Discovering similar (epi)genomics feature patterns in multiple genome browser tracks
|
|
|
- Albert Turner
- 5 years ago
- Views:
Transcription
1 Genomic Computing Politecnico di Milano Dipartimento di Elettronica, Informazione e Bioingegneria Discovering similar (epi)genomics feature patterns in multiple genome browser tracks Piero Montanari 1, Arnaud Ceol 2, Ilaria Bartolini 1, Paolo Ciaccia 1, Marco Patella 1, Stefano Ceri 3, Marco Masseroli 3 1 DISI - Università di Bologna, 2 IIT@SEMM IIT, 3 DEIB - Politecnico di Milano
2 Background 2 Next Generation Sequencing (NGS) is opening many interesting practical and theoretical computational problems Genome browsers (UCSC Genome Browser, Integrated Genome Browser (IGB)) allow: Visual inspection and identification of interesting patterns on multiple genome browser tracks Pattern: a set of (epi)genomic regions / peaks at given distances from each other in different tracks e.g.: gene regulatory DNA areas, which include heterogeneous (epi)genomic features (different histone modifications, TFBSs,, )
3 Pattern-based queries from genome browser 3 A pattern of genomic features
4 Motivation 4 Once such patterns are visually identified in a genome section: Search for pattern occurrences in whole genome: Complex computational task Currently not supported Their discovery in whole genome very important for: Biological interpretation of NGS experimental results Comprehension of biomolecular phenomena
5 Reached goal 5 We defined an optimized pattern-search algorithm to find genomic region sets that are similar to a given pattern Efficiently In large (epi)genomic / transcriptomic data sets We implemented the algorithm within an IGB plugin, named SimSearch, which allows intuitive user interaction in both: Visual selection of an interesting pattern on loaded IGB tracks Visualization of occurrences of similar patterns identified in the whole genome
6 6 Method Best-Matching Problem (BMP) is suspected to be NP-hard For the considered application: Aligned genomic data -> strictly increasing region sequences M << N (M and N: number of elements in a pattern and in the target tracks to be compared, respectively) Our proposal: a Root-element approach (R-BMP) and Windowed Dynamic Programming algorithm (WDP-BMP) to lower complexity: Best matching for each element of the pattern in the target track through a binary search Complexity only of O(N*log(N)) Applicable also to (very) large data problems
7 7 Method Base problem / model Pattern = single track Regions = points, identified by their linear genomic coordinate Patterns and tracks = sequences of integers (region coordinates) Only relative distances between regions/points are relevant Pattern Target Track A matching is a strictly increasing function f that assigns to each pattern (query) element a (target) track element Preserving element order Pattern Pattern Target Track Target Track
8 8 Method Base problem / model Pattern matching typically solved by a cost based approach, where lower cost implies higher similarity Root-element approach (R-BMP): The cost C of a matching f is the sum of squared distances relative to the first (root) matching pair Q and T: sequences of query (pattern) and target elements T f(1) Q 1 : root-distance (relative distance between root elements) Goal: find the matching with minimum cost
9 9 Method Base problem / model Example Pattern (query) Q = <1,7,10> Target track T = <3,5,9,11,13,14,18,21> A possible matching: {(1->3),(7->9),(10->21)} Root-distance = 3-1 = 2 Cost = (3-1-2) 2 + (9-7-2) 2 + ( ) 2 = 81 Best matching {(1->5),(7->11),(10->14)} Root-distance = 5-1 = 4 Cost = (5-1-4) 2 + (11-7-4) 2 + ( ) 2 = 0
10 Method Base problem / model 10 Windowed Dynamic Programming algorithm (WDP-BMP) Main result: A matching is optimal if and only if all its partial matchings have minimum cost For all possible root positions in T: They are T - Q +1 For all regions Q i in Q: Find (with binary search) the best match for Q i in T Possibly, widen the window to avoid conflicts (regions in T that are best match for multiple regions in Q) For all possible matches: Compute the cost using the partial cost obtained till then Abandon the current solution if partial cost best cost till then Overall complexity: O( T Q (log T + Q ) But O( T log T ) if Q << T (usually Q =1 10, T ~ )
11 Method Extended model Multi-track patterns Same approach is repeated for each pattern track Negative matching tracks Pattern tracks with regions that should not appear in the solution Removed from the solution search space (before search start) Partial matching tracks Pattern tracks with regions that might be missing in the results A cost is considered for those regions that remain unmatched Region features The cost function includes the (dis-)similarity between features (attributes) of query and target track regions Regions as intervals Region size modeled as a region feature Top-K distinct matchings To consider diverse results, we require that the best K results have no matched region in common 12
12 13 Validation Finding enhancer regions Complex pattern 2 single region positive tracks (H3K4me1 and H3K27ac) 2 single region negative tracks (TSS and H3K4me3) 4 single region partial tracks (CTCF, DHS, P300 and Pol2) Target data tracks about K562 cell line (AML) from ENCODE Top-100 results visually inspected by an expert 100% precision (of the resulting matchings to the pattern) All (1,651) results compared with enhancer chromatin state regions from ChromHMM tool (Broad Institute), data available in ENCODE 85.46% precision (but ChromHMM uses more histone marks)
13 SimSearch: a plugin for IGB 14 Available at: Galaxy Bed/BigBed, BigWig, GTF HTS-flow ( The Integrated Genome Browser (
14 SimSearch: define the pattern to search for Load pattern from file Create pattern by selecting IGB tracks / regions Where to search Load pre-defined pattern (e.g. CHromHMM chromatin states from Roadmap Epigenomics Consortium, Nature 2015) 15
15 SimSearch: searching for new enhancers Pradeepa et al. (Nature Genetics 2016) identified a new class of enhancer (with H3K122ac and absence of H3K27ac) in mouse Several examples validated in mouse - Homozygous deletion of group 2 putative enhancer 42 kb downstream of Leukemia inhibitory factor (Lif) gene led to reduced expression of Lif, but not of flanking Hormad2 gene - Deletion of one allele of putative enhancer 30 kb upstream Tbx3 led to downregulation of Tbx3 16
16 17 SimSearch: searching for new enhancers Look for this new pattern in human samples: 1. Load tracks of human data 2. Add tracks to the pattern 3. Add TSS (negative)
17 18 SimSearch: editing the pattern In a new track (red), see all active regions (matches) found Similar match found in human cells
18 Conclusions 19 Definition of a method for finding patterns in genomic sequences and of a dynamic programming algorithm for efficient solution Experimental evaluation found it accurate and efficient Algorithm implemented as a plugin for Integrated Genome Browser Applicable to any (epi)genomic / transcriptomic region data, e.g.: Histone Modifications (HM), Transcription Factor Binding Sites (TFBS), Single Nucleotide Polymorphisms (SNP), Differentially Expressed Genes (DEG), DNase I Hypersensitive Sites (DHS), Transcription Start Sites (TSS), or any other annotations Support for understanding biomolecular mechanisms Response to treatments, pathology onset/development, Thanks to the GenData 2020 PRIN project and GeCo ERC project!
19 20 Thank you for your attention! Any question?
