Intelligent and automatic in vivo detection and quantification of transplanted cells in MRI (Supporting Material)
|
|
|
- Cory Ellis
- 5 years ago
- Views:
Transcription
1 Intelligent and automatic in vivo detection and quantification of transplanted cells in MRI (Supporting Material) Muhammad Jamal Afridi 1, Arun Ross 1, Xiaoming Liu 1, Margaret F. Bennewitz 2, Dorela D. Shuboni 3, Erik M. Shapiro 3 1. Department of Computer Science and Engineering, Michigan State University, USA 2. Vascular Medicine Institute, University of Pittsburgh, USA 3. Department of Radiology, Michigan State University, USA Correspondence to: Erik M. Shapiro Department of Radiology, Michigan State University Radiology Building, Michigan State University 846 Service Road, East Lansing, MI Erik.Shapiro@radiology.msu.edu Submitted to Magnetic Resonance in Medicine
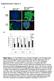
2 Supporting Material - Figure S1 Figure 1: a,b) Single cells within plane c) cell through plane d,e) two cells f) 3 connected cells g, h, i) showing cells bounded within blood vessels, with some at bifurcations; scale bars are 25 ums. Cropped from images with width of 300 um (no MPIO labeled MSCs were cropped from view).
3 Supporting Material - Figure S2 Figure 2: Similar to the concept of eigen faces, Principle Component Analysis (PCA) was utilized to extract eigen spot shapes using all of the 9 9 spot patches in the training set. The top PCA component for the spot patches obtained on three labeled rats in G A are shown here. An iteratively increasing threshold is then applied on the values of these top PCA components to extract different binary patches that are utilized as filters to capture the shape and intensity information on spot patches. Supporting Material - Figure S3 Figure 3: Binary shape filters are obtained using the top PCA components (shown in Fig. S2.). By iteratively increasing the threshold values from dark to light intensities, PCA components can result in different binary shapes. Domain experts agree that these binary filter patches represent many frequent shapes of the actual spots. All these patches are rotated and translated to obtain a large set of different shape filters. These filters are convolved with each candidate patch and the computed response is taken as a feature. A large set of these responses comprehensively captures the shape and intensity information of spot patches.
4 Supporting Material - Figure S4 Figure 4: Visual representation of context features for two context patches: While learning the definition of a spot, it may also be useful for a classifier to learn more about its surrounding context. Therefore, to capture the appearance of the patch s context, a larger patch of was extracted around the center of a candidate patch x i. Two well-known appearance descriptors in computer vision were then used to extract features: (a) Histogram of Gradients (HoG) and (b) Gist. A visual representation of HoG for two context patches is shown here. Red lines here indicate the different directions of intensity gradients whereas the lengths of the lines determine their magnitude. Supporting Material - Figure S5 Figure 5: CNN architecture used in this work. The network takes a 9x9 image patch as input for a classification task i. Each composite layer L k, k {1, 2, 3}, is composed of a convolutional layer C ik which produces the feature maps F ik and a non-linear gating function β producing the transformed feature maps Fβ ik. After passing through the composite layers, the net passes through the fully connected layer L fc which produces the output. The softmax function is then applied to the output. Note that in the context of this work, this architecture represent the model M. The weights of all the filters across its processing layers are learned using the training data.
5 Supporting Material - Figure S6 Figure 6: Some convolution filters learned by the 2nd composite layers of our deep learning approach. Usually, each filter acts as a neuron and due to co-adaptation between a large number of such neurons, highly sophisticated features are extracted that can potentially model spot shape, intensity, texture etc. Supporting Material - Figure S7 Figure 7: Transformed, average images for different source datasets.
6 Supporting Material - Methods Cell preparation A model of single, dispersed cells in the rat brain was created by intracardiac injection of magnetically labeled MSCs. Animal experiments were approved by the Yale University and Michigan State University Animal Care and Use Committees. MSCs were maintained in low glucose DMEM without L-glutamine, supplemented with 10% fetal bovine serum (FBS), 1% L-glutamine, and 1% penicillin/streptomycin. The cells were grown until 80% confluent in T25 culture flasks. Prior to injection, MSCs were labeled with fluorescent 1.63 micron MPIO (Bangs Laboratory, Flash Red) by overnight incubation at a concentration of 40 beads per cell ( 44 pg Fe/cell), followed by washing in PBS to remove uninternalized particles. This labeling scheme has previously been validated to label cells to a level of pg Fe/cell (37). Control cells were labeled with media only. Unlabeled and MPIO labeled MSCs were also stained with carboxyfluorescein succinimidyl ester (CFSE), a fluorescent dye. In brief, cells were incubated for 5 min in a concentration of 0.05mM CFSE and then washed twice with PBS containing 5% HI FBS after centrifugation at 300g for 5 min. Prior to injection, the cells were then diluted in 0.9% saline into two concentrations (n=3/group), which would deliver 200,000 cells (Set G A ) or 250,000 cells (Set G B ) per 200uL injection. Animal preparation Adult Fischer rats were anesthetized with 3% isoflurane in 90% oxygen/10% medical air and orally intubated. A mechanical ventilating unit provided regulated breathing at 65 breaths/minute. Respiratory patterns and end tidal CO 2 were monitored. After being placed in the supine position, rats were shaved along the left side of the chest, from the sternum downwards and prepped using aseptic technique. No skin incisions were made; rather, cells were injected noninvasively. 200,000 or 250,000 MPIO labeled MSCs or unlabeled MSCs (200 µl) were injected slowly into the left ventricle of the rodent heart with a 1 ml syringe attached to a 30-gauge needle. The needle was held at a 45-degree angle, placed ventrally, just left of the sternum, and lowered into the intercostal space between the 6 th and 7 th ribs. Noticeable blood flow into the syringe was used to gauge placement of the needle into the left ventricle. Animals remained under anesthesia, with ventilated breathing for hours before being imaged on MRI In vivo MRI MRI of live rats was performed on a 7.0T Bruker Biospin or 11.7T Varian system, hr after intracardiac injection of unlabeled or MPIO labeled MSCs. Separate transmit-only volume and receive-only surface coils were used. During MRI, rats remained mechanically ventilated at 65 breaths/min, under 3% isoflurane anesthesia in 90% oxygen/10% medical air. End tidal CO 2 and respiratory patterns were monitored. T 2 weighted 3D gradient echo MRI was performed at 100 micron resolution using the following imaging parameters: TR = 30 ms and TEs = 10 (7.0T) or 8 (11.7T) ms.
7 Histology Immediately following MRI, rats were transcardially perfused with warm 0.9% saline, followed by ice cold 4% paraformaldehyde solutions of ph 6.5 and ph 9.5. Brains were excised and post-fixed in 4% paraformaldehyde with 10% sucrose at 4 C. After sinking, brains were rinsed in PBS, patted dry, placed in a plastic cryomold, and immersed in TissueTek embedding compound. After a few minutes of equilibration, brains were quickly frozen on dry ice and stored at -80 C. A cryostat was used to cut 16µm sections of the brains. Slides of interest were hydrated in PBS for 10 minutes, and mounted with ProLong Gold antifade with DAPI. Fluorescent microscopic images were acquired with the GFP filter for MSCs, Texas Red filter for MPIOs, and the UV filter for DAPI. Images of cells were taken with an approximate width of 300µm across. To determine the percentage of images containing 1, 2, or 3 cells, the number of MPIO labeled cells in each image was counted for a total of 70 images. Each image was cropped to a FOV of 100 µm across for a closer view of the cells to display in Figure S1; no MPIO labeled MSCs were removed from view by cropping.
