Decoding Chromatin States with Epigenome Data Advanced Topics in Computa8onal Genomics
|
|
|
- Lillian Robbins
- 6 years ago
- Views:
Transcription
1 Decoding Chromatin States with Epigenome Data Advanced Topics in Computa8onal Genomics
2 HMMs for Decoding Chromatin States Epigene8c modifica8ons of the genome have been associated with Establishing cell iden88es during development DNA repair, replica8on Human diseases De novo discovery of chroma8n states given epigene8c marks with HMMs Emission probabili8es: which histone marks co- occur? Transi8on probabili8es: how chroma8n states are distributed spa8ally across the genome
3 Dataset Genome- wide occupancy data in human CD4 T- cells from ChIP- seq experiments 38 different histone methyla8on and acetyla8on marks Histone variant H2AZ RNA polymerase II CTCF E.g., H3K9me3 trimethylated lysine 9 of histone 3
4 HMMs for Decoding Chromatin States Hidden states for unknown chroma8n states Models with varying number of states 79 states, pruned to 51 states Histone mark data as observa8ons Data are binarized (a\er thresholding) for each window of size 200bp Binomial distribu8on for each histone mark as emission probability All histone marks are treated as independent
5 Example of Chromatin State Annotation Posterior probability of states at each locus, given data
6 Estimated Chromatin States - Emission Probabilities Emission probabili8es Genomic func8onal enrichment
7 GO Enrichment for Promoter States Although states 3-8 were promoter states, each state is enriched for genes with different GO categories
8 Comparison of Promoter States Different promoter states peak at different sites
9 Comparison of Transcribed States
10 GWAS and Chromatin States GWAS- enriched chroma8n state 33
11 Power for Discovering Chromatin States
12 Feature Selection We may not need all of the histone marks to explain the chroma8n state Feature selec8on as step- wise forward selec8on to select a subset of histone marks that describe the chroma8n state
13 Feature Selection
14 Epigenome and Gene Expression
15 Epigenome and Transcription Histone modifica8on levels can influence gene expressions Nucleosome posi8ons can influence gene expressions DNA sequence specifici8es of nucleosome and transcrip8on factor binding sites Nucleosomes as repressors Methyla8on usually represses transcrip8on
16 Key Questions Is there a quan8ta8ve rela8onship between histone modifica8ons levels and transcrip8on? Is there a subset of histone modifica8ons that predict transcrip8on becer than others? Are there different requirements for epigene8c marks for different promoter types? Do these rela8onships between histone modifica8ons and transcrip8on hold in different 8ssue types?
17 Dataset 38 histone modifica8ons and one histone variant in human CD4+ T- cells ChIP- seq data In a region of 4,001 bp surrounding the transcrip8on start sites of 14,801 RefSeq genes Gene expression levels in the CD4+ T- cells 9 histone modifica8ons in CD36+ and CD133+ cells Gene expression levels in CD36+ and CD133+ cells Histone modifica8on levels are predic8ve for gene expression. (Karlic et al., PNAS, 2010)
18 Linear Models Linear regression method Predictors: histone marks No binariza8on For genes with no histone modifica8ons for par8cular modifica8ons, add a pseudocount Responses: gene expressions Promoter regions of different genes as samples
19 Linear Models Full model including all histone modifica8ons Compute r 2 between observed gene expressions and predicted values to assess the predic8ve power of the model
20 Linear Models Selec8ng the histone modifica8ons with the most predic8ve power
21 Linear Models Selec8ng the histone modifica8ons with the most predic8ve power with BIC scores
22 Prediction Accuracy
23 Searching for Histone Modifications with the Most Predictive Power The most frequently appearing histone modifica8ons in models with 1, 2, 3 histone modifica8ons
24 Model with One Histone Modification Correla8ons between expressions and each histone modifica8on Redundancy in histone modifica8ons
25 Histone Modifications and Promoter Types Different promoter types to be considered LCPs : low CpG content promoters HCPs : high CpG content promoters Nucleosomes in HCPs almost always have H3K4me3 marks, whereas nucleosomes in LCPs carry this modifica8on only when they are expressed. Hypothesis: expression levels of genes with LCPs and HCPs can be predicted by different sets of histone modifica8ons
26 Histone Modifications and Promoter Types Experimental setup 1,779 LCPs and 7,089 HCPs in the dataset Fit different models to each of LCPs and HCPs and compare them with the model es8mated from the full dataset
27 Histone Modifications and Promoter Types
28 Considering Different Tissue Types Used the model trained on CD4+ data to predict gene expressions in CD133+ and CD36+ cells Used only those gene expressions with more than five fold differences between CD4+ and CD133+ (also between CD4+ and CD36+)
29 Nucleosome and Transcription DNA sequence mo8fs with high nucleosome binding affini8es Poten8ally related to bending DNA around the nucleosomes DNA sequence mo8fs with high transcrip8on factor binding affini8es TF concentra8on can also influence gene expression Compe88on between nucleosomes and transcrip8on factors can influence the transcrip8on
30 DNA Sequence, DNA-binding Proteins, and Gene Expression Mixture model for predic8ng gene expressions from nucleosomes and other DNA binding proteins E: gene expression C: protein configura8ons
31 DNA Sequence, DNA-binding Proteins, and Gene Expression Mixture propor8ons Mixture component models
32 Nucleosome and Transcription
33 Nucleosome and Transcription
34 Competition between Nucleosomes and Transcription Factors
35 Competition between Nucleosomes and Transcription Factors
36 Transcriptional Noise
37 Cooperative Binding Reduces Transcriptional Noise
38 Fuzzy Nucleosomes Well- posi8oned vs. fuzzy nucleosomes Can be inferred from DNA sequences In fuzzy nucleosomes, many nucleosome posi8ons are observed Well- posi8oned nucleosomes Fuzzy nucleosomes
39 Summary Histone modifica8ons contain informa8on on chroma8n states. Chroma8n states can be poten8ally decoded from epigene8c data. Epigene8cs and gene expressions histone modifica8ons can influence gene expression nucleosome posi8ons and the compe88on between TFs and nucleosome can influence gene expression
Nucleosome Positioning and Organization Advanced Topics in Computa8onal Genomics
