Association of Nosema ceranae Infection with Honey Bee (Apis meillfera) Exposome Profiles and Affected Biological Pathways
|
|
|
- Octavia Craig
- 6 years ago
- Views:
Transcription
1 Application Note Exposomics, Genomics, Bioinformatics Association of Nosema ceranae Infection with Honey Bee (Apis meillfera) Exposome Profiles and Affected Biological Pathways Authors Anthony Macherone 1,2, Robert L. Broadrup 3, Christopher Mayack 4, and Helen K. White 3 1 Life Science and Chemical Analysis Group, Agilent Technologies, Santa Clara, CA Department of Biological Chemistry, Johns Hopkins University School of Medicine, Baltimore, MD USA Abstract Foraging honey bee samples representative of 30 individual hives were collected simultaneously at the end of the 2015 foraging season. Polymerase chain reaction was used to determine if the hives were infected with Nosema ceranae a common honey bee parasite. A second sample of foraging bees was collected from each hive at the same time for discovery based exposomics using the Agilent 7890B GC combined with an Agilent 7200B GC/Q TOF system. Significance testing and biological pathway analyses were conducted using Agilent MassProfiler Professional and Pathway Architect software. Significance testing determined that exposome profiles were associated with N. ceranae infection. This study illustrates how the integration of exposomics with genetic disease profiling yields a powerful multi omics platform for discovery and generation of new hypotheses for targeted follow up studies. 3 Department of Chemistry, Haverford College, Haverford, PA USA 4 Department of Biology, Swarthmore College, Swarthmore, PA USA
2 Introduction Over the past decade, the western honey bee (Apis mellifera) population has succumbed to a host of environmental stressors including exposure to parasites, persistent organic pollutants, poor nutrition, and loss of habitat. These stressors represent environmental exposures that adversely affect the health of bees at the hive level. Current honey bee research offers insight into the reasons for the global health decline of honey bees, but this complex combination of stressors has made it difficult to pinpoint key features of disease causality. The exposome represents cumulative exposures over individuals lifetime. It includes, but is not limited to, environmental pollutants, diet, drugs, and the consequent alterations of internal biochemistry. The exposome paradigm associates exposures with biological response pathways, and offers a mechanism to investigate the nongenetic causative factors of chronic disease. To address the highly complex problem of honey bee population decline, we hypothesized that integrating exposome chemical profiles with semiquantitative PCR (sq-pcr), and analyzing the resulting multi-omics dataset with bioinformatics tools may identify chemical profiles and dysregulation of biological pathways statistically associated (p <0.05) with disease and eventual collapse of a bee colony. Experimental Sample Collection Biological samples were collected from 30 noncommercial apiary hives in seven geographical locations located in southeastern Pennsylvania. Sixty to one hundred bees were collected from each hive in 50 ml disposable FalconTM tubes, and immediately frozen in dry ice. All samples were stored in a 80 C freezer prior to analysis. Sample extraction Samples for mass spectrometry were extracted by pulverizing 3 g of bees in 27 ml of water/acetonitrile/acetic acid (44 %:55 %:1 %). A modified QuEChERS extraction was performed as follows: to each sample was added 250 mg carbon (p/n ), 500 primary secondary amine (PSA, p/n ), 6 g of magnesium sulfate (p/n ), and 1.5 g sodium acetate (p/n ). The tubes were sealed, shaken and centrifuged at 3,000 rpm for 5 minutes. A 2-mL aliquot of the supernatant from each sample was applied to a conditioned SPE cartridge containing 250 mg carbon (p/n ). Analytes were eluted from the SPE cartridge with 4 ml acetone/toluene (70:30 v/v) and reduced in volume using a gentle stream of nitrogen. Approximately 2 ml of each concentrated sample was transferred to 2-mL autosampler vials prior to GC-TOF analysis. DNA Extraction and Semiquantitative PCR Samples for PCR screening were prepared by adding 6 ml of RNase free water to 30 bees. The bees were macerated with a tissue grinder to thoroughly homogenize the contents. To semiquantify Nosema ceranae, a PCR reaction with a RpS5 reference gene was performed based on methods adapted from the HBRC method1. Briefly, a 150-µL aliquot of the homogenate was added to 300 µl of a 1:1 mixture of phenol/chloroform. The solution was centrifuged at 13,000 rpm for 5 minutes, and the supernatant was added to another 300 µl of the 1:1 phenol/chloroform solution in a 1.5-mL microcentrifuge tube. The supernatant was drawn once again and transferred to a 1.5 ml microcentrifuge tube. This was followed by the addition of 30 µl sodium acetate and 600 µl of 95 % ethanol to precipitate the DNA overnight at 20 C. A PCR reaction combined 1 µl of a 10 mm solution of each primer, 1.5 µl of 10x PCR buffer, 0.5 µl of 10 mm deoxynucleotide triphosphate (dntp), 0.2 µl of a 25 mm magnesium chloride solution, 0.2 µl of 5 U/µL Taq DNA polymerase (New England BioLabs, Ipswich, MA), and 2 µl of template DNA. The PCR thermocycler program was 94 C for 2.5 minutes, followed by 10 cycles of: 15 seconds at 94 C, 30 seconds at 61.8 C, and 45 seconds at 72 C. Then, 20 cycles of: 15 seconds at 94 C, 30 seconds at 61.8 C, and 50 seconds at 72 C, followed by an extension step at 72 C for 7 minutes, and a final hold step of 4 C. PCR products were identified by a 3 % agarose gel using electrophoresis, and confirmed using the Agilent 2100 Bioanalyzer. Open-source ImageJ was used to semiquantify N. ceranae based on product band intensity from the gel electrophoresis 2. 2
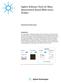
3 GC-TOF Mass Spectrometry An Agilent 7890B GC combined with an Agilent 7200B GC/Q-TOF system was used for discovery-based chemical profiling of bee exposomes. The GC was configured with a 40 m 0.25 mm, 0.25 µm DB5-MS DuraGuard column (Agilent J&W G) operated at 1.2 ml/minute helium in constant flow mode. A 0.2 µl pulsed splitless injection was made into a 250 C isothermal split/splitless inlet. The oven program was 80 C (1 minute), then 10 C/min to 310 C (6 minutes). The transfer line temperature was 300 C. The mass spectrometer was operated in electron ionization, high resolution TOF mode. The source and quadrupole (RF only) temperatures were 275 C and 150 C, respectively. High resolution, accurate mass (HRAM) spectral information was collected at 5 Hz over a mass range of 50 Da to 800 Da. Data Analysis To extract chemical features from the raw mass spectra, chromatographic deconvolution was performed using Agilent MassHunter Unknowns Analysis B software package. Briefly, chemical features were identified if the calculated signal-to-noise of a chromatographic peak was >3:1, and at least three extracted ion chromatograms could be aligned based on peak shape and retention time. Each identified feature was tested against a known commercial spectral library (NIST11) 3 for mass, number of ions, and ion ratio similarities. If a match score >0.7 was determined, the feature was annotated with chemical names, CAS number, and other pertinent information. Nonannotated chemical features were excluded from statistical analyses. Instrument Conditions Column Gas flow Injection Parameter Inlet temperature Oven program Value Transfer line temperature 300 C MS Mode Source temperature 275 C Quadrupole temperature Statistical Significance Testing and Pathway Analyses The multi-omic dataset was statistically analyzed using Agilent MassProfiler Professional (MPP) and Pathway Architect software. The variables included N. ceranae Load (ranging from 0.00 to 40.02), and Phenotype (healthy, unhealthy, collapsed, unknown) determined by organoleptic observation at the time of sample collection. The data were aligned in MPP, allowing for a retention time variation of 0.15 minutes and a mass extraction window of ±20 ppm. All raw data were Log 2 transformed. The baseline of the data was determined using the baseline to the median of all log-transformed response values across all samples. Significance testing was performed using built-in features of MPP, first Filter Agilent J&W DB5-MS DuraGuard, 40 m 0.25 mm, 0.25 μm ( G) Helium, 1.2 ml/min, constant flow mode 0.2 μl pulsed splitless 250 C, isothermal split/splitless inlet 80 C (1 minute), then 10 C/min to 310 C (6 minutes) EI, high res TOF 150 C, RF only by Flags, wherein chemical entities were retained if they were present in at least two samples, followed by Filter by Frequency to retain chemical entities that appear in 100 % of samples. One-way ANOVA of the reduced dataset yielded 20 chemical entities with a p-value <0.05 and a fold-change (distance from median value) >2. These were screened against known Apis mellifera biological pathways downloaded from the Kyoto Encyclopedia of Genes and Genomes (KEGG) database 4,5 using Agilent Pathway Architect tools. Figure 1 illustrates how the N. ceranae data are connected to deconvoluted mass spectra and analyzed in MPP to identify chemical entities, associations, and the biological pathways affected in the disease state. 3