8 Supporting Material - Source CNN Selection This section explains how the terms J i and U i, utilized in the CNN selection criterion, are computed. As mentioned in the paper, the proposed CNN architecture can be seen as a sequence of functions and therefor for a specific task i, it can written as: M i = (f ui f (u 1)i f (u 2)i,..., f 1i ). [1] where, f (u 1)i denotes the fully connected layer whose output is subjected to a softmax function in f ui. f (u 2)i represents a non-linear activation function whereas f (u 3)i denotes the deepest convolutional layer. The CNN learned on small MRI data M already captures the information available in the target s training data X. Therefore the goal of transferring weights (also called learned feature representations) from a source CNN is to capture that information which is not already accounted for by M and may be relevant in distinguishing a spot patch from non-spot patches. Intuitively, a source ranking criterion can be based on : 1. how different is the learned information (CNN weights) of a source CNN M i from the target CNN M? If the source CNN M i provides the exact same information as M then a transfer from such a source may not result in any performance improvement. Hence difference between the learned weights of the two CNNs is a desirable property. Previous research shows that the feature representations (weights) learned in the initial convolutional layers are more generic and may be the same irrespective of the learning task. However, the deeper convolutional layer are more task specific. Therefore, the difference between two CNNs, i.e., M and M i is denoted by J i which is given by J i = W (u 3)x W (u 3)i 2 2. [2] where W (u 3)i denotes a vector containing all the weights in the deepest convolutional layer f (u 3)i of the CNN M i. Similarly, W (u 3)x denotes the deepest convolutional layer weights of the CNN M. 2. how discriminating a source CNN M i is for the target task? This determines how relevant a source CNN is to the target task despite its difference with M. This can be measured by computing U i for each source CNN M i. U i = Y M i (X) 2 2. [3] where Y denotes a vector of labels on training data X and M i (X) represents the vector of predicted outputs resulting when the source CNN M i operates on the target s limited training data X as a test set.
Intelligent and Automatic In Vivo Detection and Quantification of Transplanted Cells in MRI
 FULL PAPER Magnetic Resonance in Medicine 00:000 000 (2016) Intelligent and Automatic In Vivo Detection and Quantification of Transplanted Cells in MRI Muhammad Jamal Afridi, 1 Arun Ross, 1 Xiaoming Liu,
FULL PAPER Magnetic Resonance in Medicine 00:000 000 (2016) Intelligent and Automatic In Vivo Detection and Quantification of Transplanted Cells in MRI Muhammad Jamal Afridi, 1 Arun Ross, 1 Xiaoming Liu,
Section of Morphological and Behavioral Neuroscience Protocol for low density primary neuron cultures
 Institute of Pharmacology and Toxicology Section of Morphological and Behavioral Neuroscience Protocol for low density primary neuron cultures B. Lardi-Studler, C. Sidler, J.-M. Fritschy Revised March
Institute of Pharmacology and Toxicology Section of Morphological and Behavioral Neuroscience Protocol for low density primary neuron cultures B. Lardi-Studler, C. Sidler, J.-M. Fritschy Revised March
Murine in vivo CD8 + T Cell Killing Assay Myoungjoo V. Kim 1*, Weiming Ouyang 2, Will Liao 3, Michael Q. Zhang 4 and Ming O. Li 5
 Murine in vivo CD8 + T Cell Killing Assay Myoungjoo V. Kim 1*, Weiming Ouyang 2, Will Liao 3, Michael Q. Zhang 4 and Ming O. Li 5 1 Department of Immunobiology, Yale University School of Medicine, New
Murine in vivo CD8 + T Cell Killing Assay Myoungjoo V. Kim 1*, Weiming Ouyang 2, Will Liao 3, Michael Q. Zhang 4 and Ming O. Li 5 1 Department of Immunobiology, Yale University School of Medicine, New
Example Protocol for the Culture of the SW620 Cell Line on Alvetex Scaffold in Well Insert and Well Plate Formats
 Introduction: Alvetex Scaffold is currently available in four different cell culture formats: 24-well plate (AVP006), 12-well plate (AVP002), 6-well insert (AVP004), and 12-well insert (AVP005). 24-well
Introduction: Alvetex Scaffold is currently available in four different cell culture formats: 24-well plate (AVP006), 12-well plate (AVP002), 6-well insert (AVP004), and 12-well insert (AVP005). 24-well
Supporting Information
 Electronic Supplementary Material (ESI) for Lab on a Chip. This journal is The Royal Society of Chemistry 2014 Supporting Information Cell culture. C2C12 cells (ATCC CRL 1772) were cultured in 75 cm 2
Electronic Supplementary Material (ESI) for Lab on a Chip. This journal is The Royal Society of Chemistry 2014 Supporting Information Cell culture. C2C12 cells (ATCC CRL 1772) were cultured in 75 cm 2
3D Endothelial Pericyte Coculturing Kit Product Description Kit Components
 3D Endothelial Pericyte Coculturing Kit (3D-EPC) Cat #8728 Product Description Blood vessels are responsible for transporting blood throughout the body as part of the circulatory system. Their main components
3D Endothelial Pericyte Coculturing Kit (3D-EPC) Cat #8728 Product Description Blood vessels are responsible for transporting blood throughout the body as part of the circulatory system. Their main components
CFSE Cell Division Assay Kit
 CFSE Cell Division Assay Kit Item No. 10009853 www.caymanchem.com Customer Service 800.364.9897 Technical Support 888.526.5351 1180 E. Ellsworth Rd Ann Arbor, MI USA TABLE OF CONTENTS GENERAL INFORMATION
CFSE Cell Division Assay Kit Item No. 10009853 www.caymanchem.com Customer Service 800.364.9897 Technical Support 888.526.5351 1180 E. Ellsworth Rd Ann Arbor, MI USA TABLE OF CONTENTS GENERAL INFORMATION
ProLong Live Antifade Reagent
 ProLong Live Antifade Reagent Catalog nos. P36974, P36975 Table. Contents and storage Product* Amount Catalog no. Storage ProLong Live Antifade Reagent 5 ml ml P36974 P36975 2 C or 2 8 C Protect from light
ProLong Live Antifade Reagent Catalog nos. P36974, P36975 Table. Contents and storage Product* Amount Catalog no. Storage ProLong Live Antifade Reagent 5 ml ml P36974 P36975 2 C or 2 8 C Protect from light
PROTOCOL. Wheat Germ Agglutinin Staining of 3D Cultures Grown on Alvetex Scaffold and Alvetex Strata Introduction
 Page 1 1.0. Introduction Wheat Germ Agglutinin (WGA) is a well-known lectin originally noted for its differential ability to agglutinate cancerous versus normal cells [1] and later identified as a plasma
Page 1 1.0. Introduction Wheat Germ Agglutinin (WGA) is a well-known lectin originally noted for its differential ability to agglutinate cancerous versus normal cells [1] and later identified as a plasma
Frozen tissue section
 IHC Protocol - Frozen Tissue Author : Dan Souw Immunohistochemistry on Frozen tissues IHC Protocol - Frozen Tissue: An introduction This is the second post in a series on immunohistochemistry (IHC). The
IHC Protocol - Frozen Tissue Author : Dan Souw Immunohistochemistry on Frozen tissues IHC Protocol - Frozen Tissue: An introduction This is the second post in a series on immunohistochemistry (IHC). The