The ENCODE Encyclopedia. & Variant Annotation Using RegulomeDB and HaploReg
 The ENCODE Encyclopedia & Variant Annotation Using RegulomeDB and HaploReg Jill E. Moore Weng Lab University of Massachusetts Medical School October 10, 2015 Where s the Encyclopedia? ENCODE: Encyclopedia
The ENCODE Encyclopedia & Variant Annotation Using RegulomeDB and HaploReg Jill E. Moore Weng Lab University of Massachusetts Medical School October 10, 2015 Where s the Encyclopedia? ENCODE: Encyclopedia
Introduction to NGS analyses
 Introduction to NGS analyses Giorgio L Papadopoulos Institute of Molecular Biology and Biotechnology Bioinformatics Support Group 04/12/2015 Papadopoulos GL (IMBB, FORTH) IMBB NGS Seminar 04/12/2015 1
Introduction to NGS analyses Giorgio L Papadopoulos Institute of Molecular Biology and Biotechnology Bioinformatics Support Group 04/12/2015 Papadopoulos GL (IMBB, FORTH) IMBB NGS Seminar 04/12/2015 1
What Can the Epigenome Teach Us About Cellular States and Diseases?
 What Can the Epigenome Teach Us About Cellular States and Diseases? (a computer scientist s view) Luca Pinello Outline Epigenetic: the code over the code What can we learn from epigenomic data? Resources
What Can the Epigenome Teach Us About Cellular States and Diseases? (a computer scientist s view) Luca Pinello Outline Epigenetic: the code over the code What can we learn from epigenomic data? Resources
Genome 541 Gene regulation and epigenomics Lecture 3 Integrative analysis of genomics assays
 Genome 541 Gene regulation and epigenomics Lecture 3 Integrative analysis of genomics assays Please consider both the forward and reverse strands (i.e. reverse compliment sequence). You do not need to
Genome 541 Gene regulation and epigenomics Lecture 3 Integrative analysis of genomics assays Please consider both the forward and reverse strands (i.e. reverse compliment sequence). You do not need to
NGS Approaches to Epigenomics
 I519 Introduction to Bioinformatics, 2013 NGS Approaches to Epigenomics Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Contents Background: chromatin structure & DNA methylation Epigenomic
I519 Introduction to Bioinformatics, 2013 NGS Approaches to Epigenomics Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Contents Background: chromatin structure & DNA methylation Epigenomic
Epigenetics and DNase-Seq
 Epigenetics and DNase-Seq BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 2018 Anthony Gitter gitter@biostat.wisc.edu These slides, excluding third-party material, are licensed under CC BY-NC 4.0 by Anthony
Epigenetics and DNase-Seq BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 2018 Anthony Gitter gitter@biostat.wisc.edu These slides, excluding third-party material, are licensed under CC BY-NC 4.0 by Anthony
Genome 541! Unit 4, lecture 3! Genomics assays
 Genome 541! Unit 4, lecture 3! Genomics assays I d like a bit more background on the assays and bioterminology.!! The phantom peak concept was confusing.! I didn t quite understand what the phantom peak
Genome 541! Unit 4, lecture 3! Genomics assays I d like a bit more background on the assays and bioterminology.!! The phantom peak concept was confusing.! I didn t quite understand what the phantom peak
About Strand NGS. Strand Genomics, Inc All rights reserved.
 About Strand NGS Strand NGS-formerly known as Avadis NGS, is an integrated platform that provides analysis, management and visualization tools for next-generation sequencing data. It supports extensive
About Strand NGS Strand NGS-formerly known as Avadis NGS, is an integrated platform that provides analysis, management and visualization tools for next-generation sequencing data. It supports extensive
From Variants to Pathways: Agilent GeneSpring GX s Variant Analysis Workflow
 From Variants to Pathways: Agilent GeneSpring GX s Variant Analysis Workflow Technical Overview Import VCF Introduction Next-generation sequencing (NGS) studies have created unanticipated challenges with
From Variants to Pathways: Agilent GeneSpring GX s Variant Analysis Workflow Technical Overview Import VCF Introduction Next-generation sequencing (NGS) studies have created unanticipated challenges with
Introduction to RNA-Seq in GeneSpring NGS Software
 Introduction to RNA-Seq in GeneSpring NGS Software Dipa Roy Choudhury, Ph.D. Strand Scientific Intelligence and Agilent Technologies Learn more at www.genespring.com Introduction to RNA-Seq In a few years,
Introduction to RNA-Seq in GeneSpring NGS Software Dipa Roy Choudhury, Ph.D. Strand Scientific Intelligence and Agilent Technologies Learn more at www.genespring.com Introduction to RNA-Seq In a few years,
The ChIP-Seq project. Giovanna Ambrosini, Philipp Bucher. April 19, 2010 Lausanne. EPFL-SV Bucher Group
 The ChIP-Seq project Giovanna Ambrosini, Philipp Bucher EPFL-SV Bucher Group April 19, 2010 Lausanne Overview Focus on technical aspects Description of applications (C programs) Where to find binaries,
The ChIP-Seq project Giovanna Ambrosini, Philipp Bucher EPFL-SV Bucher Group April 19, 2010 Lausanne Overview Focus on technical aspects Description of applications (C programs) Where to find binaries,
Supplementary Table 1: Oligo designs. A list of ATAC-seq oligos used for PCR.
 Ad1_noMX: Ad2.1_TAAGGCGA Ad2.2_CGTACTAG Ad2.3_AGGCAGAA Ad2.4_TCCTGAGC Ad2.5_GGACTCCT Ad2.6_TAGGCATG Ad2.7_CTCTCTAC Ad2.8_CAGAGAGG Ad2.9_GCTACGCT Ad2.10_CGAGGCTG Ad2.11_AAGAGGCA Ad2.12_GTAGAGGA Ad2.13_GTCGTGAT
Ad1_noMX: Ad2.1_TAAGGCGA Ad2.2_CGTACTAG Ad2.3_AGGCAGAA Ad2.4_TCCTGAGC Ad2.5_GGACTCCT Ad2.6_TAGGCATG Ad2.7_CTCTCTAC Ad2.8_CAGAGAGG Ad2.9_GCTACGCT Ad2.10_CGAGGCTG Ad2.11_AAGAGGCA Ad2.12_GTAGAGGA Ad2.13_GTCGTGAT
Go to Bottom Left click WashU Epigenome Browser. Click
 Now you are going to look at the Human Epigenome Browswer. It has a more sophisticated but weirder interface than the UCSC Genome Browser. All the data that you will view as tracks is in reality just files
Now you are going to look at the Human Epigenome Browswer. It has a more sophisticated but weirder interface than the UCSC Genome Browser. All the data that you will view as tracks is in reality just files
Understanding Transcripts in UCSC Genome Browser
 Understanding Transcripts in UCSC Genome Browser Cline and Kent, Nature Biotechnology 27, 153-155 (2009) 1 Gene Annotations 2 Gene Annotations 2 3 Galaxy Databases are not analyses tools. Databases are
Understanding Transcripts in UCSC Genome Browser Cline and Kent, Nature Biotechnology 27, 153-155 (2009) 1 Gene Annotations 2 Gene Annotations 2 3 Galaxy Databases are not analyses tools. Databases are
Galaxy Platform For NGS Data Analyses
 Galaxy Platform For NGS Data Analyses Weihong Yan wyan@chem.ucla.edu Collaboratory Web Site http://qcb.ucla.edu/collaboratory http://collaboratory.lifesci.ucla.edu Workshop Outline ü Day 1 UCLA galaxy
Galaxy Platform For NGS Data Analyses Weihong Yan wyan@chem.ucla.edu Collaboratory Web Site http://qcb.ucla.edu/collaboratory http://collaboratory.lifesci.ucla.edu Workshop Outline ü Day 1 UCLA galaxy