4 Samples and Variables Sample Average Nosema load Hive health Unknown Unknown Healthy Healthy Unknown Unknown Healthy Collapsed Collapsed Healthy Unknown Unknown Unknown Healthy Unknown Sample Average Nosema load Hive health Unknown Healthy Healthy Collapsed Healthy Healthy Healthy Unknown Unknown Collapsed Unknown Healthy Collapsed Unhealthy Unknown Counts GC-TOF Acquisition time (min) MH Deconvolution MPP/PA Figure 1. Connecting multi-omic data in Agilent MPP. N. ceranae load and hive health conditions for each sample are connected to the corresponding deconvoluted mass spectral data and chemical entity annotations in MPP. Quality and statistical testing is performed within the MPP environment to identify chemicals significantly associated with infection and hive health. Pathway Architect features are used to interrogate known honey bee biological pathways with the chemical entity data subset. Dysregulated chemical entities within each pathway are determined and saved for interpretation by the investigator. 4
5 Results and Discussion The N. ceranae load data were approximately log-normal distributed. Significance testing identified multiple chemical biomarkers that may affect honey bee health. Many of the annotated chemicals were associated with known biological pathways of the western honey bee including phenylalanine metabolism, caffeine metabolism, fatty acid biosynthesis, and nicotinate and nicotinamide metabolism. For example, in the Ubiquinone and Other Terpenoid Quinone Biosynthesis Pathway, a putative direct relationship between N. ceranae load and β-tocopherol and γ-tocopherol was identified. Therein, lower N. ceranae loads align with a down-regulation of these tocopherols, and higher loads align with increased tocopherol abundance (Table 1). In the Nicotinate and Nicotinamide Pathway, it was determined that nicotinamide levels in unhealthy hives was significantly upregulated. Many of the associated chemicals were found at relative concentration levels (represented by ion abundance) 10-fold higher than the median value over all samples (Figure 2). When comparing healthy to unhealthy hives, niacinamide, and semioxamazide (oxalic acid, a common treatment for diseased hives) were significantly associated (p << ) with N. ceranae infection. Other chemicals moderately associated with N. ceranae infection when comparing healthy to unhealthy hives include: 1-methyl-1H-imidazole (p = ), a histidine and histamine mimetic; 1 -tridecyn-4-ol (p = ), a 13-carbon alcohol potentially associated with the chemical communication of the bees; 5-hydroxy-4,7-dimethoxyflavinone (p = ), derived from flower pollen; and tris(2,4-di-tert-butylphenyl)phosphate (p = 0.021), a known plasticizer. Table 1. Ubiquinone and other terpenoid-quinone biosynthesis. β-tocopherol and γ-tocopherol are downregulated (blue) in hives with very low to no N. ceranae infection as determined by sq-pcr, while hives with higher levels of infection show upregulated levels of these tocopherols (red). Log 10 p Entity Averaged Nosema ceranae Load (Log 2 transformed) β-tocopherol γ-tocopherol Collapsed versus healthy Unhealthy versus healthy Unknown versus healthy Oxalic acid Niacinamide ±2 fold change Log 10 (fold change) p = Figure 2. Pseudo-Manhattan plot. Log 2 (transformed p values) versus log(transformed fold-change) for collapsed versus healthy, unhealthy versus healthy, and unknown versus healthy hives. Vertical dotted lines illustrate ±2 log 2 normalized relative concentration fold-change. The horizontal line represents the log 2 (normalized α) for p = (0.1 % occurs by chance). Significance testing identified a chemical entity list of 20 chemicals significantly (p<0.05) associated with N. ceranae infection. When comparing this list in the collapsed, unhealthy, or unknown hive health against healthy hives, most of the chemical entities in the collapsed and unknown hives fall within the ±2-fold change window. However, comparing unhealthy hives health against healthy hives, most entities have >10-fold change, with statistical significance (p <0.001). 5
6 Conclusion The initial results from this study tend to confirm the hypotheses that as parasite infestation increases, there is a measurable change in the exposome chemical profile. The identified chemicals can be associated with known biological pathways of the western honey bees. We acknowledge that the small sample size of the pilot study may not elicit the appropriate effect needed to detect more subtle associations with changes in honey bee health. We caution that these results must be confirmed with more extensive longitudinal sampling and follow-up targeted hypothesis-driven studies. References 1. Hamiduzzaman, M. M.; Guzman Novoa, E.; Goodwin, P. H. A multiplex PCR assay to diagnose and quantify Nosema infections in honey bees (Apis mellifera). J. Invertebr. Pathol. 2010, 105, Schneider, C. A.; Rasband, W. S.; Eliceiri, K. W. Nature Methods 2012, 9, 7, National Institute of Standards and Technology, NIST Standard Reference Database 1A. Retrieved February 17, 2013, from NIST.gov: gov/srd/nist1a. cfm# Kanehisa, M.; et al. Nucleic Acids Res. 2016, 44, D457-D Kanehisa, M.; Goto, S. Nucleic Acids Res. 2000, 28, For Research Use Only. Not for use in diagnostic procedures. This information is subject to change without notice. Agilent Technologies, Inc Printed in the USA, September 21, EN
The Use of High Resolution Accurate Mass GC/Q-TOF and Chemometrics in the Identification of Environmental Pollutants in Wastewater Effluents
 The Use of High Resolution Accurate Mass GC/Q-TOF and Chemometrics in the Identification of Environmental Pollutants in Wastewater s Application Note Environmental Authors Anthony Gravell and Praveen Kutty
The Use of High Resolution Accurate Mass GC/Q-TOF and Chemometrics in the Identification of Environmental Pollutants in Wastewater s Application Note Environmental Authors Anthony Gravell and Praveen Kutty
Metabolic Changes in Lung Tissue of Tuberculosis-Infected Mice Using GC/Q-TOF with Low Energy EI
 Metabolic Changes in Lung Tissue of Tuberculosis-Infected Mice Using GC/Q-TOF with Low Energy EI Application Brief Authors Mª Fernanda Rey-Stolle 1, Vineel P. Reddy 2, Santiago Angulo 1, Adrie J.C. Steyn
Metabolic Changes in Lung Tissue of Tuberculosis-Infected Mice Using GC/Q-TOF with Low Energy EI Application Brief Authors Mª Fernanda Rey-Stolle 1, Vineel P. Reddy 2, Santiago Angulo 1, Adrie J.C. Steyn
Metabolomics of Carbon-Fixing Mutants of Cyanobacteria by GC/Q-TOF
 Metabolomics of Carbon-Fixing Mutants of Cyanobacteria by GC/Q-TOF Application Note Biofuel Authors Dong Hee Chung, Christine Rabinovitch-Deere, and Shota Atsumi Department of Chemistry University of California-Davis,
Metabolomics of Carbon-Fixing Mutants of Cyanobacteria by GC/Q-TOF Application Note Biofuel Authors Dong Hee Chung, Christine Rabinovitch-Deere, and Shota Atsumi Department of Chemistry University of California-Davis,
Using the Blood Exposome to Discover Causes of Disease
 Using the Blood Exposome to Discover Causes of Disease Technical Overview Clinical Research Authors Stephen M. Rappaport, Ph.D. Center for Exposure Biology University of California, Berkeley Anthony Macherone,
Using the Blood Exposome to Discover Causes of Disease Technical Overview Clinical Research Authors Stephen M. Rappaport, Ph.D. Center for Exposure Biology University of California, Berkeley Anthony Macherone,
Expand Your Research with Metabolomics and Proteomics. Christine Miller Omics Market Manager ASMS 2017
 Expand Your Research with Metabolomics and Proteomics Christine Miller Omics Market Manager ASMS 2017 New Additions to Agilent Omics Workflows Acquisition Ion Mobility Q-TOF IM All Ions MS/MS Find Features
Expand Your Research with Metabolomics and Proteomics Christine Miller Omics Market Manager ASMS 2017 New Additions to Agilent Omics Workflows Acquisition Ion Mobility Q-TOF IM All Ions MS/MS Find Features
EVALUATION OF D DIFFERENT EXTRACTION METHODS OF PESTICIDE RESIDUES IN SPICES BY
 EVALUATION OF DIFFERENT EXTRACTION METHODS OF PESTICIDE RESIDU UES IN SPICES BY LC-ToF-MS and GC-ToF-MS 1 Content 1. Aim and scope... 3 2. Short description... 3 3. 4. 5. 6. Consumables... 3 Chemicals...