PCCS Growth Media, Cell Tagging, Cell Separation Final Assignment. Igneris Rosado-Erazo. Panama College of Cell Science
 Running Head: Growth Media, Cell Tagging, Cell Separation PCCS Growth Media, Cell Tagging, Cell Separation Final Assignment Igneris Rosado-Erazo Panama College of Cell Science In partial fulfillment of
Running Head: Growth Media, Cell Tagging, Cell Separation PCCS Growth Media, Cell Tagging, Cell Separation Final Assignment Igneris Rosado-Erazo Panama College of Cell Science In partial fulfillment of
Leaf-inspired Artificial Microvascular Networks (LIAMN) for Threedimensional
 Electronic Supplementary Material (ESI) for RSC Advances. This journal is The Royal Society of Chemistry 2015 Supporting Information Leaf-inspired Artificial Microvascular Networks (LIAMN) for Threedimensional
Electronic Supplementary Material (ESI) for RSC Advances. This journal is The Royal Society of Chemistry 2015 Supporting Information Leaf-inspired Artificial Microvascular Networks (LIAMN) for Threedimensional
Axol Guide to Performing Immunocytochemistry (ICC) Application Protocol Version 2.0
 Axol Guide to Performing Immunocytochemistry (ICC) Application Protocol Version 2.0 Table of Contents General ICC Protocol 3 Synaptic Marker ICC Protocol 5 Recommended Markers 6 Technical Support 7 2 General
Axol Guide to Performing Immunocytochemistry (ICC) Application Protocol Version 2.0 Table of Contents General ICC Protocol 3 Synaptic Marker ICC Protocol 5 Recommended Markers 6 Technical Support 7 2 General
Adult Neuron Isolation Protocol (R. Kaletsky 9/2014)
 Notes before starting: Adult Neuron Isolation Protocol (R. Kaletsky 9/2014) Begin protocol ~45min before sort time to minimize time between cell harvesting and sorting Prepare pronase solution immediately
Notes before starting: Adult Neuron Isolation Protocol (R. Kaletsky 9/2014) Begin protocol ~45min before sort time to minimize time between cell harvesting and sorting Prepare pronase solution immediately
Immunofluorescent staining and flow cytometric analysis of cells
 UCAN-U 0003 Version 1 November 2010 Page 1 of 7 CMCI (Center for molecular and Cellular Intervention) University Medical Center Utrecht Written By Name Function Date Signature Mark Klein Lab manager Conformation
UCAN-U 0003 Version 1 November 2010 Page 1 of 7 CMCI (Center for molecular and Cellular Intervention) University Medical Center Utrecht Written By Name Function Date Signature Mark Klein Lab manager Conformation
Supplementary Figure 1 A green: cytokeratin 8
 Supplementary Figure 1 A green: cytokeratin 8 green: α-sma red: α-sma blue: DAPI blue: DAPI Panc-1 Panc-1 Panc-1+hPSC Panc-1+hPSC monoculture coculture B Suppl. Figure 1: A, Immunofluorescence staining
Supplementary Figure 1 A green: cytokeratin 8 green: α-sma red: α-sma blue: DAPI blue: DAPI Panc-1 Panc-1 Panc-1+hPSC Panc-1+hPSC monoculture coculture B Suppl. Figure 1: A, Immunofluorescence staining
Materials and Methods Materials Required for Fixing, Embedding and Sectioning. OCT embedding matrix (Thermo Scientific, LAMB/OCT)
 Page 1 Introduction Tissue freezing and sectioning is a rapid method of generating tissue samples (cryosections) for histological analysis, and obviates the need for wax embedding. The method is popular
Page 1 Introduction Tissue freezing and sectioning is a rapid method of generating tissue samples (cryosections) for histological analysis, and obviates the need for wax embedding. The method is popular
Metzger Lab Protocol Book EF May 2003
 I. Preparation for cell isolation: A. Make sure the following items are autoclaved: 1. Glassware* (1) 100 ml beaker (3) wide mouthed 1 liter bottles (1) 100 mm glass petri dish (1) 60 mm sigma coated petri
I. Preparation for cell isolation: A. Make sure the following items are autoclaved: 1. Glassware* (1) 100 ml beaker (3) wide mouthed 1 liter bottles (1) 100 mm glass petri dish (1) 60 mm sigma coated petri
LHCN-M2 cell culture, differentiation treatment, and cross-linking protocol.
 Cell Growth Protocol and Differentiation treatment for the LHCN-M2 Cell Line From: HudsonAlpha/Caltech ENCODE group Date: February 17, 2011 Prepared by: Chun-Hong Zhu and Woodring E. Wright (typed by Brian
Cell Growth Protocol and Differentiation treatment for the LHCN-M2 Cell Line From: HudsonAlpha/Caltech ENCODE group Date: February 17, 2011 Prepared by: Chun-Hong Zhu and Woodring E. Wright (typed by Brian
Immunofluorescence of organoids embedded in Basement Membrane Matrix
 Immunofluorescence of organoids embedded in Basement Membrane Matrix Sol Degese 1, Gabe Benton 1 1 Organoid Resource Lab (ORL), Trevigen, Inc., 8405 Helgerman Court, Gaithersburg, MD 20877 Introduction
Immunofluorescence of organoids embedded in Basement Membrane Matrix Sol Degese 1, Gabe Benton 1 1 Organoid Resource Lab (ORL), Trevigen, Inc., 8405 Helgerman Court, Gaithersburg, MD 20877 Introduction
L6 Cell Growth Protocol
 L6 Cell Growth Protocol Background: Parental L6 cells were subcloned for high fusion (1, 2). Materials: 1. α-minimal Essential Medium (α-mem) Life Technologies #12571-063 2. Fetal Bovine Serum (FBS) Life
L6 Cell Growth Protocol Background: Parental L6 cells were subcloned for high fusion (1, 2). Materials: 1. α-minimal Essential Medium (α-mem) Life Technologies #12571-063 2. Fetal Bovine Serum (FBS) Life
HepG2 Labeling and Cell Growth Protocol for IROA Phenotypic Metabolic Profiling
 HepG2 Labeling and Cell Growth Protocol for IROA Phenotypic Metabolic Profiling Candice Z. Ulmer 1, Felice A. de Jong 2, Chris Beecher 1, Timothy J. Garrett 3, Richard A. Yost 1,3 1 Department of Chemistry,
HepG2 Labeling and Cell Growth Protocol for IROA Phenotypic Metabolic Profiling Candice Z. Ulmer 1, Felice A. de Jong 2, Chris Beecher 1, Timothy J. Garrett 3, Richard A. Yost 1,3 1 Department of Chemistry,
Monoclonal antibodies
 Monoclonal antibodies 1. Boost the best-responded mouse 3-5 days before fusion. 2. Build up frozen stock of myeloma cells (NS-1 or P 3 U 1 etc.) in MEM + 10% FCS containing 0.1
Monoclonal antibodies 1. Boost the best-responded mouse 3-5 days before fusion. 2. Build up frozen stock of myeloma cells (NS-1 or P 3 U 1 etc.) in MEM + 10% FCS containing 0.1
VDL602.2 RAPID ASSAY FOR DETERMINING ADENOVIRAL VECTOR TITER
 1. Purpose 1.1. The purpose of this protocol is to determine the number of infectious adenoviral particles. 1.2. The starting material is purified adenoviral vectors. 1.3. This protocol is based upon the
1. Purpose 1.1. The purpose of this protocol is to determine the number of infectious adenoviral particles. 1.2. The starting material is purified adenoviral vectors. 1.3. This protocol is based upon the
Neural Stem Cell Characterization Kit
 Neural Stem Cell Characterization Kit Catalog No. SCR019 FOR RESEARCH USE ONLY Not for use in diagnostic procedures USA & Canada Phone: +1(800) 437-7500 Fax: +1 (951) 676-9209 Europe +44 (0) 23 8026 2233
Neural Stem Cell Characterization Kit Catalog No. SCR019 FOR RESEARCH USE ONLY Not for use in diagnostic procedures USA & Canada Phone: +1(800) 437-7500 Fax: +1 (951) 676-9209 Europe +44 (0) 23 8026 2233