 Nucleosome Positioning and Organization 02-715 Advanced Topics in Computa8onal Genomics Nucleosome Core Nucleosome Core and Linker 147 bp DNA wrapping around nucleosome core Varying lengths of linkers
Nucleosome Positioning and Organization 02-715 Advanced Topics in Computa8onal Genomics Nucleosome Core Nucleosome Core and Linker 147 bp DNA wrapping around nucleosome core Varying lengths of linkers
Making Deep Learning Understandable for Analyzing Sequen;al Data about Gene Regula;on. Dr. Yanjun Qi 2017/11/26
 Making Deep Learning Understandable for Analyzing Sequen;al Data about Gene Regula;on Dr. Yanjun Qi 2017/11/26 Roadmap ² Background of Machine Learning ² Background of Sequen?al Data about Gene Regula?on
Making Deep Learning Understandable for Analyzing Sequen;al Data about Gene Regula;on Dr. Yanjun Qi 2017/11/26 Roadmap ² Background of Machine Learning ² Background of Sequen?al Data about Gene Regula?on
2/19/13. Contents. Applications of HMMs in Epigenomics
 2/19/13 I529: Machine Learning in Bioinformatics (Spring 2013) Contents Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2013 Background:
2/19/13 I529: Machine Learning in Bioinformatics (Spring 2013) Contents Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2013 Background:
Applications of HMMs in Epigenomics
 I529: Machine Learning in Bioinformatics (Spring 2013) Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2013 Contents Background:
I529: Machine Learning in Bioinformatics (Spring 2013) Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2013 Contents Background:
Next- genera*on Sequencing. Lecture 13
 Next- genera*on Sequencing Lecture 13 ChIP- seq Applica*ons iden%fy sequence varia%ons DNA- seq Iden%fy Pathogens RNA- seq Kahvejian et al, 2008 Protein-DNA interaction DNA is the informa*on carrier of
Next- genera*on Sequencing Lecture 13 ChIP- seq Applica*ons iden%fy sequence varia%ons DNA- seq Iden%fy Pathogens RNA- seq Kahvejian et al, 2008 Protein-DNA interaction DNA is the informa*on carrier of
Gene Regulatory Networks Computa.onal Genomics Seyoung Kim
 Gene Regulatory Networks 02-710 Computa.onal Genomics Seyoung Kim Transcrip6on Factor Binding Transcrip6on Control Gene transcrip.on is influenced by Transcrip.on factor binding affinity for the regulatory
Gene Regulatory Networks 02-710 Computa.onal Genomics Seyoung Kim Transcrip6on Factor Binding Transcrip6on Control Gene transcrip.on is influenced by Transcrip.on factor binding affinity for the regulatory
Understanding transcriptional regulation by integrative analysis of transcription factor binding data
 Understanding transcriptional regulation by integrative analysis of transcription factor binding data Cheng et al. 2012 Shu Yang Feb. 21, 2013 1 / 26 Introduction 2 / 26 DNA-binding Proteins sequence-specific
Understanding transcriptional regulation by integrative analysis of transcription factor binding data Cheng et al. 2012 Shu Yang Feb. 21, 2013 1 / 26 Introduction 2 / 26 DNA-binding Proteins sequence-specific
V23: the histone code
 V23: the histone code The DNA of eukaryotic organisms is packaged into chromatin, whose basic repeating unit is the nucleosome. A nucleosome is formed by wrapping 147 base pairs of DNA twice around an
V23: the histone code The DNA of eukaryotic organisms is packaged into chromatin, whose basic repeating unit is the nucleosome. A nucleosome is formed by wrapping 147 base pairs of DNA twice around an
NGS Approaches to Epigenomics
 I519 Introduction to Bioinformatics, 2013 NGS Approaches to Epigenomics Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Contents Background: chromatin structure & DNA methylation Epigenomic
I519 Introduction to Bioinformatics, 2013 NGS Approaches to Epigenomics Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Contents Background: chromatin structure & DNA methylation Epigenomic
Lecture: Genetic Basis of Complex Phenotypes Advanced Topics in Computa8onal Genomics
 Lecture: Genetic Basis of Complex Phenotypes 02-715 Advanced Topics in Computa8onal Genomics Genome Polymorphisms A Human Genealogy TCGAGGTATTAAC The ancestral chromosome From SNPS TCGAGGTATTAAC TCTAGGTATTAAC
Lecture: Genetic Basis of Complex Phenotypes 02-715 Advanced Topics in Computa8onal Genomics Genome Polymorphisms A Human Genealogy TCGAGGTATTAAC The ancestral chromosome From SNPS TCGAGGTATTAAC TCTAGGTATTAAC
ChIP-seq analysis 2/28/2018
 ChIP-seq analysis 2/28/2018 Acknowledgements Much of the content of this lecture is from: Furey (2012) ChIP-seq and beyond Park (2009) ChIP-seq advantages + challenges Landt et al. (2012) ChIP-seq guidelines
ChIP-seq analysis 2/28/2018 Acknowledgements Much of the content of this lecture is from: Furey (2012) ChIP-seq and beyond Park (2009) ChIP-seq advantages + challenges Landt et al. (2012) ChIP-seq guidelines
Introduc)on to ChIP- Seq data analyses
 Introduc)on to ChIP- Seq data analyses 7 th BBBC, February 2014 Outline Introduc8on to ChIP- seq experiment: Biological mo8va8on. Experimental procedure. Method and sohware for ChIP- seq peak calling:
Introduc)on to ChIP- Seq data analyses 7 th BBBC, February 2014 Outline Introduc8on to ChIP- seq experiment: Biological mo8va8on. Experimental procedure. Method and sohware for ChIP- seq peak calling:
Epigenetics and DNase-Seq
 Epigenetics and DNase-Seq BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 2018 Anthony Gitter gitter@biostat.wisc.edu These slides, excluding third-party material, are licensed under CC BY-NC 4.0 by Anthony
Epigenetics and DNase-Seq BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 2018 Anthony Gitter gitter@biostat.wisc.edu These slides, excluding third-party material, are licensed under CC BY-NC 4.0 by Anthony
The ENCODE Encyclopedia. & Variant Annotation Using RegulomeDB and HaploReg
 The ENCODE Encyclopedia & Variant Annotation Using RegulomeDB and HaploReg Jill E. Moore Weng Lab University of Massachusetts Medical School October 10, 2015 Where s the Encyclopedia? ENCODE: Encyclopedia
The ENCODE Encyclopedia & Variant Annotation Using RegulomeDB and HaploReg Jill E. Moore Weng Lab University of Massachusetts Medical School October 10, 2015 Where s the Encyclopedia? ENCODE: Encyclopedia
Charles Girardot, Furlong Lab. MACS, CisGenome, SISSRs and other peak calling algorithms: differences and practical use
 Charles Girardot, Furlong Lab MACS, CisGenome, SISSRs and other peak calling algorithms: differences and practical use ChIP-Seq signal properties Only 5 ends of ChIPed fragments are sequenced Shifted read
Charles Girardot, Furlong Lab MACS, CisGenome, SISSRs and other peak calling algorithms: differences and practical use ChIP-Seq signal properties Only 5 ends of ChIPed fragments are sequenced Shifted read