EVALUATION OF DIFFERENT EXTRACTION METHODS OF PESTICIDE RESIDU UES IN SPICES BY LC-ToF-MS and GC-ToF-MS 1 Content 1. Aim and scope... 3 2. Short description... 3 3. 4. 5. 6. Consumables... 3 Chemicals...
Analysis of 16 Pesticide Residues in Broccoli Using CarbonX dspe QuEChERS AOAC Kits Using GC/MS
 Analysis of 16 Pesticide Residues in Broccoli Using CarbonX dspe QuEChERS AOAC Kits Using GC/MS Authors: Conor Smith, Doug Fryer United Science Corp. 15911 Furuby Rd. Center City, MN 55012 USA Abstract
Analysis of 16 Pesticide Residues in Broccoli Using CarbonX dspe QuEChERS AOAC Kits Using GC/MS Authors: Conor Smith, Doug Fryer United Science Corp. 15911 Furuby Rd. Center City, MN 55012 USA Abstract
The Agilent Metabolomics Dynamic MRM Database and Method
 The Agilent Metabolomics Dynamic MRM Database and Method White Paper Author Mark Sartain Agilent Technologies, Inc. Santa Clara, California, USA Introduction A major challenge in metabolomics is achieving
The Agilent Metabolomics Dynamic MRM Database and Method White Paper Author Mark Sartain Agilent Technologies, Inc. Santa Clara, California, USA Introduction A major challenge in metabolomics is achieving
Reduced carryover and increased reproducibility in the analysis of cold pressed pink grapefruit essential oil
 Application Note Food Testing and Agriculture Reduced carryover and increased reproducibility in the analysis of cold pressed pink grapefruit essential oil Using an Agilent J&W DB-HeavyWAX GC column Author
Application Note Food Testing and Agriculture Reduced carryover and increased reproducibility in the analysis of cold pressed pink grapefruit essential oil Using an Agilent J&W DB-HeavyWAX GC column Author
Agilent Solutions for Metabolomics YOUR PATH TO SUCCESS
 Agilent Solutions for Metabolomics YOUR PATH TO SUCCESS UNDERSTANDING METABOLOMICS Agilent is the leading global metabolomics vendor, offering our customers a broad array of cutting-edge instrumentation
Agilent Solutions for Metabolomics YOUR PATH TO SUCCESS UNDERSTANDING METABOLOMICS Agilent is the leading global metabolomics vendor, offering our customers a broad array of cutting-edge instrumentation
A Highly Accurate Mass Profiling Approach to Protein Biomarker Discovery Using HPLC-Chip/ MS-Enabled ESI-TOF MS
 Application Note PROTEOMICS METABOLOMICS GENOMICS INFORMATICS GLYILEVALCYSGLUGLNALASERLEUASPARG CYSVALLYSPROLYSPHETYRTHRLEUHISLYS A Highly Accurate Mass Profiling Approach to Protein Biomarker Discovery
Application Note PROTEOMICS METABOLOMICS GENOMICS INFORMATICS GLYILEVALCYSGLUGLNALASERLEUASPARG CYSVALLYSPROLYSPHETYRTHRLEUHISLYS A Highly Accurate Mass Profiling Approach to Protein Biomarker Discovery
A Rapid Method for Trace Analysis of Organophosphorus Pesticides in Drinking Water
 A Rapid Method for Trace Analysis of Organophosphorus Pesticides in Drinking Water Application Note Environmental Authors Min Cai and Yun Zou Agilent Technologies Co. Ltd, 412 Ying Lun Road Waigaoqiao
A Rapid Method for Trace Analysis of Organophosphorus Pesticides in Drinking Water Application Note Environmental Authors Min Cai and Yun Zou Agilent Technologies Co. Ltd, 412 Ying Lun Road Waigaoqiao
Agilent Solutions for Metabolomics EMERGING INSIGHTS
 Agilent Solutions for Metabolomics EMERGING INSIGHTS Understanding Metabolic Fingerprints A Powerful Way to Investigate Biology What is Metabolomics? Metabolomics is the study of the metabolite content
Agilent Solutions for Metabolomics EMERGING INSIGHTS Understanding Metabolic Fingerprints A Powerful Way to Investigate Biology What is Metabolomics? Metabolomics is the study of the metabolite content
Accurately Identify and Quantify One Hundred Pesticides in a Single GC Run
 Accurately Identify and Quantify One Hundred Pesticides in a Single GC Run Application Note Author Jessica Westland Agilent Technologies, Inc. Abstract A selected target compound list of 195 various pesticides
Accurately Identify and Quantify One Hundred Pesticides in a Single GC Run Application Note Author Jessica Westland Agilent Technologies, Inc. Abstract A selected target compound list of 195 various pesticides
Nitrosamines Analysis in Drinking Water Using GC/MS/MS Meeting Equivalence to EPA Method 521
 Application Note Environmental Nitrosamines Analysis in Drinking Water Using GC/MS/MS Meeting Equivalence to EPA Method 521 Using Agilent 71 and 7 triple quadrupole GC/MS systems Authors Andy Eaton, Charles
Application Note Environmental Nitrosamines Analysis in Drinking Water Using GC/MS/MS Meeting Equivalence to EPA Method 521 Using Agilent 71 and 7 triple quadrupole GC/MS systems Authors Andy Eaton, Charles
New Diagnostic Method for Pancreatic Cancer by Profiling of Metabolites in Serum Using GC/MS
 C146-E 6-E156 New Diagnostic Method for Pancreatic Cancer by Profiling of Metabolites in Serum Using GC/MS Dr. Masaru Yoshida, Division of Metabolomics Research, Division of Gastroenterology, The Integrated
C146-E 6-E156 New Diagnostic Method for Pancreatic Cancer by Profiling of Metabolites in Serum Using GC/MS Dr. Masaru Yoshida, Division of Metabolomics Research, Division of Gastroenterology, The Integrated
Improving GC-MS Method Robustness and Cycle Times Using Capillary Flow Technology and Backflushing
 Improving GC-MS Method Robustness and Cycle Times Using Capillary Flow Technology and Backflushing Application Note Environmental Author Chris Sandy Agilent Technologies, Inc. UK and Ireland Sales Headquarters
Improving GC-MS Method Robustness and Cycle Times Using Capillary Flow Technology and Backflushing Application Note Environmental Author Chris Sandy Agilent Technologies, Inc. UK and Ireland Sales Headquarters
Multiresidue Pesticide Analysis with the Agilent Intuvo 9000 GC and Agilent 7000 Series Mass Spectrometer
 Multiresidue Pesticide Analysis with the Agilent Intuvo GC and Agilent 7 Series Mass Spectrometer Application Note Author Rebecca Veeneman, PhD and Joan Stevens, PhD Agilent Technologies, Inc. Abstract
Multiresidue Pesticide Analysis with the Agilent Intuvo GC and Agilent 7 Series Mass Spectrometer Application Note Author Rebecca Veeneman, PhD and Joan Stevens, PhD Agilent Technologies, Inc. Abstract
Instrumental Solutions for Metabolomics
 Instrumental Solutions for Metabolomics Dr. Desislav Donchev US14702384-16012196010132-S01-A01.tif 300 400 500 600 700 800 3100 3200 3300 3400 3500 3600 3700 3800 Color Scale R = [52 1484] G = [26 514]
Instrumental Solutions for Metabolomics Dr. Desislav Donchev US14702384-16012196010132-S01-A01.tif 300 400 500 600 700 800 3100 3200 3300 3400 3500 3600 3700 3800 Color Scale R = [52 1484] G = [26 514]
Agilent Consumables Improve Performance with Shimadzu GC/MS
 Agilent Consumables Improve Performance with Shimadzu GC/MS Application Note Environmental & Food Safety Author Ken Lynam Agilent Technologies, Inc. Abstract Leak-free performance over 150 heat cycles
Agilent Consumables Improve Performance with Shimadzu GC/MS Application Note Environmental & Food Safety Author Ken Lynam Agilent Technologies, Inc. Abstract Leak-free performance over 150 heat cycles
Discovery of Imidacloprid Metabolites in Onions Using Mass Profiler Professional
 Discovery of Imidacloprid Metabolites in Onions Using Mass Profiler Professional Application Note Food Authors Imma Ferrer and E. Michael Thurman Center for Environmental Mass Spectrometry Department of
Discovery of Imidacloprid Metabolites in Onions Using Mass Profiler Professional Application Note Food Authors Imma Ferrer and E. Michael Thurman Center for Environmental Mass Spectrometry Department of
Agilent Software Tools for Mass Spectrometry Based Multi-omics Studies
 Agilent Software Tools for Mass Spectrometry Based Multi-omics Studies Technical Overview Introduction The central dogma for biological information flow is expressed as a series of chemical conversions
Agilent Software Tools for Mass Spectrometry Based Multi-omics Studies Technical Overview Introduction The central dogma for biological information flow is expressed as a series of chemical conversions