Leveraging variant annotations for WGS data analysis
 1 Deepti Jain (jaind@uw.edu) and Ben Heavner (bheavner@uw.edu) Leveraging variant annotations for WGS data analysis Deepti Jain & Ben Heavner Data coordinating center for Trans-Omics for Precision Medicine
1 Deepti Jain (jaind@uw.edu) and Ben Heavner (bheavner@uw.edu) Leveraging variant annotations for WGS data analysis Deepti Jain & Ben Heavner Data coordinating center for Trans-Omics for Precision Medicine
Multi-omics in biology: integration of omics techniques
 31/07/17 Летняя школа по биоинформатике 2017 Multi-omics in biology: integration of omics techniques Konstantin Okonechnikov Division of Pediatric Neurooncology German Cancer Research Center (DKFZ) 2 Short
31/07/17 Летняя школа по биоинформатике 2017 Multi-omics in biology: integration of omics techniques Konstantin Okonechnikov Division of Pediatric Neurooncology German Cancer Research Center (DKFZ) 2 Short
Understanding transcriptional regulation by integrative analysis of transcription factor binding data
 Understanding transcriptional regulation by integrative analysis of transcription factor binding data Cheng et al. 2012 Shu Yang Feb. 21, 2013 1 / 26 Introduction 2 / 26 DNA-binding Proteins sequence-specific
Understanding transcriptional regulation by integrative analysis of transcription factor binding data Cheng et al. 2012 Shu Yang Feb. 21, 2013 1 / 26 Introduction 2 / 26 DNA-binding Proteins sequence-specific
Introduction to Bioinformatics
 Introduction to Bioinformatics Richard Corbett Canada s Michael Smith Genome Sciences Centre Vancouver, British Columbia June 28, 2017 Our mandate is to advance knowledge about cancer and other diseases
Introduction to Bioinformatics Richard Corbett Canada s Michael Smith Genome Sciences Centre Vancouver, British Columbia June 28, 2017 Our mandate is to advance knowledge about cancer and other diseases
Browser Exercises - I. Alignments and Comparative genomics
 Browser Exercises - I Alignments and Comparative genomics 1. Navigating to the Genome Browser (GBrowse) Note: For this exercise use http://www.tritrypdb.org a. Navigate to the Genome Browser (GBrowse)
Browser Exercises - I Alignments and Comparative genomics 1. Navigating to the Genome Browser (GBrowse) Note: For this exercise use http://www.tritrypdb.org a. Navigate to the Genome Browser (GBrowse)
GMQL - Biological examples
 GMQL - Biological examples This section of GMQL documentation collects several examples where GMQL is used to answer practical questions/tasks of biological and clinical interest. For each example, after
GMQL - Biological examples This section of GMQL documentation collects several examples where GMQL is used to answer practical questions/tasks of biological and clinical interest. For each example, after
The ENCODE Encyclopedia. Zhiping Weng University of Massachusetts Medical School ENCODE 2016: Research Applications and Users Meeting June 8, 2016
 The ENCODE Encyclopedia Zhiping Weng University of Massachusetts Medical School ENCODE 2016: Research Applications and Users Meeting June 8, 2016 The Encyclopedia Of DNA Elements Consortium Goals: - Catalog
The ENCODE Encyclopedia Zhiping Weng University of Massachusetts Medical School ENCODE 2016: Research Applications and Users Meeting June 8, 2016 The Encyclopedia Of DNA Elements Consortium Goals: - Catalog
2/19/13. Contents. Applications of HMMs in Epigenomics
 2/19/13 I529: Machine Learning in Bioinformatics (Spring 2013) Contents Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2013 Background:
2/19/13 I529: Machine Learning in Bioinformatics (Spring 2013) Contents Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2013 Background:
Applications of HMMs in Epigenomics
 I529: Machine Learning in Bioinformatics (Spring 2013) Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2013 Contents Background:
I529: Machine Learning in Bioinformatics (Spring 2013) Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2013 Contents Background:
2/10/17. Contents. Applications of HMMs in Epigenomics
 2/10/17 I529: Machine Learning in Bioinformatics (Spring 2017) Contents Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2017 Background:
2/10/17 I529: Machine Learning in Bioinformatics (Spring 2017) Contents Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2017 Background:
MODULE TSS1: TRANSCRIPTION START SITES INTRODUCTION (BASIC)
 MODULE TSS1: TRANSCRIPTION START SITES INTRODUCTION (BASIC) Lesson Plan: Title JAMIE SIDERS, MEG LAAKSO & WILSON LEUNG Identifying transcription start sites for Peaked promoters using chromatin landscape,
MODULE TSS1: TRANSCRIPTION START SITES INTRODUCTION (BASIC) Lesson Plan: Title JAMIE SIDERS, MEG LAAKSO & WILSON LEUNG Identifying transcription start sites for Peaked promoters using chromatin landscape,
Nature Genetics: doi: /ng Supplementary Figure 1. ChIP-seq genome browser views of BRM occupancy at previously identified BRM targets.
 Supplementary Figure 1 ChIP-seq genome browser views of BRM occupancy at previously identified BRM targets. Gene structures are shown underneath each panel. Supplementary Figure 2 pref6::ref6-gfp complements
Supplementary Figure 1 ChIP-seq genome browser views of BRM occupancy at previously identified BRM targets. Gene structures are shown underneath each panel. Supplementary Figure 2 pref6::ref6-gfp complements
Deep learning frameworks for regulatory genomics and epigenomics
 Deep learning frameworks for regulatory genomics and epigenomics Chuan Sheng Foo Avanti Shrikumar Nicholas Sinnott- Armstrong ANSHUL KUNDAJE Genetics, Computer science Stanford University Johnny Israeli
Deep learning frameworks for regulatory genomics and epigenomics Chuan Sheng Foo Avanti Shrikumar Nicholas Sinnott- Armstrong ANSHUL KUNDAJE Genetics, Computer science Stanford University Johnny Israeli
Reads to Discovery. Visualize Annotate Discover. Small DNA-Seq ChIP-Seq Methyl-Seq. MeDIP-Seq. RNA-Seq. RNA-Seq.
 Reads to Discovery RNA-Seq Small DNA-Seq ChIP-Seq Methyl-Seq RNA-Seq MeDIP-Seq www.strand-ngs.com Analyze Visualize Annotate Discover Data Import Alignment Vendor Platforms: Illumina Ion Torrent Roche
Reads to Discovery RNA-Seq Small DNA-Seq ChIP-Seq Methyl-Seq RNA-Seq MeDIP-Seq www.strand-ngs.com Analyze Visualize Annotate Discover Data Import Alignment Vendor Platforms: Illumina Ion Torrent Roche
Complete Sample to Analysis Solutions for DNA Methylation Discovery using Next Generation Sequencing
 Complete Sample to Analysis Solutions for DNA Methylation Discovery using Next Generation Sequencing SureSelect Human/Mouse Methyl-Seq Kyeong Jeong PhD February 5, 2013 CAG EMEAI DGG/GSD/GFO Agilent Restricted
Complete Sample to Analysis Solutions for DNA Methylation Discovery using Next Generation Sequencing SureSelect Human/Mouse Methyl-Seq Kyeong Jeong PhD February 5, 2013 CAG EMEAI DGG/GSD/GFO Agilent Restricted
Ensembl Funcgen: A Database and API for Epigenomics and Gene Regulation Data.