Kit Components (Included) Cat # # of vials Product Name Quantity Storage Human Renal Cortical Epithelial Cells (HRCEpiC)
 3D Renal Tubule Formation Kit 3D-RTF Cat # 3D-4110 Product Description The human kidney is frequently exposed to drugs and toxic compounds, which can result in nephrotoxicity. A drug s uptake and clearance
3D Renal Tubule Formation Kit 3D-RTF Cat # 3D-4110 Product Description The human kidney is frequently exposed to drugs and toxic compounds, which can result in nephrotoxicity. A drug s uptake and clearance
Immunofluorescence Confocal Microscopy of 3D Cultures Grown on Alvetex
 Immunofluorescence Confocal Microscopy of 3D Cultures Grown on Alvetex 1.0. Introduction Immunofluorescence uses the recognition of cellular targets by fluorescent dyes or antigen-specific antibodies coupled
Immunofluorescence Confocal Microscopy of 3D Cultures Grown on Alvetex 1.0. Introduction Immunofluorescence uses the recognition of cellular targets by fluorescent dyes or antigen-specific antibodies coupled
Adenovirus Titration Kit
 Adenovirus Titration Kit Catalog # LF-RK0001(1 kit) Immunostaining method for Quantitative Detection of Adenovirus For research use only Not for diagnostic or therapeutic procedures AbFrontier Science
Adenovirus Titration Kit Catalog # LF-RK0001(1 kit) Immunostaining method for Quantitative Detection of Adenovirus For research use only Not for diagnostic or therapeutic procedures AbFrontier Science
Single Molecule FISH on Mouse Tissue Sections
 Single Molecule FISH on Mouse Tissue Sections Shalev Itzkovitz, December 2012 Tissue preparation Solutions and s 10X PBS (Ambion #AM9625) 4% formaldehyde or paraformaldehyde in PBS Cryoprotecting solution:
Single Molecule FISH on Mouse Tissue Sections Shalev Itzkovitz, December 2012 Tissue preparation Solutions and s 10X PBS (Ambion #AM9625) 4% formaldehyde or paraformaldehyde in PBS Cryoprotecting solution:
ab BrdU Immunohistochemistry Kit
 ab125306 - BrdU Immunohistochemistry Kit Instructions for Use For the detection and localization of bromodeoxyuridine incorporated into newly synthesized DNA of actively proliferating cells. This product
ab125306 - BrdU Immunohistochemistry Kit Instructions for Use For the detection and localization of bromodeoxyuridine incorporated into newly synthesized DNA of actively proliferating cells. This product
Immunofluorescence Staining Protocol for 3 Well Chamber, removable
 Immunofluorescence Staining Protocol for 3 Well Chamber, removable This Application Note presents a simple protocol for the cultivation, fixation, and staining of cells using the 3 Well Chamber, removable.
Immunofluorescence Staining Protocol for 3 Well Chamber, removable This Application Note presents a simple protocol for the cultivation, fixation, and staining of cells using the 3 Well Chamber, removable.
Proliferation of Human Mononuclear Cells with CFSE
 UCANU 0005 Version 01, November 2010 Page 1 of 7 CMCI (Center for molecular and Cellular Intervention) University Medical Center Utrecht Written By Name Function Date Signature Mark Klein Lab manager Conformation
UCANU 0005 Version 01, November 2010 Page 1 of 7 CMCI (Center for molecular and Cellular Intervention) University Medical Center Utrecht Written By Name Function Date Signature Mark Klein Lab manager Conformation
CytoSelect 24-Well Wound Healing Assay
 Product Manual CytoSelect 24-Well Wound Healing Assay Catalog Number CBA-120 CBA-120-5 24 assays 5 x 24 assays FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction Wounded tissue initiates
Product Manual CytoSelect 24-Well Wound Healing Assay Catalog Number CBA-120 CBA-120-5 24 assays 5 x 24 assays FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction Wounded tissue initiates
User Manual. OriCell TM Mesenchymal Stem Cell Osteogenic Differentiation Medium. Cat. No. GUXMX-90021
 User Manual OriCell TM Mesenchymal Stem Cell Osteogenic Differentiation Cat. No. GUXMX-90021 PRODUCT DESCRIPTION: OriCell TM Mesenchymal Stem Cell Osteogenic Differentiation consists of optimized Mesenchymal
User Manual OriCell TM Mesenchymal Stem Cell Osteogenic Differentiation Cat. No. GUXMX-90021 PRODUCT DESCRIPTION: OriCell TM Mesenchymal Stem Cell Osteogenic Differentiation consists of optimized Mesenchymal
PROTOCOL TO PREPARE PLANTAR FOOTSKIN FOR MORPHOMETRY. I. Removal and Fixation of Plantar Skin (see video)
 PROTOCOL TO PREPARE PLANTAR FOOTSKIN FOR MORPHOMETRY I. Removal and Fixation of Plantar Skin (see video) 1. Sacrifice the animal a. Anaesthetize the animal by placing in a closed chamber with isoflurane.
PROTOCOL TO PREPARE PLANTAR FOOTSKIN FOR MORPHOMETRY I. Removal and Fixation of Plantar Skin (see video) 1. Sacrifice the animal a. Anaesthetize the animal by placing in a closed chamber with isoflurane.
Supporting Information
 Supporting Information Deng et al. 10.1073/pnas.1515692112 SI Materials and Methods FPLC. All fusion proteins were expressed and purified through a three-step FPLC purification protocol, as described (20),
Supporting Information Deng et al. 10.1073/pnas.1515692112 SI Materials and Methods FPLC. All fusion proteins were expressed and purified through a three-step FPLC purification protocol, as described (20),
ab CFSE Fluorescent Cell Labeling Kit
 ab113853 CFSE Fluorescent Cell Labeling Kit Instructions for Use For the durable fluorescent labeling of live cells for fluorescent microscopy and flow cytometry, population growth studies and within sample
ab113853 CFSE Fluorescent Cell Labeling Kit Instructions for Use For the durable fluorescent labeling of live cells for fluorescent microscopy and flow cytometry, population growth studies and within sample
VDL300.3 PRODUCTION OF RETROVIRAL VECTOR BY TRANSIENT TRANSFECTION
 1. Purpose 1.1. The purpose of this protocol is to produce retrovirus from transfected plasmid DNA. 1.2. This procedure is routinely performed in the Vector Development Laboratory (VDL) following Good
1. Purpose 1.1. The purpose of this protocol is to produce retrovirus from transfected plasmid DNA. 1.2. This procedure is routinely performed in the Vector Development Laboratory (VDL) following Good
Supporting Information
 Supporting Information Alenina et al. 10.1073/pnas.0810793106 SI Materials and Methods In Vivo Brain Magnetic Resonance Imaging (MRI). In vivo brain MRI was performed in 6-month-old Tph2 / and control
Supporting Information Alenina et al. 10.1073/pnas.0810793106 SI Materials and Methods In Vivo Brain Magnetic Resonance Imaging (MRI). In vivo brain MRI was performed in 6-month-old Tph2 / and control
SOP 3 Processing of Ascites and Pleural Fluid for Cell Culture and Xenografts
 Texas Cancer Cell Repository Cell Culture and Xenograft Repository http://cancer.ttuhsc.edu www.txccr.org www.cogcell.org SOP 3 Processing of Ascites and Pleural Fluid for Cell Culture and Xenografts Receiving
Texas Cancer Cell Repository Cell Culture and Xenograft Repository http://cancer.ttuhsc.edu www.txccr.org www.cogcell.org SOP 3 Processing of Ascites and Pleural Fluid for Cell Culture and Xenografts Receiving
Nature Protocols: doi: /nprot Supplementary Figure 1. Shear stress determination in microvascular networks.