AGILENT SOLUTIONS FOR METABOLOMICS
 YOUR Path To Succcess AGILENT SOLUTIONS FOR METABOLOMICS Understanding METABOLOMICS Agilent is the leading global solution vendor for measuring metabolism, offering researchers a broad array of cutting-edge
YOUR Path To Succcess AGILENT SOLUTIONS FOR METABOLOMICS Understanding METABOLOMICS Agilent is the leading global solution vendor for measuring metabolism, offering researchers a broad array of cutting-edge
Application of Agilent AdvanceBio Desalting-RP Cartridges for LC/MS Analysis of mabs A One- and Two-dimensional LC/MS Study
 Application of Agilent AdvanceBio Desalting-RP Cartridges for LC/MS Analysis of mabs A One- and Two-dimensional LC/MS Study Application note Biotherapeutics and Biologics Authors Suresh Babu C.V., Anne
Application of Agilent AdvanceBio Desalting-RP Cartridges for LC/MS Analysis of mabs A One- and Two-dimensional LC/MS Study Application note Biotherapeutics and Biologics Authors Suresh Babu C.V., Anne
Analysis of Nucleosides Using an Agilent Infinity II High Speed UHPLC with the 6130 Single Quadrupole Mass Selective Detector
 Analysis of Nucleosides Using an Agilent Infinity II High Speed UHPLC with the 6130 Single Quadrupole Mass Selective Detector Application Note Clinical Research Author Patrick Cronan Agilent Technologies,
Analysis of Nucleosides Using an Agilent Infinity II High Speed UHPLC with the 6130 Single Quadrupole Mass Selective Detector Application Note Clinical Research Author Patrick Cronan Agilent Technologies,
Clone Selection Using the Agilent 1290 Infinity Online 2D-LC/MS Solution
 Clone Selection Using the Agilent 9 Infinity Online D-LC/MS Solution Application Note Biopharmaceuticals Authors Abstract Ravindra Gudihal and This Application Note describes the Agilent 9 Infinity online
Clone Selection Using the Agilent 9 Infinity Online D-LC/MS Solution Application Note Biopharmaceuticals Authors Abstract Ravindra Gudihal and This Application Note describes the Agilent 9 Infinity online
FFPE DNA Extraction Protocol
 FFPE DNA Extraction Protocol Introduction The number of archival formalin-fixed paraffin embedded (FFPE) samples is in the millions, providing an invaluable repository of information for genetic analysis.
FFPE DNA Extraction Protocol Introduction The number of archival formalin-fixed paraffin embedded (FFPE) samples is in the millions, providing an invaluable repository of information for genetic analysis.
AGILENT S BIOINFORMATICS ANALYSIS SOFTWARE
 ACCELERATING PROGRESS IS IN OUR GENES AGILENT S BIOINFORMATICS ANALYSIS SOFTWARE GENESPRING GENE EXPRESSION (GX) MASS PROFILER PROFESSIONAL (MPP) PATHWAY ARCHITECT (PA) See Deeper. Reach Further. BIOINFORMATICS
ACCELERATING PROGRESS IS IN OUR GENES AGILENT S BIOINFORMATICS ANALYSIS SOFTWARE GENESPRING GENE EXPRESSION (GX) MASS PROFILER PROFESSIONAL (MPP) PATHWAY ARCHITECT (PA) See Deeper. Reach Further. BIOINFORMATICS
Low-cost DNA extraction for use in TILLING and Ecotilling assays
 Low-cost DNA extraction for use in TILLING and Ecotilling assays Version 1.3 (January 31, 2013) Plant Breeding and Genetics Laboratory Prepared by Bernhard Hofinger and Bradley Till 1. MATERIALS Company
Low-cost DNA extraction for use in TILLING and Ecotilling assays Version 1.3 (January 31, 2013) Plant Breeding and Genetics Laboratory Prepared by Bernhard Hofinger and Bradley Till 1. MATERIALS Company
METHOD TRANSLATION OF HJ FOR INTUVO
 METHOD TRANSLATION OF HJ679-2013 FOR INTUVO Translating a 530 µm id Column Method to 250 µm or 320 µm column Methods for the Agilent Intuvo 9000 GC Introduction HJ method 679-2013 describes the determination
METHOD TRANSLATION OF HJ679-2013 FOR INTUVO Translating a 530 µm id Column Method to 250 µm or 320 µm column Methods for the Agilent Intuvo 9000 GC Introduction HJ method 679-2013 describes the determination
Benefits of 2D-LC/MS/MS in Pharmaceutical Bioanalytics
 Application Note Small Molecule Pharmaceuticals Benefits of 2D-LC/MS/MS in Pharmaceutical Bioanalytics Avoiding Matrix Effects Increasing Detection Sensitivity Authors Jonas Dinser Daiichi-Sankyo Europe
Application Note Small Molecule Pharmaceuticals Benefits of 2D-LC/MS/MS in Pharmaceutical Bioanalytics Avoiding Matrix Effects Increasing Detection Sensitivity Authors Jonas Dinser Daiichi-Sankyo Europe
Genomic Sequencing. Genomic Sequencing. Maj Gen (R) Suhaib Ahmed, HI (M)
 Maj Gen (R) Suhaib Ahmed, HI (M) The process of determining the sequence of an unknown DNA is called sequencing. There are many approaches for DNA sequencing. In the last couple of decades automated Sanger
Maj Gen (R) Suhaib Ahmed, HI (M) The process of determining the sequence of an unknown DNA is called sequencing. There are many approaches for DNA sequencing. In the last couple of decades automated Sanger
Stool DNA Isolation Kit
 Product Information Norgen s provides a convenient and rapid method to isolate total DNA from fresh or frozen stool samples. The universal protocol conveniently allows for the isolation of total genomic
Product Information Norgen s provides a convenient and rapid method to isolate total DNA from fresh or frozen stool samples. The universal protocol conveniently allows for the isolation of total genomic
The Agilent Total RNA Isolation Kit
 Better RNA purity. Better data. Without DNase treatment. The Agilent Total RNA Isolation Kit Now you can isolate highly purified, intact RNA without DNase treatment Introducing the Agilent Total RNA Isolation
Better RNA purity. Better data. Without DNase treatment. The Agilent Total RNA Isolation Kit Now you can isolate highly purified, intact RNA without DNase treatment Introducing the Agilent Total RNA Isolation
Method Translation in Liquid Chromatography
 Method Translation in Liquid Chromatography Technical Overview Abstract Ronald E. Majors Agilent Technologies, Inc. 2850 Centerville Rd Wilmington, DE 19808 USA With the recent emphasis on high performance
Method Translation in Liquid Chromatography Technical Overview Abstract Ronald E. Majors Agilent Technologies, Inc. 2850 Centerville Rd Wilmington, DE 19808 USA With the recent emphasis on high performance
SuperTCRExpress TM Human TCR Vβ Repertoire CDR3 Diversity Determination (Spectratyping) and Quantitative Analysis Kit
 SuperTCRExpress TM Human TCR Vβ Repertoire CDR3 Diversity Determination (Spectratyping) and Quantitative Analysis Kit Cat. No. H0521 Size: 2 sets (22 Vβ families/each, with enzymes) H0522 Size: 4 sets
SuperTCRExpress TM Human TCR Vβ Repertoire CDR3 Diversity Determination (Spectratyping) and Quantitative Analysis Kit Cat. No. H0521 Size: 2 sets (22 Vβ families/each, with enzymes) H0522 Size: 4 sets
FFPE DNA Extraction kit
 FFPE DNA Extraction kit Instruction Manual FFPE DNA Extraction kit Cat. No. C20000030, Format 50 rxns Version 1 / 14.08.13 Content Introduction 4 Overview of FFPE DNA extraction workflow 4 Kit contents
FFPE DNA Extraction kit Instruction Manual FFPE DNA Extraction kit Cat. No. C20000030, Format 50 rxns Version 1 / 14.08.13 Content Introduction 4 Overview of FFPE DNA extraction workflow 4 Kit contents
Isolation of High-Purity Total Cellular RNA from Muscle Tissues Using the Agilent Total RNA Isolation Mini Kit Application
 Isolation of High-Purity Total Cellular RNA from Muscle Tissues Using the Total RNA Isolation Mini Kit Application Genomics Author Christopher Rizzo Technologies, Inc. 2850 Centerville Road Wilmington,
Isolation of High-Purity Total Cellular RNA from Muscle Tissues Using the Total RNA Isolation Mini Kit Application Genomics Author Christopher Rizzo Technologies, Inc. 2850 Centerville Road Wilmington,
Optimizing Productivity and Reliability for Monocyclic Aromatic Hydrocarbon Purity Analysis According to ASTM D7504 on the Agilent 8890 GC System
 Application Note Petrochemicals Optimizing Productivity and Reliability for Monocyclic Aromatic Hydrocarbon Purity Analysis According to ASTM D50 on the Agilent 0 GC System Authors Jie Pan, Lukas Wieder,