 Ensembl Funcgen: A Database and API for Epigenomics and Gene Regulation Data. Nathan Johnson Ensembl Regulation EBI is an Outstation of the European Molecular Biology Laboratory.! Workshop Overview http://www.ebi.ac.uk/~njohnson/courses/23.05.2013-
Ensembl Funcgen: A Database and API for Epigenomics and Gene Regulation Data. Nathan Johnson Ensembl Regulation EBI is an Outstation of the European Molecular Biology Laboratory.! Workshop Overview http://www.ebi.ac.uk/~njohnson/courses/23.05.2013-
Introduction to human genomics and genome informatics
 Introduction to human genomics and genome informatics Session 1 Prince of Wales Clinical School Dr Jason Wong ARC Future Fellow Head, Bioinformatics & Integrative Genomics Adult Cancer Program, Lowy Cancer
Introduction to human genomics and genome informatics Session 1 Prince of Wales Clinical School Dr Jason Wong ARC Future Fellow Head, Bioinformatics & Integrative Genomics Adult Cancer Program, Lowy Cancer
Introduction to genome biology
 Introduction to genome biology Lisa Stubbs We ve found most genes; but what about the rest of the genome? Genome size* 12 Mb 95 Mb 170 Mb 1500 Mb 2700 Mb 3200 Mb #coding genes ~7000 ~20000 ~14000 ~26000
Introduction to genome biology Lisa Stubbs We ve found most genes; but what about the rest of the genome? Genome size* 12 Mb 95 Mb 170 Mb 1500 Mb 2700 Mb 3200 Mb #coding genes ~7000 ~20000 ~14000 ~26000
ChIP-Seq, mrna-seq, & Resequencing via the Genboree Workench
 ChIP-Seq, mrna-seq, & Resequencing via the Genboree Workench Chromatin ImmunopreciPitation Sequencing ChIP-Seq Peak Calling Transcription Factors Histone modifications H3K4me3, H3K27me3, H3K36me3, H3K9me3,
ChIP-Seq, mrna-seq, & Resequencing via the Genboree Workench Chromatin ImmunopreciPitation Sequencing ChIP-Seq Peak Calling Transcription Factors Histone modifications H3K4me3, H3K27me3, H3K36me3, H3K9me3,
A Brief History. Bootstrapping. Bagging. Boosting (Schapire 1989) Adaboost (Schapire 1995)
 A Brief History Bootstrapping Bagging Boosting (Schapire 1989) Adaboost (Schapire 1995) What s So Good About Adaboost Improves classification accuracy Can be used with many different classifiers Commonly
A Brief History Bootstrapping Bagging Boosting (Schapire 1989) Adaboost (Schapire 1995) What s So Good About Adaboost Improves classification accuracy Can be used with many different classifiers Commonly
ChIP-seq and RNA-seq. Farhat Habib
 ChIP-seq and RNA-seq Farhat Habib fhabib@iiserpune.ac.in Biological Goals Learn how genomes encode the diverse patterns of gene expression that define each cell type and state. Protein-DNA interactions
ChIP-seq and RNA-seq Farhat Habib fhabib@iiserpune.ac.in Biological Goals Learn how genomes encode the diverse patterns of gene expression that define each cell type and state. Protein-DNA interactions
ChIP-seq and RNA-seq
 ChIP-seq and RNA-seq Biological Goals Learn how genomes encode the diverse patterns of gene expression that define each cell type and state. Protein-DNA interactions (ChIPchromatin immunoprecipitation)
ChIP-seq and RNA-seq Biological Goals Learn how genomes encode the diverse patterns of gene expression that define each cell type and state. Protein-DNA interactions (ChIPchromatin immunoprecipitation)
Comparative eqtl analyses within and between seven tissue types suggest mechanisms underlying cell type specificity of eqtls
 Comparative eqtl analyses within and between seven tissue types suggest mechanisms underlying cell type specificity of eqtls, Duke University Christopher D Brown, University of Pennsylvania November 9th,
Comparative eqtl analyses within and between seven tissue types suggest mechanisms underlying cell type specificity of eqtls, Duke University Christopher D Brown, University of Pennsylvania November 9th,
SNPs - GWAS - eqtls. Sebastian Schmeier
 SNPs - GWAS - eqtls s.schmeier@gmail.com http://sschmeier.github.io/bioinf-workshop/ 17.08.2015 Overview Single nucleotide polymorphism (refresh) SNPs effect on genes (refresh) Genome-wide association
SNPs - GWAS - eqtls s.schmeier@gmail.com http://sschmeier.github.io/bioinf-workshop/ 17.08.2015 Overview Single nucleotide polymorphism (refresh) SNPs effect on genes (refresh) Genome-wide association
Biology 644: Bioinformatics
 Processes Activation Repression Initiation Elongation.... Processes Splicing Editing Degradation Translation.... Transcription Translation DNA Regulators DNA-Binding Transcription Factors Chromatin Remodelers....
Processes Activation Repression Initiation Elongation.... Processes Splicing Editing Degradation Translation.... Transcription Translation DNA Regulators DNA-Binding Transcription Factors Chromatin Remodelers....