Introduc)on to Chroma)n IP sequencing (ChIP- seq) data analysis
 Introduc)on to Chroma)n IP sequencing (ChIP- seq) data analysis Introduc)on to Bioinforma)cs Using NGS Data 27 January 2016 Agata Smialowska BILS, SciLife Lab, Stockholm University Chroma)n state and gene
Introduc)on to Chroma)n IP sequencing (ChIP- seq) data analysis Introduc)on to Bioinforma)cs Using NGS Data 27 January 2016 Agata Smialowska BILS, SciLife Lab, Stockholm University Chroma)n state and gene
From reads to results: differen1al expression analysis with RNA seq. Alicia Oshlack Bioinforma1cs Division Walter and Eliza Hall Ins1tute
 From reads to results: differen1al expression analysis with RNA seq Alicia Oshlack Bioinforma1cs Division Walter and Eliza Hall Ins1tute Purported benefits and opportuni1es of RNA seq All transcripts are
From reads to results: differen1al expression analysis with RNA seq Alicia Oshlack Bioinforma1cs Division Walter and Eliza Hall Ins1tute Purported benefits and opportuni1es of RNA seq All transcripts are
Genome 541! Unit 4, lecture 3! Genomics assays
 Genome 541! Unit 4, lecture 3! Genomics assays I d like a bit more background on the assays and bioterminology.!! The phantom peak concept was confusing.! I didn t quite understand what the phantom peak
Genome 541! Unit 4, lecture 3! Genomics assays I d like a bit more background on the assays and bioterminology.!! The phantom peak concept was confusing.! I didn t quite understand what the phantom peak
2/10/17. Contents. Applications of HMMs in Epigenomics
 2/10/17 I529: Machine Learning in Bioinformatics (Spring 2017) Contents Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2017 Background:
2/10/17 I529: Machine Learning in Bioinformatics (Spring 2017) Contents Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2017 Background:
Genome 541 Gene regulation and epigenomics Lecture 3 Integrative analysis of genomics assays
 Genome 541 Gene regulation and epigenomics Lecture 3 Integrative analysis of genomics assays Please consider both the forward and reverse strands (i.e. reverse compliment sequence). You do not need to
Genome 541 Gene regulation and epigenomics Lecture 3 Integrative analysis of genomics assays Please consider both the forward and reverse strands (i.e. reverse compliment sequence). You do not need to
Introduction to ChIP Seq data analyses. Acknowledgement: slides taken from Dr. H
 Introduction to ChIP Seq data analyses Acknowledgement: slides taken from Dr. H Wu @Emory ChIP seq: Chromatin ImmunoPrecipitation it ti + sequencing Same biological motivation as ChIP chip: measure specific
Introduction to ChIP Seq data analyses Acknowledgement: slides taken from Dr. H Wu @Emory ChIP seq: Chromatin ImmunoPrecipitation it ti + sequencing Same biological motivation as ChIP chip: measure specific
Introduc)on to Chroma)n IP sequencing (ChIP- seq) data analysis
 Introduc)on to Chroma)n IP sequencing (ChIP- seq) data analysis Introduc)on to Bioinforma)cs using NGS data Linköping, 21 April 2016 Agata Smialowska BILS / NBIS, SciLifeLab, Stockholm University Chroma)n
Introduc)on to Chroma)n IP sequencing (ChIP- seq) data analysis Introduc)on to Bioinforma)cs using NGS data Linköping, 21 April 2016 Agata Smialowska BILS / NBIS, SciLifeLab, Stockholm University Chroma)n
RNAseq / ChipSeq / Methylseq and personalized genomics
 RNAseq / ChipSeq / Methylseq and personalized genomics 7711 Lecture Subhajyo) De, PhD Division of Biomedical Informa)cs and Personalized Biomedicine, Department of Medicine University of Colorado School
RNAseq / ChipSeq / Methylseq and personalized genomics 7711 Lecture Subhajyo) De, PhD Division of Biomedical Informa)cs and Personalized Biomedicine, Department of Medicine University of Colorado School
A Brief History. Bootstrapping. Bagging. Boosting (Schapire 1989) Adaboost (Schapire 1995)
 A Brief History Bootstrapping Bagging Boosting (Schapire 1989) Adaboost (Schapire 1995) What s So Good About Adaboost Improves classification accuracy Can be used with many different classifiers Commonly
A Brief History Bootstrapping Bagging Boosting (Schapire 1989) Adaboost (Schapire 1995) What s So Good About Adaboost Improves classification accuracy Can be used with many different classifiers Commonly
Deep learning frameworks for regulatory genomics and epigenomics
 Deep learning frameworks for regulatory genomics and epigenomics Chuan Sheng Foo Avanti Shrikumar Nicholas Sinnott- Armstrong ANSHUL KUNDAJE Genetics, Computer science Stanford University Johnny Israeli
Deep learning frameworks for regulatory genomics and epigenomics Chuan Sheng Foo Avanti Shrikumar Nicholas Sinnott- Armstrong ANSHUL KUNDAJE Genetics, Computer science Stanford University Johnny Israeli
The ChIP-Seq project. Giovanna Ambrosini, Philipp Bucher. April 19, 2010 Lausanne. EPFL-SV Bucher Group
 The ChIP-Seq project Giovanna Ambrosini, Philipp Bucher EPFL-SV Bucher Group April 19, 2010 Lausanne Overview Focus on technical aspects Description of applications (C programs) Where to find binaries,
The ChIP-Seq project Giovanna Ambrosini, Philipp Bucher EPFL-SV Bucher Group April 19, 2010 Lausanne Overview Focus on technical aspects Description of applications (C programs) Where to find binaries,
Pathway Analysis in other data types
 Pathway Analysis in other data types Alison Motsinger-Reif, PhD Associate Professor Bioinforma
Pathway Analysis in other data types Alison Motsinger-Reif, PhD Associate Professor Bioinforma
Title: Genome-Wide Predictions of Transcription Factor Binding Events using Multi- Dimensional Genomic and Epigenomic Features Background
 Title: Genome-Wide Predictions of Transcription Factor Binding Events using Multi- Dimensional Genomic and Epigenomic Features Team members: David Moskowitz and Emily Tsang Background Transcription factors
Title: Genome-Wide Predictions of Transcription Factor Binding Events using Multi- Dimensional Genomic and Epigenomic Features Team members: David Moskowitz and Emily Tsang Background Transcription factors
Introduc)on to ChIP- seq data analysis
 Introduc)on to ChIP- seq data analysis ENAR 2014 Hao Wu, Emory University Outline Introduc=on to ChIP- seq experiment: biological mo=va=on and experimental procedure. Method and sokware for ChIP- seq peak
Introduc)on to ChIP- seq data analysis ENAR 2014 Hao Wu, Emory University Outline Introduc=on to ChIP- seq experiment: biological mo=va=on and experimental procedure. Method and sokware for ChIP- seq peak
What Can the Epigenome Teach Us About Cellular States and Diseases?