 Supplementary Figure 1 Shear stress determination in microvascular networks. A pressure drop of ~4 mmh2o is applied across the vascular network using a suspension of 2 micron red polystyrene spheres in
Supplementary Figure 1 Shear stress determination in microvascular networks. A pressure drop of ~4 mmh2o is applied across the vascular network using a suspension of 2 micron red polystyrene spheres in
TUNEL Apoptosis Detection Kit (Green Fluorescence)
 TUNEL Apoptosis Detection Kit (Green Fluorescence) Item NO. KTA2010 Product Name TUNEL Apoptosis Detection Kit (Green Fluorescence) ATTENTION For laboratory research use only. Not for clinical or diagnostic
TUNEL Apoptosis Detection Kit (Green Fluorescence) Item NO. KTA2010 Product Name TUNEL Apoptosis Detection Kit (Green Fluorescence) ATTENTION For laboratory research use only. Not for clinical or diagnostic
Nonspecific binding of 10 nm Cy5-labeled DinB on nine different surfaces, measured by the number of DinB spots over an imaging area of 2,500 µm 2.
 Supplementary Figure 1 Nonspecific binding of 10 nm Cy5-labeled DinB on nine different surfaces, measured by the number of DinB spots over an imaging area of 2,500 µm 2. Free DinB was washed out using
Supplementary Figure 1 Nonspecific binding of 10 nm Cy5-labeled DinB on nine different surfaces, measured by the number of DinB spots over an imaging area of 2,500 µm 2. Free DinB was washed out using
BrdU Immunohistochemistry Kit Instruction Manual
 BrdU Immunohistochemistry Kit Instruction Manual Features Easy to use system Reagents titered for success Proven protocol Ordering Information Catalog Number X1545K Size 50 Slides Format Immunohistochemistry
BrdU Immunohistochemistry Kit Instruction Manual Features Easy to use system Reagents titered for success Proven protocol Ordering Information Catalog Number X1545K Size 50 Slides Format Immunohistochemistry
Standard Operating Procedure
 Standard Operating Procedure Title Subtitle NANoREG Work package/task: Owner and co-owner(s) HTS Comet Assay with and without FPG - 20 wells Comet assay with and without FPG using Trevigen 20-well slides
Standard Operating Procedure Title Subtitle NANoREG Work package/task: Owner and co-owner(s) HTS Comet Assay with and without FPG - 20 wells Comet assay with and without FPG using Trevigen 20-well slides
ab DCFDA Cellular ROS Detection Assay Kit
 Version 12 Last updated 2 February 2018 ab113851 DCFDA Cellular ROS Detection Assay Kit For measurement of reactive oxygen species (ROS) in cells This product is for research use only and is not intended
Version 12 Last updated 2 February 2018 ab113851 DCFDA Cellular ROS Detection Assay Kit For measurement of reactive oxygen species (ROS) in cells This product is for research use only and is not intended
CFSE Cell Division Assay Kit
 CFSE Cell Division Assay Kit Catalog Number KA1302 100 assays Version: 02 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Intended Use... 3 Background... 3 General Information...
CFSE Cell Division Assay Kit Catalog Number KA1302 100 assays Version: 02 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Intended Use... 3 Background... 3 General Information...
Supplementary Information. Trojan Horse Nanotheranostics with Dual Transformability and Multifunctionality. for Highly Effective Cancer Treatment
 Supplementary Information Trojan Horse Nanotheranostics with Dual Transformability and Multifunctionality for Highly Effective Cancer Treatment Xue et. al. Supplementary Figure 1. Synthetic route of PhD
Supplementary Information Trojan Horse Nanotheranostics with Dual Transformability and Multifunctionality for Highly Effective Cancer Treatment Xue et. al. Supplementary Figure 1. Synthetic route of PhD
Buffer (20X) Detergent (10X) NADH Reagent 1 (20X) Reagent 2 (100X) 96-well microplate (12 strips) 1
 PROTOCOL Complex I Enzyme Activity Microplate Assay Kit 1850 Millrace Drive, Suite 3A Eugene, Oregon 97403 MS141 Rev.1 DESCRIPTION Complex I Enzyme Activity Microplate Assay Kit Included in this kit is
PROTOCOL Complex I Enzyme Activity Microplate Assay Kit 1850 Millrace Drive, Suite 3A Eugene, Oregon 97403 MS141 Rev.1 DESCRIPTION Complex I Enzyme Activity Microplate Assay Kit Included in this kit is
ebioscience BrdU Kit for IHC/ICC Colorimetric Catalog Number: RUO: For Research Use Only. Not for use in diagnostic procedures.
 Page 1 of 1 ebioscience BrdU Kit for IHC/ICC Colorimetric Catalog Number: 8800-6599 RUO: For Research Use Only. Not for use in diagnostic procedures. Product Information Contents: ebioscience BrdU Kit
Page 1 of 1 ebioscience BrdU Kit for IHC/ICC Colorimetric Catalog Number: 8800-6599 RUO: For Research Use Only. Not for use in diagnostic procedures. Product Information Contents: ebioscience BrdU Kit
Propagation of H7 hesc From: UW (John Stamatoyannopoulos) ENCODE group Date: 12/17/2009 Prepared By: S. Paige/S. Hansen (UW)
 Propagation of H7 hesc From: UW (John Stamatoyannopoulos) ENCODE group Date: 12/17/2009 Prepared By: S. Paige/S. Hansen (UW) Growth and Harvest Modifications Addendum to: Propagation of H7 hesc from UW
Propagation of H7 hesc From: UW (John Stamatoyannopoulos) ENCODE group Date: 12/17/2009 Prepared By: S. Paige/S. Hansen (UW) Growth and Harvest Modifications Addendum to: Propagation of H7 hesc from UW
Supporting Information
 Supporting Information hoi et al. 1.173/pnas.1187918 SI Materials and Methods Histology. Mice were killed under deep anesthesia by transcardial perfusion with PS, followed by fixation with 4% formalin
Supporting Information hoi et al. 1.173/pnas.1187918 SI Materials and Methods Histology. Mice were killed under deep anesthesia by transcardial perfusion with PS, followed by fixation with 4% formalin
Paramagnetic nanoemulsions with unified signal for sensitive 19 F MRI. cell tracking
 Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2018 Paramagnetic nanoemulsions with unified signal for sensitive 19 F MRI cell tracking Qiaoli Peng,
Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2018 Paramagnetic nanoemulsions with unified signal for sensitive 19 F MRI cell tracking Qiaoli Peng,
RPCI 001 v.003 In vitro Intracellular Cytokine Staining With and Without Stimulation