RNA-SEQUENCING ANALYSIS
 RNA-SEQUENCING ANALYSIS Joseph Powell SISG- 2018 CONTENTS Introduction to RNA sequencing Data structure Analyses Transcript counting Alternative splicing Allele specific expression Discovery APPLICATIONS
RNA-SEQUENCING ANALYSIS Joseph Powell SISG- 2018 CONTENTS Introduction to RNA sequencing Data structure Analyses Transcript counting Alternative splicing Allele specific expression Discovery APPLICATIONS
V23: the histone code
 V23: the histone code The DNA of eukaryotic organisms is packaged into chromatin, whose basic repeating unit is the nucleosome. A nucleosome is formed by wrapping 147 base pairs of DNA twice around an
V23: the histone code The DNA of eukaryotic organisms is packaged into chromatin, whose basic repeating unit is the nucleosome. A nucleosome is formed by wrapping 147 base pairs of DNA twice around an
Figure S1. nuclear extracts. HeLa cell nuclear extract. Input IgG IP:ORC2 ORC2 ORC2. MCM4 origin. ORC2 occupancy
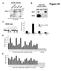 A nuclear extracts B HeLa cell nuclear extract Figure S1 ORC2 (in kda) 21 132 7 ORC2 Input IgG IP:ORC2 32 ORC C D PRKDC ORC2 occupancy Directed against ORC2 C-terminus (sc-272) MCM origin 2 2 1-1 -1kb
A nuclear extracts B HeLa cell nuclear extract Figure S1 ORC2 (in kda) 21 132 7 ORC2 Input IgG IP:ORC2 32 ORC C D PRKDC ORC2 occupancy Directed against ORC2 C-terminus (sc-272) MCM origin 2 2 1-1 -1kb
A survey on computational methods for enhancer and enhancer target predictions Introduction
 A survey on computational methods for enhancer and enhancer target predictions Qin Cao and Kevin Y. Yip Department of Computer Science and Engineering, The Chinese University of Hong Kong, Shatin, New
A survey on computational methods for enhancer and enhancer target predictions Qin Cao and Kevin Y. Yip Department of Computer Science and Engineering, The Chinese University of Hong Kong, Shatin, New
Predicting tissue specific cis-regulatory modules in the human genome using pairs of co-occurring motifs
 BMC Bioinformatics This Provisional PDF corresponds to the article as it appeared upon acceptance. Fully formatted PDF and full text (HTML) versions will be made available soon. Predicting tissue specific
BMC Bioinformatics This Provisional PDF corresponds to the article as it appeared upon acceptance. Fully formatted PDF and full text (HTML) versions will be made available soon. Predicting tissue specific
Week 1 BCHM 6280 Tutorial: Gene specific information using NCBI, Ensembl and genome viewers
 Week 1 BCHM 6280 Tutorial: Gene specific information using NCBI, Ensembl and genome viewers Web resources: NCBI database: http://www.ncbi.nlm.nih.gov/ Ensembl database: http://useast.ensembl.org/index.html
Week 1 BCHM 6280 Tutorial: Gene specific information using NCBI, Ensembl and genome viewers Web resources: NCBI database: http://www.ncbi.nlm.nih.gov/ Ensembl database: http://useast.ensembl.org/index.html
Nature Genetics: doi: /ng Supplementary Figure 1
 Supplementary Figure 1 Characterization of Hi-C/CHi-C dynamics and enhancer identification. (a) Scatterplot of Hi-C read counts supporting contacts between domain boundaries. Contacts enclosing domains
Supplementary Figure 1 Characterization of Hi-C/CHi-C dynamics and enhancer identification. (a) Scatterplot of Hi-C read counts supporting contacts between domain boundaries. Contacts enclosing domains
user s guide Question 1
 Question 1 How does one find a gene of interest and determine that gene s structure? Once the gene has been located on the map, how does one easily examine other genes in that same region? doi:10.1038/ng966
Question 1 How does one find a gene of interest and determine that gene s structure? Once the gene has been located on the map, how does one easily examine other genes in that same region? doi:10.1038/ng966
Supplementary Figures
 Supplementary Figures A B Supplementary Figure 1. Examples of discrepancies in predicted and validated breakpoint coordinates. A) Most frequently, predicted breakpoints were shifted relative to those derived
Supplementary Figures A B Supplementary Figure 1. Examples of discrepancies in predicted and validated breakpoint coordinates. A) Most frequently, predicted breakpoints were shifted relative to those derived
Whole Transcriptome Analysis of Illumina RNA- Seq Data. Ryan Peters Field Application Specialist
 Whole Transcriptome Analysis of Illumina RNA- Seq Data Ryan Peters Field Application Specialist Partek GS in your NGS Pipeline Your Start-to-Finish Solution for Analysis of Next Generation Sequencing Data
Whole Transcriptome Analysis of Illumina RNA- Seq Data Ryan Peters Field Application Specialist Partek GS in your NGS Pipeline Your Start-to-Finish Solution for Analysis of Next Generation Sequencing Data
SeattleSNPs Interactive Tutorial: Database Inteface Entrez, dbsnp, HapMap, Perlegen
 SeattleSNPs Interactive Tutorial: Database Inteface Entrez, dbsnp, HapMap, Perlegen The tutorial is designed to take you through the steps necessary to access SNP data from the primary database resources:
SeattleSNPs Interactive Tutorial: Database Inteface Entrez, dbsnp, HapMap, Perlegen The tutorial is designed to take you through the steps necessary to access SNP data from the primary database resources:
Mapping long-range promoter contacts in human cells with high-resolution capture Hi-C
 CORRECTION NOTICE Nat. Genet. 47, 598 606 (2015) Mapping long-range promoter contacts in human cells with high-resolution capture Hi-C Borbala Mifsud, Filipe Tavares-Cadete, Alice N Young, Robert Sugar,
CORRECTION NOTICE Nat. Genet. 47, 598 606 (2015) Mapping long-range promoter contacts in human cells with high-resolution capture Hi-C Borbala Mifsud, Filipe Tavares-Cadete, Alice N Young, Robert Sugar,
DNA. bioinformatics. epigenetics methylation structural variation. custom. assembly. gene. tumor-normal. mendelian. BS-seq. prediction.
 Epigenomics T TM activation SNP target ncrna validation metagenomics genetics private RRBS-seq de novo trio RIP-seq exome mendelian comparative genomics DNA NGS ChIP-seq bioinformatics assembly tumor-normal
Epigenomics T TM activation SNP target ncrna validation metagenomics genetics private RRBS-seq de novo trio RIP-seq exome mendelian comparative genomics DNA NGS ChIP-seq bioinformatics assembly tumor-normal
Analysis of ChIP-seq data with R / Bioconductor
 Analysis of ChIP-seq data with R / Bioconductor Martin Morgan Bioconductor / Fred Hutchinson Cancer Research Center Seattle, WA, USA 8-10 June 2009 ChIP-seq Chromatin immunopreciptation to enrich sample
Analysis of ChIP-seq data with R / Bioconductor Martin Morgan Bioconductor / Fred Hutchinson Cancer Research Center Seattle, WA, USA 8-10 June 2009 ChIP-seq Chromatin immunopreciptation to enrich sample
Finding Enrichments of Functional Annotations for Disease- Associated Single-Nucleotide Polymorphisms
 Finding Enrichments of Functional Annotations for Disease- Associated Single-Nucleotide Polymorphisms Steven Homberg 11/10/13 1 Abstract Computational analysis of SNP-disease associations from GWAS as
Finding Enrichments of Functional Annotations for Disease- Associated Single-Nucleotide Polymorphisms Steven Homberg 11/10/13 1 Abstract Computational analysis of SNP-disease associations from GWAS as
arxiv: v1 [q-bio.gn] 11 Oct 2012
![arxiv: v1 [q-bio.gn] 11 Oct 2012 arxiv: v1 [q-bio.gn] 11 Oct 2012](/thumbs/96/129046863.jpg) 1 arxiv:121.3294v1 [q-bio.gn] 11 Oct 12 Integrative modeling of eqtls and cis-regulatory elements suggest mechanisms underlying cell type specificity of eqtls Christopher D Brown 1,2, Lara M Mangravite