Application Note Petrochemicals Optimizing Productivity and Reliability for Monocyclic Aromatic Hydrocarbon Purity Analysis According to ASTM D50 on the Agilent 0 GC System Authors Jie Pan, Lukas Wieder,
Multi-Residue Pesticide Analysis in Ginseng Powder: Optimized Cleanup After QuEChERS Extraction for UPLC-MS/MS And GC-MS/MS Analysis
 Multi-Residue Pesticide Analysis in Ginseng Powder: Optimized Cleanup After QuEChERS Extraction for UPLC-MS/MS And GC-MS/MS Analysis Michael S. Young, Kim Van Tran, and Jeremy C. Shia Waters Corporation,
Multi-Residue Pesticide Analysis in Ginseng Powder: Optimized Cleanup After QuEChERS Extraction for UPLC-MS/MS And GC-MS/MS Analysis Michael S. Young, Kim Van Tran, and Jeremy C. Shia Waters Corporation,
Validation Data for the Analysis of Chlorotalonil in Leek Using Mini-Luke Method Followed by GC-QqQ-MS/MS
 Validation Data for the Analysis of Chlorotalonil in Leek Using Mini-Luke Method Followed by GC-QqQ-MS/MS CONTENTS 1. Aim and Scope 2 2. Short Description 2 3. Apparatus and Consumables 2 4. Chemicals
Validation Data for the Analysis of Chlorotalonil in Leek Using Mini-Luke Method Followed by GC-QqQ-MS/MS CONTENTS 1. Aim and Scope 2 2. Short Description 2 3. Apparatus and Consumables 2 4. Chemicals
An Examination of the Presence, Formation, and Transformation of Volatile Halogenated Organic Species in Wastewater Extracts Using GC-ICP-MS
 An Examination of the Presence, Formation, and Transformation of Volatile Halogenated Organic Species in Wastewater Extracts Using GC-ICP-MS Application Note Environmental Authors Armando Durazo and Shane
An Examination of the Presence, Formation, and Transformation of Volatile Halogenated Organic Species in Wastewater Extracts Using GC-ICP-MS Application Note Environmental Authors Armando Durazo and Shane
Direct Injection of Fish Oil for the GC-ECD Analysis of PCBs: Results Using a Deans Switch With Backflushing. Application. Author.
 Direct Injection of Fish Oil for the GC-ECD Analysis of PCBs: Results Using a Deans Switch With Backflushing Application Environmental and Pharmaceutical Author Philip L. Wylie Agilent Technologies, Inc.
Direct Injection of Fish Oil for the GC-ECD Analysis of PCBs: Results Using a Deans Switch With Backflushing Application Environmental and Pharmaceutical Author Philip L. Wylie Agilent Technologies, Inc.
Low cost and non-toxic genomic DNA extraction for use in molecular marker studies.
 Low cost and non-toxic genomic DNA extraction for use in molecular marker studies. Version 1.4, February 28 th, 2013. Prepared by Bernhard Hofinger, Owen Huynh and Brad Till. 1. OBJECTIVE To develop and
Low cost and non-toxic genomic DNA extraction for use in molecular marker studies. Version 1.4, February 28 th, 2013. Prepared by Bernhard Hofinger, Owen Huynh and Brad Till. 1. OBJECTIVE To develop and
Quant One Step RT-PCR Kit
 1. Quant One Step RT-PCR Kit For fast and sensitive one-step RT-PCR www.tiangen.com/en RT121221 Quant One Step RT-PCR Kit Kit Contents Cat. no. KR113 Contents Hotmaster Taq Polymerase (2.5 U/μl) Quant
1. Quant One Step RT-PCR Kit For fast and sensitive one-step RT-PCR www.tiangen.com/en RT121221 Quant One Step RT-PCR Kit Kit Contents Cat. no. KR113 Contents Hotmaster Taq Polymerase (2.5 U/μl) Quant
Precise Characterization of Intact Monoclonal Antibodies by the Agilent 6545XT AdvanceBio LC/Q-TOF
 Precise Characterization of Intact Monoclonal Antibodies by the Agilent 6545XT AdvanceBio LC/Q-TOF Application Note Author David L. Wong Agilent Technologies, Inc. Santa Clara, CA, USA Introduction Monoclonal
Precise Characterization of Intact Monoclonal Antibodies by the Agilent 6545XT AdvanceBio LC/Q-TOF Application Note Author David L. Wong Agilent Technologies, Inc. Santa Clara, CA, USA Introduction Monoclonal
Analysis of Monoclonal Antibody (mab) Using Agilent 1290 Infinity LC System Coupled to Agilent 6530 Accurate-Mass Quadrupole Time-of-Flight (Q-TOF)
 Analysis of Monoclonal Antibody (mab) Using Agilent 9 Infinity LC System Coupled to Agilent Accurate-Mass Quadrupole Time-of-Flight (Q-TOF) Application Note Authors Ravindra Gudihal and Suresh Babu CV
Analysis of Monoclonal Antibody (mab) Using Agilent 9 Infinity LC System Coupled to Agilent Accurate-Mass Quadrupole Time-of-Flight (Q-TOF) Application Note Authors Ravindra Gudihal and Suresh Babu CV
Monitoring of Mammalian Cell Culture Media with HILIC LC/MS
 Application Note Metabolomics Monitoring of Mammalian Cell Culture Media with HILIC LC/MS Authors Jordy J. Hsiao, Genevieve Van de Bittner, Te Wei Chu, Oscar G. Potter, and Hongfeng Yin Agilent Technologies,
Application Note Metabolomics Monitoring of Mammalian Cell Culture Media with HILIC LC/MS Authors Jordy J. Hsiao, Genevieve Van de Bittner, Te Wei Chu, Oscar G. Potter, and Hongfeng Yin Agilent Technologies,
Separation of Native Monoclonal Antibodies and Identification of Charge Variants:
 Separation of Native Monoclonal Antibodies and Identification of Charge Variants: Teamwork of the Agilent 31 OFFGEL Fractionator, Agilent 21 Bioanalyzer and Agilent LC/MS Systems Application Note Biosimilar
Separation of Native Monoclonal Antibodies and Identification of Charge Variants: Teamwork of the Agilent 31 OFFGEL Fractionator, Agilent 21 Bioanalyzer and Agilent LC/MS Systems Application Note Biosimilar
Stool Nucleic Acid Isolation Kit Product 45600
 3430 Schmon Parkway Thorold, ON, Canada L2V 4Y6 Phone: 866-667-4362 (905) 227-8848 Fax: (905) 227-1061 Email: techsupport@norgenbiotek.com Stool Nucleic Acid Isolation Kit Product 45600 Product Insert
3430 Schmon Parkway Thorold, ON, Canada L2V 4Y6 Phone: 866-667-4362 (905) 227-8848 Fax: (905) 227-1061 Email: techsupport@norgenbiotek.com Stool Nucleic Acid Isolation Kit Product 45600 Product Insert
MMLV Reverse Transcriptase 1st-Strand cdna Synthesis Kit
 MMLV Reverse Transcriptase 1st-Strand cdna Synthesis Kit Cat. No. MM070150 Available exclusively thru Lucigen. lucigen.com/epibio www.lucigen.com MA265E MMLV Reverse Transcriptase 1st-Strand cdna Synthesis
MMLV Reverse Transcriptase 1st-Strand cdna Synthesis Kit Cat. No. MM070150 Available exclusively thru Lucigen. lucigen.com/epibio www.lucigen.com MA265E MMLV Reverse Transcriptase 1st-Strand cdna Synthesis
Guide-it sgrna In Vitro Transcription and Screening Systems User Manual
 Clontech Laboratories, Inc. Guide-it sgrna In Vitro Transcription and Screening Systems User Manual Cat. Nos. 631438, 631439 & 631440 (042114) Clontech Laboratories, Inc. A Takara Bio Company 1290 Terra
Clontech Laboratories, Inc. Guide-it sgrna In Vitro Transcription and Screening Systems User Manual Cat. Nos. 631438, 631439 & 631440 (042114) Clontech Laboratories, Inc. A Takara Bio Company 1290 Terra
High-Purity Bacterial RNA Isolated with the Agilent Total RNA Isolation Mini Kit Application
 High-Purity Bacterial RNA Isolated with the Total RNA Isolation Mini Kit Application Gene Expression Author Christopher Rizzo Claudia A. Robbins Rhonda R. Taylor Technologies, Inc. 285 Centerville Road
High-Purity Bacterial RNA Isolated with the Total RNA Isolation Mini Kit Application Gene Expression Author Christopher Rizzo Claudia A. Robbins Rhonda R. Taylor Technologies, Inc. 285 Centerville Road
SeCore. GSSP Kits. Instructions for Use. 1 Research Use Only
 SeCore GSSP Kits Instructions for Use 1 SeCore GSSP Kits Instructions for Use Invitrogen SeCore GSSP (Group Specific Sequencing Products) are designed to resolve ambiguous HLA heterozygous combinations,
SeCore GSSP Kits Instructions for Use 1 SeCore GSSP Kits Instructions for Use Invitrogen SeCore GSSP (Group Specific Sequencing Products) are designed to resolve ambiguous HLA heterozygous combinations,
EPIGENTEK. EpiQuik Tissue Chromatin Immunoprecipitation Kit. Base Catalog # P-2003 PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE
