Gene Expression Classification: Decision Trees vs. SVMs
|
|
|
- Doris Kennedy
- 5 years ago
- Views:
Transcription
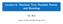
1 Gene Expression Classification: Decision Trees vs. SVMs Xiaojing Yuan and Xiaohui Yuan and Fan Yang and Jing Peng and Bill P. Buckles Electrical Engineering and Computer Science Department Tulane University New Orleans, Louisiana 70118, U.S.A {yuan, yuanx, yangf, jp, Abstract In this article, we compare decision trees (DT) and support vector machines (SVM) in classifying gene expressions. With the explosion of genome research, tremendous amount of data have been made available and a deep insight study becomes demanding. Among various kinds of gene analysis approaches being developed, sequence based gene expression classification shows the importance due to its ability to identify existence of some specific gene pieces. In this article, we focus on two major categories of classification methods, namely decision trees and support vector machines. By comparing various versions of decision tree algorithms, SVMs, and a particular SVM that integrates structural information of the gene sequence, it is shown that the structural information does help in achieving better performance with respect to the classification accuracy. Introduction A gene is a functional unit along the DNA of a chromosome. Nucleotide sequencing and other physical studies have shown that between genes there are sequences of DNA, where functions are mostly unknown. Therefore, classifying gene sequences is fundamental in gene expression analysis. (Mount 2001) The strong sequence similarity often implies functional and structural relationships. In particular, sequence classification algorithms can be applied to problems such as determining whether a DNA sequence is part of a gene-coding or a non-coding region, identifying the introns or exons of a eukaryotic gene sequence, predicting the secondary structure of a protein, and assigning a protein sequence to a specific protein family. Various machine learning techniques have been applied to gene expression classification,(deshpande & Karypis 2002; Kuramochi & Karypis 2001; Lesh, Zaki, & Ogihara 1999; Ohler et al. 1999; Yuan et al. 2002) among which decision tree algorithms and support vector machines are widely recognized as the most effective and efficient approaches dealing with that huge amount of data. (Kuramochi & Karypis 2001; Wang et al. 1999) However, few if any studies have been conducted that compare their performance over such Copyright c 2003, American Association for Artificial Intelligence ( All rights reserved. particular kind of data. In addition, whether the characteristics of the gene data can be employed to improve the classification performance remains an open question. In this paper, we compare the performance of various kinds of decision tree algorithms, namely traditional decision trees (such as C4.5), bagged (bagging) decision trees, and boosted (boosting) decision trees, and SVM classifier with respect to conditions such as varying training set size and different number of possible classes. The experiments show that these methods generate good classifiers and handle large amount of data very well. Although decision tree algorithms may suffer from over fitting, both bagged (bagging) and boosted (boosting) decision trees perform as well as SVMs and even better in some cases. However, due to the characteristics of gene data, a SVM Bank is created to integrate the structural knowledge of a gene into the classifier. The classification is determined by the weighted majority vote of SVMs from the bank and the improvement on performance is clearly shown by the experiments. The rest of this article is organized as follows. A brief review on decision trees and SVMs, as well as ensemble techniques such as bagging and boosting are given in the next section. A further extension of SVMs integrating gene structural characteristics is also described. The experiments are carried out with widely used gene data sets and followed by discussion and conclusion. Background Decision Tree Decision tree learning is a method for approximating discrete-valued target functions, in which the learned function is represented by a decision tree. (Mitchell 1997) Decision trees classify instances by sorting them down the tree from the root to some leaf node, which provides the classification of the instance. Each node in the tree specifies the test of some attribute of the instance, and each branch descending from that node corresponds to one of the possible values for this attribute. An instance is classified by starting at the root node of the tree, testing the attribute specified by this node, then moving down the tree branch corresponding to the value of the attribute in the given example. This process is then repeated for the sub-tree rooted at the new node. Some widely used algorithms include ID3, ASSISTANT, and C4.5. (Quinlan 1986; 1993) Most algorithms that have 92 FLAIRS 2003
2 been developed for learning decision trees are variations on a core algorithm that employs a top-down and greedy search through the space of possible decision trees. The central issue is selecting which attribute to test at each node in the tree. This is determined based on measures, such as information gain, that evaluate how well a single node classifies the training examples. The Bagging algorithm votes classifiers built from different bootstrap samples, which are generated by uniformly sampling N replacement instances from the original training set (of size N). (Breiman 1994; 1996) The new training set has the same size as that of the original training set, but is more randomized in that some instances may not appeared while others appear more than once. For trials t, t = {1, 2,...,T}, bootstrap sample sets D 1,D 2,...,D T are generated and a classifier C i is built by a learning system for each bootstrap sample set D i. The final classifier C is built from C 1,C 2,...,C T by majority vote, so that the result classification is the class that output most often by the sub-classifiers. Boosting is another ensemble technique to improve the performance of a weak learning system. A weak learning system is a system whose prediction accuracy is at least 50It is proven that the error rate of the boosted classifier C on the given examples approaches zero exponentially as number of trials T increases, and the error rate of trials is less than 0.5. (Freund & Schapire 1999) Therefore, a succession of weak classifiers are boosted to a strong classifier, which is usually more accurate than the best weak classifier is on the given training data. Support Vector Machine Support vector machines are introduced to solve two-class pattern recognition problems using the Structural Risk Minimization principle. (Vapnik 1995; 1998; Burges 1998; Cristianini & Shawe-Taylor 2000; Scholkopf, Burges, & Smola 1999) Given a training set in a vector space, SVMs find the best decision hyperplane that separates two classes. The quality of a decision hyperplane is determined by the distance (i.e. margin) between two hyperplanes defined by support vectors. The best decision hyperplane is the one that maximizes this margin. Figure 1 shows these concepts in two-dimensional feature space. By defining the hyperplane in this fashion, SVM is able to generalize to unseen instances quite effectively. SVM extends its applicability on the linearly non-separable data sets by either using soft margin hyperplanes, or by mapping the original data vectors into a higher dimensional space in which the data points are linearly separable. The mapping to higher dimensional spaces is done using appropriate kernels such as Gaussian kernel and polynomial kernel. Integrating Gene Structural Information Gene Representation A gene can be defined as a region of the chromosomal DNA that can be transcribed into a functional RNA during development. DNA is constructed with four nitrogenous bases: adenine, guanine, cytosine, and thymine, which are referred by A, G, C, and T respectively. In turn, the gene data are most commonly arranged in sequence as illustrated below. Figure 1: Classification with SVMs.... GAAGTCCAGTTGCTACAGGCCCTGGAG ACTGACATGCCAAAGTACCATCCGAAGGG AGCCCCGTGCTCGGCAGCCCCGATGCTCT AAACTAGGAAGTGGCAGCATGTCAGAC... From computer science point of view, to analyze gene data, it is necessary to convert from symbolic representation to numerical representation, especially when applying methods such as neural networks and SVMs. Obviously, there are infinite ways of mappings. The question is will simple ways, such as assigning each nucleotide a two-digit number, make classification difference from using a complicated coding strategy? To study the potential differences, we compare the following two coding schemes with Hamming distance. Scheme 1: A = 00, C=01,G=10,T=11 Scheme 2: A = 0001, C = 0010, G = 0100, T = 1000 Sequences: {A, A, G, T, C, C, A, G, T, T, G, C} {A, G, T, A, C, C, A, T, T, C, G, G} The Hamming distance using the first coding scheme is 8, while it is 12 using the second coding scheme. It is clear that the coding scheme affects the value of similarity measurement. Integrating Gene Structural Information into SVM Given a DNA sequence, since there is no a priori information on how each nucleotide relates to each other, it is important to preserve the sequential structure in it. From above discussion, it is obvious that different coding schemes affect the similarity measure, thus may affect the learned classifier. This requires that when translating symbols to numerical representation, no bias should be introduced by these coding schemes. The best choice is to treat these nucleotides is to consider these bases as equally correlated individual. It is suggested that the ideal mapping should be as shown in Figure 2. (Xu & Buckles 2002) However, with any set of values, it is infeasible to guarantee an unbiased assign- FLAIRS