1 arxiv:121.3294v1 [q-bio.gn] 11 Oct 12 Integrative modeling of eqtls and cis-regulatory elements suggest mechanisms underlying cell type specificity of eqtls Christopher D Brown 1,2, Lara M Mangravite
Supplementary Figure 1 Strategy for parallel detection of DHSs and adjacent nucleosomes
 Supplementary Figure 1 Strategy for parallel detection of DHSs and adjacent nucleosomes DNase I cleavage DNase I DNase I digestion Sucrose gradient enrichment Small Large F1 F2...... F9 F1 F1 F2 F3 F4
Supplementary Figure 1 Strategy for parallel detection of DHSs and adjacent nucleosomes DNase I cleavage DNase I DNase I digestion Sucrose gradient enrichment Small Large F1 F2...... F9 F1 F1 F2 F3 F4
BCHM 6280 Tutorial: Gene specific information using NCBI, Ensembl and genome viewers
 BCHM 6280 Tutorial: Gene specific information using NCBI, Ensembl and genome viewers Web resources: NCBI database: http://www.ncbi.nlm.nih.gov/ Ensembl database: http://useast.ensembl.org/index.html UCSC
BCHM 6280 Tutorial: Gene specific information using NCBI, Ensembl and genome viewers Web resources: NCBI database: http://www.ncbi.nlm.nih.gov/ Ensembl database: http://useast.ensembl.org/index.html UCSC
ChIP-seq data analysis with Chipster. Eija Korpelainen CSC IT Center for Science, Finland
 ChIP-seq data analysis with Chipster Eija Korpelainen CSC IT Center for Science, Finland chipster@csc.fi What will I learn? Short introduction to ChIP-seq Analyzing ChIP-seq data Central concepts Analysis
ChIP-seq data analysis with Chipster Eija Korpelainen CSC IT Center for Science, Finland chipster@csc.fi What will I learn? Short introduction to ChIP-seq Analyzing ChIP-seq data Central concepts Analysis
A WEB-BASED TOOL FOR GENOMIC FUNCTIONAL ANNOTATION, STATISTICAL ANALYSIS AND DATA MINING
 A WEB-BASED TOOL FOR GENOMIC FUNCTIONAL ANNOTATION, STATISTICAL ANALYSIS AND DATA MINING D. Martucci a, F. Pinciroli a,b, M. Masseroli a a Dipartimento di Bioingegneria, Politecnico di Milano, Milano,
A WEB-BASED TOOL FOR GENOMIC FUNCTIONAL ANNOTATION, STATISTICAL ANALYSIS AND DATA MINING D. Martucci a, F. Pinciroli a,b, M. Masseroli a a Dipartimento di Bioingegneria, Politecnico di Milano, Milano,
Introduction to genome biology
 Introduction to genome biology Lisa Stubbs Deep transcritpomes for traditional model species from ENCODE (and modencode) Deep RNA-seq and chromatin analysis on 147 human cell types, as well as tissues,
Introduction to genome biology Lisa Stubbs Deep transcritpomes for traditional model species from ENCODE (and modencode) Deep RNA-seq and chromatin analysis on 147 human cell types, as well as tissues,
An Analytical Upper Bound on the Minimum Number of. Recombinations in the History of SNP Sequences in Populations
 An Analytical Upper Bound on the Minimum Number of Recombinations in the History of SNP Sequences in Populations Yufeng Wu Department of Computer Science and Engineering University of Connecticut Storrs,
An Analytical Upper Bound on the Minimum Number of Recombinations in the History of SNP Sequences in Populations Yufeng Wu Department of Computer Science and Engineering University of Connecticut Storrs,
Decoding Chromatin States with Epigenome Data Advanced Topics in Computa8onal Genomics
 Decoding Chromatin States with Epigenome Data 02-715 Advanced Topics in Computa8onal Genomics HMMs for Decoding Chromatin States Epigene8c modifica8ons of the genome have been associated with Establishing
Decoding Chromatin States with Epigenome Data 02-715 Advanced Topics in Computa8onal Genomics HMMs for Decoding Chromatin States Epigene8c modifica8ons of the genome have been associated with Establishing
IBM Research Report. Genomic Regions Tools for High-throughput Analytics in Genomics
 RC25125 (W1103-072) March 10, 2011 Other IBM Research Report Genomic Regions Tools for High-throughput Analytics in Genomics A. Tsirigos, N. Haiminen, E. Bilal IBM Research Division Thomas J. Watson Research
RC25125 (W1103-072) March 10, 2011 Other IBM Research Report Genomic Regions Tools for High-throughput Analytics in Genomics A. Tsirigos, N. Haiminen, E. Bilal IBM Research Division Thomas J. Watson Research
Nature Genetics: doi: /ng Supplementary Figure 1. H3K27ac HiChIP enriches enhancer promoter-associated chromatin contacts.
 Supplementary Figure 1 H3K27ac HiChIP enriches enhancer promoter-associated chromatin contacts. (a) Schematic of chromatin contacts captured in H3K27ac HiChIP. (b) Loop call overlap for cohesin HiChIP
Supplementary Figure 1 H3K27ac HiChIP enriches enhancer promoter-associated chromatin contacts. (a) Schematic of chromatin contacts captured in H3K27ac HiChIP. (b) Loop call overlap for cohesin HiChIP
Frumkin, 2e Part 1: Methods and Paradigms. Chapter 6: Genetics and Environmental Health
 Frumkin, 2e Part 1: Methods and Paradigms Chapter 6: Genetics and Environmental Health Genetics Genetics, the study of individual genes, has expanded to include genomics, which is the study of all the
Frumkin, 2e Part 1: Methods and Paradigms Chapter 6: Genetics and Environmental Health Genetics Genetics, the study of individual genes, has expanded to include genomics, which is the study of all the
Personal and population genomics of human regulatory variation
 Personal and population genomics of human regulatory variation Benjamin Vernot,, and Joshua M. Akey Department of Genome Sciences, University of Washington, Seattle, Washington 98195, USA Ting WANG 1 Brief
Personal and population genomics of human regulatory variation Benjamin Vernot,, and Joshua M. Akey Department of Genome Sciences, University of Washington, Seattle, Washington 98195, USA Ting WANG 1 Brief
Title: Genome-Wide Predictions of Transcription Factor Binding Events using Multi- Dimensional Genomic and Epigenomic Features Background
 Title: Genome-Wide Predictions of Transcription Factor Binding Events using Multi- Dimensional Genomic and Epigenomic Features Team members: David Moskowitz and Emily Tsang Background Transcription factors
Title: Genome-Wide Predictions of Transcription Factor Binding Events using Multi- Dimensional Genomic and Epigenomic Features Team members: David Moskowitz and Emily Tsang Background Transcription factors
A Supervised Learning Approach to the Prediction of Hi-C Data
 A Supervised Learning Approach to the Prediction of Hi-C Data Tyler Derr YUE Lab The Department of Biochemistry & Molecular Biology Institute for Personalized Medicine Pennsylvania State University College
A Supervised Learning Approach to the Prediction of Hi-C Data Tyler Derr YUE Lab The Department of Biochemistry & Molecular Biology Institute for Personalized Medicine Pennsylvania State University College
Large-scale imputation of epigenomic datasets for systematic annotation of diverse human tissues
 Large-scale imputation of epigenomic datasets for systematic annotation of diverse human tissues The MIT Faculty has made this article openly available. Please share how this access benefits you. Your
Large-scale imputation of epigenomic datasets for systematic annotation of diverse human tissues The MIT Faculty has made this article openly available. Please share how this access benefits you. Your
Integrative Genomics 3b. Systems Biology and Epigenetics
 2018 Integrative Genomics 3b. Systems Biology and Epigenetics ggibson.gt@gmail.com http://www.gibsongroup.biology.gatech.edu Content of the Lecture 1. Immuno-Transcriptomics 2. Epigenome Projects from
2018 Integrative Genomics 3b. Systems Biology and Epigenetics ggibson.gt@gmail.com http://www.gibsongroup.biology.gatech.edu Content of the Lecture 1. Immuno-Transcriptomics 2. Epigenome Projects from
Nature Genetics: doi: /ng Supplementary Figure 1. Analysis of DHS STARR-seq in the P5424 cell line.