 EpiQuik Tissue Chromatin Immunoprecipitation Kit Base Catalog # P-2003 PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE The EpiQuik Tissue Chromatin Immunoprecipitation Kit is suitable for combining the specificity
EpiQuik Tissue Chromatin Immunoprecipitation Kit Base Catalog # P-2003 PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE The EpiQuik Tissue Chromatin Immunoprecipitation Kit is suitable for combining the specificity
Amplicon Library Preparation Method Manual. GS FLX Titanium Series October 2009
 GS FLX Titanium Series 1. Workflow 3. Procedure The procedure to prepare Amplicon libraries is shown in Figure 1. It consists of a PCR amplification, performed using special Fusion Primers for the Genome
GS FLX Titanium Series 1. Workflow 3. Procedure The procedure to prepare Amplicon libraries is shown in Figure 1. It consists of a PCR amplification, performed using special Fusion Primers for the Genome
5 min Soil DNA/RNA Preservation and Extraction Kit
 5 min Soil DNA/RNA Preservation and Extraction Kit Biofactories 5 min Soil DNA/RNA Preservation and Extraction Kit provides the fastest method for the storage/preservation and isolation/purification of
5 min Soil DNA/RNA Preservation and Extraction Kit Biofactories 5 min Soil DNA/RNA Preservation and Extraction Kit provides the fastest method for the storage/preservation and isolation/purification of
Comparison of Different Brands of Carrier Ampholytes for Monoclonal Antibody Charge Heterogeneity Analysis by Capillary Isoelectric Focusing
 Comparison of Different Brands of Carrier Ampholytes for Monoclonal Antibody Charge Heterogeneity Analysis by Capillary Isoelectric Focusing Application Note Biotherapeutics & Biosimilars Author Christian
Comparison of Different Brands of Carrier Ampholytes for Monoclonal Antibody Charge Heterogeneity Analysis by Capillary Isoelectric Focusing Application Note Biotherapeutics & Biosimilars Author Christian
Monitoring for Microcystins in Raw Water Supply Reservoirs Using the Agilent 6410 Triple Quadrupole LC/MS
 Monitoring for Microcystins in Raw Water Supply Reservoirs Using the Agilent 641 Triple Quadrupole LC/MS Application Note Environmental Author Toni Hall Wessex Water Bath, UK Abstract A method for the
Monitoring for Microcystins in Raw Water Supply Reservoirs Using the Agilent 641 Triple Quadrupole LC/MS Application Note Environmental Author Toni Hall Wessex Water Bath, UK Abstract A method for the
MassHunter Profinder: Batch Processing Software for High Quality Feature Extraction of Mass Spectrometry Data
 MassHunter Profinder: Batch Processing Software for High Quality Feature Extraction of Mass Spectrometry Data Technical Overview Introduction LC/MS metabolomics workflows typically involve the following
MassHunter Profinder: Batch Processing Software for High Quality Feature Extraction of Mass Spectrometry Data Technical Overview Introduction LC/MS metabolomics workflows typically involve the following
Ribonucleic acid (RNA)
 Ribonucleic acid (RNA) Ribonucleic acid (RNA) is more often found in nature as a singlestrand folded onto itself. Cellular organisms use messenger RNA (mrna) to convey genetic information (using the nitrogenous
Ribonucleic acid (RNA) Ribonucleic acid (RNA) is more often found in nature as a singlestrand folded onto itself. Cellular organisms use messenger RNA (mrna) to convey genetic information (using the nitrogenous
HCV Genotype Primer Kit
 Instruction Manual for HCV Genotype Primer Kit HCV Genotype Determination Kit for Research Purpose Thoroughly read this instruction manual before use of this kit Background Study of nucleotide sequence
Instruction Manual for HCV Genotype Primer Kit HCV Genotype Determination Kit for Research Purpose Thoroughly read this instruction manual before use of this kit Background Study of nucleotide sequence
Using the Pulsed Flame Photometric Detector (PFPD) for Low-level Analysis of Organophosphorus Pesticides
 Application Note 25390206 Using the Pulsed Flame Photometric Detector (PFPD) for Low-level Analysis of Organophosphorus Pesticides Keywords Model 5380 Organophosphorus Pesticide PFPD PFPDView Phosphorus
Application Note 25390206 Using the Pulsed Flame Photometric Detector (PFPD) for Low-level Analysis of Organophosphorus Pesticides Keywords Model 5380 Organophosphorus Pesticide PFPD PFPDView Phosphorus
100 mg. 18 g 10 g. 5 g. 2.5 g 1.5 g 2 g 4 g
 3430 Schmon Parkway Thorold, ON, Canada L2V 4Y6 Phone: 866-667-4362 (905) 227-8848 Fax: (905) 227-1061 Email: techsupport@norgenbiotek.com Plant/Fungi DNA Isolation Kit Product # 26200, 26250 Product Insert
3430 Schmon Parkway Thorold, ON, Canada L2V 4Y6 Phone: 866-667-4362 (905) 227-8848 Fax: (905) 227-1061 Email: techsupport@norgenbiotek.com Plant/Fungi DNA Isolation Kit Product # 26200, 26250 Product Insert
Labeling Protocol for mytags Immortal Libraries
 5840 Interface Drive, Suite 101 Ann Arbor MI 48103 1 (734) 998 0751 techsupport@arborbiosci.com Labeling Protocol for mytags Immortal Libraries March 2018 Version 1.5 Contents Reagents and Equipment...
5840 Interface Drive, Suite 101 Ann Arbor MI 48103 1 (734) 998 0751 techsupport@arborbiosci.com Labeling Protocol for mytags Immortal Libraries March 2018 Version 1.5 Contents Reagents and Equipment...
5 min Soil RNA Preservation and Extraction Kit
 5 min Soil RNA Preservation and Extraction Kit Biofactories 5 min Soil RNA Preservation and Extraction Kit provides the fastest method for the storage/preservation and isolation/purification of total RNA
5 min Soil RNA Preservation and Extraction Kit Biofactories 5 min Soil RNA Preservation and Extraction Kit provides the fastest method for the storage/preservation and isolation/purification of total RNA
PlantDirect TM Multiplex PCR System
 PlantDirect TM Multiplex PCR System Technical Manual No. 0178 Version 10112010 I Description.. 1 II Applications 2 III Key Features.. 3 IV Shipping and Storage. 3 V Simplified Procedures. 3 VI Detailed
PlantDirect TM Multiplex PCR System Technical Manual No. 0178 Version 10112010 I Description.. 1 II Applications 2 III Key Features.. 3 IV Shipping and Storage. 3 V Simplified Procedures. 3 VI Detailed
Puzzled About LCMS? Sample. Sensitivity in Mass Spec Analysis. Prep. Adapting LC-UV. Optimizing and Maintaining Your Mass Spec LCMS
 Puzzled About LCMS? Sensitivity in Mass Spec Analysis Sample Prep Adapting LC-UV Optimizing and Maintaining Your Mass Spec To LCMS Paul Altiero Alex Ucci Applications Phone Support Chemistry and Supplies
Puzzled About LCMS? Sensitivity in Mass Spec Analysis Sample Prep Adapting LC-UV Optimizing and Maintaining Your Mass Spec To LCMS Paul Altiero Alex Ucci Applications Phone Support Chemistry and Supplies
qpcr Kit, DNA-free Product components 100 rxn 250 rxn Product description
 qpcr Kit, DNA-free For the PCR detection and identification of bacterial and fungal DNA using custom primers Product code A8514 Product components 100 rxn 250 rxn A 2.5x mastermix (3 mm MgCl 2 final concentration)
qpcr Kit, DNA-free For the PCR detection and identification of bacterial and fungal DNA using custom primers Product code A8514 Product components 100 rxn 250 rxn A 2.5x mastermix (3 mm MgCl 2 final concentration)
BiOstic Bacteremia DNA Isolation Kit Sample Catalog No S (For use with cultured blood for the enrichment of bacteria)
 BiOstic Bacteremia DNA Isolation Kit Sample Catalog No. 12240-S (For use with cultured blood for the enrichment of bacteria) Information for Ordering Product Catalog No. Quantity 12240-50 50 preps Instruction
BiOstic Bacteremia DNA Isolation Kit Sample Catalog No. 12240-S (For use with cultured blood for the enrichment of bacteria) Information for Ordering Product Catalog No. Quantity 12240-50 50 preps Instruction
Protein and transcriptome quantitation using BD AbSeq Antibody-Oligonucleotide
 Protein and transcriptome quantitation using BD AbSeq Antibody-Oligonucleotide technology and the 10X Genomics Chromium Single Cell Gene Expression Solution Jocelyn G. Olvera, Brigid S. Boland, John T.