3 ment. To approach this ideal model, we take into account all the possible encoding schemes and build a SVM bank. The four nucleotides, A, C, G, T, formed a base so that a gene sequence can be fully represented by their combinations. Thus, for gene sequence analysis, a maximum of 24 coding combinations are necessary, as shown in Table 1. These 24 coding schemes can be used for any application that can be fully described by four characteristics and their combinations, which could be a very large number. It is noted that even though coding combination may increase exponentially when the underline base increases, there are several ways to simplify the coding schemes. Figure 2: The ideal mapping structure. Keep symbols equally apart. Table 1: Possible coding combinations, assuming number basis is ( ). Nuc. A G C T Nuc. A G C T Using different coding schemes, 24 support vector machines are built from training data, S = {s 1,s 2,...,s 24 }, which can be considered as a SVM Bank designed for gene expression analysis. To predict an unlabelled sequence, each SVM, s i (i = {1, 2,...,24}) is applied and gives its classification result p i. The final judgment is determined by the majority vote on all predictions. In addition, weights, W = {w 1,w 2,...,w 24 }, are assigned to SVMs according to their accuracy during prediction. Initially, an equal weight is assigned to each SVM (Here, w i =1is used in our experiments.) Each weight is dynamically updated with adjustment δ based on the correctness of each prediction. Let the final classification on a unlabeled instance be p. w i = w i { + φ(s i ) δ 1 pi = p and φ(s i )= 1 otherwise For reference purpose, we call such SVM as Gene Structural SVM (GS-SVM). Experiments In this section, the experiments on the classification of two gene sequence data sets, namely ALU gene and splice junctions, are performed and followed by the comparison and discussion. Data Set Description Two widely used gene sequence analysis sets, ALU gene sequences and splice junction sequence, are used as test beds, which are available for download at (ALU 2002; Blake & Merz 1998). (Jurka et al. 1993; Wang et al. 1999) ALU gene is the most abundant single sequence in the entire human genome. The data set we use has 327 positive instances which are downloaded from (ALU 2002; Blake & Merz 1998) and 654 negative instances. These negative instances were generated randomly while keeping the nucleotide frequencies unchanged. (Wang et al. 1999) The sequence length varies from 69bp to 379bp (where bp stands for base pair, a biological metric for gene length.) Splice junctions (SJ) are sections on a DNA sequence at which superfluous DNA is removed during the process of protein creation in higher level organisms. There exist two forms of boundaries: (1) the transition from intron to exon, called acceptor, and (2) the transition from exon to intron, called donor. All the sequences are in length of 60bp, among which 767 sequences contain acceptor, 768 sequences contain donor, and 1655 sequences contain neither of them. In all experiments, training sets are randomly selected from the original data set without replacement according to specified percentage. The remaining data will be used as testing set. Table 2 summarizes the properties of the experimental data sets. Table 2: Properties of the experimental data sets. Set Cases Classes Positive Negative ALU Positive Set Cases Classes Acceptor Donor Neg. SJ Experimental Setup Both bagging and boosting decision trees are developed by extending C4.5 algorithm. (Quinlan 1993) The implementation of SVM for our experiments is developed by S. R. Gunn. (Gunn 1998) Considering possible errors made during data collection stage, a soft margin SVM is used. 94 FLAIRS 2003
4 The kernel we employed is a 6th order polynomial function. For both Bagging and Boosting, the parameter T governing the number of classifiers generated was set at ten. This is based on the results from (Breiman 1994; 1996) that most of the performance improvement from bagging is evident within the first ten trials. Ten complete tests were carried out for each training set. In each classification experiment, Training sets are randomly selected from original data set and the training set size range from 10% to 90% of original data set. The corresponding remaining data sets are used as testing set. All parameters of C4.5 maintained their default values, and the pruned trees were used for decision trees. However, there are several issues regarding the choice of the training set. First, extreme situations, such as all training instances are negative or positive examples, will compromise learning the classifier. Second, margin samples may missed from training set. To minimize the possible misleading caused by the training data, ten experiments were conducted for each experiment using same data sets and parameters. The average of the prediction accuracy is used in the performance comparison. Three sets of experiments were done to systematically compare the classification performance of different ensemble methods (bagging or boosting), different encoding methods (symbolic or genestructural based), and in general, different learning methods (decision trees or SVMs). Results and Discussion Figure 3 illustrates the classification performance with splice junctions sequence. Without considering gene structural information, the accuracy of decision tree algorithms and SVM algorithm are very close, especially when using less than 50% data as training set. In some cases, bagging and boosting are better than SVMs. When increasing training set size, the accuracy of SVMs continues to improve while decision tree based classifiers are less satisfactory for little improvement and they show degradation due to over fitting. Also, it is shown that by incorporating gene structural information, GS-SVM classifiers gain improvement for all population sizes, where it gives the best performance than any of others most of time. Figure 3: Classification performance with splice junctions sequence. Figure 4 illustrates the classification performance with ALU gene sequence data. It is clear that the gene structural information plays an important role in learning the classifier and an average accuracy gain is over 70%. Classification performance for ALU gene se- Figure 4: quence. Table 3 lists the error rate improvement of GS-SVM in terms of Average Error Rate (AER) of decision trees and SVMs and in terms of average error rate of SVMs. The second column lists the classification error rate of GS-SVM. The next two columns are average error rate and the improvement of GS-SVM compared to AER. The last two columns show the error rate of SVM classifier and GS- SVM s performance gain. Table 3: Error rate reduction of GS-SVM. Splice junctions sequence TDR GS-SVM AER Impro. SVM Impro. 10% % % 20% % % 30% % % 40% % % 50% % % 60% % % 70% % % 80% % % 90% % % ALU gene sequence TDR GS-SVM AER Impro. SVM Impro. 10% % % 20% % % 30% % % 40% % % 50% % % 60% % % 70% % % 80% % % 90% % % The performance improvement of GS-SVM relies on the FLAIRS
5 fact that although there are some co-occurrences of nucleotides to form a gene, little knowledge is present. By assuming an equal correlation between nucleotides, given a coding base, 24 identical combinations exist. Obviously, encoding scheme changes the information distribution, i.e. the co-occurrence of basic representative units. In addition, genes have their own features, or we can say, the regularity of both occurrence of nucleotides and spatial relationship determine if it is a gene or not and which gene it is. Encoding nucleotides is essentially mapping the sequence to a different space. This process may increase the discrimination, within one encoding scheme, or among several encoding schemes. Also, which encoding schemes play a more important role varies from gene to gene. Therefore, a weighted majority voting circumvents this by learning the weights along predictions. Conclusions In this article, we study the performance of the classification of gene sequence data with decision trees and SVMs. From our experiments, it is clearly shown that these methods generate good classifiers and handle large amount of data very well. Although decision tree algorithms suffer from over fitting difficulty, both bagging and boosting decision trees perform as well as or close to SVM and even better in some cases. However, due the data pre-processing requirements, i.e. converting symbolic to numeric representation, bias is inevitable. To circumvent this, a weighted voting scheme using SVM Bank is described, which is inspired by the gene structural characteristics. A 24-SVM bank is learned from data sets independently and a voting weight is assigned to each SVM and updated according to the classification performance on each prediction. The promising results with experimental data suggest that incorporating structural information greatly improves the classifier. Acknowledgment We would like to thank Ibrahim Gokcen and Jian Zhang for discussions and insights on preparing this article. We would also like to thank Zujia Xu and Kun Zhang for their help on developing software. References ALU Alu gene data. ftp://ncbi.nlm.nih.gov/ pub/ jmc/ alu/ ALU.327.dna.ref. Blake, C. L., and Merz, C. J Uci: Repository of machine learning databases. Breiman, L Heuristic of instability in model