 What Can the Epigenome Teach Us About Cellular States and Diseases? (a computer scientist s view) Luca Pinello Outline Epigenetic: the code over the code What can we learn from epigenomic data? Resources
What Can the Epigenome Teach Us About Cellular States and Diseases? (a computer scientist s view) Luca Pinello Outline Epigenetic: the code over the code What can we learn from epigenomic data? Resources
ChIP-Seq Tools. J Fass UCD Genome Center Bioinformatics Core Wednesday September 16, 2015
 ChIP-Seq Tools J Fass UCD Genome Center Bioinformatics Core Wednesday September 16, 2015 What s the Question? Where do Transcription Factors (TFs) bind genomic DNA 1? (Where do other things bind DNA or
ChIP-Seq Tools J Fass UCD Genome Center Bioinformatics Core Wednesday September 16, 2015 What s the Question? Where do Transcription Factors (TFs) bind genomic DNA 1? (Where do other things bind DNA or
ChIP-Seq Data Analysis. J Fass UCD Genome Center Bioinformatics Core Wednesday 15 June 2015
 ChIP-Seq Data Analysis J Fass UCD Genome Center Bioinformatics Core Wednesday 15 June 2015 What s the Question? Where do Transcription Factors (TFs) bind genomic DNA 1? (Where do other things bind DNA
ChIP-Seq Data Analysis J Fass UCD Genome Center Bioinformatics Core Wednesday 15 June 2015 What s the Question? Where do Transcription Factors (TFs) bind genomic DNA 1? (Where do other things bind DNA
ChIP-Seq Data Analysis. J Fass UCD Genome Center Bioinformatics Core Wednesday December 17, 2014
 ChIP-Seq Data Analysis J Fass UCD Genome Center Bioinformatics Core Wednesday December 17, 2014 What s the Question? Where do Transcription Factors (TFs) bind genomic DNA 1? (Where do other things bind
ChIP-Seq Data Analysis J Fass UCD Genome Center Bioinformatics Core Wednesday December 17, 2014 What s the Question? Where do Transcription Factors (TFs) bind genomic DNA 1? (Where do other things bind
How it All Works. Sample. Data analysis. Library Prepara>on. Sequencing
 Library PREP How it All Works Extract DNA Fragment Sample Data analysis Sequencing Library Prepara>on Polymerase Chain Reaction Polymerase Chain Reaction Polymerase Chain Reaction Polymerase Chain Reaction
Library PREP How it All Works Extract DNA Fragment Sample Data analysis Sequencing Library Prepara>on Polymerase Chain Reaction Polymerase Chain Reaction Polymerase Chain Reaction Polymerase Chain Reaction
ChIP. November 21, 2017
 ChIP November 21, 2017 functional signals: is DNA enough? what is the smallest number of letters used by a written language? DNA is only one part of the functional genome DNA is heavily bound by proteins,
ChIP November 21, 2017 functional signals: is DNA enough? what is the smallest number of letters used by a written language? DNA is only one part of the functional genome DNA is heavily bound by proteins,
University of Groningen
 University of Groningen Accounting for immunoprecipitation efficiencies in the statistical analysis of ChIP-seq data Bao, Yanchun; Vinciotti, Veronica; Wit, Ernst; 't Hoen, Peter A C Published in: Bmc
University of Groningen Accounting for immunoprecipitation efficiencies in the statistical analysis of ChIP-seq data Bao, Yanchun; Vinciotti, Veronica; Wit, Ernst; 't Hoen, Peter A C Published in: Bmc
Perm-seq: Mapping Protein-DNA Interactions in Segmental Duplication and Highly Repetitive Regions of Genomes with Prior- Enhanced Read Mapping
 RESEARCH ARTICLE Perm-seq: Mapping Protein-DNA Interactions in Segmental Duplication and Highly Repetitive Regions of Genomes with Prior- Enhanced Read Mapping Xin Zeng 1,BoLi 2, Rene Welch 1, Constanza
RESEARCH ARTICLE Perm-seq: Mapping Protein-DNA Interactions in Segmental Duplication and Highly Repetitive Regions of Genomes with Prior- Enhanced Read Mapping Xin Zeng 1,BoLi 2, Rene Welch 1, Constanza
Bio5488 Practice Midterm (2018) 1. Next-gen sequencing
 1. Next-gen sequencing 1. You have found a new strain of yeast that makes fantastic wine. You d like to sequence this strain to ascertain the differences from S. cerevisiae. To accurately call a base pair,
1. Next-gen sequencing 1. You have found a new strain of yeast that makes fantastic wine. You d like to sequence this strain to ascertain the differences from S. cerevisiae. To accurately call a base pair,
Bioinformatics of Transcriptional Regulation
 Bioinformatics of Transcriptional Regulation Carl Herrmann IPMB & DKFZ c.herrmann@dkfz.de Wechselwirkung von Maßnahmen und Auswirkungen Einflussmöglichkeiten in einem Dialog From genes to active compounds
Bioinformatics of Transcriptional Regulation Carl Herrmann IPMB & DKFZ c.herrmann@dkfz.de Wechselwirkung von Maßnahmen und Auswirkungen Einflussmöglichkeiten in einem Dialog From genes to active compounds
Measuring Protein-DNA interactions
 Measuring Protein-DNA interactions How is Biological Complexity Achieved? Mediated by Transcription Factors (TFs) 2 Transcription Factors are genetic switches 3 Regulation of Gene Expression by Transcription
Measuring Protein-DNA interactions How is Biological Complexity Achieved? Mediated by Transcription Factors (TFs) 2 Transcription Factors are genetic switches 3 Regulation of Gene Expression by Transcription
Large-scale imputation of epigenomic datasets for systematic annotation of diverse human tissues
 Large-scale imputation of epigenomic datasets for systematic annotation of diverse human tissues The MIT Faculty has made this article openly available. Please share how this access benefits you. Your
Large-scale imputation of epigenomic datasets for systematic annotation of diverse human tissues The MIT Faculty has made this article openly available. Please share how this access benefits you. Your
Week6 The molecular basis for growth and reproduc9on
 Week6 The molecular basis for growth and reproduc9on 1. Review: Central dogma (and beyond) 2. Structure of DNA and its replica9on 3. Transla9ng the gene9c code 4. Using ribosomal RNA to infer the evolu9onary
Week6 The molecular basis for growth and reproduc9on 1. Review: Central dogma (and beyond) 2. Structure of DNA and its replica9on 3. Transla9ng the gene9c code 4. Using ribosomal RNA to infer the evolu9onary
Supplementary Table 1: Oligo designs. A list of ATAC-seq oligos used for PCR.
 Ad1_noMX: Ad2.1_TAAGGCGA Ad2.2_CGTACTAG Ad2.3_AGGCAGAA Ad2.4_TCCTGAGC Ad2.5_GGACTCCT Ad2.6_TAGGCATG Ad2.7_CTCTCTAC Ad2.8_CAGAGAGG Ad2.9_GCTACGCT Ad2.10_CGAGGCTG Ad2.11_AAGAGGCA Ad2.12_GTAGAGGA Ad2.13_GTCGTGAT
Ad1_noMX: Ad2.1_TAAGGCGA Ad2.2_CGTACTAG Ad2.3_AGGCAGAA Ad2.4_TCCTGAGC Ad2.5_GGACTCCT Ad2.6_TAGGCATG Ad2.7_CTCTCTAC Ad2.8_CAGAGAGG Ad2.9_GCTACGCT Ad2.10_CGAGGCTG Ad2.11_AAGAGGCA Ad2.12_GTAGAGGA Ad2.13_GTCGTGAT
Green Center Computational Core ChIP- Seq Pipeline, Just a Click Away
 Green Center Computational Core ChIP- Seq Pipeline, Just a Click Away Venkat Malladi Computational Biologist Computational Core Cecil H. and Ida Green Center for Reproductive Biology Science Introduc
Green Center Computational Core ChIP- Seq Pipeline, Just a Click Away Venkat Malladi Computational Biologist Computational Core Cecil H. and Ida Green Center for Reproductive Biology Science Introduc
MCDB 1041 Class 21 Splicing and Gene Expression
 MCDB 1041 Class 21 Splicing and Gene Expression Learning Goals Describe the role of introns and exons Interpret the possible outcomes of alternative splicing Relate the generation of protein from DNA to
MCDB 1041 Class 21 Splicing and Gene Expression Learning Goals Describe the role of introns and exons Interpret the possible outcomes of alternative splicing Relate the generation of protein from DNA to
CS273B: Deep learning for Genomics and Biomedicine
 CS273B: Deep learning for Genomics and Biomedicine Lecture 2: Convolutional neural networks and applications to functional genomics 09/28/2016 Anshul Kundaje, James Zou, Serafim Batzoglou Outline Anatomy
CS273B: Deep learning for Genomics and Biomedicine Lecture 2: Convolutional neural networks and applications to functional genomics 09/28/2016 Anshul Kundaje, James Zou, Serafim Batzoglou Outline Anatomy
DOUBLE-STRAND DNA BREAK PREDICTION USING EPIGENOME MARKS AT KILOBASE RESOLUTION
 DOUBLE-STRAND DNA BREAK PREDICTION USING EPIGENOME MARKS AT KILOBASE RESOLUTION Raphaël MOURAD, Assist. Prof. Centre de Biologie Intégrative Université Paul Sabatier, Toulouse III INTRODUCTION Double-strand
DOUBLE-STRAND DNA BREAK PREDICTION USING EPIGENOME MARKS AT KILOBASE RESOLUTION Raphaël MOURAD, Assist. Prof. Centre de Biologie Intégrative Université Paul Sabatier, Toulouse III INTRODUCTION Double-strand
ChIP-seq and RNA-seq. Farhat Habib
 ChIP-seq and RNA-seq Farhat Habib fhabib@iiserpune.ac.in Biological Goals Learn how genomes encode the diverse patterns of gene expression that define each cell type and state. Protein-DNA interactions
ChIP-seq and RNA-seq Farhat Habib fhabib@iiserpune.ac.in Biological Goals Learn how genomes encode the diverse patterns of gene expression that define each cell type and state. Protein-DNA interactions
Genome 373: Gene Predic/on I. Doug Fowler
 Genome 373: Gene Predic/on I Doug Fowler Outline Review of gene structure Scale of the problem Solu;ons Empirical methods Ab ini&o predic;on What is a gene? A locatable region of genomic sequence, corresponding
Genome 373: Gene Predic/on I Doug Fowler Outline Review of gene structure Scale of the problem Solu;ons Empirical methods Ab ini&o predic;on What is a gene? A locatable region of genomic sequence, corresponding
Nature Genetics: doi: /ng Supplementary Figure 1. H3K27ac HiChIP enriches enhancer promoter-associated chromatin contacts.