Protein and transcriptome quantitation using BD AbSeq Antibody-Oligonucleotide technology and the 10X Genomics Chromium Single Cell Gene Expression Solution Jocelyn G. Olvera, Brigid S. Boland, John T.
Mass Spectrometry Analysis of Liquid Chromatography Fractions using Ettan LC MS System
 GUIDE TO LC MS - December 21 1 Spectrometry Analysis of Liquid Chromatography Fractions using Ettan LC MS System Henrik Wadensten, Inger Salomonsson, Staffan Lindqvist, Staffan Renlund, Amersham Biosciences,
GUIDE TO LC MS - December 21 1 Spectrometry Analysis of Liquid Chromatography Fractions using Ettan LC MS System Henrik Wadensten, Inger Salomonsson, Staffan Lindqvist, Staffan Renlund, Amersham Biosciences,
DNA Extraction DNA Extraction (small scale) using CTAB method
 DNA Extraction DNA Extraction (small scale) using CTAB method This method is relatively simple, and has been used successfully with a wide range of monocot and dicot species. The method may be used with
DNA Extraction DNA Extraction (small scale) using CTAB method This method is relatively simple, and has been used successfully with a wide range of monocot and dicot species. The method may be used with
The Five Key Elements of a Successful Metabolomics Study
 The Five Key Elements of a Successful Metabolomics Study Metabolomics: Completing the Biological Picture Metabolomics is offering new insights into systems biology, empowering biomarker discovery, and
The Five Key Elements of a Successful Metabolomics Study Metabolomics: Completing the Biological Picture Metabolomics is offering new insights into systems biology, empowering biomarker discovery, and
Functional Genomic Study of Exogenous n-butanol Stress in Escherichia coli. Supplementary Data
 Functional Genomic Study of Exogenous n-butanol Stress in Escherichia coli Supplementary Data Complete Microarray Data: http://www.microbesonline.org/cgi-bin/microarray/viewexp.cgi?expid=1319 or http://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=gse16973
Functional Genomic Study of Exogenous n-butanol Stress in Escherichia coli Supplementary Data Complete Microarray Data: http://www.microbesonline.org/cgi-bin/microarray/viewexp.cgi?expid=1319 or http://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=gse16973
CHROMATOGRAPHY/MASS SPECTROMETRY
 DETERMINATION OF 1,4-DIOXANE IN DRINKING WATER BY SOLID PHASE EXTRACTION (SPE) AND GAS CHROMATOGRAPHY/MASS SPECTROMETRY (GC/MS) WITH SELECTED ION MONITORING (SIM)* EPA Method 522 Part Number: EU52112M6
DETERMINATION OF 1,4-DIOXANE IN DRINKING WATER BY SOLID PHASE EXTRACTION (SPE) AND GAS CHROMATOGRAPHY/MASS SPECTROMETRY (GC/MS) WITH SELECTED ION MONITORING (SIM)* EPA Method 522 Part Number: EU52112M6
Presto Soil DNA Extraction Kit
 Instruction Manual Ver. 02.23.17 For Research Use Only Presto Soil DNA Extraction Kit Advantages SLD004 (4 Preparation Sample Kit) SLD050 (50 Preparation Kit) SLD100 (100 Preparation Kit) Sample: 250-500
Instruction Manual Ver. 02.23.17 For Research Use Only Presto Soil DNA Extraction Kit Advantages SLD004 (4 Preparation Sample Kit) SLD050 (50 Preparation Kit) SLD100 (100 Preparation Kit) Sample: 250-500
Integrating workflows and Bioinformatics for Exposome and Environmental Health Research ADVANCING EXPOSOMICS RESEARCH
 Integrating workflows and Bioinformatics for Exposome and Environmental Health Research ADVANCING EXPOSOMICS RESEARCH EXPOSOMICS AND ENVIRONMENTAL HEALTH RESEARCH Agilent empowers exposome and environmental
Integrating workflows and Bioinformatics for Exposome and Environmental Health Research ADVANCING EXPOSOMICS RESEARCH EXPOSOMICS AND ENVIRONMENTAL HEALTH RESEARCH Agilent empowers exposome and environmental
Purpose: To sequence 16S microbial DNA isolated from animal fecal matter and perform compositional analysis of bacterial communities.
 Purpose: To sequence 16S microbial DNA isolated from animal fecal matter and perform compositional analysis of bacterial communities. Reference: This protocol was adapted from: A custom and streamlined
Purpose: To sequence 16S microbial DNA isolated from animal fecal matter and perform compositional analysis of bacterial communities. Reference: This protocol was adapted from: A custom and streamlined
Monitoring for Microcystins in Raw Water Supply Reservoirs Using the Agilent 6410 Triple Quadrupole LC/MS
 Monitoring for Microcystins in Raw Water Supply Reservoirs Using the Agilent 641 Triple Quadrupole LC/MS Application Note Environmental Author Toni Hall Wessex Water Bath, UK Abstract A method for the
Monitoring for Microcystins in Raw Water Supply Reservoirs Using the Agilent 641 Triple Quadrupole LC/MS Application Note Environmental Author Toni Hall Wessex Water Bath, UK Abstract A method for the
Functional Genomics Research Stream. Research Meeting: November 8, 2011 cdna Library Construction for RNA-Seq
 Functional Genomics Research Stream Research Meeting: November 8, 2011 cdna Library Construction for RNA-Seq Lab Issues Don t leave out boxes of water + ethidium bromide Minimize tip boxes in phenol waste
Functional Genomics Research Stream Research Meeting: November 8, 2011 cdna Library Construction for RNA-Seq Lab Issues Don t leave out boxes of water + ethidium bromide Minimize tip boxes in phenol waste
BRCA MAQ USER GUIDE Version 1.0
 BRCA MAQ USER GUIDE Version 1.0 For Copy Number Analysis Assays IFU604 v170797 www.multiplicom.com 2012 Multiplicom NV, all rights reserved Table of Contents Introduction to Multiplex Amplicon Quantification...2
BRCA MAQ USER GUIDE Version 1.0 For Copy Number Analysis Assays IFU604 v170797 www.multiplicom.com 2012 Multiplicom NV, all rights reserved Table of Contents Introduction to Multiplex Amplicon Quantification...2
Introduction. Benefits of the SWATH Acquisition Workflow for Metabolomics Applications
 SWATH Acquisition Improves Metabolite Coverage over Traditional Data Dependent Techniques for Untargeted Metabolomics A Data Independent Acquisition Technique Employed on the TripleTOF 6600 System Zuzana
SWATH Acquisition Improves Metabolite Coverage over Traditional Data Dependent Techniques for Untargeted Metabolomics A Data Independent Acquisition Technique Employed on the TripleTOF 6600 System Zuzana
T7-Based RNA Amplification Protocol (in progress)
 T7-Based RNA Amplification Protocol (in progress) Jacqueline Ann Lopez (modifications) Amy Cash & Justen Andrews INTRODUCTION T7 RNA Amplification, a technique originally developed in the laboratory of
T7-Based RNA Amplification Protocol (in progress) Jacqueline Ann Lopez (modifications) Amy Cash & Justen Andrews INTRODUCTION T7 RNA Amplification, a technique originally developed in the laboratory of
A Direct Column-Performance Comparison for Rapid Contract Laboratory Program (CLP) Pesticide Analysis. Application. Authors. Abstract.