selection. Technical report, Statistics Department, University of California. Breiman, L Bagging predictors. Machine Learning 24: Burges, C. J. C A tutorial on support vector machines for pattern recognition. Data Mining and Knowledge Discovery 2(2): Cristianini, N., and Shawe-Taylor, J An Introduction to Support Vector Machine. Cambridge University Press. Deshpande, M., and Karypis, G Evaluation of techniques for classifying biological sequences. In Proceedings of the 2002 Pacific Asia Conference on Knowledge Discovery and Data Mining, Freund, Y., and Schapire, R. E A short introduction to boosting. Journal of Japaneses Society for Artificial Intelligence 14(5): Gunn, S. R Support vector machines for classification and regression. Technical report, University of Southampton. Jurka, J.; Kaplan, D. J.; Duncan, C. H.; Walichiewicz, J.; Milosavljevic, A.; Murali, G.; and Solus, J. F Identification and characterization of new human medium reiteration frequency repeats. Nucleic Acids Res Kuramochi, M., and Karypis, G Gene classification using expression profiles: A feasibility study. In IEEE International Conference on Data Mining, Lesh, N.; Zaki, M. J.; and Ogihara, M Mining features for sequence classification. In 5th ACM SIGKDD International Conference on Knowledge Discovery and Data Mining (KDD), Mitchell, T. M Machine Learning. McGraw-Hill Companies, Inc. Mount, D. W Bioinformatics: Sequence and Genome Analysis. Cold Spring Harbor Laboratory Press. Ohler, U.; Harbeck, S.; Niemann, H.; Noth, E.; and Reese, M. G Interpolated markov chains for eukaryotic promoter recognition. Bioinformatics 15(5). Quinlan, J. R Induction of decision trees. Maching Learning 1(1): Quinlan, J. R C4.5.: Programs for Machine Learning. San Mateo, CA: Morgan Kaufmann. Scholkopf, C.; Burges, J.; and Smola, A Advances in Kernel methods: Support Vector Learning. MIT Press. Vapnik, V The Nature of Statistical Learning Theory. New York: Springer Verlag. Vapnik, V Statistical Learning Theory. New York: Wiley. Wang, J. T. L.; Rozen, S.; Shapiro, B. A.; Shasha, D.; Wang, Z.; and Yin, M New techniques for dna sequence classification. Journal of Computational Biology 6(2): Xu, Z., and Buckles, B Dna sequence classification by using support vector machine. EECS, Tulane University. Yuan, X.; Buckles, B. P.; Yuan, Z.; and Zhang, J Mining negative association rules. In Proceedings of the 7th IEEE Symposium on Computers and Communications. 96 FLAIRS 2003
Identifying Splice Sites Of Messenger RNA Using Support Vector Machines
 Identifying Splice Sites Of Messenger RNA Using Support Vector Machines Paige Diamond, Zachary Elkins, Kayla Huff, Lauren Naylor, Sarah Schoeberle, Shannon White, Timothy Urness, Matthew Zwier Drake University
Identifying Splice Sites Of Messenger RNA Using Support Vector Machines Paige Diamond, Zachary Elkins, Kayla Huff, Lauren Naylor, Sarah Schoeberle, Shannon White, Timothy Urness, Matthew Zwier Drake University
Detecting and Pruning Introns for Faster Decision Tree Evolution
 Detecting and Pruning Introns for Faster Decision Tree Evolution Jeroen Eggermont and Joost N. Kok and Walter A. Kosters Leiden Institute of Advanced Computer Science Universiteit Leiden P.O. Box 9512,
Detecting and Pruning Introns for Faster Decision Tree Evolution Jeroen Eggermont and Joost N. Kok and Walter A. Kosters Leiden Institute of Advanced Computer Science Universiteit Leiden P.O. Box 9512,
Application of Decision Trees in Mining High-Value Credit Card Customers
 Application of Decision Trees in Mining High-Value Credit Card Customers Jian Wang Bo Yuan Wenhuang Liu Graduate School at Shenzhen, Tsinghua University, Shenzhen 8, P.R. China E-mail: gregret24@gmail.com,
Application of Decision Trees in Mining High-Value Credit Card Customers Jian Wang Bo Yuan Wenhuang Liu Graduate School at Shenzhen, Tsinghua University, Shenzhen 8, P.R. China E-mail: gregret24@gmail.com,
Tree Depth in a Forest
 Tree Depth in a Forest Mark Segal Center for Bioinformatics & Molecular Biostatistics Division of Bioinformatics Department of Epidemiology and Biostatistics UCSF NUS / IMS Workshop on Classification and
Tree Depth in a Forest Mark Segal Center for Bioinformatics & Molecular Biostatistics Division of Bioinformatics Department of Epidemiology and Biostatistics UCSF NUS / IMS Workshop on Classification and
Study on the Application of Data Mining in Bioinformatics. Mingyang Yuan
 International Conference on Mechatronics Engineering and Information Technology (ICMEIT 2016) Study on the Application of Mining in Bioinformatics Mingyang Yuan School of Science and Liberal Arts, New
International Conference on Mechatronics Engineering and Information Technology (ICMEIT 2016) Study on the Application of Mining in Bioinformatics Mingyang Yuan School of Science and Liberal Arts, New
BIOINFORMATICS THE MACHINE LEARNING APPROACH
 88 Proceedings of the 4 th International Conference on Informatics and Information Technology BIOINFORMATICS THE MACHINE LEARNING APPROACH A. Madevska-Bogdanova Inst, Informatics, Fac. Natural Sc. and
88 Proceedings of the 4 th International Conference on Informatics and Information Technology BIOINFORMATICS THE MACHINE LEARNING APPROACH A. Madevska-Bogdanova Inst, Informatics, Fac. Natural Sc. and
Gene Identification in silico
 Gene Identification in silico Nita Parekh, IIIT Hyderabad Presented at National Seminar on Bioinformatics and Functional Genomics, at Bioinformatics centre, Pondicherry University, Feb 15 17, 2006. Introduction
Gene Identification in silico Nita Parekh, IIIT Hyderabad Presented at National Seminar on Bioinformatics and Functional Genomics, at Bioinformatics centre, Pondicherry University, Feb 15 17, 2006. Introduction
Classification of DNA Sequences Using Convolutional Neural Network Approach
 UTM Computing Proceedings Innovations in Computing Technology and Applications Volume 2 Year: 2017 ISBN: 978-967-0194-95-0 1 Classification of DNA Sequences Using Convolutional Neural Network Approach
UTM Computing Proceedings Innovations in Computing Technology and Applications Volume 2 Year: 2017 ISBN: 978-967-0194-95-0 1 Classification of DNA Sequences Using Convolutional Neural Network Approach
DNAFSMiner: A Web-Based Software Toolbox to Recognize Two Types of Functional Sites in DNA Sequences
 DNAFSMiner: A Web-Based Software Toolbox to Recognize Two Types of Functional Sites in DNA Sequences Huiqing Liu Hao Han Jinyan Li Limsoon Wong Institute for Infocomm Research, 21 Heng Mui Keng Terrace,
DNAFSMiner: A Web-Based Software Toolbox to Recognize Two Types of Functional Sites in DNA Sequences Huiqing Liu Hao Han Jinyan Li Limsoon Wong Institute for Infocomm Research, 21 Heng Mui Keng Terrace,
Introduction to Bioinformatics
 Introduction to Bioinformatics If the 19 th century was the century of chemistry and 20 th century was the century of physic, the 21 st century promises to be the century of biology...professor Dr. Satoru
Introduction to Bioinformatics If the 19 th century was the century of chemistry and 20 th century was the century of physic, the 21 st century promises to be the century of biology...professor Dr. Satoru
Proposal for ISyE6416 Project
 Profit-based classification in customer churn prediction: a case study in banking industry 1 Proposal for ISyE6416 Project Profit-based classification in customer churn prediction: a case study in banking
Profit-based classification in customer churn prediction: a case study in banking industry 1 Proposal for ISyE6416 Project Profit-based classification in customer churn prediction: a case study in banking
Customer Relationship Management in marketing programs: A machine learning approach for decision. Fernanda Alcantara
 Customer Relationship Management in marketing programs: A machine learning approach for decision Fernanda Alcantara F.Alcantara@cs.ucl.ac.uk CRM Goal Support the decision taking Personalize the best individual
Customer Relationship Management in marketing programs: A machine learning approach for decision Fernanda Alcantara F.Alcantara@cs.ucl.ac.uk CRM Goal Support the decision taking Personalize the best individual
ARTIFICIAL IMMUNE SYSTEM CLASSIFICATION OF MULTIPLE- CLASS PROBLEMS
 1 ARTIFICIAL IMMUNE SYSTEM CLASSIFICATION OF MULTIPLE- CLASS PROBLEMS DONALD E. GOODMAN, JR. Mississippi State University Department of Psychology Mississippi State, Mississippi LOIS C. BOGGESS Mississippi
1 ARTIFICIAL IMMUNE SYSTEM CLASSIFICATION OF MULTIPLE- CLASS PROBLEMS DONALD E. GOODMAN, JR. Mississippi State University Department of Psychology Mississippi State, Mississippi LOIS C. BOGGESS Mississippi
A Brief History. Bootstrapping. Bagging. Boosting (Schapire 1989) Adaboost (Schapire 1995)
 A Brief History Bootstrapping Bagging Boosting (Schapire 1989) Adaboost (Schapire 1995) What s So Good About Adaboost Improves classification accuracy Can be used with many different classifiers Commonly
A Brief History Bootstrapping Bagging Boosting (Schapire 1989) Adaboost (Schapire 1995) What s So Good About Adaboost Improves classification accuracy Can be used with many different classifiers Commonly
Predictive Analytics Using Support Vector Machine
 International Journal for Modern Trends in Science and Technology Volume: 03, Special Issue No: 02, March 2017 ISSN: 2455-3778 http://www.ijmtst.com Predictive Analytics Using Support Vector Machine Ch.Sai
International Journal for Modern Trends in Science and Technology Volume: 03, Special Issue No: 02, March 2017 ISSN: 2455-3778 http://www.ijmtst.com Predictive Analytics Using Support Vector Machine Ch.Sai
Predicting Toxicity of Food-Related Compounds Using Fuzzy Decision Trees
