruzzo, J. Davison, H.
 Analyzing ChIP-seq data R. Gentleman, D. Sarkar, S. Tapscott, Y. Cao, Z. Yao, M. Lawrence, P. Aboyoun, M. Morgan, L. Ruzzo, J. Davison, H. Pages Biological Motivation Chromatin-immunopreciptation followed
Analyzing ChIP-seq data R. Gentleman, D. Sarkar, S. Tapscott, Y. Cao, Z. Yao, M. Lawrence, P. Aboyoun, M. Morgan, L. Ruzzo, J. Davison, H. Pages Biological Motivation Chromatin-immunopreciptation followed
ChampionChIP Quick, High Throughput Chromatin Immunoprecipitation Assay System
 ChampionChIP Quick, High Throughput Chromatin Immunoprecipitation Assay System Liyan Pang, Ph.D. Application Scientist 1 Topics to be Covered Introduction What is ChIP-qPCR? Challenges Facing Biological
ChampionChIP Quick, High Throughput Chromatin Immunoprecipitation Assay System Liyan Pang, Ph.D. Application Scientist 1 Topics to be Covered Introduction What is ChIP-qPCR? Challenges Facing Biological
RIPSeeker: a statistical package for identifying protein-associated transcripts from RIP-seq experiments
 RIPSeeker: a statistical package for identifying protein-associated transcripts from RIP-seq experiments Yue Li yueli@cs.toronto.edu April 4, 2013 1 Introduction Ribonucleoprotein (RNP) immunoprecipitation
RIPSeeker: a statistical package for identifying protein-associated transcripts from RIP-seq experiments Yue Li yueli@cs.toronto.edu April 4, 2013 1 Introduction Ribonucleoprotein (RNP) immunoprecipitation
ChIP-seq analysis 2/28/2018
 ChIP-seq analysis 2/28/2018 Acknowledgements Much of the content of this lecture is from: Furey (2012) ChIP-seq and beyond Park (2009) ChIP-seq advantages + challenges Landt et al. (2012) ChIP-seq guidelines
ChIP-seq analysis 2/28/2018 Acknowledgements Much of the content of this lecture is from: Furey (2012) ChIP-seq and beyond Park (2009) ChIP-seq advantages + challenges Landt et al. (2012) ChIP-seq guidelines
A Protein Secondary Structure Prediction Method Based on BP Neural Network Ru-xi YIN, Li-zhen LIU*, Wei SONG, Xin-lei ZHAO and Chao DU
 2017 2nd International Conference on Artificial Intelligence: Techniques and Applications (AITA 2017 ISBN: 978-1-60595-491-2 A Protein Secondary Structure Prediction Method Based on BP Neural Network Ru-xi
2017 2nd International Conference on Artificial Intelligence: Techniques and Applications (AITA 2017 ISBN: 978-1-60595-491-2 A Protein Secondary Structure Prediction Method Based on BP Neural Network Ru-xi
RSVSim an R/Bioconductor package for the simulation of structural variations
 RSVSim an R/Bioconductor package for the simulation of structural variations Christoph Bartenhagen October 30, 2017 Contents 1 Introduction 1 1.1 Loading the package......................................
RSVSim an R/Bioconductor package for the simulation of structural variations Christoph Bartenhagen October 30, 2017 Contents 1 Introduction 1 1.1 Loading the package......................................
What Can the Epigenome Teach Us About Cellular States and Diseases?