 A Direct Column-Performance Comparison for Rapid Contract Laboratory Program (CLP) Pesticide Analysis Application Environmental Authors Ken Lynam and John W. Henderson, Jr. Agilent Technologies, Inc. 2850
A Direct Column-Performance Comparison for Rapid Contract Laboratory Program (CLP) Pesticide Analysis Application Environmental Authors Ken Lynam and John W. Henderson, Jr. Agilent Technologies, Inc. 2850
Molecular Methods in Microbial Ecology
 Molecular Methods in Microbial Ecology Kristin Gribble 508-289-7194 kgribble@mbl.edu Lillie 305 Tuesday 10/23/18 Introduction, Extraction of DNA from Winogradsky columns Run DNA products on gel Thursday
Molecular Methods in Microbial Ecology Kristin Gribble 508-289-7194 kgribble@mbl.edu Lillie 305 Tuesday 10/23/18 Introduction, Extraction of DNA from Winogradsky columns Run DNA products on gel Thursday
5 min Insect DNA/RNA Preservation and Extraction Kit
 5 min Insect DNA/RNA Preservation and Extraction Kit Biofactories 5 min Insect DNA/RNA Preservation and Extraction Kit provides the fastest method for the storage/preservation and isolation/purification
5 min Insect DNA/RNA Preservation and Extraction Kit Biofactories 5 min Insect DNA/RNA Preservation and Extraction Kit provides the fastest method for the storage/preservation and isolation/purification
N-Glycan Profiling Analysis of a Monoclonal Antibody Using UHPLC/FLD/Q-TOF
 N-Glycan Profiling Analysis of a Monoclonal Antibody Using UHPLC/FLD/Q-TOF Application Note Authors Xianming Liu, Wei Zhang, Yi Du, Sheng Yin, Hong Que, and Weichang Zhou WuXi AppTec iopharmaceuticals
N-Glycan Profiling Analysis of a Monoclonal Antibody Using UHPLC/FLD/Q-TOF Application Note Authors Xianming Liu, Wei Zhang, Yi Du, Sheng Yin, Hong Que, and Weichang Zhou WuXi AppTec iopharmaceuticals
Xevo G2-S QTof and TransOmics: A Multi-Omics System for the Differential LC/MS Analysis of Proteins, Metabolites, and Lipids
 Xevo G2-S QTof and TransOmics: A Multi-Omics System for the Differential LC/MS Analysis of Proteins, Metabolites, and Lipids Ian Edwards, Jayne Kirk, and Joanne Williams Waters Corporation, Manchester,
Xevo G2-S QTof and TransOmics: A Multi-Omics System for the Differential LC/MS Analysis of Proteins, Metabolites, and Lipids Ian Edwards, Jayne Kirk, and Joanne Williams Waters Corporation, Manchester,
Polyskope 1.0 Multiplex Pathogen Detection Assay
 Polyskope 1.0 Multiplex Pathogen Detection Assay User Guide Test for the real-time simultaneous PCR detection of E.coli O157 STEC, Salmonella spp. and Listeria monocytogenes in food and environmental samples
Polyskope 1.0 Multiplex Pathogen Detection Assay User Guide Test for the real-time simultaneous PCR detection of E.coli O157 STEC, Salmonella spp. and Listeria monocytogenes in food and environmental samples
Agilent s NEW MassHunter Profinder
 Agilent s NEW MassHunter Profinder The Most Advanced Batch Feature Extraction Software for Metabolomics Theodore Sana, Ph.D. Metabolomics Marketing Manager 1 MassHunter Profinder A Batch Feature Extraction
Agilent s NEW MassHunter Profinder The Most Advanced Batch Feature Extraction Software for Metabolomics Theodore Sana, Ph.D. Metabolomics Marketing Manager 1 MassHunter Profinder A Batch Feature Extraction
LunaScript RT SuperMix Kit
 POLYMERASES & AMPLIFICATION LunaScript RT SuperMix Kit Instruction Manual NEB #E3010S/L 25/100 reactions Version 1.0 1/18 be INSPIRED drive DISCOVERY stay GENUINE This product is intended for research
POLYMERASES & AMPLIFICATION LunaScript RT SuperMix Kit Instruction Manual NEB #E3010S/L 25/100 reactions Version 1.0 1/18 be INSPIRED drive DISCOVERY stay GENUINE This product is intended for research
Agilent Anion-Exchange Media for Proteins
 Agilent Anion-Exchange Media for Proteins Technical Overview Introduction Agilent PL-SAX combines rigid macroporous polystyrene/divinylbenzene (PS/DVB) polymer matrix and chemically stable quaternized
Agilent Anion-Exchange Media for Proteins Technical Overview Introduction Agilent PL-SAX combines rigid macroporous polystyrene/divinylbenzene (PS/DVB) polymer matrix and chemically stable quaternized
TIANamp Marine Animals DNA Kit
 TIANamp Marine Animals DNA Kit For isolation of genomic DNA from marine animal tissues www.tiangen.com/en DP121221 TIANamp Marine Animals DNA Kit (Spin Column) Cat. no. DP324 Kit Contents Contents DP324-02
TIANamp Marine Animals DNA Kit For isolation of genomic DNA from marine animal tissues www.tiangen.com/en DP121221 TIANamp Marine Animals DNA Kit (Spin Column) Cat. no. DP324 Kit Contents Contents DP324-02
SOP for Quantitation of the Hepatitis C Virus (HCV) Genome by RT-PCR
 Virus Bank SOP-HCV-001 1. Scope SOP for Quantitation of the Hepatitis C Virus (HCV) Genome by RT-PCR 1.1 This procedure describes a method for quantitation of the HCV genome in HCV-infected cells by RT-PCR.
Virus Bank SOP-HCV-001 1. Scope SOP for Quantitation of the Hepatitis C Virus (HCV) Genome by RT-PCR 1.1 This procedure describes a method for quantitation of the HCV genome in HCV-infected cells by RT-PCR.
NEBNext Magnesium RNA Fragmentation Module
 SAMPLE PREPARATION NEBNext Magnesium RNA Fragmentation Module Instruction Manual NEB #E6150S 200 reactions NEBNext Magnesium RNA Fragmentation Module Table of Contents: Description....2 Applications....2
SAMPLE PREPARATION NEBNext Magnesium RNA Fragmentation Module Instruction Manual NEB #E6150S 200 reactions NEBNext Magnesium RNA Fragmentation Module Table of Contents: Description....2 Applications....2
Average Yields* Yeast DNA Yeast RNA Time to Complete 10 Purifications * Yield will vary depending on the type of sample processed
 3430 Schmon Parkway Thorold, ON, Canada L2V 4Y6 Phone: 866-667-4362 (905) 227-8848 Fax: (905) 227-1061 Email: techsupport@norgenbiotek.com Fungi/Yeast RNA/DNA Purification Kit Product # 35800 Product Insert
3430 Schmon Parkway Thorold, ON, Canada L2V 4Y6 Phone: 866-667-4362 (905) 227-8848 Fax: (905) 227-1061 Email: techsupport@norgenbiotek.com Fungi/Yeast RNA/DNA Purification Kit Product # 35800 Product Insert
Jet Stream Proteomics for Sensitive and Robust Standard Flow LC/MS
 Jet Stream Proteomics for Sensitive and Robust Standard Flow LC/MS Technical Overview Authors Yanan Yang, Vadiraj hat, and Christine Miller Agilent Technologies, Inc. Santa Clara, California Introduction
Jet Stream Proteomics for Sensitive and Robust Standard Flow LC/MS Technical Overview Authors Yanan Yang, Vadiraj hat, and Christine Miller Agilent Technologies, Inc. Santa Clara, California Introduction
Telomerase Activity Quantification qpcr Assay Kit (TAQ) Catalog # reactions
 Telomerase Activity Quantification qpcr Assay Kit (TAQ) Catalog #8928 100 reactions Product Description Telomeres are repetitive nucleotide elements at the ends of chromosomes that protect chromosomes
Telomerase Activity Quantification qpcr Assay Kit (TAQ) Catalog #8928 100 reactions Product Description Telomeres are repetitive nucleotide elements at the ends of chromosomes that protect chromosomes
iplex HS Melanoma Panel
 iplex HS Melanoma Panel This quick reference guide gives the assay protocol for the iplex HS Melanoma Panel. For complete information on equipment and reagents required, laboratory work areas and guidelines,
iplex HS Melanoma Panel This quick reference guide gives the assay protocol for the iplex HS Melanoma Panel. For complete information on equipment and reagents required, laboratory work areas and guidelines,