 Predicting Toxicity of Food-Related Compounds Using Fuzzy Decision Trees Daishi Yajima, Takenao Ohkawa, Kouhei Muroi, and Hiromasa Imaishi Abstract Clarifying the interaction between cytochrome P450 (P450)
Predicting Toxicity of Food-Related Compounds Using Fuzzy Decision Trees Daishi Yajima, Takenao Ohkawa, Kouhei Muroi, and Hiromasa Imaishi Abstract Clarifying the interaction between cytochrome P450 (P450)
2 Maria Carolina Monard and Gustavo E. A. P. A. Batista
 Graphical Methods for Classifier Performance Evaluation Maria Carolina Monard and Gustavo E. A. P. A. Batista University of São Paulo USP Institute of Mathematics and Computer Science ICMC Department of
Graphical Methods for Classifier Performance Evaluation Maria Carolina Monard and Gustavo E. A. P. A. Batista University of São Paulo USP Institute of Mathematics and Computer Science ICMC Department of
Course Information. Introduction to Algorithms in Computational Biology Lecture 1. Relations to Some Other Courses
 Course Information Introduction to Algorithms in Computational Biology Lecture 1 Meetings: Lecture, by Dan Geiger: Mondays 16:30 18:30, Taub 4. Tutorial, by Ydo Wexler: Tuesdays 10:30 11:30, Taub 2. Grade:
Course Information Introduction to Algorithms in Computational Biology Lecture 1 Meetings: Lecture, by Dan Geiger: Mondays 16:30 18:30, Taub 4. Tutorial, by Ydo Wexler: Tuesdays 10:30 11:30, Taub 2. Grade:
Introduction to Algorithms in Computational Biology Lecture 1
 Introduction to Algorithms in Computational Biology Lecture 1 Background Readings: The first three chapters (pages 1-31) in Genetics in Medicine, Nussbaum et al., 2001. This class has been edited from
Introduction to Algorithms in Computational Biology Lecture 1 Background Readings: The first three chapters (pages 1-31) in Genetics in Medicine, Nussbaum et al., 2001. This class has been edited from
Comparison of Different Independent Component Analysis Algorithms for Sales Forecasting
 International Journal of Humanities Management Sciences IJHMS Volume 2, Issue 1 2014 ISSN 2320 4044 Online Comparison of Different Independent Component Analysis Algorithms for Sales Forecasting Wensheng
International Journal of Humanities Management Sciences IJHMS Volume 2, Issue 1 2014 ISSN 2320 4044 Online Comparison of Different Independent Component Analysis Algorithms for Sales Forecasting Wensheng
Today. Last time. Lecture 5: Discrimination (cont) Jane Fridlyand. Oct 13, 2005
 Biological question Experimental design Microarray experiment Failed Lecture : Discrimination (cont) Quality Measurement Image analysis Preprocessing Jane Fridlyand Pass Normalization Sample/Condition
Biological question Experimental design Microarray experiment Failed Lecture : Discrimination (cont) Quality Measurement Image analysis Preprocessing Jane Fridlyand Pass Normalization Sample/Condition
CONSTRUCTIVE INDUCTION IN BIO-SEQUENCES
 Key words bioinformatics, DNA, classification Rafał BIEDRZYCKI, Jarosław ARABAS + CONSTRUCTIVE INDUCTION IN BIO-SEQUENCES This paper is devoted to analysis of DNA sequences. The aim of the analysis is
Key words bioinformatics, DNA, classification Rafał BIEDRZYCKI, Jarosław ARABAS + CONSTRUCTIVE INDUCTION IN BIO-SEQUENCES This paper is devoted to analysis of DNA sequences. The aim of the analysis is
Representation in Supervised Machine Learning Application to Biological Problems
 Representation in Supervised Machine Learning Application to Biological Problems Frank Lab Howard Hughes Medical Institute & Columbia University 2010 Robert Howard Langlois Hughes Medical Institute What
Representation in Supervised Machine Learning Application to Biological Problems Frank Lab Howard Hughes Medical Institute & Columbia University 2010 Robert Howard Langlois Hughes Medical Institute What
Using Decision Tree to predict repeat customers
 Using Decision Tree to predict repeat customers Jia En Nicholette Li Jing Rong Lim Abstract We focus on using feature engineering and decision trees to perform classification and feature selection on the
Using Decision Tree to predict repeat customers Jia En Nicholette Li Jing Rong Lim Abstract We focus on using feature engineering and decision trees to perform classification and feature selection on the
E-Commerce Sales Prediction Using Listing Keywords
 E-Commerce Sales Prediction Using Listing Keywords Stephanie Chen (asksteph@stanford.edu) 1 Introduction Small online retailers usually set themselves apart from brick and mortar stores, traditional brand
E-Commerce Sales Prediction Using Listing Keywords Stephanie Chen (asksteph@stanford.edu) 1 Introduction Small online retailers usually set themselves apart from brick and mortar stores, traditional brand
Water Treatment Plant Decision Making Using Rough Multiple Classifier Systems
 Available online at www.sciencedirect.com Procedia Environmental Sciences 11 (2011) 1419 1423 Water Treatment Plant Decision Making Using Rough Multiple Classifier Systems Lin Feng 1, 2 1 College of Computer
Available online at www.sciencedirect.com Procedia Environmental Sciences 11 (2011) 1419 1423 Water Treatment Plant Decision Making Using Rough Multiple Classifier Systems Lin Feng 1, 2 1 College of Computer
DNA is normally found in pairs, held together by hydrogen bonds between the bases
 Bioinformatics Biology Review The genetic code is stored in DNA Deoxyribonucleic acid. DNA molecules are chains of four nucleotide bases Guanine, Thymine, Cytosine, Adenine DNA is normally found in pairs,
Bioinformatics Biology Review The genetic code is stored in DNA Deoxyribonucleic acid. DNA molecules are chains of four nucleotide bases Guanine, Thymine, Cytosine, Adenine DNA is normally found in pairs,
Lecture 6: Decision Tree, Random Forest, and Boosting
 Lecture 6: Decision Tree, Random Forest, and Boosting Tuo Zhao Schools of ISyE and CSE, Georgia Tech CS764/ISYE/CSE 674: Machine Learning/Computational Data Analysis Tree? Tuo Zhao Lecture 6: Decision
Lecture 6: Decision Tree, Random Forest, and Boosting Tuo Zhao Schools of ISyE and CSE, Georgia Tech CS764/ISYE/CSE 674: Machine Learning/Computational Data Analysis Tree? Tuo Zhao Lecture 6: Decision
Introduction to Microarray Data Analysis and Gene Networks. Alvis Brazma European Bioinformatics Institute
 Introduction to Microarray Data Analysis and Gene Networks Alvis Brazma European Bioinformatics Institute A brief outline of this course What is gene expression, why it s important Microarrays and how
Introduction to Microarray Data Analysis and Gene Networks Alvis Brazma European Bioinformatics Institute A brief outline of this course What is gene expression, why it s important Microarrays and how
Original article: IDENTIFYING DNA SPLICE SITES USING PATTERNS STATISTICAL PROPERTIES AND FUZZY NEURAL NETWORKS
 Original article: IDENTIFYING DNA SPLICE SITES USING PATTERNS STATISTICAL PROPERTIES AND FUZZY NEURAL NETWORKS Essam Al-Daoud Computer Science Department, Faculty of Science and Information Technology,
Original article: IDENTIFYING DNA SPLICE SITES USING PATTERNS STATISTICAL PROPERTIES AND FUZZY NEURAL NETWORKS Essam Al-Daoud Computer Science Department, Faculty of Science and Information Technology,
Telomerase Gene Prediction Using Support Vector Machines
 Telomerase Gene Prediction Using Support Vector Machines David Luper 1 and Spandana Makeneni 2 1 Computer Science Department, UGA, Athens, GA, USA 2 Institute of Bioinformatics, Complex Carbohydrate Research
Telomerase Gene Prediction Using Support Vector Machines David Luper 1 and Spandana Makeneni 2 1 Computer Science Department, UGA, Athens, GA, USA 2 Institute of Bioinformatics, Complex Carbohydrate Research
Computational gene finding
 Computational gene finding Devika Subramanian Comp 470 Outline (3 lectures) Lec 1 Lec 2 Lec 3 The biological context Markov models and Hidden Markov models Ab-initio methods for gene finding Comparative
Computational gene finding Devika Subramanian Comp 470 Outline (3 lectures) Lec 1 Lec 2 Lec 3 The biological context Markov models and Hidden Markov models Ab-initio methods for gene finding Comparative
Smart water in urban distribution networks: limited financial capacity and Big Data analytics
 Urban Water II 63 Smart water in urban distribution networks: limited financial capacity and Big Data analytics A. Candelieri 1 & F. Archetti 1,2 1 Consorzio Milano Ricerche, Italy 2 Department of Computer
Urban Water II 63 Smart water in urban distribution networks: limited financial capacity and Big Data analytics A. Candelieri 1 & F. Archetti 1,2 1 Consorzio Milano Ricerche, Italy 2 Department of Computer
A Two-stage Architecture for Stock Price Forecasting by Integrating Self-Organizing Map and Support Vector Regression
 Georgia State University ScholarWorks @ Georgia State University Computer Information Systems Faculty Publications Department of Computer Information Systems 2008 A Two-stage Architecture for Stock Price
Georgia State University ScholarWorks @ Georgia State University Computer Information Systems Faculty Publications Department of Computer Information Systems 2008 A Two-stage Architecture for Stock Price
Computational gene finding
 Computational gene finding Devika Subramanian Comp 470 Outline (3 lectures) Lec 1 Lec 2 Lec 3 The biological context Markov models and Hidden Markov models Ab-initio methods for gene finding Comparative
Computational gene finding Devika Subramanian Comp 470 Outline (3 lectures) Lec 1 Lec 2 Lec 3 The biological context Markov models and Hidden Markov models Ab-initio methods for gene finding Comparative
From Ordinal Ranking to Binary Classification
 From Ordinal Ranking to Binary Classification Hsuan-Tien Lin Learning Systems Group, California Institute of Technology Talk at Caltech CS/IST Lunch Bunch March 4, 2008 Benefited from joint work with Dr.