 Immune Tolerance Network RPCI 001 v.003 Author: Paul Wallace and Earl Timm, Director, RPCI Laboratory of Flow Cytometry Approved by: Paul Wallace, Director, RPCI Laboratory of Flow Cytometry 1.0 Title
Immune Tolerance Network RPCI 001 v.003 Author: Paul Wallace and Earl Timm, Director, RPCI Laboratory of Flow Cytometry Approved by: Paul Wallace, Director, RPCI Laboratory of Flow Cytometry 1.0 Title
EUCOMM Protocol for ES cells
 EUCOMM Protocol for ES cells 1. SNL Feeder Cells 1.1. Preparing inactivated SNL Feeder Cells 1) Thaw one vial of SNL cells (approximately1.5-2 x 10 6 cells per vial) in a 37 C water bath and dilute into
EUCOMM Protocol for ES cells 1. SNL Feeder Cells 1.1. Preparing inactivated SNL Feeder Cells 1) Thaw one vial of SNL cells (approximately1.5-2 x 10 6 cells per vial) in a 37 C water bath and dilute into
3T3-L1 Differentiation Protocol
 3T3-L1 Differentiation Protocol Written by Eisuke Kato on 2013/10/09 MATERIALS Dulbecco's Modified Eagles Medium (D-MEM) (High Glucose) with L-Glutamine and Phenol Red High glucose (Wako Chem 044-29765,
3T3-L1 Differentiation Protocol Written by Eisuke Kato on 2013/10/09 MATERIALS Dulbecco's Modified Eagles Medium (D-MEM) (High Glucose) with L-Glutamine and Phenol Red High glucose (Wako Chem 044-29765,
Materials and Methods Materials Required for Fixing, Embedding and Sectioning
 Page 1 Introduction Immunofluorescence uses the recognition of cellular targets by fluorescent dyes or antigenspecific antibodies coupled to fluorophores. Depending on the antibody or dye used, proteins,
Page 1 Introduction Immunofluorescence uses the recognition of cellular targets by fluorescent dyes or antigenspecific antibodies coupled to fluorophores. Depending on the antibody or dye used, proteins,
Cytotrophoblast Isolation from Term Placenta References:
 Cytotrophoblast Isolation from Term Placenta References: CY Kuo, et al. Placental Basement Membrane Proteins are Required for Effective Cytotrophoblast Invasion in a 3D Bioprinted Placenta Model. Journal
Cytotrophoblast Isolation from Term Placenta References: CY Kuo, et al. Placental Basement Membrane Proteins are Required for Effective Cytotrophoblast Invasion in a 3D Bioprinted Placenta Model. Journal
SUPPLEMENTARY NOTE 3. Supplememtary Note 3, Wehr et al., Monitoring Regulated Protein-Protein Interactions Using Split-TEV 1
 SUPPLEMENTARY NOTE 3 Fluorescent Proteolysis-only TEV-Reporters We generated TEV reporter that allow visualizing TEV activity at the membrane and in the cytosol of living cells not relying on the addition
SUPPLEMENTARY NOTE 3 Fluorescent Proteolysis-only TEV-Reporters We generated TEV reporter that allow visualizing TEV activity at the membrane and in the cytosol of living cells not relying on the addition
Predictable Heating and Positive MRI Contrast from
 Predictable Heating and Positive MRI Contrast from a Mesoporous Silica-Coated Iron Oxide Nanoparticle Katie R. Hurley a, Hattie L. Ring ab, Michael Etheridge cd, Jinjin Zhang be, Zhe Gao ad, Qi Shao c,
Predictable Heating and Positive MRI Contrast from a Mesoporous Silica-Coated Iron Oxide Nanoparticle Katie R. Hurley a, Hattie L. Ring ab, Michael Etheridge cd, Jinjin Zhang be, Zhe Gao ad, Qi Shao c,
ab CFSE Fluorescent Cell Labeling Kit
 ab113853 CFSE Fluorescent Cell Labeling Kit Instructions for Use For the durable fluorescent labeling of live cells for fluorescent microscopy and flow cytometry, population growth studies and within sample
ab113853 CFSE Fluorescent Cell Labeling Kit Instructions for Use For the durable fluorescent labeling of live cells for fluorescent microscopy and flow cytometry, population growth studies and within sample
BD IMag. Streptavidin Particles Plus - DM. Technical Data Sheet. Product Information
 Technical Data Sheet Streptavidin Particles Plus - DM Product Information Material Number: Size: Storage Buffer: 557812 5 ml Aqueous buffered solution containing BSA and 0.09% sodium azide. Description
Technical Data Sheet Streptavidin Particles Plus - DM Product Information Material Number: Size: Storage Buffer: 557812 5 ml Aqueous buffered solution containing BSA and 0.09% sodium azide. Description
StemXVivo. Mesoderm Kit. Catalog Number SC030B. Reagents for the differentiation of human pluripotent stem cells into mesoderm.
 StemXVivo Mesoderm Kit Catalog Number SC030B Reagents for the differentiation of human pluripotent stem cells into mesoderm. This package insert must be read in its entirety before using this product.
StemXVivo Mesoderm Kit Catalog Number SC030B Reagents for the differentiation of human pluripotent stem cells into mesoderm. This package insert must be read in its entirety before using this product.
Poietics Rat Mesenchymal Stem Cells Instructions for Use
 Poietics Rat Mesenchymal Stem Cells Instructions for Use Lonza Walkersville, Inc. www.lonza.com scientific.support@lonza.com Scientific Support: 800-521-0390 Document # IS-PT-2505 08/09 Walkersville, MD
Poietics Rat Mesenchymal Stem Cells Instructions for Use Lonza Walkersville, Inc. www.lonza.com scientific.support@lonza.com Scientific Support: 800-521-0390 Document # IS-PT-2505 08/09 Walkersville, MD
ab JC1 - Mitochondrial Membrane Potential Assay Kit
 ab113850 JC1 - Mitochondrial Membrane Potential Assay Kit Instructions for Use For the measurement of mitochondrial membrane potential by fluorescence plate reader This product is for research use only
ab113850 JC1 - Mitochondrial Membrane Potential Assay Kit Instructions for Use For the measurement of mitochondrial membrane potential by fluorescence plate reader This product is for research use only
Cellartis MSC Xeno-Free Culture Medium
 Cat. # Y50200 For Research Use Cellartis MSC Xeno-Free Culture Medium Product Manual Table of Contents I. Description... 3 II. Contents... 3 III. Storage... 3 IV. Precautions... 3 V. Materials Required
Cat. # Y50200 For Research Use Cellartis MSC Xeno-Free Culture Medium Product Manual Table of Contents I. Description... 3 II. Contents... 3 III. Storage... 3 IV. Precautions... 3 V. Materials Required
Biosensis P1TM Polar Lipid Tracing Reagent. Catalogue Number: TR-600-P1. For research use only, not for use in clinical and diagnostic procedures.