 Supplementary Figure 1 H3K27ac HiChIP enriches enhancer promoter-associated chromatin contacts. (a) Schematic of chromatin contacts captured in H3K27ac HiChIP. (b) Loop call overlap for cohesin HiChIP
Supplementary Figure 1 H3K27ac HiChIP enriches enhancer promoter-associated chromatin contacts. (a) Schematic of chromatin contacts captured in H3K27ac HiChIP. (b) Loop call overlap for cohesin HiChIP
RNA-SEQUENCING ANALYSIS
 RNA-SEQUENCING ANALYSIS Joseph Powell SISG- 2018 CONTENTS Introduction to RNA sequencing Data structure Analyses Transcript counting Alternative splicing Allele specific expression Discovery APPLICATIONS
RNA-SEQUENCING ANALYSIS Joseph Powell SISG- 2018 CONTENTS Introduction to RNA sequencing Data structure Analyses Transcript counting Alternative splicing Allele specific expression Discovery APPLICATIONS
ChIP-Seq, mrna-seq, & Resequencing via the Genboree Workench
 ChIP-Seq, mrna-seq, & Resequencing via the Genboree Workench Chromatin ImmunopreciPitation Sequencing ChIP-Seq Peak Calling Transcription Factors Histone modifications H3K4me3, H3K27me3, H3K36me3, H3K9me3,
ChIP-Seq, mrna-seq, & Resequencing via the Genboree Workench Chromatin ImmunopreciPitation Sequencing ChIP-Seq Peak Calling Transcription Factors Histone modifications H3K4me3, H3K27me3, H3K36me3, H3K9me3,
RNAseq Applications in Genome Studies. Alexander Kanapin, PhD Wellcome Trust Centre for Human Genetics, University of Oxford
 RNAseq Applications in Genome Studies Alexander Kanapin, PhD Wellcome Trust Centre for Human Genetics, University of Oxford RNAseq Protocols Next generation sequencing protocol cdna, not RNA sequencing
RNAseq Applications in Genome Studies Alexander Kanapin, PhD Wellcome Trust Centre for Human Genetics, University of Oxford RNAseq Protocols Next generation sequencing protocol cdna, not RNA sequencing
Gene Expression Microarrays. For microarrays, purity of the RNA was further assessed by
 Supplemental Methods Gene Expression Microarrays. For microarrays, purity of the RNA was further assessed by an Agilent 2100 Bioanalyzer. 500 ng of RNA was reverse transcribed into crna and biotin-utp
Supplemental Methods Gene Expression Microarrays. For microarrays, purity of the RNA was further assessed by an Agilent 2100 Bioanalyzer. 500 ng of RNA was reverse transcribed into crna and biotin-utp
DNA, Genes and their Regula4on. Me#e Voldby Larsen PhD, Assistant Professor
 DNA, Genes and their Regula4on Me#e Voldby Larsen PhD, Assistant Professor Learning Objecer this talk, you should be able to Account for the structure of DNA and RNA including their similari
DNA, Genes and their Regula4on Me#e Voldby Larsen PhD, Assistant Professor Learning Objecer this talk, you should be able to Account for the structure of DNA and RNA including their similari
Sta$s$cs for Genomics ( )
 Sta$s$cs for Genomics (140.688) Instructor: Jeff Leek Website: http://www.biostat.jhsph.edu/~jleek/teaching/2011/genomics/ Class Times: MW, 10:30AM-11:50AM + R Lab TBA Grading: 20% Reading Assignments,
Sta$s$cs for Genomics (140.688) Instructor: Jeff Leek Website: http://www.biostat.jhsph.edu/~jleek/teaching/2011/genomics/ Class Times: MW, 10:30AM-11:50AM + R Lab TBA Grading: 20% Reading Assignments,
Integrating diverse datasets improves developmental enhancer prediction
 1 Integrating diverse datasets improves developmental enhancer prediction Genevieve D. Erwin 1, Rebecca M. Truty 1, Dennis Kostka 2, Katherine S. Pollard 1,3, John A. Capra 4 1 Gladstone Institute of Cardiovascular
1 Integrating diverse datasets improves developmental enhancer prediction Genevieve D. Erwin 1, Rebecca M. Truty 1, Dennis Kostka 2, Katherine S. Pollard 1,3, John A. Capra 4 1 Gladstone Institute of Cardiovascular
Complete Sample to Analysis Solutions for DNA Methylation Discovery using Next Generation Sequencing
 Complete Sample to Analysis Solutions for DNA Methylation Discovery using Next Generation Sequencing SureSelect Human/Mouse Methyl-Seq Kyeong Jeong PhD February 5, 2013 CAG EMEAI DGG/GSD/GFO Agilent Restricted
Complete Sample to Analysis Solutions for DNA Methylation Discovery using Next Generation Sequencing SureSelect Human/Mouse Methyl-Seq Kyeong Jeong PhD February 5, 2013 CAG EMEAI DGG/GSD/GFO Agilent Restricted
Transcription start site classification
 Transcription start site classification Max Libbrecht, Matt Fisher, Roy Frostig, Hrysoula Papadakis, Anshul Kundaje, Serafim Batzoglou December 11, 2009 Abstract Understanding the mechanisms of gene expression
Transcription start site classification Max Libbrecht, Matt Fisher, Roy Frostig, Hrysoula Papadakis, Anshul Kundaje, Serafim Batzoglou December 11, 2009 Abstract Understanding the mechanisms of gene expression
Chapter 17 Lecture. Concepts of Genetics. Tenth Edition. Regulation of Gene Expression in Eukaryotes
 Chapter 17 Lecture Concepts of Genetics Tenth Edition Regulation of Gene Expression in Eukaryotes Chapter Contents 17.1 Eukaryotic Gene Regulation Can Occur at Any of the Steps Leading from DNA to Protein
Chapter 17 Lecture Concepts of Genetics Tenth Edition Regulation of Gene Expression in Eukaryotes Chapter Contents 17.1 Eukaryotic Gene Regulation Can Occur at Any of the Steps Leading from DNA to Protein
Computa(onal Approaches to Promoter Analysis: Part I. Shifra Ben Dor May 2010
 Computa(onal Approaches to Promoter Analysis: Part I Shifra Ben Dor May 2010 Why use a computer to study promoters? Wet techniques for promoter discovery are (me and material consuming Planning your experiments
Computa(onal Approaches to Promoter Analysis: Part I Shifra Ben Dor May 2010 Why use a computer to study promoters? Wet techniques for promoter discovery are (me and material consuming Planning your experiments
Ensembl Funcgen: A Database and API for Epigenomics and Gene Regulation Data.