 Supplementary Figure 1 Analysis of DHS STARR-seq in the P5424 cell line. (a) Luciferase enhancer assays of proximal DHSs defined as active or inactive enhancers by STARR-seq in P5424 cells. For each candidate,
Supplementary Figure 1 Analysis of DHS STARR-seq in the P5424 cell line. (a) Luciferase enhancer assays of proximal DHSs defined as active or inactive enhancers by STARR-seq in P5424 cells. For each candidate,
Lecture 5: Regulation
 Machine Learning in Computational Biology CSC 2431 Lecture 5: Regulation Instructor: Anna Goldenberg Central Dogma of Biology Transcription DNA RNA protein Process of producing RNA from DNA Constitutive
Machine Learning in Computational Biology CSC 2431 Lecture 5: Regulation Instructor: Anna Goldenberg Central Dogma of Biology Transcription DNA RNA protein Process of producing RNA from DNA Constitutive
Perm-seq: Mapping Protein-DNA Interactions in Segmental Duplication and Highly Repetitive Regions of Genomes with Prior- Enhanced Read Mapping
 RESEARCH ARTICLE Perm-seq: Mapping Protein-DNA Interactions in Segmental Duplication and Highly Repetitive Regions of Genomes with Prior- Enhanced Read Mapping Xin Zeng 1,BoLi 2, Rene Welch 1, Constanza
RESEARCH ARTICLE Perm-seq: Mapping Protein-DNA Interactions in Segmental Duplication and Highly Repetitive Regions of Genomes with Prior- Enhanced Read Mapping Xin Zeng 1,BoLi 2, Rene Welch 1, Constanza
a previous report, Lee and colleagues showed that Oct4 and Sox2 bind to DXPas34 and Xite
 SUPPLEMENTARY INFORMATION doi:1.138/nature9496 a 1 Relative RNA levels D12 female ES cells 24h Control sirna 24h sirna Oct4 5 b,4 Oct4 Xist (wt) Xist (mut) Tsix 3' (wt) Tsix 5' (wt) Beads Oct4,2 c,2 Xist
SUPPLEMENTARY INFORMATION doi:1.138/nature9496 a 1 Relative RNA levels D12 female ES cells 24h Control sirna 24h sirna Oct4 5 b,4 Oct4 Xist (wt) Xist (mut) Tsix 3' (wt) Tsix 5' (wt) Beads Oct4,2 c,2 Xist
Introduction to the UCSC genome browser
 Introduction to the UCSC genome browser Dominik Beck NHMRC Peter Doherty and CINSW ECR Fellow, Senior Lecturer Lowy Cancer Research Centre, UNSW and Centre for Health Technology, UTS SYDNEY NSW AUSTRALIA
Introduction to the UCSC genome browser Dominik Beck NHMRC Peter Doherty and CINSW ECR Fellow, Senior Lecturer Lowy Cancer Research Centre, UNSW and Centre for Health Technology, UTS SYDNEY NSW AUSTRALIA
Machine learning applications in genomics: practical issues & challenges. Yuzhen Ye School of Informatics and Computing, Indiana University
 Machine learning applications in genomics: practical issues & challenges Yuzhen Ye School of Informatics and Computing, Indiana University Reference Machine learning applications in genetics and genomics
Machine learning applications in genomics: practical issues & challenges Yuzhen Ye School of Informatics and Computing, Indiana University Reference Machine learning applications in genetics and genomics
Aaditya Khatri. Abstract
 Abstract In this project, Chimp-chunk 2-7 was annotated. Chimp-chunk 2-7 is an 80 kb region on chromosome 5 of the chimpanzee genome. Analysis with the Mapviewer function using the NCBI non-redundant database
Abstract In this project, Chimp-chunk 2-7 was annotated. Chimp-chunk 2-7 is an 80 kb region on chromosome 5 of the chimpanzee genome. Analysis with the Mapviewer function using the NCBI non-redundant database
Reference genomes and common file formats
 Reference genomes and common file formats Overview Reference genomes and GRC Fasta and FastQ (unaligned sequences) SAM/BAM (aligned sequences) Summarized genomic features BED (genomic intervals) GFF/GTF
Reference genomes and common file formats Overview Reference genomes and GRC Fasta and FastQ (unaligned sequences) SAM/BAM (aligned sequences) Summarized genomic features BED (genomic intervals) GFF/GTF
Multi-scale Deep Tensor Factorization Learns a Latent Representation of the Human Epigenome
 Multi-scale Deep Tensor Factorization Learns a Latent Representation of the Human Epigenome Jacob Schreiber Paul G. Allen School of Computer Science University of Washington jmschreiber91 @jmschrei @jmschreiber91
Multi-scale Deep Tensor Factorization Learns a Latent Representation of the Human Epigenome Jacob Schreiber Paul G. Allen School of Computer Science University of Washington jmschreiber91 @jmschrei @jmschreiber91
Reference genomes and common file formats
 Reference genomes and common file formats Dóra Bihary MRC Cancer Unit, University of Cambridge CRUK Functional Genomics Workshop September 2017 Overview Reference genomes and GRC Fasta and FastQ (unaligned
Reference genomes and common file formats Dóra Bihary MRC Cancer Unit, University of Cambridge CRUK Functional Genomics Workshop September 2017 Overview Reference genomes and GRC Fasta and FastQ (unaligned
Measuring Protein-DNA interactions
 Measuring Protein-DNA interactions How is Biological Complexity Achieved? Mediated by Transcription Factors (TFs) 2 Transcription Factors are genetic switches 3 Regulation of Gene Expression by Transcription
Measuring Protein-DNA interactions How is Biological Complexity Achieved? Mediated by Transcription Factors (TFs) 2 Transcription Factors are genetic switches 3 Regulation of Gene Expression by Transcription
Charles Girardot, Furlong Lab. MACS, CisGenome, SISSRs and other peak calling algorithms: differences and practical use
 Charles Girardot, Furlong Lab MACS, CisGenome, SISSRs and other peak calling algorithms: differences and practical use ChIP-Seq signal properties Only 5 ends of ChIPed fragments are sequenced Shifted read
Charles Girardot, Furlong Lab MACS, CisGenome, SISSRs and other peak calling algorithms: differences and practical use ChIP-Seq signal properties Only 5 ends of ChIPed fragments are sequenced Shifted read
In silico identification of transcriptional regulatory regions
 In silico identification of transcriptional regulatory regions Martti Tolvanen, IMT Bioinformatics, University of Tampere Eija Korpelainen, CSC Jarno Tuimala, CSC Introduction (Eija) Program Retrieval
In silico identification of transcriptional regulatory regions Martti Tolvanen, IMT Bioinformatics, University of Tampere Eija Korpelainen, CSC Jarno Tuimala, CSC Introduction (Eija) Program Retrieval
The University of California, Santa Cruz (UCSC) Genome Browser
 The University of California, Santa Cruz (UCSC) Genome Browser There are hundreds of available userselected tracks in categories such as mapping and sequencing, phenotype and disease associations, genes,
The University of California, Santa Cruz (UCSC) Genome Browser There are hundreds of available userselected tracks in categories such as mapping and sequencing, phenotype and disease associations, genes,
Defining a Registry of Candidate Regulatory Elements to Interpret Disease Associated Genetic Variation
 University of Massachusetts Medical School escholarship@umms GSBS Dissertations and Theses Graduate School of Biomedical Sciences 10-10-2017 Defining a Registry of Candidate Regulatory Elements to Interpret