 Biosensis P1TM Polar Lipid Tracing Reagent Catalogue Number: TR-600-P1 For research use only, not for use in clinical and diagnostic procedures. Version: TR-600-P1/v2/Sep2018 Biosensis Pty Ltd., 51 West
Biosensis P1TM Polar Lipid Tracing Reagent Catalogue Number: TR-600-P1 For research use only, not for use in clinical and diagnostic procedures. Version: TR-600-P1/v2/Sep2018 Biosensis Pty Ltd., 51 West
Mesenchymal Stem/Stromal Cells
 Mesenchymal Stem/Stromal Cells ABOUT US Irvine Scientific, a member of JX Holdings group, is a worldwide leader in the design, manufacture and distribution of medical devices, including Cell Therapy, Industrial
Mesenchymal Stem/Stromal Cells ABOUT US Irvine Scientific, a member of JX Holdings group, is a worldwide leader in the design, manufacture and distribution of medical devices, including Cell Therapy, Industrial
idisco+ protocol Recommendations for sample handling Buffers The following are given as a general guideline and may vary for specific applications.
 idisco+ protocol Recommendations for sample handling The following are given as a general guideline and may vary for specific applications. Buffers Sample type Incubation time ( n ) Solution volume Embryonic:
idisco+ protocol Recommendations for sample handling The following are given as a general guideline and may vary for specific applications. Buffers Sample type Incubation time ( n ) Solution volume Embryonic:
Preparation of Mouse Bone Marrow Stromal Cells
 Preparation of Mouse Bone Marrow Stromal Cells A single-step stem cell purification method using adhesion to cell culture plastic was employed as described in the Reference. Briefly, neonatal and adult
Preparation of Mouse Bone Marrow Stromal Cells A single-step stem cell purification method using adhesion to cell culture plastic was employed as described in the Reference. Briefly, neonatal and adult
CHO-Tet herg Cell Line
 F B SYS GmbH CHO-Tet herg Cell Line Specification Sheet B SYS GmbH B SYS GmbH CHO-Tet herg Cells Page 2 TABLE OF CONTENTS 1 BACKGROUND...3 1.1 Drug-induced QT Prolongation...3 1.2 Regulatory Issues...3
F B SYS GmbH CHO-Tet herg Cell Line Specification Sheet B SYS GmbH B SYS GmbH CHO-Tet herg Cells Page 2 TABLE OF CONTENTS 1 BACKGROUND...3 1.1 Drug-induced QT Prolongation...3 1.2 Regulatory Issues...3
BrdU IHC Kit. For the detection and localization of bromodeoxyuridine incorporated into newly synthesized DNA of actively proliferating cells
 K-ASSAY BrdU IHC Kit For the detection and localization of bromodeoxyuridine incorporated into newly synthesized DNA of actively proliferating cells Cat. No. KT-077 For Research Use Only. Not for Use in
K-ASSAY BrdU IHC Kit For the detection and localization of bromodeoxyuridine incorporated into newly synthesized DNA of actively proliferating cells Cat. No. KT-077 For Research Use Only. Not for Use in
Phagocytosis Assay Kit (IgG PE)
 Phagocytosis Assay Kit (IgG PE) Item No. 600540 www.caymanchem.com Customer Service 800.364.9897 Technical Support 888.526.5351 1180 E. Ellsworth Rd Ann Arbor, MI USA TABLE OF CONTENTS GENERAL INFORMATION
Phagocytosis Assay Kit (IgG PE) Item No. 600540 www.caymanchem.com Customer Service 800.364.9897 Technical Support 888.526.5351 1180 E. Ellsworth Rd Ann Arbor, MI USA TABLE OF CONTENTS GENERAL INFORMATION
SUPPLEMENTARY INFORMATION
 Biosynthesis of Luminescent Quantum Dots in an Earthworm S.R. Stürzenbaum, a# M. Hoeckner, a# A. Panneerselvam, b J. Levitt, b J.-S. Bouillard, b S. Taniguchi, b L.-A. Dailey, d R. Ahmad Khanbeigi, d E.
Biosynthesis of Luminescent Quantum Dots in an Earthworm S.R. Stürzenbaum, a# M. Hoeckner, a# A. Panneerselvam, b J. Levitt, b J.-S. Bouillard, b S. Taniguchi, b L.-A. Dailey, d R. Ahmad Khanbeigi, d E.
Reviewed: Hamilton. Contents; Overview. 2.0 Methods 3.0 Notes 4.0 Acknowledgements & References
 Microarray Core UCSF Comprehensive Cancer Center Standard Operating Procedure Title: Array CGH Hybridization Protocol HumArray 3.2 SOP No.: MC023QA Version: 5 Date: 5-12-06 Page No.: 1 of 5 Authors: Albertson,
Microarray Core UCSF Comprehensive Cancer Center Standard Operating Procedure Title: Array CGH Hybridization Protocol HumArray 3.2 SOP No.: MC023QA Version: 5 Date: 5-12-06 Page No.: 1 of 5 Authors: Albertson,
Stellaris FISH Probes Protocols and Storage
 Stellaris FISH Probes Protocols and Storage Catalog No. SMF-2035-1 Product Name Stellaris FISH Probes, Human MALAT1 with Quasar 570 Dye Product Description Product consists of Quasar 570-labeled oligos
Stellaris FISH Probes Protocols and Storage Catalog No. SMF-2035-1 Product Name Stellaris FISH Probes, Human MALAT1 with Quasar 570 Dye Product Description Product consists of Quasar 570-labeled oligos
Addendum B cell ELISPOT assay
 !"#$%&'()*+!"#$#""%&'( )*('&+,&-./$01.& U-CyTech BV 21$&3$.1$/#"%45 Yalelaan 48!&6)78)98:*)&*;0?.$0180@A& P +31.30.253.5960 &&&8>0?.$0180@A F
!"#$%&'()*+!"#$#""%&'( )*('&+,&-./$01.& U-CyTech BV 21$&3$.1$/#"%45 Yalelaan 48!&6)78)98:*)&*;0?.$0180@A& P +31.30.253.5960 &&&8>0?.$0180@A F
ab Mitochondrial Viability Assay
 ab129732 Mitochondrial Viability Assay Instructions for Use For measuring mitochondrial/cellular viability in high throughput This product is for research use only and is not intended for diagnostic use.
ab129732 Mitochondrial Viability Assay Instructions for Use For measuring mitochondrial/cellular viability in high throughput This product is for research use only and is not intended for diagnostic use.
A Supersandwich Fluorescence in Situ Hybridization (SFISH) Strategy. for Highly Sensitive and Selective mrna Imaging in Tumor Cells
 Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2015 Electronic Supplementary Information (ESI) A Supersandwich Fluorescence in Situ Hybridization (SFISH)
Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2015 Electronic Supplementary Information (ESI) A Supersandwich Fluorescence in Situ Hybridization (SFISH)
Protocols for Hematopoietic Differentiation of Murine ES Cells (in 6 or 24 well plates) Media Preparation
 Protocols for Hematopoietic Differentiation of Murine ES Cells (in 6 or 24 well plates) Media Preparation A) Embryonic Stem Cell Medium (ES) Dulbecco s modified Eagle s Medium (DMEM) with high glucose
Protocols for Hematopoietic Differentiation of Murine ES Cells (in 6 or 24 well plates) Media Preparation A) Embryonic Stem Cell Medium (ES) Dulbecco s modified Eagle s Medium (DMEM) with high glucose
Real-Time Monitoring of Arsenic Trioxide Release and Delivery
 Real-Time Monitoring of Arsenic Trioxide Release and Delivery by Activatable T 1 Imaging Zhenghuan Zhao, Xiaomin Wang, Zongjun Zhang, Hui Zhang, Hanyu Liu, Xianglong Zhu, Hui Li, Xiaoqin Chi, Zhenyu Yin,
Real-Time Monitoring of Arsenic Trioxide Release and Delivery by Activatable T 1 Imaging Zhenghuan Zhao, Xiaomin Wang, Zongjun Zhang, Hui Zhang, Hanyu Liu, Xianglong Zhu, Hui Li, Xiaoqin Chi, Zhenyu Yin,
Tube Formation Assays in µ-slide Angiogenesis. 1. General Information. Application Note 19
 Tube Formation Assays in µ-slide Angiogenesis Contents 1. General Information... 1 2. Material... 2 3. Work Flow Overview... 3 4. Preparation of the Gel and the Slide... 4 4.1. Gel Application... 4 4.2.