From Ordinal Ranking to Binary Classification Hsuan-Tien Lin Learning Systems Group, California Institute of Technology Talk at Caltech CS/IST Lunch Bunch March 4, 2008 Benefited from joint work with Dr.
Data Mining for Biological Data Analysis
 Data Mining for Biological Data Analysis Data Mining and Text Mining (UIC 583 @ Politecnico di Milano) References Data Mining Course by Gregory-Platesky Shapiro available at www.kdnuggets.com Jiawei Han
Data Mining for Biological Data Analysis Data Mining and Text Mining (UIC 583 @ Politecnico di Milano) References Data Mining Course by Gregory-Platesky Shapiro available at www.kdnuggets.com Jiawei Han
A Direct Marketing Framework to Facilitate Data Mining Usage for Marketers: A Case Study in Supermarket Promotions Strategy
 A Direct Marketing Framework to Facilitate Data Mining Usage for Marketers: A Case Study in Supermarket Promotions Strategy Adel Flici (0630951) Business School, Brunel University, London, UK Abstract
A Direct Marketing Framework to Facilitate Data Mining Usage for Marketers: A Case Study in Supermarket Promotions Strategy Adel Flici (0630951) Business School, Brunel University, London, UK Abstract
Automatic Facial Expression Recognition
 Automatic Facial Expression Recognition Huchuan Lu, Pei Wu, Hui Lin, Deli Yang School of Electronic and Information Engineering, Dalian University of Technology Dalian, Liaoning Province, China lhchuan@dlut.edu.cn
Automatic Facial Expression Recognition Huchuan Lu, Pei Wu, Hui Lin, Deli Yang School of Electronic and Information Engineering, Dalian University of Technology Dalian, Liaoning Province, China lhchuan@dlut.edu.cn
Ranking Potential Customers based on GroupEnsemble method
 Ranking Potential Customers based on GroupEnsemble method The ExceedTech Team South China University Of Technology 1. Background understanding Both of the products have been on the market for many years,
Ranking Potential Customers based on GroupEnsemble method The ExceedTech Team South China University Of Technology 1. Background understanding Both of the products have been on the market for many years,
Understanding the Drivers of Negative Electricity Price Using Decision Tree
 2017 Ninth Annual IEEE Green Technologies Conference Understanding the Drivers of Negative Electricity Price Using Decision Tree José Carlos Reston Filho Ashutosh Tiwari, SMIEEE Chesta Dwivedi IDAAM Educação
2017 Ninth Annual IEEE Green Technologies Conference Understanding the Drivers of Negative Electricity Price Using Decision Tree José Carlos Reston Filho Ashutosh Tiwari, SMIEEE Chesta Dwivedi IDAAM Educação
Bio-inspired Computing for Network Modelling
 ISSN (online): 2289-7984 Vol. 6, No.. Pages -, 25 Bio-inspired Computing for Network Modelling N. Rajaee *,a, A. A. S. A Hussaini b, A. Zulkharnain c and S. M. W Masra d Faculty of Engineering, Universiti
ISSN (online): 2289-7984 Vol. 6, No.. Pages -, 25 Bio-inspired Computing for Network Modelling N. Rajaee *,a, A. A. S. A Hussaini b, A. Zulkharnain c and S. M. W Masra d Faculty of Engineering, Universiti
ABSTRACT COMPUTER EVOLUTION OF GENE CIRCUITS FOR CELL- EMBEDDED COMPUTATION, BIOTECHNOLOGY AND AS A MODEL FOR EVOLUTIONARY COMPUTATION
 ABSTRACT COMPUTER EVOLUTION OF GENE CIRCUITS FOR CELL- EMBEDDED COMPUTATION, BIOTECHNOLOGY AND AS A MODEL FOR EVOLUTIONARY COMPUTATION by Tommaso F. Bersano-Begey Chair: John H. Holland This dissertation
ABSTRACT COMPUTER EVOLUTION OF GENE CIRCUITS FOR CELL- EMBEDDED COMPUTATION, BIOTECHNOLOGY AND AS A MODEL FOR EVOLUTIONARY COMPUTATION by Tommaso F. Bersano-Begey Chair: John H. Holland This dissertation
Annotating the Genome (H)
 Annotating the Genome (H) Annotation principles (H1) What is annotation? In general: annotation = explanatory note* What could be useful as an annotation of a DNA sequence? an amino acid sequence? What
Annotating the Genome (H) Annotation principles (H1) What is annotation? In general: annotation = explanatory note* What could be useful as an annotation of a DNA sequence? an amino acid sequence? What
Lecture 10. Ab initio gene finding
 Lecture 10 Ab initio gene finding Uses of probabilistic sequence Segmentation models/hmms Multiple alignment using profile HMMs Prediction of sequence function (gene family models) ** Gene finding ** Review
Lecture 10 Ab initio gene finding Uses of probabilistic sequence Segmentation models/hmms Multiple alignment using profile HMMs Prediction of sequence function (gene family models) ** Gene finding ** Review
Basics of RNA-Seq. (With a Focus on Application to Single Cell RNA-Seq) Michael Kelly, PhD Team Lead, NCI Single Cell Analysis Facility
 2018 ABRF Meeting Satellite Workshop 4 Bridging the Gap: Isolation to Translation (Single Cell RNA-Seq) Sunday, April 22 Basics of RNA-Seq (With a Focus on Application to Single Cell RNA-Seq) Michael Kelly,
2018 ABRF Meeting Satellite Workshop 4 Bridging the Gap: Isolation to Translation (Single Cell RNA-Seq) Sunday, April 22 Basics of RNA-Seq (With a Focus on Application to Single Cell RNA-Seq) Michael Kelly,
Algorithms in Bioinformatics
 Algorithms in Bioinformatics Sami Khuri Department of Computer Science San José State University San José, California, USA khuri@cs.sjsu.edu www.cs.sjsu.edu/faculty/khuri Outline Central Dogma of Molecular
Algorithms in Bioinformatics Sami Khuri Department of Computer Science San José State University San José, California, USA khuri@cs.sjsu.edu www.cs.sjsu.edu/faculty/khuri Outline Central Dogma of Molecular
EVOLUTIONARY ALGORITHMS AT CHOICE: FROM GA TO GP EVOLŪCIJAS ALGORITMI PĒC IZVĒLES: NO GA UZ GP
 ISSN 1691-5402 ISBN 978-9984-44-028-6 Environment. Technology. Resources Proceedings of the 7 th International Scientific and Practical Conference. Volume I1 Rēzeknes Augstskola, Rēzekne, RA Izdevniecība,
ISSN 1691-5402 ISBN 978-9984-44-028-6 Environment. Technology. Resources Proceedings of the 7 th International Scientific and Practical Conference. Volume I1 Rēzeknes Augstskola, Rēzekne, RA Izdevniecība,
PREDICTION OF CONCRETE MIX COMPRESSIVE STRENGTH USING STATISTICAL LEARNING MODELS
 Journal of Engineering Science and Technology Vol. 13, No. 7 (2018) 1916-1925 School of Engineering, Taylor s University PREDICTION OF CONCRETE MIX COMPRESSIVE STRENGTH USING STATISTICAL LEARNING MODELS
Journal of Engineering Science and Technology Vol. 13, No. 7 (2018) 1916-1925 School of Engineering, Taylor s University PREDICTION OF CONCRETE MIX COMPRESSIVE STRENGTH USING STATISTICAL LEARNING MODELS
What is Bioinformatics? Bioinformatics is the application of computational techniques to the discovery of knowledge from biological databases.