 What Can the Epigenome Teach Us About Cellular States and Diseases? (a computer scientist s view) Luca Pinello Outline Epigenetic: the code over the code What can we learn from epigenomic data? Resources
What Can the Epigenome Teach Us About Cellular States and Diseases? (a computer scientist s view) Luca Pinello Outline Epigenetic: the code over the code What can we learn from epigenomic data? Resources
ChIP-Seq Data Analysis: Identification of Protein DNA Binding Sites with SISSRs Peak-Finder
 Chapter 20 ChIP-Seq Data Analysis: Identification of Protein DNA Binding Sites with SISSRs Peak-Finder Leelavati Narlikar and Raja Jothi Abstract Protein DNA interactions play key roles in determining
Chapter 20 ChIP-Seq Data Analysis: Identification of Protein DNA Binding Sites with SISSRs Peak-Finder Leelavati Narlikar and Raja Jothi Abstract Protein DNA interactions play key roles in determining
Galaxy Platform For NGS Data Analyses
 Galaxy Platform For NGS Data Analyses Weihong Yan wyan@chem.ucla.edu Collaboratory Web Site http://qcb.ucla.edu/collaboratory http://collaboratory.lifesci.ucla.edu Workshop Outline ü Day 1 UCLA galaxy
Galaxy Platform For NGS Data Analyses Weihong Yan wyan@chem.ucla.edu Collaboratory Web Site http://qcb.ucla.edu/collaboratory http://collaboratory.lifesci.ucla.edu Workshop Outline ü Day 1 UCLA galaxy
annotationtools Alexandre Kuhn October 30, 2017
 annotationtools Alexandre Kuhn October 30, 2017 annotationtools is an R package for the annotation of DNA microarray experiments and for the cross-platform and cross-species integration of gene expression
annotationtools Alexandre Kuhn October 30, 2017 annotationtools is an R package for the annotation of DNA microarray experiments and for the cross-platform and cross-species integration of gene expression
ChroMoS Guide (version 1.2)
 ChroMoS Guide (version 1.2) Background Genome-wide association studies (GWAS) reveal increasing number of disease-associated SNPs. Since majority of these SNPs are located in intergenic and intronic regions
ChroMoS Guide (version 1.2) Background Genome-wide association studies (GWAS) reveal increasing number of disease-associated SNPs. Since majority of these SNPs are located in intergenic and intronic regions
An Introduction to the package geno2proteo
 An Introduction to the package geno2proteo Yaoyong Li January 24, 2018 Contents 1 Introduction 1 2 The data files needed by the package geno2proteo 2 3 The main functions of the package 3 1 Introduction
An Introduction to the package geno2proteo Yaoyong Li January 24, 2018 Contents 1 Introduction 1 2 The data files needed by the package geno2proteo 2 3 The main functions of the package 3 1 Introduction
Figure S1. nuclear extracts. HeLa cell nuclear extract. Input IgG IP:ORC2 ORC2 ORC2. MCM4 origin. ORC2 occupancy
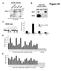 A nuclear extracts B HeLa cell nuclear extract Figure S1 ORC2 (in kda) 21 132 7 ORC2 Input IgG IP:ORC2 32 ORC C D PRKDC ORC2 occupancy Directed against ORC2 C-terminus (sc-272) MCM origin 2 2 1-1 -1kb
A nuclear extracts B HeLa cell nuclear extract Figure S1 ORC2 (in kda) 21 132 7 ORC2 Input IgG IP:ORC2 32 ORC C D PRKDC ORC2 occupancy Directed against ORC2 C-terminus (sc-272) MCM origin 2 2 1-1 -1kb
Tutorial. Bisulfite Sequencing. Sample to Insight. September 15, 2016
 Bisulfite Sequencing September 15, 2016 Sample to Insight CLC bio, a QIAGEN Company Silkeborgvej 2 Prismet 8000 Aarhus C Denmark Telephone: +45 70 22 32 44 www.clcbio.com support-clcbio@qiagen.com Bisulfite
Bisulfite Sequencing September 15, 2016 Sample to Insight CLC bio, a QIAGEN Company Silkeborgvej 2 Prismet 8000 Aarhus C Denmark Telephone: +45 70 22 32 44 www.clcbio.com support-clcbio@qiagen.com Bisulfite
Chapter 17 Lecture. Concepts of Genetics. Tenth Edition. Regulation of Gene Expression in Eukaryotes
 Chapter 17 Lecture Concepts of Genetics Tenth Edition Regulation of Gene Expression in Eukaryotes Chapter Contents 17.1 Eukaryotic Gene Regulation Can Occur at Any of the Steps Leading from DNA to Protein
Chapter 17 Lecture Concepts of Genetics Tenth Edition Regulation of Gene Expression in Eukaryotes Chapter Contents 17.1 Eukaryotic Gene Regulation Can Occur at Any of the Steps Leading from DNA to Protein
Genome 541 Gene regulation and epigenomics Lecture 3 Integrative analysis of genomics assays
 Genome 541 Gene regulation and epigenomics Lecture 3 Integrative analysis of genomics assays Please consider both the forward and reverse strands (i.e. reverse compliment sequence). You do not need to
Genome 541 Gene regulation and epigenomics Lecture 3 Integrative analysis of genomics assays Please consider both the forward and reverse strands (i.e. reverse compliment sequence). You do not need to
Package goseq. R topics documented: December 23, 2017
 Package goseq December 23, 2017 Version 1.30.0 Date 2017/09/04 Title Gene Ontology analyser for RNA-seq and other length biased data Author Matthew Young Maintainer Nadia Davidson ,
Package goseq December 23, 2017 Version 1.30.0 Date 2017/09/04 Title Gene Ontology analyser for RNA-seq and other length biased data Author Matthew Young Maintainer Nadia Davidson ,