 Ensembl Funcgen: A Database and API for Epigenomics and Gene Regulation Data. Nathan Johnson Ensembl Regulation EBI is an Outstation of the European Molecular Biology Laboratory.! Workshop Overview http://www.ebi.ac.uk/~njohnson/courses/23.05.2013-
Ensembl Funcgen: A Database and API for Epigenomics and Gene Regulation Data. Nathan Johnson Ensembl Regulation EBI is an Outstation of the European Molecular Biology Laboratory.! Workshop Overview http://www.ebi.ac.uk/~njohnson/courses/23.05.2013-
Whole Transcriptome Analysis of Illumina RNA- Seq Data. Ryan Peters Field Application Specialist
 Whole Transcriptome Analysis of Illumina RNA- Seq Data Ryan Peters Field Application Specialist Partek GS in your NGS Pipeline Your Start-to-Finish Solution for Analysis of Next Generation Sequencing Data
Whole Transcriptome Analysis of Illumina RNA- Seq Data Ryan Peters Field Application Specialist Partek GS in your NGS Pipeline Your Start-to-Finish Solution for Analysis of Next Generation Sequencing Data
PPAR-γ - Peroxisome proliferator-ac4vated receptor gamma
 PPAR-γ - Peroxisome proliferator-ac4vated receptor gamma PPAR-γ - Peroxisome proliferator-ac4vated receptor gamma C/EBPalpha - CCAAT/enhancer-binding protein alpha C/EBPα contains: - a func)onally related
PPAR-γ - Peroxisome proliferator-ac4vated receptor gamma PPAR-γ - Peroxisome proliferator-ac4vated receptor gamma C/EBPalpha - CCAAT/enhancer-binding protein alpha C/EBPα contains: - a func)onally related
Galaxy Platform For NGS Data Analyses
 Galaxy Platform For NGS Data Analyses Weihong Yan wyan@chem.ucla.edu Collaboratory Web Site http://qcb.ucla.edu/collaboratory http://collaboratory.lifesci.ucla.edu Workshop Outline ü Day 1 UCLA galaxy
Galaxy Platform For NGS Data Analyses Weihong Yan wyan@chem.ucla.edu Collaboratory Web Site http://qcb.ucla.edu/collaboratory http://collaboratory.lifesci.ucla.edu Workshop Outline ü Day 1 UCLA galaxy
Gene expression analysis. Biosciences 741: Genomics Fall, 2013 Week 5. Gene expression analysis
 Gene expression analysis Biosciences 741: Genomics Fall, 2013 Week 5 Gene expression analysis From EST clusters to spotted cdna microarrays Long vs. short oligonucleotide microarrays vs. RT-PCR Methods
Gene expression analysis Biosciences 741: Genomics Fall, 2013 Week 5 Gene expression analysis From EST clusters to spotted cdna microarrays Long vs. short oligonucleotide microarrays vs. RT-PCR Methods
Bi 8 Lecture 3. DNA to RNA: Transcription and Splicing. Katalin Fejes Toth 12 January 2016
 Bi 8 Lecture 3 DNA to RNA: Transcription and Splicing Katalin Fejes Toth 12 January 2016 Alberts 6 th edi5on: Ch. 6: pp. 301-333 Readings for this lecture (Alberts 5 th edi5on equivalents: Ch. 6: pp. 329-366)
Bi 8 Lecture 3 DNA to RNA: Transcription and Splicing Katalin Fejes Toth 12 January 2016 Alberts 6 th edi5on: Ch. 6: pp. 301-333 Readings for this lecture (Alberts 5 th edi5on equivalents: Ch. 6: pp. 329-366)
Computational Analysis of Ultra-high-throughput sequencing data: ChIP-Seq
 Computational Analysis of Ultra-high-throughput sequencing data: ChIP-Seq Philipp Bucher Wednesday January 21, 2009 SIB graduate school course EPFL, Lausanne Data flow in ChIP-Seq data analysis Level 1:
Computational Analysis of Ultra-high-throughput sequencing data: ChIP-Seq Philipp Bucher Wednesday January 21, 2009 SIB graduate school course EPFL, Lausanne Data flow in ChIP-Seq data analysis Level 1:
Cis ac$ng transcrip$onal elements with nega$ve regulatory func$on
 Cis ac$ng transcrip$onal elements with nega$ve regulatory func$on Laura Elnitski, PhD Na$onal Human Genome Research Ins$tute Na$onal Ins$tutes of Health 2 Non coding regulatory signals affec$ng transcrip$on
Cis ac$ng transcrip$onal elements with nega$ve regulatory func$on Laura Elnitski, PhD Na$onal Human Genome Research Ins$tute Na$onal Ins$tutes of Health 2 Non coding regulatory signals affec$ng transcrip$on
Chapter 24: Promoters and Enhancers
 Chapter 24: Promoters and Enhancers A typical gene transcribed by RNA polymerase II has a promoter that usually extends upstream from the site where transcription is initiated the (#1) of transcription
Chapter 24: Promoters and Enhancers A typical gene transcribed by RNA polymerase II has a promoter that usually extends upstream from the site where transcription is initiated the (#1) of transcription
MODULE TSS1: TRANSCRIPTION START SITES INTRODUCTION (BASIC)
 MODULE TSS1: TRANSCRIPTION START SITES INTRODUCTION (BASIC) Lesson Plan: Title JAMIE SIDERS, MEG LAAKSO & WILSON LEUNG Identifying transcription start sites for Peaked promoters using chromatin landscape,
MODULE TSS1: TRANSCRIPTION START SITES INTRODUCTION (BASIC) Lesson Plan: Title JAMIE SIDERS, MEG LAAKSO & WILSON LEUNG Identifying transcription start sites for Peaked promoters using chromatin landscape,
GATCGTGCACGATCTCGGCAATTCGGGATGCCGGCTCGTCACCGGTCGCT
 Problem. (pts) A. (5pts) Your colleague professor Eugene Mathew Lateed generated a genome-wide DNA methylation map for normal colon cells using MRE-seq and MeDIP-seq. In an intergenic region, he found
Problem. (pts) A. (5pts) Your colleague professor Eugene Mathew Lateed generated a genome-wide DNA methylation map for normal colon cells using MRE-seq and MeDIP-seq. In an intergenic region, he found
Introduction to genome biology
 Introduction to genome biology Lisa Stubbs Deep transcritpomes for traditional model species from ENCODE (and modencode) Deep RNA-seq and chromatin analysis on 147 human cell types, as well as tissues,
Introduction to genome biology Lisa Stubbs Deep transcritpomes for traditional model species from ENCODE (and modencode) Deep RNA-seq and chromatin analysis on 147 human cell types, as well as tissues,
The ENCODE Encyclopedia. Zhiping Weng University of Massachusetts Medical School ENCODE 2016: Research Applications and Users Meeting June 8, 2016
 The ENCODE Encyclopedia Zhiping Weng University of Massachusetts Medical School ENCODE 2016: Research Applications and Users Meeting June 8, 2016 The Encyclopedia Of DNA Elements Consortium Goals: - Catalog
The ENCODE Encyclopedia Zhiping Weng University of Massachusetts Medical School ENCODE 2016: Research Applications and Users Meeting June 8, 2016 The Encyclopedia Of DNA Elements Consortium Goals: - Catalog
A survey on computational methods for enhancer and enhancer target predictions Introduction
 A survey on computational methods for enhancer and enhancer target predictions Qin Cao and Kevin Y. Yip Department of Computer Science and Engineering, The Chinese University of Hong Kong, Shatin, New
A survey on computational methods for enhancer and enhancer target predictions Qin Cao and Kevin Y. Yip Department of Computer Science and Engineering, The Chinese University of Hong Kong, Shatin, New
Gene expression DNA RNA. Protein DNA. Replication. Initiation Elongation Processing Export. DNA RNA Protein. Transcription. Degradation.