 What is Bioinformatics? Bioinformatics is the application of computational techniques to the discovery of knowledge from biological databases. Bioinformatics is the marriage of molecular biology with computer
What is Bioinformatics? Bioinformatics is the application of computational techniques to the discovery of knowledge from biological databases. Bioinformatics is the marriage of molecular biology with computer
An Efficient and Effective Immune Based Classifier
 Journal of Computer Science 7 (2): 148-153, 2011 ISSN 1549-3636 2011 Science Publications An Efficient and Effective Immune Based Classifier Shahram Golzari, Shyamala Doraisamy, Md Nasir Sulaiman and Nur
Journal of Computer Science 7 (2): 148-153, 2011 ISSN 1549-3636 2011 Science Publications An Efficient and Effective Immune Based Classifier Shahram Golzari, Shyamala Doraisamy, Md Nasir Sulaiman and Nur
BUSINESS DATA MINING (IDS 572) Please include the names of all team-members in your write up and in the name of the file.
 BUSINESS DATA MINING (IDS 572) HOMEWORK 4 DUE DATE: TUESDAY, APRIL 10 AT 3:20 PM Please provide succinct answers to the questions below. You should submit an electronic pdf or word file in blackboard.
BUSINESS DATA MINING (IDS 572) HOMEWORK 4 DUE DATE: TUESDAY, APRIL 10 AT 3:20 PM Please provide succinct answers to the questions below. You should submit an electronic pdf or word file in blackboard.
Lecture 2: Central Dogma of Molecular Biology & Intro to Programming
 Lecture 2: Central Dogma of Molecular Biology & Intro to Programming Central Dogma of Molecular Biology Proteins: workhorse molecules of biological systems Proteins are synthesized from the genetic blueprints
Lecture 2: Central Dogma of Molecular Biology & Intro to Programming Central Dogma of Molecular Biology Proteins: workhorse molecules of biological systems Proteins are synthesized from the genetic blueprints
Introduction to Cellular Biology and Bioinformatics. Farzaneh Salari
 Introduction to Cellular Biology and Bioinformatics Farzaneh Salari Outline Bioinformatics Cellular Biology A Bioinformatics Problem What is bioinformatics? Computer Science Statistics Bioinformatics Mathematics...
Introduction to Cellular Biology and Bioinformatics Farzaneh Salari Outline Bioinformatics Cellular Biology A Bioinformatics Problem What is bioinformatics? Computer Science Statistics Bioinformatics Mathematics...
Homework 4. Due in class, Wednesday, November 10, 2004
 1 GCB 535 / CIS 535 Fall 2004 Homework 4 Due in class, Wednesday, November 10, 2004 Comparative genomics 1. (6 pts) In Loots s paper (http://www.seas.upenn.edu/~cis535/lab/sciences-loots.pdf), the authors
1 GCB 535 / CIS 535 Fall 2004 Homework 4 Due in class, Wednesday, November 10, 2004 Comparative genomics 1. (6 pts) In Loots s paper (http://www.seas.upenn.edu/~cis535/lab/sciences-loots.pdf), the authors
Automated Empirical Selection of Rule Induction Methods Based on Recursive Iteration of Resampling Methods
 Automated Empirical Selection of Rule Induction Methods Based on Recursive Iteration of Resampling Methods Shusaku Tsumoto, Shoji Hirano, and Hidenao Abe Department of Medical Informatics, Faculty of Medicine,
Automated Empirical Selection of Rule Induction Methods Based on Recursive Iteration of Resampling Methods Shusaku Tsumoto, Shoji Hirano, and Hidenao Abe Department of Medical Informatics, Faculty of Medicine,
Methods and tools for exploring functional genomics data
 Methods and tools for exploring functional genomics data William Stafford Noble Department of Genome Sciences Department of Computer Science and Engineering University of Washington Outline Searching for
Methods and tools for exploring functional genomics data William Stafford Noble Department of Genome Sciences Department of Computer Science and Engineering University of Washington Outline Searching for
Copyr i g ht 2012, SAS Ins titut e Inc. All rights res er ve d. ENTERPRISE MINER: ANALYTICAL MODEL DEVELOPMENT
 ENTERPRISE MINER: ANALYTICAL MODEL DEVELOPMENT ANALYTICAL MODEL DEVELOPMENT AGENDA Enterprise Miner: Analytical Model Development The session looks at: - Supervised and Unsupervised Modelling - Classification
ENTERPRISE MINER: ANALYTICAL MODEL DEVELOPMENT ANALYTICAL MODEL DEVELOPMENT AGENDA Enterprise Miner: Analytical Model Development The session looks at: - Supervised and Unsupervised Modelling - Classification
An Analytical Upper Bound on the Minimum Number of. Recombinations in the History of SNP Sequences in Populations
 An Analytical Upper Bound on the Minimum Number of Recombinations in the History of SNP Sequences in Populations Yufeng Wu Department of Computer Science and Engineering University of Connecticut Storrs,
An Analytical Upper Bound on the Minimum Number of Recombinations in the History of SNP Sequences in Populations Yufeng Wu Department of Computer Science and Engineering University of Connecticut Storrs,
Feature Selection of Gene Expression Data for Cancer Classification: A Review
 Available online at www.sciencedirect.com ScienceDirect Procedia Computer Science 50 (2015 ) 52 57 2nd International Symposium on Big Data and Cloud Computing (ISBCC 15) Feature Selection of Gene Expression
Available online at www.sciencedirect.com ScienceDirect Procedia Computer Science 50 (2015 ) 52 57 2nd International Symposium on Big Data and Cloud Computing (ISBCC 15) Feature Selection of Gene Expression
Feature Selection Algorithms in Classification Problems: An Experimental Evaluation
 Feature Selection Algorithms in Classification Problems: An Experimental Evaluation MICHAEL DOUMPOS, ATHINA SALAPPA Department of Production Engineering and Management Technical University of Crete University
Feature Selection Algorithms in Classification Problems: An Experimental Evaluation MICHAEL DOUMPOS, ATHINA SALAPPA Department of Production Engineering and Management Technical University of Crete University
Prediction of Success or Failure of Software Projects based on Reusability Metrics using Support Vector Machine
 Prediction of Success or Failure of Software Projects based on Reusability Metrics using Support Vector Machine R. Sathya Assistant professor, Department of Computer Science & Engineering Annamalai University
Prediction of Success or Failure of Software Projects based on Reusability Metrics using Support Vector Machine R. Sathya Assistant professor, Department of Computer Science & Engineering Annamalai University
A Forecasting Methodology Using Support Vector Regression and Dynamic Feature Selection
 Journal of Information & Knowledge Management, Vol. 5, No. 4 (2006) 329 335 c ikms & World Scientific Publishing Co. A Forecasting Methodology Using Support Vector Regression and Dynamic Feature Selection
Journal of Information & Knowledge Management, Vol. 5, No. 4 (2006) 329 335 c ikms & World Scientific Publishing Co. A Forecasting Methodology Using Support Vector Regression and Dynamic Feature Selection
PDGA: the Primal-Dual Genetic Algorithm
 P: the Primal-Dual Genetic Algorithm Shengxiang Yang Department of Computer Science University of Leicester University Road, Leicester LE1 7RH, UK Email: syang@mcsleacuk Abstract Genetic algorithms (GAs)
P: the Primal-Dual Genetic Algorithm Shengxiang Yang Department of Computer Science University of Leicester University Road, Leicester LE1 7RH, UK Email: syang@mcsleacuk Abstract Genetic algorithms (GAs)
Finding Regularity in Protein Secondary Structures using a Cluster-based Genetic Algorithm
 Finding Regularity in Protein Secondary Structures using a Cluster-based Genetic Algorithm Yen-Wei Chu 1,3, Chuen-Tsai Sun 3, Chung-Yuan Huang 2,3 1) Department of Information Management 2) Department
Finding Regularity in Protein Secondary Structures using a Cluster-based Genetic Algorithm Yen-Wei Chu 1,3, Chuen-Tsai Sun 3, Chung-Yuan Huang 2,3 1) Department of Information Management 2) Department
Decision Tree Learning. Richard McAllister. Outline. Overview. Tree Construction. Case Study: Determinants of House Price. February 4, / 31
 1 / 31 Decision Decision February 4, 2008 2 / 31 Decision 1 2 3 3 / 31 Decision Decision Widely Used Used for approximating discrete-valued functions Robust to noisy data Capable of learning disjunctive
1 / 31 Decision Decision February 4, 2008 2 / 31 Decision 1 2 3 3 / 31 Decision Decision Widely Used Used for approximating discrete-valued functions Robust to noisy data Capable of learning disjunctive