University of Massachusetts Medical School escholarship@umms GSBS Dissertations and Theses Graduate School of Biomedical Sciences 10-10-2017 Defining a Registry of Candidate Regulatory Elements to Interpret
PRIMEGENSw3 User Manual
 PRIMEGENSw3 User Manual PRIMEGENSw3 is Web Server version of PRIMEGENS program to automate highthroughput primer and probe design. It provides three separate utilities to select targeted regions of interests
PRIMEGENSw3 User Manual PRIMEGENSw3 is Web Server version of PRIMEGENS program to automate highthroughput primer and probe design. It provides three separate utilities to select targeted regions of interests
Gene expression analysis. Biosciences 741: Genomics Fall, 2013 Week 5. Gene expression analysis
 Gene expression analysis Biosciences 741: Genomics Fall, 2013 Week 5 Gene expression analysis From EST clusters to spotted cdna microarrays Long vs. short oligonucleotide microarrays vs. RT-PCR Methods
Gene expression analysis Biosciences 741: Genomics Fall, 2013 Week 5 Gene expression analysis From EST clusters to spotted cdna microarrays Long vs. short oligonucleotide microarrays vs. RT-PCR Methods
Core vertebrate long range cis-regulatory interactions revealed by zebrafish human comparative genomics
 Core vertebrate long range cis-regulatory interactions revealed by zebrafish human comparative genomics Yves Clément & Hugues Roest Crollius GTGC Lyon 01/07/2016 Enhancers as long range cis-regulatory
Core vertebrate long range cis-regulatory interactions revealed by zebrafish human comparative genomics Yves Clément & Hugues Roest Crollius GTGC Lyon 01/07/2016 Enhancers as long range cis-regulatory
Result Tables The Result Table, which indicates chromosomal positions and annotated gene names, promoter regions and CpG islands, is the best way for
 Result Tables The Result Table, which indicates chromosomal positions and annotated gene names, promoter regions and CpG islands, is the best way for you to discover methylation changes at specific genomic
Result Tables The Result Table, which indicates chromosomal positions and annotated gene names, promoter regions and CpG islands, is the best way for you to discover methylation changes at specific genomic
ChIP-Seq Data Analysis. J Fass UCD Genome Center Bioinformatics Core Wednesday December 17, 2014
 ChIP-Seq Data Analysis J Fass UCD Genome Center Bioinformatics Core Wednesday December 17, 2014 What s the Question? Where do Transcription Factors (TFs) bind genomic DNA 1? (Where do other things bind
ChIP-Seq Data Analysis J Fass UCD Genome Center Bioinformatics Core Wednesday December 17, 2014 What s the Question? Where do Transcription Factors (TFs) bind genomic DNA 1? (Where do other things bind
Integrating diverse datasets improves developmental enhancer prediction
 1 Integrating diverse datasets improves developmental enhancer prediction Genevieve D. Erwin 1, Rebecca M. Truty 1, Dennis Kostka 2, Katherine S. Pollard 1,3, John A. Capra 4 1 Gladstone Institute of Cardiovascular
1 Integrating diverse datasets improves developmental enhancer prediction Genevieve D. Erwin 1, Rebecca M. Truty 1, Dennis Kostka 2, Katherine S. Pollard 1,3, John A. Capra 4 1 Gladstone Institute of Cardiovascular
Rapid Transcriptome Characterization for a nonmodel organism using 454 pyrosequencing
 Rapid Transcriptome Characterization for a nonmodel organism using 454 pyrosequencing "#$%&'()*+,"(-*."#$%&/.,"*01*0.,(%-*.&0("2*01*3,$,45,"-*4#66&*71** 3"#)(82,"-*2&9:)($*)1*"(03&"2-*#)66(*.(8$6#*;
Rapid Transcriptome Characterization for a nonmodel organism using 454 pyrosequencing "#$%&'()*+,"(-*."#$%&/.,"*01*0.,(%-*.&0("2*01*3,$,45,"-*4#66&*71** 3"#)(82,"-*2&9:)($*)1*"(03&"2-*#)66(*.(8$6#*;
Escape from epigenetic silencing of lactase gene expression triggered by a single nucleotide change
 Escape from epigenetic silencing of lactase gene expression triggered by a single nucleotide change Dallas M. Swallow and Jesper T. Troelsen The importance of subtle gene regulation and epigenetics in
Escape from epigenetic silencing of lactase gene expression triggered by a single nucleotide change Dallas M. Swallow and Jesper T. Troelsen The importance of subtle gene regulation and epigenetics in
MAKING WHOLE GENOME ALIGNMENTS USABLE FOR BIOLOGISTS. EXAMPLES AND SAMPLE ANALYSES.
 MAKING WHOLE GENOME ALIGNMENTS USABLE FOR BIOLOGISTS. EXAMPLES AND SAMPLE ANALYSES. Table of Contents Examples 1 Sample Analyses 5 Examples: Introduction to Examples While these examples can be followed
MAKING WHOLE GENOME ALIGNMENTS USABLE FOR BIOLOGISTS. EXAMPLES AND SAMPLE ANALYSES. Table of Contents Examples 1 Sample Analyses 5 Examples: Introduction to Examples While these examples can be followed
Genome 373: Mapping Short Sequence Reads II. Doug Fowler
 Genome 373: Mapping Short Sequence Reads II Doug Fowler The final Will be in this room on June 6 th at 8:30a Will be focused on the second half of the course, but will include material from the first half
Genome 373: Mapping Short Sequence Reads II Doug Fowler The final Will be in this room on June 6 th at 8:30a Will be focused on the second half of the course, but will include material from the first half
Human Genetic Variation. Ricardo Lebrón Dpto. Genética UGR
 Human Genetic Variation Ricardo Lebrón rlebron@ugr.es Dpto. Genética UGR What is Genetic Variation? Origins of Genetic Variation Genetic Variation is the difference in DNA sequences between individuals.
Human Genetic Variation Ricardo Lebrón rlebron@ugr.es Dpto. Genética UGR What is Genetic Variation? Origins of Genetic Variation Genetic Variation is the difference in DNA sequences between individuals.
DOUBLE-STRAND DNA BREAK PREDICTION USING EPIGENOME MARKS AT KILOBASE RESOLUTION
 DOUBLE-STRAND DNA BREAK PREDICTION USING EPIGENOME MARKS AT KILOBASE RESOLUTION Raphaël MOURAD, Assist. Prof. Centre de Biologie Intégrative Université Paul Sabatier, Toulouse III INTRODUCTION Double-strand
DOUBLE-STRAND DNA BREAK PREDICTION USING EPIGENOME MARKS AT KILOBASE RESOLUTION Raphaël MOURAD, Assist. Prof. Centre de Biologie Intégrative Université Paul Sabatier, Toulouse III INTRODUCTION Double-strand
Experimental validation of candidates of tissuespecific and CpG-island-mediated alternative polyadenylation in mouse
 Karin Fleischhanderl; Martina Fondi Experimental validation of candidates of tissuespecific and CpG-island-mediated alternative polyadenylation in mouse 108 - Biotechnologie Abstract --- Keywords: Alternative
Karin Fleischhanderl; Martina Fondi Experimental validation of candidates of tissuespecific and CpG-island-mediated alternative polyadenylation in mouse 108 - Biotechnologie Abstract --- Keywords: Alternative
CS273B: Deep Learning in Genomics and Biomedicine. Recitation 1 30/9/2016
 CS273B: Deep Learning in Genomics and Biomedicine. Recitation 1 30/9/2016 Topics Genetic variation Population structure Linkage disequilibrium Natural disease variants Genome Wide Association Studies Gene
CS273B: Deep Learning in Genomics and Biomedicine. Recitation 1 30/9/2016 Topics Genetic variation Population structure Linkage disequilibrium Natural disease variants Genome Wide Association Studies Gene