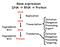 Gene expression DNA RNA Protein DNA DNA Degradation RNA Degradation Protein Replication Transcription Translation Initiation Elongation Processing Export Initiation Elongation Processing Targeting Chapter
Gene expression DNA RNA Protein DNA DNA Degradation RNA Degradation Protein Replication Transcription Translation Initiation Elongation Processing Export Initiation Elongation Processing Targeting Chapter
PS#2: We will begin at 10:30 am with Problem 1 in BH212. Assignments Problem 1- group 4 Problem 2- group 3 Problem 3- group 2 Problem 4- group 1
 PS#2: We will begin at 10:30 am with Problem 1 in BH212 Assignments Problem 1- group 4 Problem 2- group 3 Problem 3- group 2 Problem 4- group 1 Time limit: 25 min per group. The beginning: Designate one
PS#2: We will begin at 10:30 am with Problem 1 in BH212 Assignments Problem 1- group 4 Problem 2- group 3 Problem 3- group 2 Problem 4- group 1 Time limit: 25 min per group. The beginning: Designate one
ChIP-seq data analysis with Chipster. Eija Korpelainen CSC IT Center for Science, Finland
 ChIP-seq data analysis with Chipster Eija Korpelainen CSC IT Center for Science, Finland chipster@csc.fi What will I learn? Short introduction to ChIP-seq Analyzing ChIP-seq data Central concepts Analysis
ChIP-seq data analysis with Chipster Eija Korpelainen CSC IT Center for Science, Finland chipster@csc.fi What will I learn? Short introduction to ChIP-seq Analyzing ChIP-seq data Central concepts Analysis
Spatial Clustering of Multivariate Genomic and Epigenomic Information
 Spatial Clustering of Multivariate Genomic and Epigenomic Information Rami Jaschek and Amos Tanay Department of Computer Science and Applied Mathematics. The Weizmann Institute of Science, Rehovot, 76100
Spatial Clustering of Multivariate Genomic and Epigenomic Information Rami Jaschek and Amos Tanay Department of Computer Science and Applied Mathematics. The Weizmann Institute of Science, Rehovot, 76100
Caroline Townsend December 2012 Biochem 218 A critical review of ChIP-seq enrichment analysis tools
 Caroline Townsend December 2012 Biochem 218 A critical review of ChIP-seq enrichment analysis tools Introduction Transcriptional regulation, chromatin states, and genome stability pathways are largely
Caroline Townsend December 2012 Biochem 218 A critical review of ChIP-seq enrichment analysis tools Introduction Transcriptional regulation, chromatin states, and genome stability pathways are largely
DNA:CHROMATIN INTERACTIONS
 DNA:CHROMATIN INTERACTIONS Exploring transcription factor binding and the epigenomic landscape Chris Seward Introductions Cell and Developmental Biology PhD Candidate in Dr. Lisa Stubbs Laboratory Currently
DNA:CHROMATIN INTERACTIONS Exploring transcription factor binding and the epigenomic landscape Chris Seward Introductions Cell and Developmental Biology PhD Candidate in Dr. Lisa Stubbs Laboratory Currently
ChIP-seq and RNA-seq
 ChIP-seq and RNA-seq Biological Goals Learn how genomes encode the diverse patterns of gene expression that define each cell type and state. Protein-DNA interactions (ChIPchromatin immunoprecipitation)
ChIP-seq and RNA-seq Biological Goals Learn how genomes encode the diverse patterns of gene expression that define each cell type and state. Protein-DNA interactions (ChIPchromatin immunoprecipitation)
methylpipe: a library for the analysis of base- resolu5on DNA methyla5on data
 methylpipe: a library for the analysis of base- resolu5on DNA methyla5on data Bioconductor European Developers' Workshop 2012 University of Zurich MaGa Pelizzola - Center for Genomic Science of IIT@SEMM
methylpipe: a library for the analysis of base- resolu5on DNA methyla5on data Bioconductor European Developers' Workshop 2012 University of Zurich MaGa Pelizzola - Center for Genomic Science of IIT@SEMM
Editorial. Current Computational Models for Prediction of the Varied Interactions Related to Non-Coding RNAs
 Editorial Current Computational Models for Prediction of the Varied Interactions Related to Non-Coding RNAs Xing Chen 1,*, Huiming Peng 2, Zheng Yin 3 1 School of Information and Electrical Engineering,
Editorial Current Computational Models for Prediction of the Varied Interactions Related to Non-Coding RNAs Xing Chen 1,*, Huiming Peng 2, Zheng Yin 3 1 School of Information and Electrical Engineering,
Go to Bottom Left click WashU Epigenome Browser. Click
 Now you are going to look at the Human Epigenome Browswer. It has a more sophisticated but weirder interface than the UCSC Genome Browser. All the data that you will view as tracks is in reality just files
Now you are going to look at the Human Epigenome Browswer. It has a more sophisticated but weirder interface than the UCSC Genome Browser. All the data that you will view as tracks is in reality just files
Huijuan Feng, Shining Ma,Chao Ye & Zhixing Feng
 Huijuan Feng, Shining Ma,Chao Ye & Zhixing Feng Background-Author introduction Research interest: Methods for gene mapping of complex traits Inference of population structure from genetic data Genome variation
Huijuan Feng, Shining Ma,Chao Ye & Zhixing Feng Background-Author introduction Research interest: Methods for gene mapping of complex traits Inference of population structure from genetic data Genome variation
Discovering gene regulatory control using ChIP-chip and ChIP-seq. Part 1. An introduction to gene regulatory control, concepts and methodologies
 Discovering gene regulatory control using ChIP-chip and ChIP-seq Part 1 An introduction to gene regulatory control, concepts and methodologies Ian Simpson ian.simpson@.ed.ac.uk http://bit.ly/bio2links
Discovering gene regulatory control using ChIP-chip and ChIP-seq Part 1 An introduction to gene regulatory control, concepts and methodologies Ian Simpson ian.simpson@.ed.ac.uk http://bit.ly/bio2links
Many transcription factors! recognize DNA shape
 Many transcription factors! recognize DN shape Katie Pollard! Gladstone Institutes USF Division of Biostatistics, Institute for Human Genetics, and Institute for omputational Health Sciences ENODE Users
Many transcription factors! recognize DN shape Katie Pollard! Gladstone Institutes USF Division of Biostatistics, Institute for Human Genetics, and Institute for omputational Health Sciences ENODE Users
Integrating Diverse Datasets Improves Developmental Enhancer Prediction
 Integrating Diverse Datasets Improves Developmental Enhancer Prediction Genevieve D. Erwin 1,2, Nir Oksenberg 2,3, Rebecca M. Truty 1, Dennis Kostka 4, Karl K. Murphy 2,3, Nadav Ahituv 2,3, Katherine S.