Classification Model for Intent Mining in Personal Website Based on Support Vector Machine
 , pp.145-152 http://dx.doi.org/10.14257/ijdta.2016.9.2.16 Classification Model for Intent Mining in Personal Website Based on Support Vector Machine Shuang Zhang, Nianbin Wang School of Computer Science
, pp.145-152 http://dx.doi.org/10.14257/ijdta.2016.9.2.16 Classification Model for Intent Mining in Personal Website Based on Support Vector Machine Shuang Zhang, Nianbin Wang School of Computer Science
A Novel Splice Site Prediction Method using Support Vector Machine
 Journal of Computational Information Systems 9: 20 (2013) 8053 8060 Available at http://www.jofcis.com A Novel Splice Site Prediction Method using Support Vector Machine Dan WEI 1,2, Huiling ZHANG 2, Yanjie
Journal of Computational Information Systems 9: 20 (2013) 8053 8060 Available at http://www.jofcis.com A Novel Splice Site Prediction Method using Support Vector Machine Dan WEI 1,2, Huiling ZHANG 2, Yanjie
Data Mining and Applications in Genomics
 Data Mining and Applications in Genomics Lecture Notes in Electrical Engineering Volume 25 For other titles published in this series, go to www.springer.com/series/7818 Sio-Iong Ao Data Mining and Applications
Data Mining and Applications in Genomics Lecture Notes in Electrical Engineering Volume 25 For other titles published in this series, go to www.springer.com/series/7818 Sio-Iong Ao Data Mining and Applications
USES THE BAGGING ALGORITHM OF CLASSIFICATION METHOD WITH WEKA TOOL FOR PREDICTION TECHNIQUE
 USES THE BAGGING ALGORITHM OF CLASSIFICATION METHOD WITH WEKA TOOL FOR PREDICTION TECHNIQUE 1 POOJA SHRIVASTAVA, 2 MANOJ SHUKLA 1 Computer Science and Engineering, Jayoti Vidyapeeth Women s University,
USES THE BAGGING ALGORITHM OF CLASSIFICATION METHOD WITH WEKA TOOL FOR PREDICTION TECHNIQUE 1 POOJA SHRIVASTAVA, 2 MANOJ SHUKLA 1 Computer Science and Engineering, Jayoti Vidyapeeth Women s University,
Biology. Biology. Slide 1 of 39. End Show. Copyright Pearson Prentice Hall
 Biology Biology 1 of 39 12-3 RNA and Protein Synthesis 2 of 39 Essential Question What is transcription and translation and how do they take place? 3 of 39 12 3 RNA and Protein Synthesis Genes are coded
Biology Biology 1 of 39 12-3 RNA and Protein Synthesis 2 of 39 Essential Question What is transcription and translation and how do they take place? 3 of 39 12 3 RNA and Protein Synthesis Genes are coded
Biology. Biology. Slide 1 of 39. End Show. Copyright Pearson Prentice Hall
 Biology Biology 1 of 39 12-3 RNA and Protein Synthesis 2 of 39 12 3 RNA and Protein Synthesis Genes are coded DNA instructions that control the production of proteins. Genetic messages can be decoded by
Biology Biology 1 of 39 12-3 RNA and Protein Synthesis 2 of 39 12 3 RNA and Protein Synthesis Genes are coded DNA instructions that control the production of proteins. Genetic messages can be decoded by
Forecasting Electricity Consumption with Neural Networks and Support Vector Regression
 Available online at www.sciencedirect.com Procedia - Social and Behavioral Sciences 58 ( 2012 ) 1576 1585 8 th International Strategic Management Conference Forecasting Electricity Consumption with Neural
Available online at www.sciencedirect.com Procedia - Social and Behavioral Sciences 58 ( 2012 ) 1576 1585 8 th International Strategic Management Conference Forecasting Electricity Consumption with Neural
Machine learning applications in genomics: practical issues & challenges. Yuzhen Ye School of Informatics and Computing, Indiana University
 Machine learning applications in genomics: practical issues & challenges Yuzhen Ye School of Informatics and Computing, Indiana University Reference Machine learning applications in genetics and genomics
Machine learning applications in genomics: practical issues & challenges Yuzhen Ye School of Informatics and Computing, Indiana University Reference Machine learning applications in genetics and genomics
CS6716 Pattern Recognition
 CS6716 Pattern Recognition Aaron Bobick School of Interactive Computing Administrivia Shray says the problem set is close to done Today chapter 15 of the Hastie book. Very few slides brought to you by
CS6716 Pattern Recognition Aaron Bobick School of Interactive Computing Administrivia Shray says the problem set is close to done Today chapter 15 of the Hastie book. Very few slides brought to you by
A Protein Secondary Structure Prediction Method Based on BP Neural Network Ru-xi YIN, Li-zhen LIU*, Wei SONG, Xin-lei ZHAO and Chao DU
 2017 2nd International Conference on Artificial Intelligence: Techniques and Applications (AITA 2017 ISBN: 978-1-60595-491-2 A Protein Secondary Structure Prediction Method Based on BP Neural Network Ru-xi
2017 2nd International Conference on Artificial Intelligence: Techniques and Applications (AITA 2017 ISBN: 978-1-60595-491-2 A Protein Secondary Structure Prediction Method Based on BP Neural Network Ru-xi
Breast Cancer Diagnostic Factors Elimination via Evolutionary Neural Network Pruning
 Breast Cancer Diagnostic Factors Elimination via Evolutionary Neural Network Pruning Adam V. Adamopoulos Democritus University of Thrace, Department of Medicine, Medical Physics Laboratory, 681 00, Alexandroupolis,
Breast Cancer Diagnostic Factors Elimination via Evolutionary Neural Network Pruning Adam V. Adamopoulos Democritus University of Thrace, Department of Medicine, Medical Physics Laboratory, 681 00, Alexandroupolis,
Correcting Sampling Bias in Structural Genomics through Iterative Selection of Underrepresented Targets
 Correcting Sampling Bias in Structural Genomics through Iterative Selection of Underrepresented Targets Kang Peng Slobodan Vucetic Zoran Obradovic Abstract In this study we proposed an iterative procedure
Correcting Sampling Bias in Structural Genomics through Iterative Selection of Underrepresented Targets Kang Peng Slobodan Vucetic Zoran Obradovic Abstract In this study we proposed an iterative procedure
March 9, Hidden Markov Models and. BioInformatics, Part I. Steven R. Dunbar. Intro. BioInformatics Problem. Hidden Markov.
 and, and, March 9, 2017 1 / 30 Outline and, 1 2 3 4 2 / 30 Background and, Prof E. Moriyama (SBS) has a Seminar SBS, Math, Computer Science, Statistics Extensive use of program "HMMer" Britney (Hinds)
and, and, March 9, 2017 1 / 30 Outline and, 1 2 3 4 2 / 30 Background and, Prof E. Moriyama (SBS) has a Seminar SBS, Math, Computer Science, Statistics Extensive use of program "HMMer" Britney (Hinds)
Prediction of noncoding RNAs with RNAz
 Prediction of noncoding RNAs with RNAz John Dzmil, III Steve Griesmer Philip Murillo April 4, 2007 What is non-coding RNA (ncrna)? RNA molecules that are not translated into proteins Size range from 20
Prediction of noncoding RNAs with RNAz John Dzmil, III Steve Griesmer Philip Murillo April 4, 2007 What is non-coding RNA (ncrna)? RNA molecules that are not translated into proteins Size range from 20
Promoter Prediction in DNA Sequences
 Promoter Prediction in DNA Sequences A Thesis Submitted to the Faculty of Department of Computer Science and Engineering National Sun Yat-sen University by Jih-Wei Huang In Partial Fulfillment of the Requirements
Promoter Prediction in DNA Sequences A Thesis Submitted to the Faculty of Department of Computer Science and Engineering National Sun Yat-sen University by Jih-Wei Huang In Partial Fulfillment of the Requirements
Outline. Introduction to ab initio and evidence-based gene finding. Prokaryotic gene predictions
 Outline Introduction to ab initio and evidence-based gene finding Overview of computational gene predictions Different types of eukaryotic gene predictors Common types of gene prediction errors Wilson
Outline Introduction to ab initio and evidence-based gene finding Overview of computational gene predictions Different types of eukaryotic gene predictors Common types of gene prediction errors Wilson
Proactive Data Mining Using Decision Trees
 2012 IEEE 27-th Convention of Electrical and Electronics Engineers in Israel Proactive Data Mining Using Decision Trees Haim Dahan and Oded Maimon Dept. of Industrial Engineering Tel-Aviv University Tel
2012 IEEE 27-th Convention of Electrical and Electronics Engineers in Israel Proactive Data Mining Using Decision Trees Haim Dahan and Oded Maimon Dept. of Industrial Engineering Tel-Aviv University Tel
DNA. translation. base pairing rules for DNA Replication. thymine. cytosine. amino acids. The building blocks of proteins are?