Tube Formation Assays in µ-slide Angiogenesis Contents 1. General Information... 1 2. Material... 2 3. Work Flow Overview... 3 4. Preparation of the Gel and the Slide... 4 4.1. Gel Application... 4 4.2.
Supplementary File 3: DNA and RNA isolation
 Supplementary File 3: DNA and RNA isolation Q-CROC-02 Biopsy protocol For the purposes of this protocol, four needle core biopsies (NCBs) of lymph node tissue are isolated from each patient using a 16G
Supplementary File 3: DNA and RNA isolation Q-CROC-02 Biopsy protocol For the purposes of this protocol, four needle core biopsies (NCBs) of lymph node tissue are isolated from each patient using a 16G
AdEasy Virus Purification Kits and 5 100
 AdEasy Virus Purification Kits 2 100 and 5 100 Instruction Manual Catalog #240243 (2 preps 100 ml), #240244 (5 preps 100 ml) Revision F.0 For Research Use Only. Not for use in diagnostic procedures. 240243-12
AdEasy Virus Purification Kits 2 100 and 5 100 Instruction Manual Catalog #240243 (2 preps 100 ml), #240244 (5 preps 100 ml) Revision F.0 For Research Use Only. Not for use in diagnostic procedures. 240243-12
Plasmid DNA transfection of SW480 human colorectal cancer cells with the Biontex K2 Transfection System
 Plasmid DNA transfection of human colorectal cancer cells with the Biontex K2 Transfection System Stephanie Hehlgans and Franz Rödel, Department of Radiotherapy and Oncology, Goethe- University Frankfurt,
Plasmid DNA transfection of human colorectal cancer cells with the Biontex K2 Transfection System Stephanie Hehlgans and Franz Rödel, Department of Radiotherapy and Oncology, Goethe- University Frankfurt,
ScienCell. Human Epidermal Keratinocytes-adult (HEK-a) Catalog Number: Research Laboratories
 ScienCell TM Research Laboratories Human Epidermal Keratinocytes-adult (HEK-a) Catalog Number: 2110 Cell Specification The epithelial layer of the skin provides an essential function as a protective barrier
ScienCell TM Research Laboratories Human Epidermal Keratinocytes-adult (HEK-a) Catalog Number: 2110 Cell Specification The epithelial layer of the skin provides an essential function as a protective barrier
Adaptation of hpscs from Different Culture Systems to Essential 8 Medium or Essential 8 Flex Medium
 Adaptation of hpscs from Different Culture Systems to Essential 8 Medium or Essential 8 Flex Medium Publication Part Number MAN0013768 Revision A.0 Introduction This protocol describes the transition of
Adaptation of hpscs from Different Culture Systems to Essential 8 Medium or Essential 8 Flex Medium Publication Part Number MAN0013768 Revision A.0 Introduction This protocol describes the transition of
Immunofluorescence and phalloidin labeling of mammalian cells
 Immunofluorescence and phalloidin labeling of mammalian cells 2 Contents Materials for immunofluorescence and phalloidin labeling of mammalian cells...1 Immunofluorescence-labelling on cultivated adherent
Immunofluorescence and phalloidin labeling of mammalian cells 2 Contents Materials for immunofluorescence and phalloidin labeling of mammalian cells...1 Immunofluorescence-labelling on cultivated adherent
Preparation for 3D cell culture on Alvetex Scaffold. 1. 3T3 cells were routinely maintained in T-75 flasks.
 Page 1 Introduction Alvetex Scaffold is currently available in four different cell culture formats: 24-well plate (AVP006), 12-well plate (AVP002), 6-well insert (AVP004), and 12-well insert (AVP005).
Page 1 Introduction Alvetex Scaffold is currently available in four different cell culture formats: 24-well plate (AVP006), 12-well plate (AVP002), 6-well insert (AVP004), and 12-well insert (AVP005).
TR DISTRIBUTION STATEMENT A: Approved for public release; distribution is unlimited.
 Chapter 14. Intracellular detection of viral transcription and replication using RNA FISH i. Summary/Abstract Many hemorrhagic fever viruses require BSL-3 or 4 laboratory containment for study. The necessary
Chapter 14. Intracellular detection of viral transcription and replication using RNA FISH i. Summary/Abstract Many hemorrhagic fever viruses require BSL-3 or 4 laboratory containment for study. The necessary
ReproRNA -OKSGM is a non-integrating, self-replicating RNA-based reprogramming vector for generating induced pluripotent stem (ips)
 Kit for generating ips cells using ReproRNA -OKSGM, a non-integrating, self-replicating RNA reprogramming vector Product Description ReproRNA -OKSGM is a non-integrating, self-replicating RNA-based reprogramming
Kit for generating ips cells using ReproRNA -OKSGM, a non-integrating, self-replicating RNA reprogramming vector Product Description ReproRNA -OKSGM is a non-integrating, self-replicating RNA-based reprogramming
USER GUIDE. Introduction. Catalog No. C10423, C10723
 USER GUIDE CellEvent Caspase-3/7 Green Detection Reagent Catalog No. C10423, C10723 Pub. No. MAN0003556 Rev. B.0 Table 1. Contents and storage Material C10423 Amount C10723 Concentration Storage* CellEvent
USER GUIDE CellEvent Caspase-3/7 Green Detection Reagent Catalog No. C10423, C10723 Pub. No. MAN0003556 Rev. B.0 Table 1. Contents and storage Material C10423 Amount C10723 Concentration Storage* CellEvent
High throughput screening: Huh-7 cells were seeded into 96-well plate (2000
 1 SUPPLEMENTARY INFORMATION METHODS 6 7 8 9 1 11 1 1 1 1 16 17 18 19 High throughput screening: Huh-7 cells were seeded into 96-well plate ( cells/well) and infected with MOI of DENV-. One hour post-infection
1 SUPPLEMENTARY INFORMATION METHODS 6 7 8 9 1 11 1 1 1 1 16 17 18 19 High throughput screening: Huh-7 cells were seeded into 96-well plate ( cells/well) and infected with MOI of DENV-. One hour post-infection
All quality control test results are reported on a lot specific Certificate of Analysis which is available at or upon request.
 PRIME-XV NPC Expansion XSFM PRIME-XV NPC Expansion XSFM is a xeno-and serum-free medium optimized for the cultivation and expansion of neural progenitor cells (NPC) that are able to maintain their ability
PRIME-XV NPC Expansion XSFM PRIME-XV NPC Expansion XSFM is a xeno-and serum-free medium optimized for the cultivation and expansion of neural progenitor cells (NPC) that are able to maintain their ability
Human Pluripotent Stem Cell Functional Identification Kit
 Human Pluripotent Stem Cell Functional Identification Kit Catalog Number SC027B Reagents for the identification of human pluripotent stem cells by in vitro functional differentiation. This package insert
Human Pluripotent Stem Cell Functional Identification Kit Catalog Number SC027B Reagents for the identification of human pluripotent stem cells by in vitro functional differentiation. This package insert
Acoustically targeted chemogenetics for the non-invasive control of neural circuits
 SUPPLEMENTARY INFORMATION Articles https://doi.org/10.1038/s41551-018-0258-2 In the format provided by the authors and unedited. Acoustically targeted chemogenetics for the non-invasive control of neural
SUPPLEMENTARY INFORMATION Articles https://doi.org/10.1038/s41551-018-0258-2 In the format provided by the authors and unedited. Acoustically targeted chemogenetics for the non-invasive control of neural