Integrating Diverse Datasets Improves Developmental Enhancer Prediction Genevieve D. Erwin 1,2, Nir Oksenberg 2,3, Rebecca M. Truty 1, Dennis Kostka 4, Karl K. Murphy 2,3, Nadav Ahituv 2,3, Katherine S.
Predicting tissue specific cis-regulatory modules in the human genome using pairs of co-occurring motifs
 BMC Bioinformatics This Provisional PDF corresponds to the article as it appeared upon acceptance. Fully formatted PDF and full text (HTML) versions will be made available soon. Predicting tissue specific
BMC Bioinformatics This Provisional PDF corresponds to the article as it appeared upon acceptance. Fully formatted PDF and full text (HTML) versions will be made available soon. Predicting tissue specific
Discovery and characterization of chromatin states for systematic annotation of the human genome
 a n a ly s i s Discovery and characterization of chromatin states for systematic annotation of the human genome Jason Ernst 1,2 & Manolis Kellis 1,2 21 Nature America, Inc. All rights reserved. A plethora
a n a ly s i s Discovery and characterization of chromatin states for systematic annotation of the human genome Jason Ernst 1,2 & Manolis Kellis 1,2 21 Nature America, Inc. All rights reserved. A plethora
Introduction to human genomics and genome informatics
 Introduction to human genomics and genome informatics Session 1 Prince of Wales Clinical School Dr Jason Wong ARC Future Fellow Head, Bioinformatics & Integrative Genomics Adult Cancer Program, Lowy Cancer
Introduction to human genomics and genome informatics Session 1 Prince of Wales Clinical School Dr Jason Wong ARC Future Fellow Head, Bioinformatics & Integrative Genomics Adult Cancer Program, Lowy Cancer
A n a ly s i s. Discovery and characterization of chromatin states for systematic annotation of the human genome. Jason Ernst 1,2 & Manolis Kellis 1,2
 A n a ly s i s Discovery and characterization of chromatin states for systematic annotation of the human genome Jason Ernst 1,2 & Manolis Kellis 1,2 A plethora of epigenetic modifications have been described
A n a ly s i s Discovery and characterization of chromatin states for systematic annotation of the human genome Jason Ernst 1,2 & Manolis Kellis 1,2 A plethora of epigenetic modifications have been described
Why DNA Isn t Your Destiny
 Technical Journal Club April 7 2015 Why DNA Isn t Your Destiny Despoina Goniotaki Aguzzi Group GENOME: a genome is an organism s complete set of DNA, including all of its genes. Each genome contains all
Technical Journal Club April 7 2015 Why DNA Isn t Your Destiny Despoina Goniotaki Aguzzi Group GENOME: a genome is an organism s complete set of DNA, including all of its genes. Each genome contains all
Discovering gene regulatory control using ChIP-chip and ChIP-seq. An introduction to gene regulatory control, concepts and methodologies
 Discovering gene regulatory control using ChIP-chip and ChIP-seq An introduction to gene regulatory control, concepts and methodologies Ian Simpson ian.simpson@.ed.ac.uk bit.ly/bio2_2012 The Central Dogma
Discovering gene regulatory control using ChIP-chip and ChIP-seq An introduction to gene regulatory control, concepts and methodologies Ian Simpson ian.simpson@.ed.ac.uk bit.ly/bio2_2012 The Central Dogma
Wednesday, November 22, 17. Exons and Introns
 Exons and Introns Introns and Exons Exons: coded regions of DNA that get transcribed and translated into proteins make up 5% of the genome Introns and Exons Introns: non-coded regions of DNA Must be removed
Exons and Introns Introns and Exons Exons: coded regions of DNA that get transcribed and translated into proteins make up 5% of the genome Introns and Exons Introns: non-coded regions of DNA Must be removed
ECS 234: Genomic Data Integration ECS 234
 : Genomic Data Integration Heterogeneous Data Integration DNA Sequence Microarray Proteomics >gi 12004594 gb AF217406.1 Saccharomyces cerevisiae uridine nucleosidase (URH1) gene, complete cds ATGGAATCTGCTGATTTTTTTACCTCACGAAACTTATTAAAACAGATAATTTCCCTCATCTGCAAGGTTG
: Genomic Data Integration Heterogeneous Data Integration DNA Sequence Microarray Proteomics >gi 12004594 gb AF217406.1 Saccharomyces cerevisiae uridine nucleosidase (URH1) gene, complete cds ATGGAATCTGCTGATTTTTTTACCTCACGAAACTTATTAAAACAGATAATTTCCCTCATCTGCAAGGTTG
Understanding transcriptional regulation by integrative analysis of transcription factor binding data
 Research Understanding transcriptional regulation by integrative analysis of transcription factor binding data Chao Cheng, 1,2 Roger Alexander, 1,2 Renqiang Min, 1,2 Jing Leng, 2 Kevin Y. Yip, 1,2,3 Joel
Research Understanding transcriptional regulation by integrative analysis of transcription factor binding data Chao Cheng, 1,2 Roger Alexander, 1,2 Renqiang Min, 1,2 Jing Leng, 2 Kevin Y. Yip, 1,2,3 Joel
Lecture 21: Epigenetics Nurture or Nature? Chromatin DNA methylation Histone Code Twin study X-chromosome inactivation Environemnt and epigenetics
 Lecture 21: Epigenetics Nurture or Nature? Chromatin DNA methylation Histone Code Twin study X-chromosome inactivation Environemnt and epigenetics Epigenetics represents the science for the studying heritable
Lecture 21: Epigenetics Nurture or Nature? Chromatin DNA methylation Histone Code Twin study X-chromosome inactivation Environemnt and epigenetics Epigenetics represents the science for the studying heritable
A Prac'cal Guide to NCBI BLAST
 A Prac'cal Guide to NCBI BLAST Leonardo Mariño-Ramírez NCBI, NIH Bethesda, USA June 2018 1 NCBI Search Services and Tools Entrez integrated literature and molecular databases Viewers BLink protein similarities
A Prac'cal Guide to NCBI BLAST Leonardo Mariño-Ramírez NCBI, NIH Bethesda, USA June 2018 1 NCBI Search Services and Tools Entrez integrated literature and molecular databases Viewers BLink protein similarities