 2 strands, has the 5-carbon sugar deoxyribose, and has the nitrogen base Thymine. The actual process of assembling the proteins on the ribosome is called? DNA translation Adenine pairs with Thymine, Thymine
2 strands, has the 5-carbon sugar deoxyribose, and has the nitrogen base Thymine. The actual process of assembling the proteins on the ribosome is called? DNA translation Adenine pairs with Thymine, Thymine
EECS730: Introduction to Bioinformatics
 EECS730: Introduction to Bioinformatics Lecture 08: Gene finding aatgcatgcggctatgctaatgcatgcggctatgctaagctgggatccgatgacaatgcatgcggctatgctaatgcatgcggc tatgcaagctgggatccgatgactatgctaagctgggatccgatgacaatgcatgcggctatgctaatgaatggtcttgggatt
EECS730: Introduction to Bioinformatics Lecture 08: Gene finding aatgcatgcggctatgctaatgcatgcggctatgctaagctgggatccgatgacaatgcatgcggctatgctaatgcatgcggc tatgcaagctgggatccgatgactatgctaagctgggatccgatgacaatgcatgcggctatgctaatgaatggtcttgggatt
The study of the structure, function, and interaction of cellular proteins is called. A) bioinformatics B) haplotypics C) genomics D) proteomics
 Human Biology, 12e (Mader / Windelspecht) Chapter 21 DNA Which of the following is not a component of a DNA molecule? A) a nitrogen-containing base B) deoxyribose sugar C) phosphate D) phospholipid Messenger
Human Biology, 12e (Mader / Windelspecht) Chapter 21 DNA Which of the following is not a component of a DNA molecule? A) a nitrogen-containing base B) deoxyribose sugar C) phosphate D) phospholipid Messenger
Videos. Lesson Overview. Fermentation
 Lesson Overview Fermentation Videos Bozeman Transcription and Translation: https://youtu.be/h3b9arupxzg Drawing transcription and translation: https://youtu.be/6yqplgnjr4q Objectives 29a) I can contrast
Lesson Overview Fermentation Videos Bozeman Transcription and Translation: https://youtu.be/h3b9arupxzg Drawing transcription and translation: https://youtu.be/6yqplgnjr4q Objectives 29a) I can contrast
Using AI to Make Predictions on Stock Market
 Using AI to Make Predictions on Stock Market Alice Zheng Stanford University Stanford, CA 94305 alicezhy@stanford.edu Jack Jin Stanford University Stanford, CA 94305 jackjin@stanford.edu 1 Introduction
Using AI to Make Predictions on Stock Market Alice Zheng Stanford University Stanford, CA 94305 alicezhy@stanford.edu Jack Jin Stanford University Stanford, CA 94305 jackjin@stanford.edu 1 Introduction
Generational and steady state genetic algorithms for generator maintenance scheduling problems
 Generational and steady state genetic algorithms for generator maintenance scheduling problems Item Type Conference paper Authors Dahal, Keshav P.; McDonald, J.R. Citation Dahal, K. P. and McDonald, J.
Generational and steady state genetic algorithms for generator maintenance scheduling problems Item Type Conference paper Authors Dahal, Keshav P.; McDonald, J.R. Citation Dahal, K. P. and McDonald, J.
Random forest for gene selection and microarray data classification
 www.bioinformation.net Hypothesis Volume 7(3) Random forest for gene selection and microarray data classification Kohbalan Moorthy & Mohd Saberi Mohamad* Artificial Intelligence & Bioinformatics Research
www.bioinformation.net Hypothesis Volume 7(3) Random forest for gene selection and microarray data classification Kohbalan Moorthy & Mohd Saberi Mohamad* Artificial Intelligence & Bioinformatics Research
Genomes: What we know and what we don t know
 Genomes: What we know and what we don t know Complete draft sequence 2001 October 15, 2007 Dr. Stefan Maas, BioS Lehigh U. What we know Raw genome data The range of genome sizes in the animal & plant kingdoms!
Genomes: What we know and what we don t know Complete draft sequence 2001 October 15, 2007 Dr. Stefan Maas, BioS Lehigh U. What we know Raw genome data The range of genome sizes in the animal & plant kingdoms!
Lecture 2: Biology Basics Continued. Fall 2018 August 23, 2018
 Lecture 2: Biology Basics Continued Fall 2018 August 23, 2018 Genetic Material for Life Central Dogma DNA: The Code of Life The structure and the four genomic letters code for all living organisms Adenine,
Lecture 2: Biology Basics Continued Fall 2018 August 23, 2018 Genetic Material for Life Central Dogma DNA: The Code of Life The structure and the four genomic letters code for all living organisms Adenine,
Research of Product Design based on Improved Genetic Algorithm
 , pp. 45-50 http://dx.doi.org/10.14257/ijhit.2016.9.6.04 Research of Product Design based on Improved Genetic Algorithm Li Ma (Zhejiang Industry Polytechnic College Shaoxing Zhejiang 312000 China) zjsxmali@sina.com
, pp. 45-50 http://dx.doi.org/10.14257/ijhit.2016.9.6.04 Research of Product Design based on Improved Genetic Algorithm Li Ma (Zhejiang Industry Polytechnic College Shaoxing Zhejiang 312000 China) zjsxmali@sina.com
New Methods for Splice Site Recognition
 New Methods for Splice Site Recognition S. Sonnenburg, G. Rätsch, A. Jagota, and K.-R. Müller Fraunhofer FIRST, Kekuléstr. 7, 12489 Berlin, Germany Australian National University, Canberra, ACT 0200, Australia
New Methods for Splice Site Recognition S. Sonnenburg, G. Rätsch, A. Jagota, and K.-R. Müller Fraunhofer FIRST, Kekuléstr. 7, 12489 Berlin, Germany Australian National University, Canberra, ACT 0200, Australia
Predicting Popularity of Messages in Twitter using a Feature-weighted Model
 International Journal of Advanced Intelligence Volume 0, Number 0, pp.xxx-yyy, November, 20XX. c AIA International Advanced Information Institute Predicting Popularity of Messages in Twitter using a Feature-weighted
International Journal of Advanced Intelligence Volume 0, Number 0, pp.xxx-yyy, November, 20XX. c AIA International Advanced Information Institute Predicting Popularity of Messages in Twitter using a Feature-weighted
Index Terms: Customer Loyalty, SVM, Data mining, Classification, Gaussian kernel
 Volume 4, Issue 12, December 2014 ISSN: 2277 128X International Journal of Advanced Research in Computer Science and Software Engineering Research Paper Available online at: www.ijarcsse.com Gaussian Kernel
Volume 4, Issue 12, December 2014 ISSN: 2277 128X International Journal of Advanced Research in Computer Science and Software Engineering Research Paper Available online at: www.ijarcsse.com Gaussian Kernel
Gene Regulation 10/19/05
 10/19/05 Gene Regulation (formerly Gene Prediction - 2) Gene Prediction & Regulation Mon - Overview & Gene structure review: Eukaryotes vs prokaryotes Wed - Regulatory regions: Promoters & enhancers -
10/19/05 Gene Regulation (formerly Gene Prediction - 2) Gene Prediction & Regulation Mon - Overview & Gene structure review: Eukaryotes vs prokaryotes Wed - Regulatory regions: Promoters & enhancers -
Bagged Ensembles of Support Vector Machines for Gene Expression Data Analysis
 Bagged Ensembles of Support Vector Machines for Gene Expression Data Analysis Giorgio Valentini INFM, Istituto Nazionale di Fisica della Materia, DSI, Dip. di Scienze dell Informazione Università degli
Bagged Ensembles of Support Vector Machines for Gene Expression Data Analysis Giorgio Valentini INFM, Istituto Nazionale di Fisica della Materia, DSI, Dip. di Scienze dell Informazione Università degli
DNA Gene Expression Classification with Ensemble Classifiers Optimized by Speciated Genetic Algorithm
 DNA Gene Expression Classification with Ensemble Classifiers Optimized by Speciated Genetic Algorithm Kyung-Joong Kim and Sung-Bae Cho Department of Computer Science, Yonsei University, 134 Shinchon-dong,
DNA Gene Expression Classification with Ensemble Classifiers Optimized by Speciated Genetic Algorithm Kyung-Joong Kim and Sung-Bae Cho Department of Computer Science, Yonsei University, 134 Shinchon-dong,
Chapter 3 Top Scoring Pair Decision Tree for Gene Expression Data Analysis
 Chapter 3 Top Scoring Pair Decision Tree for Gene Expression Data Analysis Marcin Czajkowski and Marek Krȩtowski Abstract Classification problems of microarray data may be successfully performed with approaches
Chapter 3 Top Scoring Pair Decision Tree for Gene Expression Data Analysis Marcin Czajkowski and Marek Krȩtowski Abstract Classification problems of microarray data may be successfully performed with approaches
